Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q1 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 20 days after submission; acceptance to publication is undertaken in 2.9 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about Microorganisms.
- Companion journal for Microorganisms include: Applied Microbiology and Bacteria.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
4.2 (2024);
5-Year Impact Factor:
4.6 (2024)
Latest Articles
Infectious Spondylodiscitis of Bacterial Causes in Adults: Epidemiology, Pathophysiology, Diagnostic and Treatment Challenges
Microorganisms 2026, 14(5), 1110; https://doi.org/10.3390/microorganisms14051110 - 13 May 2026
Abstract
Spinal infections in general, and infectious spondylodiscitis in particular, are increasingly diagnosed in the Western world, in recent decades. This rise in incidence is associated with an ageing population and with an increased availability of accurate diagnostic modalities. Even so, due to the
[...] Read more.
Spinal infections in general, and infectious spondylodiscitis in particular, are increasingly diagnosed in the Western world, in recent decades. This rise in incidence is associated with an ageing population and with an increased availability of accurate diagnostic modalities. Even so, due to the non-specific nature of clinical manifestations, and of the implicated blood and serum markers, there is a risk of underdiagnosis or misdiagnosis of the disease in its initial stages. Ionizing radiation methods, such as plain radiography (X-ray) and computed tomography (CT), are also not reliable in the early stages of the diseases, and the golden standard of imagistic diagnosis, magnetic resonance imaging (MRI), is not always available or requested. Still, MRI remains the most reliable method in most cases where there is a need for differential diagnosis with other pathologies, namely Andersson lesions, destructive spondyloarthropathy, erosive osteochondritis, micro-crystalline spondylitis, Modic 1 lesion, Charcot spinal arthropathy, osteoporotic fractures, SAPHO syndrome with spinal involvement, and Schmorl’s nodes. Infectious spondylodiscitis is caused by bacteria, and, less frequently, by fungi. Rare cases of parasitic causes have also been reported in the literature. Infectious spondylodiscitis of bacterial causes may be pyogenic, more frequently caused by Staphylococcus spp. or Streptococcus spp., or granulomatous, usually caused by Mycobacterium tuberculosis complex (MTBC) or from classical brucellosis. In all these cases, therapy may be conservative, with antibiotics, or surgical, when the former fails or in patients with significant spinal instability or other neurological manifestations. There are various surgical approaches, each with its own drawbacks, and usually used according to the preference of the attending physician. Even in cases of surgical treatment, antibiotic administration is prolonged, and it is important for a proper scheme to be selected based on antimicrobial susceptibility testing. However, given that in many cases, the causative agent cannot be identified, empirical treatment must be initiated. Finally, newer approaches, including the incorporation of antimicrobial substances, may offer better solutions for improving treatment and rehabilitation outcomes.
Full article
(This article belongs to the Special Issue The Epidemiology, Clinical Aspects, Treatment and Preventive Strategies for Infectious Diseases in the Era of Antimicrobial Resistance (AMR))
Open AccessReview
The Silent Spillover Threat: Nipah Virus Epidemiology, Pathogenesis, Clinical Manifestations, and Advances in Therapeutics and Vaccine Development
by
Elli-Panagiota Magklara, Maria Kkirgia, Andreas G. Tsantes, Petros Ioannou, Alexandra Mpakosi, Vasiliki Mougiou, Zoi Iliodromiti, Theodora Boutsikou, Nicoletta Iacovidou and Rozeta Sokou
Microorganisms 2026, 14(5), 1109; https://doi.org/10.3390/microorganisms14051109 - 13 May 2026
Abstract
Nipah virus (NiV) is an animal-borne RNA virus of the genus Henipavirus that poses a significant global health threat. This threat is driven by the virus’s high mortality rate, its capacity to cause epidemics, and the lack of licensed therapeutic interventions or vaccines.
[...] Read more.
Nipah virus (NiV) is an animal-borne RNA virus of the genus Henipavirus that poses a significant global health threat. This threat is driven by the virus’s high mortality rate, its capacity to cause epidemics, and the lack of licensed therapeutic interventions or vaccines. Since its initial identification during the 1998–1999 outbreak in Malaysia and Singapore, recurrent episodes have occurred primarily in Bangladesh and India, with mortality rates frequently exceeding 70%. Fruit bats of the genus Pteropus serve as the biological host for the virus. Transmission to humans occurs via contact with infected wildlife, consumption of contaminated products, such as freshly harvested date palm sap, or direct person-to-person exposure. Other modes of transmission, such as transplacentally or via breast milk, are still under investigation. The clinical presentation of NiV infection varies widely, from mild flu-like symptoms to life-threatening respiratory disease and acute encephalitis. It frequently attacks the nervous system, which can lead to coma, permanent neurological damage, or relapsing encephalitis. The virus enters host cells via ephrin-B2/B3 receptors, enabling systemic dissemination and infiltration of the central nervous system. Diagnosis relies primarily on RT-PCR and serological assays, and virus isolation requires high-containment laboratories. Management remains largely supportive, as no approved antiviral therapy exists. Experimental agents, such as remdesivir, favipiravir, and monoclonal antibodies such as m102.4, have shown promise in preclinical studies. Multiple vaccine platforms—including subunit, viral vector, mRNA, and nanoparticle-based approaches—are under development, though none is yet licensed for human use. Strengthened surveillance, infection control measures, and continued research are essential to mitigate the threat posed by this emerging pathogen. This review summarizes current knowledge on NiV, including its virology, epidemiology, pathogenesis, transmission, and recent progress in therapeutic and vaccine development.
Full article
(This article belongs to the Section Virology)
Open AccessArticle
Fluoroquinolone-Induced Metabolic Dysregulation and Oxidative Stress Orchestrate Bacterial Demise
by
Caiyuan Zhou, Jing Sun, Yihan Luo, Fang Wang, Luqi Li, Tong Wu, Peng Xie, Chenxi Liu, Yibin Hu, Leilei Sun and Chengbao Wang
Microorganisms 2026, 14(5), 1108; https://doi.org/10.3390/microorganisms14051108 - 13 May 2026
Abstract
The bactericidal mechanisms of fluoroquinolones extend beyond their canonical inhibition of DNA topoisomerases, yet the associated metabolic perturbations remain incompletely understood. In this study, we systematically investigated the metabolic responses of Escherichia coli to three representative FQs—ofloxacin, enrofloxacin, and ciprofloxacin—using untargeted UPLC–Q Exactive
[...] Read more.
