-
 Neurotransmitter Systems in Alzheimer’s Disease
Neurotransmitter Systems in Alzheimer’s Disease -
 From Molecules to Meaning: Integrating Neuropeptides, Sociostasis, and Hormesis in the Brain–Heart Axis
From Molecules to Meaning: Integrating Neuropeptides, Sociostasis, and Hormesis in the Brain–Heart Axis -
 Transcriptional Divergence of Conserved Starch Metabolism Genes During Grain Filling in Indica and Japonica Rice
Transcriptional Divergence of Conserved Starch Metabolism Genes During Grain Filling in Indica and Japonica Rice -
 Different Effects of Antioxidants Against Ionizing Radiation: An Experimental Model of Micronuclei
Different Effects of Antioxidants Against Ionizing Radiation: An Experimental Model of Micronuclei -
 Anti-Biofilm Activity of Combinations of Cinnamic Acid and Its Derivatives with Cloxacillin Against Methicillin-Resistant Staphylococcus epidermidis
Anti-Biofilm Activity of Combinations of Cinnamic Acid and Its Derivatives with Cloxacillin Against Methicillin-Resistant Staphylococcus epidermidis
Journal Description
Current Issues in Molecular Biology
Current Issues in Molecular Biology
is an international, scientific, peer-reviewed, open access journal on molecular biology, published monthly online by MDPI (from Volume 43, Issue 1 - 2021).
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PMC, PubMed, Embase, CAPlus / SciFinder, FSTA, AGRIS, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.3 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names are published annually in the journal.
Impact Factor:
3.0 (2024);
5-Year Impact Factor:
3.2 (2024)
Latest Articles
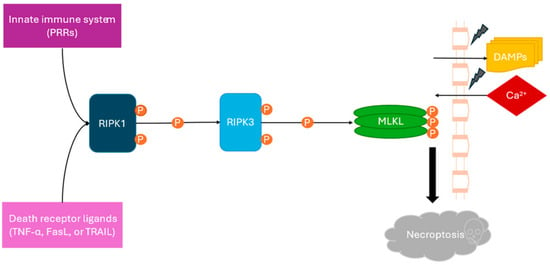
Necroptosis in SJS/TEN: RIPK1 and RIPK3 Expression and Implications for Disease Pathogenesis
Curr. Issues Mol. Biol. 2026, 48(5), 540; https://doi.org/10.3390/cimb48050540 - 21 May 2026
Abstract
Necroptosis has been implicated in the pathogenesis of Stevens–Johnson syndrome and toxic epidermal necrolysis (SJS/TEN), with prior studies demonstrating tissue-level involvement of receptor-interacting protein kinases RIPK1 and RIPK3. However, their systemic expression in the circulatory compartment remains incompletely characterized. The objective of this
[...] Read more.
Necroptosis has been implicated in the pathogenesis of Stevens–Johnson syndrome and toxic epidermal necrolysis (SJS/TEN), with prior studies demonstrating tissue-level involvement of receptor-interacting protein kinases RIPK1 and RIPK3. However, their systemic expression in the circulatory compartment remains incompletely characterized. The objective of this study is to evaluate circulating levels of RIPK1 and RIPK3 in patients with SJS/TEN and explore their potential association with diseases. Serum samples from patients with SJS/TEN and control groups were analyzed for RIPK1 and RIPK3 levels using ELISA. Group differences were assessed using non-parametric statistical methods. Circulating levels of RIPK1 and RIPK3 were elevated in patients with SJS/TEN compared with controls. These findings were consistent across analyses; however, variability within groups and overlap between cohorts were observed. These results suggest an association between increased circulating RIPK1 and RIPK3 levels and SJS/TEN. Given the limited sample size, heterogeneous control populations, and lack of functional or phosphorylation-specific assays, these findings should be considered exploratory. Further studies incorporating larger cohorts and mechanistic validation are needed to clarify the role of necroptosis-related pathways in the systemic manifestations of SJS/TEN.
Full article
(This article belongs to the Special Issue Pathogenesis, Diagnosis and Treatment of Skin and Sexually Transmitted Diseases)
►
Show Figures
Open AccessEditorial
Editorial for the Special Issue “The Role of Bioactives in Inflammation, 2nd Edition”
by
Chan-Yen Kuo
Curr. Issues Mol. Biol. 2026, 48(5), 539; https://doi.org/10.3390/cimb48050539 - 21 May 2026
Abstract
Inflammation is increasingly recognized as a dynamic and interconnected regulatory network rather than a linear signaling cascade, in which immune signaling, oxidative stress, metabolic adaptation, and tissue-specific microenvironmental factors collectively shape inflammatory responses [...]
Full article
(This article belongs to the Special Issue The Role of Bioactives in Inflammation, 2nd Edition)
Open AccessArticle
Hesperetin-7-O-Glucuronide Improves Endothelial Cell Function Through Improving NO/ET-1 Balance and Reducing Oxidative Stress via miRNAs
by
Lu Li, Kexin Ji, Fengqi Du, Nini Jin, He Li and Xinqi Liu
Curr. Issues Mol. Biol. 2026, 48(5), 538; https://doi.org/10.3390/cimb48050538 - 21 May 2026
Abstract
Citrus flavonoid intake is associated with beneficial effects on endothelial function. Our previous randomized control trial demonstrated that the concentration of Hesperetin-7-O-glucuronide (H7G) was positively correlated with the improvement in endothelial function in overweight and obese participants following blood orange juice consumption. To
[...] Read more.
Citrus flavonoid intake is associated with beneficial effects on endothelial function. Our previous randomized control trial demonstrated that the concentration of Hesperetin-7-O-glucuronide (H7G) was positively correlated with the improvement in endothelial function in overweight and obese participants following blood orange juice consumption. To explore the underlying mechanism by which H7G improves endothelial function, we investigated the regulation of H7G on endothelial function in a permanent human endothelial cell line (EA. hy926 cells) under normal and oxidative conditions treated with high-oxidation low-density lipoprotein. The results indicated that H7G improved the expression of nitric oxide synthase 3 (NOS3), heme oxygenase 1 (HMOX1) ad glutamate cysteine ligase catalytic (GCLC), and inhibited the expression of endothelin-1 (EDN1), through the upregulation of miR-660-5p and inhibition of miR-21-5p. In summary, H7G improves endothelial cell function via the upregulation of miR-660-5p and the inhibition of miR-21-5p.
