Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS), Brazilian Society for Virology (BSV) and Global Virus Network (GVN) are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.2 days after submission; acceptance to publication is undertaken in 2.7 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
3.5 (2024);
5-Year Impact Factor:
3.7 (2024)
Latest Articles
Targeted Protein Degradation Strategies in DNA Virus Research
Viruses 2026, 18(6), 658; https://doi.org/10.3390/v18060658 (registering DOI) - 9 Jun 2026
Abstract
DNA viruses rely extensively on host cellular machinery, including replication factors and transcriptional systems, to persist after infection. These mechanisms make studying and targeting DNA viral proteins challenging, as they also play key roles in mammalian processes. Traditional strategies include CRISPR-mediated gene disruption
[...] Read more.
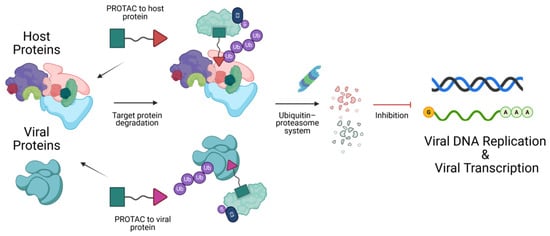
DNA viruses rely extensively on host cellular machinery, including replication factors and transcriptional systems, to persist after infection. These mechanisms make studying and targeting DNA viral proteins challenging, as they also play key roles in mammalian processes. Traditional strategies include CRISPR-mediated gene disruption and small interfering RNA (siRNA) to target host proteins. However, Proteolysis Targeting Chimeras (PROTACs) offer a novel strategy by enabling the selective and rapid degradation of specific viral or host proteins involved in the DNA viral lifecycle. PROTACs are heterobifunctional molecules composed of three key components: a ligand that binds the target protein, a chemical linker, and a ligand that recruits an E3 ubiquitin ligase. By simultaneously binding both the target protein and the E3 ligase, PROTACs form a ternary complex. This proximity enables the E3 ligase to ubiquitinate the target protein, marking it for recognition and subsequent degradation by the intracellular proteasome. This approach represents a promising avenue for targeting previously undruggable proteins and improving therapeutic outcomes in virus-associated malignancies. In this perspective, we describe studies that use PROTACs as tools to modulate host proteins to investigate DNA viral processes with temporal control of host protein expression, as well as the use of PROTACs as antivirals to directly target DNA viral proteins. We also provide a detailed chart summarizing known host-targeting PROTACs and their potential applications across different stages of DNA viral lifecycles, highlighting opportunities for future DNA virus research.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►
Show Figures
Open AccessArticle
Mapping the TAR vRNA Interaction with HIV-1 Integrase
by
Jian Sun, Rahul Yadav, Tolga Catmakas, Luke Fisher, Nicholas C. Fitzkee and Jacques J. Kessl
Viruses 2026, 18(6), 657; https://doi.org/10.3390/v18060657 (registering DOI) - 9 Jun 2026
Abstract
A series of critical interactions within the viral core between viral RNA (vRNA) and HIV-1 Integrase (IN) has previously been reported. In these studies, contact points between vRNA and IN were identified using RNA-seq and MS-based protein foot-printing approaches. Several IN amino acids
[...] Read more.
A series of critical interactions within the viral core between viral RNA (vRNA) and HIV-1 Integrase (IN) has previously been reported. In these studies, contact points between vRNA and IN were identified using RNA-seq and MS-based protein foot-printing approaches. Several IN amino acids located in its C-terminal domain (CTD) were found to be essential for vRNA binding, and their alanine substitution severely impacted the correct morphogenesis of the mature viral core. Here, we have used the TAR element to extend these studies by performing a comprehensive mapping of the interaction by deploying RNA crosslinking and NMR methodologies. Together, these approaches were able to identify additional contact points between the TAR vRNA and IN. Our results reveal several new basic amino acids located in the IN CTD critical for the vRNA-IN interaction, viral replication and correct morphology of the mature viral core.
Full article
(This article belongs to the Special Issue Molecular Insights into HIV-1 Infection)
►▼
Show Figures
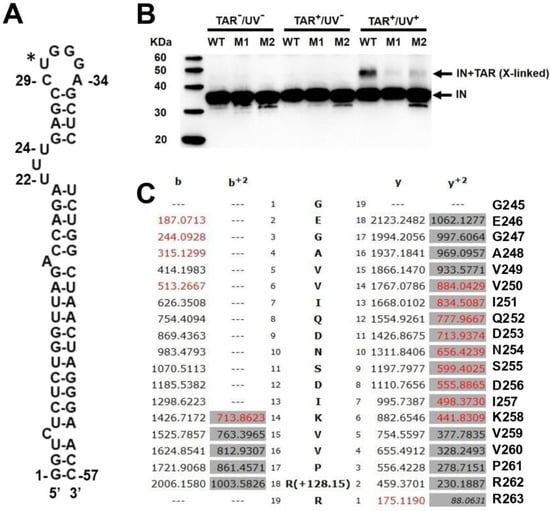
Figure 1
Open AccessArticle
ViroBioTree: A Tree-Structured Biological Evidence Retrieval Framework for Viral Protein Function Annotation
by
Tinglian Lai, Fuguo Liu, Guodong Li and Liyan Hua
Viruses 2026, 18(6), 656; https://doi.org/10.3390/v18060656 (registering DOI) - 9 Jun 2026
Abstract
Accurate viral protein function annotation is essential for genomic surveillance, yet conventional retrieval-augmented generation (RAG) pipelines often fragment biological evidence into fixed-length text chunks, disrupting relationships among ORFs, annotations, structural domains, sequence motifs, residue mappings, and model-derived attention evidence. We propose ViroBioTree, a
[...] Read more.
