-
 Snail Slime vs. Glycolic Acid: Cell Insights into Skin and Endothelial Effects
Snail Slime vs. Glycolic Acid: Cell Insights into Skin and Endothelial Effects -
 PFAS Impair Protamine/DNA Binding and May Promote Sperm DNA Damage
PFAS Impair Protamine/DNA Binding and May Promote Sperm DNA Damage -
 Zinc Therapy in Mild Cognitive Impairment: Cognitive Stabilization in Pharmacodynamically Responsive Patients in the ZINCAiD Trial
Zinc Therapy in Mild Cognitive Impairment: Cognitive Stabilization in Pharmacodynamically Responsive Patients in the ZINCAiD Trial -
 Deciphering the Fasciola hepatica Glycocode in Host–Parasite Interactions
Deciphering the Fasciola hepatica Glycocode in Host–Parasite Interactions -
 Loss of Mucin-Type O-Glycans Leads to Synaptic Defects in SLC35A2-CDG Fly Model
Loss of Mucin-Type O-Glycans Leads to Synaptic Defects in SLC35A2-CDG Fly Model
Journal Description
Biomolecules
Biomolecules
is an international, peer-reviewed, open access journal on structures and functions of bioactive and biogenic substances, molecular mechanisms with biological and medical implications as well as biomaterials and their applications, published monthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Biochemistry and Molecular Biology) / CiteScore - Q1 (Biochemistry)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.9 days after submission; acceptance to publication is undertaken in 2.9 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Sections: published in 15 topical sections.
- Testimonials: See what our editors and authors say about Biomolecules.
- Companion journal: Receptors.
Impact Factor:
4.8 (2024);
5-Year Impact Factor:
5.6 (2024)
Latest Articles
Early Recurrence of HCC Is Driven by Inflammation-Related HIF-1α Independent Angiogenesis Rather than Hypoxia-Induced Immune Escape
Biomolecules 2026, 16(5), 723; https://doi.org/10.3390/biom16050723 (registering DOI) - 14 May 2026
Abstract
Background: Hepatocellular carcinoma (HCC) shows a high rate of early recurrence after curative resection, indicating a critical contribution of tumor microenvironment-driven molecular mechanisms. Early recurrence of hepatocellular carcinoma is defined as recurrence within 6 months after curative resection, with a prevalence exceeding 30%.
[...] Read more.
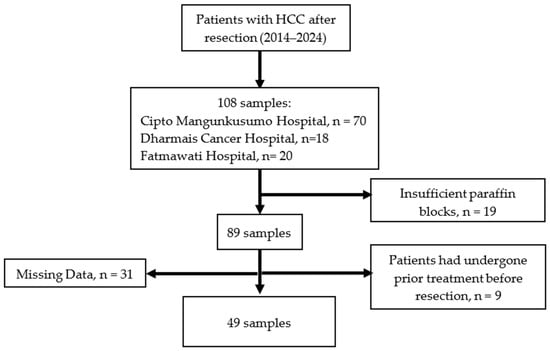
Background: Hepatocellular carcinoma (HCC) shows a high rate of early recurrence after curative resection, indicating a critical contribution of tumor microenvironment-driven molecular mechanisms. Early recurrence of hepatocellular carcinoma is defined as recurrence within 6 months after curative resection, with a prevalence exceeding 30%. Hypoxia signaling and immune dysregulation have been implicated, yet their compartment-specific relevance remains unclear. Methods: This multicenter nested case–control study included 49 HCC patients to evaluate associations between hypoxia-inducible factor-1 alpha (HIF-1α), vascular endothelial growth factor (VEGF), tumor-infiltrating lymphocytes (TILs), CD4+ T cells, CD8+ T cells, regulatory T cells (Tregs), programmed cell death protein 1 (PD-1), and programmed death-ligand 1 (PD-L1) and early recurrence after resection. TIL density was assessed using hematoxylin and eosin staining, while immunohistochemistry was performed to quantify intratumoral and peritumoral expression of the studied markers. Receiver operating characteristic (ROC) curve analysis was used to evaluate the predictive performance. Recurrence-free survival (RFS) was analyzed using the Kaplan–Meier, and independent predictors were identified using multivariate Cox proportional hazards regression. Results: Early recurrence occurred in 11 of 49 patients (22.4%) of Child–Pugh A patients. Recurrent tumors were characterized by elevated VEGF expression despite absent HIF-1α, alongside significant depletion of intratumoral TILs (HR 5.02; 95% CI 1.09–23.26), CD4+ (HR 7.68; 95% CI 1.66–35.60) and CD8+ cells (HR 6.68; 95% CI 1.77–25.23) and reduced peritumoral CD8+ infiltration (HR 4.20; 95% CI 1.11–15.91). Multivariable analysis identified low intratumoral CD4+ (HR 7.98; 95% CI 1.63–39.07) and reduced peritumoral CD8+ expression (HR 4.98; 95% CI 1.14–21.70) as independent predictors, whereas HIF-1α, VEGF, Treg, PD-1, and PD-L1 were not significantly associated. Conclusions: Early HCC recurrence shows HIF-1α-independent angiogenesis alongside spatial immune depletion, supporting integrated immune profiling over single angiogenic markers.
Full article
(This article belongs to the Special Issue Tumor Microenvironment, Immunology and Precision Medicine of Liver Cancer)
►
Show Figures
Open AccessReview
Biomolecular Interfaces in Targeted Nano-Drug Delivery: Molecular Recognition, Signaling Modulation, and Translational Pathways
by
Zeyu Wang, Lixia Dai, Zhen Zhu and Xiaofei Shang
Biomolecules 2026, 16(5), 722; https://doi.org/10.3390/biom16050722 (registering DOI) - 14 May 2026
Abstract
Traditional pharmacotherapy is often constrained by suboptimal bioavailability and systemic toxicity. Biomolecularly inspired nano-drug delivery systems (nano-DDS) have emerged as precise platforms to overcome these barriers by orchestrating molecular interactions at the bio-nano interface. This review systematically evaluates the molecular recognition mechanisms and
[...] Read more.
Traditional pharmacotherapy is often constrained by suboptimal bioavailability and systemic toxicity. Biomolecularly inspired nano-drug delivery systems (nano-DDS) have emerged as precise platforms to overcome these barriers by orchestrating molecular interactions at the bio-nano interface. This review systematically evaluates the molecular recognition mechanisms and biochemical principles governing nano-DDS performance. We systematically evaluate how passive targeting relies on the EPR effect—dictated by the nanocarrier’s physicochemical properties—and how active targeting exploits ligand-receptor affinity to enhance cellular uptake. Special emphasis is placed on bioresponsive strategies that utilize pathological cues—such as pH gradients, redox potential, and enzymatic activity—for intelligent, on-demand drug release. Furthermore, we discuss structure-function relationships in lipid, polymeric, and biologically derived systems, highlighting their roles in modulating therapeutic signaling in oncology and inflammatory diseases. Finally, translational hurdles and emerging AI-driven molecular design strategies are critically examined.
Full article
(This article belongs to the Special Issue Advances in Nano-Based Drug Delivery: Unveiling the Next Frontier)
►▼
Show Figures
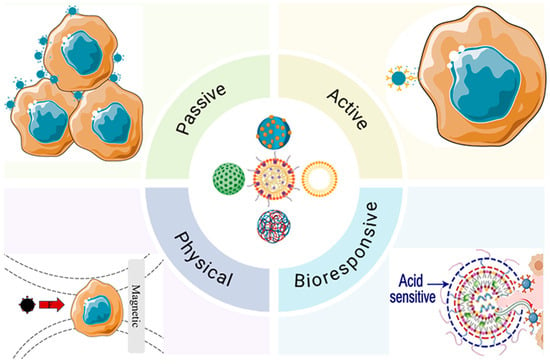
Figure 1
Open AccessArticle
Oxidative–Nitrosative Stress and Routine Biochemical Parameters in Amyotrophic Lateral Sclerosis: Associations with Clinical Status and Disease Duration—A Pilot Study
by
Pavlína Malá, Nela Váňová, Ondřej Malý and Oldřich Vyšata
Biomolecules 2026, 16(5), 721; https://doi.org/10.3390/biom16050721 (registering DOI) - 13 May 2026
Abstract
Background: This pilot study examined whether oxidative–nitrosative stress is associated with clinical status in amyotrophic lateral sclerosis (ALS). We analyzed associations between plasma markers of oxidative–nitrosative imbalance and ALSFRS–R, disease duration, survival, and routine biochemical parameters. Methods: Twenty-nine ALS patients fulfilling the Gold
[...] Read more.
