Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS), Brazilian Society for Virology (BSV) and Global Virus Network (GVN) are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Virology/Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.2 days after submission; acceptance to publication is undertaken in 2.7 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journal: Zoonotic Diseases.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
3.5 (2024);
5-Year Impact Factor:
3.7 (2024)
Latest Articles
Genomic Characterization and Evolutionary Dynamics of SARS-CoV-2 Lineage NB.1.8.1 in Thailand, 2025
Viruses 2026, 18(4), 450; https://doi.org/10.3390/v18040450 (registering DOI) - 8 Apr 2026
Abstract
SARS-CoV-2 continues to cause recurrent waves in the post-pandemic period, yet genomic data from Southeast Asia remain limited for several emerging Omicron lineages, including NB.1.8.1. In this study, routine acute respiratory infection (ARI) surveillance performed in Bangkok, Thailand, from January to December 2025
[...] Read more.
SARS-CoV-2 continues to cause recurrent waves in the post-pandemic period, yet genomic data from Southeast Asia remain limited for several emerging Omicron lineages, including NB.1.8.1. In this study, routine acute respiratory infection (ARI) surveillance performed in Bangkok, Thailand, from January to December 2025 was integrated with real-time RT-PCR testing and complete spike-gene sequencing for lineage assignment and evolutionary analysis. Among 4756 ARI specimens, 473 (9.9%) tested positive for SARS-CoV-2. Positivity increased in late April, peaked in May (epidemiological week 21; 58.4%), and declined through late June. Lineage typing was successful for 165/473 positive samples (34.9%), identifying 16 Pango lineages. Early 2025 showed heterogeneous circulation, including XEC- and XEC.8-related lineages, whereas NB.1.8.1 predominated during the main wave, accounting for 92.4% of typed cases in May and 89.8% in June. No recombination signals meeting predefined criteria were detected in the spike dataset. The mean spike substitution rate was estimated at 1.11 × 10−3 substitutions/site/year (95% HPD, 9.13 × 10−4–1.31 × 10−3), and the major Thai-containing NB.1.8.1 clade had an estimated tMRCA of 17 July 2024. These findings show that routine ARI surveillance combined with spike-based genomics can provide timely insights into SARS-CoV-2 circulation, lineage replacement, and ongoing viral evolution in Thailand.
Full article
(This article belongs to the Special Issue Molecular Epidemiology of SARS-CoV-2, 4th Edition)
Open AccessBrief Report
Aujeszky’s Disease in a Grey Wolf (Canis lupus) in Poland
by
Wojciech Wójcik, Anna Didkowska, Blanka Orłowska, Sabina Nowak, Bartosz Sell, Krzysztof Anusz, Florian Pfaff and Bernd Hoffmann
Viruses 2026, 18(4), 449; https://doi.org/10.3390/v18040449 (registering DOI) - 8 Apr 2026
Abstract
Aujeszky’s disease (AD), caused by suid herpesvirus 1 (pseudorabies virus, PRV), is a highly contagious infection primarily affecting swine, with wild boars serving as an important reservoir in Europe. Spillover infections in non-suid species, including carnivores, are rare but typically fatal and of
[...] Read more.
Aujeszky’s disease (AD), caused by suid herpesvirus 1 (pseudorabies virus, PRV), is a highly contagious infection primarily affecting swine, with wild boars serving as an important reservoir in Europe. Spillover infections in non-suid species, including carnivores, are rare but typically fatal and of epidemiological concern. This study presents the first case of AD in a grey wolf (Canis lupus) in Central Europe with genomic characterization. The 8-month-old wolf was found in the Carpathians (SE Poland), moribund with acute neurological signs, and euthanized for animal welfare reasons. Necropsy revealed no pathognomonic gross lesions. Molecular analyses of tissues confirmed the presence of PRV DNA using real-time PCR, and virus isolation was successful. Genomic analysis revealed that the PRV isolate clustered within genotype I, the predominant circulating genotype in Europe. However, due to the limited availability of reference PRV genome sequences from European wildlife, the precise geographic origin and transmission pathways of this strain could not be fully resolved. In the presented case, wild boars were considered a possible source of infection. This highlights the potential for PRV transmission to apex predators. This study emphasizes the importance of systematic surveillance of PRV in wildlife and the need for expanded genomic databases of PRV strains. Full-genome sequencing is crucial for improving the understanding of PRV transmission.
Full article
(This article belongs to the Special Issue Herpesviruses and Associated Diseases, 2nd Edition)
Open AccessArticle
ScRNA-Seq and BCR Analysis of Murine Immune Responses to Inactivated DHAV-1 as a Model Antigen
by
Yaru Fan, Saisai Zhao, Yafei Qin, Guocheng Liu, Linyu Cui, Siming Zhu, Youxiang Diao, Dalin He and Yi Tang
Viruses 2026, 18(4), 448; https://doi.org/10.3390/v18040448 - 8 Apr 2026
Abstract
Currently, the B-cell response patterns induced by viral antigens in avian disease models and their detailed immunological characteristics still require comprehensive elucidation at the single-cell level. In this study, we employed single-cell sequencing (scRNA-seq) and B cell library technology to conduct an in-depth
[...] Read more.
Currently, the B-cell response patterns induced by viral antigens in avian disease models and their detailed immunological characteristics still require comprehensive elucidation at the single-cell level. In this study, we employed single-cell sequencing (scRNA-seq) and B cell library technology to conduct an in-depth analysis of B cells in the spleens of mice with inactivated duck hepatitis A virus type 1 (DHAV-1) as model antigen. This study aimed to investigate the immunological characteristics of the virus antigen in the mouse model and characteristics of B-Cell Receptors. The results showed that the DHAV-1 group had distinct changes in splenic B cell subset counts, proportions, and intercellular communication. Additionally, an increased trend in communication strength between Gm26917+B and Gm11837+B cells was observed, with enriched expression of C-X-C motif chemokine ligand (CXCL) and lymphotoxin (LT) detected in the DHAV-1 group. Furthermore, the DHAV-1 group exhibited a prominent combination of the IGHV1 family and IGHV3-1/IGHJ3 in the heavy (H) chain variable region. Compared with the CK group (negative control group), the amino acid sequence length and diversity of the CDR3 region in the DHAV-1 group exhibited a decreasing trend. In summary, our findings characterize the immunological features of splenic B cells in mice after immunization with inactivated DHAV-1, and provide a preliminary characterization of DHAV-1-induced B cell transcriptional states and BCR repertoire features, generating testable hypotheses for subsequent mechanistic investigations of B cell-mediated immune responses to viral antigens.