Accurate viral protein function annotation is essential for genomic surveillance, yet conventional retrieval-augmented generation (RAG) pipelines often fragment biological evidence into fixed-length text chunks, disrupting relationships among ORFs, annotations, structural domains, sequence motifs, residue mappings, and model-derived attention evidence. We propose ViroBioTree, a tree-structured biological evidence retrieval framework for downstream viral protein evidence review rather than a new primary annotation classifier. Built as an evidence organization layer on ViralMultiNet-derived ORF-level predictions and annotations, ViroBioTree converts sequence, annotation, structure, and attention evidence into typed biological nodes and traceable edges, then performs deterministic multi-channel recall, evidence-aware reranking, balanced TopK selection, rule-based verification, and node-cited report generation. In a demo benchmark, ViroBioTree achieved its strongest deterministic proxy performance on structure-explanation tasks, with Precision@K = 1.0, Recall@K = 1.0, and diversity = 0.52; these values reflect expected node-type and tag agreement rather than independent biological correctness. A bounded full-scale SARS-CoV-2 index contained 39,800 ORF rows, 80,000 attention records, 199,418 nodes, and 495,886 edges. In a stratified full20k diagnostic evaluation, ViroBioTree showed task-dependent advantages over LlamaIndex vector retrieval for conflict detection, evidence retrieval, and structure explanation, while LlamaIndex remained competitive or stronger for annotation-rich function annotation. A cross-family Influenza A Virus (IAV) diagnostic audit showed that the schema can represent IAV evidence namespaces while explicitly exposing missing formal ORF inputs, missing attention evidence, and unavailable residue/PDB assertions. Supplementary robustness, external sanity-check, diversity-risk, expert-evaluation, domain-tool positioning, and cross-family audit analyses supported traceability, report quality, and conservative evidence handling, but also showed that stable Precision@K under query perturbation does not necessarily imply stable retrieved evidence sets. ViroBioTree operates offline and deterministically, but does not address raw-read assembly, base calling, primary ORF prediction, or wet-lab validation. Its results should be interpreted as proxy and expert-reviewed evidence for traceable viral protein evidence retrieval and report generation rather than as direct validation of biological function annotation.
Full article
(This article belongs to the Section General Virology)
►▼
Show Figures
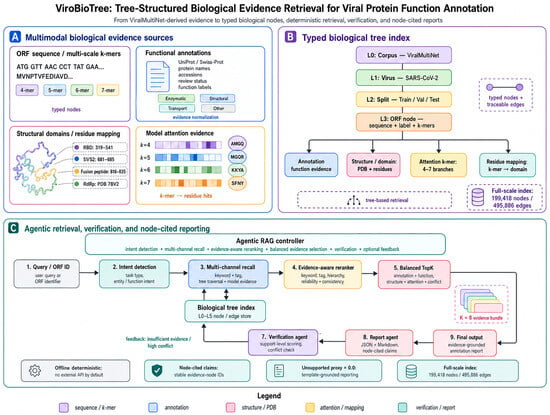
Figure 1
Open AccessReview
Reprogramming Rotavirus: Reverse Genetics-Driven Design of Viral Vector Platforms
by
Ke Li, Xiao Wei, Xuanze Ouyang, Xiafei Liu, Pengdi Chai, Yu Bai and Zhaojun Duan
Viruses 2026, 18(6), 655; https://doi.org/10.3390/v18060655 (registering DOI) - 9 Jun 2026
Abstract
Rotaviruses remain the leading cause of severe dehydrating diarrhea and associated mortality in children under five years of age worldwide. The successful global rollout of live-attenuated oral rotavirus vaccines has dramatically reduced rotavirus gastroenteritis morbidity and mortality, unequivocally demonstrating their excellent safety profile
[...] Read more.
Rotaviruses remain the leading cause of severe dehydrating diarrhea and associated mortality in children under five years of age worldwide. The successful global rollout of live-attenuated oral rotavirus vaccines has dramatically reduced rotavirus gastroenteritis morbidity and mortality, unequivocally demonstrating their excellent safety profile and potent induction of mucosal immunity. These attributes highlight the substantial potential of rotaviruses as novel oral mucosal vaccine vectors. Recent breakthroughs in reverse genetics, particularly the establishment of a fully plasmid-based system in 2017, have enabled precise insertion and stable expression of foreign antigens at targeted genomic loci. This advance has opened a viable pathway for developing multivalent oral mucosal vaccines. This review traces the historical development of rotavirus reverse genetics and summarizes the latest progress in its application as a vaccine vector platform. We focus on key strategies for foreign gene insertion and immunological outcomes in animal models, while critically evaluating persistent challenges—virus rescue efficiency, genetic stability of inserts, and limitations of current animal models—and outlining the rational design framework for RV-based vectors with improved stability, expression efficiency, and immunogenicity.
Full article
(This article belongs to the Special Issue Advances in Viral Vectors for Gene and Cell Therapy)
►▼
Show Figures
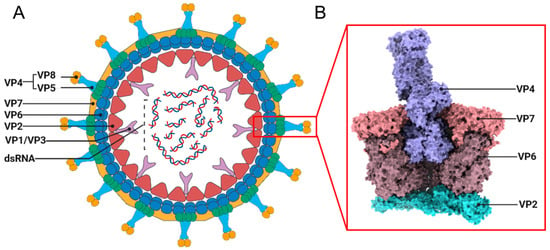
Figure 1
Open AccessArticle
Cloning and Characterization of an Endolysin LysSA120 as a Potential Staphylococcus Biofilm-Removing Agent
by
Natalia N. Golosova, Yana A. Khlusevich, Bogdana I. Kravchuk, Lyudmila A. Emelyanova, Anna V. Sushko, Yulia N. Kozlova, Nina V. Tikunova and Andrey L. Matveev
Viruses 2026, 18(6), 654; https://doi.org/10.3390/v18060654 (registering DOI) - 9 Jun 2026
Abstract
Staphylococcus aureus, including methicillin- and vancomycin-resistant variants (MRSA, VRSA), causes infections that are increasingly difficult to treat with conventional antibiotics. One of the approaches to developing new therapeutics to treat staphylococcal infections is the use of bacteriophages specific to these bacteria or
[...] Read more.
Staphylococcus aureus, including methicillin- and vancomycin-resistant variants (MRSA, VRSA), causes infections that are increasingly difficult to treat with conventional antibiotics. One of the approaches to developing new therapeutics to treat staphylococcal infections is the use of bacteriophages specific to these bacteria or the lytic enzymes of such bacteriophages, which are capable of hydrolyzing the cell walls of these bacteria. Phage-encoded endolysins offer an alternative promising class of antimicrobial agents. In this study, LysSA120, a 250-amino-acid endolysin encoded by the S. aureus podophage vB_SauP_120, was cloned, expressed in Escherichia coli, and characterized. The domain organization and tertiary structure of LysSA120 were predicted. Recombinant LysSA120 hydrolyzed cell walls were obtained from S. aureus, S. epidermidis, S. haemolyticus, S. warneri, S. auricularis and S. saprophyticus. It was shown that treatment of S. aureus planktonic cells with endolysin LysSA120 led to reduced viability. Furthermore, LysSA120 could hydrolyze mature biofilm formed by VRSA. The lytic spectrum and antibiofilm activity of LysSA120 warrant its further evaluation as an enzybiotic against drug-resistant staphylococcal infections.
