-
 Probiotic and Postbiotic Interactions of Lactobacillus Strains with Candida albicans: Antifungal Effects Through Microbial Competition
Probiotic and Postbiotic Interactions of Lactobacillus Strains with Candida albicans: Antifungal Effects Through Microbial Competition -
 IBS and SIBO: Gut Microbiota, Pathophysiology, and Non-Pharmacological Interventions
IBS and SIBO: Gut Microbiota, Pathophysiology, and Non-Pharmacological Interventions -
 Impact of an Antimicrobial Stewardship Program on Antibiotic Consumption, Bacterial Susceptibility, and Costs in a High-Complexity Public Hospital
Impact of an Antimicrobial Stewardship Program on Antibiotic Consumption, Bacterial Susceptibility, and Costs in a High-Complexity Public Hospital
Journal Description
Antibiotics
Antibiotics
is an international, peer-reviewed, open access journal on all aspects of antibiotics, published monthly online by MDPI. The Croatian Pharmacological Society (CPS) and Northern Greece Society of Medical Biopathology (EIBBE) are affiliated with Antibiotics, and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Infectious Diseases) / CiteScore - Q1 (General Pharmacology, Toxicology and Pharmaceutics )
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.4 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
4.6 (2024);
5-Year Impact Factor:
4.9 (2024)
Latest Articles
Clinical Considerations of Amikacin Pharmacotherapy in Adults—A Narrative Review with Focus on Safety and TDM
Antibiotics 2026, 15(6), 534; https://doi.org/10.3390/antibiotics15060534 (registering DOI) - 24 May 2026
Abstract
Background: Amikacin remains a key agent in the treatment of severe and complicated infections due to its bactericidal activity and low risk of Clostridioides difficile infection. It retains activity against most aerobic Gram-negative bacteria, including multidrug-resistant Enterobacterales and Pseudomonas. However, its use is
[...] Read more.
Background: Amikacin remains a key agent in the treatment of severe and complicated infections due to its bactericidal activity and low risk of Clostridioides difficile infection. It retains activity against most aerobic Gram-negative bacteria, including multidrug-resistant Enterobacterales and Pseudomonas. However, its use is limited by nephrotoxicity and ototoxicity. Methods: This narrative review evaluates clinical indications, pharmacokinetic and pharmacodynamic properties, dosing strategies, therapeutic drug monitoring (TDM), and safety profile of amikacin in adult patients based on 56 selected publications. A total of 24 articles were identified through database searches (PubMed and Embase), complemented by 32 additional sources to provide clinical and pharmacological context. Results: Available evidence demonstrates considerable uncertainty regarding the comparative effectiveness of different monitoring strategies. Lower trough concentrations are generally associated with reduced nephrotoxicity; however, an optimal safety threshold has not been clearly established. Guideline-recommended targets vary substantially and are supported by low-quality evidence. Amikacin pharmacokinetics, tissue penetration and toxicity are influenced by patient-specific factors, including critical illness, renal function variability, and concomitant nephrotoxic therapy, particularly vancomycin. Ototoxicity remains an additional clinically relevant concern. Conclusions: Current evidence suggests that uniform dosing and monitoring paradigms are insufficient. Patient-tailored strategies integrating TDM and mitigation of modifiable risk factors are required. Prospective studies comparing monitoring regimens are needed to optimize the safe clinical use of amikacin and inform future guideline development.
Full article
(This article belongs to the Special Issue Feature Reviews in Antibiotic Therapy for Infectious Diseases 2026)
Open AccessArticle
Carbapenem-Resistant Klebsiella pneumoniae: Carbapenemase Production, Antibiotic Resistance and Treatment Options, in an Infectious Diseases Hospital from Romania
by
Alexandra Cireșă, Gabriel-Adrian Popescu, Daniela Tălăpan, Mihai Octavian Dan and Cristina Popescu
Antibiotics 2026, 15(6), 533; https://doi.org/10.3390/antibiotics15060533 (registering DOI) - 24 May 2026
Abstract
Background/Objectives: Carbapenem-resistant Klebsiella pneumoniae (CRKP) is of great concern because of the difficulties encountered in the management of infections it may cause. This study aims to identify possible difficulties in the management of K. pneumoniae infections in the current context of antibiotic resistance,
[...] Read more.
Background/Objectives: Carbapenem-resistant Klebsiella pneumoniae (CRKP) is of great concern because of the difficulties encountered in the management of infections it may cause. This study aims to identify possible difficulties in the management of K. pneumoniae infections in the current context of antibiotic resistance, particularly regarding carbapenem resistance. Methods: This is a retrospective, cross-sectional study that analyses epidemiological, clinical and bacteriological features identified in all patients with CRKP infections/colonization admitted during 2024 in an infectious diseases hospital. Results: Carbapenemase-producing K. pneumoniae isolates were co-harboring NDM+OXA-48 in 55.2% of cases. NDM+OXA-48-producing K. pneumoniae (116 isolates, 55.2%) was correlated with high resistance to aztreonam (100%, p = 0.01), ceftazidime–avibactam (100%, p < 0.01), trimethoprim–sulfamethoxazole (99.1%, p < 0.01), gentamycin (94.8%, p < 0.01), amikacin (93.8%, p < 0.01), colistin (79.8%, p < 0.01). OXA-48-producing K. pneumoniae (29 isolates, 13.8%) was correlated with lower resistance to ceftazidime–avibactam (11.5%, p < 0.01), amikacin (48.1%, p < 0.01), colistin (51.7%, p = 0.01), and gentamycin (65.5%, p < 0.01). We found in vitro synergistic effects of ceftazidime/avibactam + aztreonam for 32/32 CRKP isolates and of colistin + tigecycline for 12/14 CRKP isolates. Higher recurrence of CRKP infections was recorded in patients with urinary tract conditions (RR = 11.58, 95%CI: 1.58–81.91) and upper urinary tract devices (RR = 3.53, 95% CI: 1.72–7.22). In this study, adequate antibiotic treatment, compared to excessive antibiotic treatment in CRKP infections, was associated with shorter treatment duration (p = 0.02) and shorter length of hospitalization (p = 0.04). Conclusions: In our study, CRKP is frequently coharboring NDM+OXA-48, having limited treatment options. Implementing new treatment strategies, testing antibiotic synergies for older antibiotics in order to identify alternative treatment options and avoiding unnecessary carbapenem consumption are essential for decreasing the burden of CRKP infections.
Full article
(This article belongs to the Special Issue The Issue of Multidrug-Resistant Pathogens in Nosocomial Infections, 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
Colistin Resistance in Acinetobacter baumannii Clinical Isolates from Bahrain: Evaluation of Detection Methods and Clonal Relationships
by
Zainab Husain Salman, Mohd Shadab, Zainab Salman Saleh, Nouf Al-Rashed and Mohammad Shahid
Antibiotics 2026, 15(6), 532; https://doi.org/10.3390/antibiotics15060532 (registering DOI) - 23 May 2026
Abstract
Background: Acinetobacter baumannii (A. baumannii) is a critical-priority pathogen of major concern in healthcare settings. Colistin remains a last-resort antibiotic for multidrug-resistant (MDR) A. baumannii infections; however, resistance is increasingly reported worldwide yet remains understudied in Bahrain. Reliable
[...] Read more.