Full article
(This article belongs to the Special Issue Humoral Immune Response to Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
Characterization of a Goose-Origin Avian Orthoreovirus with Interferon Suppression Activity
by
Yijia Liu, Yong Li, Yingxuan Xie, Mei Wang, Boxuan Yin, Changyan Li, Lilin Zhang, Deping Hua, Junwei Liu, Xintian Zheng and Jinhai Huang
Viruses 2026, 18(4), 447; https://doi.org/10.3390/v18040447 - 8 Apr 2026
Abstract
The emergence of variant strains of Avian orthoreovirus (ARV) has caused substantial economic losses in the poultry industry worldwide, but the molecular features of goose-origin strains and viral transmission among different avian species remain poorly understood. Here, we describe a goose-origin avian orthoreovirus
[...] Read more.
The emergence of variant strains of Avian orthoreovirus (ARV) has caused substantial economic losses in the poultry industry worldwide, but the molecular features of goose-origin strains and viral transmission among different avian species remain poorly understood. Here, we describe a goose-origin avian orthoreovirus strain, SD0407, associated with growth retardation and joint swelling. Complete genome analysis identified ten double-stranded RNA segments. Sequence comparison indicated that SD0407 is closely related to previously reported duck-origin reovirus strains. Phylogenetic and recombination analyses showed that most segments clustered with duck-origin strains, indicating close genetic relatedness among waterfowl-origin orthoreoviruses. Sequence and structural analysis of the σC attachment protein revealed ten unique amino acid substitutions, including D250 within the DE-loop region involved in receptor-binding. Molecular docking suggested that σC interacts with the conserved AnxA2-S100A10 heterotetrameric receptor complex, providing a possible structural basis for receptor compatibility across avian species. Although SD0407 replicated efficiently in goose embryo fibroblasts, it did not induce expression of type I, II or III interferons. Transcriptome profiling revealed weak activation of innate immune signaling and downregulation of metabolic and cytoskeletal genes, consistent with effective suppression of antiviral responses. These findings demonstrate that SD0407 combines structural variability with immune evasion to enhance host adaptability and underscore the importance of sustained ARV surveillance in waterfowl populations.
Full article
(This article belongs to the Special Issue Avian Reovirus)
►▼
Show Figures

Figure 1
Open AccessArticle
Induction Cisplatin, Docetaxel, and 5-Fluorouracil Versus Platinum Doublet in Human Papillomavirus-Associated Oropharyngeal Cancer
by
Ashwin Ganti, Patrick W. McGarrah, Harry Fuentes Bayne, Casey Fazer-Posorske, Binav Baral, Daniel J. Ma, Scott C. Lester, David M. Routman, Michelle A. Neben-Wittich, Jessica M. Wilson, Daniel L. Price, Eric J. Moore, Kendall K. Tasche, Katie M. Van Abel, Linda X. Yin, Nathan R. Foster and Katharine Price
Viruses 2026, 18(4), 446; https://doi.org/10.3390/v18040446 - 8 Apr 2026
Abstract
Management of locoregionally advanced human papillomavirus-positive oropharyngeal squamous cell carcinoma (HPV(+) OPSCC) can include induction chemotherapy followed by definitive chemoradiation. The standard induction regimen of cisplatin, docetaxel, and 5-fluorouracil (TPF) is associated with high toxicity. Given the chemosensitivity of HPV(+) OPSCC, platinum doublets
[...] Read more.
Management of locoregionally advanced human papillomavirus-positive oropharyngeal squamous cell carcinoma (HPV(+) OPSCC) can include induction chemotherapy followed by definitive chemoradiation. The standard induction regimen of cisplatin, docetaxel, and 5-fluorouracil (TPF) is associated with high toxicity. Given the chemosensitivity of HPV(+) OPSCC, platinum doublets are frequently used as induction therapy with potentially less toxicity. This retrospective study aimed to compare outcomes between treatment-naive HPV(+) OPSCC patients receiving induction TPF and those receiving an induction platinum doublet. Data collected included tumor characteristics, response after chemoradiation, hospitalization rates, and overall survival (OS). Fifty-five patients (18 TPF and 37 platinum doublet) were included. There was no significant difference in response after completion of definitive chemoradiation (TPF: CR 83.3%, PR 5.6%, progression or metastasis 11.1% vs. platinum doublet: CR 75.7%, PR 16.2%, progression or metastasis 8.1%; p = 0.5241). There were also no differences in hospitalizations for adverse events (38.9% in TPF vs. 40.5% in platinum doublet, p = 0.907) or recurrence (11.1% in TPF vs. 2.7% in platinum doublet, p = 0.198). The 5-year OS was 84.6% in the TPF group and 81.5% in the platinum doublet group (p = 0.581). Induction platinum doublet regimens offer comparable OS, response, and hospitalization rates to TPF in locally advanced HPV(+) OPSCC. Induction with a platinum doublet may be a viable de-escalation strategy for patients who are not candidates for TPF.
Full article
(This article belongs to the Special Issue Advancements in Immunotherapy for Human Papillomavirus)
►▼
Show Figures
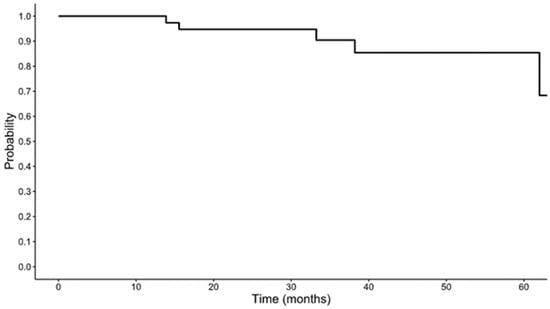
Figure 1
Open AccessArticle
A Newly Established ELISA for the Surveillance of Rift Valley Fever in Dromedary Camels and Their Owners, Kenya 2018
by
Shannon L. M. Whitmer, Jessica Rowland, Emir Talundzic, Deborah Cannon, Aridth Gibbons, Cynthia Ombok, Jennifer L. Harcourt, Natalie J. Thornburg, Clayton Onyango, Peninah Munyua, Elizabeth Hunsperger, Isaac Ngere, M. Kariuki Njenga, Caroline Ochieng, Mathew Muturi, Joel M. Montgomery, Marc-Alain Widdowson and John D. Klena
Viruses 2026, 18(4), 445; https://doi.org/10.3390/v18040445 - 8 Apr 2026
Abstract
In 2024 Kenya had a population of 4.78 million camels that contributed to the livelihoods of pastoralist communities in northern Kenya. Previous studies in Kenya, Saudi Arabia and eastern Africa demonstrated high seroprevalence of Middle East respiratory syndrome coronavirus (MERS-CoV)-specific antibodies in dromedary
[...] Read more.