Full article
(This article belongs to the Special Issue Biotechnological Applications of Phage and Phage-Derived Proteins 2025-2026)
►▼
Show Figures
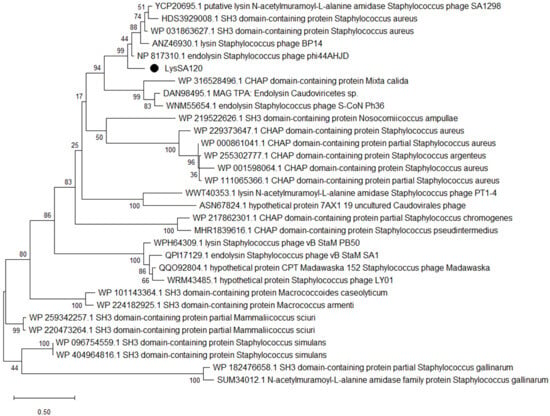
Figure 1
Open AccessArticle
Evaluation of Safety, Immunogenicity and Efficacy of an Inactivated Bovine Viral Diarrhea Virus (BVDV-1) Vaccine Candidate in Cattle
by
Semmannan Kalaiyarasu, Niranjan Mishra, Shashi Bhusan Sudhakar, Vijendra Pal Singh and Aniket Sanyal
Viruses 2026, 18(6), 653; https://doi.org/10.3390/v18060653 (registering DOI) - 8 Jun 2026
Abstract
Bovine viral diarrhea (BVD) is a globally significant disease that adversely affects cattle health and productivity, including in India. It is caused by three bovine pestiviruses: bovine viral diarrhea virus 1 (BVDV-1), BVDV-2, and HoBi-like pestivirus (HoBiPeV), which belong to the Pestivirus genus
[...] Read more.
Bovine viral diarrhea (BVD) is a globally significant disease that adversely affects cattle health and productivity, including in India. It is caused by three bovine pestiviruses: bovine viral diarrhea virus 1 (BVDV-1), BVDV-2, and HoBi-like pestivirus (HoBiPeV), which belong to the Pestivirus genus within the Flaviviridae family. Despite the prevalence of all three pestivirus species in India, no commercial vaccine based on the local circulating strain is currently available. This study evaluates the safety, immunogenicity, and protective efficacy of an inactivated whole-virus BVD vaccine, based on an Indian BVDV-1 strain. The virus was propagated in MDBK cells, inactivated using 3 mM binary ethylenimine (BEI) for 24 h at 37 °C, and formulated with Montanide ISA 61 VG (SEPPIC) in a 50:50 water-in-oil emulsion. Vaccine safety was confirmed in both guinea pigs and bovine calves, with no adverse effects observed. Immunogenicity testing in guinea pigs (n = 6) showed neutralizing antibody titres up to 9 log2 (1/512). In calves aged 9–12 months (n = 3), the vaccine elicited strong humoral and cell-mediated immune responses, with mean neutralizing antibody titres against the homologous BVDV-1 strain reaching 14 log2 (1/16,384). Neutralizing antibody levels remained detectable for up to 12 months post vaccination with sustained mean titres of 7 log2 (1/128). Notably, titres reported to be adequate for fetal protection (≥9 log2 or ≥1/512 were maintained for five months following vaccination. Challenge studies demonstrated complete protection of vaccinated calves against homologous BVDV-1 acute infection. In addition, the vaccine conferred partial cross-protection against heterologous strains including BVDV-2 and HoBiPeV. In a field trial involving 125 cattle, 74% of animals developed protective neutralizing titres (≥7 log2 or ≥1/128), while 48% achieved titres reported to be adequate for fetal protection (9 log2 or 1/512). Furthermore, 92% of vaccinated cattle maintained neutralizing antibody titres of at least 6 log2 (≥1/64) for up to six months post-booster vaccination. A strong positive correlation was observed between guinea pig and bovine antibody responses (R2 = 0.6809; p < 0.0001), indicating the potential of guinea pigs as a predictive model. Vaccine stability was confirmed for up to 8 months when stored at 4 °C, as demonstrated by the immunogenicity in guinea pigs. Collectively, these findings demonstrate that the locally developed inactivated BVDV-1 vaccine is safe, highly immunogenic, and capable of providing protective immunity against BVDV-1 infection, supporting its potential use in BVD control programs in India.
Full article
(This article belongs to the Special Issue Pestivirus 2026)
►▼
Show Figures
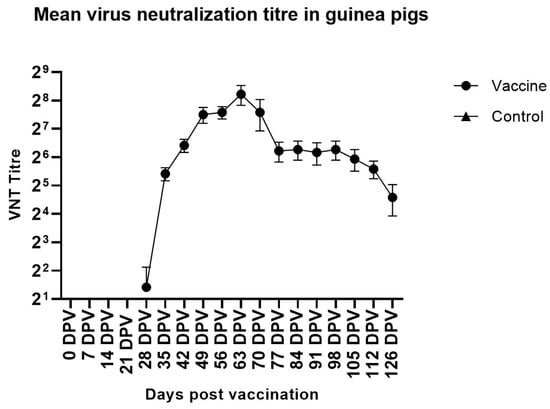
Figure 1
Open AccessArticle
Hantavirus Seroprevalence in the Population of Saint Petersburg and the Leningrad Region, Russia
by
Tatiana Arbuzova, Dmitry Naydenov, Regina Baimova, Alena Khalilova, Denis Sarksyan, Konstantin Manakhov, Tamara Ginevskaia, Margarita Popova, Ekaterina Klyuchnikova, Svetlana Egorova, Vladimir Dedkov and Areg Totolian
Viruses 2026, 18(6), 652; https://doi.org/10.3390/v18060652 - 6 Jun 2026
Abstract
The aim of the study was to assess the seroprevalence of hantaviruses, the causative agents of hemorrhagic fever with renal syndrome (HFRS), and their distribution to socio-demographic characteristics among the populations of Saint Petersburg and the Leningrad Region. A total of 4464 samples
[...] Read more.
The aim of the study was to assess the seroprevalence of hantaviruses, the causative agents of hemorrhagic fever with renal syndrome (HFRS), and their distribution to socio-demographic characteristics among the populations of Saint Petersburg and the Leningrad Region. A total of 4464 samples were analyzed, including 2265 samples from residents of Saint Petersburg and 2199 samples from residents of the Leningrad Region. Blood plasma samples were tested for specific immunoglobulin G (IgG) antibodies using enzyme-linked immunosorbent assay (ELISA). Blood samples were collected in 2023 from randomly selected volunteers. Hantavirus seroprevalence in Saint Petersburg was 5.39%, while in the Leningrad Region, it was 8.55%. In both regions, the highest proportion of seropositive individuals was found among volunteers aged ≥70 years, whereas the lowest seroprevalence was observed in the 1–17-year age group (inclusive). Seroprevalence was significantly higher in men than in women in both regions. The seroprevalence values identified in this study are comparable to those reported in similar studies in areas with a high incidence of HFRS. These findings may indicate that the true incidence of HFRS may be significantly higher than officially registered in Saint Petersburg and the Leningrad Region.
