Journal Description
Microbiology Research
Microbiology Research
is an international, peer-reviewed, open access journal published monthly online by MDPI (from Volume 11, Issue 2 - 2020).
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), Embase, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 20.2 days after submission; acceptance to publication is undertaken in 5.8 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
2.2 (2024);
5-Year Impact Factor:
2.1 (2024)
Latest Articles
Optimization of IAA Production by Halotolerant Vreelandella titanicae J113 Through Fermentation Process Engineering with Response Surface Methodology
Microbiol. Res. 2026, 17(5), 95; https://doi.org/10.3390/microbiolres17050095 (registering DOI) - 12 May 2026
Abstract
►
Show Figures
Soil salinization is a significant environmental factor limiting agricultural production. Developing salt–alkali-tolerant microbial resources is important for the improvement of saline–alkali land. Plant growth-promoting rhizobacteria stimulate crop growth by producing the plant growth hormone indole-3-acetic acid (IAA), but their fermentation process under salt
[...] Read more.
Soil salinization is a significant environmental factor limiting agricultural production. Developing salt–alkali-tolerant microbial resources is important for the improvement of saline–alkali land. Plant growth-promoting rhizobacteria stimulate crop growth by producing the plant growth hormone indole-3-acetic acid (IAA), but their fermentation process under salt stress still needs optimization. Single-factor experiments and response surface methodology (RSM) were used to systematically optimize the fermentation conditions of the salt–alkali-tolerant Vreelandella titanicae J113. Key influencing factors were screened using the single-factor experiment design, and optimal process parameters were determined using the Box–Behnken design. IAA production and cell biomass were used as evaluation indicators to study the interactions of carbon sources, nitrogen sources, inorganic salts, temperature, cultivation time, and inoculum size. The optimal fermentation process was obtained: starch concentration 17.5 g/L, NaCl concentration 32.5 g/L, yeast extract 5 g/L, cultivation temperature 30 °C, inoculum size 3%, and cultivation time 144 h. After optimization, IAA production reached 23.02 μg/mL, an increase of 115% compared with before optimization. Salt stress experiments showed that the strain could still maintain high IAA production under 3% NaCl, demonstrating good salt tolerance. Maize seed germination experiments demonstrated that the optimized fermentation broth significantly promoted seed germination and seedling growth under salt stress conditions, with root length, fibrous root number, and fresh weight increasing by 61–86%, 137–200%, and 25–57%, respectively, compared to the control group. This study established an efficient IAA fermentation process for the salt–alkali-tolerant Vreelandella titanicae J113, providing technical support for developing microbial plant growth regulators suitable for saline–alkali land. The optimized strain exhibits excellent growth-promoting potential under salt stress conditions, offering favorable application prospects.
Full article
Open AccessReview
Engineering Escherichia coli for Aromatic Compound Biosynthesis: Integrating Metabolic Engineering and Synthetic Biology
by
Silvana M. Tapia-Cabrera, Adelfo Escalante and Francisco Bolívar
Microbiol. Res. 2026, 17(5), 94; https://doi.org/10.3390/microbiolres17050094 (registering DOI) - 9 May 2026
Abstract
►▼
Show Figures
Aromatic compounds derived from the shikimate (SHK) pathway constitute a diverse class of high-value molecules with applications in the pharmaceutical, food, cosmetic, and chemical industries. In microbial systems, particularly Escherichia coli, this pathway links central carbon metabolism (CCM) to the biosynthesis of
[...] Read more.
Aromatic compounds derived from the shikimate (SHK) pathway constitute a diverse class of high-value molecules with applications in the pharmaceutical, food, cosmetic, and chemical industries. In microbial systems, particularly Escherichia coli, this pathway links central carbon metabolism (CCM) to the biosynthesis of L-tyrosine (L-Tyr), L-phenylalanine (L-Phe), and L-tryptophan (L-Trp), which serve as key precursors for structurally diverse metabolites. Over the past decades, metabolic engineering strategies have focused on increasing precursor availability, relieving feedback inhibition, and eliminating competing pathways. More recently, advances in synthetic biology have enabled dynamic control of metabolic flux through pathway modularization, genome-scale interventions, and regulatory circuit design. In this review, we provide a comprehensive overview of the engineering of E. coli for aromatic compound biosynthesis, highlighting key developments in the optimization of the SHK pathway and its major metabolic nodes chorismate, L-Tyr, L-Phe, and L-Trp. We examine emerging approaches, including CRISPR-based regulation, biosensor-driven dynamic control, membrane engineering, and synthetic microbial consortia. Despite significant progress, challenges related to pathway regulation, cofactor balance, metabolic burden, and product toxicity remain critical bottlenecks. Integrating metabolic engineering with synthetic biology is driving the development of programmable, scalable microbial platforms for the efficient bioproduction of aromatic compounds.
Full article