Full article
(This article belongs to the Special Issue Biologically Active Compounds: Sources, Mechanisms of Action, and Applications)
►▼
Show Figures
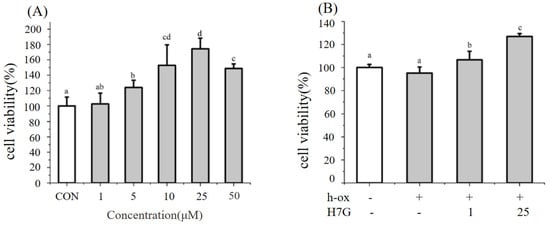
Figure 1
Open AccessEditorial
Editorial for Special Issue “Osteoclastogenesis and Osteogenesis: Physiological and Molecular Responses to Xenobiotics and Biomaterials”
by
Maria Giovanna Rizzo
Curr. Issues Mol. Biol. 2026, 48(5), 537; https://doi.org/10.3390/cimb48050537 - 21 May 2026
Abstract
We are pleased to present this Special Issue of Current Issues in Molecular Biology, entitled “Osteoclastogenesis and Osteogenesis: Physiological and Molecular Responses to Xenobiotics and Biomaterials” [...]
Full article
(This article belongs to the Special Issue Osteoclastogenesis and Osteogenesis: Physiological and Molecular Responses to Xenobiotics and Biomaterials)
Open AccessArticle
An Iron–Complement Network Model of Thromboinflammation and Humoral Immune Remodeling in Severe COVID-19
by
Zhen Chen, Shanshan Wang and Yuzong Chen
Curr. Issues Mol. Biol. 2026, 48(5), 536; https://doi.org/10.3390/cimb48050536 - 21 May 2026
Abstract
Severe COVID-19 is characterized by profound thromboinflammatory and immune disturbances, but the network-level relationships among complement–coagulation dysregulation, humoral immune remodeling, and iron-associated immune regulation remain incompletely understood. Here, we performed integrative proteomic and transcriptomic analyses across peripheral blood and lung microenvironments using weighted
[...] Read more.
Severe COVID-19 is characterized by profound thromboinflammatory and immune disturbances, but the network-level relationships among complement–coagulation dysregulation, humoral immune remodeling, and iron-associated immune regulation remain incompletely understood. Here, we performed integrative proteomic and transcriptomic analyses across peripheral blood and lung microenvironments using weighted gene co-expression network analysis (WGCNA), differential network analysis (DiNA), and immune deconvolution. Proteomic network analysis identified a disease-associated module enriched in complement activation, coagulation cascades, platelet degranulation, and acute inflammatory responses. Hub proteins, including C9, LBP, vWF, and F11, were prioritized based on module association and intramodular connectivity. Notably, C9 and LBP were repeatedly identified across WGCNA, DiNA, and differential expression analyses, underscoring their robust association with severe COVID-19-associated molecular network remodeling. Transcriptomic and CIBERSORTx-based immune deconvolution analyses showed altered immune-cell composition in blood and lung tissues, including B-cell and plasma-cell-associated changes. Notably, TFRC displayed cell-type-associated expression changes in naïve B cells and plasma cells, suggesting a potential link between iron-associated immune regulation and humoral immune remodeling. Collectively, these computational findings highlight coordinated complement–coagulation dysregulation, humoral immune remodeling, and TFRC-associated iron-related immune alterations in severe COVID-19, and prioritize TFRC, C9, and LBP as candidate molecular indicators requiring further experimental and clinical validation.
Full article
(This article belongs to the Special Issue Molecular and Real-World Evidence Research of Respiratory Diseases and Infections, 2nd Edition)
►▼
Show Figures
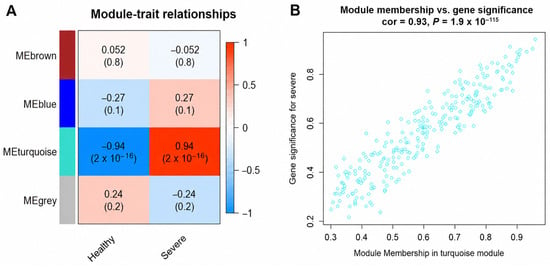
Figure 1
Open AccessReview
Triple-Negative Breast Cancer: Molecular Subtypes; Immune Escape; Limitations of Current Immunotherapy; and the BTLA/HVEM/CD160 Axis as an Emerging Target
by
Bernardo L. Rapoport, Ronald Anderson, Daniel van Tonder, Teresa Smit, Theresa M. Rossouw, Carol-Ann Benn and Helen C. Steel
Curr. Issues Mol. Biol. 2026, 48(5), 535; https://doi.org/10.3390/cimb48050535 - 20 May 2026
Abstract
Triple-negative breast cancer is an aggressive and heterogeneous type of invasive breast cancer (BC) in which the cancer cells lack estrogen and progesterone receptors, as well as expression of the human epidermal growth factor 2 protein. This cancer tends to grow and spread
[...] Read more.
Triple-negative breast cancer is an aggressive and heterogeneous type of invasive breast cancer (BC) in which the cancer cells lack estrogen and progesterone receptors, as well as expression of the human epidermal growth factor 2 protein. This cancer tends to grow and spread faster than other BC subtypes, and is associated with a poor prognosis due to early visceral and neurological recurrences. Multidisciplinary management includes surgery, chemotherapy, radiation therapy, and immunotherapy with targeted immune checkpoint inhibitors (ICIs). The introduction of ICIs has improved outcomes in patients with TNBC, particularly in the metastatic and neoadjuvant settings. Despite these advances, a significant proportion of patients either do not respond to treatment or develop resistance to it. TNBC mortality remains high, underscoring the urgent need to identify novel prognostic and predictive biomarkers to overcome resistance to immunotherapy. Following a brief overview of the clinical features and established biomarkers of TNBC, the current review focuses on immune checkpoint proteins (ICPs) beyond PD-1 and PD-L1, and on the potential use of soluble ICPs rather than the well-established membrane-bound assays. These soluble ICPs are produced through the alternative splicing of messenger (m)RNA or the cleavage/shedding of membrane-bound proteins. This is followed by an overview of current treatment and novel predictive targets in TNBC. Additionally, the involvement of the B- and T-lymphocyte attenuator (BTLA)/herpes virus entry mediator (HVEM)/CD160 pathway and its role in the pathogenesis of BC and TNBC are reviewed, highlighting the potential use of BTLA and HVEM as biomarkers.