Background: Acinetobacter baumannii (A. baumannii) is a critical-priority pathogen of major concern in healthcare settings. Colistin remains a last-resort antibiotic for multidrug-resistant (MDR) A. baumannii infections; however, resistance is increasingly reported worldwide yet remains understudied in Bahrain. Reliable detection methods and understanding clonal dissemination are essential for infection control. Objectives: This study aimed to (1) determine the rate of colistin resistance in 102 clinical A. baumannii isolates from Bahrain, (2) evaluate the diagnostic performance of the colistin agar test (CAT) and E-test against broth microdilution (BMD method), and (3) assess clonal relationships using BOX-PCR fingerprinting. Methods: 102 clinical isolates from multiple hospitals in Bahrain underwent susceptibility testing via the BMD method, CAT, and E-test; screening for mcr-1 to mcr-5 genes; and BOX-PCR DNA fingerprinting. Results: Colistin resistance was detected in 14.7% of isolates by BMD method, higher than regional and global averages. All resistant isolates were mcr-negative, suggesting chromosomally mediated resistance. CAT showed 86.7% sensitivity, 98.8% specificity, and a 13.3% very major error rate. The E-test failed to detect resistant isolates (very major error 100%). BOX-PCR revealed predominant clonal relatedness with intra- and inter-hospital spread. Conclusions: Colistin resistance in A. baumannii from Bahrain exceeds regional and global levels, likely driven by chromosomal mechanisms under selective pressure. The BMD method remains the gold standard for colistin testing, while CAT may serve as a screening tool requiring confirmation. Strengthened stewardship and infection control measures are vital to contain dissemination.
Full article
Open AccessArticle
Effectiveness and Safety of First-Line Empirical Eradication Therapy with Rebamipide: Results from the European Registry on Helicobacter pylori Management (Hp-EuReg)
by
Dmitry S. Bordin, Dmitrii N. Andreev, Sayar R. Abdulkhakov, Irina N. Voynovan, Igor G. Bakulin, Natalia V. Bakulina, Ludmila G. Vologzhanina, Boris D. Starostin, Aiman S. Sarsenbayeva, Tatyana A. Ilchishina, Larisa V. Tarasova, Alla G. Kononova, Sergey A. Alekseenko, Oleg V. Zaytsev, Natalya N. Dekhnich, Pavel O. Bogomolov, Galina N. Tarasova, Olga A. Kolokolnikova, Igor B. Khlynov, Ekaterina Yu. Plotnikova, Natalia V. Baryshnikova, Maria A. Livzan, Marina F. Osipenko, Natalya V. Bakanova, Ekaterina V. Partsvania-Vinogradova, Margarita V. Chebotareva, Rustam A. Abdulkhakov, Inna V. Savilova, Anastasia G. Sushilova, Sergey V. Kolbasnikov, Alsu R. Khurmatullina, Igor V. Maev, Maria O. Tishchenko, Rimma E. Petrova, Ludmila V. Morkovkina, Yulia V. Tsyganova, Мargarita D. Kozlova, Aleksei A. Iakovlev, Alexander S. Volkov, Elizaveta S. Moroz, Olga V. Gaus, Singla Alba, Oriol Farrés, Leticia Moreira, Pablo Parra, Olga P. Nyssen, Francis Megraud, Colm O’Morain and Javier P. Gisbertadd
Show full author list
remove
Hide full author list
Antibiotics 2026, 15(6), 531; https://doi.org/10.3390/antibiotics15060531 (registering DOI) - 23 May 2026
Abstract
Background/Objective: Eradication of Helicobacter pylori typically uses acid-suppressing agents plus antimicrobials, but treatment failure may reach 20–30%. Adjunctive rebamipide has been proposed as a strategy to improve treatment outcomes. This study aimed to assess the effectiveness and safety of first-line empirical therapy when
[...] Read more.
Background/Objective: Eradication of Helicobacter pylori typically uses acid-suppressing agents plus antimicrobials, but treatment failure may reach 20–30%. Adjunctive rebamipide has been proposed as a strategy to improve treatment outcomes. This study aimed to assess the effectiveness and safety of first-line empirical therapy when rebamipide was added to eradication regimens. Methods: Treatment-naïve patient data were collected between January 2020 and August 2024 from the European Registry on Helicobacter pylori Management (Hp-EuReg), an international, prospective, multicentre, observational study. Rebamipide-containing regimens were analysed and compared with those without rebamipide. Effectiveness was assessed using a modified intention-to-treat approach. Results: Overall, 4581 patients (mean age 48 ± 15 years, 64% women) from 15 Russian cities were included. From those, 820 patients received eradication therapy with rebamipide and 3761 without rebamipide. Overall effectiveness was significantly higher with rebamipide (731/763, 96%) than without it (2785/3059, 91%) (p = 0.0002). Rebamipide-containing triple therapy consisting of a proton-pump inhibitor (PPI), clarithromycin and amoxicillin was associated with effectiveness above 90%, while rebamipide-containing quadruple therapy consisting of a PPI, clarithromycin, amoxicillin and bismuth yielded a 97% eradication rate, compared with 79% and 95%, respectively, without rebamipide (p < 0.001). Adverse events were significantly more frequent in patients receiving rebamipide-containing regimens (46% vs. 29%, p < 0.0001), though none were serious or caused discontinuation. Compliance was similar in both groups (99% vs. 98.5%; p > 0.05). Conclusions: In this observational registry cohort, first-line empirical triple therapy with amoxicillin–clarithromycin, and quadruple therapy with amoxicillin–clarithromycin–bismuth, both prescribed with rebamipide, were associated with higher eradication rates than regimens without rebamipide.
Full article
(This article belongs to the Section Antibiotic Therapy in Infectious Diseases)
Open AccessArticle
Comparison of Antimicrobial Susceptibility Patterns of Bacterial Isolates from Blood, Urine, and Lower Respiratory Tract Specimens Between Elderly Patients in Long-Term Care Hospitals and Community-Acquired Infections: A Retrospective Study
by
Kye Won Choe, Sumi Yoon, Yong Kwan Lim, Hongkyung Kim, Mi-Kyung Lee and Oh Joo Kweon
Antibiotics 2026, 15(6), 530; https://doi.org/10.3390/antibiotics15060530 - 22 May 2026
Abstract
Background/Objectives: Patients in long-term care hospitals (LTCHs) are at increased risk of harboring antimicrobial-resistant organisms due to frequent healthcare exposure and multiple comorbidities. This retrospective observational study aimed to compare the antimicrobial susceptibility of bacterial isolates from LTCH-onset infections (LTCHIs) with those from
[...] Read more.
Background/Objectives: Patients in long-term care hospitals (LTCHs) are at increased risk of harboring antimicrobial-resistant organisms due to frequent healthcare exposure and multiple comorbidities. This retrospective observational study aimed to compare the antimicrobial susceptibility of bacterial isolates from LTCH-onset infections (LTCHIs) with those from community-acquired infections (CAIs) in elderly patients. Methods: This study was conducted at a 700-bed urban tertiary university hospital and included patients aged ≥65 years with positive cultures for bacteremia, lower respiratory tract infections (LRTIs), or urinary tract infections (UTIs) within 48 h of admission. Medical records, including antimicrobial susceptibility test results, were reviewed for a total of 1780 patients and their isolates. Antimicrobial susceptibility patterns were compared between LTCHI and CAI patients. Results: Patients with LTCHI exhibited significantly higher antimicrobial non-susceptibility than those with CAIs across multiple pathogens and antimicrobial classes (p < 0.05). In bacteremia, Staphylococcus aureus, Escherichia coli, and Klebsiella pneumoniae from LTCHI cases showed increased non-susceptibility to β-lactams and fluoroquinolones. In LRTIs, Pseudomonas aeruginosa and Acinetobacter baumannii demonstrated high non-susceptibility to carbapenems (52.9% and 90%, respectively) and aminoglycosides. In UTIs, LTCHI isolates exhibited broader resistance among Enterobacterales and P. aeruginosa. Notably, the proportion of multidrug-resistant organisms, including carbapenem-resistant Enterobacterales (15.4–50.0%) and carbapenem-resistant Acinetobacter baumannii (90.5%), was substantially higher in the LTCHI group across all infection sites. Conclusions: Elderly patients with LTCHI are more likely to harbor antimicrobial-resistant pathogens than those with CAIs. Careful consideration of LTCHI origin is therefore essential for empirical antibiotic selection and for strategies aimed at limiting further resistance.