Figure 1
Open AccessArticle
Molecular Epidemiology of Drug-Resistant Mycobacterium tuberculosis: Mutation Profiles and Resistance Associations
by
Mandlenkosi Manika, Lindiwe Modest Faye, Ntandazo Dlatu and Mojisola Clara Hosu
Microbiol. Res. 2026, 17(5), 93; https://doi.org/10.3390/microbiolres17050093 - 8 May 2026
Abstract
Background: The global burden of drug-resistant Mycobacterium tuberculosis continues to threaten tuberculosis control efforts, largely due to the emergence and transmission of resistance-associated genetic mutations. Molecular epidemiology provides critical insights into mutation profiles and resistance associations, yet the interplay among key mutations and
[...] Read more.
Background: The global burden of drug-resistant Mycobacterium tuberculosis continues to threaten tuberculosis control efforts, largely due to the emergence and transmission of resistance-associated genetic mutations. Molecular epidemiology provides critical insights into mutation profiles and resistance associations, yet the interplay among key mutations and their contributions to complex resistance patterns remains poorly understood, particularly in high-burden settings. Methods: A retrospective, cross-sectional, laboratory-based design was used to analyze 111 phenotypically confirmed drug-resistant isolates. Molecular drug susceptibility testing (DST) for first- and second-line anti-tuberculosis drugs was performed at the National Health Laboratory Service (NHLS) TB reference laboratory. Drug-resistance profiles were classified according to World Health Organization (WHO) definitions. Descriptive and inferential statistical analyses were conducted to determine mutation frequencies, co-occurrence patterns, and associations with resistance profiles. Results: rpoB (D435V 38.7%; S450L 36.0%) and katG (S315T 80.2%) mutations predominated, forming the core molecular basis of MDR-TB, while 15% harbored inhA promoter mutations associated with low-level isoniazid resistance. The most frequent combinations included rpoB S450L with katG S315T and rpoB D435V with katG S315T, consistent with multidrug-resistant tuberculosis (MDR-TB) profiles. Nearly 48% showed dual resistance to fluoroquinolones and second-line injectables. Conclusion: This study highlights the predominance of resistance-associated mutations and their co-occurrence patterns in shaping MDR-TB profiles in the study setting. The observed burden of second-line drug resistance underscores the importance of comprehensive resistance testing. These findings support the use of mutation profiling for rapid diagnosis and informed treatment decisions, while emphasizing the need for ongoing local surveillance to guide TB control efforts.
Full article
(This article belongs to the Topic Multidrug Resistance Across Pathogens: Fungi, Bacteria, Parasites, and Viruses)
Open AccessSystematic Review
Molecular and Proteomic Determinants of Trypanosoma cruzi Adaptation Within Triatomine Vectors: Insights from Current Experimental Models
by
Jessy T. Santana, Berenice González-Rete, Elia Torres-Gutiérrez, Juliana Cordeiro Cardoso, Cláudia Moura Melo and Paz M. S. Salazar-Schettino
Microbiol. Res. 2026, 17(5), 92; https://doi.org/10.3390/microbiolres17050092 - 8 May 2026
Abstract
Trypanosoma cruzi exhibits complex genetic diversity, organized into seven distinct typing units. To complete its life cycle, the parasite must adapt to the digestive tract of various species of triatomine bugs. This systematic review aimed to understand the molecular adaptation mechanisms of T.
[...] Read more.
Trypanosoma cruzi exhibits complex genetic diversity, organized into seven distinct typing units. To complete its life cycle, the parasite must adapt to the digestive tract of various species of triatomine bugs. This systematic review aimed to understand the molecular adaptation mechanisms of T. cruzi in relation to different vector species, systematizing knowledge on vector competence. Following PRISMA guidelines, 18 experimental studies (published between 1995 and 2025) were selected from the ScienceDirect, PubMed, Scopus, and Web of Science databases, focusing on the parasite–vector interface and proteomic analyses. There was a predominance of studies conducted in Brazil (66.67%), using the Rhodnius prolixus model (72.22%) and the TcI strain (clone Dm28c). The evolution of methodological approaches reflects a transition from classical techniques, such as SDS-PAGE, to high-throughput omics strategies, including LC-MS/MS and gene editing tools such as CRISPR. The findings were organized into key biological processes, including parasite adhesion mediated by perimicrovillar membrane components, glycoinositolphospholipids (GIPLs), and mucins; the influence of the metabolic and nutritional microenvironment, particularly hemoglobin-derived peptides and glucose availability; and the role of intestinal redox conditions in triggering metacyclogenesis. Overall, the available evidence suggests that T. cruzi adaptation within triatomine vectors is a multifactorial process driven by proteomic reprogramming and post-transcriptional regulation in response to environmental signals within the vector gut. However, this understanding is largely derived from studies based on Rhodnius prolixus and TcI strains, which limits the generalization of these mechanisms across other triatomine species and parasite lineages.
Full article
(This article belongs to the Special Issue Host–Microbe Interactions in Health and Disease)
►▼
Show Figures

Figure 1
Open AccessArticle
Impact of Pre-Existing Uterine Microbiome on Pregnancy Success After Embryo Transfer in Cattle
by
Nilton Luis Murga Valderrama, Gleni T. Segura, Jakson Ch Del Solar, Hugo Frias, Ana C. Romani, Deiner J. Gongora-Bardales, Ulises S. Quispe-Gutierrez, Carla Maria Ordinola-Ramirez, Richard C. Polveiro, Dielson da S. Vieira, Jorge Luis Maicelo Quintana and Rainer M. Lopez Lapa
Microbiol. Res. 2026, 17(5), 91; https://doi.org/10.3390/microbiolres17050091 - 8 May 2026
Abstract
►▼
Show Figures
The uterine microbiome plays a critical role in maintaining pH balance, modulating the immune system, and influencing fertility, especially in artificial breeding contexts. This study examined the impact of uterine microbiota on pregnancy success in cows following embryo transfer (ET), using Illumina 16S
[...] Read more.
The uterine microbiome plays a critical role in maintaining pH balance, modulating the immune system, and influencing fertility, especially in artificial breeding contexts. This study examined the impact of uterine microbiota on pregnancy success in cows following embryo transfer (ET), using Illumina 16S rRNA gene sequencing of the V4 hypervariable region of samples collected from the uterine horn (UH) and the uterine body (UB) of cows during the estrous cycle preceding synchronization for ET in the Amazon region. Microbiomes from the uterine horn (UH) and the uterine body (UB) were analyzed before embryo transfer. Cows that became pregnant (UH-P and UB-P) and those that did not (UH-NP and UB-NP) were compared. Fifteen cows were grouped as follows: UB-P (three), UB-NP (five), UH-P (three), and UH-NP (four). Linear discriminant analysis effect size and heat tree analyses identified Sphingobacterium and Stenotrophomonas spp. as significantly enriched in the UB-P and UH-NP groups, respectively. Additionally, non-pregnant cows exhibited more distinctive genera than pregnant ones. These findings suggest that cows achieving pregnancy have lower microbial diversity and fewer potentially pathogenic genera. This study contributes to the emerging field of pre-pregnancy uterine microbiome research in cattle, offering evidence that microbial composition may influence reproductive success, and highlights specific taxa as potential biomarkers for pregnancy outcomes following embryo transfer.
Full article