Full article
(This article belongs to the Section Biochemistry, Molecular and Cellular Biology)
►▼
Show Figures

Figure 1
Open AccessReview
Alpha-Fetoprotein as a Biomarker in Pregnancy: From Genetic Disorders to Obstetric Complications
by
Shaqraa Musawi
Curr. Issues Mol. Biol. 2026, 48(5), 534; https://doi.org/10.3390/cimb48050534 - 20 May 2026
Abstract
Alpha-fetoprotein (AFP) is a glycoprotein primarily produced by the fetal liver and yolk sac during development. It is a multifaceted biomarker with significant applications in the prenatal screening of congenital abnormalities, cancer, and other disorders. The level of AFP in maternal blood may
[...] Read more.
Alpha-fetoprotein (AFP) is a glycoprotein primarily produced by the fetal liver and yolk sac during development. It is a multifaceted biomarker with significant applications in the prenatal screening of congenital abnormalities, cancer, and other disorders. The level of AFP in maternal blood may indicate several obstetric concerns and complications during pregnancy. Atypical AFP levels are commonly utilized as a biomarker for detecting fetal anomalies, placental complications, and other pregnancy-related issues. These findings raise concerns regarding the effectiveness of screening maternal serum alpha-fetoprotein (MS-AFP) as a primary indicator of pregnancy problems and underscore the need for further investigation into the functional role of AFP throughout pregnancy. The measurement of MS-AFP has been utilized for the past four decades. It is anticipated that MS-AFP measurement will continue to be utilized as a component of integrated or sequential tests for chromosomal abnormalities and may serve as a prognostic indicator for adverse obstetric outcomes. Critically, whether AFP functions solely as a passive marker or plays active biological roles in pregnancy physiology and pathology remains unresolved, necessitating additional mechanistic investigation and discourse. This review consolidates critical data from numerous studies on AFP, focusing specifically on its diagnostic and prognostic applications for congenital abnormalities and problems during pregnancy. This review also identifies key research gaps regarding the functional biology of AFP, particularly whether AFP functions as a passive biomarker or an active participant in the pathophysiology of adverse pregnancy outcomes.
Full article
(This article belongs to the Special Issue Targeted Therapies and Biomarker Discovery in Health and Disease)
►▼
Show Figures

Figure 1
Open AccessEditorial
Editorial for Special Issue “Molecular Research in Vaccinology and Vaccine Development”
by
Attila Farsang
Curr. Issues Mol. Biol. 2026, 48(5), 533; https://doi.org/10.3390/cimb48050533 - 20 May 2026
Abstract
Infectious diseases have always posed a significant threat to both humankind and our livestock [...]
Full article
(This article belongs to the Special Issue Molecular Research in Vaccinology and Vaccine Development)
Open AccessReview
AI and Machine Learning for Proteomics-Driven Drug Discovery: Methods, Tools, and Best Practices
by
Suman Basak
Curr. Issues Mol. Biol. 2026, 48(5), 532; https://doi.org/10.3390/cimb48050532 - 20 May 2026
Abstract
Proteomics has become central to pharmacological research by providing quantitative readouts of protein abundance, post-translational modifications, interactions, and spatial context. However, proteomic datasets are high-dimensional, heterogeneous, and frequently affected by missingness, batch effects, and limited cohort size. Artificial intelligence (AI) and machine learning
[...] Read more.
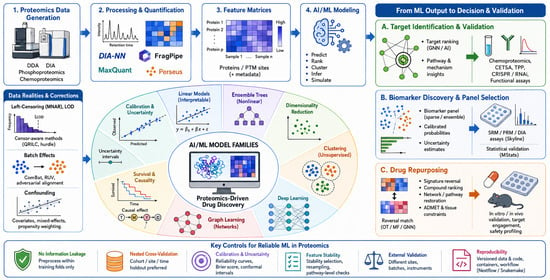
Proteomics has become central to pharmacological research by providing quantitative readouts of protein abundance, post-translational modifications, interactions, and spatial context. However, proteomic datasets are high-dimensional, heterogeneous, and frequently affected by missingness, batch effects, and limited cohort size. Artificial intelligence (AI) and machine learning (ML) can help convert these complex data into decision-relevant outputs for target identification, biomarker discovery, pharmacodynamic monitoring, and drug repurposing. This review critically compares supervised learning, ensemble methods, dimensionality reduction, clustering, deep learning, graph learning, survival modeling, causal inference, and calibration approaches in proteomics-driven drug discovery. We also summarize major software ecosystems for mass-spectrometry processing, targeted assays, spectrum prediction, phosphoproteomics, structure modeling, and reproducible workflows. Emphasis is placed on model selection, benchmarking, missing-data handling, batch correction, interpretability, uncertainty, experimental validation, and translational readiness. Finally, we highlight emerging directions, including contrastive learning, diffusion models, graph-based integration, and federated analytics.
Full article
(This article belongs to the Special Issue Multi-Omics Integration for Precision Medicine: From Pathogenesis, Diagnosis to Treatment)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Screening the Combination of Gemcitabine, Clomipramine, and Resveratrol in HL-60 Leukemia Cells
by
Burcu Biltekin, Yusuf Elgormus and Ayhan Bilir
Curr. Issues Mol. Biol. 2026, 48(5), 531; https://doi.org/10.3390/cimb48050531 - 19 May 2026
Abstract
Background and Objectives: Potential anti-neoplastic effects of resveratrol, which has antioxidant features combined with clomipramine, which has antineoplastic features, or with gemcitabine, used as a nucleoside analog widely used in chemotherapy, were evaluated together and individually on the HL-60 leukemia cells in
[...] Read more.
