-
 Neurotransmitter Systems in Alzheimer’s Disease
Neurotransmitter Systems in Alzheimer’s Disease -
 From Molecules to Meaning: Integrating Neuropeptides, Sociostasis, and Hormesis in the Brain–Heart Axis
From Molecules to Meaning: Integrating Neuropeptides, Sociostasis, and Hormesis in the Brain–Heart Axis -
 Transcriptional Divergence of Conserved Starch Metabolism Genes During Grain Filling in Indica and Japonica Rice
Transcriptional Divergence of Conserved Starch Metabolism Genes During Grain Filling in Indica and Japonica Rice -
 Different Effects of Antioxidants Against Ionizing Radiation: An Experimental Model of Micronuclei
Different Effects of Antioxidants Against Ionizing Radiation: An Experimental Model of Micronuclei -
 Anti-Biofilm Activity of Combinations of Cinnamic Acid and Its Derivatives with Cloxacillin Against Methicillin-Resistant Staphylococcus epidermidis
Anti-Biofilm Activity of Combinations of Cinnamic Acid and Its Derivatives with Cloxacillin Against Methicillin-Resistant Staphylococcus epidermidis
Journal Description
Current Issues in Molecular Biology
Current Issues in Molecular Biology
is an international, scientific, peer-reviewed, open access journal on molecular biology, published monthly online by MDPI (from Volume 43, Issue 1 - 2021).
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PMC, PubMed, Embase, CAPlus / SciFinder, FSTA, AGRIS, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.3 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names are published annually in the journal.
Impact Factor:
3.0 (2024);
5-Year Impact Factor:
3.2 (2024)
Latest Articles
Genomic Analysis Highlights the Misinterpretation of Acquired Aminoglycoside Resistance Genes in Deinococcus radiodurans
Curr. Issues Mol. Biol. 2026, 48(5), 505; https://doi.org/10.3390/cimb48050505 (registering DOI) - 14 May 2026
Abstract
Aminoglycoside resistance is commonly mediated by enzymatic modification, target alteration, or efflux mechanisms; however, acquired resistance has not been characterized in radiation-resistant Deinococcus species. Here, we investigated the occurrence and genomic context of acquired aminoglycoside resistance genes in all publicly available Deinococcus radiodurans
[...] Read more.
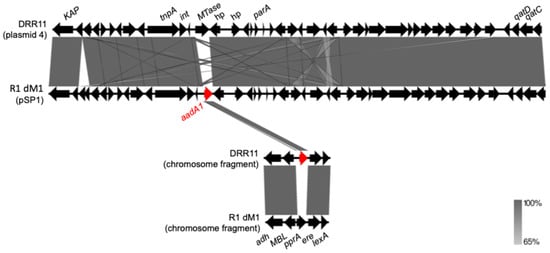
Aminoglycoside resistance is commonly mediated by enzymatic modification, target alteration, or efflux mechanisms; however, acquired resistance has not been characterized in radiation-resistant Deinococcus species. Here, we investigated the occurrence and genomic context of acquired aminoglycoside resistance genes in all publicly available Deinococcus radiodurans genomes. A total of 19 genomes were screened using ResFinder and CARD, followed by comparative genomic analyses. The aadA1 gene was identified in two genomes, being located on the plasmid pSP1 in strain R1 dM1, a known shuttle vector used for genetic manipulation. In contrast, aadA1 was found on a chromosomal contig in strain DRR11, suggesting a possible assembly artifact. Additionally, the aph(3′)-Ia gene was detected in three genomes within a conserved chromosomal region that lacks this gene in reference strains. Sequence similarity analyses indicated that aph(3′)-Ia is associated with laboratory vectors, being consistent with a potential non-natural origin. Considering the high recombination capacity and genomic plasticity of D. radiodurans, these findings suggest that the detected aminoglycoside resistance genes may be derived from laboratory constructs, potentially combined with assembly inconsistencies or chromosomal integration events. Therefore, this study highlights the importance of integrating genomic context with molecular and evolutionary plausibility to avoid misinterpretation of antimicrobial resistance in extremophiles and model organisms, and underscores the importance of complementary raw-read analyses to distinguish natural acquisition from technical or laboratory-derived origins.
Full article
(This article belongs to the Section Bioinformatics and Systems Biology)
►
Show Figures
Open AccessCorrection
Correction: Lai et al. Association Between Catenin Beta-1 (CTNNB1) Gene Polymorphisms and Non-Traumatic Osteonecrosis of the Femoral Head (ONFH). Curr. Issues Mol. Biol. 2026, 48, 164
by
I-Chang Lai, De-Yi Liu, Shih-Chan Hsu and Shu-Jui Kuo
Curr. Issues Mol. Biol. 2026, 48(5), 504; https://doi.org/10.3390/cimb48050504 (registering DOI) - 14 May 2026
Abstract
In the published article [...]
Full article
(This article belongs to the Section Bioinformatics and Systems Biology)
Open AccessArticle
Phytochemical Analysis, GC-MS Chemical Profiling, and In Vitro Antidiabetic Evaluation of South African Momordica balsamina Linn Leaf Extracts and Its Effects on Oxidative Stress Modulation
by
Buang Matseke, Daniel Tswaledi and Kokoette Bassey
Curr. Issues Mol. Biol. 2026, 48(5), 503; https://doi.org/10.3390/cimb48050503 (registering DOI) - 13 May 2026
Abstract
Background: Momordica balsamina L. is widely used in traditional medicine for the management of diabetes in South Africa and globally. This study evaluated the in vitro antidiabetic and cytotoxic effects of M. balsamina leaf extracts and identified bioactive compounds potentially responsible for its
[...] Read more.
Background: Momordica balsamina L. is widely used in traditional medicine for the management of diabetes in South Africa and globally. This study evaluated the in vitro antidiabetic and cytotoxic effects of M. balsamina leaf extracts and identified bioactive compounds potentially responsible for its activity. Methods: Leaves were sequentially extracted using solvents of increasing polarity. Phytochemical composition was determined using standard colorimetric assays, while gas chromatography–mass spectrometry (GC–MS) was employed for compound identification. Antioxidant activity was evaluated using dot blot, DPPH radical scavenging, hydrogen peroxide scavenging, and ferric reducing power assays. Antidiabetic potential was assessed using α-amylase, α-glucosidase, and β-glucosidase inhibitory assays, with acarbose as the reference drug. Cytotoxicity was determined by using the MTT assay on Vero and HEK-293 cell lines. Results: Phytochemical screening revealed alkaloids, flavonoids, terpenoids, saponins, glycosides, and steroids. GC–MS analysis identified compounds associated with antidiabetic activity, including vanillin, 2,4-di-tert-butylphenol, oleic acid, phytol, and hexadecenoic acid. All extracts exhibited antioxidant activity, with the ethyl acetate extract showing the strongest effect. Enzyme inhibition was concentration dependent. The dichloromethane and ethyl acetate extracts showed stronger α-amylase inhibition (IC50 = 0.149 and 0.146 mg/mL) than acarbose (0.209 mg/mL). For α-glucosidase, acarbose showed the highest activity, while extracts displayed moderate inhibition. In β-glucosidase assays, both extracts were more active than acarbose. Both extracts were non-cytotoxic up to 500 µg/mL. Conclusions: These findings support the traditional use of M. balsamina and highlight its potential as a safe source of antidiabetic agents, warranting further investigation.