Full article
(This article belongs to the Special Issue Advances in Research on Emerging and Zoonotic Diseases)
►▼
Show Figures
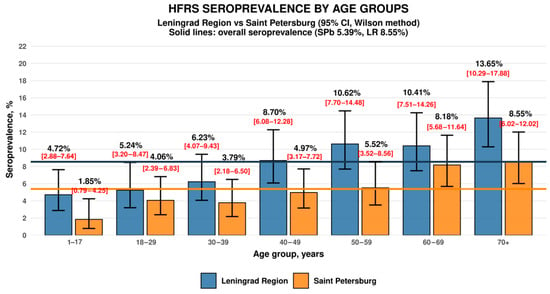
Figure 1
Open AccessArticle
Co-Circulation of Divergent Strains Supports Vector-Mediated Transmission of Rodent Hepacivirus J (Orthohepacivirus glareoli)
by
Sarah Marmorosch, Thomas Anton von Graffenried, Rainer G. Ulrich and Gerald Heckel
Viruses 2026, 18(6), 651; https://doi.org/10.3390/v18060651 - 5 Jun 2026
Abstract
Orthohepacivirus glareoli (RHVJ, Hepaciviridae) and its natural host, the bank vole (Myodes glareolus), have been proposed as a model system for human hepatitis C research, yet the mode of transmission remains largely unknown. Here, we investigated spatial patterns of RHVJ
[...] Read more.
Orthohepacivirus glareoli (RHVJ, Hepaciviridae) and its natural host, the bank vole (Myodes glareolus), have been proposed as a model system for human hepatitis C research, yet the mode of transmission remains largely unknown. Here, we investigated spatial patterns of RHVJ sequence diversity and evolutionary relationships using novel sequences from dense regional sampling alongside all published NS3 gene sequences. Phylogenetic analyses revealed mostly local clustering of RHVJ sequences, resulting in isolation-by-distance patterns at regional geographic scales. This suggests primarily local transmission of the virus. However, nucleotide sequence divergence of up to 19% within bank vole populations is difficult to reconcile with local transmission of RHVJ alone, implying that mechanisms beyond local evolution shape the extensive sequence diversity of RHVJ at local scales. Using spatially explicit computer simulations of sequence evolution, we contrasted the phylogenetic relationships resulting from exclusively short-distance transmission, e.g., from vole to vole, with those resulting when rare long-distance transmission events were included. The latter produced phylogenetic patterns comparable to those of RHVJ, including monophyletic clustering of samples from distant locations and unresolved basal nodes. We suggest that the transmission of RHVJ likely involves at least occasionally a vector, as the mobility of its natural rodent host is strongly limited.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
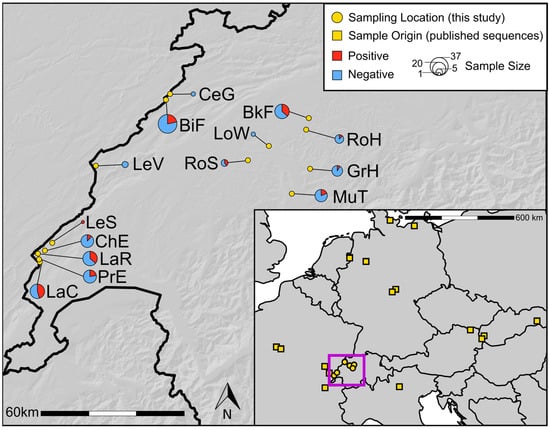
Figure 1
Open AccessEditorial
Respiratory Viruses, Self-Diagnosis, Early Treatment and Prophylaxis
by
Anna Puigdellívol-Sánchez, Celia Lozano-Paz, Roger Valls-Foix, Ignacio Morán-Blanco, Ignasi Calicó-Bosch and Mariana Castells
Viruses 2026, 18(6), 650; https://doi.org/10.3390/v18060650 - 4 Jun 2026
Abstract
Respiratory viruses are a common cause of hospital admissions during the winter season worldwide [...]
Full article
(This article belongs to the Special Issue Advances in Respiratory Viruses Research: From Basic Studies to Public Health)
Open AccessArticle
HMPV Impairs Macrophage Phagocytosis Through a Replication-Dependent Mechanism Associated with Reduced CD36 Expression and the Viral G Protein
by
Iván Martínez-Espinoza, Pius I. Babawale, Basel Abuaita and Antonieta Guerrero-Plata
Viruses 2026, 18(6), 649; https://doi.org/10.3390/v18060649 - 4 Jun 2026
Abstract
Human metapneumovirus (HMPV) is a major cause of respiratory infections, but its impact on macrophage antibacterial functions remains poorly understood. Macrophages play a crucial role in host defense through phagocytosis, and impairment of this function may increase susceptibility to secondary infections. Here, we
[...] Read more.
Human metapneumovirus (HMPV) is a major cause of respiratory infections, but its impact on macrophage antibacterial functions remains poorly understood. Macrophages play a crucial role in host defense through phagocytosis, and impairment of this function may increase susceptibility to secondary infections. Here, we show that HMPV infection of THP-1-derived macrophages significantly reduces bacterial uptake in a replication-dependent manner. This effect was restricted to infected cells and was not recapitulated by cell-free supernatants, indicating a cell-intrinsic mechanism. HMPV infection was also associated with reduced expression of the scavenger receptor CD36. Viral gene knockdown studies further implicated the HMPV G protein in this phenotype, as silencing the G protein restored phagocytic function. Analysis of single-cell RNA-sequencing datasets from HMPV-infected mouse lungs revealed reduced CD36 expression and broader alterations in phagocytosis-associated gene programs across lung macrophage subsets. Supporting these observations, expression of Cd36 and Marco was reduced in lung tissue from HMPV-infected mice. Overall, these findings demonstrate that HMPV impairs macrophage-mediated bacterial uptake through a replication-dependent, cell-intrinsic mechanism and identify reduced scavenger receptor expression and the viral G protein as factors associated with this phenotype. These alterations may contribute to increase susceptibility to secondary bacterial infections during HMPV infection.