Background: This pilot study examined whether oxidative–nitrosative stress is associated with clinical status in amyotrophic lateral sclerosis (ALS). We analyzed associations between plasma markers of oxidative–nitrosative imbalance and ALSFRS–R, disease duration, survival, and routine biochemical parameters. Methods: Twenty-nine ALS patients fulfilling the Gold Coast diagnostic criteria were enrolled. Plasma levels of 3-nitrotyrosine (3–NT), 8-oxo-2′-deoxyguanosine (8–oxodG), malondialdehyde (MDA), glutathione (GSH), non-protein thiols (NP–SH), and non-protein disulfides (NP–SS–NP), as well as creatinine, urea, uric acid and BMI, were measured. Associations with ALSFRS–R and disease duration were evaluated using non-parametric correlation analyses and second-order polynomial regression (adjusted R2), while survival was explored using Kaplan–Meier analysis and multivariable Cox regression. Given the modest sample, we considered statistical power and applied Benjamini–Hochberg false discovery rate (FDR) correction within marker families. Results: At the uncorrected significance level, 3–NT showed a positive correlation with ALSFRS–R and a negative correlation with disease duration, and NP–SH correlated negatively with disease duration; however, these associations did not remain significant after FDR correction (FDR-adjusted p ≥ 0.099). Other oxidative–nitrosative markers and biochemical parameters showed no robust relationships with clinical measures. In Cox models, 3–NT was not significantly associated with survival (HR 3.44 per 1 nM, 95% CI 0.25–47.97, p = 0.358), whereas older age predicted higher mortality (HR 1.05 per year, 95% CI 1.00–1.10, p = 0.036). Conclusions: 3–NT and NP–SH exhibited the strongest trends among the investigated markers, but their clinical associations in this small cross-sectional cohort remain exploratory and require confirmation in larger longitudinal studies.
Full article
(This article belongs to the Special Issue Molecular Biomarkers of Oxidative and Nitrosative Stress: Mechanisms, Detection, and Clinical Significance)
►▼
Show Figures

Figure 1
Open AccessArticle
The Multifunctional Peptide AP10W Enhances Skin Wound Healing Through Macrophage Reprogramming and Angiogenesis
by
Cuiling Xuan, Zixuan Liu, Peng Zhang, Bojian Liu, Zhiqin Gao and Fei Wu
Biomolecules 2026, 16(5), 720; https://doi.org/10.3390/biom16050720 (registering DOI) - 13 May 2026
Abstract
Skin wound healing is a complex and highly coordinated biological process involving inflammation, cell migration and proliferation, angiogenesis, extracellular matrix remodeling and tissue regeneration. While the zebrafish-derived antimicrobial peptide AP10W exhibits broad-spectrum antimicrobial properties, its potential in tissue repair remains unexplored. Herein, we
[...] Read more.
Skin wound healing is a complex and highly coordinated biological process involving inflammation, cell migration and proliferation, angiogenesis, extracellular matrix remodeling and tissue regeneration. While the zebrafish-derived antimicrobial peptide AP10W exhibits broad-spectrum antimicrobial properties, its potential in tissue repair remains unexplored. Herein, we demonstrate that AP10W possesses intrinsic wound-healing capabilities, providing a preliminary investigation into its underlying mechanisms. In this study, using a full-thickness murine wound model and in vitro cell-based assays to evaluate the effects of AP10W on fibroblasts, keratinocytes, endothelial cells, and macrophages, we found that AP10W significantly promoted fibroblast and keratinocyte migration and proliferation. Furthermore, it enhanced endothelial cell motility, survival, and tube formation, while upregulating key pro-angiogenic factors, including Vascular endothelial growth factor A (VEGFA), Platelet-derived growth factor (PDGF), and Fibroblast growth factor 2 (FGF2). Concurrently, AP10W drove macrophage reprogramming from a pro-inflammatory M1 phenotype toward a pro-healing M2 state, as evidenced by upregulated Arginase-1 (Arg-1) and Interleukin-10 (Il-10) expression, alongside attenuated Tumor necrosis factor-alpha (Tnf-α), Interleukin-1 beta (Il-1β), Interleukin-6 (Il-6), and Inducible nitric oxide synthase (iNOS) levels. In vivo, the topical application of AP10W accelerated wound closure, markedly improving re-epithelialization, collagen deposition, vascularization, tissue perfusion, and skin appendage regeneration. Preliminary mechanistic studies revealed that AP10W increased YAP expression and nuclear translocation; conversely, the pharmacological inhibition of YAP significantly abrogated these pro-healing effects. Collectively, our findings identify AP10W as a multifunctional peptide with potent wound-healing properties, positioning it as a promising candidate for wound therapy.
Full article
(This article belongs to the Section Natural and Bio-derived Molecules)
►▼
Show Figures
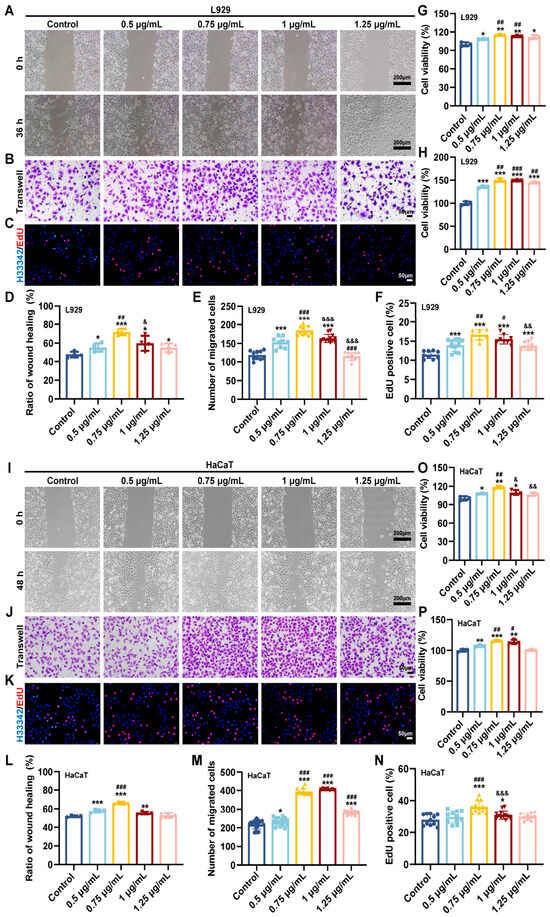
Figure 1
Open AccessReview
TIF1 Family Proteins as Modulators of Cell Death: Mechanisms and Therapeutic Opportunities
by
Dong Yang and Yuchen Chen
Biomolecules 2026, 16(5), 719; https://doi.org/10.3390/biom16050719 (registering DOI) - 13 May 2026
Abstract
Regulated cell death is essential for development, tissue homeostasis, host defense, and disease. Beyond apoptosis, it is now clear that other forms of cell death, including ferroptosis, pyroptosis, and necroptosis, also contribute to pathology, often in interconnected rather than isolated ways. Within this
[...] Read more.
Regulated cell death is essential for development, tissue homeostasis, host defense, and disease. Beyond apoptosis, it is now clear that other forms of cell death, including ferroptosis, pyroptosis, and necroptosis, also contribute to pathology, often in interconnected rather than isolated ways. Within this broader framework, the transcriptional intermediary factor 1 (TIF1) family, comprising TRIM24, TRIM28, TRIM33, and TRIM66, has emerged as an important group of regulators linking stress adaptation, cell-state control, and cell death susceptibility. Although these proteins belong to the same family, they influence cell death through distinct and context-dependent mechanisms. Across the TIF1 family, apoptosis is by far the most extensively studied cell death phenotype, whereas links to ferroptosis, pyroptosis, and necroptosis remain more limited, more context dependent, and more unevenly distributed across individual members. Cell death often becomes evident when TIF1-dependent stress-buffering programs are disrupted, highlighting both their biological importance and potential therapeutic relevance. At the same time, family-level differences are emerging, while the underlying mechanisms remain incompletely understood, and recent advances in this field have not been synthesized. This review summarizes how TIF1 family members intersect with different cell death programs, discusses emerging translational opportunities and challenges, and highlights key mechanistic questions for future study.