In 2024 Kenya had a population of 4.78 million camels that contributed to the livelihoods of pastoralist communities in northern Kenya. Previous studies in Kenya, Saudi Arabia and eastern Africa demonstrated high seroprevalence of Middle East respiratory syndrome coronavirus (MERS-CoV)-specific antibodies in dromedary camels, as well as sporadic transmission of MERS-CoV from camels to humans. Based on the MERS-CoV data and the very close contact between owners and their camels in northern Kenya, we speculated that camels may also transmit other zoonotic viruses, such as Rift Valley fever virus (RVFV). In this study, 493 camel and 197 human sera were collected in Marsabit, Kenya, through a cross-sectional survey in 2018 and analyzed for the presence of RVFV IgG antibodies using a laboratory-developed indirect enzyme-linked immunosorbent assay (ELISA). Overall, 15.6% of camels and 7.6% of humans were RVFV IgG-positive; IgG-positive camels were predominantly females in large population herds and IgG-positive humans were engaged in farming-related activities and were greater than 18 years old. Of the eight location groups sampled, two had high camel (site 2 and site 6) and two had high human (site 5 and site 6) RVFV seropositivity rates. These data suggest that camelids, such as dromedary camels, may serve as amplifying hosts for vector-borne zoonotic diseases, such as RVFV, and that humans with frequent farming and camel meat, milk, or camel product contact may have increased risk for RVFV exposure or infection.
Full article
(This article belongs to the Special Issue Rift Valley Fever Virus: New Insights into a One Health Archetype)
►▼
Show Figures

Figure 1
Open AccessArticle
Neomycin Exhibits Immunomodulatory and Antiviral Activity Against Influenza B Virus
by
Ekaterina Romanovskaya-Romanko, Marina Plotnikova, Anna-Polina Shurygina, Marina Shuklina, Sergey Klotchenko, Zhanna Buzitskaya, Dmitry Lioznov and Marina Stukova
Viruses 2026, 18(4), 444; https://doi.org/10.3390/v18040444 - 7 Apr 2026
Abstract
Viral infections remain a global public health challenge. Stimulating the innate immune system is a potent therapeutic strategy that promotes pathogen clearance, directly impacting disease severity and clinical outcomes. Interferons and interferon-stimulated genes (ISGs) are critical components of this antiviral defense system. Neomycin,
[...] Read more.
Viral infections remain a global public health challenge. Stimulating the innate immune system is a potent therapeutic strategy that promotes pathogen clearance, directly impacting disease severity and clinical outcomes. Interferons and interferon-stimulated genes (ISGs) are critical components of this antiviral defense system. Neomycin, an aminoglycoside antibiotic, can induce ISG expression and help establish an antiviral state. In this study, we demonstrated that neomycin induces the production of pro-inflammatory cytokines (IL1β, TNFα, IL6, GM-CSF, and IFN-γ) in peripheral mononuclear blood cells (PBMCs) and activates key antiviral ISGs, including MxA, OAS1, and IRF7. The protein expression profiles elicited by neomycin were comparable to those induced by poly(I:C). Intranasal delivery of neomycin to CBA and BALB/c mice induced various ISGs in both the respiratory tract and splenic tissues. Prophylactic administration of neomycin significantly inhibited influenza B virus replication in the lung and nasal turbinates of CBA mice in a sublethal infection model. Overall, our data suggest that neomycin, when used prophylactically alone or combined with other antiviral strategies, shows considerable potential for the attenuation of influenza B virus infections.
Full article
(This article belongs to the Section Viral Immunology, Vaccines, and Antivirals)
►▼
Show Figures

Figure 1
Open AccessFeature PaperArticle
Metatranscriptomic Analysis of Tick Virome Diversity in Hebei Province, China
by
Minghao Geng, Xueqi Wang, Xiaoxia Huang, Yan Li, Yamei Wei, Yanan Cai, Jiandong Li, Caixiao Jiang, Wei Wu, Shiyou Liu, Nana Guo, Xinyang Zhang, Wentao Wu, Guangyue Han, Xu Han, Tiezhu Liu, Qi Li and Shiwen Wang
Viruses 2026, 18(4), 443; https://doi.org/10.3390/v18040443 - 7 Apr 2026
Abstract
Ticks serve as primary vectors for a wide array of RNA viruses, yet the diversity and distribution of tick-associated RNA viruses remain incompletely characterized in Hebei province. To address this gap, we conducted a systematic metatranscriptomic investigation of 986 ticks representing six species,
[...] Read more.
Ticks serve as primary vectors for a wide array of RNA viruses, yet the diversity and distribution of tick-associated RNA viruses remain incompletely characterized in Hebei province. To address this gap, we conducted a systematic metatranscriptomic investigation of 986 ticks representing six species, collected from the diverse ecological landscapes of Hebei Province in northern China. Our analysis recovered 25 complete or near-complete viral genomes spanning 12 families, including Phenuiviridae, Flaviviridae, and Nairoviridae. Of critical public health significance, we identified Severe Fever with Thrombocytopenia Syndrome Virus (SFTSV) in both Haemaphysalis longicornis and Dermacentor nuttalli. Phylogenetic reconstruction revealed marked geographic stratification where strains from the coastal plains clustered with the dominant Genotype F, while those from the mountainous north formed a characteristic and divergent lineage phylogenetically linked to isolates from Inner Mongolia. Furthermore, a novel viral agent provisionally named Zhangjiakou Hepacivirus was discovered in Haemaphysalis japonica. This virus shared less than 80% nucleotide identity with the rodent-associated Hepacivirus P, consistent with a rodent origin and possible cross-species transmission. Collectively, these findings reveal descriptive variation associated with vector identity, physiological status, and ecological context in shaping viral evolution and underscore the need for continuous metagenomic surveillance to mitigate emerging tick-borne disease risks within a One Health framework.
Full article
(This article belongs to the Special Issue Zoonotic and Vector-Borne Viral Diseases: 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
LL-37 Inhibits EV71 Infection by Upregulating STAC via the EGFR-ERK Signaling Pathway
by
Jiaqi Zhang, Hanlin Zhang, Yi Chen, Hanfei Liu, Shuhuang Peng, Jiwei Zhao, Zhe Luan, Yujian Zhang, Meng Dong, Wanzhu Jin and Gang Sun
Viruses 2026, 18(4), 442; https://doi.org/10.3390/v18040442 - 7 Apr 2026
Abstract
LL-37, a 37-amino acid human-derived antimicrobial peptide, was shown in our earlier clinical study to shorten the negative conversion time of the Omicron BA.5.1.3 variant of SARS-CoV-2. In this work, we investigated the broad mechanism of LL-37 by examining its inhibitory effect on
[...] Read more.