Full article
(This article belongs to the Special Issue Antibiotic Resistance in Geriatric Emergency Patients: Challenges and Strategies for Optimal Management)
Open AccessArticle
Multiplex PCR-Based Detection of Eight Carbapenemase Genes and Their Clinical Characteristics in Urinary Tract Infections
by
Nishadi Jayathilaka, Upeksha Kulasekara, Dilini Nakkawita, Dharshan De Silva, Samanmalee Gunasekara and Thamarasi Senaratne
Antibiotics 2026, 15(6), 529; https://doi.org/10.3390/antibiotics15060529 - 22 May 2026
Abstract
Background: The emergence and spread of urinary carbapenem-resistant organisms (CROs) are a major public health concern, particularly in Sri Lanka. Therefore, we aimed to detect and genotypically characterize CROs in urinary tract infections (UTIs) and their clinical outcomes. Methods: Urinary CROs were collected
[...] Read more.
Background: The emergence and spread of urinary carbapenem-resistant organisms (CROs) are a major public health concern, particularly in Sri Lanka. Therefore, we aimed to detect and genotypically characterize CROs in urinary tract infections (UTIs) and their clinical outcomes. Methods: Urinary CROs were collected from two hospitals in Sri Lanka from January to December, 2023. Among 7640 urine samples, 100 CROs were identified by disk diffusion method, and 99 were detected by BD PheonixTM automated system. The presence of eight carbapenemase genes; blaKPC, blaNDM, blaVIM, blaIMP, blaOXA-23, blaOXA-48, blaOXA-51, and blaOXA-58, among 97 CROs was detected by a multiplex PCR kit. Results: Out of 99 urinary CROs, K. pneumoniae (33.3%; n = 33/97) was the most common species. Among the 97 isolates tested by PCR, a single carbapenemase gene was detected in 35.05% (34/97), while two or more genes co-occurred in 39.18% (38/97). The most frequently identified gene was blaOXA-51 (47.4%), followed by blaOXA-58 (41.2%). Most patients (95.74%; n = 90/97) showed clinical improvement within seven days of treatment. Among the 93 patients discharged and followed for three months, 74.20% (n = 69/93) experienced at least one mild UTI recurrence. A total of 10 patients died during the study period. Of which, four (40%) during hospitalization and six (60%) during follow-up, though none of the deaths were attributed to UTIs. Conclusions: K. pneumoniae, showed the highest carbapenemase gene diversity. Recurrent UTIs were observed during the follow-up period. Continuous surveillance and implementation of targeted infection control programs are needed to minimize further emergence and spread of carbapenemase genes.
Full article
(This article belongs to the Section Mechanism and Evolution of Antibiotic Resistance)
Open AccessArticle
Epidemiology of Clostridioides difficile Infection in Argentina and Associated Risk Factors Evaluated Through a Meta-Analysis
by
Angela María Barbero, Nicolás Diego Moriconi, Sabina Palma, Josefina Celano, María Gracia Balbi, Lorenzo Sebastián Morro, María Martina Calvo Zarlenga, Jorgelina Suárez, María Guadalupe Martínez, Mónica Graciela Machain, Carlos Gabriel Altamiranda, Gabriel Erbiti, Rodrigo Emanuel Hernández Del Pino and Virginia Pasquinelli
Antibiotics 2026, 15(6), 528; https://doi.org/10.3390/antibiotics15060528 - 22 May 2026
Abstract
Background: Clostridioides difficile is classified within the first 18 threats for antimicrobial resistance and is the leading cause of hospital-acquired enteric infection. Community-associated cases have notably increased in recent decades, highlighting that accurate and up-to-date statistics characterizing the epidemiology of C. difficile infection
[...] Read more.
Background: Clostridioides difficile is classified within the first 18 threats for antimicrobial resistance and is the leading cause of hospital-acquired enteric infection. Community-associated cases have notably increased in recent decades, highlighting that accurate and up-to-date statistics characterizing the epidemiology of C. difficile infection (CDI) are critical. Methods: We conducted a retrospective (2019–2023) case-control study evaluating the prevalence of CDI in 249 stool samples from hospitalized patients in the sanitary region III of Buenos Aires, Argentina. The presence of C. difficile was detected by combining EIA, PCR, and toxigenic culture via a diagnostic algorithm. Clinical and demographic data from patients were analyzed to identify CDI-associated risk factors. We also conducted a systematic review and a meta-analysis contrasting our results with 38 studies selected from different countries. Results: One in five patients presented C. difficile as the etiological agent of diarrhea. Eighty percent of the CDI+ cases carried toxigenic strains, with a third of cases associated with community environments. Age ≥ 69 years, previous use of antibiotics, previous hospitalization, and previous episodes of CDI emerged as predisposing factors for CDI in our study cohort. In an exploratory evaluation of clinical data, CDI+ patients showed higher leukocytes and platelets counts, a decreased basophil count, and increased urea concentration. At the global level, the meta-analysis reinforced advanced age, previous consumption of antibiotics, previous consumption of proton pump inhibitors, previous hospitalization, and previous CDI as risk factors for CDI. Conclusions: These results emphasize the importance of continued epidemiological surveillance of CDI. Our findings confirm previously described risk factors, both in our cohort and at the global level. Exploratory alterations in laboratory parameters were observed, although their clinical relevance and specificity require further investigation.
Full article
(This article belongs to the Special Issue Clostridioides difficile Infection, 3rd Edition)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Antarctic Fish Antimicrobial Peptides Active Against Bacterial and Viral Pathogens of Aquacultural Importance
by
Federica Massaro, Luana Cortinovis, Romy Lucon Xiccato, Eleonora Fiocchi, Amedeo Manfrin, Anna Rita Taddei, Paolo Roberto Saraceni, Fernando Porcelli, Anna Toffan and Francesco Buonocore
Antibiotics 2026, 15(6), 527; https://doi.org/10.3390/antibiotics15060527 - 22 May 2026
Abstract
Background/Objectives: The aquaculture industry represents a fundamental food sector. One of the main limiting factors for this sector is related to bacterial diseases, for which antibiotics have been widely used worldwide for decades. In recent years, a more conscious approach to the use
[...] Read more.