Full article
(This article belongs to the Special Issue Advancing Innovations in Biomedical Research for Translational Applications)
►▼
Show Figures
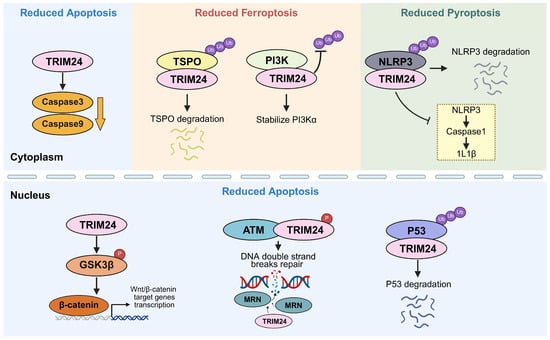
Figure 1
Open AccessReview
Oxidative Stress in Dry Eye Disease: Molecular Mechanisms and Emerging Therapeutic Strategies
by
Tingting Tang, Jiaxin Yang and Hongbo Yin
Biomolecules 2026, 16(5), 718; https://doi.org/10.3390/biom16050718 (registering DOI) - 13 May 2026
Abstract
►▼
Show Figures
Dry eye disease (DED) is a chronic inflammatory disorder of the ocular surface, characterized by tear film homeostasis imbalance, with aging being identified as a crucial independent risk factor. Oxidative stress, which refers to the excessive production of reactive oxygen species (ROS) and
[...] Read more.
Dry eye disease (DED) is a chronic inflammatory disorder of the ocular surface, characterized by tear film homeostasis imbalance, with aging being identified as a crucial independent risk factor. Oxidative stress, which refers to the excessive production of reactive oxygen species (ROS) and reactive nitrogen substances during mitochondrial metabolism and the weakened protective effect of antioxidants, plays a central role in this process. With aging, the mitochondrial function of ocular surface tissues, such as the corneal epithelium, meibomian glands, and lacrimal glands, declines. Concurrently, the activity of endogenous antioxidant enzymes (such as superoxide dismutase and glutathione peroxidase) decreases, and the levels of tear antioxidants such as lactoferrin also decrease. These age-related changes collectively lead to excessive accumulation of ROS, triggering oxidative stress that directly damages biomacromolecules in ocular surface cells and impairs the stability of the tear film. Furthermore, we have summarized the current therapeutic strategies for oxidative stress in DED, including both conventional antioxidants and emerging approaches such as eye drops based on nanoenzymes, thermosensitive hydrogels, intense pulsed light therapy, and drug-eluting contact lenses. By combining the new progress in the delivery systems of biomaterials-based drugs with mechanism-guided interventions, this review systematically establishes the intimate functional linkages between mitochondrial dysfunction, oxidative stress, and the pathogenesis of DED and focuses on elaborating the translational potential of advanced biomaterials-based antioxidant regimens, aiming to provide novel foundations and insights theoretical for the development of more effective and precise therapeutic strategies for DED.
Full article

Figure 1
Open AccessFeature PaperArticle
Spiny Mice Show a Profibrotic Epicardial Mesothelial Response to Hypoxic Injury Comparable to C57BL/6 Mice
by
Konstantin Dergilev, Aleria Dolgodvorova, Zoya Tsokolaeva, Irina Iarushkina, Irina Beloglazova, Yulia Goltseva and Yelena Parfyonova
Biomolecules 2026, 16(5), 717; https://doi.org/10.3390/biom16050717 (registering DOI) - 13 May 2026
Abstract
Epicardial mesothelium plays a pivotal role in postinfarction cardiac repair by generating fibroblasts, producing extracellular matrix, and releasing paracrine mechanisms. However, interspecies differences have not been sufficiently studied, particularly in in vivo models of scar-free healing such as the African spiny mouse (
[...] Read more.
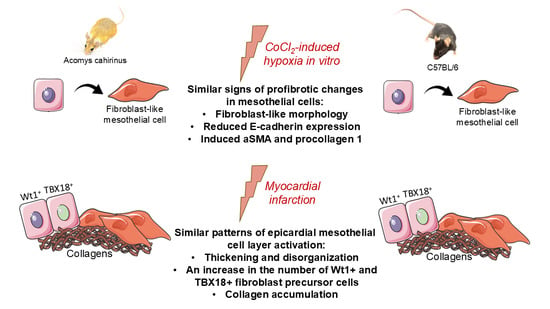
Epicardial mesothelium plays a pivotal role in postinfarction cardiac repair by generating fibroblasts, producing extracellular matrix, and releasing paracrine mechanisms. However, interspecies differences have not been sufficiently studied, particularly in in vivo models of scar-free healing such as the African spiny mouse (Acomys cahirinus). This study aimed to compare the profibrotic response of epicardial mesothelial cells (MCs) from Acomys and C57BL/6 mice to hypoxic stress, a key factor in postinfarction recovery. We isolated epicardial MCs from the African spiny mouse (Acomys cahirinus), a species with documented cardiac regenerative capabilities, and from C57BL/6 laboratory mice. Using a CoCl2-induced hypoxia model in vitro, we assessed cell viability, morphological changes, and expression of epithelial and fibroblast markers. In vivo, following experimental myocardial infarction (MI), we evaluated tissue hypoxia (pimonidazole adducts), epicardial activation (layer thickness, Wt1+ and TBX18+ progenitor cells), and collagen accumulation. The study was conducted using real-time PCR, Western blotting, immunohistochemical analysis and microscopic examination. In vitro, MCs from both species exhibited an epithelial-like phenotype under normoxic conditions, expressing E-cadherin and cytokeratin 18. Hypoxia (200 µM CoCl2) induced a comparable response in both Acomys and C57BL/6 cells, characterized by a shift to a spindle-shaped, fibroblast-like morphology, decreased E-cadherin expression, and increased pro-collagen 1 and α-SMA expression. Following MI, both species exhibited similarly extensive hypoxic areas affecting the epicardial zone. Epicardial activation dynamics were comparable: from day 3 post-MI, epicardial thickness increased significantly, and Wt1+ and TBX18+ progenitor cells accumulated, peaking during the first week. Collagen accumulation in the epicardial region was similar between species, although the number of Wt1+ cells was higher in C57BL/6 on day 7. Despite the well-known superior regenerative capacity of spiny mice, epicardial MCs from Acomys and C57BL/6 demonstrated similar signs of profibrotic responses to hypoxic stimulation both in vitro and following MI. These findings suggest that species-specific regenerative outcomes may not be attributable to differential acute epicardial sensitivity to hypoxia, but rather to downstream mechanisms or additional factors influencing the cardiac repair process. This study provides the first characterization of Acomys epicardial MCs and establishes a foundation for further investigation of evolutionarily conserved and species-specific mechanisms of cardiac regeneration.
Full article
(This article belongs to the Special Issue New Insights into Mesothelial Cells)
►▼
Show Figures
Graphical abstract
Open AccessArticle
High-Affinity Nanobody Against the LEDGF PWWP Domain Inhibits Chromatin Binding In Vitro
by
Thibault Vantieghem, Sofie Jansen, Thatcher Zinabu Akele, Pieterjan Van Maele, Sam Noppen, Dominique Schols, Maarten Dewilde, Zeger Debyser and Sergei V. Strelkov
Biomolecules 2026, 16(5), 716; https://doi.org/10.3390/biom16050716 (registering DOI) - 13 May 2026
Abstract
Background and objectives: The PWWP domain of lens epithelium-derived growth factor p75 (LEDGF/p75) mediates chromatin engagement through recognition of histone H3 lysine 36 di- and trimethylation (H3K36me2/3) and nucleosomal DNA. LEDGF/p75 plays a role in multiple human diseases. In particular, its interaction with
[...] Read more.