The bactericidal mechanisms of fluoroquinolones extend beyond their canonical inhibition of DNA topoisomerases, yet the associated metabolic perturbations remain incompletely understood. In this study, we systematically investigated the metabolic responses of Escherichia coli to three representative FQs—ofloxacin, enrofloxacin, and ciprofloxacin—using untargeted UPLC–Q Exactive Orbitrap–MS-based metabolomics. Bacterial cells were exposed to bactericidal concentrations (2 × MIC) for a single-time point (1 h), followed by comprehensive metabolomic profiling with six biological replicates per group. Our findings demonstrate that FQ-induced metabolic reprogramming serves as a primary driver of oxidative stress and nucleic acid damage, rather than a mere secondary effect. All three FQs induced substantial metabolic reprogramming characterized by disruptions in nucleotide biosynthesis, central carbon metabolism, and redox-related pathways, with notable drug-specific differences. Ciprofloxacin exhibited the most pronounced suppression of energy metabolism and antioxidant systems, whereas ofloxacin and enrofloxacin showed partial compensatory metabolic responses. Consistently, intracellular ROS levels were significantly elevated in all treatment groups, and this effect was attenuated by antioxidant supplementation. Furthermore, increased accumulation of 8-hydroxydeoxyguanosine and 8-hydroxyguanosine confirmed the occurrence of oxidative DNA and RNA damage. Collectively, these findings indicate that FQs induce distinct metabolic perturbations that are closely associated with oxidative stress and nucleic acid damage, providing a metabolic perspective on their bactericidal activity and suggesting potential targets for metabolic adjuvant strategies.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►▼
Show Figures
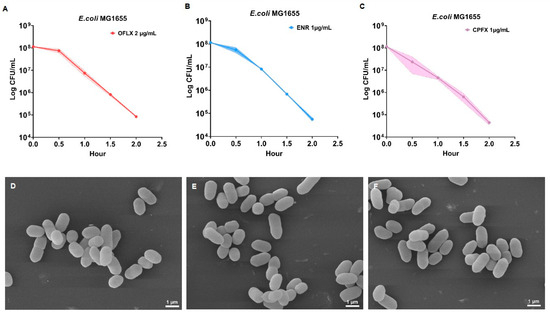
Figure 1
Open AccessArticle
Mechanism and Control of Black Spot Deterioration on Lacquered Architectural Components of Dajue Temple
by
Sifan Ai, Yu Wang, Jiao Pan, Gang Hu and Ruiting Zhao
Microorganisms 2026, 14(5), 1107; https://doi.org/10.3390/microorganisms14051107 - 13 May 2026
Abstract
Dajue Temple, a representative ancient architectural heritage in North China, houses numerous lacquered wooden components of exceptional historical and artistic value. Prolonged environmental exposure causes severe dark discoloration and black spotting on lacquer surfaces, threatening their structural integrity. This first investigation into the
[...] Read more.
Dajue Temple, a representative ancient architectural heritage in North China, houses numerous lacquered wooden components of exceptional historical and artistic value. Prolonged environmental exposure causes severe dark discoloration and black spotting on lacquer surfaces, threatening their structural integrity. This first investigation into the damage identifies the spots as microbial in origin, with Cladosporium spp. as the primary agent driving deterioration and possessing wood-degrading capabilities. Antifungal tests show that thymol, clove essential oil, and nano-silver gel are all effective inhibitors. We proposed targeted, relic-friendly microbial control strategies tailored for ancient lacquered wooden components. These findings provided scientific guidance for the sustainable conservation and restoration of lacquered architectural elements in historic temples and comparable cultural heritage sites. In future work, environmental monitoring and the biocides’ compatibility should be involved, which will help to clarify microbe–environment interactions, enable early warning of biodeterioration risks and explore the wood-friendly biocides.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures
Figure 1
Open AccessArticle
Scalp Microbiota Dysbiosis in Seborrheic Alopecia and Restoration Following Herbal Extract Shampoo Intervention
by
Jing Feng, Jiancong Huang, Shaolu Zhou, Xianghai Chen, Gang Zhou, Xudong Wang, Xia Wen, Qingshan Shi, Pianjuan Guo, Qiongfei Li and Xiaobao Xie
Microorganisms 2026, 14(5), 1106; https://doi.org/10.3390/microorganisms14051106 - 13 May 2026
Abstract
Seborrheic alopecia (SA) is one of the most common forms of hair loss with a complex pathogenesis involving multiple etiological factors. Although the scalp microbiome has been implicated in various scalp disorders, its specific role in the development and progression of SA remains
[...] Read more.
Seborrheic alopecia (SA) is one of the most common forms of hair loss with a complex pathogenesis involving multiple etiological factors. Although the scalp microbiome has been implicated in various scalp disorders, its specific role in the development and progression of SA remains incompletely understood. To characterize the scalp microbiome in SA, we performed high-throughput sequencing of the 16S rRNA gene and ITS region on scalp samples from 41 Chinese SA participants before and after a 12-week intervention with a shampoo containing herbal extracts (ginger root, Polygonum multiflorum, and Platycladus orientalis leaf) and 29 healthy controls. The untreated SA group exhibited significant microbial dysbiosis compared to the healthy controls, characterized by reduced bacterial and fungal alpha diversity and increased relative abundances of Staphylococcus, Cutibacterium, and Malassezia. LEfSe analysis confirmed the significant enrichment of these three genera. Correlation network analysis revealed a substantial restructuring of microbial interactions in the untreated SA group: Staphylococcus and Malassezia lost all positive correlations with other genera, whereas Cutibacterium displayed relatively stable topological relationships. Following the 12-week intervention, the treated SA group showed significant clinical improvement (reduced hair loss and scalp sebum content), along with a restoration of microbial diversity to levels comparable to the healthy group and a normalization of the abundances of Staphylococcus and Malassezia. Our study confirms the critical role of scalp microecological dysbiosis in SA pathogenesis and identifies Staphylococcus and Malassezia as key taxa strongly associated with this dysbiosis. These findings provide a theoretical foundation for developing microbiome-targeted strategies for SA treatment and support the use of multi-targeted, plant-based interventions to restore microbial homeostasis and promote hair growth.
Full article
(This article belongs to the Section Microbiomes)
►▼
Show Figures
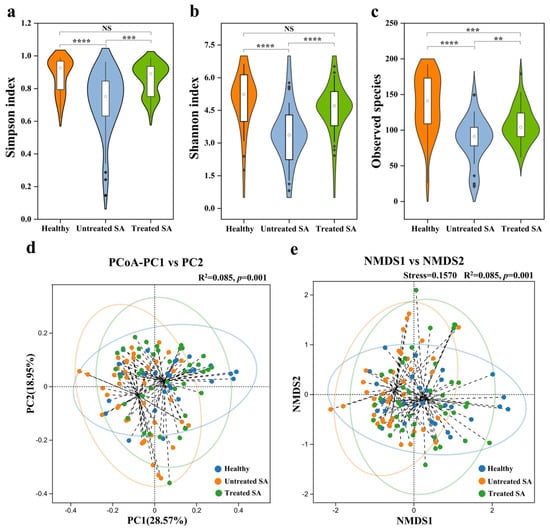
Figure 1
Open AccessArticle
Early-Life Iron Exposure Influences Long-Term Gut Microbiota Recovery After Intestinal Dysbiosis
by
Thibault Maumy, Claire McCartney, Ayodeji Samuel Ajayi, Claire Gerkins, Gabriela Fragoso, Annie Calvé and Manuela M. Santos
Microorganisms 2026, 14(5), 1105; https://doi.org/10.3390/microorganisms14051105 - 13 May 2026
Abstract
Host–microbiota interactions during the neonatal window of opportunity have gained significant interest as factors influencing long-term health. Factors such as nutrient availability may shape the microbial community, and iron is no exception to this rule. Although the use of iron supplementation is widespread
[...] Read more.
