Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS), Brazilian Society for Virology (BSV) and Global Virus Network (GVN) are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Virology/Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.2 days after submission; acceptance to publication is undertaken in 2.7 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
3.5 (2024);
5-Year Impact Factor:
3.7 (2024)
Latest Articles
Genomic Evidence for Novel Introduction and Intra-Host Diversity of DENV-2 in Dar es Salaam, Tanzania
Viruses 2026, 18(5), 585; https://doi.org/10.3390/v18050585 - 21 May 2026
Abstract
Dengue virus (DENV) poses a growing risk in Tanzania, yet its genetic diversity in mosquito populations remains poorly understood. Using Nanopore sequencing, we recovered full coding sequences from six DENV-2 positive mosquito pools collected in Dar es Salaam outside recognized outbreak periods. Phylogenetic
[...] Read more.
Dengue virus (DENV) poses a growing risk in Tanzania, yet its genetic diversity in mosquito populations remains poorly understood. Using Nanopore sequencing, we recovered full coding sequences from six DENV-2 positive mosquito pools collected in Dar es Salaam outside recognized outbreak periods. Phylogenetic analysis placed these sequences in a distinct monophyletic clade within genotype II, separate from strains linked to Tanzania’s 2014 outbreak. Instead, they clustered with Asian lineages and showed the closest relatedness to DENV-2 strains from Kenya (2013) and India (2014), with divergence estimated to have occurred around 2010. Variant profiling identified 212 low-frequency intra-pool variants, predominantly non-synonymous changes in the NS3, NS4B, and NS5 coding regions. These results suggest a previously unrecognized introduction of genotype II that is now circulating silently within local mosquito populations. Our findings highlight the value of genomic surveillance in mosquito vectors for early detection of arboviral threats, even in the absence of reported human cases.
Full article
(This article belongs to the Special Issue Current Trends in Arbovirus Outbreaks and Research)
►
Show Figures
Open AccessArticle
Infectome Landscape of Rodents and Shrews in Guangdong Province Reveals Diverse Pathogens with Zoonotic Potential in Wildlife
by
Yukun Lin, Fenxiang Li, Peiyu Liang, Yangzi Zhou, Lihua Zhang, Wudi Zhou, Yufeng Liang, Ruolan Yu, Wei Yang, Zhijian Zhou, Zeliang Wei, Jian He, Jingzhe Jiang and Huacheng Yan
Viruses 2026, 18(5), 584; https://doi.org/10.3390/v18050584 - 21 May 2026
Abstract
Rodents and shrews are important reservoir hosts due to their close association with human activities and their role in carrying various zoonotic pathogens. Recently, meta-transcriptomic sequencing has become a powerful tool for surveilling and screening novel pathogens from wild animals. However, many of
[...] Read more.
Rodents and shrews are important reservoir hosts due to their close association with human activities and their role in carrying various zoonotic pathogens. Recently, meta-transcriptomic sequencing has become a powerful tool for surveilling and screening novel pathogens from wild animals. However, many of these studies focused only on the diversity and genetic evolution of viruses from wildlife, while ignoring non-viral pathogens such as bacterial and eukaryotic microorganisms. Here, we performed a comprehensive infectome analysis of 227 tissue samples collected from 42 rodents and 16 shrews across six cities of Guangdong Province, China. We identified 34 viral families, including 23 mammalian viruses. Phylogenetic analysis revealed a henipavirus from the kidneys of shrews closely related to the Langya virus with potential infection risks to humans. Additionally, two potential pathogenic bacteria and 12 eukaryotic pathogens from six genera were found, showing clearer organ tropism than viruses. Interestingly, a moderate positive abundance correlation between Usmuvirus newyorkense and Trichinella suggested a potential virus–parasite association. We used machine learning models to evaluate the zoonotic potential of the obtained viruses, which indicated that 15 of 23 viral species were high risk for human infection. These findings provide important insight into the substantial zoonotic threat posed by pathogens circulating in wild small mammals in southern China and highlight the necessity for persistent wildlife pathogen surveillance.
Full article
(This article belongs to the Section Animal Viruses)
Open AccessArticle
Influence of Sampling Strategies and Disease Prevalence on SARS-CoV-2 Detection Dynamics in Wastewater Surveillance
by
Siti Aishah Rashid, Mohd Ishtiaq Anasir, Fadly Syah Arsad, Nurul Farehah Shahrir, Khayri Azizi Kamel, Sakshaleni Rajendiran, Nurul Amalina Khairul Hasni, Mohamad Iqbal Mazeli, Yuvaneswary Veloo, Syahidiah Syed Abu Thahir, Wan Rozita Wan Mahiyuddin, Khor Bee Chin, Alijah Mohd Aris, Redzuan Zainudin, Rafiza Shaharudin and Raheel Nazakat
Viruses 2026, 18(5), 583; https://doi.org/10.3390/v18050583 - 21 May 2026
Abstract
Background: Wastewater-based surveillance (WBS) has emerged as a valuable tool for population-level monitoring of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) transmission, yet the interplay between sampling strategies and disease prevalence in shaping detection performance remains ambiguous. We investigated how grab and composite
[...] Read more.