Background/Objectives: The aquaculture industry represents a fundamental food sector. One of the main limiting factors for this sector is related to bacterial diseases, for which antibiotics have been widely used worldwide for decades. In recent years, a more conscious approach to the use of antimicrobials within the framework of the One Health approach has increased the need for alternatives capable of helping with disease management while avoiding the onset of antimicrobial resistance phenomena. Antimicrobial peptides, which have a broad spectrum of action against pathogens, are a promising solution. Methods: In this work, we investigated the capability of three peptides (Trematocine, Chionodracine, and Cnd-m3) isolated from Antarctic fish to target bacterial and viral pathogens affecting aquaculture. Successively, we investigated their cytotoxicity versus a continuous embryonic cell line (DLEC) derived from European sea bass and their haemolytic activity against fish erythrocytes. Moreover, we evaluated their immunomodulatory effect. Results: Regarding antibacterial properties, Cnd-m3 was identified as the best peptide, demonstrating good bactericidal and bacteriostatic activity against various bacterial strains, including Lactococcus garvieae. Concerning this bacterium, ANS permeability assays showed that the Cnd-m3 peptide has a great ability to interact with its outer membrane, while TEM analysis revealed that the peptide, after destabilization of the cell membrane, interacts with nucleic structures. Considering the antiviral activity, Trematocine was effective against two tested pathogenic enveloped viruses. Moreover, the toxicity of Trematocine and Cnd-m3 was evaluated by investigating their cytotoxicity against a cell line derived from Dicentrarchus labrax and haemolysis against sea bass erythrocytes. Both revealed good selectivity towards pathogens at the lowest concentration. Finally, Cnd-m3 manifested light in vitro immunomodulatory properties. Conclusions: Overall, these data provide a solid basis for future studies assessing the potential applications of two of the tested peptides in aquaculture.
Full article
(This article belongs to the Section Antimicrobial Peptides)
►▼
Show Figures

Figure 1
Open AccessReview
Bacterial Persister Cells as Evolutionary Catalysts of Antibiotic Resistance: Mechanisms, Clinical Implications, and Therapeutic Strategies
by
Tae-Jong Kim
Antibiotics 2026, 15(6), 526; https://doi.org/10.3390/antibiotics15060526 - 22 May 2026
Abstract
Antibiotic resistance is a growing global health threat. However, its evolution cannot be fully understood without considering antibiotic tolerance and persistence. Persister cells are phenotypic variants that survive lethal antibiotic exposure without heritable resistance, primarily through growth arrest, metabolic slowdown, and stress-adaptive states.
[...] Read more.
Antibiotic resistance is a growing global health threat. However, its evolution cannot be fully understood without considering antibiotic tolerance and persistence. Persister cells are phenotypic variants that survive lethal antibiotic exposure without heritable resistance, primarily through growth arrest, metabolic slowdown, and stress-adaptive states. Although persistence has been viewed as a transient survival phenomenon, increasing evidence suggests that it may also have a genetic basis by preserving populations during antibiotic-induced bottlenecks and enabling regrowth, mutation, and selection under certain conditions. This review examines the molecular mechanisms underlying persister formation, including toxin–antitoxin systems, stringent-response signaling, ATP depletion, translational arrest, and stress-response networks. We discuss how persistence contributes to antibiotic tolerance in biofilms, host environments, and recurrent infections, and how repeated antibiotic exposure may promote stepwise evolution from phenotypic survival to stable resistance in specific contexts. Evidence from experimental evolution, clinical observations, and system-level analyses supports a potential but context-dependent link between persistence and resistance. We also highlight therapeutic strategies targeting persister cells, including antipersister compounds, metabolic activation, combination therapies, bacteriophages, and alternative approaches. Finally, we outline future research directions, emphasizing single-cell technologies, systems biology, longitudinal clinical studies, and evolution-informed treatment design. A comprehensive understanding of persistence and its evolutionary implications is essential for improving treatment efficacy and limiting the emergence of long-term antibiotic resistance.
Full article
(This article belongs to the Special Issue Feature Reviews in Mechanism and Evolution of Antibiotic Resistance 2026)
►▼
Show Figures

Figure 1
Open AccessArticle
Sustainability and Impact of an Antimicrobial Stewardship Program on Broad-Spectrum Antibiotic Consumption in South Korea: A 14-Month Extended Follow-Up Study
by
Tae-Hoon No and Kyeong Min Jo
Antibiotics 2026, 15(6), 525; https://doi.org/10.3390/antibiotics15060525 - 22 May 2026
Abstract
Background: Antimicrobial stewardship programs (ASPs) are critical for promoting rational antibiotic use. While early implementation outcomes have been reported, extended follow-up sustainability and the impact on high-priority broad-spectrum antibiotics in South Korean secondary/tertiary hospitals require further validation. This study aimed to evaluate the
[...] Read more.
Background: Antimicrobial stewardship programs (ASPs) are critical for promoting rational antibiotic use. While early implementation outcomes have been reported, extended follow-up sustainability and the impact on high-priority broad-spectrum antibiotics in South Korean secondary/tertiary hospitals require further validation. This study aimed to evaluate the extended outcomes and sustainability of an ASP over a 14-month period. Methods: This retrospective, single-center study analyzed ASP activities from January 2025 to February 2026 at a tertiary hospital in South Korea. Interventions included prospective audit and feedback (PAF) for restricted antibiotics and recommendations for prolonged prescriptions (≥14 days). Primary outcomes were the monthly rejection rate of restricted antibiotics and the acceptance rate of ASP interventions. Secondary outcomes included the days of therapy (DOT) per 1000 patient–days for meropenem and piperacillin/tazobactam (Pip/Taz). Results: During the 14-month period, the ASP intervention acceptance rate increased significantly from a mean of 72.0% in the implementation phase (January–April 2025) to 81.2% in the stabilization phase (May 2025–February 2026) (p = 0.035). The DOT for Pip/Taz decreased significantly from 169.4 to 151.8 per 1000 patient–days (p = 0.002), with a significant negative correlation identified between the intervention acceptance rate and Pip/Taz consumption (r = −0.625, p = 0.017). Although overall meropenem DOT showed seasonal fluctuations without reaching statistical significance across phases, a year-over-year comparison revealed a 7.5% reduction in meropenem DOT (January–February 2025: 54.8 vs. January–February 2026: 50.7 per 1000 patient–days). The rejection rate for restricted antibiotics declined from 3.8% to 2.6%, suggesting that clinicians increasingly self-regulated inappropriate prescribing attempts. Conclusions: The ASP demonstrated extended follow-up sustainability with a significant reduction in the consumption of key broad-spectrum antibiotics. A progressive increase in clinician acceptance of ASP interventions from 72.0% to 81.2%, combined with a concurrent decline in the restricted antibiotic rejection rate, reflected a measurable shift in institutional prescribing culture and confirmed the successful transition to a stabilized program. These findings support the necessity of sustained multidisciplinary ASPs, even in resource-limited settings, to combat antimicrobial resistance effectively.
Full article
(This article belongs to the Special Issue Antibacterial Utilisation from Different Perspectives: An Effective Tool to Develop and Evaluate Antimicrobial Stewardship Interventions)
►▼
Show Figures

Figure 1
Open AccessCase Report
Severe Mycoplasma pneumoniae Pneumonia During the 2023–2024 European Re-Emergence: Why Severity Does Not Predict Macrolide Resistance
by
Enrico Perugini, Ludovica Ferrari, Marco Iannetta, Barbara Bartolini, Valentina Dimartino, Marco Favaro, Carla Fontana and Loredana Sarmati
Antibiotics 2026, 15(5), 524; https://doi.org/10.3390/antibiotics15050524 - 21 May 2026
Abstract
Background: Following a significant decline during the 2020–2021 SARS-CoV-2 pandemic, Mycoplasma pneumoniae (MP) experienced a resurgence across Europe in 2023–2024. Although macrolide-resistant MP has increased globally, severe disease can occur even in the absence of resistance, which highlights the importance of rapid molecular
[...] Read more.