Full article
(This article belongs to the Special Issue Viral Strategies to Regulate Host Immunity or Signal Pathways, 2nd Edition)
►▼
Show Figures
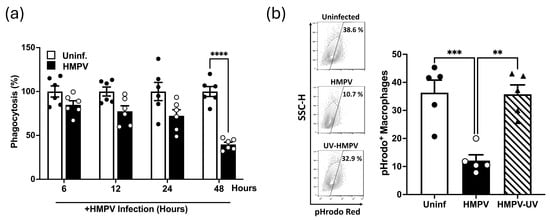
Figure 1
Open AccessArticle
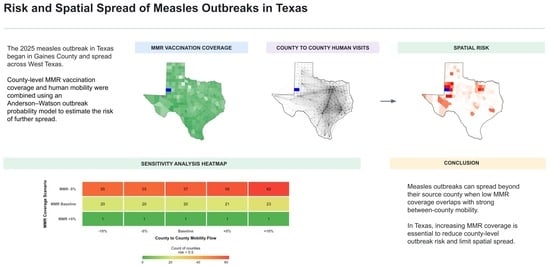
Risk and Spatial Spread of a Measles Outbreak in Texas
by
Martial Loth Ndeffo-Mbah, Sina Mokhtar, Abhishek Pandey and Chad Richard Wells
Viruses 2026, 18(6), 648; https://doi.org/10.3390/v18060648 - 4 Jun 2026
Abstract
In January 2025, a measles outbreak was reported in Gaines County, Texas, and subsequently spread to other counties and states. However, investigations into the geographic spread of this outbreak remain limited. We developed a measles transmission model parameterized with 2020–2024 measles–mumps–rubella (MMR) vaccination
[...] Read more.
In January 2025, a measles outbreak was reported in Gaines County, Texas, and subsequently spread to other counties and states. However, investigations into the geographic spread of this outbreak remain limited. We developed a measles transmission model parameterized with 2020–2024 measles–mumps–rubella (MMR) vaccination coverage and human mobility data in Texas. We conducted sensitivity analyses to evaluate how variation in model parameters affects outcomes. We compared model predictions to data from the 2025 measles outbreak and simulated scenarios for outbreak originating in different low vaccination counties. We found that an outbreak originating in Gaines County would have at least 80% probability of directly generating local outbreaks in eight neighboring counties in West Texas. The spatial spread was highly sensitive to the basic reproduction number (
(This article belongs to the Special Issue Current: Measles Outbreak, a Global Situation)
►▼
Show Figures
Graphical abstract
Open AccessReview
The Begomovirus Disease Tetrahedron: Weeds as the Missing Dimension in Virus Epidemiology
by
Marjia Tabassum, Thuy T. B. Vo, Nattanong Bupi, Muhammad Amir Qureshi, Hyo-Jin Im, Min-Kwan Kim, Imankul Assem, S. M. Hemayet Jahan, Li-Long Pan, Giuseppe Parrella, Peter Palukaitis, Taek-Kyun Lee and Sukchan Lee
Viruses 2026, 18(6), 647; https://doi.org/10.3390/v18060647 - 4 Jun 2026
Abstract
Begomoviruses are among the most destructive plant viruses, causing substantial yield losses across diverse cropping systems. Their epidemiological success is driven by high genetic plasticity, broad host range, and efficient transmission by the whitefly vector Bemisia tabaci. Traditional epidemiological models based on
[...] Read more.
Begomoviruses are among the most destructive plant viruses, causing substantial yield losses across diverse cropping systems. Their epidemiological success is driven by high genetic plasticity, broad host range, and efficient transmission by the whitefly vector Bemisia tabaci. Traditional epidemiological models based on the classical disease triangle (virus–host–vector) fail to fully capture the ecological and evolutionary complexity of begomovirus pathosystems. Increasing evidence highlights the critical role of non-cultivated plants, particularly weeds, as persistent reservoirs that maintain viral populations during off seasons, facilitate recombination, and act as primary inoculum sources for subsequent outbreaks. Here, we propose the Begomovirus Disease Tetrahedron, an integrative framework that expands the disease triangle by incorporating weeds as a fourth essential component. We synthesize current knowledge on begomovirus adaptive evolution, including genome plasticity, noncanonical protein functions, and virus–vector mutualism, alongside key ecological drivers such as seasonal dynamics, agricultural intensification, and landscape connectivity. By integrating molecular, ecological, and epidemiological perspectives, this framework provides a comprehensive understanding of begomovirus emergence and persistence, offering new insights for the development of sustainable and ecologically informed disease management strategies.
Full article
(This article belongs to the Special Issue Molecular and Biological Virus-Plant-Insect Vector Interactions)
►▼
Show Figures

Figure 1
Open AccessArticle
Comparative Analysis of Transcription Factor Binding Sites in the Long Control Region Across Human Papillomavirus Types
by
Derrin Bright and Juan I. Fuxman Bass
Viruses 2026, 18(6), 646; https://doi.org/10.3390/v18060646 - 4 Jun 2026
Abstract
Human papillomaviruses (HPVs) comprise more than 200 types associated with diverse clinical outcomes, ranging from benign lesions caused by low-risk types to cancers driven by high-risk types. These differences are partly driven by variation in the Long Control Region (LCR), a non-coding element
[...] Read more.
Human papillomaviruses (HPVs) comprise more than 200 types associated with diverse clinical outcomes, ranging from benign lesions caused by low-risk types to cancers driven by high-risk types. These differences are partly driven by variation in the Long Control Region (LCR), a non-coding element that regulates viral gene expression through interactions with viral and host transcription factors (TFs). Although individual TF binding sites have been mapped in a few well-studied HPV types, the broader regulatory differences between high-risk and low-risk HPVs remain poorly defined. Here, we systematically analyzed LCR sequences from 207 HPV types using TF motif scanning and identified 104 TFs with significantly different binding site densities between risk groups. Integration with TCGA transcriptomics data showed that 50 of 69 TF enriched in high-risk types are expressed in HPV-positive head and neck tumors (HNSC) and 53 in HPV-positive cervical tumors (CESC). Analysis of published ChIP-seq datasets further confirmed LCR occupancy for seven of these TFs in HPV18-positive cells. In addition, conservation analysis across clinical isolates of HPV-16 and HPV-18 identified highly conserved TF binding sites overlapping multiple high-risk-enriched TF motifs, suggesting functional constraint on key regulatory elements. Together, these findings reveal distinct TF binding landscapes associated with HPV risk groups and identify candidate host regulators that may contribute to differences in viral transcriptional programs and oncogenic potential across HPV types.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
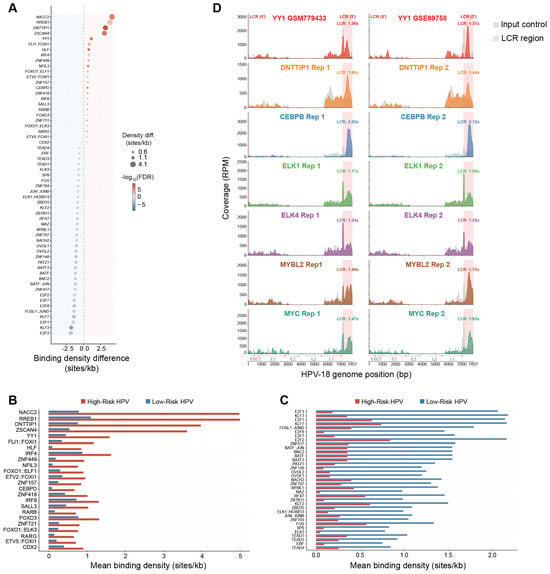
Figure 1
Open AccessArticle
In Silico Saturation-Mutagenesis-Based Genomic Mutation Risk Assessment for Enterovirus B
by
Linglin Wang, Jiajie Tang, Yongtao Jia, Xiaoxiang Tong, Xiaofeng Ying, Qin Chen and Changzheng Dong
Viruses 2026, 18(6), 645; https://doi.org/10.3390/v18060645 - 3 Jun 2026
Abstract
Enterovirus B (EVB) is the most prevalent species of human enteroviruses, responsible for a wide range of diseases, including hand, foot, and mouth disease, viral meningitis, myocarditis, and neonatal sepsis, imposing a significant disease burden primarily on children. Coxsackievirus B (CVB1-6) and various
[...] Read more.