Background: Wastewater-based surveillance (WBS) has emerged as a valuable tool for population-level monitoring of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) transmission, yet the interplay between sampling strategies and disease prevalence in shaping detection performance remains ambiguous. We investigated how grab and composite sampling influence SARS-CoV-2 ribonucleic acid (RNA) detection dynamics and predictive lag times across high- and low-prevalence communities in Selangor, Malaysia. Methods: A 28-week longitudinal study was conducted in Selangor, Malaysia, comparing grab and composite wastewater sampling in communities with high and low Coronavirus disease 2019 (COVID-19) prevalence. SARS-CoV-2 RNA in 348 samples was quantified using digital Reverse Transcription Polymerase Chain Reaction (RT-dPCR), and viral lineages were characterized by Nanopore sequencing. Detection sensitivity and lead times relative to reported cases were evaluated. Results: In low-prevalence settings, grab sampling showed higher detection sensitivity than composite sampling (92.0% vs. 70.0%), whereas both methods achieved similarly high detection in high-prevalence areas (>97.0%). Lag-time analysis indicated that grab sampling in high-prevalence settings was significantly associated with case trends at potential two-week lead (p = 0.024), while composite sampling in low-prevalence settings showed the strongest association at a potential one-week lead (p = 0.0022). Overall, lag structures varied by both sampling strategy and prevalence context. Both sampling approaches captured the replacement of Omicron sublineages (XBB.1.5, XBB.1.9.1, XBB.1.16) and identified additional circulating variants, including EG.5, that were not captured in the available clinical sequencing dataset during the same period. Conclusions: These findings reveal that local transmission intensity is associated with the utility of different sampling designs. Context-specific optimization of WBS sampling strategies enhances sensitivity, reduces detection lag, and strengthens early warning and genomic-tracking capacity in public health surveillance frameworks.
Full article
(This article belongs to the Special Issue Wastewater-Based Epidemiology and Viral Surveillance)
►▼
Show Figures
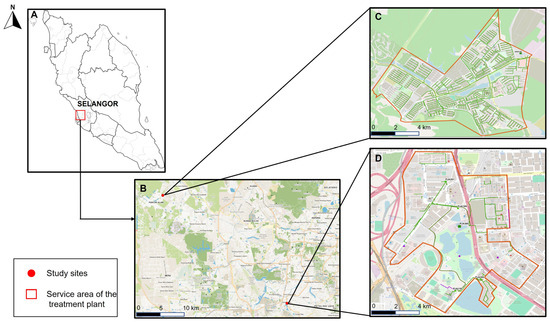
Figure 1
Open AccessArticle
Study on the Protective Efficacy of the Japanese Encephalitis Live Attenuated Vaccine SA14-14-2 Against Newly Isolated Genotype I Japanese Encephalitis Viruses
by
Shuai Shang, Qikai Yin, Tingyi Che, Xinzhu Wang, Qi Su, Shihong Fu, Hongshan Xu, Yongxin Yu, Qunying Mao, Huanyu Wang and Xinyu Liu
Viruses 2026, 18(5), 582; https://doi.org/10.3390/v18050582 - 21 May 2026
Abstract
Japanese encephalitis virus (JEV) comprises a single serotype but can be classified into five genotypes (genotypes I–V, GI–GV) based on nucleic acid sequences. Historically, genotype III (GIII) was the predominant strain. However, since the 21st century, genotype I (GI) rapidly replaced GIII as
[...] Read more.
Japanese encephalitis virus (JEV) comprises a single serotype but can be classified into five genotypes (genotypes I–V, GI–GV) based on nucleic acid sequences. Historically, genotype III (GIII) was the predominant strain. However, since the 21st century, genotype I (GI) rapidly replaced GIII as the major genotype in China, Southeast Asia, and other regions. The live attenuated vaccine (LAV) SA14-14-2, licensed in China in 1988, was successfully exported to 13 countries, with cumulative vaccinations exceeding one billion doses. The vaccine seed virus SA14-14-2 belonged to genotype III. Whether this GIII-based vaccine provided sufficient protection against the currently circulating GI strains warranted systematic investigation. In this study, recent JEV isolates collected from China were subjected to genotypic analysis, followed by comprehensive evaluations including protective efficacy against challenge and serum neutralizing antibody levels. The results indicated that, despite antigenic differences between GIII and GI strains, no significant differences in protective efficacy post-challenge were observed. The SA14-14-2 LAV remained effective in preventing GI strain infection.