Background: Following a significant decline during the 2020–2021 SARS-CoV-2 pandemic, Mycoplasma pneumoniae (MP) experienced a resurgence across Europe in 2023–2024. Although macrolide-resistant MP has increased globally, severe disease can occur even in the absence of resistance, which highlights the importance of rapid molecular characterization for clinical purposes. In this context, clinical severity is often improperly used as a surrogate marker of macrolide resistance, potentially driving unnecessary antibiotic escalation. Methods: We report a severe MP pneumonia occurring during the 2023–2024 resurgence and evaluate macrolide resistance through a rapid two-step workflow (Real Time-PCR screening for A2063G/A2064G followed by confirmatory 23S rRNA sequencing), to assess whether severity predicts resistance and to support antibiotic stewardship. Results: The patient developed acute hypoxic respiratory failure (PaO2 54.9 mmHg; P/F ratio 110), extensive centrilobular micronodules on chest CT imaging, significant systemic inflammation and elevated liver enzymes. Respiratory support was escalated from a Venturi mask to a high-flow nasal cannula and BiPAP. MP infection was confirmed by multiplex Real Time-PCR (RT-PCR) and supported by positive IgM/IgG serology. RT-PCR targeting A2063G/A2064G mutations revealed no resistance-associated variants, and Sanger sequencing of an 807 bp 23S rRNA fragment confirmed a wild-type genotype. Despite severe hypoxemic respiratory failure, no resistance-associated variants were detected, documenting a clear severity–genotype mismatch. Clinical and radiological improvement followed second-line antibiotic therapy. Conclusions: Severe MP pneumonia can occur despite the absence of macrolide resistance. During MP re-emergence, clinical severity should not be used to infer macrolide resistance. Integrating nucleic acid amplification test (NAAT) diagnosis with rapid genotyping/confirmatory 23S rRNA sequencing can prevent misclassification, reduce unwarranted broad-spectrum escalation, and strengthen antimicrobial stewardship decisions.
Full article
(This article belongs to the Special Issue Epidemiology, Clinical Microbiology and Antimicrobial Therapy: A Shared Effort against Infectious Diseases, 2nd Edition)
Open AccessArticle
The Antimicrobial Mechanism of Geraniol Against Penicillium polonicum and Its Application in Fresh-Cut Yam
by
Na Feng, Wei Yang, Xiaoyang Zhang, Yusha He, Min Zhang and Na Wang
Antibiotics 2026, 15(5), 523; https://doi.org/10.3390/antibiotics15050523 - 21 May 2026
Abstract
Background: Plant essential oils are extensively utilized for their antimicrobial properties; however, the specific antifungal mechanisms of certain compounds are not well characterized. Geraniol, a naturally occurring monoterpene alcohol approved for use in foods, demonstrates potential efficacy against spoilage fungi, yet detailed mechanistic
[...] Read more.
Background: Plant essential oils are extensively utilized for their antimicrobial properties; however, the specific antifungal mechanisms of certain compounds are not well characterized. Geraniol, a naturally occurring monoterpene alcohol approved for use in foods, demonstrates potential efficacy against spoilage fungi, yet detailed mechanistic insights are lacking. Methods: In this study, we determined the minimum inhibitory concentration (MIC) and minimum fungicidal concentration (MFC) of geraniol against P. polonicum. We assessed the underlying mechanisms by evaluating membrane integrity, intracellular leakage, reactive oxygen species (ROS), antioxidant enzymes (superoxide dismutase [SOD], peroxidase [POD], catalase [CAT]), malondialdehyde (MDA) levels, ATP content, and ATPase activity. Inoculated yam slices were exposed to geraniol vapor, and we monitored sensory, physicochemical, enzymatic, and microbial parameters. Results: Geraniol exhibited a minimum inhibitory concentration/minimum fungicidal concentration (MIC/MFC) of 0.3 mL/L. It disrupted cellular membranes, induced leakage, generated ROS, and caused lipid peroxidation, leading to elevated levels of malondialdehyde (MDA). Additionally, geraniol activated antioxidant enzymes and impaired energy metabolism. Fumigation with geraniol dose-dependently delayed the deterioration of yam, reduced weight loss, preserved texture and color, inhibited polyphenol oxidase (PPO) and POD activities, enhanced CAT and SOD activities, lowered MDA levels, and suppressed bacterial growth. Conclusions: Geraniol inhibits P. polonicum through multiple mechanisms, including membrane disruption, oxidative stress, and interference with energy metabolism, thereby effectively preserving the quality of fresh-cut yam and demonstrating potential as a natural preservative.
Full article
(This article belongs to the Special Issue Natural Compounds as Antimicrobial Agents, 3rd Edition)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Occurrence and Antimicrobial Susceptibility Pattern of Clinical Escherichia coli Isolates from Dogs in Grenada, West Indies
by
Erika Brewer, Kaitlin Law, Bhumika Sharma, Andy Alhassan, Erica Hazel-Ann Brathwaite, Wayne Sylvester and Kamashi Kumar
Antibiotics 2026, 15(5), 522; https://doi.org/10.3390/antibiotics15050522 - 21 May 2026
Abstract
►▼
Show Figures
Background/objectives: Infections caused by multidrug-resistant (MDR) bacteria are becoming increasingly difficult to treat with recommended antimicrobials. Considering the critical and growing challenge of antimicrobial resistance (AMR), this study aims to evaluate the antimicrobial susceptibility patterns of Escherichia coli clinical isolates from dogs
[...] Read more.
Background/objectives: Infections caused by multidrug-resistant (MDR) bacteria are becoming increasingly difficult to treat with recommended antimicrobials. Considering the critical and growing challenge of antimicrobial resistance (AMR), this study aims to evaluate the antimicrobial susceptibility patterns of Escherichia coli clinical isolates from dogs in Grenada. This research project consists of two distinct studies: a retrospective analysis of AMR in canine E. coli isolates collected between 2010 and 2020, and a cross-sectional study characterizing the genotypic AMR profiles of E. coli isolates obtained between April and June 2023. Methods: A retrospective analysis of antibacterial sensitivity test (ABST) reports from canine clinical samples submitted to the Small Animal Clinic at St. George’s University (SGU), St. George’s, Grenada, between 2010 and 2020 revealed a notable prevalence of AMR among canine E. coli isolates. To further investigate the underlying mechanisms of this resistance, the study analyzed canine E. coli isolates that exhibited phenotypic resistance in ABST assays. These isolates were subsequently screened for AMR-associated genes using polymerase chain reaction (PCR) and next-generation sequencing (NGS). Results: The retrospective study identified 153 canine clinical isolates positive for E. coli. The antimicrobial drugs, imipenem, cefotaxime and ciprofloxacin were found to be highly effective against these isolates. However, a gradual increase in AMR was observed for amoxicillin–clavulanic acid (34.88%), ampicillin–sulbactam (17.31%), cephalexin (43.08%), cefpodoxime (22.31%), cephalothin (68.42%), and doxycycline (37.04%). In the prospective study, PCR analysis of resistant canine E. coli isolates detected the tetA (577 bp) and blaTEM (686 bp) genes. These AMR determinants were further confirmed through analysis of NGS reads and assembled contigs. Additionally, NGS-based predictions identified genes associated with resistance to aminoglycosides and potentiated sulfonamides. Conclusions: This study demonstrates that E. coli from dogs in Grenada exhibits resistance to tetracycline and several β-lactam antimicrobials. These findings underscore the need for rational antimicrobial stewardship and continuous AMR surveillance in small animal practice within the region.