Figure 1
Open AccessArticle
Detection and Host Range Investigation of Phytopythium helicoides and Phytopythium palingenes
by
Petya Koeva Christova
Microbiol. Res. 2026, 17(5), 90; https://doi.org/10.3390/microbiolres17050090 (registering DOI) - 30 Apr 2026
Abstract
Phytopythium species are water molds that have been divided as a separate group of oomycetes about 10 years ago. They are associated with diverse environments worldwide, but the ecological function of most of the species is still under investigation. In the present study,
[...] Read more.
Phytopythium species are water molds that have been divided as a separate group of oomycetes about 10 years ago. They are associated with diverse environments worldwide, but the ecological function of most of the species is still under investigation. In the present study, isolation and characterization of Pp. helicoides and Pp. palingenes are described. Both Phytopythium isolates originate from an aquatic environment and were derived from infected leaves. A host range of Pp. helicoides and Pp. palingenes using pathogenicity tests with leaves and cuttings from trees, bushes, perennial and herbaceous plants that belong to 15 different families was studied. Out of 21 tested plants, 18 were susceptible to infection with Pp. helicoides and nine were negatively affected by Pp. palingenes. Pp. helicoides is distinguished by higher aggressiveness and a wider range of potential host species. These results indicate that the impact of pathogenic species from the genus Phytopythium on forests and other natural ecosystems could be much more significant than is currently known.
Full article
(This article belongs to the Special Issue Advances in Plant–Pathogen Interactions)
►▼
Show Figures

Figure 1
Open AccessArticle
Enterococcus durans Secretome Modulates Interleukins Gene Expressions in Intestinal Epithelial Cells Challenged by Staphylococcus aureus Secretome: In Vitro Study on the HT-29 Cell Line
by
Egidia Costanzi, Giovanna Traina, Marco Misuraca, Donia Msakni, Giada Sgaravizzi, Musafiri Karama, Ebtesam Al-Olayan, Saeed El-Ashram, Marcelo Martinez-Barbitta, Massimo Zerani and Beniamino T. Cenci-Goga
Microbiol. Res. 2026, 17(5), 89; https://doi.org/10.3390/microbiolres17050089 (registering DOI) - 30 Apr 2026
Abstract
The present study examined the effect of Enterococcus durans cell-free supernatant (CFS) on interleukin (IL)-8, -10 and -1β gene expressions in the intestinal cell line HT-29 treated with Staphylococcus aureus CFS. HT-29 cells were incubated with E. durans CFS or S. aureus
[...] Read more.
The present study examined the effect of Enterococcus durans cell-free supernatant (CFS) on interleukin (IL)-8, -10 and -1β gene expressions in the intestinal cell line HT-29 treated with Staphylococcus aureus CFS. HT-29 cells were incubated with E. durans CFS or S. aureus CFS, or S. aureus CFS plus E. durans CFS. All concentrations of E. durans CFS did not show cytotoxicity, while the highest treatment (44.9 μg/mL) with S. aureus CFS induced significant cell death. S. aureus CFS did not modify IL-1β gene expression, while E. durans CFS alone or in combination with S. aureus CFS reduced it. Treatment with S. aureus CFS induced greater expression of the IL-8 gene compared to S. aureus CFS plus E. durans CFS. S. aureus CFS alone or in combination with E. durans CFS increased the expression of the IL-10 gene, while E. durans CFS alone did not modify it. These results suggest a potential protective role of the E. durans secretome in mitigating the inflammatory environment in intestinal cells. This treatment could be useful to protect against possible contact with dangerous soluble microbial products present in food.
Full article
(This article belongs to the Special Issue Cell Culture Advances in Food Microbiology: Innovative In Vitro Models for Safety, Quality, and Functionality)
►▼
Show Figures

Figure 1
Open AccessCommunication
Serological Investigation of Infectious Bovine Rhinotracheitis in Dromedary Camels and Dairy Herds in Tunisia: Preliminary Results
by
Stefano Petrini, Mohamed Methnani, Cecilia Righi, Khaled El Hicheri, Cristina Casciari, Aida Tatli, Ben Smida Boubaker, Elena Tinelli, Sana Kacem, Claudia Pellegrini, Roberto Sabato, Francesco Feliziani and Giovanni Pezzotti
Microbiol. Res. 2026, 17(5), 88; https://doi.org/10.3390/microbiolres17050088 - 29 Apr 2026
Abstract
Livestock farming represents a key economic activity in the Tataouine Governorate of southern Tunisia, where cattle and dromedary camels coexist. Varicellovirus bovinealpha1 (BoAHV-1), the etiological agent of infectious bovine rhinotracheitis (IBR), primarily affects cattle, while its circulation in camelids remains poorly understood. Following
[...] Read more.
Livestock farming represents a key economic activity in the Tataouine Governorate of southern Tunisia, where cattle and dromedary camels coexist. Varicellovirus bovinealpha1 (BoAHV-1), the etiological agent of infectious bovine rhinotracheitis (IBR), primarily affects cattle, while its circulation in camelids remains poorly understood. Following recent European Union regulations requiring BoAHV-1 surveillance in multiple animal species, this short communication reports serological findings from dairy cattle and dromedary herds in southern Tunisia. In March 2024, serum samples were collected from four non-vaccinated farms, including two intensive Friesian dairy cattle herds and two extensive dromedary herds (50 animals each). Serum samples from all animals were tested for BoAHV-1 antibodies using competitive commercial gB- and gE-based enzyme-linked immunosorbent assays (c-ELISA) and confirmed by virus neutralization test (VNT). Antibodies against BoAHV-1 were detected in cattle from both dairy farms, with low seroprevalence and neutralizing antibody titers, indicating past or ongoing exposure. In contrast, all dromedary samples tested seronegative by both c-ELISA and VNT. These findings confirm BoAHV-1 circulation in cattle in the Tataouine region and its absence in dromedaries at sampling. Further studies involving larger sample sizes and molecular investigations are required to clarify the potential role of camelids in BoAHV-1 epidemiology in southern Tunisia.
Full article
(This article belongs to the Special Issue Emerging and Re-Emerging Viruses in Animal Health: Pathogenesis, Diagnosis, and One Health Strategies)
Open AccessReview
Rapid Antimicrobial Susceptibility Testing (AST): Overview of New Commercially Available Automated Phenotypic Tools for Minimum Inhibitory Concentration (MIC) Determination
by
Giorgia Piccinini, Antonio Curtoni, Alessandro Bondi, Mattia Genco, Fabio Longo, Carlotta Polizzi, Paolo Valesella, Silvia Corcione, Francesco Giuseppe De Rosa and Cristina Costa
Microbiol. Res. 2026, 17(5), 87; https://doi.org/10.3390/microbiolres17050087 - 29 Apr 2026
Abstract
Antimicrobial resistance (AMR) represents one of the most urgent global health threats, significantly impacting patient outcomes, healthcare systems, and economic sustainability. Rapid and accurate antimicrobial susceptibility testing (AST) are essential to guide targeted therapy, reduce inappropriate antimicrobial use, and support antimicrobial stewardship programs.
[...] Read more.
Antimicrobial resistance (AMR) represents one of the most urgent global health threats, significantly impacting patient outcomes, healthcare systems, and economic sustainability. Rapid and accurate antimicrobial susceptibility testing (AST) are essential to guide targeted therapy, reduce inappropriate antimicrobial use, and support antimicrobial stewardship programs. However, conventional phenotypic AST methods, including broth microdilution, disk diffusion, agar dilution, and gradient strip tests, remain labor-intensive and require prolonged turnaround times, often delaying optimal therapeutic decisions. Although automated commercial platforms such as VITEK 2, BD Phoenix, MicroScan WalkAway, and Sensititre ARIS have improved laboratory workflow and standardization, they still rely on culture-based approaches and typically require 16–36 h to generate minimum inhibitory concentration (MIC) results. In recent years, several innovative rapid phenotypic AST technologies have emerged, aiming to significantly shorten the time to susceptibility results while maintaining high accuracy. This review provides an overview of currently available rapid automated phenotypic platforms for MIC determination, including VITEK® Reveal™, ASTar, FASTinov®AST, QuickMIC®, and the Accelerate Pheno® system. These systems employ advanced technologies such as volatile organic compound detection, flow cytometry, microfluidics, real-time imaging, and morphokinetic cellular analysis to deliver susceptibility results within a few hours directly from positive blood cultures. We summarize their technical principles, antibiotics and pathogens included, performances, and current limitations. Overall, the implementation of rapid phenotypic AST tools has the potential to substantially improve clinical decisions, optimize antimicrobial therapy, and contribute to fight AMR.
Full article
(This article belongs to the Special Issue Antimicrobial Resistance: New Diagnostic Strategies)
►▼
Show Figures