Host–microbiota interactions during the neonatal window of opportunity have gained significant interest as factors influencing long-term health. Factors such as nutrient availability may shape the microbial community, and iron is no exception to this rule. Although the use of iron supplementation is widespread during infancy, this micronutrient is known to have profound effects on gut microbiota. This study aimed to determine how early-life iron supplementation shapes gut microbiota composition and whether it influences recovery from gut dysbiosis later in life. Three-week-old female C57BL/6 mice were fed an iron-excess diet for five weeks during the critical period of microbiota establishment. After a two-week washout period to normalize luminal iron content, dysbiosis was induced using either dextran sulfate sodium-induced acute colitis or antibiotic treatment. Mice were then allowed an 8-week recovery period. Gut microbiota composition was longitudinally analyzed by 16S rRNA gene sequencing of fecal samples. Early-life iron supplementation induced durable alterations in gut microbiota composition, with differences persisting even after luminal iron normalization (β-diversity, PERMANOVA p < 0.01). At the endpoint, mice exposed to an iron-sufficient diet remained significantly more distant from their baseline compared to the excess iron group in both the dextran sulfate sodium (33% greater distance) and antibiotic (41% greater distance) models (both p < 0.05). Notably, this pattern was not observed when supplementation occurred in adulthood. In the dextran sulfate sodium model, mice that received an iron-sufficient diet during early life showed an expansion of the probiotic Ligilactobacillus murinus, which positively correlated with fecal succinate levels. Conversely, in the antibiotic-induced model, early-life exposure to an iron-sufficient diet was associated with a more pronounced dysbiosis characterized by a nearly two-fold-greater loss in α-diversity compared to 500 ppm mice (∆Shannon: 1.98 ± 0.22 vs. 1.02 ± 0.25, p < 0.01). These findings suggest that early-life iron supplementation influences long-term host–microbiota interactions and recovery from gut dysbiosis.
Full article
(This article belongs to the Special Issue Effects of Diet and Nutrition on Gut Microbiota)
►▼
Show Figures
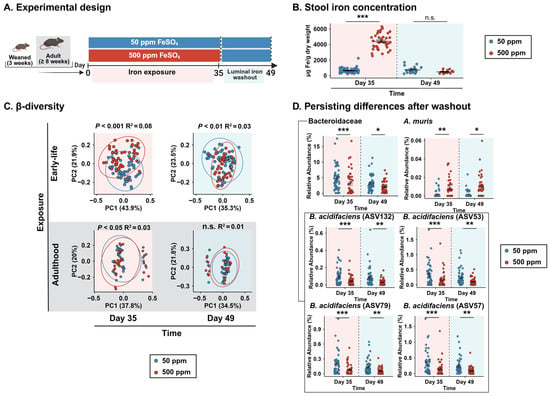
Figure 1
Open AccessReview
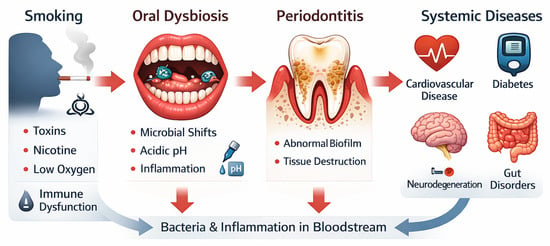
Tobacco-Induced Oral Dysbiosis and Microbial Shifts: A Narrative Review of Their Role in Systemic Inflammation and Disease
by
Glenda M. Davison, Tandi Matsha, Shanel Raghubeer, Stanton Hector, Saarah Davids and Yvonne Prince
Microorganisms 2026, 14(5), 1104; https://doi.org/10.3390/microorganisms14051104 - 13 May 2026
Abstract
The oral cavity is home to a diverse community of microbiota comprising bacteria, viruses, protozoa, and fungi. These microorganisms inhabit several oral niches and play a significant role in supporting both oral and systemic health. The fine balance between the microbial communities can
[...] Read more.
The oral cavity is home to a diverse community of microbiota comprising bacteria, viruses, protozoa, and fungi. These microorganisms inhabit several oral niches and play a significant role in supporting both oral and systemic health. The fine balance between the microbial communities can be influenced by genetics and environmental factors, potentially leading to dysbiosis. Alterations in the oral microbiota have been implicated in periodontitis, chronic inflammation, and systemic disease. Tobacco has been identified as a major player in altering the oral microenvironment and disturbing the balance between potentially pathogenic and beneficial commensals. The resulting dysbiosis promotes inflammation and assists in the passage of pathogenic microorganisms into the blood system. This narrative review examines current evidence linking the use of tobacco with the dominance of pathogenic oral bacteria and a dysfunctional immune response. We explore how the chemicals and toxins in cigarettes promote a reduction in oxygen and cause changes in the abundance of anaerobic bacteria. After discussing the mechanistic pathways leading to periodontitis and the entry of microorganisms into the circulation, the review will interrogate previous studies and identify opportunities and priorities for future research.
Full article
(This article belongs to the Special Issue Microbiomes in Human Health and Diseases)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Isolation, Identification, and Functional Characterization of a Rhizosphere Bacterium Promoting the Growth of Alsophila spinulosa
by
Jiya Wu, Weicheng Yang, Xiaona Zhang, Xianyu Li, Bibo Zhou, Tianyu Liang and Fen Liu
Microorganisms 2026, 14(5), 1103; https://doi.org/10.3390/microorganisms14051103 - 13 May 2026
Abstract
Alsophila spinulosa is a tree fern designated as a second-class nationally protected species in China and valued for its medicinal and ornamental properties. Its slow growth and susceptibility to environmental stresses pose challenges to its cultivation. Plant-growth-promoting rhizobacteria (PGPR) can enhance plant development
[...] Read more.
Alsophila spinulosa is a tree fern designated as a second-class nationally protected species in China and valued for its medicinal and ornamental properties. Its slow growth and susceptibility to environmental stresses pose challenges to its cultivation. Plant-growth-promoting rhizobacteria (PGPR) can enhance plant development by producing phytohormones, such as indole-3-acetic acid (IAA). In this study, 39 IAA-producing strains were isolated from the rhizosphere of A. spinulosa. Morphological and molecular analyses identified the highest IAA-producing strain, R74, as Burkholderia pyrrocinia. Its optimal inoculum age was determined to be 12–20 h, and its optimal culture conditions for IAA production were 24 h of incubation, 32 °C and pH 7.0. Whole-genome sequencing revealed that the genome of strain R74 is 8,347,169 bp in size with a GC content of 67%, comprising 7543 genetic elements. Further genomic analysis showed that IAA biosynthesis in R74 involves the tryptophan side-chain oxidase (TSO) pathway and the tryptophan-independent pathway. Pot experiments revealed that inoculation with R74 increased the height, root length, stem diameter, and biomass of A. spinulosa seedlings. It also increased antioxidant enzyme activities, elevated soluble protein and chlorophyll contents, and reduced malondialdehyde levels. This study provides an empirical basis for the development of Burkholderia-based biofertilizers to promote A. spinulosa growth.