Full article
(This article belongs to the Special Issue Advances in Phytochemical Research: Molecular Pathways in Health and Disease)
►▼
Show Figures
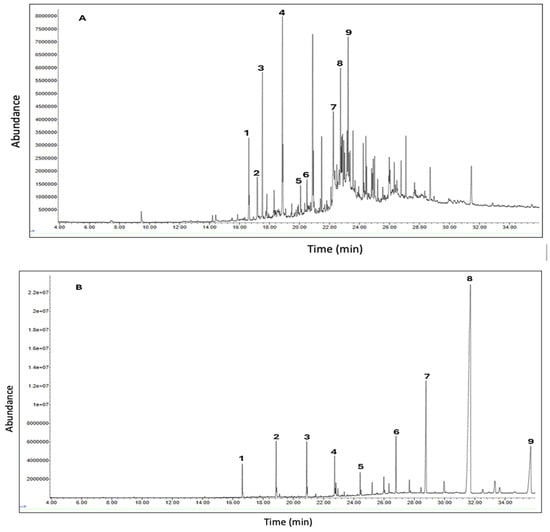
Figure 1
Open AccessReview
Molecular Subtypes of Pancreatic Cancer: A Review of the Literature
by
Jakub Wnuk, Wiktoria Skowron, Anna Długaszek, Joanna Sadurska, Łukasz Pietrzyński, Jacek Kabut and Iwona Gisterek-Grocholska
Curr. Issues Mol. Biol. 2026, 48(5), 502; https://doi.org/10.3390/cimb48050502 (registering DOI) - 13 May 2026
Abstract
Pancreatic cancer (PC) is considered one of the deadliest cancers worldwide, and the number of PC-related deaths is expected to increase. Early diagnosis of PC is crucial for improving treatment outcomes. Despite improvements in overall survival (OS) in metastatic and unresectable PC due
[...] Read more.
Pancreatic cancer (PC) is considered one of the deadliest cancers worldwide, and the number of PC-related deaths is expected to increase. Early diagnosis of PC is crucial for improving treatment outcomes. Despite improvements in overall survival (OS) in metastatic and unresectable PC due to systemic therapy, there is still a need to search for novel therapies and factors predictive of response to treatment. Cancer profiling based on genome sequencing can be used to develop targeted therapies and improve prognostics and treatment outcomes. Therefore, this review was conducted to evaluate the clinical value of molecular subtyping in pancreatic cancer as a prognostic and predictive factor in pancreatic cancer. Due to its limitations, including the lack of a registered protocol and risk of bias assessment for the included studies and those whose results were not included, this study should be considered a narrative review with a structured search strategy rather than a systematic review.
Full article
(This article belongs to the Special Issue Molecular Approaches in Early Detection and Targeted Therapies for Pancreatic Cancer)
►▼
Show Figures
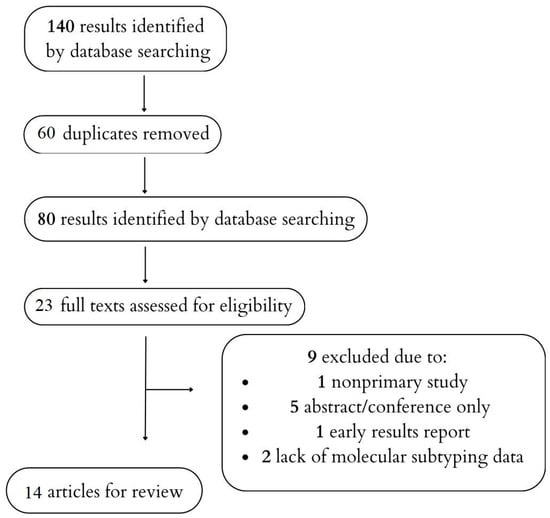
Figure 1
Open AccessArticle
Sodium Butyrate Attenuates Isoprenaline-Induced Myocardial Injury via Restoring the Gut–Heart Axis and Suppressing TLR4/NF-κB Signaling
by
Hazrat Bilal, Imran Khan, Ayesha Yaseen, Xiaopeng Zhang, Xuexue Liu, Jian Zhao, Jing Li, Ata Ur Rehman, Lei Sun and Xiao Yu
Curr. Issues Mol. Biol. 2026, 48(5), 501; https://doi.org/10.3390/cimb48050501 (registering DOI) - 13 May 2026
Abstract
The gut–heart axis plays a role in cardiac injury due to the disruption of barriers, endotoxemia, and inflammatory signaling; yet, it is not clear whether sodium butyrate (SB) is capable of alleviating isoprenaline-induced myocardial injury through coordinated intestinal, microbial, and metabolic restoration. This
[...] Read more.
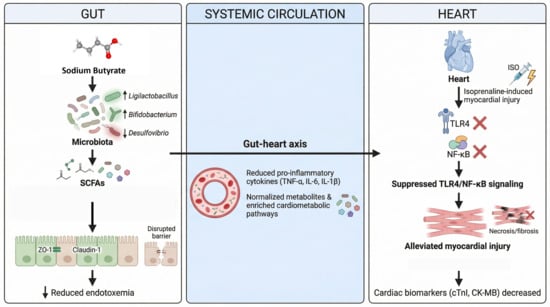
The gut–heart axis plays a role in cardiac injury due to the disruption of barriers, endotoxemia, and inflammatory signaling; yet, it is not clear whether sodium butyrate (SB) is capable of alleviating isoprenaline-induced myocardial injury through coordinated intestinal, microbial, and metabolic restoration. This study used male Sprague-Dawley rats, which were grouped into control, control + SB, isoprenaline (ISO)-induced myocardial injury, and ISO + SB groups. We evaluated cardiac biomarkers of injury, oxidative stress, histopathologic, intestinal barrier (16S rRNA sequencing), and serum metabolomics (LC-MS). SB treatment decreased serum cardiac troponin I, creatine kinase-MB, and lactate dehydrogenase; relieved oxidative stress; and lowered myocardial necrosis and fibrosis. It re-established colonic architecture, upregulated the expression of ZO-1 (zonula occludens-1) and claudin-1, and reduced endotoxin in the bloodstream. SB also prevented the production of proinflammatory cytokines such as TNF-α, IL-6, and IL-1β; cardiac TLR4; IκBα degradation; and NF-κB p 65 phosphorylation. In addition, SB altered the gut microbiota in favor of beneficial commensals, including Ligilactobacillus and Bifidobacterium, and reduced Desulfovibrio. It normalized key circulating metabolites and enriched cardiometabolic pathways, and the patterns of correlation suggested the coordinated remodeling of the microbiome–metabolome. These findings reveal that SB prevents myocardial injury caused by ISO through strengthening gut barrier protection, alleviating endotoxemia, inhibiting TLR4/NF-κB, and remodeling the microbiome–metabolome axis, indicating its potential for use as a gut-targeted cardioprotective intervention.