LL-37, a 37-amino acid human-derived antimicrobial peptide, was shown in our earlier clinical study to shorten the negative conversion time of the Omicron BA.5.1.3 variant of SARS-CoV-2. In this work, we investigated the broad mechanism of LL-37 by examining its inhibitory effect on non-enveloped virus Enterovirus 71 (EV71). LL-37 treatment dose-dependently reduced EV71 viral RNA abundance, suppressed virus-encoded protein expression, and decreased infectious titers, acting predominantly at a post-entry stage of the viral life cycle. Transcriptomic analysis revealed that the SH3 and cysteine-rich domain protein (Stac) was uniquely upregulated by LL-37 irrespective of EV71 infection. Short hairpin RNA (shRNA)-mediated Stac silencing significantly enhanced EV71 infection, while Stac overexpression markedly reduced it. Furthermore, we found that LL-37 activates the EGFR–ERK signaling pathway, leading to time-dependent upregulation of Stac expression. These findings uncover a novel host-directed mechanism by which LL-37 combats EV71 infection and suggests a potential therapeutic use of LL-37 against non-enveloped viral disease.
Full article
(This article belongs to the Special Issue Virus-Host Interactions: From Mechanisms to Therapeutics)
►▼
Show Figures

Figure 1
Open AccessArticle
Surveillance of Seasonal Influenza Viruses in Kazakhstan (2020–2025)
by
Tatyana Glebova, Nailya Klivleyeva, Nuray Ongarbayeva, Assem Baimukhametova, Nurbol Saktaganov, Mereke Kalkozhayeva, Kobey Karamendin, Indira Ibragimova, Madisha Sagatova, Aknur Mutaliyeva, Altynay Gabiden and Richard Webby
Viruses 2026, 18(4), 441; https://doi.org/10.3390/v18040441 - 7 Apr 2026
Abstract
Influenza viruses are significant causes of acute respiratory infections, often leading to severe health issues and mortality. These viruses undergo continuous mutations and genetic reassortments, resulting in annual epidemics and potential pandemics. The A(H3N2) strains exhibit high genetic and antigenic variability, that influence
[...] Read more.
Influenza viruses are significant causes of acute respiratory infections, often leading to severe health issues and mortality. These viruses undergo continuous mutations and genetic reassortments, resulting in annual epidemics and potential pandemics. The A(H3N2) strains exhibit high genetic and antigenic variability, that influence vaccine efficacy. This study aimed to assess the prevalence of influenza viruses, including A(H3N2) strains, in Kazakhstan during 2020–2025. The study used nasopharyngeal swab and serum samples obtained from patients. The presence of influenza virus antigens in nasopharyngeal swabs was analyzed using real-time polymerase chain reaction. The level of specific antibodies in the blood serum was determined using the hemagglutination inhibition reaction and enzyme-linked immunosorbent assay methods. Influenza A/H1N1, A/H3N2 and B viruses were diagnosed using real-time PCR. Antibodies to A/H1N1pdm09, A/H3N2 and B were detected in serological studies. Our studies revealed a trend toward seasonal patterns in influenza A viruses circulation. Therefore, it was established that the A/H3N2 strains dominated in Kazakhstan during the 2021–2022 and 2023–2024 epidemic seasons. The 2023–2024 strains belong to the specific genetic clade J.2 or 3C.2a1b.2a.2a.3a.1. These studies confirmed the role of influenza viruses in the etiology of respiratory infections and emphasized the need to continue monitoring their spread in Kazakhstan.
Full article
(This article belongs to the Section General Virology)
►▼
Show Figures
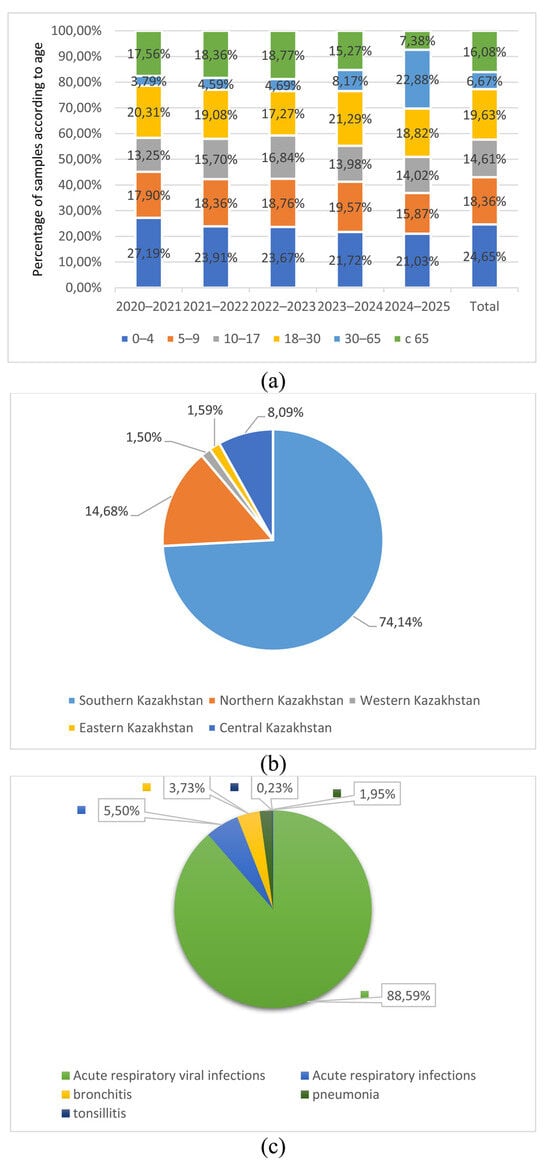
Figure 1
Open AccessArticle
Intranasal Administration of SARS-CoV-2 ORF8 Accessory Protein Increases Blood Pressure and Oxidative Stress in Different Tissues of Male BALB/c Mice
by
Karla A. Pavon-Martinez, Giovani Visoso-Carvajal, Rebeca Campi-Caballero, Jazmín García-Machorro and Judith Espinosa-Raya
Viruses 2026, 18(4), 440; https://doi.org/10.3390/v18040440 - 5 Apr 2026
Abstract
SARS-CoV-2 is the etiological agent responsible for COVID-19. While most research has focused on structural proteins, the accessory protein Open Reading Frame 8 (ORF8) has attracted attention for its role in immune evasion and the induction of a cytokine storm. Although the exact
[...] Read more.