Full article
(This article belongs to the Section Plant Microbe Interactions)
►▼
Show Figures
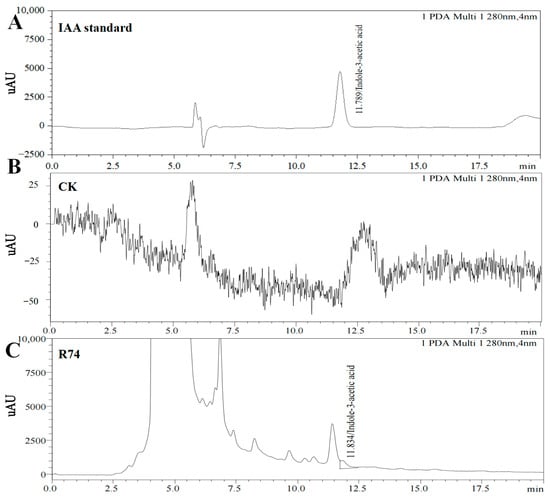
Figure 1
Open AccessArticle
Mutation-Tolerant Inhibition of HIV-1 Integrase Strand Transfer by Secondary Metabolites from the Endophytic Fungus Alternaria alternata PO4PR2
by
Ndzalo Mashabela, Darian Naidu, Ernest Oduro-Kwateng and Nompumelelo P. Mkhwanazi
Microorganisms 2026, 14(5), 1102; https://doi.org/10.3390/microorganisms14051102 - 13 May 2026
Abstract
Endophytic fungi are promising sources of novel antiviral compounds, and the crude extract from Alternaria alternata PO4PR2 has previously shown anti-HIV-1 activity. This study evaluated its efficacy against integrase strand-transfer inhibitor (INSTI)-resistant HIV-1 and its mechanism of action. Key resistance mutations (Y143H, G118R,
[...] Read more.
Endophytic fungi are promising sources of novel antiviral compounds, and the crude extract from Alternaria alternata PO4PR2 has previously shown anti-HIV-1 activity. This study evaluated its efficacy against integrase strand-transfer inhibitor (INSTI)-resistant HIV-1 and its mechanism of action. Key resistance mutations (Y143H, G118R, N155H, and R263K) were introduced into the HIV-1 pNL4.3 clone via site-directed mutagenesis and confirmed through Sanger sequencing. Viral infectivity was assessed in TZM-bl cells, while cytotoxicity was measured using an MTT assay. Antiviral activity was determined through a luciferase-based assay, and integration inhibition was evaluated using integrase activity assays and Alu-gag nested PCR. The extract demonstrated potent inhibition of resistant mutants, with low IC50 values (0.02971–0.1652 μg/mL), and showed minimal cytotoxicity (CC50 = 300 μg/mL), maintaining over 80% cell viability. It inhibited integrase activity by 67%, specifically targeting the strand-transfer step, and significantly reduced integrated viral DNA. Molecular docking of 14 compounds identified coumarin derivatives as key bioactive metabolites, exhibiting mutation-tolerant binding within the integrase catalytic pocket. Overall, these findings highlight PO4PR2 as a promising source of compounds for developing new therapies targeting drug-resistant HIV-1 integrase.
Full article
(This article belongs to the Section Virology)
►▼
Show Figures
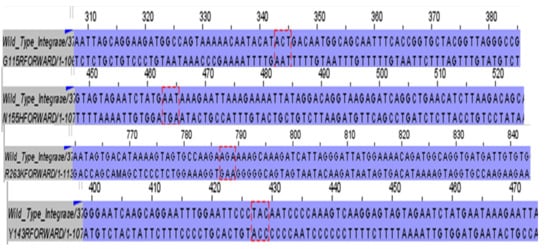
Figure 1
Open AccessArticle
From the Carp Gut to Plastic Solutions: Hafnia Strain from Cyprinus carpio Demonstrates Robust Degradation of Synthetic Polymers
by
Mina Popovic, Boris Rajicic and Neveka Rajic
Microorganisms 2026, 14(5), 1101; https://doi.org/10.3390/microorganisms14051101 - 13 May 2026
Abstract
The accumulation of polyethylene (PE) in aquatic ecosystems represents a significant environmental challenge due to the polymer’s high molecular weight and chemical stability. This study investigates the biodegradation potential of Hafnia paralvei UUNT_MP29, a bacterial strain isolated from the gut of common carp
[...] Read more.
The accumulation of polyethylene (PE) in aquatic ecosystems represents a significant environmental challenge due to the polymer’s high molecular weight and chemical stability. This study investigates the biodegradation potential of Hafnia paralvei UUNT_MP29, a bacterial strain isolated from the gut of common carp (Cyprinus carpio), for low-density polyethylene (LDPE). Initial screening on LDPE-emulsified agar confirmed extracellular enzymatic activity through the formation of distinct clear zones. Quantitative analysis showed a cumulative mass loss of 24.10% by Day 16, with the most intensive degradation occurring between Days 4 and 8, which closely correlated with maximum bacterial count (CFU/mL). Kinetic modeling indicated that the degradation followed a first-order rate law (R2 = 0.9269), with a rate constant (k) of 0.2991 days−1 and a remarkably short half-life (t1/2) of 2.32 days. Structural characterization via FTIR spectroscopy demonstrated oxidative transformation, evidenced by a reduction in sp3 C-H stretching and the emergence of C-O/C-O-C functional groups. SEM micrographs further confirmed extensive bio-deterioration, including surface pitting and macroscale erosion. Thermal analysis (TGA/DTG) supported these findings, showing a significant 10.95 °C decrease in the maximum degradation temperature (Tmax), indicating a reduction in polymer chain length. These results suggest that H. paralvei UUNT_MP29 is a highly efficient agent for the rapid breakdown of polyethylene and highlight the potential of aquatic gut microbiota as reservoirs for plastic-degrading biotechnologies.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures
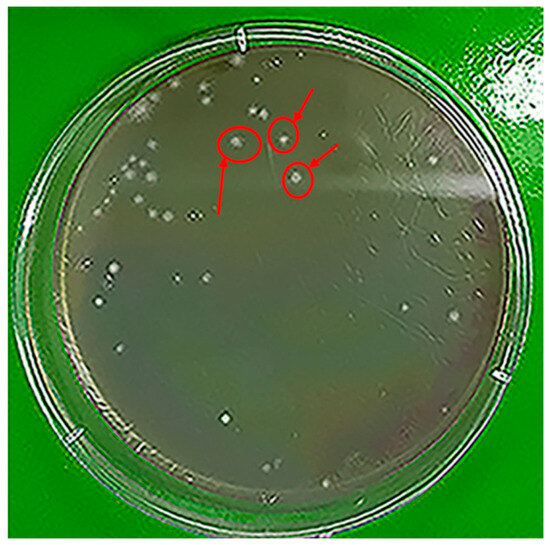
Figure 1
Open AccessEditorial
Editorial for the Special Issue Research on Biology of Dinoflagellates
by
Tania Islas-Flores, Estefanía Morales-Ruiz and Marco A. Villanueva
Microorganisms 2026, 14(5), 1100; https://doi.org/10.3390/microorganisms14051100 - 13 May 2026
Abstract
Protists arose about 1 [...]