Background and objectives: The PWWP domain of lens epithelium-derived growth factor p75 (LEDGF/p75) mediates chromatin engagement through recognition of histone H3 lysine 36 di- and trimethylation (H3K36me2/3) and nucleosomal DNA. LEDGF/p75 plays a role in multiple human diseases. In particular, its interaction with HIV-1 integrase enables viral genome integration. However, the LEDGF PWWP domain remains difficult to target with small molecules as it lacks optimally shaped binding pockets. Here, we report the generation of high-affinity nanobodies (Nbs) to investigate the structure and function of this domain. Methods: Camelids were immunized with recombinant LEDGF PWWP domain, and immune phage display libraries were screened for affinity. Selected Nbs were recombinantly expressed in E. coli and purified. Their interaction with the PWWP domain of LEDGF and its close homolog HRP-2 was characterized using size-exclusion chromatography and surface plasmon resonance. Structural characterization of the Nbs was performed using X-ray crystallography. Functional effects on chromatin engagement were evaluated using an AlphaScreen assay. Results: Nine sequence-distinct Nbs were identified, seven of which were confirmed to bind the LEDGF PWWP domain with nanomolar affinities. Five Nbs also bound the HRP-2 domain, consistent with conserved functional surfaces, while two showed reduced affinity. The crystal structures of two Nbs (NbC03 and NbH10) confirmed there were canonical immunoglobulin folds, while the latter additionally revealed a domain-swapped dimer. Moreover, NbH10 dose-dependently inhibited the interaction between full-length LEDGF/p75 and H3K36me3-modified nucleosomes in vitro. Conclusions: This work establishes a validated panel of Nbs targeting the LEDGF PWWP domain and identifies one Nb capable of functionally disrupting the LEDGF–chromatin interaction. These Nbs serve as valuable tools for functional studies and structure-based drug design.
Full article
(This article belongs to the Section Molecular Biology)
►▼
Show Figures
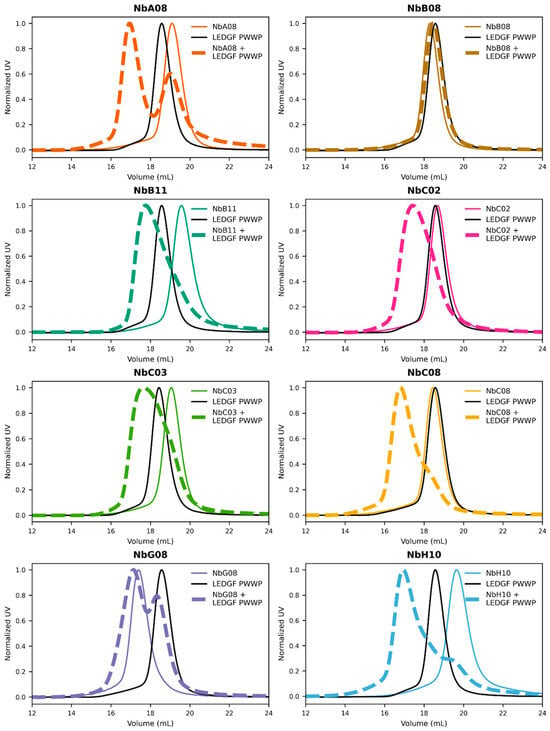
Figure 1
Open AccessArticle
Structure-Based Rational Design of TcAgo from Thermogladius calderae
by
Xiaochen Xie, Wanping Chen, Shi Chen, Tianxin Cai, Chendi Zhang, Jie Chen, Zhenni Xu, Zhuang Li, Longyu Wang and Lixin Ma
Biomolecules 2026, 16(5), 715; https://doi.org/10.3390/biom16050715 (registering DOI) - 13 May 2026
Abstract
Thermophilic prokaryotic Argonaute proteins (pAgos) have emerged as powerful tools for nucleic acid manipulation, with applications in nucleic acid detection, and DNA assembly. However, their strong dependence on high-temperature catalytic activity limits their utility under moderate conditions. TcAgo, a thermophilic Argonaute nuclease from
[...] Read more.
Thermophilic prokaryotic Argonaute proteins (pAgos) have emerged as powerful tools for nucleic acid manipulation, with applications in nucleic acid detection, and DNA assembly. However, their strong dependence on high-temperature catalytic activity limits their utility under moderate conditions. TcAgo, a thermophilic Argonaute nuclease from Thermogladius calderae, exhibits efficient DNA-guided target DNA cleavage above 80 °C, yet its structural basis and catalytic mechanism remain unclear. In this study, we attempted to analyze the structure of the TcAgo ternary complex and performed rational engineering based on its structure and characteristics of cold-adapted enzymes. A mutant, mTcAgo (K574G, D577G), was obtained with enhanced activity at moderate temperatures. Compared with the wild type, mTcAgo exhibited significantly improved cleavage activity toward both DNA and RNA targets at 37 °C. It utilized multiple guide types, including 5′OH- and 5′P-modified DNA and RNA guides, with a preference for 5′P-gDNA. mTcAgo displayed optimal activity at pH 7–8, broad salt tolerance, and an extended catalytic temperature range from 37 °C to 95 °C. Notably, it retained high activity after incubation at 90 °C, with a melting temperature of ~88 °C, and efficiently cleaved GC-rich targets under low Mg2+ conditions. These results demonstrate that rational cold-adaptation engineering can expand the functional temperature range of thermophilic pAgos, providing a promising strategy for developing versatile nucleic acid tools.
Full article
(This article belongs to the Section Biomacromolecules: Proteins, Nucleic Acids and Carbohydrates)
►▼
Show Figures
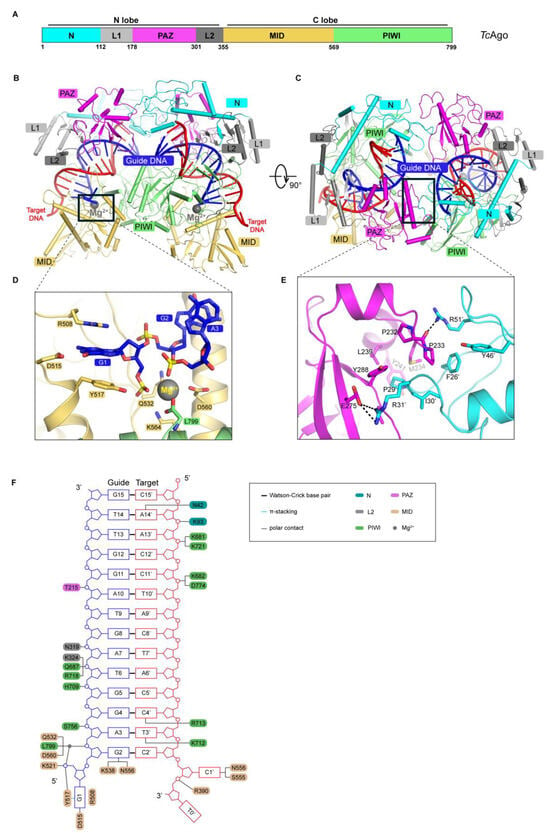
Figure 1
Open AccessArticle
Targeting Protein–Protein Interactions (PPIs) to Drive Functional Annotation: An Integrative Methodology to Study Senescence-Associated PPIs Using TMPRSS11a as Model Interactor
by
Roberto Rosales-Rojas, Christian Fernández, Mariela González-Avendaño, Ariela Vergara-Jaque and Mónica Cáceres
Biomolecules 2026, 16(5), 714; https://doi.org/10.3390/biom16050714 (registering DOI) - 13 May 2026
Abstract
Protein–protein interactions (PPIs) play a central role in regulating cellular processes. However, the identification and characterization of senescence-associated PPIs remain challenging. In this study, we developed and evaluated an integrative methodology based on selective proteomics to identify PPIs associated with cellular senescence induced
[...] Read more.
Protein–protein interactions (PPIs) play a central role in regulating cellular processes. However, the identification and characterization of senescence-associated PPIs remain challenging. In this study, we developed and evaluated an integrative methodology based on selective proteomics to identify PPIs associated with cellular senescence induced by TMPRSS11a. We started from isolated proteins that co-immunoprecipitated with TMPRSS11a and were subsequently identified by mass spectrometry. Building on this dataset, we implemented a workflow combining selective proteomics, structural bioinformatics, and experimental validation. Using this approach, we investigated the interaction between the transmembrane serine protease TMPRSS11a and the chaperone HSPA8. Structural bioinformatics analyses were performed to identify potential residues involved in the interaction interface, and the proximity of the TMPRSS11a–HSPA8 complex was evaluated using an in vitro proximity ligation assay. Our results provide evidence for an interaction between TMPRSS11a and HSPA8 and suggest its association with an enhanced senescence response. Overall, this study presents a workflow to investigate senescence-associated PPIs from proteomics-derived candidate proteins.