Figure 1
Open AccessArticle
Molecular Epidemiology, Hematobiochemical Alterations, and Oxidative Stress-Induced Genotoxicity of Equine Trypanosomiasis in Pakistan
by
Waqas Ahmad, Naeem Rasool, Qurat ul Ain, Usama Bin Naeem, Muhammad Azeem, Umbreen Anwar, Tehreem Fayyaz, Zeba Amjad, Muhammad Yasin Tipu and Mehmood Ahmad
Microbiol. Res. 2026, 17(5), 86; https://doi.org/10.3390/microbiolres17050086 (registering DOI) - 27 Apr 2026
Abstract
►▼
Show Figures
Trypanosoma evansi (T. evansi) infection poses a significant health threat to equines. This study was aimed to assess the prevalence, risk factors, hematobiochemical alterations, and oxidative stress-mediated genotoxicity associated with equine trypanosomiasis in the Rahim Yar Khan District. This cross-sectional study
[...] Read more.
Trypanosoma evansi (T. evansi) infection poses a significant health threat to equines. This study was aimed to assess the prevalence, risk factors, hematobiochemical alterations, and oxidative stress-mediated genotoxicity associated with equine trypanosomiasis in the Rahim Yar Khan District. This cross-sectional study was conducted on 384 equines from October 2024 to September 2025. Blood samples were collected for thin blood film microscopy and PCR assay using RoTat 1.2 primers. Hematological indices were analyzed with an automated hematology analyzer; serum biochemical parameters were quantified via standard assays. Oxidative stress markers, including malondialdehyde (MDA), catalase (CAT), superoxide dismutase (SOD), and reduced glutathione (GSH), were also measured. Genotoxicity was evaluated using the alkaline comet assay. Statistical analyses included the chi-square test, logistic regression, and independent t-tests. T. evansi was detected in 5.99% of samples by microscopy and 10.16% by PCR, with no significant association with species, age, or sex. Infected equines exhibited significant reductions in hemoglobin (5.4 ± 0.6 vs. 10.8 ± 0.5 g/dL; p < 0.0001), total serum protein (2.1 ± 0.3 vs. 5.8 ± 0.2 g/dL; p < 0.0001), albumin, and globulin, alongside elevated hepatic enzymes, blood urea nitrogen, and creatinine (all p < 0.01). Oxidative stress was confirmed by increased MDA (p < 0.0001) and decreased CAT activity (p < 0.001). Genotoxicity was significantly higher in infected animals (genetic damage index; 1.12 ± 0.08 vs. 0.40 ± 0.01; p < 0.01). This study provides the first integrated assessment of molecular epidemiology and oxidative stress-mediated genotoxicity in equines in this region, suggesting the pathogenic impact of the infection and targeted diagnostics for disease management strategies.
Full article