Background and Objectives: Potential anti-neoplastic effects of resveratrol, which has antioxidant features combined with clomipramine, which has antineoplastic features, or with gemcitabine, used as a nucleoside analog widely used in chemotherapy, were evaluated together and individually on the HL-60 leukemia cells in this in vitro screening study. Materials and Methods: HL-60 cells were treated with gemcitabine, clomipramine, resveratrol, or their combinations at concentrations ranging from 1 to 200 µM. Cell viability was assessed at 24, 48, and 72 h using the trypan blue exclusion method, and results are expressed as a percentage of time-matched untreated controls. Cell proliferation was further evaluated by bromodeoxyuridine (BrdU) immunohistochemical labeling. All experiments were performed in triplicate, and statistical analyses were conducted using one-way analysis of variance (ANOVA) with post hoc comparisons. Results: Gemcitabine markedly reduced HL-60 cell viability at all concentrations and time points (p < 0.001), indicating strong time-dependent cytotoxicity, with a significant drop in BrdU proliferation index at 48 h (p < 0.001). Clomipramine exhibited a biphasic response: high concentrations decreased viability (p < 0.05), while low concentrations allowed partial recovery by 72 h. Resveratrol showed concentration-dependent cytotoxicity, with reduced viability at high concentration and near-control levels at low concentration by 72 h; BrdU indices remained significantly lower than control (p < 0.001). Combination treatments with gemcitabine showed no additive cytotoxic or antiproliferative effects (p > 0.05). A transient enhanced effect was observed in the clomipramine + resveratrol group at 24 h (p < 0.01 vs. clomipramine; p < 0.05 vs. gemcitabine). Conclusions: Gemcitabine, clomipramine, and resveratrol all exhibited inhibitory effects on cell proliferation in HL-60 cell cultures. However, the combination treatments did not show additional cytotoxicity or additive effects. These findings suggest that while each of these compounds individually has the potential to inhibit cell growth, their combined application does not enhance the cytotoxic effects beyond those observed with single treatments. These findings highlight the necessity of a rational approach when considering novel drug combinations.
Full article
(This article belongs to the Special Issue Novel Drugs and Natural Products Discovery—2nd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
AWmeta Empowers Adaptively Weighted Transcriptomic Meta-Analysis
by
Yanshi Hu, Zixuan Wang, Yueming Hu, Cong Feng, Qiuyu Fang and Ming Chen
Curr. Issues Mol. Biol. 2026, 48(5), 530; https://doi.org/10.3390/cimb48050530 - 19 May 2026
Abstract
Transcriptomic meta-analysis enhances biological veracity and reproducibility of differentially expressed genes (DEGs) by integrating multiple independent studies, yet prevailing p-value or effect-size integration approaches exhibit limited power to resolve subtle yet vital gene signatures. This study presents AWmeta, an adaptively weighted framework
[...] Read more.
Transcriptomic meta-analysis enhances biological veracity and reproducibility of differentially expressed genes (DEGs) by integrating multiple independent studies, yet prevailing p-value or effect-size integration approaches exhibit limited power to resolve subtle yet vital gene signatures. This study presents AWmeta, an adaptively weighted framework that unifies both meta-analytical paradigms for the first time. Benchmarked on 35 Parkinson’s and Crohn’s disease datasets spanning diverse tissues and adaptively down-weighting underpowered studies, AWmeta yields higher-fidelity DEGs with markedly reduced false positives and achieves more truthful gene differential expression quantification across individual studies at both gene and study levels over the random-effects model (REM). Resilience experiments demonstrate AWmeta’s remarkable stability and robustness against external and internal perturbations. Crucially, AWmeta prioritizes more tissue-contextual genes of Parkinson’s and Crohn’s disease with genuine pathological importance than those from REM and constituent studies. Functional enrichment analysis further verifies that these screened gene signatures capture higher contextual coherence in all analyzed disease tissues. AWmeta harmonizes heterogeneous transcriptomic datasets into reliable DEG identification and mechanistic insights, serving as an indispensable tool for precision transcriptomic integration.
Full article
(This article belongs to the Section Bioinformatics and Systems Biology)
►▼
Show Figures

Graphical abstract
Open AccessReview
How Polyphenol Metabolites Spatiotemporally Reprogram Transcription Factors and Human Proteostasis: A Metabolite-Centric Framework
by
José Manuel Pérez de la Lastra, Celia María Curieses Andrés, Elena Bustamante Munguira, Celia Andrés Juan and Eduardo Pérez Lebeña
Curr. Issues Mol. Biol. 2026, 48(5), 529; https://doi.org/10.3390/cimb48050529 (registering DOI) - 19 May 2026
Abstract
Polyphenols act in humans through authentic metabolites, including regio-isomeric glucuronides/sulphates, O-methylated forms, and microbiota products (urolithins, γ-valerolactones, equol), that reach targets by spatiotemporally gated exposure. Vectorial transport (MRP2/BCRP/P-gp), enterohepatic cycling, and β-glucuronidase hubs create early, surface-proximal microbursts of aglycone/catechol, whereas microbiota metabolites arrive
[...] Read more.