SARS-CoV-2 is the etiological agent responsible for COVID-19. While most research has focused on structural proteins, the accessory protein Open Reading Frame 8 (ORF8) has attracted attention for its role in immune evasion and the induction of a cytokine storm. Although the exact mechanisms underlying viral pathogenicity remain to be elucidated, oxidative stress has been proposed as a key contributing factor. In this study, we evaluated the effect of intranasal administration of ORF8 on arterial blood pressure and the antioxidant system in different organs of male BALB/c mice at 2- or 8 weeks post-administration. A significant increase in blood pressure and renal total antioxidant capacity was observed in the 8-week group, and decreased catalase activity in the prefrontal cortex was observed in the 2-week group. These findings suggest that ORF8 may contribute to long-term renal alterations and potentially to mechanism relevant to cognitive dysfunction associated with COVID-19.
Full article
(This article belongs to the Special Issue SARS-CoV-2, COVID-19 Pathologies, Long COVID, and Anti-COVID Vaccines)
►▼
Show Figures

Figure 1
Open AccessReview
Decoding Self vs. Non-Self: Alphavirus Cap0 Recognition and Immune Evasion
by
Santiago E. Faraj and Claudia V. Filomatori
Viruses 2026, 18(4), 439; https://doi.org/10.3390/v18040439 - 5 Apr 2026
Abstract
Host receptors can detect traces of non-self-pathogenic RNAs within a sea of cellular mRNA molecules. In host cells, mRNA cap methylation occurs in the nucleus, generating Cap1 and Cap2 structures (m7GpppNm and m7GpppNmNm, respectively). By contrast, alphavirus genomes carry
[...] Read more.
Host receptors can detect traces of non-self-pathogenic RNAs within a sea of cellular mRNA molecules. In host cells, mRNA cap methylation occurs in the nucleus, generating Cap1 and Cap2 structures (m7GpppNm and m7GpppNmNm, respectively). By contrast, alphavirus genomes carry a Cap0 structure (m7GpppN), which lacks 2′-O-methylation. This difference in the structure of the host and viral caps serves as a molecular signature that enables discrimination between self and non-self RNAs. Several host immune sensors, such as RIG-I and IFIT1, recognize the alphavirus Cap0 structure and trigger an antiviral response to restrict viral replication. It has been proposed that IFIT1 sequesters aberrant RNAs, preventing their translation by host ribosomes and blocking viral protein synthesis. However, alphaviruses have evolved molecular strategies to circumvent IFIT1-mediated restriction and facilitate infection in mammalian cells. One such strategy involves the folding of a 5′ RNA structure that hides the cap from host immune sensors. This highlights the dynamic interplay between viral evasion tactics and host immune defenses. This review will discuss how specific modifications at the 5′ end of alphavirus RNA modulate host defenses and how a deeper understanding of the virus–host interaction may inform the development of novel vaccine strategies.
Full article
(This article belongs to the Special Issue Advances in Understanding Viral Pathogenesis and Host Immune Responses to Arboviruses and Respiratory Viruses)
►▼
Show Figures
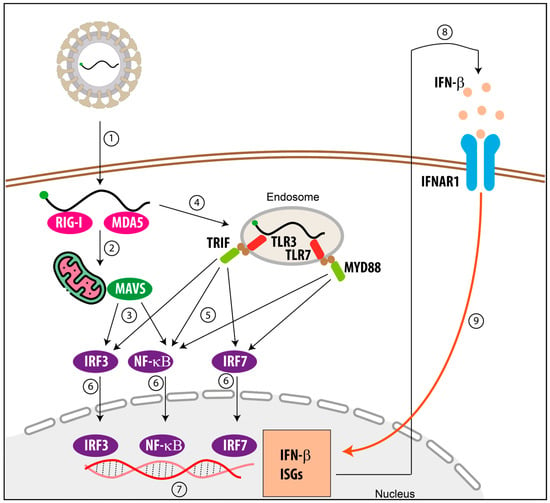
Figure 1
Open AccessArticle
EMCV Non-Structural Protein 2C Antagonizes cGAS-STING-Mediated Type I Interferon Signaling via Promoting K48-Linked Polyubiquitination and Degradation of STING
by
Rongrong Cheng, Pingan Dong, Wei Xing, Hongyuan Jin, Tingting Ma, Jingying Xie, Yanqiao Wen, Bixiu Su, Xiangrong Li and Ruofei Feng
Viruses 2026, 18(4), 438; https://doi.org/10.3390/v18040438 - 5 Apr 2026
Abstract
The cyclic GMP-AMP synthase-stimulator of interferon genes (cGAS-STING) pathway serves as a central innate immune signaling axis in host defense against DNA virus infections, and RNA viruses have also evolved diverse strategies to counteract this pathway. Encephalomyocarditis virus (EMCV), a zoonotic RNA virus,
[...] Read more.
The cyclic GMP-AMP synthase-stimulator of interferon genes (cGAS-STING) pathway serves as a central innate immune signaling axis in host defense against DNA virus infections, and RNA viruses have also evolved diverse strategies to counteract this pathway. Encephalomyocarditis virus (EMCV), a zoonotic RNA virus, utilizes its 2C protein to antagonize RIG-I-like receptor-mediated type I interferon signaling and induce autophagic degradation of calcium binding and coiled-coil domain 2, thereby evading host antiviral immunity. However, the precise molecular mechanism by which EMCV 2C protein modulates the cGAS-STING pathway remains incompletely understood. Herein, we show that EMCV infection reduces the expression of cGAS and STING proteins, and its 2C protein significantly suppresses the production of IFN-β triggered by poly(dA:dT) or viral infection, as well as the mRNA expression of interferon-stimulated genes. Mechanistically, 2C protein binds to STING via its ATPase domain and facilitates K48-linked polyubiquitination and proteasomal degradation of STING, while dominantly interfering STING translocation to the Golgi apparatus and the formation of STING-TBK1-IRF3 complex, thereby blocking STING-mediated IFN-β signal transduction at multiple levels. This study reveals a novel mechanism by which the EMCV 2C protein suppresses the host antiviral response by targeting STING and promoting its ubiquitination and degradation. This finding deepens understanding of the immune evasion mechanism of EMCV and provides a theoretical foundation for the development of antiviral therapies targeting the 2C protein of picornaviruses.