Full article
(This article belongs to the Special Issue Molecules at Play in Cardiovascular Diseases)
►▼
Show Figures
Graphical abstract
Open AccessArticle
DNA Barcoding and Comparative Chloroplast Marker Performance in Endemic Plants of Crete (Greece)
by
Dimitra Ioannidou, Ioulietta Samartza, Georgios Tsoktouridis, Andreas D. Drouzas and Nikos Krigas
Curr. Issues Mol. Biol. 2026, 48(5), 500; https://doi.org/10.3390/cimb48050500 (registering DOI) - 13 May 2026
Abstract
Crete, a major Mediterranean biodiversity hotspot, hosts many local endemic, threatened and/or protected plant taxa (species and subspecies). Besides their ecological and conservation significance, these unique phytogenetic resources hold significant economic potential for sustainable utilization. Since DNA barcoding is critical for conservation, taxonomy,
[...] Read more.
Crete, a major Mediterranean biodiversity hotspot, hosts many local endemic, threatened and/or protected plant taxa (species and subspecies). Besides their ecological and conservation significance, these unique phytogenetic resources hold significant economic potential for sustainable utilization. Since DNA barcoding is critical for conservation, taxonomy, and plant-derived product authentication, we studied 15 local Cretan endemic taxa using three chloroplast DNA (cpDNA) regions (rbcL, trnL, trnH-psbA). A comparative analysis against GenBank (NCBI) records revealed significant new data: (i) the first genetic information for five taxa (Centaurea redempta subsp. redempta, Galium fruticosum, Micromeria hispida, Salix kaptarae, Teucrium cuneifolium); (ii) new marker-specific sequences for seven taxa (Helichrysum heldreichii, Scutellaria hirta, Sesleria doerfleri, Staehelina petiolata, Teucrium alpestre, Campanula pelviformis, Phlomis lanata); and (iii) novel genotypes of already represented markers for three species (Phlomis lanata, Scutellaria sieberi, Staehelina petiolata). Phylogenetic analyses were performed for all three molecular markers across selected members of Scutellaria section Scutellaria, Teucrium section Polium, and Campanula section Quinqueloculares. The overall results indicated that, amongst the studied species, the trnH-psbA marker is more suitable for species-level identification, whereas the rbcL and trnL markers were more helpful to genus-level identification within Lamiaceae and Campanulaceae. These results enrich the DNA barcoding reference library and form a concrete contribution towards the protection, conservation and traceability of Crete’s unique botanical heritage.
Full article
(This article belongs to the Special Issue Molecular Breeding and Genetics Research in Plants—3rd Edition)
►▼
Show Figures

Figure 1
Open AccessReview
Oncolytic Herpes Simplex Virus for Glioblastoma: Molecular Engineering, Tumor Microenvironment Barriers, and Clinical Translation
by
Jiayu Liu, Yuxin Wang, Zhao Gao, Tongtan Liu, Ao Xu, Wenxuan Li, Mei Li, Xiaomeng Song, Baorui Guo, Huadong Wang, Wenying Lv and Jianning Zhang
Curr. Issues Mol. Biol. 2026, 48(5), 499; https://doi.org/10.3390/cimb48050499 (registering DOI) - 13 May 2026
Abstract
Glioblastoma (GBM) remains the most aggressive primary malignant brain tumor in adults, with limited survival benefit from the current standard of care consisting of maximal safe resection, radiotherapy, and temozolomide-based chemotherapy. The highly infiltrative growth pattern, profound intratumoral heterogeneity, and strongly immunosuppressive tumor
[...] Read more.
Glioblastoma (GBM) remains the most aggressive primary malignant brain tumor in adults, with limited survival benefit from the current standard of care consisting of maximal safe resection, radiotherapy, and temozolomide-based chemotherapy. The highly infiltrative growth pattern, profound intratumoral heterogeneity, and strongly immunosuppressive tumor microenvironment together contribute to therapeutic resistance and frequent recurrence. In this context, oncolytic herpes simplex virus (oHSV) has emerged as a promising therapeutic platform for glioblastoma because of its dual capacity to directly lyse tumor cells and stimulate antitumor immune responses. In addition, the large viral genome and well-characterized biology of herpes simplex virus enable extensive genetic engineering to improve tumor selectivity, safety, and immunomodulatory function. In this review, we summarize the molecular design strategies that have driven the development of oHSV for glioblastoma, including attenuation of neurovirulence, enhancement of tumor-selective replication, and arming with immune-stimulatory transgenes. We further discuss the major biological barriers within the GBM tumor microenvironment that continue to limit therapeutic efficacy, with particular attention given to representative engineered oHSV platforms and the lessons learned from preclinical and early-phase clinical studies. A dedicated section examines these barriers in detail, including restricted intratumoral viral spread, antiviral innate immunity, and immunosuppressive myeloid cell dominance. We also review current efforts to improve outcomes through rational combination strategies with radiotherapy, immune checkpoint blockade, cytokine modulation, and other multimodal approaches. Although encouraging advances have been achieved, the clinical translation of oHSV therapy for glioblastoma still faces substantial challenges in patient selection, delivery optimization, response assessment, and treatment integration. A deeper understanding of virus–host–tumor interactions and more precise engineering of viral platforms may help unlock the full potential of oHSV-based therapy. Overall, oHSV represents one of the most compelling translational approaches in glioblastoma and provides a valuable framework for the development of mechanism-driven viro-immunotherapy in neuro-oncology.