Full article
(This article belongs to the Special Issue Research on Biology of Dinoflagellates)
Open AccessArticle
Effect of Aeration Rate Redistribution on Nitrogen Removal Performance of a Novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System
by
Zichun Yan, Shuichao Fan, Wankai Yan, Haopeng Ma and Tianhao Zhao
Microorganisms 2026, 14(5), 1099; https://doi.org/10.3390/microorganisms14051099 - 13 May 2026
Abstract
To address the problems of short-circuit flow and dead zones, complicated operation and control caused by intermittent influent, and the mismatch between aeration rate and oxygen demand in the Cyclic Activated Sludge System (CASS), a novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System (MCFCASS)
[...] Read more.
To address the problems of short-circuit flow and dead zones, complicated operation and control caused by intermittent influent, and the mismatch between aeration rate and oxygen demand in the Cyclic Activated Sludge System (CASS), a novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System (MCFCASS) was developed. This system operated in continuous-flow mode, and the aeration rate of each compartment was redistributed using a mathematical model. The results show that the plug flow ratio of the MCFCASS reactor increased from 18.75% to 31.25% compared with the CASS reactor. After aeration rate redistribution, the average total nitrogen (TN) removal efficiency of the MCFCASS reactor rose from 83.34% to 86.80%, and the effluent TN concentration consistently met the Grade I-A limit (15 mg/L) specified in the Discharge Standard of Pollutants for Municipal Wastewater Treatment Plant (GB 18918-2002). The average removal efficiencies of chemical oxygen demand (COD) and ammonium nitrogen (NH4+-N) increased from 91.58% and 93.39% to 92.98% and 94.57%, respectively. Microbial community analysis revealed that after aeration rate redistribution, the relative abundances of Pseudomonadota, Bacteroidota, and Bacillota in the pre-reaction zone of MCFCASS were 39.17%, 17.78%, and 10.33%, respectively. In addition, the abundances of some functional bacterial groups in the first and fourth compartments of the main reaction zone shifted adaptively in response to the aeration rate redistribution, consistent with the trends in pollutant removal contributions in these compartments. Hierarchical clustering and principal coordinate analysis (PCoA) further indicated that aeration rate redistribution influenced the microbial community structure. The above laboratory-scale optimization results may provide a preliminary reference for aeration control and improvement of denitrification performance in similar processes.
Full article
(This article belongs to the Collection Feature Papers in Environmental Microbiology)
►▼
Show Figures
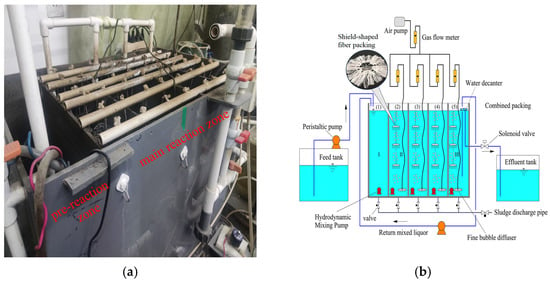
Figure 1
Open AccessArticle
Lectin-Based Antiviral Strategies for Porcine Reproductive and Respiratory Syndrome Virus 2 Infection: Griffithsin Suppresses Viral Replication In Vitro and Reduces Early Viremia In Vivo
by
Darshana Kadekar, Deepak Velayudhan, Ester Vinyeta, Jianqiang Zhang, Ethan Aljets, Veeraya Bamrung, Panchan Sitthicharoenchai, Alyona Michael, Keith Frogue, Meng Heng, Amy Liu, Cristina Bongiorni, Manasi Bhate, David A. Estell, Chong Shen and Charlotte Poulsen
Microorganisms 2026, 14(5), 1098; https://doi.org/10.3390/microorganisms14051098 - 12 May 2026
Abstract
Porcine reproductive and respiratory syndrome virus (PRRSV) remains a major challenge to swine production worldwide. Current vaccines have limited efficacy against genetically diverse PRRSV strains. Therefore, strategies with alternative modes of action—such as antiviral approaches that target conserved virus–host interactions, including viral attachment
[...] Read more.
Porcine reproductive and respiratory syndrome virus (PRRSV) remains a major challenge to swine production worldwide. Current vaccines have limited efficacy against genetically diverse PRRSV strains. Therefore, strategies with alternative modes of action—such as antiviral approaches that target conserved virus–host interactions, including viral attachment and entry, rather than relying solely on adaptive immune responses—are needed. We first evaluated the in vitro effect of griffithsin (GRFT), a high-mannose-binding lectin, in the monkey kidney cell line MARC-145. Cells were pre-treated with GRFT (50–200 µg/mL) prior to PRRSV infection, after which cell morphology and viral RNA replication (measured by RT-qPCR) were assessed. Pre-treatment with 100–200 µg/mL GRFT, followed by PRRSV inoculation at a multiplicity of infection of 1 or 10, reduced viral replication in MARC145 cells in a dose-dependent manner, achieving almost 100% inhibition of ORF5 and ORF7 RNA compared with untreated controls (p < 0.0001). We next investigated the in vivo effects of intranasal GRFT administration (7.5 or 15 mg/day) in pigs (n = 56). Pigs treated with 15 mg/day GRFT exhibited significantly reduced (p < 0.05) viremia 2, 4 and 7 days post-challenge, compared with untreated, challenged, and controls (log10 8.1 ± 0.2 vs. 9.0 ± 0.25, 8.2 ± 0.1 vs. 9.1 ± 0.2, and 8.9 ± 0.2 vs. 9.3 ± 0.2, respectively), along with earlier resolution of fever and a trend toward increased average daily gain over 42 days (p < 0.1). These findings are the first report of GRFT efficacy in pigs and support its potential as an antiviral strategy against PRRSV, alongside existing interventions.