Full article

Figure 1
Open AccessArticle
Evaluating the Impact of Filmarray Pneumonia Plus Panel in Therapeutic Decision-Making in Critical Patients with Suspected Respiratory Infection
by
Rosa Latorre Ibars, Sulamita Carvalho-Brugger, Paula Rodríguez Ibáñez, Montserrat Vallverdú Vidal, Silvia Iglesias Moles, Mar Miralbés Torner, Alba Bellés Bellés, Andrea Castellano, David Campi, Jesús Caballero and José Javier Trujillano Cabello
Antibiotics 2026, 15(5), 521; https://doi.org/10.3390/antibiotics15050521 - 21 May 2026
Abstract
Background: Respiratory infections in critically ill patients remain a major challenge in intensive care units (ICUs), with high morbidity and mortality. Conventional microbiological methods often fail to identify the causative pathogen promptly, particularly in patients previously exposed to antibiotics. Multiplex molecular platforms, such
[...] Read more.
Background: Respiratory infections in critically ill patients remain a major challenge in intensive care units (ICUs), with high morbidity and mortality. Conventional microbiological methods often fail to identify the causative pathogen promptly, particularly in patients previously exposed to antibiotics. Multiplex molecular platforms, such as the BioFire FilmArray® Pneumonia Panel Plus (FAPP), allow rapid detection of multiple respiratory pathogens and resistance markers, potentially improving early therapeutic decision-making. The objective of this work is to evaluate the impact of implementing FAPP on antimicrobial therapeutic decisions in critically ill patients with suspected respiratory infection. Methods: We conducted a retrospective cohort study in two mixed ICUs between 2023 and 2024. All respiratory samples in which FAPP was requested were analyzed. The results were compared with conventional cultures, and changes in antimicrobial therapy following the FAPP results were assessed, classified as escalation/initiation or de-escalation/discontinuation. Concordance between FAPP and culture was evaluated, and clinical and demographic variables were analyzed. Differences between groups were assessed using p-values obtained from the chi-square test or the Mann–Whitney test. Results: A total of 363 respiratory samples were included, 88.4% from mechanically ventilated patients. FAPP was positive in 65.3% of samples, whereas cultures were positive in 23.1%. Overall concordance between FAPP and culture was 57.3%. In 42.4% of cases, pathogens were detected exclusively by FAPP. Antimicrobial therapy was modified in 29.8% of patients, predominantly through de-escalation or discontinuation (69.4% of changes). Therapeutic modifications were more frequent in nosocomial infections and in patients with a positive FAPP result. Conclusions: The use of FAPP in critically ill patients with suspected respiratory infection provides rapid microbiological information that significantly influences antimicrobial decision-making, particularly by facilitating antibiotic de-escalation. Although discrepancies with conventional cultures remain and require careful clinical interpretation, FAPP represents a valuable tool for antimicrobial stewardship in the ICU setting.
Full article
(This article belongs to the Special Issue From Epidemiology to Intervention: Addressing the Spread of Multidrug-Resistant Microorganisms)
Open AccessArticle
Impact of PA-100 AST System Rapid Antibiotic Susceptibility Test on Antibiotic Prescription for Community-Acquired Urinary Tract Infections in Spanish Primary Care Settings
by
Lourdes Martínez-Berganza Asensio, Gonzalo Largo-Rojo, Ana Isabel Menéndez-Fernández, Carmen Solano-Villarrubia, María Fuentes-Romero and José Medina-Polo
Antibiotics 2026, 15(5), 520; https://doi.org/10.3390/antibiotics15050520 - 21 May 2026
Abstract
Background/Objectives: This intervention study compared the impact of the PA-100 AST System (PA-100) with the standard of care on antibiotic-prescribing behaviour for community-acquired urinary tract infections in a Spanish primary care setting. Methods: Women seeking care for symptoms of uncomplicated urinary
[...] Read more.
Background/Objectives: This intervention study compared the impact of the PA-100 AST System (PA-100) with the standard of care on antibiotic-prescribing behaviour for community-acquired urinary tract infections in a Spanish primary care setting. Methods: Women seeking care for symptoms of uncomplicated urinary tract infections were recruited based on the last digit of their regional personal identification number in a control (no PA-100 result available) or intervention (PA-100 result available) arm. Differences in antibiotic-prescribing behaviour were analysed using Fisher’s exact test, with the sample size powered to detect a change in prescription in ≥6% of patients. Results: Availability of the PA-100 revealed resistance to fosfomycin in 21.5% of confirmed infections. This significantly shifted prescription away from fosfomycin towards nitrofurantoin and amoxycillin/clavulanic acid (p < 0.001). In accordance with local guidelines, fosfomycin was the most frequently prescribed antibiotic in the control arm (65.9%), whereas a significantly lower rate (37.7%) was observed in the intervention arm. Conclusions: The PA-100 shows potential to support antimicrobial stewardship by enabling targeted antibiotic treatment at the first visit and improving care in primary care settings.
Full article
(This article belongs to the Section Antibiotic Therapy in Infectious Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Expanding Diagnostic Options for Pediatric Meningitis: BCID2 Testing Results on Cerebrospinal Fluid After a Negative Meningitis/Encephalitis Panel
by
Venere Cortazzo, Lorenza Romani, Gianluca Vrenna, Maia De Luca, Marilena Agosta, Martina Rossitto, Valeria Fox, Barbara Lucignano, Manuela Onori, Stefania Mercadante, Vito Tommaso, Laura Lancella, Stefania Bernardi, Mara Pisani, Alessandra Salvatori, Alberto Villani, Massimiliano Raponi, Carlo Federico Perno and Paola Bernaschi
Antibiotics 2026, 15(5), 519; https://doi.org/10.3390/antibiotics15050519 - 21 May 2026
Abstract
Background: Rapid etiological diagnosis of bacterial meningitis is crucial in children, as delays can lead to neurological sequelae. The BioFire FilmArray Meningitis/Encephalitis (ME) panel is widely used on cerebrospinal fluid (CSF), but its target spectrum may miss healthcare-associated or multidrug-resistant pathogens. We evaluated
[...] Read more.
Background: Rapid etiological diagnosis of bacterial meningitis is crucial in children, as delays can lead to neurological sequelae. The BioFire FilmArray Meningitis/Encephalitis (ME) panel is widely used on cerebrospinal fluid (CSF), but its target spectrum may miss healthcare-associated or multidrug-resistant pathogens. We evaluated the diagnostic performance and stewardship-oriented clinical impact of off-label BioFire FilmArray Blood Culture Identification 2 (BCID2) testing on CSF from pediatric patients with suspected bacterial CNS infection and negative ME results. Methods: We retrospectively analyzed CSF samples collected between January 2023 and March 2025 at a tertiary pediatric hospital. In ME-negative cases with persistent suspicion and abnormal CSF parameters, BCID2 was performed off-label on residual CSF aliquots after routine testing, without additional sampling. We assessed pathogen detection, agreement with culture, resistance-gene identification, and documented stewardship actions. Results: Among 76 ME-negative CSF samples tested with BCID2, 23 (30.3%) were positive, all involving organisms not included in the ME panel. BCID2 was concordant with culture in 19/23 cases (82.6%); 4/23 (17.4%) were BCID2-positive/culture-negative, consistent with reduced culture sensitivity in frequently pretreated cases. Resistance genes (VIM, vanA/B, CTX-M) were detected in 30.4% of BCID2-positive samples. Overall agreement with culture was 94.7% (PPA 100%, NPA 93.0%). Escalation was documented in 13/23 episodes (56.5%), discontinuation in 2/23 (8.7%), and confirmation in 9/23 (39.1%), with no de-escalation events; clinical outcomes were not systematically available. Conclusions: In selected ME-negative pediatric cases with abnormal CSF profiles, BCID2 testing on residual CSF provided rapid, clinically meaningful microbiological information that may support antimicrobial optimization.