Polyphenols act in humans through authentic metabolites, including regio-isomeric glucuronides/sulphates, O-methylated forms, and microbiota products (urolithins, γ-valerolactones, equol), that reach targets by spatiotemporally gated exposure. Vectorial transport (MRP2/BCRP/P-gp), enterohepatic cycling, and β-glucuronidase hubs create early, surface-proximal microbursts of aglycone/catechol, whereas microbiota metabolites arrive systemically 6–24 h later. Signalling emerges from a continuum of weak noncovalent modulation, conditionally gated electrophile/redox relays (catechol → o-quinone, reversible Michael adduction plus signalling-range H2O2), and PTM cascades (phosphorylation → acylation → proteostasis) that reprogram NRF2/Keap1, NF-κB/IKK, AMPK/MAPK/PI3K-Akt, SIRT1/HDACs, PPARγ, AhR, and TFEB according to where and when metabolites appear. We provide methods and standards to dose isomer-resolved metabolites at physiological free concentrations (nM-low µM) in transport-competent systems, with PK-informed sampling across seconds–minutes, 15/60/240 min, and 6–24 h, and we outline a research agenda (reference panels, spatial exposure atlases, metabotype-stratified trials, safety windows). Framed this way, polyphenols shift from vague “antioxidants” to programmable dietary signals that enable precision nutrition targeting transcription-factor and proteostasis programmes in vivo.
Full article
(This article belongs to the Special Issue Mechanistic Insights into the Antioxidant Activity of Bioorganic Molecules: A Chemical Perspective)
►▼
Show Figures

Figure 1
Open AccessReview
Hormonal Dysregulation and Neuroinflammation in Endometriosis: Convergent Druggable Pathways
by
Ioana-Laura Olteanu, Ciprian Pușcașu, Corina Andrei and Anca Zanfirescu
Curr. Issues Mol. Biol. 2026, 48(5), 528; https://doi.org/10.3390/cimb48050528 - 19 May 2026
Abstract
Endometriosis is a chronic, estrogen-dependent disorder defined by ectopic endometrial-like tissue growth, persistent inflammation, and aberrant innervation. Emerging evidence indicates that disease progression and symptom severity are driven by a reciprocal interaction between hormonal dysregulation and neuroinflammatory signaling. This narrative review synthesizes human-based
[...] Read more.
Endometriosis is a chronic, estrogen-dependent disorder defined by ectopic endometrial-like tissue growth, persistent inflammation, and aberrant innervation. Emerging evidence indicates that disease progression and symptom severity are driven by a reciprocal interaction between hormonal dysregulation and neuroinflammatory signaling. This narrative review synthesizes human-based mechanistic and clinical evidence on the hormonal–neuroinflammatory interface in endometriosis, drawing on peer-reviewed publications retrieved from PubMed and Scopus through November 2025. The publications comprised studies using data from patient-derived tissues, primary endometriotic cells, and clinical cohorts. Several convergent molecular nodes at this interface were identified: the prostaglandin E2–prostaglandin E receptor 2/prostaglandin E receptor 4–aromatase axis, estrogen receptor beta—nuclear factor kappa B signaling, interleukin-6/signal transducer and activator of transcription 3-mediated fibrosis, neurotrophin pathways, transient receptor potential channels (TRPV1/TRPA1), and neurokinin 1 receptor signaling. In this integrated model, endocrine dysfunction fuels neuroinflammation, which in turn impairs steroid responsiveness. This cycle explains the frequent pain–lesion mismatch and the persistence of symptoms despite standard hormonal suppression. Targeting these druggable interface pathways enables better patient stratification and more effective combination therapies for endometriosis.
Full article
(This article belongs to the Special Issue Molecular Pathways and Therapeutic Targets in Endometriosis)
►▼
Show Figures

Figure 1
Open AccessArticle
In Silico Analysis of Polycyclic Aromatic Hydrocarbon (PAH) Degrader from Bordetella petrii Strain P003 Isolated from Contaminated Oil of Kuwait
by
Abrar Akbar, Rita Rahmeh, Mohamed Kishk and Anisha Shajan
Curr. Issues Mol. Biol. 2026, 48(5), 527; https://doi.org/10.3390/cimb48050527 - 18 May 2026
Abstract
Bordetella petrii is an environmentally versatile Gram-negative bacterium with hydrocarbon-degrading capabilities, yet its genetic and metabolic characteristics remain poorly characterized. This study investigated the genomic features of a PAH-degrading Bordetella petrii strain P003 isolated from contaminated oil in Kuwait using bioinformatic approaches. The
[...] Read more.
Bordetella petrii is an environmentally versatile Gram-negative bacterium with hydrocarbon-degrading capabilities, yet its genetic and metabolic characteristics remain poorly characterized. This study investigated the genomic features of a PAH-degrading Bordetella petrii strain P003 isolated from contaminated oil in Kuwait using bioinformatic approaches. The genome of B. petrii P003 was sequenced and analyzed for genomic islands, comparative genomics, and PAH degradation pathways. The draft genome assembly of B. petrii P003 was 5,011,660 bp with 49 contigs and 68.67% GC content. It contained 4687 coding sequences, 5 rRNAs, and 56 tRNAs. Prediction of genomic islands (GIs) revealed that strain P003 possessed 99 GIs, whereas the reference B. pertii DSM 12,804 had 58 unique GIs. Comparative genomics showed 279 locally collinear blocks with the reference strain. The P003 genome encoded multiple genes involved in PAH, naphthalene, and benzoate degradation pathways, including an 8-gene PAH operon (pht4, ph2, pht5, pht3, pcaG, pcaH, nahAb/nagAb/ndoA/nbzA). We found that pcaG and pcaH encode the enzymes responsible for the breakdown of PAH, protocatechuate 3,4-dioxygenase, alpha and beta subunits (EC: 1.13.11.3). The genomic analysis of B. petrii P003 provides insights into its PAH degradation capabilities and potential for bioremediation applications. The strain possesses an expanded repertoire of aromatic compound degradation genes compared to reference strains, suggesting enhanced metabolic versatility for degrading environmental pollutants.
Full article
(This article belongs to the Section Molecular Microbiology)
►▼
Show Figures

Figure 1
Open AccessReview
Reduced CAG Repeats in the Androgen Receptor Gene May Independently Cause Polycystic Ovarian Syndrome
by
Rhea Sharma and Daniel H. Shain
Curr. Issues Mol. Biol. 2026, 48(5), 526; https://doi.org/10.3390/cimb48050526 - 18 May 2026
Abstract
Polycystic ovarian syndrome (PCOS) affects over 116 million women globally and is typically linked with excess androgens such as testosterone. Many patients, however, display classic PCOS symptoms despite normal serum androgen. One proposed mechanism for these cases involves a shortened CAG (i.e., encodes
[...] Read more.