Full article
(This article belongs to the Special Issue Advanced Research in Glioblastoma and Neuroblastoma)
►▼
Show Figures
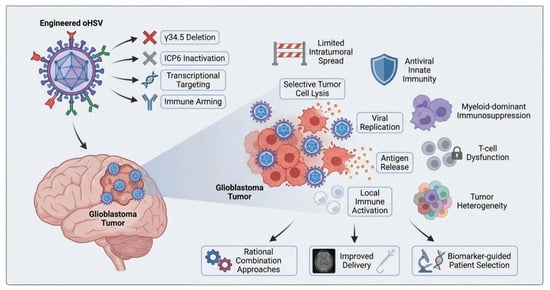
Figure 1
Open AccessReview
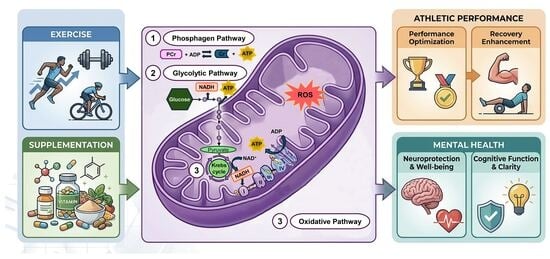
Revision of Energy Metabolism Adaptations in High-Level Athletes: From Physical Performance Enhancement to Potential Therapeutic Targets in Mental Disorders
by
Ane Larrea, Mariyem Naji, Marina Gulak, Maria Torrecilla and Gabriel Barreda-Gómez
Curr. Issues Mol. Biol. 2026, 48(5), 498; https://doi.org/10.3390/cimb48050498 (registering DOI) - 11 May 2026
Abstract
High-level athletic performance requires the implementation of personalized strategies based on the analysis of metabolic pathways involved in energy production: phosphagen, glycolytic, and oxidative pathways. In this context, mitochondria play an essential role as the central regulator of energy production, being closely linked
[...] Read more.
High-level athletic performance requires the implementation of personalized strategies based on the analysis of metabolic pathways involved in energy production: phosphagen, glycolytic, and oxidative pathways. In this context, mitochondria play an essential role as the central regulator of energy production, being closely linked to these three pathways. Exercise boosts cellular respiration, which can also be optimized by nutritional interventions and targeted supplementation, promoting mitochondrial biogenesis, reducing oxidative stress and increasing ATP production. These metabolic adaptations improve athletic performance, accelerate recovery processes, and reduce the risk of injury, adapting to the physiological characteristics of each athlete. Moreover, some of these metabolic adaptations converge on specific targets whose expression or activity is also altered in mental disorders. Therefore, the aim of this review is to analyze mitochondrial adaptations induced by exercise and supplementation, evaluating their impact on the phosphagen, glycolytic, and oxidative metabolic pathways, as well as their relationship with optimizing performance and recovery in high-level athletes, with special attention to their potential application to mental health.
Full article
(This article belongs to the Special Issue Recent Advances in Energy Metabolism)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Application of Raman Spectroscopy to Rapid Discrimination of Autochthonous Lactic Acid Bacteria Isolated from Goat Cheese
by
Ana Yanina Bustos, Juan José Carol Paz, Jorge Nicolás Gómez and Ana Estela Ledesma
Curr. Issues Mol. Biol. 2026, 48(5), 497; https://doi.org/10.3390/cimb48050497 (registering DOI) - 11 May 2026
Abstract
The rapid characterization of lactic acid bacteria (LAB) with probiotic and technological properties is crucial for functional food design. In this study, fourteen LAB strains belonging to the species Lactiplantibacillus (L.) plantarum, Lentilactobacillus (L.) parabuchneri, and Leuconostoc
[...] Read more.
The rapid characterization of lactic acid bacteria (LAB) with probiotic and technological properties is crucial for functional food design. In this study, fourteen LAB strains belonging to the species Lactiplantibacillus (L.) plantarum, Lentilactobacillus (L.) parabuchneri, and Leuconostoc (L.) mesenteroides were differentiated using Raman spectroscopy. By integrating Principal Component Analysis (PCA) and Linear Discriminant Analysis (LDA), we achieved a clear inter-generic separation while simultaneously enabling the intra-specific grouping of L. plantarum strains. Our results demonstrate that the Raman spectral fingerprint, coupled with supervised chemometric models, successfully categorized the strains into three distinct clusters based on their macromolecular profiles. Specifically, the analysis provided high-resolution differentiation between genera and, more importantly, allowed for the fine-scale clustering of diverse L. plantarum isolates. This highlights Raman spectroscopy as a robust, non-destructive tool for the rapid identification and taxonomic classification of LAB, offering a high-throughput alternative to traditional molecular methods for strain-level discrimination.
Full article
(This article belongs to the Special Issue Molecular Dynamics Simulations in Structural Biology: From Proteins to Complex Biomolecular Systems)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Potential Prognostic and Metastatic Implications of MACC1 and MMP8 in Colorectal Cancer
by
Hilal Oğuz Soydinç, Sena Şen, Murat Serilmez and Senem Karabulut
Curr. Issues Mol. Biol. 2026, 48(5), 496; https://doi.org/10.3390/cimb48050496 (registering DOI) - 11 May 2026
Abstract
Colorectal cancer (CRC) remains a major cause of cancer-related morbidity and mortality worldwide. Metastasis regulators and matrix metalloproteinases have been implicated in tumor progression; however, their clinical significance in CRC remains incompletely defined. In this study, the prognostic value of MACC1 and MMP8
[...] Read more.
Colorectal cancer (CRC) remains a major cause of cancer-related morbidity and mortality worldwide. Metastasis regulators and matrix metalloproteinases have been implicated in tumor progression; however, their clinical significance in CRC remains incompletely defined. In this study, the prognostic value of MACC1 and MMP8 expression levels was investigated. A total of 140 patients diagnosed with CRC and 48 healthy controls were included. Serum levels of MACC1 and MMP8 were measured using ELISA. Clinicopathological parameters were recorded, and their associations with biomarker expression were analyzed. Both MACC1 and MMP8 levels demonstrated moderate diagnostic performance with comparable area under the curve values. A strong positive correlation between MACC1 and MMP8 expression was observed. MACC1 expression was significantly associated with metastasis status and tumor stage, whereas MMP8 expression was associated with tumor localization. In survival analyses, established clinicopathological factors, particularly tumor stage and metastasis status, were identified as the primary determinants of overall survival. In multivariate analysis, tumor stage remained the only consistent independent prognostic factor, while MMP8 showed a modest independent association in a separate model. MACC1 did not retain independent prognostic significance. Although MACC1 and MMP8 may have diagnostic and biological relevance in CRC, their prognostic utility appears limited compared to established clinical parameters. Further large-scale prospective studies are needed.
Full article
(This article belongs to the Section Molecular Medicine)
►▼
Show Figures

Figure 1
Open AccessArticle
In Vitro Anti-Inflammatory Activity and Molecular Docking Analysis of Compounds Isolated from Beyeria viscosa
by
Hamza Shahid, James P. Flood, Feng Li, Xian Zhou, Gerald Münch and Ritesh Raju
Curr. Issues Mol. Biol. 2026, 48(5), 495; https://doi.org/10.3390/cimb48050495 (registering DOI) - 10 May 2026
Abstract
Inflammation contributes to the progression of numerous chronic diseases, and plants are a rich source of bioactive secondary metabolites. In this study, bioassay-guided isolation of the previously unexplored Australian native plant Beyeria viscosa (Labill.) Miq. yielded eleven known compounds (1–11
[...] Read more.