Full article
(This article belongs to the Special Issue The Pathogenesis, Epidemiology and Diagnosis of Infectious Diseases in Livestock Animals, 2nd Edition)
►▼
Show Figures
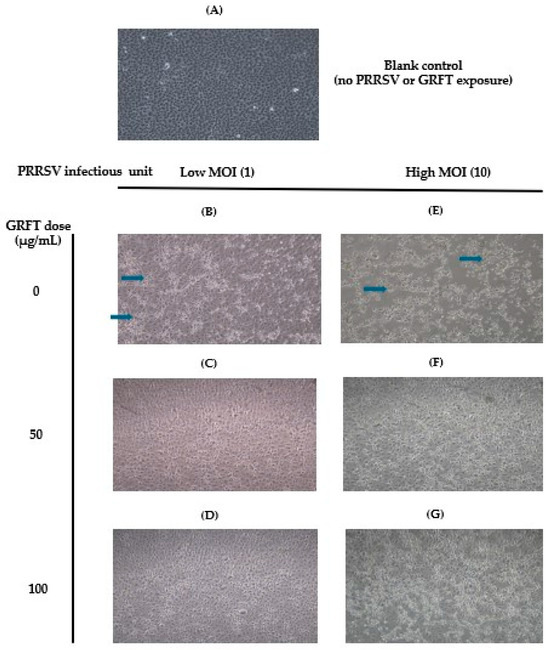

Figure 1
Open AccessArticle
Whole-Genome Sequencing and Genomic Characterization of a Multi-Drug Resistant Phenotype of Listeria monocytogenes Isolated from Pet Food
by
Antonia Mataragka, Marios Mataragas, Nikolaos Tzimotoudis, Ioannis Galiatsatos, Panagiota Stathopoulou, Spiros Paramithiotis, John Ikonomopoulos and Nikolaos D. Andritsos
Microorganisms 2026, 14(5), 1097; https://doi.org/10.3390/microorganisms14051097 - 12 May 2026
Abstract
Listeria monocytogenes is already a well-known foodborne bacterial pathogen, ubiquitously dispersed not only in the food production environment but also in the primary animal production environment as well. The present study performed whole-genome characterization of the multidrug-resistant (MDR) L. monocytogenes strain BF11, previously
[...] Read more.
Listeria monocytogenes is already a well-known foodborne bacterial pathogen, ubiquitously dispersed not only in the food production environment but also in the primary animal production environment as well. The present study performed whole-genome characterization of the multidrug-resistant (MDR) L. monocytogenes strain BF11, previously isolated from raw pet food and phenotypically described for antimicrobial resistance. To this end, the genomic analysis performed on the isolate confirmed the pathogen’s designation as a serotype 1/2b strain belonging to ST5 and CC5 (Lineage I), carrying multiple MDR genes, stress-related genes, and mobile genetic elements, despite the absence of plasmids. The strain is phylogenetically closely related to Lineage I epidemic strains (e.g., F2365), as it has a full-length inlA and a functional prfA, rendering it capable of invading human cells and marking its high virulence. Overall, this strain may represent a potentially novel genomic profile when core genome multilocus sequence typing (cgMLST) is used, although further data from additional isolates would be required to confirm its classification within a new Complex Type, while displaying a hybrid unique profile. It is an evolved ST5 L. monocytogenes strain that has acquired genetic material conferring a “clinical signature” (Lineage I-like) and an extensive resistance network. Therefore, presence of L. monocytogenes strain BF11 in pet food is alarming, since such hybrid strains often evade surveillance monitoring as they do not fit strictly into classical categories, posing a serious food safety and public health threat in the concept of One Health.
Full article
(This article belongs to the Special Issue Advances in Foodborne Pathogenic Bacteria Microcosm Mapping and Its Interrelation Within the Food Production Setting)
►▼
Show Figures
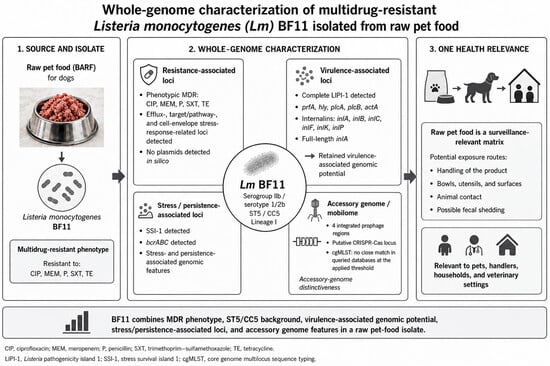
Graphical abstract
Open AccessReview
Bird–Borrelia Interactions: A Historical Review and Their Significance for Human Disease Ecology
by
András P. Bózsik, Dömötör M. László and Borisz Egri
Microorganisms 2026, 14(5), 1096; https://doi.org/10.3390/microorganisms14051096 - 12 May 2026
Abstract
Research increasingly identifies wild birds, particularly long-distance migratory species, as epidemiologically relevant hosts and vectors for tick-borne Borrelia species that pose risks to both avian and human health. This review contextualizes avian-associated Borrelia research historically and microbiologically, showing the role of avian hosts
[...] Read more.
Research increasingly identifies wild birds, particularly long-distance migratory species, as epidemiologically relevant hosts and vectors for tick-borne Borrelia species that pose risks to both avian and human health. This review contextualizes avian-associated Borrelia research historically and microbiologically, showing the role of avian hosts in the ecology of agents causing relapsing fever and Lyme borreliosis. We identify key publications that trace the evolution of Borrelia research—from early microscopic observations of spirochetes to the modern molecular and serological evidence. The review collects literature on the process by which Borrelia gained early scientific attention due to its characteristic morphology and elevated bloodstream concentrations during septicemic phases, which enabled early etiological links between the microbe and disease. It follows the recognition of avian spirochetosis caused by Borrelia anserina and charts the shift in focus after the discovery of Borrelia burgdorferi sensu lato (Subgen. novum recomm. Borreliella, Lyme-group Borrelia). Publications listed show that birds can transport infected human-parasitic ticks over long distances and, in certain bird species, selectively amplify Lyme-group Borrelia species, especially Borrelia garinii, which has the highest temperature tolerance and is thus potentially viable in avian hosts. The literature supports the role of birds in maintaining and disseminating Borrelia infections and infected ticks across continents.
Full article
(This article belongs to the Special Issue Borreliosis and Other Tick-Borne Diseases in the Northern and Southern Hemispheres, Second Edition)
Open AccessSystematic Review
Structure and Function of the Dental Plaque Microbiome in Eubiosis: A Systematic Review of Ethnic-Racial Influences
by
Edisson Ronaldo Duran Yunga and María de Lourdes Rodriguez Coyago
Microorganisms 2026, 14(5), 1095; https://doi.org/10.3390/microorganisms14051095 - 12 May 2026
Abstract
While a conserved core microbiome is shared across healthy individuals, significant interindividual taxonomic variation exists; however, the specific influence of genetic ancestry on supragingival plaque structure in eubiosis remains unclear. This systematic review analyzed evidence regarding taxonomic variations in supragingival plaque associated with
[...] Read more.
While a conserved core microbiome is shared across healthy individuals, significant interindividual taxonomic variation exists; however, the specific influence of genetic ancestry on supragingival plaque structure in eubiosis remains unclear. This systematic review analyzed evidence regarding taxonomic variations in supragingival plaque associated with ethnicity in systemically healthy populations. A search was conducted in PubMed, Scopus, ScienceDirect, and Scielo following PRISMA 2020 guidelines, covering literature up to October 2025. Cross-sectional studies using genomic sequencing or metagenomics were included, with quality assessed via the GRADE system. Six studies met eligibility criteria. Results identified a universal core microbiome structurally dominated by Corynebacterium spp. and Streptococcus spp. However, distinct ethnic-specific taxonomic signatures emerged, such as the enrichment of Fusobacterium spp. in African Americans and Corynebacterium spp. in Caucasians, alongside the exclusive presence of Sneathia spp. in Burmese individuals. Although a basal microbial architecture necessary for homeostasis exists, ethnicity acts as a biological filter defining distinctive bacterial profiles and differential susceptibilities. These findings suggest that while the core microbiome is conserved, the composition of peripheral species in the dental plaque hedgehog structure varies according to ancestry. This supports a transition from standardized dental care to personalized medicine oriented towards the patient’s biological heritage.