Polycystic ovarian syndrome (PCOS) affects over 116 million women globally and is typically linked with excess androgens such as testosterone. Many patients, however, display classic PCOS symptoms despite normal serum androgen. One proposed mechanism for these cases involves a shortened CAG (i.e., encodes glutamine) repeat length in the androgen receptor (AR) gene, which increases AR activity without elevating testosterone. Fewer glutamine repeats alter the AR’s N-terminal domain and may contribute to strengthened interactions with co-activators and enhanced transcription of androgen-regulated genes. Heightened AR activity in hypothalamus neurons stimulates increased pulsatile release of gonadotropin-releasing hormone (GnRH), which disrupts pituitary secretion dynamics and favors luteinizing hormone (LH) over follicle-stimulating hormone (FSH). This altered LH/FSH ratio leads to impaired folliculogenesis, anovulation and other hallmark PCOS symptoms. Targeting AR activity directly, for example by using compounds that covalently modify the AR N-terminal domain to suppress activity, may therefore offer a more precise treatment strategy for PCOS.
Full article
(This article belongs to the Section Biochemistry, Molecular and Cellular Biology)
►▼
Show Figures

Figure 1
Open AccessArticle
Elevated Tumor HIF-1α Expression Correlates with Advanced Pathological Stage Following Neoadjuvant Concurrent Chemoradiotherapy in Esophageal Squamous Cell Carcinoma
by
Hsin-Yi Shih and Chien-Chih Chen
Curr. Issues Mol. Biol. 2026, 48(5), 525; https://doi.org/10.3390/cimb48050525 - 18 May 2026
Abstract
Tumor hypoxia has been implicated in treatment resistance and disease progression in esophageal squamous cell carcinoma (ESCC), yet its relationship with post-neoadjuvant pathological staging remains unclear. This study evaluated the association between hypoxia-inducible factor-1α (HIF-1α) expression and pathological stage following neoadjuvant concurrent chemoradiotherapy
[...] Read more.
Tumor hypoxia has been implicated in treatment resistance and disease progression in esophageal squamous cell carcinoma (ESCC), yet its relationship with post-neoadjuvant pathological staging remains unclear. This study evaluated the association between hypoxia-inducible factor-1α (HIF-1α) expression and pathological stage following neoadjuvant concurrent chemoradiotherapy (CCRT). We retrospectively analyzed 55 patients with ESCC treated with standardized neoadjuvant CCRT followed by curative esophagectomy. Immunohistochemical staining was performed on surgical specimens to assess tumor (HIF-T%) and stromal (HIF-N%) HIF-1α expression, and correlations with postoperative pathological stage were analyzed. Tumor HIF-1α expression was significantly higher in patients with pathological stage III disease compared with stage I–II disease (40% vs. 15%, p = 0.023). Increasing trends in tumor HIF-T% were observed across higher T and N classifications, although these did not reach statistical significance. Stromal HIF-1α expression was not associated with pathological stage. These findings demonstrate that elevated tumor HIF-1α expression is associated with advanced pathological stage following neoadjuvant CCRT in ESCC, supporting the role of hypoxia-related signaling in treatment resistance. HIF-1α may serve as a clinically relevant biomarker of residual disease burden, although further validation in larger cohorts is warranted.
Full article
(This article belongs to the Special Issue Molecular Markers of Tumor Response and Toxicity of Antitumor Therapy)
►▼
Show Figures

Figure A1
Open AccessArticle
Semaglutide Induces Oxidative Stress and Differentially Modulates mTOR-Dependent Growth and Invasion in Human Trophoblast Cell Models: Implications for Placental Function
by
Elizabeth Thurmond, Eliza J. Roeth, Kristen Noyes, Madeline Boyer, Ethan Evans, Benjamin T. Bikman, Paul R. Reynolds and Juan A. Arroyo
Curr. Issues Mol. Biol. 2026, 48(5), 524; https://doi.org/10.3390/cimb48050524 - 18 May 2026
Abstract
Semaglutide, a long-acting glucagon-like peptide-1 receptor agonist (GLP-1RA), has transformed obesity and diabetes management. However, its expanding use among reproductive-age women raises concerns about potential effects on early placental development. We examined semaglutide’s impact on two human trophoblast cell lines: Swan71 (invasive extravillous)
[...] Read more.
Semaglutide, a long-acting glucagon-like peptide-1 receptor agonist (GLP-1RA), has transformed obesity and diabetes management. However, its expanding use among reproductive-age women raises concerns about potential effects on early placental development. We examined semaglutide’s impact on two human trophoblast cell lines: Swan71 (invasive extravillous) and BeWO (syncytiotrophoblast-like). Cells were treated with semaglutide (100 nM) for 24 h, and proliferation, viability, mitochondrial respiration, oxidative stress, signaling pathways, and invasiveness were evaluated. Semaglutide significantly reduced proliferation in Swan71 cells and increased it in BeWO cells, with no significant change in viability for Swan71 and a slight increase for BeWO. Western blot analysis revealed altered phosphorylation of key signaling proteins, including mTOR, p70S6K, 4EBP1, AKT, and ERK, as well as increased AMPK phosphorylation, indicating a shift toward catabolic signaling. Reactive oxygen species (ROS) accumulation increased markedly, accompanied by altered oxygen consumption rates—reduced in Swan71 cells and elevated in BeWO cells. Functionally, semaglutide suppressed Swan71 invasion through Matrigel by approximately three-fold. These findings suggest that semaglutide induces oxidative and metabolic stress in trophoblasts and is associated with altered mTOR-mediated signaling and reduced invasive potential. Such cellular alterations may contribute to compromised placental development and uterine vascular remodeling if exposure occurs near conception. While clinical data remain limited, this study provides mechanistic insight supporting caution in the use of semaglutide during the periconception period and underscores the need for targeted reproductive safety studies.