Inflammation contributes to the progression of numerous chronic diseases, and plants are a rich source of bioactive secondary metabolites. In this study, bioassay-guided isolation of the previously unexplored Australian native plant Beyeria viscosa (Labill.) Miq. yielded eleven known compounds (1–11). The chemical structures of these compounds were identified by detailed spectroscopic data analysis, and definitive structural confirmation was established using single-crystal X-ray crystallography for fritillebic acid (8) and herbacetin 3,7,8-trimethyl ether (5) for the first time. All compounds were first screened for nitric oxide (NO) inhibitory activity and cytotoxicity in lipopolysaccharide (LPS) and interferon (IFN)-γ-stimulated RAW 264.7 macrophages, and the active NO inhibitors were further assessed for tumor necrosis factor-α (TNF-α) and interleukin-6 (IL-6) inhibition. Notably, compounds 5, 8, and 10 were evaluated for NO inhibitory activity for the first time, with compound 8 being the most potent (IC50 = 8.8 ± 1.3 μM), compound 10 showing moderate potency (IC50 = 12.2 ± 8.8 μM), and compound 5 being inactive. Among all tested compounds, fritillebic acid (8) emerged as the most active constituent, showing strong NO inhibition and moderate suppression of TNF-α and IL-6 production; therefore, it was further assessed in LPS-stimulated N-11 microglial cells, where it retained NO inhibitory activity (IC50 = 12.3 ± 0.5 μM) with a favorable activity–cytotoxicity profile (LC50 = 107.9 ± 1.9 μM). Consistent with the promising activity, molecular docking of compound 8 showed strong receptor-binding affinity with selected inflammation-related targets. Moreover, preliminary structure–activity relationship analysis of all isolated compounds suggested that substitution and oxygenation patterns may influence NO inhibitory potency. Overall, these findings identify fritillebic acid as the major anti-inflammatory lead from B. viscosa and highlight Australian native plants as a source of bioactive secondary metabolites.
Full article
(This article belongs to the Special Issue Biologically Active Compounds: Sources, Mechanisms of Action, and Applications)
Open AccessArticle
Immune-Enhancing Effects of Polygonatum cyrtonema Polysaccharides in Immunodeficient Zebrafish
by
Daoyuan Li, Jie Wang, Naifu Chen and Naidong Chen
Curr. Issues Mol. Biol. 2026, 48(5), 494; https://doi.org/10.3390/cimb48050494 (registering DOI) - 9 May 2026
Abstract
To evaluate the immune-enhancing effects of Polygonatum cyrtonema polysaccharides in vivo, an immunodeficiency zebrafish model was established by microinjecting vinorelbine tartrate into the caudal vein. Effects of the polysaccharides (500, 1000 and 2000 μg/mL) on neutrophil counts were assessed in Tg (mpx:GFP) zebrafish.
[...] Read more.
To evaluate the immune-enhancing effects of Polygonatum cyrtonema polysaccharides in vivo, an immunodeficiency zebrafish model was established by microinjecting vinorelbine tartrate into the caudal vein. Effects of the polysaccharides (500, 1000 and 2000 μg/mL) on neutrophil counts were assessed in Tg (mpx:GFP) zebrafish. Transcriptome sequencing was employed to investigate the immunomodulatory effects of the polysaccharides. The results revealed a dose-dependent increase in neutrophil counts following treatment with the polysaccharides. Transcriptomic profiling identified 1286 DEGs across the three comparison groups. GO and KEGG enrichment analyses indicated that the polysaccharides could modulate immune-related pathways in the zebrafish model. Two enriched KEGG pathways, including the MAPK signaling and the mTOR signaling pathway, were utilized to analyze immune-related gene expression. To validate RNA-seq data, qRT-PCR was performed on selected DEGs, including il1b, crk, fgf10b, atp6v1aa, and eif4e1c. The results confirmed that the expression patterns of these genes were consistent with the RNA-seq data. Within the tested concentrations (500, 1000 and 2000 μg/mL), the polysaccharides exhibited a dose-dependent immunostimulatory effect, with the highest immunostimulatory response observed at 2000 μg/mL. The molecular level primarily involves the enhancement of neutrophil function through the modulation of multiple immune-related pathways. These findings provide a theoretical basis for the potential application of Polygonatum cyrtonema polysaccharides as a natural immunomodulatory agent.
Full article
(This article belongs to the Section Bioorganic Chemistry and Medicinal Chemistry)
Open AccessReview
Microbial Dysbiosis in Photodermatoses: Formation, Pathogenesis and Intervention Strategies
by
Lanhai Zhong, Tian Wang, Lu Tang, Jiande Han, Qun Zhao and Naiyu Lin
Curr. Issues Mol. Biol. 2026, 48(5), 493; https://doi.org/10.3390/cimb48050493 (registering DOI) - 9 May 2026
Abstract
Recent studies have reported skin microbiome dysbiosis in patients with photodermatoses, featuring enriched Staphylococcus aureus colonization and decreased microbiome diversity. We propose that ultraviolet radiation (UVR), along with atypical antimicrobial peptides, may exert selective pressure on the skin microbiome, while cytokine dysregulation and
[...] Read more.
Recent studies have reported skin microbiome dysbiosis in patients with photodermatoses, featuring enriched Staphylococcus aureus colonization and decreased microbiome diversity. We propose that ultraviolet radiation (UVR), along with atypical antimicrobial peptides, may exert selective pressure on the skin microbiome, while cytokine dysregulation and a reduction in commensal bacteria amplify microbial dysbiosis. Dysbiotic microorganisms further release pathogen-associated patterns and virulence factors, and activate tissue-resident memory T cells, which collectively contribute to local inflammation. These mechanisms establish the skin microbiome as a potential target for early intervention. Potential therapeutic strategies may include antibiotics, phototherapy, bleach baths, phage therapy, and microbiota-based therapies. This review integrates current findings from microbial ecology, molecular biology, and host immunology to outline a conceptual framework linking UVR exposure, microbiome alterations, and cutaneous immune responses, while emphasizing the current limitations and evidence gaps in this field.
Full article
(This article belongs to the Special Issue Exploring Molecular Pathways in Skin Health and Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Molecular Target Discovery and Systemic Mechanism Analysis of Teriflunomide for Dry Eye Disease
by
Yang Chen, Weiran Lin, Wei Feng, Wenyuan Li and Lianhao Song
Curr. Issues Mol. Biol. 2026, 48(5), 492; https://doi.org/10.3390/cimb48050492 (registering DOI) - 9 May 2026
Abstract
Background: Dry eye disease (DED) is a multifactorial ocular surface disorder characterized by tear film instability, inflammation, and neurosensory abnormalities. Current therapies remain limited by slow onset and suboptimal efficacy. Teriflunomide, an immunomodulatory agent approved for multiple sclerosis, has shown therapeutic potential in
[...] Read more.