Full article
(This article belongs to the Section Viral Immunology, Vaccines, and Antivirals)
►▼
Show Figures

Figure 1
Open AccessArticle
New Species of Kronosvirus Bacteriophages That Infect Caulobacter Strain CBR1
by
Tannaz Mohammadi, Austin Millwood, Maherah Shaik and Bert Ely
Viruses 2026, 18(4), 437; https://doi.org/10.3390/v18040437 - 5 Apr 2026
Abstract
Caulobacter segnis strain CBR1 enhances the germination rate and total biomass of Arabidopsis plants. Since bacteriophages are usually present in the rhizosphere, we isolated three additional bacteriophages, designated TMCBR2, TMCBR3, and W2, that can replicate using CBR1 as the host strain. The genome
[...] Read more.
Caulobacter segnis strain CBR1 enhances the germination rate and total biomass of Arabidopsis plants. Since bacteriophages are usually present in the rhizosphere, we isolated three additional bacteriophages, designated TMCBR2, TMCBR3, and W2, that can replicate using CBR1 as the host strain. The genome sizes and morphologies of the new phages were similar to those of the previously isolated Kronos phage, and when we compared the nucleotide sequence of these new phage genomes, we found only minor differences among the four phage genomes. Pairwise sequence comparisons indicated that the phage genomes should be grouped into three separate species within Kronosvirus genus. Interestingly, we found that most of the genomic variation occurred among genes of unknown function within a 10 kb region of the 42 kb genomes with little variation in the remaining 32 kb which contains the genes known to be important for phage propagation.
Full article
(This article belongs to the Special Issue Bacteriophage Diversity, 2nd Edition)
►▼
Show Figures
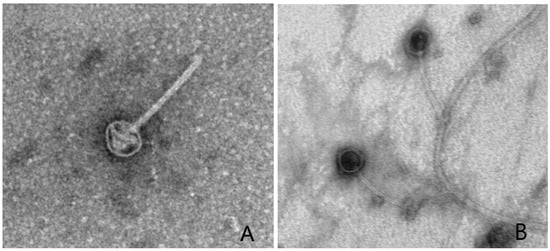
Figure 1
Open AccessReview
The Molecular Biology and Replication Cycle of Infectious Pancreatic Necrosis Virus
by
Daniela Espinoza, Jorge Gómez, Ana María Sandino, Sebastián Gonzalez-Catrilelbún and Andrea Rivas-Aravena
Viruses 2026, 18(4), 436; https://doi.org/10.3390/v18040436 - 3 Apr 2026
Abstract
Infectious pancreatic necrosis virus (IPNV), a member of the family Birnaviridae, is a major pathogen of farmed salmonids and an important model in fish virology. Despite its small genome, which encodes only five viral proteins, IPNV exhibits complex molecular processes that govern
[...] Read more.
Infectious pancreatic necrosis virus (IPNV), a member of the family Birnaviridae, is a major pathogen of farmed salmonids and an important model in fish virology. Despite its small genome, which encodes only five viral proteins, IPNV exhibits complex molecular processes that govern genome expression, replication, and particle assembly. Comprehensive descriptions of the molecular biology and replication cycle of IPNV were largely established in reviews published in the mid-1990s, whereas more recent reviews have primarily focused on virulence determinants, epidemiology, or host–virus interactions. This review provides an updated synthesis of available experimental knowledge on the molecular biology of IPNV by integrating classical and recent studies addressing virion architecture, genome organization, and the functions of viral proteins. Particular attention is given to the molecular events involved in the viral replication cycle, including virus entry, genome transcription, translation and replication in the cytoplasm, polyprotein processing by the viral protease, and the coordination between genome replication and virion assembly. When appropriate, experimental observations from the related Avibirnavirus infectious bursal disease virus are considered to provide additional context for molecular mechanisms conserved within the family Birnaviridae. Together, these studies outline the current understanding of the molecular processes governing IPNV replication and morphogenesis.
Full article
(This article belongs to the Special Issue Viral Diseases of Aquatic Animals: Crustaceans, Mollusks, Fish, Sea Turtles, and Marine Mammals)
►▼
Show Figures

Figure 1
Open AccessArticle
Evidence of SARS-CoV-2 Exposure in Rodents from Rural Localities in the Yucatan Peninsula, Mexico
by
Ana Laura Vigueras-Galván, Montserrat Elemi García-Hernández, Karen Cortés-Sarabia, Oscar Del Moral-Hernández, Sofía L. Alcaraz-Estrada, Benjamín Roche, Audrey Arnal, Gerardo Suzán and Rosa Elena Sarmiento-Silva
Viruses 2026, 18(4), 435; https://doi.org/10.3390/v18040435 - 3 Apr 2026
Abstract
Zoonotic diseases involve pathogen transmission between humans and animals, with most research focused on animal-to-human spillover. However, reverse zoonosis—the transmission of pathogens from humans to animals—remains understudied despite its potential ecological and epidemiological consequences. The SARS-CoV-2 pandemic highlights this risk, as human-associated viruses
[...] Read more.
Zoonotic diseases involve pathogen transmission between humans and animals, with most research focused on animal-to-human spillover. However, reverse zoonosis—the transmission of pathogens from humans to animals—remains understudied despite its potential ecological and epidemiological consequences. The SARS-CoV-2 pandemic highlights this risk, as human-associated viruses may sporadically infect wildlife species and generate novel exposure pathways. To assess evidence of SARS-CoV-2 exposure in wildlife, we analyzed serum and rectal swab samples from rodents collected in rural localities of the Yucatan Peninsula, Mexico, between 2021 and 2022. An indirect ELISA detected antibodies against SARS-CoV-2 in 23.1% of sampled rodents. Molecular analysis detected one positive sample with a pan-coronavirus RT-PCR, though all were negative for SARS-CoV-2–specific assays. This study provides serological evidence of SARS-CoV-2 exposure in rodent communities from rural areas of Mexico and is consistent with sporadic wildlife spillback events rather than sustained transmission. The observed exposure patterns may be influenced by human activities and frequent human–wildlife interactions in heterogeneous rural landscapes. Our results underscore the need for integrated serological and genomic surveillance to better understand the ecological context of reverse zoonosis and its implications for public health.