Full article
(This article belongs to the Special Issue Epidemiology, Clinical Microbiology and Antimicrobial Therapy: A Shared Effort against Infectious Diseases, 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
Effect of Fatty Acid Chain Modification on the Self-Assembly Behavior and Antimicrobial Activity of Antimicrobial Peptides
by
Hongyan Yang, Meiqian Luo, Yutao Min, Yehuan Zheng, Yanhua Xu, Bingchao Duan, Fei Pan and Kui Lu
Antibiotics 2026, 15(5), 518; https://doi.org/10.3390/antibiotics15050518 - 20 May 2026
Abstract
Background: The overuse of traditional antimicrobial agents has accelerated the global spread of drug-resistant bacteria, posing a severe threat to global public health. Methods: In this work, a series of lipopeptides with varying fatty acid chain lengths were designed using the
[...] Read more.
Background: The overuse of traditional antimicrobial agents has accelerated the global spread of drug-resistant bacteria, posing a severe threat to global public health. Methods: In this work, a series of lipopeptides with varying fatty acid chain lengths were designed using the targeting antimicrobial peptide CL5 as the parental peptide. A variety of technical methods, including spectroscopic techniques, electron microscopy and computer simulation, were adopted to explore the self-assembly properties of the lipopeptides and their antimicrobial properties against Gram-positive and Gram-negative bacteria. Results: The results showed that lipopeptide self-assembly could be triggered by fatty acid chain modification with a carbon chain length exceeding 8 atoms, and hydrophobic interactions between fatty acid chains were the primary driving force for this process. The geometric mean of the minimum inhibitory concentrations of the lipopeptides exhibited an approximate “U”-shaped correlation with the length of the fatty acid chains. Among these lipopeptides, C8CL5–C12CL5 exhibited broad-spectrum and highly potent antimicrobial activity, with geometric means of 6.20, 5.16, and 8.00 μM against all tested bacteria, and selectivity index values of 12.26, 8.14, and 7.48, respectively. Furthermore, the lipopeptides exhibited high selectivity, rapid time-killing kinetics, as well as excellent thermal, pH and salt stability. Mechanistic studies revealed that the lipopeptides exerted antimicrobial effects through multiple pathways: disrupted bacterial cell membranes and caused the leakage of cellular contents, bound to bacterial genomic DNA, and promoted the production of reactive oxygen species. Conclusions: Collectively, lipopeptides modified with appropriate fatty acid chains exhibit broad-spectrum and highly effective antimicrobial activity, making them promising alternatives to traditional antibiotics for the treatment of bacterial infections.
Full article
(This article belongs to the Section Antimicrobial Peptides)
Open AccessArticle
Intra-Individual Variability of Vancomycin Trough Concentrations Before and After Implementation of a Standardized Operating Procedure in Orthopedic Inpatients
by
Moritz Diers, Laura Isabell Werneburg, Alexander Zeh, Natalia Gutteck, Karl-Stefan Delank and Felix Werneburg
Antibiotics 2026, 15(5), 517; https://doi.org/10.3390/antibiotics15050517 - 20 May 2026
Abstract
Background: Standardized operating procedures (SOPs) for intravenous vancomycin therapy have been shown to improve population-level trough target attainment and to reduce nephrotoxicity in orthopedic inpatients. However, mean target attainment on a population level does not capture how stably an individual patient remains within
[...] Read more.
Background: Standardized operating procedures (SOPs) for intravenous vancomycin therapy have been shown to improve population-level trough target attainment and to reduce nephrotoxicity in orthopedic inpatients. However, mean target attainment on a population level does not capture how stably an individual patient remains within the therapeutic window. Intra-individual variability of vancomycin trough concentrations has remained underreported as a patient-level quality indicator in the orthopedic stewardship literature, despite its direct clinical relevance, as alternating sub- and supratherapeutic phases compromise both efficacy and safety independently of the mean exposure. Methods: We conducted a secondary analysis of the prospectively and retrospectively collected data of the Halle Vancomycin SOP cohort. Pre-SOP (n = 58) and post-SOP (n = 23) patient cohorts were compared with respect to patient-level variability metrics, including the coefficient of variation (CV%), swing index, mean absolute successive difference (MSSD), range of trough values, zone-transition frequencies, and the proportion of “stable” patients defined as CV% below 20%. First-order Markov transition matrices were computed to characterize the directionality of trough movements between subtherapeutic, target, and supratherapeutic zones. The primary analysis was restricted to patients with at least three documented trough measurements. Results: The median CV% decreased from 43.5% (IQR 33.5–51.5) pre-SOP to 32.5% (IQR 21.9–38.6) post-SOP (Mann–Whitney U, p = 0.011). The swing index decreased from 1.09 to 0.75 (p = 0.002), and the median range of individual trough concentrations shrank from 19.1 mg/L to 13.2 mg/L (p = 0.029). The absolute number of zone transitions per patient did not differ significantly between cohorts, but their directionality differed substantially: target-zone persistence increased from 37.8% to 57.6%. Across all 403 measurements, subtherapeutic values declined from 38.5% to 26.6%, while target-zone measurements rose from 28.5% to 44.7%. In the post-SOP cohort, longer therapy duration was associated with lower CV% (Spearman ρ = −0.52, p = 0.032). Conclusions: In addition to improvements in population-level target attainment, implementation of the SOP was associated with stabilization of the individual exposure profile of orthopedic inpatients receiving intravenous vancomycin. Intra-individual variability was lower in the post-SOP cohort, and transitions between zones were more often oriented toward the target range. These findings, derived from a single-centre secondary analysis with a small post-SOP cohort, support patient-level variability metrics as a complementary quality indicator in protocolized vancomycin management and warrant prospective multicentre validation.
Full article
(This article belongs to the Section Antibiotics Use and Antimicrobial Stewardship)
►▼
Show Figures
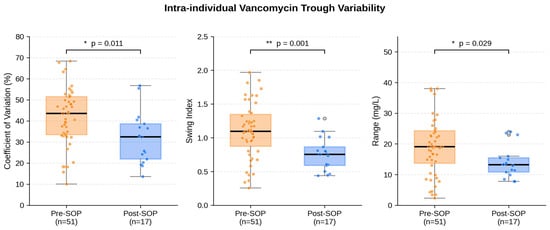
Figure 1
Open AccessArticle
Multicenter Genomic Analysis of Carbapenem-Resistant Pseudomonas aeruginosa in Austrian Community Hospitals Reveals Limited Carbapenemase Prevalence and Absence of Interhospital Clonal Spread
by
Magda Diab-Elschahawi, Tim Kirk, Susanne Häussler, Elisabeth Presterl and the PSPS Working Group
Antibiotics 2026, 15(5), 516; https://doi.org/10.3390/antibiotics15050516 - 20 May 2026
Abstract
Background/Objectives: In Europe, Pseudomonas aeruginosa is the second most common cause of ventilator-associated pneumonia in intensive care units. Intrinsic antibiotic resistance and acquired carbapenemases can lead to high mortality. To guide more targeted antimicrobial therapy and adequate infection control measures, we performed
[...] Read more.