Enterovirus B (EVB) is the most prevalent species of human enteroviruses, responsible for a wide range of diseases, including hand, foot, and mouth disease, viral meningitis, myocarditis, and neonatal sepsis, imposing a significant disease burden primarily on children. Coxsackievirus B (CVB1-6) and various echovirus (E) serotypes are the major serotypes of EVB. Since no antiviral drug or vaccine is available, it is important to strengthen monitoring, risk assessment, and early warning of genomic variations for EVB. CVB1, CVB3, E6, and E30 were selected as representative EVB serotypes for this study due to the availability of three-dimensional structures and their global prevalence. To evaluate the mutation effects of structural proteins on structural stability and receptor-binding affinity, computational saturation mutagenesis of EVB serotypes was performed using FoldX. Furthermore, based on data from deep mutational scanning for CVB3, a risk prediction model for EVB fitness was constructed by machine learning algorithms and applied to other EVB serotypes. Finally, we integrated three phenotypes—structural stability, receptor-binding affinity and fitness—to evaluate genomic variation risk of EVB and tracked the prevalence of high-risk mutants in natural viral sequences through molecular evolution analysis and mutation profiles. We identified the N-terminus and C-terminus of VP1 and the EF loop of VP2 as the EVB regions of highest genomic variation risk, and high-risk mutations had played significant roles in viral evolutionary history. These findings provide a framework for multi-phenotypic and multi-data approaches to viral risk assessment and offer insights to support the development of antiviral drugs and vaccines.
Full article
(This article belongs to the Special Issue Coxsackieviruses, Polioviruses and Associated Diseases (Second Edition))
►▼
Show Figures
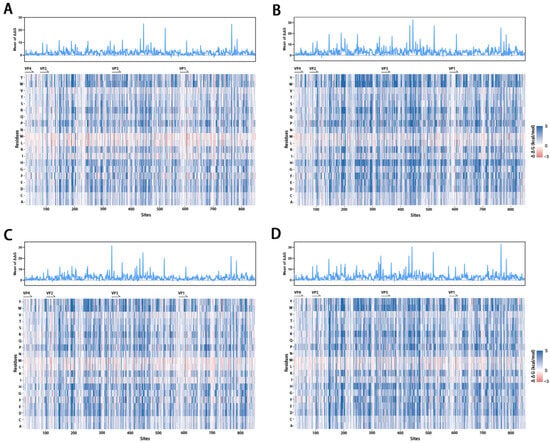
Figure 1
Open AccessArticle
One Health Genomic Surveillance at Human–Animal Interfaces in Rural Ghana Reveals Underreported Viruses of Zoonotic and Economic Concern
by
Julia E. Paoli, Nídia S. Trovão, Theophilus Odoom, Quaneeta Mohktar, Kwame Boamah Buabeng, Bright Adu, William Tasiame, Benita Anderson, Daniel Nana Yaw Tawiah-Yingar, Kuttichantran Subramaniam, Michael E. von Fricken, Gloria Ivy Mensah, Mario Mietzsch, Robert McKenna, Sherry Ama Mawuko Johnson and Carla N. Mavian
Viruses 2026, 18(6), 644; https://doi.org/10.3390/v18060644 - 3 Jun 2026
Abstract
Under a One Health framework, viruses of veterinary and zoonotic importance pose significant threats to animal and human health, food security, and livelihoods, particularly in regions with intense human–animal interactions. In West Africa, despite recent advances in surveillance programs, important gaps remain in
[...] Read more.
Under a One Health framework, viruses of veterinary and zoonotic importance pose significant threats to animal and human health, food security, and livelihoods, particularly in regions with intense human–animal interactions. In West Africa, despite recent advances in surveillance programs, important gaps remain in understanding viral diversity and cross-species transmission at wildlife–livestock interfaces. We conducted metagenomic surveillance to characterize viruses circulating across livestock, domestic animals, and wildlife in rural Ghana in 165 animals sampled across five regions. Viral RNA from serum and tissue samples was sequenced with the Illumina platform, and genomes were de novo assembled with MEGAHIT. Phylogenetic relationships were reconstructed using Bayesian approaches. We report the first genomic sequences of porcine parvovirus 3, canine parvovirus, rotavirus A genotype R16, and bovine hepacivirus subtype B from Ghana in over a decade. Phylogenetic analyses revealed intercontinental linkages between Africa and Europe for parvoviruses, persistence of hepacivirus lineages, and evidence of cross-species transmission for rotavirus. Notably, detection in apparently healthy animals highlights underrecognized circulation, gaps in vaccination effectiveness, trade-related biosecurity vulnerabilities, and the role of wildlife in viral maintenance and transmission. Our findings reveal dynamic viral diversity and connectivity across animal populations and ecological interfaces, emphasizing the fluid and interconnected nature of pathogen circulation within One Health systems. By integrating metagenomics and phylogenetics, this study provides a scalable framework for enhancing surveillance capacity, enabling the early detection of emerging threats and informing targeted strategies to mitigate zoonotic and economically important viral diseases in West Africa.