Full article
(This article belongs to the Section Viral Immunology, Vaccines, and Antivirals)
►▼
Show Figures
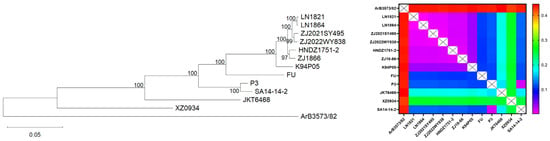
Figure 1
Open AccessArticle
Comparative Evaluation of Polymeric Nanocarriers for DNA Vaccine Delivery Against Avian Orthoavulavirus 1 in Chickens
by
Ahmed H. Khattab, Mahmoud Bayoumi, Zienab E. Eldin, Basem M. Ahmed and Haitham M. Amer
Viruses 2026, 18(5), 581; https://doi.org/10.3390/v18050581 - 21 May 2026
Abstract
Vaccination represents the cornerstone of Newcastle disease control. Nanotechnology offers a promising approach to improve the effectiveness of DNA vaccines, supporting their use as an alternative to conventional platforms. Herein, the Avian Orthoavulavirus 1 (AOAV-1) fusion (F) gene was cloned into
[...] Read more.
Vaccination represents the cornerstone of Newcastle disease control. Nanotechnology offers a promising approach to improve the effectiveness of DNA vaccines, supporting their use as an alternative to conventional platforms. Herein, the Avian Orthoavulavirus 1 (AOAV-1) fusion (F) gene was cloned into a DNA expression plasmid (pDNA). After validating the constructed pDNA-F and confirming robust intracellular protein expression in vitro, three polymeric nanoparticles (NPs)-based formulations were generated using Chitosan (Cs), poly(lactic-co-glycolic) (PLGA), and poly(amidoamine) (PAMAM)-Dendrimers. Physicochemical characterization, stability assessment, and in vitro release analysis confirmed nanoparticle formation and effective DNA incorporation. In vivo experiments were conducted to comparatively evaluate the immunogenicity, particularly the immune priming capacity, and protective efficacy of nanoparticle-based formulations and naked pDNA-F, all tested in parallel at standardized pDNA doses via intranasal (IN) and intramuscular routes. PAMAM-Dendrimers-pDNA-F IM group demonstrated superior efficacy, with 100% survival, the highest post-challenge anamnestic antibody titers, and a pronounced reduction in viral RNA shedding. PLGA-NPs-pDNA-F IN group demonstrated enhanced efficacy, with 90% survival. Naked pDNA-F surpassed the Cs-NPs-pDNA-F in both immune priming and clinical protection, with Cs-NPs-pDNA-F exhibiting the lowest overall performance. These findings highlight that DNA vaccine performance depends on both carrier type and administration route, with PAMAM dendrimers and PLGA enhancing efficacy, whereas chitosan demonstrated reduced efficacy under the tested conditions.
Full article
(This article belongs to the Section Animal Viruses)
Open AccessArticle
Whole Blood Volume-Based Absolute Quantification of HTLV-1 Proviral Load: A Comparative Method Evaluation Study
by
Gabriel O. Franco, Andreas Stocker, Eduardo M. Netto, Heliene Pereira and Carlos Brites
Viruses 2026, 18(5), 580; https://doi.org/10.3390/v18050580 - 21 May 2026
Abstract
The proviral load of human T-cell lymphotropic virus type 1 (HTLV-1) is an important biomarker associated with the monitoring and risk stratification of adult T-cell leukemia/lymphoma (ATL) and HTLV-1-associated myelopathy/tropical spastic paraparesis (HAM/TSP). However, the lack of standardized quantification methods limits its broader
[...] Read more.