Background/Objectives: In Europe, Pseudomonas aeruginosa is the second most common cause of ventilator-associated pneumonia in intensive care units. Intrinsic antibiotic resistance and acquired carbapenemases can lead to high mortality. To guide more targeted antimicrobial therapy and adequate infection control measures, we performed a multicenter study on the prevalence and genetic basis of carbapenem resistance among P. aeruginosa (CR-PA) across 17 community hospitals in Austria. Methods: During a 3-month period, we collected 621 P. aeruginosa isolates from 560 patients. Antibiotic susceptibility testing was performed according to EUCAST guidelines, and all CR-PA isolates were subjected to whole genome sequencing. Results: Antibiotic susceptibility testing revealed carbapenem resistance in 5.41% (36/621) of the investigated P. aeruginosa isolates. Only 3 produced a carbapenemase (2 Verona Integron-encoded Metallo- ß-lactamases and 1 Imipenemase Metallo-ß-lactamase) and carried a carbapenemase-encoding gene. Among the studied P. aeruginosa isolates there was a high genetic diversity, excluding a single driving epidemic lineage in the included Austrian hospitals. Conclusions: The absence of interhospital clonal dominance suggests that carbapenem resistance emerged independently in different centers, likely driven by local antibiotic selection pressures rather than regional clonal spread.
Full article
(This article belongs to the Special Issue Healthcare-Associated Infections (HAIs): Prevention, Control and Surveillance, 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessReview
Companion Dogs and Cats as Key Reservoirs of Antimicrobial Resistance: Evidence and One Health Implications
by
Balamuralikrishnan Balasubramanian, Sureshkumar Shanmugam and In Ho Kim
Antibiotics 2026, 15(5), 515; https://doi.org/10.3390/antibiotics15050515 - 19 May 2026
Abstract
Antimicrobial resistance (AMR) in companion animals is an escalating concern at the interface of veterinary medicine and public health. Dogs and cats, the most commonly treated companion species, are frequently prescribed antimicrobials for dermatological, otic, urinary, and respiratory infections—often involving drug classes that
[...] Read more.
Antimicrobial resistance (AMR) in companion animals is an escalating concern at the interface of veterinary medicine and public health. Dogs and cats, the most commonly treated companion species, are frequently prescribed antimicrobials for dermatological, otic, urinary, and respiratory infections—often involving drug classes that are critically important in human medicine. This overlap underscores the need for judicious use and integrated stewardship within a One Health framework. This narrative review synthesizes current evidence on AMR in companion animals and its implications for One Health. Studies were included if they reported AMR in dogs and cats and addressed zoonotic aspects. Staphylococcus pseudintermedius, S. aureus, Escherichia coli, Pseudomonas aeruginosa, and Enterococcus sp. are examples of clinically significant organisms that are becoming more resistant to several antibiotic classes, which can result in treatment failures and extended illness. Horizontal gene transfer facilitates the spread of resistance determinants across bacterial populations. Improved surveillance systems, prudent antibiotic use, regular culture and susceptibility testing, and enhanced antimicrobial stewardship in veterinary practice are just a few of the many strategies needed to address AMR in companion animals. The integration of companion animals into AMR surveillance, stewardship programs, and infection control strategies is essential. Coordinated One Health interventions are urgently required to mitigate the spread of AMR.
Full article
(This article belongs to the Special Issue Companion Animals: Antimicrobial Resistance in Bacterial and Fungal Pathogens Associated with Skin and Ear Infections)
►▼
Show Figures
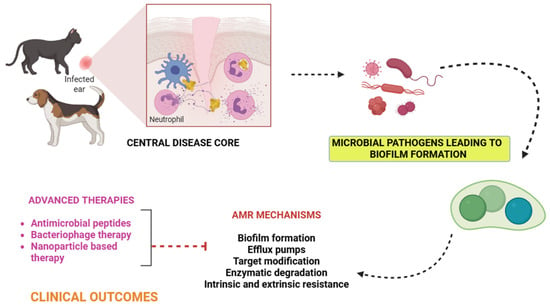
Figure 1

Journal Menu
► ▼ Journal Menu-
- Antibiotics Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Antibiotics, IJMS, Microbiology Research, Pharmaceuticals, Pharmaceutics, Nanomaterials, Microorganisms
Challenges and Future Prospects of Antibacterial Therapy, 2nd Edition
Topic Editors: Kwang-Sun Kim, Zehra EdisDeadline: 30 November 2026
Topic in
Agriculture, Animals, Antibiotics, Microorganisms, Poultry, Pathogens, Veterinary Sciences, Biologics
Advances in Infectious and Parasitic Diseases of Animals
Topic Editors: Felipe M. Salvarani, Sheyla Farhayldes Souza Domingues, Júlia Angélica Gonçalves Da SilveiraDeadline: 31 December 2026
Topic in
Analytica, Antibiotics, Bioengineering, Biosensors, Microorganisms
Spectroscopic Phenotypic Typing of Multidrug-Resistant Bacteria: Advances in FTIR and Raman Applications
Topic Editors: László Orosz, David Rodríguez TemporalDeadline: 31 January 2027
Topic in
Antioxidants, Biology, Biomedicines, Microorganisms, JoF, IJMS, Stresses, Antibiotics
Microbial Redox Biology: From Stress Response to Biotechnology
Topic Editors: Michal Letek, Volker BehrendsDeadline: 1 March 2027

Special Issues
Special Issue in
Antibiotics
Innovative and Translational Antibiosis Approaches to Tackling Protozoan Parasites: Synergy, Drug Repurposing, Natural Products, Molecular Targets, and Therapeutic Design
Guest Editor: Leonardo AcuñaDeadline: 30 May 2026
Special Issue in
Antibiotics
Antimicrobial Stewardship in Neonatal Intensive Care
Guest Editor: Chiara PoggiDeadline: 30 May 2026
Special Issue in
Antibiotics
Antibiotic Resistance in Bacterial Isolates of Animal Origin
Guest Editors: Tomasz Jarzembowski, Lidia PiechowiczDeadline: 31 May 2026
Special Issue in
Antibiotics
Antibiotic Residues, Antimicrobial Resistance and Intervention Strategies of Foodborne Pathogens, 2nd Edition
Guest Editors: Yongning Wu, Zhenling ZengDeadline: 31 May 2026
Topical Collections
Topical Collection in
Antibiotics
Antimicrobial Prescribing and Antimicrobial Use in Healthcare Settings
Collection Editors: Masayuki Maeda, Yuichi Muraki
Topical Collection in
Antibiotics
Antibiotics in Ophthalmology Practice
Collection Editor: Sanjay Marasini
Topical Collection in
Antibiotics
Staphylococcus— Molecular Pathogenesis, Virulence Regulation and Antibiotics Resistance
Collection Editor: Ewa Szczuka
Topical Collection in
Antibiotics
Synthetic and Natural Products-Based Antimicrobial and Antiparasitic Agents
Collection Editor: Antonio Eduardo Miller Crotti