Full article
(This article belongs to the Special Issue Application of Bioinformatics in Medicine, 2nd Edition)
►▼
Show Figures
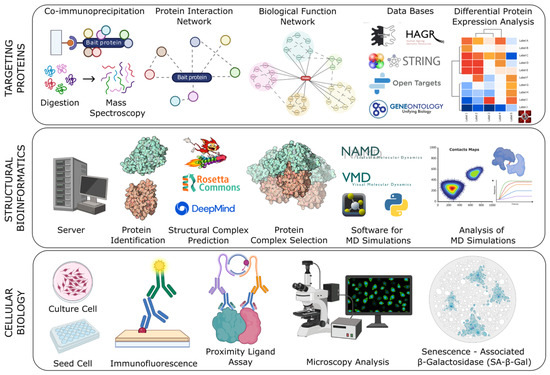
Figure 1
Open AccessArticle
Live-Cell Imaging of Microglia in Organotypic Brain Slices Using Microcontact Printing
by
Björn Y. P. Richardsen and Christian Humpel
Biomolecules 2026, 16(5), 713; https://doi.org/10.3390/biom16050713 (registering DOI) - 12 May 2026
Abstract
Microglia are brain immune cells that phagocytose cell debris and beta-amyloid plaques in patients with Alzheimer’s disease. They develop from round amoeboid cells into ramified microglia or large macrophages, which can be studied in three-dimensional organotypic mouse brain slices. In a recent publication,
[...] Read more.
Microglia are brain immune cells that phagocytose cell debris and beta-amyloid plaques in patients with Alzheimer’s disease. They develop from round amoeboid cells into ramified microglia or large macrophages, which can be studied in three-dimensional organotypic mouse brain slices. In a recent publication, we showed for the first time that we can track GFAP+ astrocytes and laminin+ vessels in organotypic brain slices using live-cell imaging . The aim of the present study was to use microcontact printing on organotypic brain slices to label microglia with Iba1 and CD11b antibodies and visualise them through live-cell imaging. We show that microglia can be easily labelled with antibodies and tracked via live-cell fluorescence microscopy for up to 20 days. Incubation in lipopolysaccharide (LPS) or granulocyte–macrophage colony-stimulating factor (GM-CSF) stimulates the migration of round amoeboid microglia, whereas interleukin-10 induces their differentiation into ramified forms. Taken together, we show the first-time live cell imaging of microglia in organotypic mouse brain slices using microcontact printing.
Full article
(This article belongs to the Section Cellular Biochemistry)
►▼
Show Figures
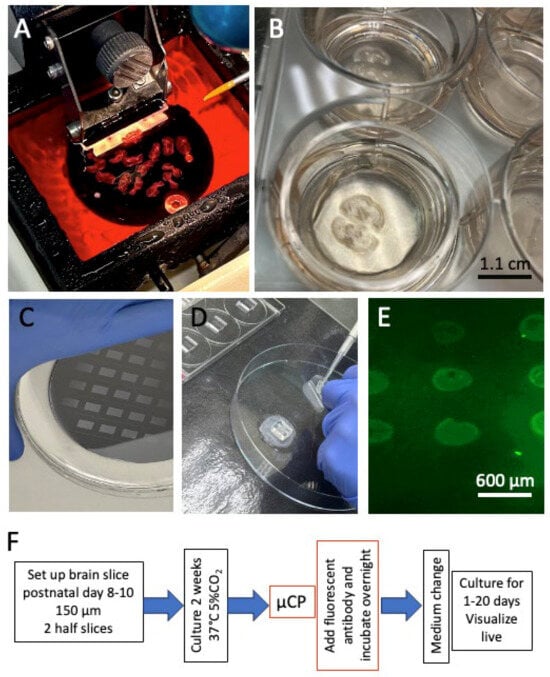

Figure 1
Open AccessCommunication
Epitranscriptomic Stability—Variable Extents of N1-Methyladenosine to N6-Methyladenosine Conversion Under Different Experimental Conditions
by
Frank Morales Shnaider, Hasna Kanan, Shrikant Patel and Norman H. L. Chiu
Biomolecules 2026, 16(5), 712; https://doi.org/10.3390/biom16050712 (registering DOI) - 12 May 2026
Abstract
The growing interest in epitranscriptomes has emphasized the need for accurate quantification of RNA modifications. However, the stability of RNA modifications under different experimental conditions remains poorly characterized. In this study, we use N1-methyladenosine (m1A) as a model for assessing stability.
[...] Read more.
The growing interest in epitranscriptomes has emphasized the need for accurate quantification of RNA modifications. However, the stability of RNA modifications under different experimental conditions remains poorly characterized. In this study, we use N1-methyladenosine (m1A) as a model for assessing stability. After exposing m1A ribonucleoside to various conditions that are commonly used in RNA protocols, the sample was analyzed using untargeted high-resolution mass spectrometry. Under alkaline or neutral pH, different extents of m1A were converted to N6-methyladenosine (m6A). Complete m1A-to-m6A conversion occurred at elevated temperatures. The m1A-to-m6A conversion was also dependent on the initial m1A concentration; the higher the concentration, the higher the rate of conversion. This poses a challenge to comparative studies if the initial amount of m1A in the control and sample of interest are not equal. No demethylation or depurination was detected. However, trace amount of N1-methylinosine was detected as a result of non-enzymatic deamination of m1A. Furthermore, the m1A-to-m6A conversion was consistently observed in a biological sample. To eliminate the bias that resulted from m1A-to-m6A conversion, the standard addition method was adopted. This report highlighted the challenges of having different extents of m1A-to-m6A conversion under specific experimental conditions and demonstrated a viable solution for resolving the issue.
Full article
(This article belongs to the Collection Feature Papers in Molecular Biomarkers)
►▼
Show Figures

Figure 1
Open AccessArticle
Biomimetic Studies on the Reactivity of Sulfur-Centered Radicals with Purine Moieties of DNA
by
Annalisa Masi, Sebastian Barata-Vallejo and Chryssostomos Chatgilialoglu
Biomolecules 2026, 16(5), 711; https://doi.org/10.3390/biom16050711 (registering DOI) - 12 May 2026
Abstract
The reaction of the HS•/S•− radical (pKa ~3.4), generated selectively from H2S by γ-irradiated N2-flushed aqueous solutions at pH 5, with purine nucleosides (dG or dA), a 10-mer double-stranded oligodeoxynucleotide (ds-ODNs), and calf thymus
[...] Read more.
The reaction of the HS•/S•− radical (pKa ~3.4), generated selectively from H2S by γ-irradiated N2-flushed aqueous solutions at pH 5, with purine nucleosides (dG or dA), a 10-mer double-stranded oligodeoxynucleotide (ds-ODNs), and calf thymus (ct) DNA was investigated, under various experimental conditions. Concurrent quantification of the four purine 5′,8-cyclo-2′-deoxynucleosides (cPu) and two 8-oxo-7,8-dihydro-2′-deoxypurines (8-oxo-Pu) by LC-MS/MS analysis using isotopomeric internal standards was achieved. The formation of 8-oxo-Pu is several tens of times larger than cPu. Mechanistic schemes for the formation of the two product groups are proposed. Hydrogen atom abstraction from C5′–H by S•− produces the cPu via cyclization of the C5′ radical onto C8, forming a new covalent bond, C5′–C8. The unexpected formation of 8-oxo-Pu should be mechanistically more complex. We propose that an S•− (coupled with H+) adds to the base rings, followed by the elimination of HS− to form the corresponding radical cation; subsequent reactions with H2O and radical disproportionation with another S•− lead to 8-oxo-Pu. A comparison of S•− with the available literature data for HO• reactivity towards ct-DNA in de-oxygenated aqueous solutions is also presented. Before the present findings, cPu lesions were attributed exclusively to HO• reactivity toward ct-DNA. The reaction of the thiyl radical (HOCH2CH2S•) with ct-DNA was also investigated, yielding results similar to those of S•− obtained under comparable experimental conditions. Our results contributed to a better understanding of DNA damage induced by reactive sulfur species (RSS), particularly the formation of purine lesions and the relative abundance of cPu versus 8-oxo-Pu.
Full article
(This article belongs to the Section Molecular Biomarkers)
►▼
Show Figures

Graphical abstract
Open AccessReview
Microenvironmental Drivers of Bone Disease in Multiple Myeloma: Oxidative Stress, Sterile Inflammation, Autophagy–Lysosomal Remodeling, and the Iron–Lipid Peroxidation Axis
by
Maria Elisa Nasso, Adele Bottaro, Manlio Fazio, Fabio Stagno, Sebastiano Gangemi and Alessandro Allegra
Biomolecules 2026, 16(5), 710; https://doi.org/10.3390/biom16050710 (registering DOI) - 12 May 2026
Abstract
Multiple myeloma profoundly remodels the bone marrow microenvironment, causing osteolytic bone disease through a persistent uncoupling of bone resorption and formation. Beyond the canonical roles of the receptor activator of nuclear factor kappa-B ligand/receptor activator of nuclear factor kappa-B/osteoprotegerin triad and Wnt antagonism,
[...] Read more.