Full article
(This article belongs to the Special Issue Controlling Zoonotic Viral Diseases from One Health Perspective 2026)
►▼
Show Figures
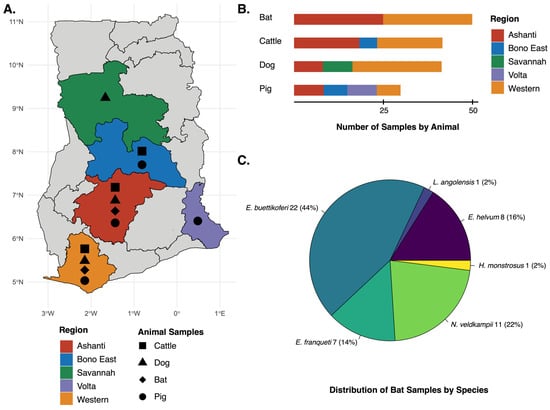
Figure 1
Open AccessTechnical Note
MGtree: A Fast and Flexible Alignment-Based Metagenomics Pipeline
by
Samantha L. Sholes, Scott Norton, Alfredo Gonzalez and John M. Gaspar
Viruses 2026, 18(6), 643; https://doi.org/10.3390/v18060643 - 3 Jun 2026
Abstract
Metagenomics analysis is a critical tool in identifying and typing viral samples to aid surveillance, clinical, epidemiological, and other workflows. Despite advances in sequencing technology and analysis pipelines, there are still limitations that lead to reduced taxonomic resolution or false positives from highly
[...] Read more.
Metagenomics analysis is a critical tool in identifying and typing viral samples to aid surveillance, clinical, epidemiological, and other workflows. Despite advances in sequencing technology and analysis pipelines, there are still limitations that lead to reduced taxonomic resolution or false positives from highly recombinant or challenging samples. Here we describe MGtree, a novel metagenomics pipeline that utilizes a combination of full-length read alignments and phylogenetic analysis to classify samples of interest. We demonstrate that MGtree accurately genotypes viral samples from challenging norovirus and HPV datasets. MGtree outperforms the popular metagenomics programs Kraken2 and Centrifuge, and it succeeds with low-input samples where de novo assembly fails. MGtree’s correct assignments across highly mutant and coinfected samples highlights its ability to resolve viral genotypes and its potential to improve classification precision in complex samples.
Full article
(This article belongs to the Section General Virology)
►▼
Show Figures
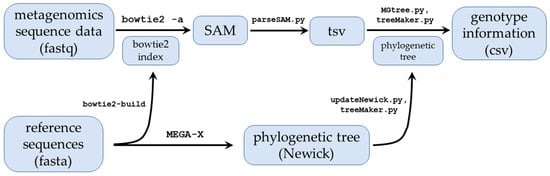
Figure 1
Open AccessReview
Broad Neutralizing Antibodies Against SARS-CoV-2: Current Progress and Engineering Strategies
by
Tianrong Jia, Zehong Huang, Ningshao Xia and Quan Yuan
Viruses 2026, 18(6), 642; https://doi.org/10.3390/v18060642 - 3 Jun 2026
Abstract
The high-frequency mutation characteristics of SARS-CoV-2 have posed formidable challenges to the development of vaccines and therapeutic agents. Neutralizing antibodies, which serve as effective tools for prevention and control, have undergone continuous updates and iterations in response to viral mutations. This article provides
[...] Read more.
The high-frequency mutation characteristics of SARS-CoV-2 have posed formidable challenges to the development of vaccines and therapeutic agents. Neutralizing antibodies, which serve as effective tools for prevention and control, have undergone continuous updates and iterations in response to viral mutations. This article provides a comprehensive review of researchers’ efforts to achieve both high neutralizing potency and high mutation tolerance in SARS-CoV-2–targeting neutralizing antibodies. Building on the characteristics of conventional antibodies directed against distinct epitopes on the S protein, it further discusses the research on nanobodies, antibody cocktails, multi-specific antibodies, and other antibody formats and engineering approaches, including artificial intelligence–enabled optimization. Each antibody-based strategy targeting SARS-CoV-2 has its own distinctive advantages and potential applications, providing an integrated perspective to support the continued development of antiviral neutralizing antibodies.
Full article
(This article belongs to the Section Coronaviruses)
►▼
Show Figures
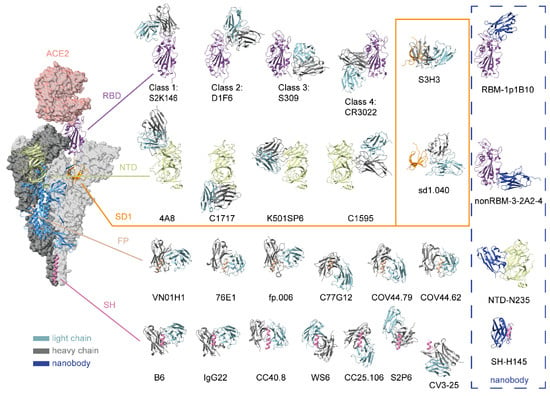
Figure 1
Open AccessArticle
Cathepsin C Restricts Influenza A Virus Replication and Is Associated with Suppression of the PI3K-AKT Signaling Pathway
by
Yansheng Zhu, Lanlan Si, Zhongzhong Cao, Huiyang Song, Aiqing Yang, Xia Wang, Yifei Qiu, Chengming Gao, Gangqiao Zhou and Pengbo Cao
Viruses 2026, 18(6), 641; https://doi.org/10.3390/v18060641 - 3 Jun 2026
Abstract
Cathepsin C (CTSC) is a major lysosomal cysteine protease characterized by its involvement in multiple essential pathological processes associated with various viral infections and pathogenesis, such as influenza virus and coronavirus. However, the antiviral spectrum of CTSC and the molecular mechanisms underlying its
[...] Read more.
Cathepsin C (CTSC) is a major lysosomal cysteine protease characterized by its involvement in multiple essential pathological processes associated with various viral infections and pathogenesis, such as influenza virus and coronavirus. However, the antiviral spectrum of CTSC and the molecular mechanisms underlying its activity remain to be fully elucidated. In this study, we demonstrate that CTSC significantly inhibits the infection of influenza A virus (IAV) H1N1 both in vitro and in vivo. Mechanistically, the antiviral function of CTSC is associated with attenuation of the PI3K-AKT signaling pathway, thereby inducing cell apoptosis and reducing inflammation, which ultimately limits the virus’s ability to hijack host resources. Taken together, our findings highlight the crucial role of CTSC in defense against H1N1 by targeting PI3K-AKT pathway and suggest a prospective antiviral target against the infection of H1N1.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Development and Application of a Fluorescence Quantitative RT-PCR Assay for Detection of Group A Rotavirus Among Pigeons in China from 2023 to 2024
by
Xiaohui Yu, Cuiyao Zhang, Lu Chen, Jingjing Wang, Yufeng Liu, Yuteng Chen, Jinping Li, Guangyu Hou, Shuang Chang, Leilei Sun and Yang Li
Viruses 2026, 18(6), 640; https://doi.org/10.3390/v18060640 - 3 Jun 2026
Abstract
Group A rotavirus (RVA), as an important pathogen causing pigeon diarrhea, has caused substantial economic losses for the pigeon breeding industry in recent years in China. To better understand the prevalence and genetic diversity of RVAs among pigeon flocks in China, a fluorescence
[...] Read more.