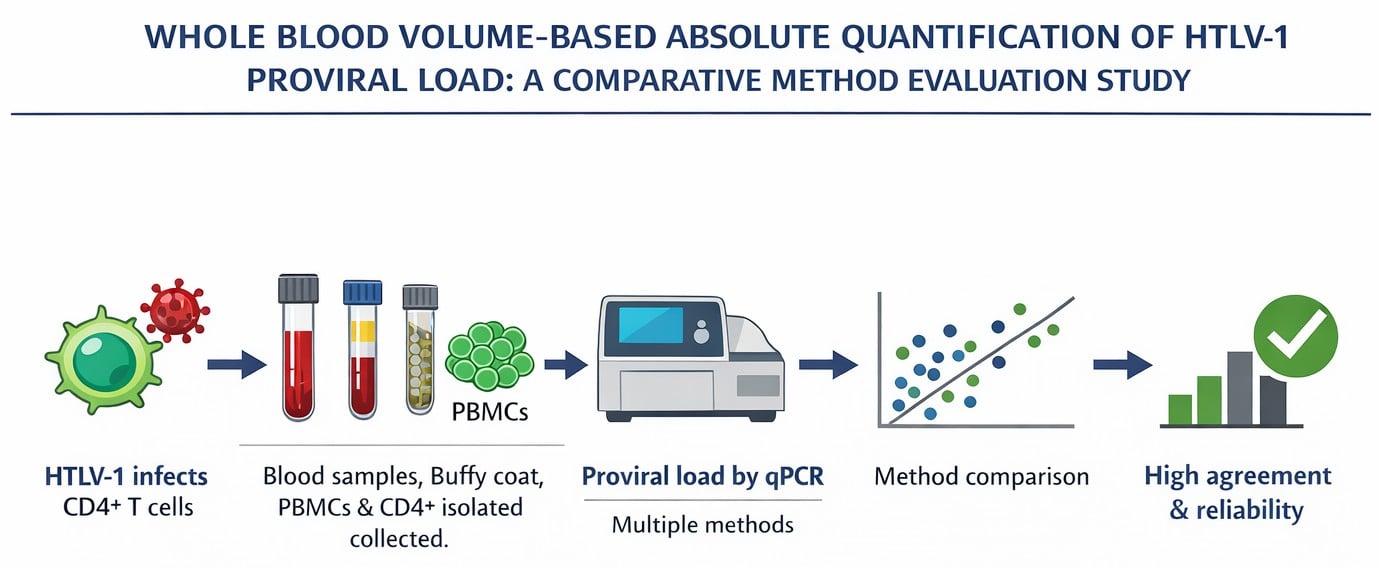
The proviral load of human T-cell lymphotropic virus type 1 (HTLV-1) is an important biomarker associated with the monitoring and risk stratification of adult T-cell leukemia/lymphoma (ATL) and HTLV-1-associated myelopathy/tropical spastic paraparesis (HAM/TSP). However, the lack of standardized quantification methods limits its broader application. This study evaluated a novel absolute quantification approach based on whole blood volume and compared its performance with established protocols. A total of 66 HTLV-1-infected individuals were analyzed using six qPCR-based methodologies, including volumetric quantification (copies/µL) by absolute quantification and the Tamegão-Lopes method, as well as normalization per 1000 cells (whole blood, buffy coat, PBMCs, and CD4+ T cells). Association and agreement were assessed using Pearson’s correlation, Bland–Altman analysis, concordance correlation coefficients (CCCs), and Deming regression. Absolute quantification showed strong correlation with both the Tamegão-Lopes method and CD4+-based quantification (r = 0.93 and 0.84, respectively; p < 0.001) and high agreement (CCC = 0.866 and 0.811, respectively), with modest systematic bias (−0.273 log10 copies/µL and 0.115 log10 copies/103 cells, respectively). Leukocyte-normalized methods showed greater discrepancies, likely due to dilution by uninfected cells. These findings show that quantification based on total blood volume is a simplified, operationally feasible alternative for assessing HTLV-1 proviral load.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Aerosol Inhalation of a Recombinant H7N9 Hemagglutinin Antigen Elicits Systemic and Mucosal Immune Responses in Mice
by
Zhuoran Hou, Han Wang, Bin Zhang, Ruixi Liu, Yuli Zhang, Ye Yang, Jianxin Wu, Xuchen Hou, Xiuguo Ge, Jun Wu and Bo Liu
Viruses 2026, 18(5), 579; https://doi.org/10.3390/v18050579 - 21 May 2026
Abstract
Highly pathogenic avian influenza A (H7N9) remains a threat to poultry health and poses a zoonotic risk, highlighting the need for vaccine antigens capable of inducing both systemic and mucosal immunity. In this study, we evaluated X33CLS-H7, a clarified cell-lysate supernatant derived from
[...] Read more.
Highly pathogenic avian influenza A (H7N9) remains a threat to poultry health and poses a zoonotic risk, highlighting the need for vaccine antigens capable of inducing both systemic and mucosal immunity. In this study, we evaluated X33CLS-H7, a clarified cell-lysate supernatant derived from glycoengineered Pichia pastoris expressing H7 hemagglutinin, in BALB/c mice following intramuscular(i.m.) injection, nebulized inhalation, or intranasal instillation. H7 expression and hemagglutination activity were confirmed by Western blotting and hemagglutination assay, respectively. Serum HA7-specific IgG and IgA responses, hemagglutination inhibition(HI) activity, H7N9 pseudovirus neutralization, bronchoalveolar lavage fluid (BALF) antibodies, and safety readouts were assessed. After two i.m. immunizations, X33CLS-H7 induced the strongest systemic antibody responses, with an HI geometric mean titer of 1:1622 95% CI, 1:1108–1:2348 and a mean log10 NT50 of 4.62. Respiratory immunization also elicited antibody responses. After four doses, high-dose nebulized delivery produced the strongest responses among the respiratory delivery regimens, with serum IgG and IgA titers of 1.02 × 105 and 2.24 × 103, respectively, an endpoint HI GMT r of 1:457 95% CI, 1:211–1:971, and a mean log10 NT50 of 3.77 compared with 2.02 in saline controls. High-dose nebulized delivery also generated detectable HA7-specific IgG and IgA responses in bronchoalveolar lavage fluid. No overt local or systemic toxicity signals were observed under the tested conditions. These findings indicate that X33CLS-H7 retains HA7-associated antigenicity and can induce systemic and respiratory mucosal antibody responses, supporting its further evaluation as a simplified and scalable H7N9 vaccine antigen candidate.