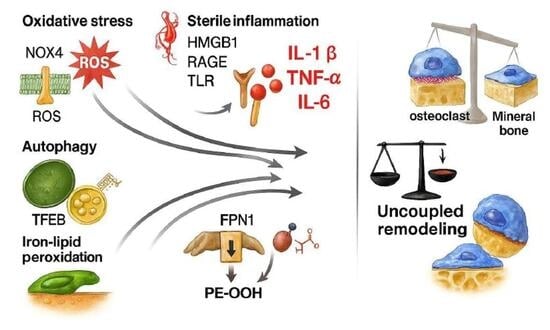
Multiple myeloma profoundly remodels the bone marrow microenvironment, causing osteolytic bone disease through a persistent uncoupling of bone resorption and formation. Beyond the canonical roles of the receptor activator of nuclear factor kappa-B ligand/receptor activator of nuclear factor kappa-B/osteoprotegerin triad and Wnt antagonism, three interdependent stress programs orchestrate the osteolytic niche. These include oxidative stress driven by mitochondrial and nicotinamide adenine dinucleotide phosphate oxidase-derived reactive oxygen species; sterile inflammation sustained by damage-associated molecular patterns, pattern-recognition receptors, and pro-inflammatory cytokines; and autophagy–lysosomal remodeling governed by transcription factor EB and the coordinated lysosomal expression and regulation network. These axes intersect with iron handling and lipid peroxidation to regulate sensitivity to ferroptotic cell death, thereby shaping osteoclast priming, osteoblast suppression, and matrix turnover. Building on these mechanistic insights, we outline a translational framework that aligns standardized bone turnover markers of formation and resorption with composite panels of oxidative and nitrosative stress. This framework also integrates modern imaging to capture structural injury and metabolically active marrow disease. We further propose a therapeutic roadmap layered on antiresorptive foundations that targets selective inhibition of nicotinamide adenine dinucleotide phosphate oxidase 4 and calibrated modulation of nuclear factor erythroid 2–related factor 2, disrupts damage-associated molecular pattern and cytokine circuits, and applies lineage- and timing-specific tuning of autophagy together with restoration of ferroportin-1 or iron chelation. This integrated strategy is designed to recouple bone remodeling and improve clinically meaningful skeletal outcomes in multiple myeloma.
Full article
(This article belongs to the Special Issue Molecular Factors Involved in Healthy and Diseased States of Skeletal Muscle: Overview and Perspectives)
►▼
Show Figures
Graphical abstract
Open AccessReview
Mechanism and Therapeutic Potential of Viral Mimicry in Cancer Immunotherapy
by
Alisha Pearl Kirkland, Mahek Shah and Charles Spruck
Biomolecules 2026, 16(5), 709; https://doi.org/10.3390/biom16050709 (registering DOI) - 12 May 2026
Abstract
►▼
Show Figures
Cancer immunotherapy has transformed oncology by harnessing the immune system to recognize and eliminate malignant cells. However, many cancers exhibit limited or variable responses to this class of treatment due to insufficient antigen presentation and impaired interferon (IFN) signaling, creating an immunologically “cold”
[...] Read more.
Cancer immunotherapy has transformed oncology by harnessing the immune system to recognize and eliminate malignant cells. However, many cancers exhibit limited or variable responses to this class of treatment due to insufficient antigen presentation and impaired interferon (IFN) signaling, creating an immunologically “cold” tumor microenvironment (TME) characterized by poor immune cell infiltration and treatment resistance. Viral mimicry has emerged as a therapeutic strategy to overcome these limitations by reactivating innate antiviral pathways within tumor cells. Viral mimicry occurs through the reactivation of endogenous retroviruses (ERVs) and other retrotransposons (e.g., LINE-1), which subsequently stimulate downstream nucleic acid sensing pathways. The resulting type I/III IFN responses restore antigen presentation and attract cytotoxic immune cells, sensitizing resistant tumors to immunotherapy. However, systemic stimulation of these pathways can trigger context-dependent inflammation and adaptive resistance, highlighting the need for temporal and spatial control. In this review, we examine the mechanistic foundation and clinical trajectory of viral mimicry, with an emphasis on its potential integration with established treatments and engineered immune cell platforms. By identifying the molecular and clinical gaps, viral mimicry can be harnessed to enhance tumor-specific immune activation and overcome treatment resistance in cancer immunotherapy.
Full article
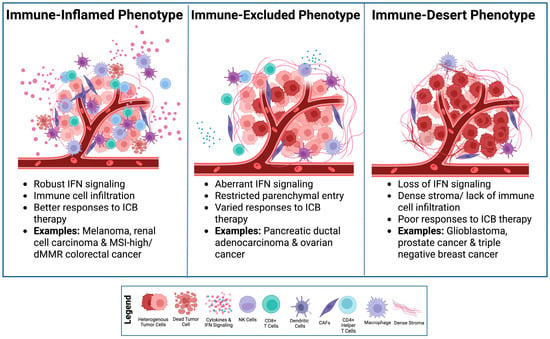
Figure 1
Open AccessArticle
Gut-Derived Lipid Mediators Orchestrate Ovarian Metabolic Homeostasis and Clutch Persistence in Aging Laying Hens via the PLA2G6-ALOX15B-AGPAT3 Axis
by
Xin Li, Xiaoliang Wang, Xia Cai, Qiang Meng, Yanyan Sun, Changsuo Yang and Junfeng Yao
Biomolecules 2026, 16(5), 708; https://doi.org/10.3390/biom16050708 (registering DOI) - 11 May 2026
Abstract
Clutch persistence, defined as the ability to sustain consecutive egg-laying cycles, is a pivotal determinant of profitability in the poultry industry, particularly for aging laying hens (≥65 weeks). However, the molecular mechanisms governing this trait remain elusive, largely due to the traditional “ovary-centric”
[...] Read more.
Clutch persistence, defined as the ability to sustain consecutive egg-laying cycles, is a pivotal determinant of profitability in the poultry industry, particularly for aging laying hens (≥65 weeks). However, the molecular mechanisms governing this trait remain elusive, largely due to the traditional “ovary-centric” paradigm that overlooks systemic regulation by the gut microbiota. To address this knowledge gap, the present study aimed to dissect the comprehensive regulatory network governing clutch persistence using integrated multi-omics analyses. A total of 20 sixty-five-week-old Rhode Island Red (RIR) laying hens with cumulative egg production exceeding 300 eggs but distinct clutch persistence were stratified into a high-clutch persistence group (HCP, ≥25 clutches, n = 10) and a low-clutch persistence group (LCPLCP, ≤15 clutches, n = 10). Multi-omics profiling, including ovarian transcriptomics, proteomics, and metabolomics; serum metabolomics; and cecal microbiota 16S rRNA sequencing was performed. Data integration and association mining were conducted via Spearman correlation analysis with stringent thresholds (r > 0.6, p < 0.01). Integrated analyses revealed a “gut–ovary axis” regulatory model mediated by a lipid mediator network, operating through a three-tiered mechanism: (1) Gut Initiation: The HCP group exhibited enriched cecal γ-Proteobacteria, which promoted biosynthesis of lipid precursors. (2) Serum Transport: Key serum lipid mediators, most notably LysoPC (22:6) (VIP = 4.5) and cholesterol ester CE (20:4), served as critical carriers transducing gut-derived signals to the ovary. (3) Ovarian Execution: These lipid signals activated a core ovarian metabolic pathway centered on the PLA2G6-ALOX15B-AGPAT3 axis, which coordinated follicular development and ovulation by supplying steroid hormone synthesis substrates, exerting anti-inflammatory effects, and stabilizing membrane structures. Collectively, this study demonstrates that gut microbiota modulates clutch persistence in aging laying hens via lipid mediators, orchestrating a systemic “gut–serum–ovary” regulatory cascade. These findings provide a novel molecular framework for extending the economic egg-laying cycle through the targeted manipulation of intestinal microbiota or serum lipid metabolism.