Background: Dry eye disease (DED) is a multifactorial ocular surface disorder characterized by tear film instability, inflammation, and neurosensory abnormalities. Current therapies remain limited by slow onset and suboptimal efficacy. Teriflunomide, an immunomodulatory agent approved for multiple sclerosis, has shown therapeutic potential in DED, but its multi-target mechanisms remain unclear. Methods: We employed an integrated computational and transcriptomic framework combining ADMET profiling, multi-dataset transcriptomic integration, and single-cell RNA sequencing (scRNA-seq) to identify disease-relevant targets. Candidate genes were further refined through molecular docking and 50 ns molecular dynamics (MD) simulations. The AetherCell virtual cell model was applied to evaluate both the concordance between target perturbation and drug-induced responses and the potential mechanistic roles of candidate targets. Results: Transcriptomic integration identified 16 consensus genes across heterogeneous DED models, which were further localized to disease-relevant epithelial and immune cell populations by scRNA-seq. Molecular simulations prioritized three core targets—CTSS, STAT1, and PTGS1—based on binding stability and affinity. AetherCell simulations demonstrated that perturbation of these targets not only recapitulated teriflunomide-induced transcriptional and pathway changes but also revealed their distinct mechanistic contributions, including epithelial barrier regulation (CTSS), microvascular and lipid homeostasis (PTGS1), and inflammation suppression coupled with tissue repair (STAT1). Conclusions: Teriflunomide exerts therapeutic effects in DED through coordinated multi-target regulation involving inflammation control, barrier restoration, and tissue repair. This study provides a rationale for novel therapeutic targets in dry eye disease, establishes a paradigm for applying virtual cell modeling to elucidate drug mechanisms, and offers a bioinformatics framework for validating drug repositioning outcomes.
Full article
(This article belongs to the Section Molecular Medicine)
►▼
Show Figures

Figure 1
Open AccessArticle
Transcriptomic Profiling Identifies Potential Prognostic Genes in Vietnamese Patients with Non-Small-Cell Lung Cancer
by
Tuan Quoc Bach, Giang Thi Chau Truong, Bang Ngoc Dao, Thang Ba Ta and Thuy Thi Bich Vo
Curr. Issues Mol. Biol. 2026, 48(5), 491; https://doi.org/10.3390/cimb48050491 (registering DOI) - 9 May 2026
Abstract
Background/Objectives: Non-small-cell lung cancer (NSCLC) is one of the most common malignancies in Vietnam, yet its molecular mechanisms remain incompletely understood. This study aimed to identify prognostic genes in Vietnamese NSCLC patients using integrative transcriptomic and bioinformatics analyses. Methods: RNA-seq data from 30
[...] Read more.
Background/Objectives: Non-small-cell lung cancer (NSCLC) is one of the most common malignancies in Vietnam, yet its molecular mechanisms remain incompletely understood. This study aimed to identify prognostic genes in Vietnamese NSCLC patients using integrative transcriptomic and bioinformatics analyses. Methods: RNA-seq data from 30 Vietnamese NSCLC patients treated at Military Hospital 103 (January 2023–April 2024) were analyzed and cross-validated with the Gene Expression Omnibus (GEO) dataset GSE140343 to identify shared differentially expressed genes (DEGs). Subsequent analyses included functional enrichment (GO and KEGG), protein–protein interaction (PPI) network construction via STRING, and module/centrality analyses to pinpoint hub genes. Finally, prognostic significance was evaluated using overall survival data from The Cancer Genome Atlas (TCGA) via the GEPIA platform. Results: A total of 1900 shared DEGs were identified, most of which were enriched in cancer-related pathways. The resulting PPI network (comprising 1528 nodes and 8185 edges) yielded eight significant modules containing 64 high-centrality candidate genes. Survival analyses demonstrated that high expression of CCNA2 and S100A12, and low expression of ADRB2, ARRB1, PTGS2, and SMAD7 were significantly associated with poor overall survival in NSCLC patients. Conclusions: These findings highlight potential biomarkers for prognosis and may inform future therapeutic strategies in Vietnamese NSCLC patients.
Full article
(This article belongs to the Special Issue Bioinformatics in Human Disease Network Analysis)
►▼
Show Figures

Figure 1
Open AccessArticle
Cytotoxic and Antimelanoma Activity of Selected 3-Methyl-1,6-diazaphenothiazines in Human Melanoma Cells—In Vitro Studies
by
Beata Morak-Młodawska, Małgorzata Jeleń, Zuzanna Rzepka, Milena Koch and Dorota Wrześniok
Curr. Issues Mol. Biol. 2026, 48(5), 490; https://doi.org/10.3390/cimb48050490 (registering DOI) - 9 May 2026
Abstract
The cytotoxic and mechanistic effects of novel 10-substituted 3-methyl-1,6-diazaphenothiazines were investigated in human melanoma models. Antiproliferative activity was evaluated in vitro using the WST-1 assay in four melanoma cell lines (A375, C32, G361, and SK-MEL-28) and normal human dermal fibroblasts (HDF). Among the
[...] Read more.
The cytotoxic and mechanistic effects of novel 10-substituted 3-methyl-1,6-diazaphenothiazines were investigated in human melanoma models. Antiproliferative activity was evaluated in vitro using the WST-1 assay in four melanoma cell lines (A375, C32, G361, and SK-MEL-28) and normal human dermal fibroblasts (HDF). Among the tested derivatives, compound 6 exhibited the most pronounced biological activity, showing the strongest growth inhibition in melanoma cells, with the lowest IC50 value against C32 cells (54 µM), while displaying lower toxicity toward normal fibroblasts. Mechanistic studies using image cytometry and immunofluorescence revealed that compound 6 profoundly disrupts melanoma cell homeostasis by suppressing cell proliferation, inducing DNA damage, and activating apoptotic cell death. These effects were accompanied by mitochondrial membrane depolarization, depletion of intracellular reduced thiols, and DNA fragmentation, indicating the involvement of oxidative stress and mitochondrial dysfunction in the observed cytotoxic response. Taken together, these results demonstrate that 10-substituted 1,6-diazaphenothiazines exert anti-melanoma activity through multiple biological mechanisms. We believe our study provides a basis for developing derivatives with optimized pharmacological properties.