Full article
(This article belongs to the Special Issue Advances in Oxidative Stress and Inflammation)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Host-Induced Gene Silencing of SmDSR32 Enhances Wheat Defense Against Sitobion miscanthi
by
Jiahui Zhang, Xue Zhong, Mingxin Cao, Jiajing Xu, Mengchao Qin, Frédéric Francis and Lanqin Xia
Curr. Issues Mol. Biol. 2026, 48(5), 523; https://doi.org/10.3390/cimb48050523 - 17 May 2026
Abstract
The grain aphid, Sitobion miscanthi, poses a serious threat to cereal crops worldwide, leading to considerable yield losses and demanding annual insecticide applications during the grain-filling stage. As a sustainable alternative, we explored host-induced gene silencing (HIGS) targeting an aphid-specific gene. In
[...] Read more.
The grain aphid, Sitobion miscanthi, poses a serious threat to cereal crops worldwide, leading to considerable yield losses and demanding annual insecticide applications during the grain-filling stage. As a sustainable alternative, we explored host-induced gene silencing (HIGS) targeting an aphid-specific gene. In this study, we identified SmDSR32, a novel gene encoding a salivary peptide in S. miscanthi, and validated its suitability for RNAi. Transgenic wheat lines expressing SmDSR32-dsRNA were generated. Aphids feeding on these lines showed a 20-fold reduction in SmDSR32 transcript levels compared with controls. This silencing disrupted normal feeding behavior in electropenetrography (EPG) analyses, characterized by a 1.94-fold prolongation of intercellular probing and a 61% shortening of phloem ingestion. Consequently, aphid performance was severely compromised, with at least a 56.7% decrease in survival, a shortening of 5 days in lifespan, and a reduction of 9–10 individuals in aphid progeny production. Impressively, upon being transferred to wild-type plants, both the surviving aphids and their progeny sustained fitness deficits, with a 30% reduction in survival still observed in the first generation. These findings validate SmDSR32 as a potent RNAi target and establish HIGS targeting essential salivary genes as a promising strategy for sustainable aphid management in wheat.
Full article
(This article belongs to the Special Issue Molecular Mechanisms of Abiotic and Biotic Stress Tolerance in Crops)
►▼
Show Figures

Figure 1
Open AccessArticle
In Vitro Effects of Amygdalin on Proliferation and Apoptosis in SH-SY5Y Neuroblastoma Cells
by
Tuba Gül and Mücahit Seçme
Curr. Issues Mol. Biol. 2026, 48(5), 522; https://doi.org/10.3390/cimb48050522 - 17 May 2026
Abstract
Background and Objectives: Neuroblastoma represents the most common extracranial solid tumor in childhood and is associated with a poor prognosis in high-risk cases. Amygdalin, a naturally occurring cyanogenic glycoside, has been reported to exhibit anti-tumor properties in various cancer models; however, its effects
[...] Read more.
Background and Objectives: Neuroblastoma represents the most common extracranial solid tumor in childhood and is associated with a poor prognosis in high-risk cases. Amygdalin, a naturally occurring cyanogenic glycoside, has been reported to exhibit anti-tumor properties in various cancer models; however, its effects on neuroblastoma cells remain insufficiently characterized. The present study was conducted with the objective of investigating the effects of amygdalin on cell proliferation, apoptosis, and invasion in SH-SY5Y neuroblastoma cells in vitro. Materials and Methods: The SH-SY5Y neuroblastoma cells were cultivated under the optimal conditions for their growth. The cytotoxic effect of amygdalin was determined using the CCK8 assay, which is dose- and time-dependent. Total RNA isolation was performed using Trizol. Subsequently, a process of cDNA synthesis was initiated. The real-time PCR method was utilized to ascertain alterations in the expression levels of mRNA molecules associated with apoptosis, namely Bax, Bcl2, caspase-3, caspase-7, caspase-8, caspase-9, caspase-10, NFkB, and invasion-related genes MMP2, MMP9, TIMP1, and TIMP3. Furthermore, alterations in NFkB levels were examined through the utilization of the ELISA method. Results: The IC50 value of amygdalin in SH-SY5Y cells was determined to be 112.7 µM at 24 h. Amygdalin demonstrated a dose-dependent cytotoxic effect on neuroblastoma cells. Furthermore, the study revealed that the drug induced apoptosis through the upregulation of BAX and BID, and the downregulation of BCL-2 and NF-κB. This process led to a reduction in cell proliferation. Furthermore, the study demonstrated an anti-invasive effect through the downregulation of MMP9 and the upregulation of TIMP1 and TIMP3. In addition, a substantial decrease in NF-κB protein concentration was observed. Conclusions: These findings demonstrate that amygdalin exerts anti-proliferative, pro-apoptotic, and anti-invasive effects in SH-SY5Y neuroblastoma cells in vitro. Amygdalin may represent a promising natural compound for further investigation as a potential therapeutic agent in neuroblastoma.
Full article
(This article belongs to the Section Biochemistry, Molecular and Cellular Biology)
►▼
Show Figures
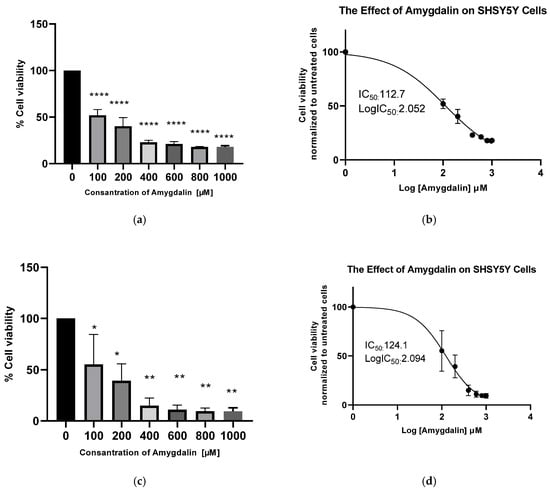
Figure 1
Open AccessArticle
Genome-Wide Analysis of bZIP Transcription Factors and Expression Patterns in Response to Shading Treatment in Taxus yunnanensis
by
Jiangtao Fan, Pengpeng Gong, Yujia Liu, Mengke Dou, Qing Li, Qiuhong Hu, Yong Wang, Gang Wang and Xiong Huang
Curr. Issues Mol. Biol. 2026, 48(5), 521; https://doi.org/10.3390/cimb48050521 - 17 May 2026
Abstract
Basic leucine zipper (bZIP) transcription factors are widely involved in plant growth, development, environmental adaptation, and secondary metabolism. However, the bZIP gene family in Taxus yunnanensis has not been systematically characterized, and its potential involvement in shading-responsive regulation of paclitaxel biosynthesis remains unclear.