Full article
(This article belongs to the Special Issue Oral Microbiomes and One Health Approach)
►▼
Show Figures
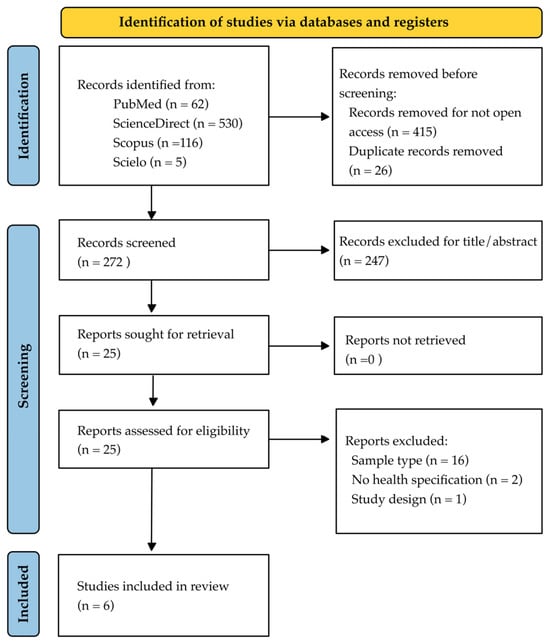
Figure 1
Open AccessArticle
Genomic and Phenotypic Insights into Carbapenem-Resistant Pseudomonas aeruginosa in the Aquatic Environments of the Tibetan Plateau
by
Dingxiang Lu, Lin Liu, Zhongwei Yang, Tianjiao Chen, Dong Yang, Danyang Shi, Shuqing Zhou, Junwen Li, Haibei Li and Min Jin
Microorganisms 2026, 14(5), 1094; https://doi.org/10.3390/microorganisms14051094 - 12 May 2026
Abstract
Carbapenem-resistant Pseudomonas aeruginosa is increasingly becoming a global health threat. However, although aquatic environments are key resistance reservoirs, data obtained from high-altitude ecosystems are scarce. Whole-genome sequencing of eight carbapenem-resistant P. aeruginosa isolates collected from aquatic environments in the Tibetan Plateau identified three
[...] Read more.
Carbapenem-resistant Pseudomonas aeruginosa is increasingly becoming a global health threat. However, although aquatic environments are key resistance reservoirs, data obtained from high-altitude ecosystems are scarce. Whole-genome sequencing of eight carbapenem-resistant P. aeruginosa isolates collected from aquatic environments in the Tibetan Plateau identified three sequence types (STs), with ST1420 predominating (62.5%, 5/8). Phylogenetic analysis revealed a close clustering of isolates with those from distant clinical settings, suggesting potential cross-habitat transmission. All studied strains were multidrug-resistant, exhibiting 100% resistance to imipenem, ceftriaxone, and trimethoprim–sulfamethoxazole. This included the PA6 strain, which showed multiple-antibiotic resistance. Eight strains harbored the intrinsic carbapenemase gene blaOXA-50. The diverse virulence-gene profiles of strains PA2, PA4, and PA6 aligned with their high pathogenicity observed both in vitro and in vivo. However, virulence genotypes sometimes did not correlate with phenotypes, revealing the limitations of relying on static genetic information alone. This study highlights the aquatic environments of the Tibetan Plateau as reservoirs of carbapenem-resistant P. aeruginosa with substantial genetic diversity and divergent pathogenic potential, underscoring their public-health relevance.
Full article
(This article belongs to the Special Issue Genomics of Marine and Aquatic Bacteria: A Focus on Novel Taxa, Diversity and Biotechnological Potential, 3rd Edition)
►▼
Show Figures

Figure 1
Open AccessReview
Avibacterium paragallinarum: Pathogenesis Mechanisms and Subunit Vaccine Development
by
Zhihua Li, Ying Liu, Zhenyi Liu, Zhaoling Jiang, Yawen Wang, Baozhu Xing, Chen Mei and Hongjun Wang
Microorganisms 2026, 14(5), 1093; https://doi.org/10.3390/microorganisms14051093 - 12 May 2026
Abstract
Avibacterium paragallinarum (A. paragallinarum) is the primary causative agent of infectious coryza in chickens. Infection often leads to growth retardation in broilers and a 10% reduction in egg production, reaching over 40% in laying hens. The problem is particularly severe under
[...] Read more.
Avibacterium paragallinarum (A. paragallinarum) is the primary causative agent of infectious coryza in chickens. Infection often leads to growth retardation in broilers and a 10% reduction in egg production, reaching over 40% in laying hens. The problem is particularly severe under intensive farming conditions, significantly jeopardizing global poultry health and farming profitability. From a ‘One Health’ perspective, this not only disrupts the stability of the food supply chain, but also increases antibiotic usage due to disease prevention and control needs, thereby aggravating antimicrobial resistance (AMR) and posing a global public health challenge. This review systematically summarizes advances in the pathogenesis of A. paragallinarum and the protective immunity induced by subunit vaccines. It focuses on the infection mechanisms of A. paragallinarum, emphasizing its colonization strategies in the infraorbital sinus and nasal epithelium of chickens, and analyzes the roles of key virulence factors such as hemagglutinin and capsule in adhesion, colonization, and immune evasion. We integrate the tissue-specific pathogenesis of A. paragallinarum with the role of respiratory commensal microbiota in facilitating infection, providing an in-depth analysis of the bacterium’s key immune evasion strategies, thus offering novel insights into host–pathogen-microbiome interactions. Concurrently, to the best of our knowledge, this review provides the first comprehensive overview of current developments in subunit vaccines and their immunoprotective properties, with special attention to limitations in eliciting mucosal immune responses. By delving into the pathogen-host interaction mechanisms, this review aims to inform the optimization of subunit vaccine design and immunization strategies. Ultimately, it seeks to establish a theoretical basis and practical framework for precise control of A. paragallinarum.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►▼
Show Figures
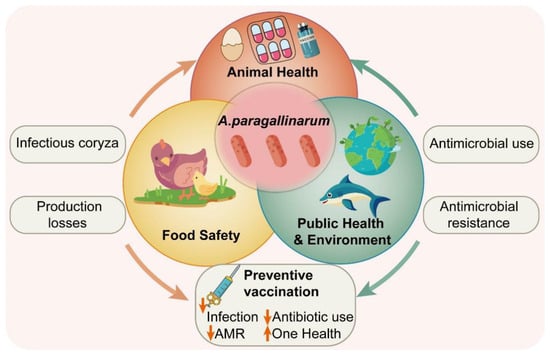
Figure 1
Open AccessArticle
Conservation Tillage-Mediated Rhizosphere Microbial Community Remodeling Drives Soil Organic Carbon Accumulation and Nitrogen and Phosphorus Transformation in Farmland
by
Haogeng Zhao, Meijuan Cheng, Shuli Wei, Gongfu Shi, Jing Fang, Huimin Shi, Qingze Liu, Yan Qu, Weijing Zhang, Fang Luo, Yu Wang, Zhanyuan Lu, Dejian Zhang and Xiaoqing Zhao
Microorganisms 2026, 14(5), 1092; https://doi.org/10.3390/microorganisms14051092 - 12 May 2026
Abstract
Conservation tillage has an influence on the cultivation and sustainable utilization of farmland. However, the microbial mechanism driving soil nutrient cycling in conservation tillage and its regulation pathway remain unclear. Based on a positioning experiment in black soil areas, this study systematically compared
[...] Read more.