Full article
(This article belongs to the Special Issue Animal Models in Emerging/Re-Emerging Infectious Diseases)
►▼
Show Figures
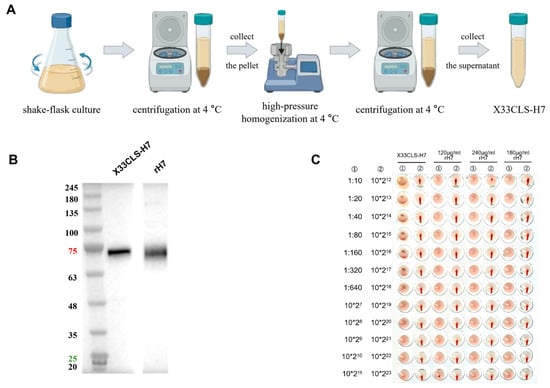
Figure 1
Open AccessArticle
Validated Quantification of HHV-8 DNA Using Inter-Convertible Plasmid and Cell-Derived Calibrators: Optimization of a Whole-Blood qPCR Assay
by
Celeste Luján Pérez, Carlos Ochoa Gamboa, Mónica Tous, Julián Hazan, Marcelo Rodríguez, Daniela Feliciotti, Lucía Irazu and Carlos Zala
Viruses 2026, 18(5), 578; https://doi.org/10.3390/v18050578 - 21 May 2026
Abstract
Human herpesvirus 8 (HHV-8) is the etiologic agent of Kaposi’s sarcoma (KS), primary effusion lymphoma (PEL), multicentric Castleman disease (MCD), and KS-associated immune reconstitution inflammatory syndrome (IRIS-KS). Quantifying HHV-8 DNA in whole blood is clinically relevant, yet laboratory practices remain heterogeneous. Here, we
[...] Read more.
Human herpesvirus 8 (HHV-8) is the etiologic agent of Kaposi’s sarcoma (KS), primary effusion lymphoma (PEL), multicentric Castleman disease (MCD), and KS-associated immune reconstitution inflammatory syndrome (IRIS-KS). Quantifying HHV-8 DNA in whole blood is clinically relevant, yet laboratory practices remain heterogeneous. Here, we developed and validated an in-house quantitative PCR (qPCR) assay targeting ORF26, optimized for whole blood. Assay calibration used plasmid, BCBL-1 cell–derived, and commercial HHV-8 DNA standards. Analytical validation was performed following the Clinical and Laboratory Standards Institute (CLSI) guidelines and the Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) guidelines and showed a 95% limit of detection of 65.7 copies/reaction, efficiencies of 90–101% (R2 > 0.99), and intra/inter-assay coefficients of variation < 6.5%. Strong correlations were observed among the three calibrators (R2 > 0.97).Clinical validation against a composite reference yielded 100% sensitivity, specificity, PPV, and NPV. Viral loads (log10 copies/mL) varied by clinical condition: classic KS and transplant-associated KS showed the lowest medians (2.30–2.23), MCD HIV− and PEL intermediate values (2.83–3.72), and epidemic KS, MCD HIV+, and IRIS-KS the highest (4.12, 4.86, and 5.03, respectively). Viremia > 5 log10 copies/mL was associated with uncontrolled E-KS, MCD HIV+, and IRIS-KS. Longitudinal follow-up revealed viral load decline paralleled clinical improvement. This validated assay provides a robust, affordable tool for HHV-8 quantification in whole blood and supports its integration into diagnostic workflows and patient monitoring.
Full article
(This article belongs to the Special Issue Herpesviruses and Associated Diseases, 2nd Edition)
►▼
Show Figures

Figure 1
Open AccessArticle
The Melon Sterol Transporter Niemann-Pick C1 Protein Is a New Interactor of Cucumber mosaic virus Movement Protein
by
Núria Real, Irene Villar, Bin Liu, Manale Gajjout, Weina Hou and Ana Montserrat Martín-Hernández
Viruses 2026, 18(5), 577; https://doi.org/10.3390/v18050577 - 20 May 2026
Abstract
Plant viruses need to use many host factors to establish infection. During the viral cycle, intracellular transport is fundamental to reach the plasmodesmata to enable cell-to-cell transport. Cucumovirus CMV (cucumber mosaic virus, CMV) can infect plants from most economically important crops. To identify
[...] Read more.