Full article
(This article belongs to the Section Lipids)
Open AccessArticle
GPIHBP1 as a Biomarker of Diabetic Polyneuropathy and Vascular Complications in Type 2 Diabetes Mellitus
by
Savelia Yordanova, Antoaneta Gateva, Diana Nikolova, Julieta Hristova and Zdravko Kamenov
Biomolecules 2026, 16(5), 707; https://doi.org/10.3390/biom16050707 (registering DOI) - 11 May 2026
Abstract
Background: Diabetic neuropathy is one of the most common chronic complications of diabetes mellitus and could lead to foot ulcerations, lower-limb amputations, increased mortality and reduced quality of life. This study examines the level of GPIHBP1 to assess its diagnostic and prognostic values
[...] Read more.
Background: Diabetic neuropathy is one of the most common chronic complications of diabetes mellitus and could lead to foot ulcerations, lower-limb amputations, increased mortality and reduced quality of life. This study examines the level of GPIHBP1 to assess its diagnostic and prognostic values across the metabolic continuum. Methods: This is an observational monocentric study, including 160 patients with type 2 diabetes mellitus, obesity without carbohydrate metabolism disorders and healthy controls. Clinical data and laboratory results were collected, and serum levels of GPIHBP1 were measured using an ELISA. The presence of DPN for the diabetes group was assessed using corneal confocal microscopy and NDS. The statistical analyses included t-tests, Pearson’s correlation analysis, and ROC analysis to explore associations and the predictive values of the biomarker. Results: The GPIHBP1 levels increased progressively, with the lowest levels observed in the control group, higher levels in patients with obesity, and the highest levels in those with diabetes mellitus. Higher GPIHBP1 levels were observed in patients with peripheral diabetic neuropathy compared to those without. GPIHBP1 demonstrated moderate discriminative performance for the presence of diabetes, diabetic neuropathy and nephropathy. GPIHBP1 levels were also associated with renal function parameters and markers of vascular involvement. After adjustment for confounders, including estimated glomerular filtration rate (eGFR), the association between GPIHBP1 and diabetic neuropathy remained statistically significant although attenuated. Higher levels were observed in patients with coronary artery disease, and a positive correlation was established with mean IMT and sudomotor dysfunction score. Conclusions: Circulating GPIHBP1 levels are associated with diabetes mellitus and its micro- and macrovascular complications, particularly diabetic neuropathy. Its measurement could enhance early diagnosis and personalized management of T2DM, and, while these findings support a potential role of GPIHBP1 as a biomarker of metabolic and vascular dysfunction, its clinical utility requires confirmation in longitudinal studies.
Full article
(This article belongs to the Section Molecular Biomarkers)
►▼
Show Figures
Figure 1
Open AccessArticle
Immunohistochemical Evaluation of Integrin β6 Expression in Triple-Negative Breast Cancer as a Predictive Biomarker for Therapeutic and Diagnostic Radionuclides
by
Muin Tuffaha, Wael Hananeh, Nikola Bangemann, Amro Tuffaha and Michael Starke
Biomolecules 2026, 16(5), 706; https://doi.org/10.3390/biom16050706 (registering DOI) - 11 May 2026
Abstract
Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype associated with limited therapeutic options and poor clinical outcomes. The aim of this research is to assess the prevalence, intensity, and distribution of integrin αvβ6 expression in TNBC using immunohistochemistry and to evaluate
[...] Read more.
Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype associated with limited therapeutic options and poor clinical outcomes. The aim of this research is to assess the prevalence, intensity, and distribution of integrin αvβ6 expression in TNBC using immunohistochemistry and to evaluate its potential as a predictive biomarker for αvβ6-targeted radionuclide therapy and other αvβ6-targeted theranostic approaches. Immunohistochemical analysis of integrin αvβ6 was performed on formalin-fixed, paraffin- embedded tumor samples from 48 patients with histologically confirmed TNBC. Staining intensity and the proportion of positive tumor cells were assessed using a semi-quantitative scoring system, and expression patterns were analyzed with regard to cellular localization and intratumoral heterogeneity. Moderate to strong integrin αvβ6 expression was observed in 43.8% of cases, with strong expression (≥50% of tumor cells) present in 25%. Expression was predominantly membranous, with occasional cytoplasmic staining, and demonstrated marked inter- and intratumoral heterogeneity. Integrin αvβ6 is frequently expressed in TNBC and represents a promising biomarker for patient selection in αvβ6-targeted radionuclide imaging and therapy. These findings provide a strong biological rationale for the clinical translation of integrin-targeted radioligands and support the development of personalized radiotheranostic strategies in TNBC.
Full article
(This article belongs to the Section Molecular Medicine)
►▼
Show Figures
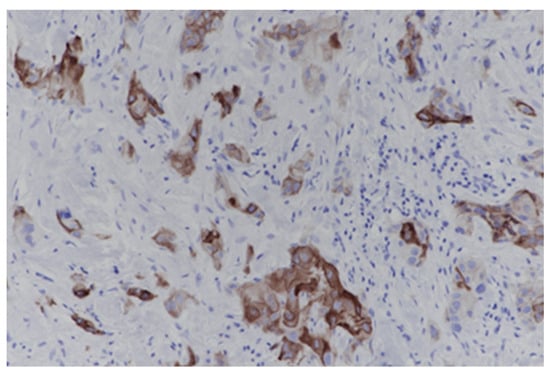
Figure 1
Open AccessReview
Plant-Derived Nanovesicles: A Comprehensive Review from Isolation to Clinical Translation—Unlocking Natural Nanocarriers for Biomedical Applications
by
Xinyan Wang, Chenchen Yuan and Rong Lu
Biomolecules 2026, 16(5), 705; https://doi.org/10.3390/biom16050705 (registering DOI) - 11 May 2026
Abstract
Plant-derived nanovesicles (PDNVs) are a class of nanoscale vesicles derived from plant tissues; they are particles with a lipid bilayer and no ability to replicate autonomously. As a type of bioactive natural nanocarrier, they demonstrate immense potential for application in 21st-century nanomedicine, skincare
[...] Read more.
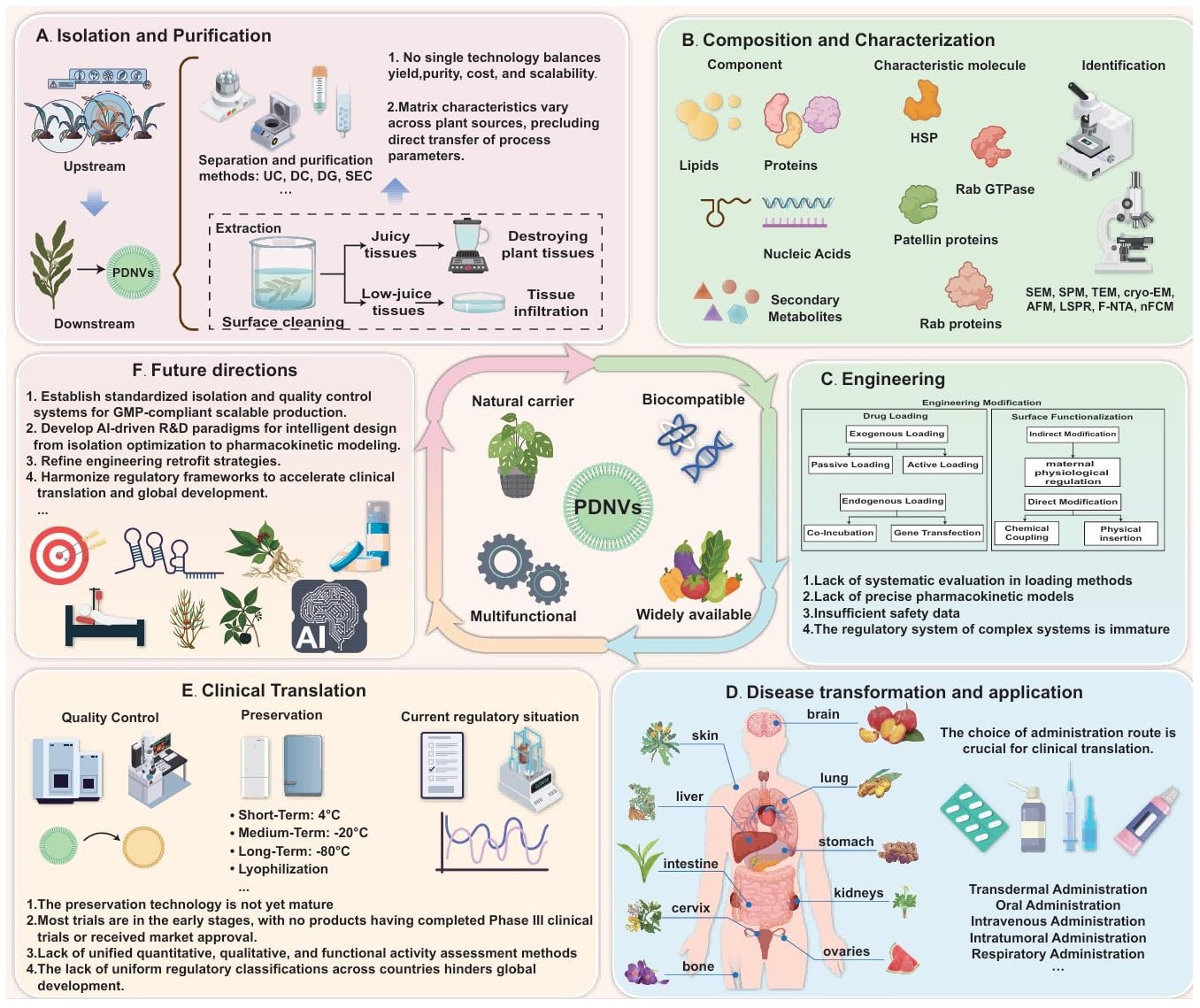