Figure 1
Open AccessArticle
Antimicrobial Effect of Slightly Acidic Hypochlorous Acid Water Against Biofilm Formed by Candida parapsilosis
by
Jun Iwahashi, Akiko Shimizu, Akinobu Togo, Hiroshi Fuketa, Kenji Gotoh, Keisuke Ohta, Norihiro Shinkai, Naohisa Kawamura and Hiroshi Watanabe
Microbiol. Res. 2026, 17(5), 85; https://doi.org/10.3390/microbiolres17050085 - 24 Apr 2026
Abstract
►▼
Show Figures
Background: Many of the pathogenic bacteria and fungi found in hospital environments form biofilms, which allow them to persist in the environment for long periods, posing a risk of hospital-acquired infections. Although the pathogens within biofilms often have reduced levels of drug susceptibility,
[...] Read more.
Background: Many of the pathogenic bacteria and fungi found in hospital environments form biofilms, which allow them to persist in the environment for long periods, posing a risk of hospital-acquired infections. Although the pathogens within biofilms often have reduced levels of drug susceptibility, the efficacy of disinfectants routinely applied against planktonic pathogens must be evaluated against biofilms as well. Our objective in this study was to determine the efficacy of treatment using slightly acidic hypochlorous acid water and to compare the results with sodium hypochlorite when both were used to disinfect Candida parapsilosis biofilms. Methods: C. parapsilosis in the planktonic or biofilm state was treated with each disinfectant. The number of viable cells that remained was determined, and scanning electron microscopy (SEM) of the disinfectant-treated biofilms was performed. Results: Compared with sodium hypochlorite, in a shorter period of time, hypochlorous acid water completely killed not only planktonic C. parapsilosis but also C. parapsilosis in a biofilm that had been formed for 72 h. SEM showed that both disinfectants were effective in removing the C. parapsilosis biofilm to some extent. Conclusions: Slightly acidic hypochlorous acid water appears to be an effective disinfectant against C. parapsilosis both in suspension and in biofilms.
Full article

Figure 1
Open AccessArticle
Differential Effects of Overexpressing WHI3, CLPP, and PMP20 on the Secretion of Human Serum Albumin and Lactoferrin in Komagataella phaffii
by
Linglin Tao, Alessandro Ruan and Shu Quan
Microbiol. Res. 2026, 17(4), 84; https://doi.org/10.3390/microbiolres17040084 - 20 Apr 2026
Abstract
►▼
Show Figures
Komagataella phaffii (formerly Pichia pastoris) is a prominent platform for recombinant protein production, yet secretion efficiency often remains a critical bottleneck. In this study, we validated three candidate genes—WHI3, CLPP, and PMP20—previously identified through genome-wide CRISPR activation screening,
[...] Read more.
Komagataella phaffii (formerly Pichia pastoris) is a prominent platform for recombinant protein production, yet secretion efficiency often remains a critical bottleneck. In this study, we validated three candidate genes—WHI3, CLPP, and PMP20—previously identified through genome-wide CRISPR activation screening, for their potential to enhance heterologous protein secretion. Overexpression of these factors under the control of the methanol-inducible AOX1 promoter increased the secretion of human serum albumin (HSA), with WHI3 and CLPP yielding improvements of 18.3% and 17.9%, respectively. Furthermore, applying this strategy to human lactoferrin (hLF) revealed that WHI3 overexpression robustly enhanced hLF secretion by approximately 70%. Comparative analysis of different promoters (AOX1, GAP, and CAT) indicated that the AOX1 promoter remains the most effective driver for these enhancers, suggesting a threshold-dependent regulatory mechanism. These results demonstrate the protein-dependent nature of secretion optimization and identify WHI3, CLPP, and PMP20 as novel, effective co-expression factors for improving recombinant protein yields in K. phaffii.
Full article

Figure 1
Open AccessArticle
Faecal Microbiota Transplantation in IL-10 Knockout Mice Reverses Increased Susceptibility to Pseudomonas aeruginosa Lung Infection
by
Natália Cristina de Melo Santos, Evandro Neves Silva, Leonardo Pereira de Araújo, Carlos Roberto Prudêncio, Rômulo Dias Novaes, Patrícia Paiva Corsetti and Leonardo Augusto de Almeida
Microbiol. Res. 2026, 17(4), 83; https://doi.org/10.3390/microbiolres17040083 - 20 Apr 2026
Abstract
Differences in the gut microbiota are directly reflected in lung–gut axis crosstalk, which may increase susceptibility to pulmonary infections, such as those caused by the bacterium Pseudomonas aeruginosa. Deficiency of the cytokine IL-10 leads to gut inflammation, and this pro-inflammatory environment is
[...] Read more.
Differences in the gut microbiota are directly reflected in lung–gut axis crosstalk, which may increase susceptibility to pulmonary infections, such as those caused by the bacterium Pseudomonas aeruginosa. Deficiency of the cytokine IL-10 leads to gut inflammation, and this pro-inflammatory environment is partly due to changes in the gut microbiota. To better understand the effects of IL-10 deficiency on the gut microbiota, the intestinal microbial composition of IL-10 KO mice was assessed, and an increase in the phyla Bacteroidetes and Proteobacteria and a decrease in the phylum Firmicutes were observed in the faeces compared with the wild-type group (WT). Additionally, IL-10 KO mice had a higher pro-inflammatory immunostimulatory caecal content. Furthermore, it was found that heterologous faecal microbiota transplantation (FMT) between groups reversed this gut imbalance. IL-10 KO mice showed greater susceptibility to acute pulmonary infection by P. aeruginosa, with a higher recovery of viable bacteria in the lung and spleen, greater tissue damage and increased expression of genes encoding pro-inflammatory cytokines in the lungs. This greater susceptibility was reversed after FMT. Taken together, these results demonstrate the role of endogenous IL-10 in the gut microbiota constitution and its importance in the pulmonary immune response against P. aeruginosa infection.
Full article
(This article belongs to the Special Issue Host–Microbe Interactions in Health and Disease)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Probiotic Potential of Saccharomyces cerevisiae var. boulardii, Weizmannia coagulans and Lacticaseibacillus rhamnosus as Commercial Supplements: In Vitro Gastrointestinal Kinetics, Pharmaceutical Stability and Antioxidant Support in Chamomile Tea
by
Eleni Alaverntian and Eugenia Papadaki
Microbiol. Res. 2026, 17(4), 82; https://doi.org/10.3390/microbiolres17040082 - 16 Apr 2026
Abstract
►▼
Show Figures
The gut microbiome plays a central role in human health, and probiotics are widely used to support microbial balance, though their efficacy depends on multiple factors. This study assessed the potential of commercial probiotics Saccharomyces cerevisiae var. boulardii, Weizmannia coagulans and Lacticaseibacillus
[...] Read more.
The gut microbiome plays a central role in human health, and probiotics are widely used to support microbial balance, though their efficacy depends on multiple factors. This study assessed the potential of commercial probiotics Saccharomyces cerevisiae var. boulardii, Weizmannia coagulans and Lacticaseibacillus rhamnosus by evaluating in vitro gastrointestinal kinetics, pharmaceutical stability, and antioxidant effects in chamomile tea. Growth across a broad pH range was modeled kinetically, while survival and inactivation were quantified in simulated gastric and intestinal fluids. Antibiotic and antifungal susceptibility was determined using disk diffusion, and antioxidant activity of fortified chamomile tea was assessed via DPPH radical scavenging. Results revealed distinct strain-dependent responses. S. cerevisiae var. boulardii and W. coagulans showed the highest gastrointestinal tolerance. The increase in fluid volume reduced survival during the gastric phase but improved survival in the intestinal phase, reflecting different stress responses. Antimicrobial susceptibility also varied, with S. cerevisiae var. boulardii exhibiting the highest resistance. Probiotic fortification enhanced chamomile tea’s antioxidant capacity, particularly for S. cerevisiae var. boulardii and L. rhamnosus. These findings provide quantitative insight into strain- and volume-dependent gastrointestinal performance, guiding the optimization of capsule formulations and the development of clean-label products combining probiotic and antioxidant benefits.
Full article