Full article
(This article belongs to the Special Issue Synanthropy and Zoonotic Viral Spillover)
►▼
Show Figures
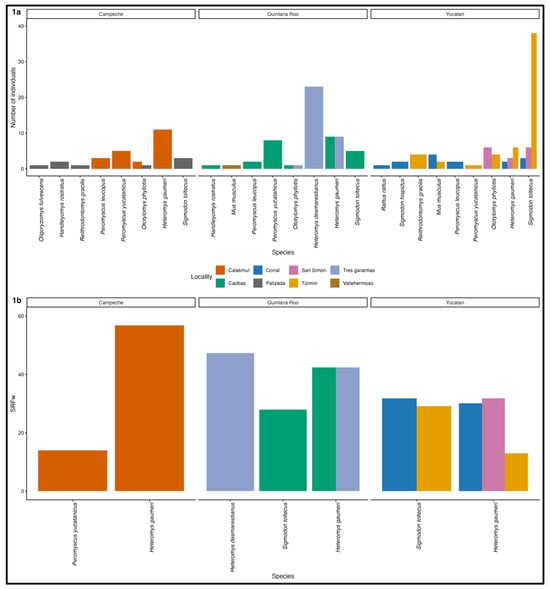
Figure 1
Open AccessArticle
Characterisation of RT Connection and RNase H Polymorphisms in HIV-1 Subtype C in Botswana
by
Boitumelo J. L. Zuze, Wonderful T. Choga, Natasha O. Moraka-Mankge, Ontlametse T. Choga, Lynnette Bhebhe, Dorcas Maruapula, Thato Phuthego, Margaret Mokomane, Sikhulile Moyo and Simani Gaseitsiwe
Viruses 2026, 18(4), 434; https://doi.org/10.3390/v18040434 - 3 Apr 2026
Abstract
Emerging evidence suggests that polymorphisms in the reverse transcriptase connection (RT-conn) and RNase H domains may contribute to resistance to reverse transcriptase inhibitors (RTIs). Here, we characterised the polymorphic landscape of the RT-conn and RNase H domains in HIV-1 subtype C (HIV-1C) from
[...] Read more.
Emerging evidence suggests that polymorphisms in the reverse transcriptase connection (RT-conn) and RNase H domains may contribute to resistance to reverse transcriptase inhibitors (RTIs). Here, we characterised the polymorphic landscape of the RT-conn and RNase H domains in HIV-1 subtype C (HIV-1C) from Botswana across the pre-ART and post-ART eras, including treatment-naïve (TN) and treatment-experienced (TE) individuals. A total of 1571 HIV-1C sequences were analysed: 76 pre-ART (≤2002) and 1495 post-ART (>2002) sequences were obtained from the Los Alamos database and the Botswana Combination Prevention Project (2013–2018). Post-ART sequences were stratified into TN (n = 1282) and TE individuals with virologic failure (TEVF, n = 213). Naturally occurring and ART-associated polymorphisms within RT-conn (aa 321–440) and RNase H (aa 441–560) were assessed. Among TN individuals, 12 polymorphisms exceeded 5% pre-ART, including R461K and L491P, while 31 polymorphisms were observed post-ART, indicating a temporal shift. Several substitutions were significantly higher in TEVF and showed a history of thymidine analogue-, tenofovir- and lamivudine/emtricitabine-based exposure. Covariant analysis identified significant co-occurrence of polymerase mutations (M184V/I, D67N) with RT-conn/RNase H substitutions (p < 0.05). These findings demonstrate HIV-1C evolution within the extended RT domains under ART pressure and support their inclusion in molecular surveillance frameworks in Botswana.
Full article
(This article belongs to the Special Issue Advances in Research on HIV Drug Resistance and Other Determinants of Treatment Success: 3rd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
The 4.4 Å Capsid Structure of the Giant Melbournevirus Belonging to the Marseilleviridae Family
by
Raymond N. Burton-Smith, Chantal Abergel, Kenta Okamoto and Kazuyoshi Murata
Viruses 2026, 18(4), 433; https://doi.org/10.3390/v18040433 - 2 Apr 2026
Abstract
Members of Marseilleviridae, a family of icosahedral giant viruses, have been identified worldwide in all types of environments. The virion shows a characteristic internal membrane extrusion at the five-fold vertices of the capsid, but its structural details need to be elucidated. We
[...] Read more.
Members of Marseilleviridae, a family of icosahedral giant viruses, have been identified worldwide in all types of environments. The virion shows a characteristic internal membrane extrusion at the five-fold vertices of the capsid, but its structural details need to be elucidated. We now report the 4.4 Å cryo-electron microscopy structure of the melbournevirus capsid by using a block-based reconstruction approach. Results: An atomic model of the major capsid protein (MCP) shows a unique cup structure on the trimer that accommodates additional proteins. A polyalanine model of the Penton base protein shows internally extended N- and C-terminals, which indirectly connect to the internal membrane extrusion. The Marseilleviruses share the same orientational organization of the MCPs as previously reported for other giant viruses, but the unique minor capsid protein components named Scaffold may be alternatively utilized to control the dimensions of the capsid during assembly as the tape measure protein.
Full article
(This article belongs to the Special Issue Nucleocytoviricota)
►▼
Show Figures
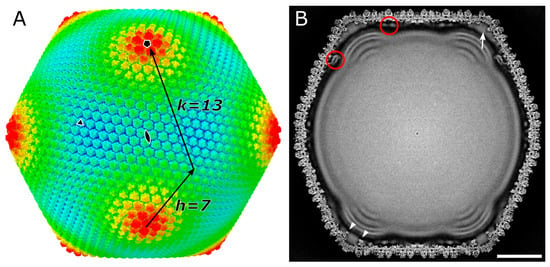
Figure 1
Open AccessArticle
Differences in Susceptibility to Cyprinid Herpesvirus 3 (CyHV-3) Infection Among Carp (Cyprinus carpio L.) Strains and Hybrids
by
Xiaona Jiang, Zhenguo Song, Chitao Li, Xuesong Hu, Yanlong Ge, Lei Cheng, Xiaodan Shi, Yaxin Di and Zhiying Jia
Viruses 2026, 18(4), 432; https://doi.org/10.3390/v18040432 - 2 Apr 2026
Abstract
Cyprinid herpesvirus 3 (CyHV-3) is a pathogen that causes high mortality in common carp (Cyprinus carpio) and koi. Common carp breeding lines with different genetic backgrounds exhibit different resistance levels to viral pathogens. This study aimed to determine the differences in
[...] Read more.