Plant viruses need to use many host factors to establish infection. During the viral cycle, intracellular transport is fundamental to reach the plasmodesmata to enable cell-to-cell transport. Cucumovirus CMV (cucumber mosaic virus, CMV) can infect plants from most economically important crops. To identify additional host proteins involved in CMV movement in melon, we used the MP as a bait to screen a Yeast two-hybrid cDNA library from CMV-infected plants and identified a Niemann-Pick C1 (NPC1) protein as a novel MP interactor. NPC1 is a transmembrane protein involved in cholesterol transport in animal cells, but also in the infection by several viruses of different families. The identified clone from the melon NPC1 gene spans from exons 25 to 28 and includes two introns. Notably, deletion of the two introns and exon 28 does not impair the interaction capacity of the remaining peptide. The identified CmNPC1 gene maps to chromosome 11. In addition, the melon genome encodes a second copy of NPC1 in chromosome 7 (CmNPC1-C7), highly similar. Functional assays revealed that the interaction domain of CmNPC1-C7 also interacts with CMV MP, suggesting that both genes could have a role in CMV infection. This study represents the first report linking NPC1 to the infection process of a plant virus, expanding our understanding of plant–virus interactions.
Full article
(This article belongs to the Special Issue Plant Virus Resistance—2nd Edition)
►▼
Show Figures
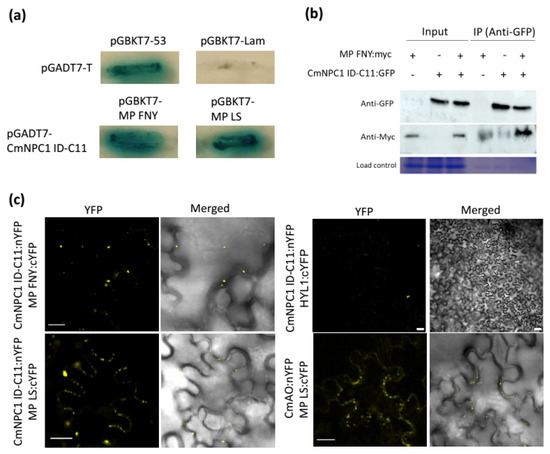
Figure 1
Open AccessData Descriptor
From Discovery to Cure—Where Are We Now? Mortality Trends in Chronic Hepatitis C: An Analysis of CDC WONDER Database (1999–2023)
by
Ashraf Ullah, Hina Wazir, Abdullah Sultany, Khalil Ur Rehman, Mohammad Ibrahim Sultani, Naeem Ahmed Khan, Saeed A. Khan, Mati Ullah Dad Ullah and Amlish Gondal
Viruses 2026, 18(5), 576; https://doi.org/10.3390/v18050576 - 20 May 2026
Abstract
Background: Hepatitis C virus (HCV) remains a major cause of preventable liver-related mortality in the United States despite highly effective direct-acting antivirals (DAAs). Contemporary assessment of mortality trends and disparities is essential for elimination efforts. Methods: Using CDC WONDER multiple cause-of-death data (1999–2023),
[...] Read more.
Background: Hepatitis C virus (HCV) remains a major cause of preventable liver-related mortality in the United States despite highly effective direct-acting antivirals (DAAs). Contemporary assessment of mortality trends and disparities is essential for elimination efforts. Methods: Using CDC WONDER multiple cause-of-death data (1999–2023), we identified HCV-related deaths using ICD-10 codes for acute and chronic HCV (B17.1, B18.2) and calculated age-adjusted mortality rates (AAMRs) per 100,000 (2000 US standard). Rates were stratified by sex, race/ethnicity, census region, and 2013 NCHS urban–rural classification. Joinpoint regression quantified temporal inflection points and annual percent changes (APCs). Results: Overall HCV-related AAMR increased from 1.8 (1999) to a peak of 5.0 (2014), then declined to 2.3 (2023), with a marked post-2014 decrease (APC −8.2%). Mortality was consistently higher in males than females (2023 rate ratio 2.57). In 2023, American Indian/Alaska Native individuals had the highest mortality (AAMR 8.7; rate ratio 3.48 vs. non-Hispanic White), followed by non-Hispanic Black individuals (AAMR 6.2; rate ratio 2.48). Mortality remained highest in the West and was higher in non-metropolitan than metropolitan counties (AAMR 2.8 vs. 2.3; rate ratio 1.22), with a slower post-2014 decline in non-metropolitan areas. Conclusions: Our findings indicate that while the DAA era has been associated with a substantial reduction in HCV-related mortality at the national level, this progress has not been uniform across all populations. Persistent excess mortality among Native American and non-Hispanic Black individuals may reflect inequities in the HCV care cascade, including screening, confirmatory testing, linkage to specialty care, insurance-related restrictions, and the high cost of antiviral therapy. These results highlight the need for policies and public health strategies that improve equitable and affordable access to curative HCV treatment.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
USP17L13 Enhances Influenza a Virus Replication by Mediating the Degradation of RIG-I and MDA5
by
Yaping Zhang, Chen Qin, Yichao Zhuang, Lei Chen, Xianying Zeng, Li Jiang, Chengjun Li, Hualan Chen and Huihui Kong
Viruses 2026, 18(5), 575; https://doi.org/10.3390/v18050575 - 20 May 2026
Abstract
The innate immune system, particularly the retinoic acid-inducible gene I (RIG-I)-like receptor (RLR) signaling pathway, is a major early defense barrier against influenza A virus infection. However, excessive immune responses can trigger lethal cytokine storms and severe immune-mediated pathology. In this study, we
[...] Read more.