Plant-derived nanovesicles (PDNVs) are a class of nanoscale vesicles derived from plant tissues; they are particles with a lipid bilayer and no ability to replicate autonomously. As a type of bioactive natural nanocarrier, they demonstrate immense potential for application in 21st-century nanomedicine, skincare and nutritional health, owing to their excellent biocompatibility, low immunogenicity and targeted delivery capabilities. However, the clinical translation of PDNVs still faces key bottlenecks, including low extraction efficiency, complex purification processes, and immature engineering modification techniques. Compared to the wealth of systematic reviews in the field of Mammalian Extracellular Vesicles (M-EVs), research on PDNVs still lacks a comprehensive exposition of its multifaceted research progress. This review endeavours to comprehensively summarise the shortcomings over the last 60 years regarding PDNV purification processes, research progress, composition and characterisation, engineering modifications, functional mechanisms, clinical translation and market regulation. It discusses the feasibility of innovative approaches such as AI deep learning technologies, interdisciplinary integration and cross-application, and outlines the latest frontiers in PDNV research. It provides comprehensive and reliable reference material for future research and application strategies regarding PDNVs, offering theoretical support and practical guidance to overcome barriers to their industrialisation. This will facilitate the transition from limited laboratory research to clinical application and drive technological innovation in the next generation of naturally derived nanomedicines.
Full article
(This article belongs to the Section Natural and Bio-derived Molecules)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Exploring Replicative Senescence and Oxidative Stress-Induced Remodelling of Mitochondrial-Associated Membranes in Human Skin Fibroblasts
by
Anne-Laure Bulteau, Gallic Beauchef, Stéphanie Chanon, Aurélie Vieille-Marchis, Julien Chlasta, Gaël Runel, Juliette Sage, Tanesha Naiken, Lauren Sobilo, Elodie Bossard, Lorene Gourguillon, Carine Nizard, Karl Pays, Laurence Canaple and Beatrice Morio
Biomolecules 2026, 16(5), 704; https://doi.org/10.3390/biom16050704 (registering DOI) - 11 May 2026
Abstract
(1) Background: Calcium transfer between the endoplasmic reticulum (ER) and mitochondria through the IP3R–VDAC1 complex at mitochondria-associated ER membranes (MAMs) is essential for cellular homeostasis. Alterations in this signalling axis have been implicated in ageing and cellular senescence. (2) Methods: We developed an
[...] Read more.
(1) Background: Calcium transfer between the endoplasmic reticulum (ER) and mitochondria through the IP3R–VDAC1 complex at mitochondria-associated ER membranes (MAMs) is essential for cellular homeostasis. Alterations in this signalling axis have been implicated in ageing and cellular senescence. (2) Methods: We developed an in vitro human dermal fibroblast (HDF) model combining replicative senescence and acute oxidative stress to investigate the role of ER–mitochondria coupling in skin ageing and to enable biomolecule screening. (3) Results: In situ proximity ligation assays revealed that replicative senescence significantly increased the number of VDAC1/IP3R complexes per cell (+85% and +72%, p < 0.01), together with elevated cellular reactive oxygen species (+47% and +74%, p < 0.05). Consistently, acute oxidative stress (50 µM t-BHP, 30 min) rapidly increased VDAC1/IP3R complexes (+48%, p < 0.001) and intra-mitochondrial calcium levels (+19%, p < 0.001). These effects persisted for 24 h post-treatment and were associated with impaired mitochondrial function (−27% in the Bioenergetic Health Index, p < 0.05). We also established a flexibility index capturing both acute and long-term adaptations and detecting the protective effects of an orchid extract. (4) Conclusions: ER–mitochondria coupling disruption via the IP3R–VDAC1 complex may contribute to oxidative stress-induced senescence and represent a key mechanism in extrinsic skin ageing.
Full article
(This article belongs to the Special Issue Mitochondrial ROS in Health and Disease: 2nd Edition)

Journal Menu
► ▼ Journal Menu-
- Biomolecules Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
14 May 2026
Meet Us at the ncRNA2026: From Molecular Mechanisms to Clinical Impact, 24–26 June 2026, Leuven, Belgium
Meet Us at the ncRNA2026: From Molecular Mechanisms to Clinical Impact, 24–26 June 2026, Leuven, Belgium

13 May 2026
Meet Us at the American Society for Virology 45th Annual Meeting (ASV), 27–30 July 2026, Minneapolis, USA
Meet Us at the American Society for Virology 45th Annual Meeting (ASV), 27–30 July 2026, Minneapolis, USA

Topics
Topic in
Chemistry, Molecules, IJMS, Biomolecules, Inorganics
Metal Ions in Health and Diseases: Current Progress and Future Challenges
Topic Editors: Massimiliano F. Peana, Carlo Santini, Maura PelleiDeadline: 31 May 2026
Topic in
Biomedicines, Biomolecules, Cancers, Cells, Hematology Reports, IJMS
Advances in Molecular Pathogenesis and Targeted Therapies for Multiple Myeloma
Topic Editors: Chung Hoow Kok, Cindy H. S. Lee, Claudio CerchioneDeadline: 3 August 2026
Topic in
Biomolecules, Chemistry, IJMS, Molecules, Pharmaceuticals
Enzymes and Enzyme Inhibitors in Drug Research
Topic Editors: Athina Geronikaki, Cosimo D. Altomare, Maria Stefania SinicropiDeadline: 11 September 2026
Topic in
Biomedicines, IJMS, Sci. Pharm., Molecules, Future Pharmacology, Biomolecules
Natural Products and Drug Discovery—2nd Edition
Topic Editors: Sonia Piacente, Marta MenegazziDeadline: 30 September 2026

Conferences
Special Issues
Special Issue in
Biomolecules
The Role of Vascular Dysfunction in Neuronal Degeneration and Cognitive Impairment (2nd Edition)
Guest Editor: David LominadzeDeadline: 15 May 2026
Special Issue in
Biomolecules
Protein Self-Assembly in Diseases and Function
Guest Editors: Anoop Arunagiri, Srivastav Ranganathan, Maria del Carmen Fernandez RamirezDeadline: 15 May 2026
Special Issue in
Biomolecules
Cancer Biology: Machine Learning and Bioinformatics
Guest Editors: Annalisa Santucci, Ottavia Spiga, Anna VisibelliDeadline: 15 May 2026
Special Issue in
Biomolecules
Neutrophils in Health and Disease: From Molecular Mechanisms to Therapies
Guest Editors: Xin Xu, Lawrence RasmussenDeadline: 16 May 2026
Topical Collections
Topical Collection in
Biomolecules
Feature Papers in Section 'Molecular Medicine'
Collection Editors: Fabio Di Domenico, Chiara Lanzillotta
Topical Collection in
Biomolecules
In Silico Drug Design for GPCRs: Big Data for Small Molecule Discovery
Collection Editor: Vsevolod Katritch
Topical Collection in
Biomolecules
TGF-Beta Signaling in Tissue Fibrosis and Cancer
Collection Editors: Paul J. Higgins, Rohan Samarakoon
Topical Collection in
Biomolecules
Feature Papers in Molecular Genetics
Collection Editor: Jürg Bähler