Figure 1
Open AccessArticle
Bioactive Secondary Metabolites and Anti-Infective Properties of Two Sordariomycetes Taxa Characterized by HR-ESI-MS Technique
by
Fatma A. Abo Nouh, Ahmed M. Abdel-Azeem, Tamer S. Abdelmoneim, Nivien A. Nafady, Saeed Mohammadi, Najeeb Ur Rehman, Hassan Moghtaderi, Moosa Al Hamadani, Saif Al-Housni, Usama Qayum and Abdullah M. S. Al-Hatmi
Microbiol. Res. 2026, 17(4), 81; https://doi.org/10.3390/microbiolres17040081 - 15 Apr 2026
Abstract
►▼
Show Figures
The emergence of antimicrobial resistance and the increasing incidence of cancer have highlighted the urgent need to develop new drugs; therefore, the discovery of new bioactive molecules is an important goal for future research. In this study, freshwater fungi isolated from submerged Phragmites
[...] Read more.
The emergence of antimicrobial resistance and the increasing incidence of cancer have highlighted the urgent need to develop new drugs; therefore, the discovery of new bioactive molecules is an important goal for future research. In this study, freshwater fungi isolated from submerged Phragmites australis from Egypt were screened for antimicrobial and cytotoxic activities. Using ITS1 and ITS4 primers, eight frequently occurring Sordariomycetes taxa were identified and were then selected for further evaluation of bioactivity. Ethyl acetate crude extracts (A–H) were evaluated for antimicrobial activity using the agar disk-diffusion method. Extracts A and E, derived from Chaetomium globosum SCUF0000404 (PX596738) and Chaetomium madrasense SCUF0000401 (PX596735), respectively, showed broad-spectrum activity at 100 mg/mL against bacterial pathogens, including Staphylococcus aureus ATCC 29213 (15.33 and 18.00 mm), Streptococcus pyogenes ATCC 19615 (11.00 mm), Escherichia coli ATCC 35218 (10.33 and 10.67 mm), Klebsiella pneumoniae ATCC 700603 (14.00 and 16.67 mm), and Pseudomonas aeruginosa ATCC 27853 (13.33 and 16.33 mm), and show antifungal activity against Candida albicans ATCC 14053 (20.33 mm), Candida krusei ATCC 6258 (15.67 and 15.33 mm), Trichosporon asahii AMS 187 (17.00 and 17.67 mm), Exserohilum rostratum AMS 1077 (34.00 and 33.67 mm), and Trichophyton indotineae AMS 180 (38.33 and 34.00 mm). Selective cytotoxic effects on the breast cancer cell line MDA-MB-231 were observed by extracts A and E at IC50 = 309 and 277 μg/mL, while non-selective cytotoxic effects on the normal HUVEC cell line were found with IC50 = 919 and 796 μg/mL, respectively. Characterization of the most effective extracts A and E by high-resolution electrospray ionization mass spectrometry (HR-ESI-MS) shows that they have a wide range of secondary metabolites, including cytochalasans, azaphilone alkaloids, steroids, terpenoids, flavonoids, and phenols. These findings underscore the chemical diversity and therapeutic potential of freshwater fungi from Egypt.
Full article

Figure 1
Open AccessArticle
Emerging Resistance in Oral Candida Isolates from Patients with Periodontal Disease
by
Claudia Berenice Tinoco-Cabral, Luis Alfonso Muñoz-Miranda, Manuel R. Kirchmayr, Vianeth Martínez-Rodríguez, Miguel Padilla-Rosas, Maricarmen Iñiguez-Moreno, Suchiquil Rangel-Velázquez, Fabiola Berenice Hernández-Reyes, Claudia Lisette Charles-Niño and Cesar Arturo Nava-Valdivia
Microbiol. Res. 2026, 17(4), 80; https://doi.org/10.3390/microbiolres17040080 - 10 Apr 2026
Abstract
Candida species can shift from commensal organisms to opportunistic pathogens. Both Candida albicans and non-albicans Candida (NAC) species colonize oral biofilms and periodontal pockets, where they may contribute to inflammation and the progression of periodontal disease. This study aimed to determine the
[...] Read more.
Candida species can shift from commensal organisms to opportunistic pathogens. Both Candida albicans and non-albicans Candida (NAC) species colonize oral biofilms and periodontal pockets, where they may contribute to inflammation and the progression of periodontal disease. This study aimed to determine the prevalence and antifungal susceptibility profiles of Candida species in individuals with different stages of periodontal disease. A cross-sectional study was conducted in 100 participants whose periodontal status was clinically evaluated. Saliva samples were cultured on chromogenic agar for yeast isolation, species identification was confirmed by MALDI-TOF MS, and antifungal susceptibility to fluconazole, clotrimazole, nystatin, and amphotericin B was assessed. Candida spp. was detected in 35% of participants, where C. albicans was the most prevalent species, followed by Nakaseomyces glabratus (formerly Candida glabrata), Candida parapsilosis, Candida dubliniensis, and Candida tropicalis. Species distribution varied according to periodontal status, with N. glabratus predominating in early periodontitis and C. albicans appeared more frequently in higher severe stages of periodontitis. Susceptibility testing showed resistance of C. albicans to clotrimazole (63.6%) and nystatin (22.7%), whereas amphotericin B and fluconazole remained effective. NAC species, particularly N. glabratus, exhibited resistance to nystatin and variable resistance to clotrimazole but remained susceptible to amphotericin B. These findings underscore the importance of early detection and personalized antifungal strategies for managing periodontal disease complicated by Candida colonization.
Full article
(This article belongs to the Special Issue Host–Microbe Interactions in Health and Disease)
►▼
Show Figures

Figure 1
Open AccessArticle
Mimotope Peptides of Salmonella Typhi AgVi Are Recognized by Anti-Vi Antigen Sera, Anti-Mimotope Peptides, and Human Sera
by
Armando Navarro-Ocaña, Armando Navarro-Cid del Prado, Ricardo Ernesto Ahumada-Cota and Ulises Hernández-Chiñas
Microbiol. Res. 2026, 17(4), 79; https://doi.org/10.3390/microbiolres17040079 - 10 Apr 2026
Abstract
►▼
Show Figures
Intestinal infections caused by Salmonella enterica serovar Typhi (S. Typhi) remain a global health concern, making preventive strategies and diagnostic tools essential. This study aimed to identify mimotope peptides of the Vi antigen using phage display and assess their recognition by
[...] Read more.
Intestinal infections caused by Salmonella enterica serovar Typhi (S. Typhi) remain a global health concern, making preventive strategies and diagnostic tools essential. This study aimed to identify mimotope peptides of the Vi antigen using phage display and assess their recognition by rabbit and 46 human sera, as well as their potential for diagnosis and immunogen design. Rabbits were immunized with the Vi antigen (AgVi) from S. Typhi ATCC 6539, and sera-derived IgG was used for phage biopanning. DNA sequences from selected phagotopes were synthesized as Salmonella mimotope peptides (SMPs), either linear or KLH-conjugated. Their reactivity was tested with ELISAs against AgVi and SMPs, using both rabbit sera and 46 human serum samples. Ten phagotopes were identified, with a consensus motif (D/G–A/V–x–P–x–x–G–x–x–x–x–x), suggesting α-helix structures. Immunization with KLH-conjugated peptides generated specific antibodies, particularly SMPVi/5 and SMPVi/10, which recognized AgVi and their respective peptides. Competitive inhibition assays confirmed that SMPVi/5 reduced the anti-AgVi binding in a dose-dependent manner. In human sera, AgVi recognition occurred in 52% of samples, while SMPVi/5 and SMPVi/10 were recognized in 45%. Overall, SMPVi/5 demonstrated immunogenicity and functional mimicry, supporting its use as a synthetic reagent for serological assays and as a candidate for immunogen design.
Full article

Figure 1
Open AccessArticle
Multidrug-Resistant and Hypervirulent Klebsiella pneumoniae from Invasive Clinical Samples: Evidence from a Tertiary-Care Hospital in India
by
Shubhangi Kansal, Kavita Gupta, Shubhneet Kaur Mamik, Neelam Taneja and Archana Angrup
Microbiol. Res. 2026, 17(4), 78; https://doi.org/10.3390/microbiolres17040078 - 8 Apr 2026
Abstract
The rise in multidrug resistance in Klebsiella pneumoniae is an alarming issue, especially in invasive infections among patients with co-morbidities. With the gain of hypervirulence traits, multidrug-resistant K. pneumoniae has led to a significant increase in chronic infections and associated mortality. This study
[...] Read more.
The rise in multidrug resistance in Klebsiella pneumoniae is an alarming issue, especially in invasive infections among patients with co-morbidities. With the gain of hypervirulence traits, multidrug-resistant K. pneumoniae has led to a significant increase in chronic infections and associated mortality. This study aims to explore the distribution of multidrug-resistant and hypervirulent (hv) K. pneumoniae in invasive infections in a tertiary care hospital. A total of 231 K. pneumoniae isolates were collected over a period of six months from invasive infections. These isolates were tested phenotypically and genotypically for the presence of antimicrobial resistance, along with molecular detection of hypervirulence determinants (iucA, rmpA, rmpA2, peg344, iroB). High levels of resistance to β-lactams, fluoroquinolones, and aminoglycosides were observed. Carbapenemase-encoding genes were widely distributed, and 22% showed the presence of at least one hypervirulence gene, most commonly iucA and rmpA. Co-carriage of resistance and hypervirulence determinants in K. pneumoniae was observed in nearly 20% of the isolates, indicating the emergence of MDR-hvKP phenotypes in the hospital setting. Mortality was significantly higher among patients infected with MDR isolates, whereas hypervirulence markers were not independently associated with mortality. The presence of MDR–hypervirulent strains remains clinically concerning and underscores the need for continued genomic surveillance.
Full article
(This article belongs to the Special Issue Klebsiella pneumoniae: Resistance, Virulence, and Epidemiology in Healthcare and Community Settings)
►▼
Show Figures