The innate immune system, particularly the retinoic acid-inducible gene I (RIG-I)-like receptor (RLR) signaling pathway, is a major early defense barrier against influenza A virus infection. However, excessive immune responses can trigger lethal cytokine storms and severe immune-mediated pathology. In this study, we performed a genome-wide CRISPR/dCas9 gene activation screen in human lung epithelial (A549) cells by using an A/Puerto Rico/8/1934 (H1N1) reporter virus, and identified the ubiquitin-specific protease USP17L13 as a novel negative regulator of innate immunity that promotes influenza virus replication. Overexpression of USP17L13 significantly enhanced the replication of multiple subtypes of influenza viruses in A549 cells, including a human pandemic H1N1 virus, seasonal H3N2 viruses, as well as a globally circulating clade, 2.3.4.4b, of the highly pathogenic avian H5N1 virus. Transcriptomic analysis demonstrated that USP17L13 suppresses host antiviral defenses by downregulating nuclear factor kappa B (NF-κB) signaling and arachidonic acid metabolism, while upregulating pathways associated with ribosomal translation and oxidative phosphorylation to facilitate viral production. Mechanistically, USP17L13 attenuates the host interferon (IFN) response by promoting the degradation of the key viral RNA sensors, RIG-I, and melanoma differentiation-associated protein 5 (MDA5). Further analysis revealed that USP17L13 is inducible by type I and type II interferons as well as inflammatory cytokines, suggesting that it may act as a negative-feedback regulator to limit excessive inflammation. Collectively, our findings identify USP17L13 as a previously unrecognized proviral host factor and provide new insight into how host deubiquitinases shape influenza virus-host interactions, with potential implications for host-directed approaches to controlling excessive inflammation during viral infection and improving influenza vaccine production.
Full article
(This article belongs to the Special Issue Avian Viruses and Antiviral Immunity)
►▼
Show Figures

Figure 1
Open AccessArticle
Targeting Host Metabolic and Epigenetic Rewiring Blocks Lytic Gammaherpesvirus Production
by
Morgan C. Jones, Tina M. Le, Connor J. Mahoney, Sara K. Hartman, Robynne D. Dona, Yennifer A. Gaspar, Sennah J. Hong, Benjamin R. Sheirbon, Thelma M. Escobar and Tracie Delgado
Viruses 2026, 18(5), 574; https://doi.org/10.3390/v18050574 - 19 May 2026
Abstract
Gammaherpesviruses are oncogenic viruses that reprogram host cell metabolism to support viral production. Among these, murine herpesvirus 68 (MHV-68) serves as a model system for studying lytic gammaherpesvirus infection and associated host cell changes. To characterize host transcriptional alterations induced throughout lytic gammaherpesvirus