[...] Read more.
Basic leucine zipper (bZIP) transcription factors are widely involved in plant growth, development, environmental adaptation, and secondary metabolism. However, the bZIP gene family in Taxus yunnanensis has not been systematically characterized, and its potential involvement in shading-responsive regulation of paclitaxel biosynthesis remains unclear. In this study, a genome-wide analysis was performed to identify and characterize the bZIP family in T. yunnanensis. Phylogenetic analysis, conserved motif and domain identification, promoter cis-element analysis, chromosomal localization, and expression profiling were conducted to investigate their structural features and regulatory potential. A total of 18 TyubZIP genes were identified and classified into 10 subfamilies. These genes exhibited variation in physicochemical properties but showed conserved structural features and nuclear localization. Promoter analysis revealed abundant light-responsive, hormone-related, and stress-related cis-elements. Expression profiling indicated tissue-specific expression patterns and diverse responses to shading treatment. WGCNA further identified candidate TyubZIP genes potentially associated with paclitaxel biosynthesis. Among them, TyuHY5 was selected for functional analysis. Subcellular localization and transcriptional assays demonstrated that TyuHY5 can bind to the promoter of TyuDBTNBT and positively regulate its activity. These findings provide the first genome-wide characterization of the bZIP family in T. yunnanensis and identify TyuHY5 as a shading-responsive candidate regulator of paclitaxel biosynthesis, providing insights that may inform the genetic improvement and cultivation strategies of Taxus for enhanced paclitaxel production.
Full article
(This article belongs to the Special Issue Functional Genomics and Comparative Genomics Analysis in Plants, 4th Edition)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- CIMB Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Editorial Office
Journal Browser
► ▼ Journal Browser-
arrow_forward_ios
Forthcoming issue
arrow_forward_ios Current issue - Volumes not published by MDPI
- Vol. 42 (2021)
- Vol. 41 (2021)
- Vol. 40 (2021)
- Vol. 39 (2020)
- Vol. 38 (2020)
- Vol. 37 (2020)
- Vol. 36 (2020)
- Vol. 35 (2020)
- Vol. 34 (2019)
- Vol. 33 (2019)
- Vol. 32 (2019)
- Vol. 31 (2019)
- Vol. 30 (2019)
- Vol. 29 (2018)
- Vol. 28 (2018)
- Vol. 27 (2018)
- Vol. 26 (2018)
- Vol. 25 (2018)
- Vol. 24 (2017)
- Vol. 23 (2017)
- Vol. 22 (2017)
- Vol. 21 (2017)
- Vol. 20 (2016)
- Vol. 19 (2016)
- Vol. 18 (2016)
- Vol. 17 (2015)
- Vol. 16 (2014)
- Vol. 15 (2013)
- Vol. 14 (2012)
- Vol. 13 (2011)
- Vol. 12 (2010)
- Vol. 11 (2009)
- Vol. 10 (2008)
- Vol. 9 (2007)
- Vol. 8 (2006)
- Vol. 7 (2005)
- Vol. 6 (2004)
- Vol. 5 (2003)
- Vol. 4 (2002)
- Vol. 3 (2001)
- Vol. 2 (2000)
- Vol. 1 (1999)
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
CIMB, IJMS, Reprod. Med., Biology, Life
Recent Research in Germ Cells
Topic Editors: Malgorzata Kloc, Jacek KubiakDeadline: 31 May 2026
Topic in
CIMB, Molecules, Pharmaceuticals, Pharmaceutics, Sci. Pharm.
Challenges and Opportunities in Drug Delivery Research, 2nd Edition
Topic Editors: Lenuta Profire, Ioana Mirela VasincuDeadline: 30 June 2026
Topic in
Cancers, CIMB, Current Oncology, Sci. Pharm., Antibodies, IJMS, IJTM
Antibody-Mediated Therapy and Other Emerging Therapies in Cancer Treatment
Topic Editors: Won Sup Lee, Yaewon Yang, Seil GoDeadline: 31 July 2026
Topic in
IJMS, Agronomy, Plants, Crops, CIMB
New Achievements in Gene Mining, Germplasm Innovation, Cultivation Technologies in Rice
Topic Editors: Shimin Zuo, Yongmei Bao, Yajie HuDeadline: 31 August 2026

Special Issues
Special Issue in
CIMB
Natural Polyphenols in Cardiovascular Disease Prevention: Molecular Mechanisms and Therapeutic Perspectives
Guest Editor: Pasquale PerroneDeadline: 31 May 2026
Special Issue in
CIMB
Cardiovascular Disease: From Molecular Mechanisms to Therapeutic Innovations
Guest Editor: Andrea MarzulloDeadline: 31 May 2026
Special Issue in
CIMB
Multiple Myeloma: From Molecular Mechanism to Diagnosis and Therapy
Guest Editor: Jari IntraDeadline: 31 May 2026
Special Issue in
CIMB
Molecular Mechanisms and Pharmacological Underlying Cardiorenal Diseases
Guest Editor: Caterina CarolloDeadline: 31 May 2026
Topical Collections
Topical Collection in
CIMB
Feature Papers in Current Issues in Molecular BiologyCollection Editor: Madhav Bhatia
Topical Collection in
CIMB
Feature Papers Collection in Molecular Microbiology
Collection Editor: Bruce Seal
Topical Collection in
CIMB
Bioinformatics Approaches to Biomedicine
Collection Editor: Giulia Fiscon
Topical Collection in
CIMB
Molecular Advances in Veterinary Pharmacology and Toxicology
Collection Editor: Chongshan Dai