Figure 1
Open AccessArticle
Preliminary Experimental Study on the Removal of Staphylococcus epidermidis and Pseudomonas aeruginosa from Surgical Instrument Surfaces Under Controlled Conditions
by
Edmar Gonçalves Pereira Filho, Stéfanne Rodrigues Rezende Ferreira, Amanda Veiga Paiva Simões, Eli Júnior Pereira Rodrigues, Iorrana Morais de Oliveira, Marillia Lima Costa, Adeliane Castro da Costa, Berendina Elsina Bouwman and Hanstter Hallison Alves Rezende
Microbiol. Res. 2026, 17(4), 77; https://doi.org/10.3390/microbiolres17040077 - 8 Apr 2026
Abstract
►▼
Show Figures
The objective of this study is to evaluate the efficiency of surgical instruments’ manual cleaning versus automated cleaning in an ultrasonic cleaner for the removal of biofilms on surgical forceps contaminated with Staphylococcus epidermidis and Pseudomonas aeruginosa. Subsequently, the residual microbial load
[...] Read more.
The objective of this study is to evaluate the efficiency of surgical instruments’ manual cleaning versus automated cleaning in an ultrasonic cleaner for the removal of biofilms on surgical forceps contaminated with Staphylococcus epidermidis and Pseudomonas aeruginosa. Subsequently, the residual microbial load was quantified through microbiological culture, aiming to evaluate the effectiveness of biofilm removal under different reprocessing conditions. Cleaning is an essential step in the processing of surgical instruments to ensure the effective removal of dirt and microorganisms. Through adhesion, microorganisms can attach to surfaces and form biofilms, organized structures surrounded by an extracellular matrix consisting of various components, which favor metabolic exchanges, adaptation, resistance, and bacterial dispersion. These biofilms increase the pathogenic potential of microorganisms, contributing to the occurrence of Healthcare-Associated Infections, and to avoid these, it is essential that preventive measures aimed at microbial reduction are adopted. Automated cleaning proved more effective than manual cleaning, and the combined approach achieved the greatest microbial reduction, though persistent contamination was still observed. The ability of adhesion and biofilm formation on the surfaces of surgical instruments is regarded as a challenge for complete microbial removal. These findings enhance the need for more rigorous reprocessing protocols and complementary strategies to ensure greater safety in the use of reusable instruments in clinical practice.
Full article
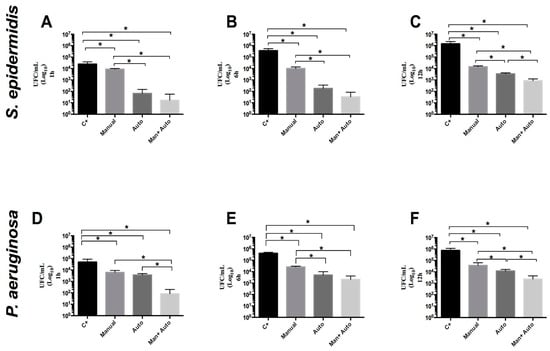
Figure 1
Open AccessArticle
Production of Surface-Active Metabolites by Bacillus sp. from Vegetable Oil-Impacted Soil: Ecological Implications and Screening Limitations
by
Eugenia Guadalupe Ortiz-Lechuga, Verónica Almaguer-Cantú, Hiram Herrera-Barquín, Karla Katiushka Solís-Arévalo, Ramón Alberto Batista-García and Katiushka Arévalo-Niño
Microbiol. Res. 2026, 17(4), 76; https://doi.org/10.3390/microbiolres17040076 - 8 Apr 2026
Abstract
►▼
Show Figures
Biosurfactant-producing microorganisms play an important ecological role in soils impacted by hydrophobic contaminants by enhancing substrate bioavailability and influencing microbial interactions. In this study, we critically evaluated the reliability of commonly used screening methods for biosurfactant detection. A total of 71 microbial isolates
[...] Read more.
Biosurfactant-producing microorganisms play an important ecological role in soils impacted by hydrophobic contaminants by enhancing substrate bioavailability and influencing microbial interactions. In this study, we critically evaluated the reliability of commonly used screening methods for biosurfactant detection. A total of 71 microbial isolates (16 bacteria and 55 fungi) were obtained from vegetable oil-contaminated soil and screened using a multi-step approach combining enzymatic assays (lipolytic and hemolytic activity) and physicochemical methods, including drop-collapse, oil spreading, emulsification index (E24), and surface tension reduction. Although 21 isolates exhibited lipolytic activity and 9 showed hemolysis, inconsistent responses among assays revealed significant limitations of individual screening methods. Only two bacterial isolates consistently tested positive across all criteria. When cultivated in mineral salt medium supplemented with hydrophobic substrates, both isolates produced stable emulsions and significantly reduced surface tension (from 54.26 mN/m to 31.46 mN/m). Substrate-dependent variation was observed for isolate C3, which showed reduced surface tension (39.63 mN/m) when grown with biodiesel. These findings highlight the risk of relying on single assays and emphasize the need for integrated screening strategies to ensure reliable detection of biosurfactant-producing microorganisms.
Full article

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Microbiology, Fermentation, Foods, Microbiology Research, Microorganisms, Nutrients
News and Updates on Probiotics
Topic Editors: Alessandra Pino, Mutamed AyyashDeadline: 30 September 2026
Topic in
Biomass, Fermentation, Microbiology Research, Microorganisms
The Utilization of Non-Grain Biomass Resources
Topic Editors: Shilei Wang, Yafan CaiDeadline: 31 October 2026
Topic in
JoF, Microbiology Research, Microorganisms, Pathogens
Pathophysiology and Clinical Management of Fungal Infections
Topic Editors: Allan J. Guimarães, Marcos de Abreu AlmeidaDeadline: 30 November 2026
Topic in
Antibiotics, Bacteria, Microbiology Research, Microorganisms, Pathogens, Parasitologia
Multidrug Resistance Across Pathogens: Fungi, Bacteria, Parasites, and Viruses
Topic Editors: Célia F. Rodrigues, Andreia S. AzevedoDeadline: 31 December 2026

Conferences
Special Issues
Special Issue in
Microbiology Research
Host–Microbe Interactions in Health and Disease
Guest Editors: Josué Juárez, Pablo Mendez-Pfeiffer, Manuel Ballesteros-MonrrealDeadline: 31 May 2026
Special Issue in
Microbiology Research
Antimicrobial Resistance: New Diagnostic Strategies
Guest Editors: Antonio Curtoni, Alessandro BondiDeadline: 30 June 2026
Special Issue in
Microbiology Research
Zoonotic Bacteria: Infection, Pathogenesis and Drugs—Second Edition
Guest Editors: Yang Wang, Shaohui WangDeadline: 30 June 2026
Special Issue in
Microbiology Research
Diagnostics and Molecular Epidemiology of Infectious Diseases in Veterinary Medicine
Guest Editor: Seyed Ali GhorashiDeadline: 30 June 2026
Topical Collections
Topical Collection in
Microbiology Research
Microbiology and Technology of Fermented Foods
Collection Editor: Salam A. Ibrahim
Topical Collection in
Microbiology Research
Public Health and Quality Aspects Related to Animal Productions
Collection Editors: Beniamino T. Cenci-Goga, Massimo Zerani
Topical Collection in
Microbiology Research
Microorganisms and Their Incredible Potential to Face Societal Challenges
Collection Editor: Mireille Fouillaud