Group A rotavirus (RVA), as an important pathogen causing pigeon diarrhea, has caused substantial economic losses for the pigeon breeding industry in recent years in China. To better understand the prevalence and genetic diversity of RVAs among pigeon flocks in China, a fluorescence quantitative RT-PCR assay for detection of RVAs was developed in this study. The fluorescence quantitative RT-PCR assay showed satisfactory specificity, sensitivity, repeatability, and reproducibility, with a limit of detection (LOD) of 1 × 102 copies/μL for RVA. Then, 645 pigeon swab samples collected from live bird markets in 10 provinces of China from 2023 to 2024 were tested for RVA with the fluorescence quantitative RT-PCR assay established in this study to characterize the current epidemiological status of pigeon-origin RVAs (PiRVAs). The results showed that the PiRVA positivity rate detected by the RT-qPCR assay was 12.24% (79/645) among pigeon flocks in China. Guizhou Province had the highest positive rate of PiRVAs (42.86%) among the 10 provinces sampled. The VP7 and VP4 gene of nine representative positive pigeon RVA samples were amplified by specific primers, and the phylogenetic analysis of these samples revealed that all RVAs among pigeons belonged to the G18P[17] genotype, though with certain genetic variations. This study established a useful tool for diagnosing Pigeon RVAs in practice, and provided valuable information to better understand the epidemiology of Pigeon RVAs among pigeon flocks in China.
Full article
(This article belongs to the Special Issue Avian Viruses and Antiviral Immunity)
►▼
Show Figures
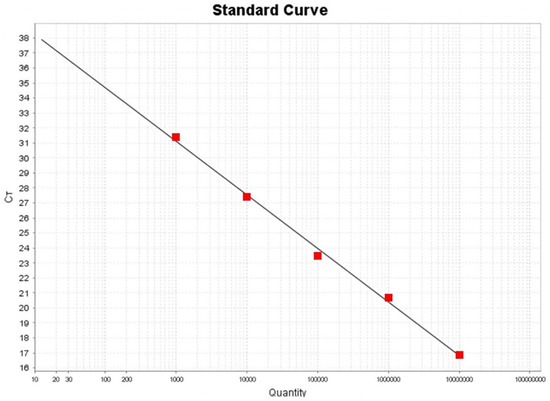
Figure 1
Open AccessReview
DNA Vaccination in Farmed Fish with a Focus on Salmonid Alphavirus Infection in Atlantic Salmon (Salmo salar L.) in Norway
by
Espen Rimstad and Øystein Evensen
Viruses 2026, 18(6), 639; https://doi.org/10.3390/v18060639 - 3 Jun 2026
Abstract
Pancreas disease (PD), caused by Salmonid alphavirus (SAV), is a notifiable disease in Atlantic salmon (Salmo salar L.) in Norway. Conventional inactivated virus vaccines have shown variable effects in mitigating the disease, and a DNA vaccine has been used over the last
[...] Read more.
Pancreas disease (PD), caused by Salmonid alphavirus (SAV), is a notifiable disease in Atlantic salmon (Salmo salar L.) in Norway. Conventional inactivated virus vaccines have shown variable effects in mitigating the disease, and a DNA vaccine has been used over the last 7–8 years, which may have resulted in the reduction in the number of reported PD cases. This manuscript provides a comprehensive overview of DNA vaccination in farmed fish, with a focus on the licensed DNA vaccine, Clynav®, against SAV3 infection. It explores the biological underpinnings of SAV infection, immune mechanisms activated by DNA vaccines, and the benefits and limitations of this approach. Although antigen processing and presentation mechanisms following DNA vaccination in fish remain incomplete, studies document robust innate responses and measurable adaptive immunity, including neutralizing antibodies, as seen in Clynav, and transcriptomic studies indicate that cell-mediated immunity is evoked under experimental conditions. Comparative trials demonstrate that DNA vaccination reduces viral load, tissue pathology, and, potentially, viral transmission, outperforming traditional oil-adjuvanted vaccines. Additionally, DNA-vaccinated fish show improved growth performance under field conditions. These findings support DNA vaccination as a promising strategy for controlling PD in salmon aquaculture, with implications for fish health, welfare, and sustainable production.
Full article
(This article belongs to the Special Issue Viral Pathogenesis and Novel Vaccines for Fish Viruses)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomedicines, Cancers, IJMS, Vaccines, Viruses, JMP
Advances in HPV-Driven Head and Neck Cancer: Pathogenesis, Treatment, and Prevention
Topic Editors: Shilpi Gupta, Bhudev Chandra Das, Prabhat KumarDeadline: 20 April 2027
Topic in
Antibiotics, Diseases, Vaccines, Viruses, Epidemiologia
HIV Prevention and Treatment: A Guide to Health and Safety
Topic Editors: Olanrewaju Oladimeji, Martha ChadyiwaDeadline: 31 December 2027

Conferences
Special Issues
Special Issue in
Viruses
Virology in Italy 2025—9th National Congress of the Italian Society for Virology
Guest Editors: Enzo Tramontano, Arnaldo Caruso, Massimiliano Galdiero, Luisa Rubino, Gabriele VaccariDeadline: 12 June 2026
Special Issue in
Viruses
Discovery, Classification, and Early Research on the Lipid-Containing, Double-Stranded RNA Bacteriophage φ6
Guest Editors: Paul Gottlieb, Aleksandra AlimovaDeadline: 15 June 2026
Special Issue in
Viruses
Role of Bacteriophage in Intestine Microbial Communities
Guest Editor: Yi DuanDeadline: 15 June 2026
Special Issue in
Viruses
Functional Structures in RNA Viruses
Guest Editors: Riccardo Delli Ponti, Gian Gaetano Tartaglia, Andrea Vandelli, Laura BrogliaDeadline: 15 June 2026
Topical Collections
Topical Collection in
Viruses
Efficacy and Safety of Antiviral Therapy
Collection Editors: Giordano Madeddu, Andrea De Vito, Agnese Colpani
Topical Collection in
Viruses
Phage Therapy
Collection Editors: Nina Chanishvili, Jean-Paul Pirnay, Mikael Skurnik