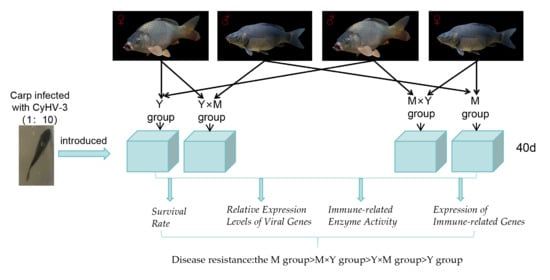
Cyprinid herpesvirus 3 (CyHV-3) is a pathogen that causes high mortality in common carp (Cyprinus carpio) and koi. Common carp breeding lines with different genetic backgrounds exhibit different resistance levels to viral pathogens. This study aimed to determine the differences in CyHV-3 disease resistance performance between the hybrid offspring (Y × M and M × Y) of the mirror carp ‘Longke 11’ (resistant to CyHV-3) and Yellow River carp, as well as the self-crossed offspring (M and Y). The M, Y × M, M × Y and Y groups were infected with CyHV-3 by immersion. The order of mortality and the duration of death for the four groups of carp were as follows: Y group > Y × M group > M × Y group > M group. Throughout the entire infection stage, the mRNA expression levels of the viral factors thymidine kinase (TK) and open reading frame 72 (ORF72) in the four groups of carp tended to first increase but then decrease. The viral factor expression evaluated on days 30 and 31 post-infection (p.i.), which was the peak of infection mortality, was the highest in the Y group and the lowest in the M group, and compared with the Y × M group, the M × Y group had considerably lower viral gene expression (p < 0.05). The immune-related enzyme activity and content levels of the four carp groups matched the patterns of viral gene expression. On day 29 p.i., a time point with high mortality, the levels of alkaline phosphatase (AKP), glutathione peroxidase (GSH-Px) and total antioxidant capacity (T-AOC) were significantly the lowest in the Y group and significantly the highest in the M group, while the Y × M group showed a significant decrease compared to the M × Y group (p < 0.05). Quantitative real-time (q-PCR) analysis revealed that interleukin-21 receptor (IL21R), interferon regulatory factor 9 (IRF9), interferon type I (IFN-I), interleukin-6 (IL-6) and microtubule-associated protein light chain 3 (LC3), exhibited an initial increase followed by a decrease among the four experimental groups of common carp. In the peak mortality period of carp in the four groups (30 days post-infection), the expression levels of IL21R, IRF9, LC3, and IFN-I were significantly the highest in the M group and significantly the lowest in the Y group, with the mRNA expression of these genes in the M × Y group being significantly higher than that in the Y × M group (p < 0.05). In contrast, IL-6 expression levels exhibited the opposite trend. In this study, the M group exhibited the greatest resistance to CyHV-3, followed by the M × Y group, whose resistance was greater than that of the Y × M group, with the Y group showing the lowest disease resistance. Our findings demonstrate that hybridization modulates resistance to CyHV-3. Furthermore, we identified conserved immune signatures common to both susceptible and resistant carp, including the activation of nonspecific immunity and the upregulation of immune-associated genes.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Eye Health and Vision Function in Adults Aging with Well-Controlled HIV
by
Alison G. Abraham, Xinxing Guo, Srijana Lawa, Aleks Mihailovic, Michael W. Plankey, Todd T. Brown, Joseph B. Margolick, Pradeep Ramulu and Seema Banerjee
Viruses 2026, 18(4), 431; https://doi.org/10.3390/v18040431 - 2 Apr 2026
Abstract
Here, we describe vision health in aging adults living with HIV (PLWH) and comparable people without HIV (PWOH) from the MACS/WIHS Combined Cohort Study (MWCCS). PLWH and PWOH aged 60 years and older were recruited from Baltimore/Washington, DC, from September 2021 to September
[...] Read more.
Here, we describe vision health in aging adults living with HIV (PLWH) and comparable people without HIV (PWOH) from the MACS/WIHS Combined Cohort Study (MWCCS). PLWH and PWOH aged 60 years and older were recruited from Baltimore/Washington, DC, from September 2021 to September 2023. Exact matching and sample weights were used to create age-balanced comparisons. Visual impairment (VA worse than 20/40 after refraction or CS worse than 1.50 logCS in the better eye) and the presence of eye pathology were assessed. We studied 74 PLWH (97% virally suppressed) and 65 PWOH, aged 61 to 79 years, 36% Black, and 87% male. For PLWH and PWOH, distance VA impairment was noted in 4% vs. 1%, respectively, and uncorrected refractive error in 15% vs. 5%. More than half had signs of dry eye disease (63% for PLWH and 51% for PWOH). About half of PLWH had developed at least an early stage of cataract, compared to 20% of PWOH. Posterior chamber abnormalities were observed in 4% and 0%, and glaucomatous changes in 19% and 25% of PLWH and PWOH, respectively. The need for eyecare was high among this sample of PLWH with viral suppression and PWOH.
Full article
(This article belongs to the Special Issue HIV and Aging)
►▼
Show Figures
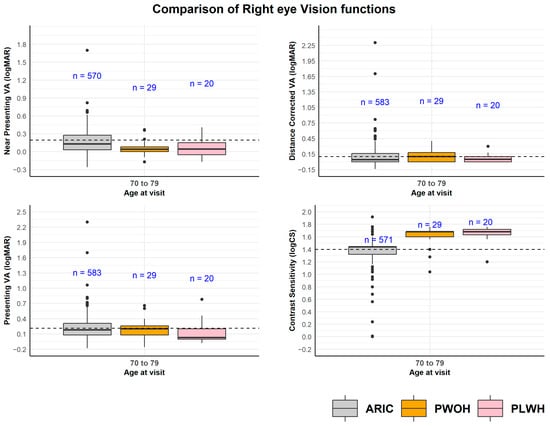
Figure 1

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics

Conferences
Special Issues
Special Issue in
Viruses
Mechanisms of Ribozyviria Transmission in Animal and Vegetal Holobionts
Guest Editors: Maurizia Rossana Brunetto, Ferruccio BoninoDeadline: 15 April 2026
Special Issue in
Viruses
Beyond Acute: Navigating Long COVID and Post-Viral Complications
Guest Editors: Maggie Bartlett, Janko Nikolich-Žugich, Rubeshan Perumal, Anders VahlneDeadline: 15 April 2026
Special Issue in
Viruses
Broadly Protective Anti-Viral Vaccines 2025
Guest Editor: Lei DengDeadline: 15 April 2026
Special Issue in
Viruses
Viral Infections in Hematopoietic Stem Cell Transplant and Cellular Therapy Recipients, 2nd Edition
Guest Editor: Sanjeet Singh DadwalDeadline: 15 April 2026
Topical Collections
Topical Collection in
Viruses
Mathematical Modeling of Viral Infection
Collection Editors: Amber M. Smith, Ruian Ke
Topical Collection in
Viruses
Efficacy and Safety of Antiviral Therapy
Collection Editors: Giordano Madeddu, Andrea De Vito, Agnese Colpani