Conservation tillage has an influence on the cultivation and sustainable utilization of farmland. However, the microbial mechanism driving soil nutrient cycling in conservation tillage and its regulation pathway remain unclear. Based on a positioning experiment in black soil areas, this study systematically compared the effects of no-tillage (NT) and moldboard tillage (MT) combined with different straw returning amounts (straw non-returning, NS; straw half-returning, HS; straw full-returning, TS) on the composition of soil carbon (C), nitrogen (N) and phosphorus (P) and focused on the role of microbial community structure succession and functional changes in soil nutrient cycling. Microbial community remodeling driven by tillage measures was mainly regulated by C and N components. Bacterial modules 2 and 4 and fungal modules 1 and 2 were key for regulating the C, N and P cycle, of which 87 bacteria and 45 fungi taxa represented the core driving microorganisms. The total amount of no-tillage straw return reduced the formation and accumulation of labile organic carbon fractions by enriching yeast-like fungi and inhibiting the expression of complex organic matter decomposition genes. Tillage mainly promoted the accumulation of labile organic carbon fractions and nutrient release by regulating the bacterial community, while no-tillage straw returning promoted the accumulation of total organic carbon and organic nitrogen fixation by promoting the fungal community. This study revealed the biological pathway of conservation tillage that drives soil nutrient cycling by regulating key microbial communities. It also provides a microbiological basis for sustainable soil management in black soil areas.
Full article
(This article belongs to the Topic Transport, Transformation and Cycling of Elements in Water and Soil and Their Response to Human Activities)
►▼
Show Figures
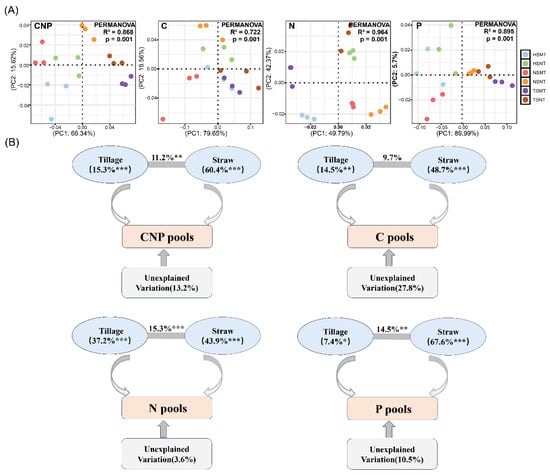
Figure 1
Open AccessReview
The Gut–Brain Axis in Post-Traumatic Stress Disorder: From Biological Mechanisms to Microbiome-Based Therapeutic Strategies—A Narrative Review
by
Eun Jin Yang and Hee Ra Park
Microorganisms 2026, 14(5), 1091; https://doi.org/10.3390/microorganisms14051091 - 11 May 2026
Abstract
Post-traumatic stress disorder (PTSD) is a debilitating psychiatric condition that impairs psychological functioning and increases susceptibility to various chronic illnesses, including inflammatory, metabolic, and cognitive disorders. Recent advances in neuroscience and microbiology have identified the brain–gut–microbiota axis as a key mediator of neuroimmune
[...] Read more.
Post-traumatic stress disorder (PTSD) is a debilitating psychiatric condition that impairs psychological functioning and increases susceptibility to various chronic illnesses, including inflammatory, metabolic, and cognitive disorders. Recent advances in neuroscience and microbiology have identified the brain–gut–microbiota axis as a key mediator of neuroimmune and neuroendocrine regulations, providing new insight into the pathophysiology of PTSD. This review synthesizes current findings from preclinical and clinical studies on gut microbiome alterations in PTSD, highlighting the underlying mechanistic pathways. Dysbiosis in PTSD is associated with immune dysregulation, altered neuroendocrine signaling, and neurotransmitter imbalances. Animal models, particularly those using the single prolonged stress paradigm, have demonstrated behavioral and microbial changes that mirror the characteristics of human PTSD. Human studies have revealed reduced abundance of beneficial bacterial taxa and increased inflammation-associated genera in patients with PTSD. Although emerging evidence supports the role of gut microbiota in PTSD, further research is needed to establish causal relationships and optimize microbiome-targeted therapies. Overall, the gut microbiome offers a novel and potentially modifiable target for the prevention and treatment of PTSD.
Full article
(This article belongs to the Section Gut Microbiota)

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Biosciences, Applied Microbiology, Fermentation, Marine Drugs, Microorganisms
Microbial Cell Factories for Natural Products
Topic Editors: Carlos Barreiro, Ana Ibáñez, José L. BarredoDeadline: 31 May 2026
Topic in
Energies, Geosciences, Hydrology, Plants, Microorganisms, Sustainability, Agriculture
Transport, Transformation and Cycling of Elements in Water and Soil and Their Response to Human Activities
Topic Editors: Jiutan Liu, Junfu DongDeadline: 30 June 2026
Topic in
Dairy, Foods, Microorganisms, Nutrients, Metabolites
The Efficacy of Probiotics and Their Functional Metabolites in Fermented Foods
Topic Editors: Rina Wu, Wenjun Liu, Zhen Wu, Feiyu AnDeadline: 31 July 2026
Topic in
Microorganisms, JCM, Biomedicines, Oral, Clinics and Practice
Periodontal and Peri-Implant Disease: Contemporary Perspectives on Pathogenesis and Precision Management
Topic Editors: Giuseppe Troiano, Vito Carlo Alberto Caponio, Mario DioguardiDeadline: 31 August 2026

Conferences
Special Issues
Special Issue in
Microorganisms
Pathobiology, Infection Biology and Control of Protozoan Parasites—the ONE HEALTH Approach
Guest Editors: Cláudia Marques, Maria João Gouveia, Susana R. SousaDeadline: 15 May 2026
Special Issue in
Microorganisms
Role of Soil Microorganisms in Agricultural Production and Environmental Biotechnologies
Guest Editor: Gianluigi CardinaliDeadline: 20 May 2026
Special Issue in
Microorganisms
Borreliosis and Other Tick-Borne Diseases in the Northern and Southern Hemispheres, Second Edition
Guest Editor: Giusto TrevisanDeadline: 30 May 2026
Special Issue in
Microorganisms
Phytoplankton Faced with Emergent Pollutants
Guest Editors: Katia Comte, Stéphanie FayolleDeadline: 31 May 2026
Topical Collections
Topical Collection in
Microorganisms
Microbial Life in Extreme Environments
Collection Editor: Ricardo Amils
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde









