-
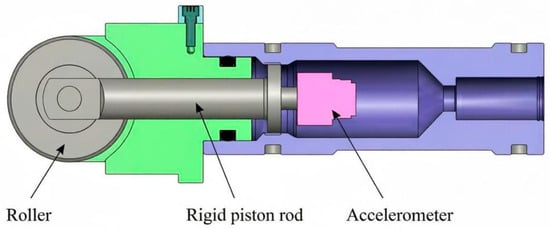
 Enhanced Reaction Time Measurement System Based on 3D Accelerometer in Athletics
Enhanced Reaction Time Measurement System Based on 3D Accelerometer in Athletics -
 Modular IoT Sensing and Data Pipeline for Digital Twin–Enabled Real-Time Aircraft Structural Health Monitoring
Modular IoT Sensing and Data Pipeline for Digital Twin–Enabled Real-Time Aircraft Structural Health Monitoring -
 Flexible Sensor Foil Based on Polymer Optical Waveguide for Haptic Assessment
Flexible Sensor Foil Based on Polymer Optical Waveguide for Haptic Assessment -
 A Codesign Framework for the Development of Next Generation Wearable Computing Systems
A Codesign Framework for the Development of Next Generation Wearable Computing Systems -
 Comparative Analysis of the Accuracy and Robustness of the Leap Motion Controller 2
Comparative Analysis of the Accuracy and Robustness of the Leap Motion Controller 2
Journal Description
Sensors
- Open Access — free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Ei Compendex, Inspec, Astrophysics Data System, and other databases.
- Journal Rank: JCR - Q2 (Instruments and Instrumentation) / CiteScore - Q1 (Instrumentation)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.8 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about Sensors.
- Companion journals for Sensors include: Chips, Targets, AI Sensors and IJMD.
- Journal Cluster of Instruments and Instrumentation: Actuators, AI Sensors, Instruments, Metrology, Micromachines and Sensors.
Latest Articles
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Deadline: 30 April 2026
Deadline: 31 May 2026
Deadline: 30 June 2026
Deadline: 31 July 2026
Conferences
Special Issues
Deadline: 10 April 2026
Deadline: 10 April 2026
Deadline: 10 April 2026
Deadline: 10 April 2026