Full article
(This article belongs to the Section Bioorganic Chemistry and Medicinal Chemistry)
►▼
Show Figures

Figure 1
Open AccessReview
Natural Products Targeting Acetylation in Bladder Cancer: Mechanistic Basis, Therapeutic Potential, and Future Perspectives
by
Wei Li, Da Liu, Qinzhamusu Yin, Yiwen Geng, Yang Liu and Yong Wang
Curr. Issues Mol. Biol. 2026, 48(5), 489; https://doi.org/10.3390/cimb48050489 (registering DOI) - 8 May 2026
Abstract
Bladder cancer remains a major clinical challenge because of its high recurrence rate, marked molecular heterogeneity, frequent progression, and limited durability of current therapeutic strategies. Increasing evidence indicates that acetylation, as a reversible and druggable epigenetic modification, plays a central role in bladder
[...] Read more.
Bladder cancer remains a major clinical challenge because of its high recurrence rate, marked molecular heterogeneity, frequent progression, and limited durability of current therapeutic strategies. Increasing evidence indicates that acetylation, as a reversible and druggable epigenetic modification, plays a central role in bladder cancer biology by linking chromatin remodeling to transcriptional regulation, DNA damage repair, metabolic adaptation, and immune modulation. Both histone and non-histone acetylation are frequently dysregulated in bladder cancer, and these alterations contribute to multiple malignant phenotypes, including sustained proliferation, defective cell-cycle control, apoptosis evasion, epithelial–mesenchymal transition, metastatic progression, and therapeutic resistance. In this review, we summarize the mechanistic basis of acetylation imbalance in bladder cancer, with particular emphasis on the roles of histone acetyltransferases, histone deacetylases, sirtuins, and acetylation-associated metabolic regulators. We further discuss the emerging evidence that natural products can modulate acetylation-related pathways in bladder cancer, mainly through targeting HDAC-dependent histone deacetylation and SIRT1-associated non-histone deacetylation. Representative compounds, including sulforaphane, erucin, puerarin, capsaicin, curcumin, trichostatin A, trichostatin C, and pinocembrin, highlight the potential of natural products to suppress tumor growth, promote apoptosis, impair migration, and enhance antitumor immunity through acetylation-related mechanisms. Beyond summarizing individual agents, the evidence was evaluated based on the integration of acetylation-related target engagement, acetylation remodeling, and bladder cancer-relevant phenotypic outcomes. The current evidence is heterogeneous. SFN/ECN, capsaicin, and pinocembrin offer the most convincing bladder cancer-specific support, whereas several other compounds remain limited by context-dependent effects, indirect pathway inference, or incomplete validation of the proposed acetylation mechanisms. These findings support an evidence-oriented translational framework that prioritizes natural products according to mechanistic robustness, bladder cancer specificity, and combination potential. Overall, acetylation-targeting natural products represent a promising but still evolving therapeutic strategy for bladder cancer, warranting further subtype-specific, mechanistically rigorous, and translationally oriented investigation.
Full article
(This article belongs to the Special Issue Natural Compounds: An Adjuvant Strategy in Cancer Management, 2nd Edition)
►▼
Show Figures
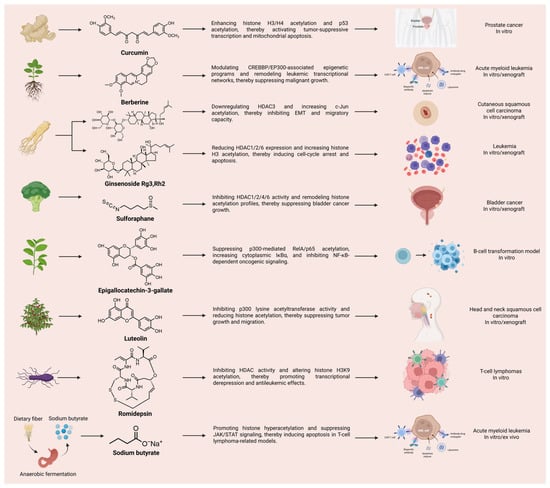
Figure 1
Open AccessReview
The Potential and Prospects of Hydrogel Applications in Traumatic Brain Injury Treatment
by
Cheng Zhong, Jie Li, Dengzhuo Liu, Xinran He, Zihao Fan, Xinxin Guo and Guangwei Wang
Curr. Issues Mol. Biol. 2026, 48(5), 488; https://doi.org/10.3390/cimb48050488 - 8 May 2026
Abstract
Traumatic brain injury (TBI) is a prevalent neurological disorder that induces severe neurological dysfunction and markedly reduces quality of life owing to its complex pathophysiology and limited therapeutic options. Conventional pharmacological and surgical interventions show restricted efficacy because of poor blood–brain barrier penetration
[...] Read more.
Traumatic brain injury (TBI) is a prevalent neurological disorder that induces severe neurological dysfunction and markedly reduces quality of life owing to its complex pathophysiology and limited therapeutic options. Conventional pharmacological and surgical interventions show restricted efficacy because of poor blood–brain barrier penetration and inability to address secondary injury cascades. In recent years, hydrogels have shown significant potential for TBI repair due to their superior biocompatibility, high water content, and ability to mimic the native extracellular matrix (ECM). This review systematically examines recent advances in hydrogel applications for TBI therapy, focusing on their roles as drug delivery platforms, stem cell scaffolds, neuroregeneration promoters, inflammation modulators, and angiogenesis facilitators. Particular emphasis is placed on the therapeutic benefits and underlying mechanisms of ECM-derived hydrogels, self-assembling peptide (SAP) hydrogels, stimuli-responsive smart hydrogels, and functionalized multicomponent systems. Current challenges and limitations in hydrogel applications are also discussed, along with future research directions, to provide scientific rationale and practical guidance for precision TBI therapy.
Full article
(This article belongs to the Special Issue The Contribution and Application of Molecular Biology in the Applied Biosciences—Focusing on Medicine, Biomaterials and Tissue Engineering Fields, 3rd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
Coix Seed Oil Ameliorates Rheumatoid Arthritis by Modulating Inflammation-Associated Metabolic Pathways
by
Yong Yang, Ying Feng, Weijie Tang, Yu Meng and Xiuping Ma
Curr. Issues Mol. Biol. 2026, 48(5), 487; https://doi.org/10.3390/cimb48050487 - 8 May 2026
Abstract
Rheumatoid arthritis (RA) is a chronic disease that primarily manifests as symmetrical joint inflammation. Although Coix Seed Oil (CSO) has demonstrated anti-inflammatory effects in RA rat models, its systemic metabolic regulatory mechanisms remain unclear. Therefore, we aimed to investigate whether CSO ameliorates RA
[...] Read more.
Rheumatoid arthritis (RA) is a chronic disease that primarily manifests as symmetrical joint inflammation. Although Coix Seed Oil (CSO) has demonstrated anti-inflammatory effects in RA rat models, its systemic metabolic regulatory mechanisms remain unclear. Therefore, we aimed to investigate whether CSO ameliorates RA by modulating inflammation-associated metabolic pathways. Ultra-High-Performance Liquid Chromatography (UHPLC)-Q Exactive HF-X-MS-based metabolomics was used to profile metabolites in the synovial tissue and serum of complete Freund’s adjuvant (CFA)-induced RA rats. Systematically altered metabolites and their associated pathways were identified using multivariate analysis and pattern recognition. CSO treatment modulated 16 RA-related biomarkers in rat synovial tissues and 12 in the serum, which mainly affected amino acids, arachidonic acids, lipids, sphingolipids, and carnitines. These metabolites were associated with eight perturbed metabolic pathways that were predominantly involved in inflammatory responses. This study demonstrated that CSO has significant anti-RA effects on pharmacodynamic activity and metabolic network regulation. Additionally, inflammation-associated metabolic pathways are closely linked to the therapeutic efficacy of CSO in RA treatment.
Full article
(This article belongs to the Section Bioinformatics and Systems Biology)
►▼
Show Figures

Figure 1
Open AccessArticle
NEK1 Promotes Ovarian Cancer Progression via p53 Suppression While Enhancing Sensitivity to Genotoxic Therapy
by
Huiyang Song, Xia Wang, Aiqing Yang, Xuejiao Ren, Xiaoqi Zhou, Yifei Qiu, Yating Cai, Chengming Gao, Gangqiao Zhou and Pengbo Cao
Curr. Issues Mol. Biol. 2026, 48(5), 486; https://doi.org/10.3390/cimb48050486 - 7 May 2026
Abstract
Ovarian cancer (OV) is a highly metastatic and recurrent malignancy with limited therapeutic options. NIMA-related kinase 1 (NEK1), a serine/threonine kinase implicated in cell cycle regulation and DNA damage response, has been associated with tumorigenesis in various cancers, yet its specific role in
[...] Read more.
Ovarian cancer (OV) is a highly metastatic and recurrent malignancy with limited therapeutic options. NIMA-related kinase 1 (NEK1), a serine/threonine kinase implicated in cell cycle regulation and DNA damage response, has been associated with tumorigenesis in various cancers, yet its specific role in OV pathogenesis remains elusive. This study systematically investigates the oncogenic function and underlying mechanisms of NEK1 in ovarian cancer. Our findings demonstrate that NEK1 promotes tumor progression both in vitro and in vivo. Mechanistically, bioinformatic and biochemical analyses reveal that NEK1 suppresses p53 signaling activity, resulting in downregulation of downstream targets p21 and PUMA, consequently attenuating cell cycle arrest and apoptosis. Importantly, NEK1-driven oncogenicity is dependent on the presence of p53 protein. Clinically, elevated NEK1 expression significantly correlates with poorer prognosis across multiple independent OV cohorts. Paradoxically, high NEK1 expression enhances radiosensitivity by impairing p53-mediated DNA damage repair. Collectively, these findings establish NEK1 as a promising prognostic biomarker and therapeutic target, with potential utility in guiding genotoxic therapy strategies for ovarian cancer patients.
Full article
(This article belongs to the Section Molecular Medicine)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- CIMB Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal Browser-
arrow_forward_ios
Forthcoming issue
arrow_forward_ios Current issue - Volumes not published by MDPI
- Vol. 42 (2021)
- Vol. 41 (2021)
- Vol. 40 (2021)
- Vol. 39 (2020)
- Vol. 38 (2020)
- Vol. 37 (2020)
- Vol. 36 (2020)
- Vol. 35 (2020)
- Vol. 34 (2019)
- Vol. 33 (2019)
- Vol. 32 (2019)
- Vol. 31 (2019)
- Vol. 30 (2019)
- Vol. 29 (2018)
- Vol. 28 (2018)
- Vol. 27 (2018)
- Vol. 26 (2018)
- Vol. 25 (2018)
- Vol. 24 (2017)
- Vol. 23 (2017)
- Vol. 22 (2017)
- Vol. 21 (2017)
- Vol. 20 (2016)
- Vol. 19 (2016)
- Vol. 18 (2016)
- Vol. 17 (2015)
- Vol. 16 (2014)
- Vol. 15 (2013)
- Vol. 14 (2012)
- Vol. 13 (2011)
- Vol. 12 (2010)
- Vol. 11 (2009)
- Vol. 10 (2008)
- Vol. 9 (2007)
- Vol. 8 (2006)
- Vol. 7 (2005)
- Vol. 6 (2004)
- Vol. 5 (2003)
- Vol. 4 (2002)
- Vol. 3 (2001)
- Vol. 2 (2000)
- Vol. 1 (1999)
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
CIMB, IJMS, Reprod. Med., Biology, Life
Recent Research in Germ Cells
Topic Editors: Malgorzata Kloc, Jacek KubiakDeadline: 31 May 2026
Topic in
CIMB, Molecules, Pharmaceuticals, Pharmaceutics, Sci. Pharm.
Challenges and Opportunities in Drug Delivery Research, 2nd Edition
Topic Editors: Lenuta Profire, Ioana Mirela VasincuDeadline: 30 June 2026
Topic in
Cancers, CIMB, Current Oncology, Sci. Pharm., Antibodies, IJMS, IJTM
Antibody-Mediated Therapy and Other Emerging Therapies in Cancer Treatment
Topic Editors: Won Sup Lee, Yaewon Yang, Seil GoDeadline: 31 July 2026
Topic in
IJMS, Agronomy, Plants, Crops, CIMB
New Achievements in Gene Mining, Germplasm Innovation, Cultivation Technologies in Rice
Topic Editors: Shimin Zuo, Yongmei Bao, Yajie HuDeadline: 31 August 2026

Conferences
Special Issues
Special Issue in
CIMB
Genetics and Epigenetics of Neurodegenerative Diseases
Guest Editor: Salvatore SacconeDeadline: 15 May 2026
Special Issue in
CIMB
Biomaterials for Disease Diagnosis and Therapy: Emerging Molecular Innovations and Applications
Guest Editor: Md Tipu SultanDeadline: 20 May 2026
Special Issue in
CIMB
Natural Phytochemicals as Modulators of Cellular Pathways and Therapeutic Targets in Disease Models
Guest Editor: Jui-Hung YenDeadline: 20 May 2026
Special Issue in
CIMB
Molecular Mechanisms and Pharmacological Underlying Cardiorenal Diseases
Guest Editor: Caterina CarolloDeadline: 31 May 2026
Topical Collections
Topical Collection in
CIMB
Feature Papers in Current Issues in Molecular BiologyCollection Editor: Madhav Bhatia
Topical Collection in
CIMB
Feature Papers Collection in Molecular Microbiology
Collection Editor: Bruce Seal
Topical Collection in
CIMB
Bioinformatics Approaches to Biomedicine
Collection Editor: Giulia Fiscon
Topical Collection in
CIMB
Molecular Advances in Veterinary Pharmacology and Toxicology
Collection Editor: Chongshan Dai