[...] Read more.
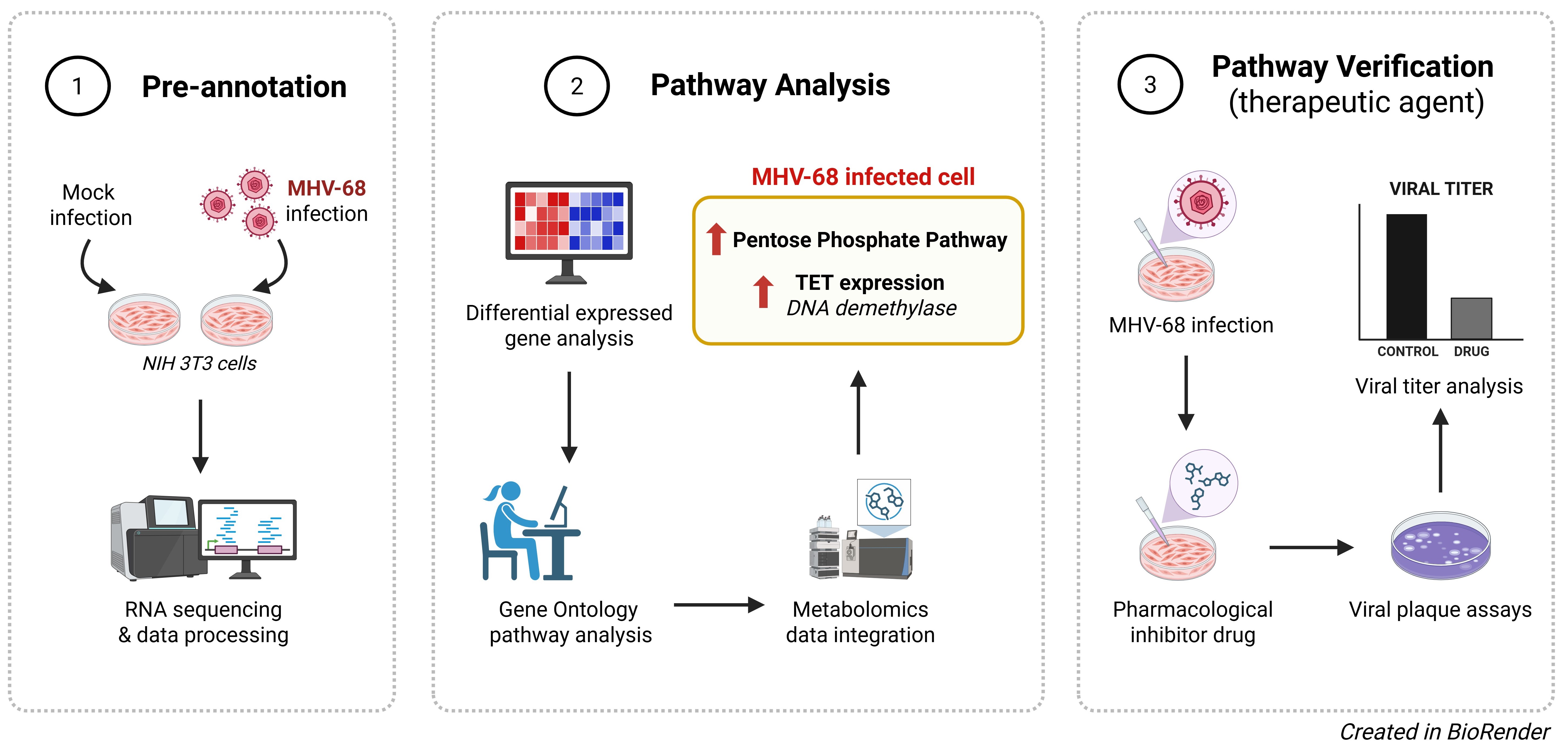
Gammaherpesviruses are oncogenic viruses that reprogram host cell metabolism to support viral production. Among these, murine herpesvirus 68 (MHV-68) serves as a model system for studying lytic gammaherpesvirus infection and associated host cell changes. To characterize host transcriptional alterations induced throughout lytic gammaherpesvirus infection and identify novel host pathways that may be therapeutically targeted, we performed temporal bulk RNA-sequencing of mock- and MHV-68-infected NIH 3T3 cells at various timepoints throughout the lytic cycle. Our analysis revealed widespread and progressive host gene expression changes, including robust innate immune pathways and extensive remodeling of metabolic gene expression. We further identified a strong activation of the pentose phosphate pathway (PPP) genes, accompanied by increased abundance in PPP metabolic intermediates. Pharmacological inhibition of the PPP with 6-aminonicotinamide (6-AN) reduced infectious virus production. Moreover, at the intersection of metabolic and transcriptional reprogramming, we identified infection-associated gene expression changes in chromatin-modulating enzymes, including Tet2, and their metabolite co-factors, such as α-KG. Pharmacological inhibition of Ten-Eleven Translocation (TET) enzymatic activity led to a marked decrease in infectious MHV-68 production. Collectively, these findings define a novel metabolic–epigenetic crosstalk that supports productive gammaherpesvirus replication and identifies host pathways that can be targeted to treat lytic gammaherpesvirus infections.
Full article
(This article belongs to the Special Issue Pharmacology of Antiviral Drugs, 2nd Edition)
►▼
Show Figures
Graphical abstract
Open AccessCorrection
Correction: Lara-Romero et al. Phylogenetic and Molecular Analysis of the Porcine Epidemic Diarrhea Virus in Mexico during the First Reported Outbreaks (2013–2017). Viruses 2024, 16, 309
by
Rocío Lara-Romero, Rebeca Martínez-Bautista, Raúl González-Martínez, Jazmín De la Luz-Armendáriz, Irma Herrera-Camacho, Nora Rosas-Murrieta, Laura Márquez-Valdelamar and José Francisco Rivera-Benítez
Viruses 2026, 18(5), 573; https://doi.org/10.3390/v18050573 - 19 May 2026
Abstract
Author Name and Order [...]
Full article
Open AccessArticle
A Multidisciplinary Healthy Aging Program in Comprehensive HIV Care: Multidomain Screening, Clinical Interventions, and Cardiometabolic Risk Management
by
Steven Y. Hong, Deborah Woodley, Megan Pao, Holly Goetz, Alejandro Alvarez, Max White, Bruce Hirsch, Edith Burns and Joseph P. McGowan
Viruses 2026, 18(5), 572; https://doi.org/10.3390/v18050572 - 19 May 2026
Abstract
Background: People living with HIV (PLWH) are increasingly reaching older ages due to the success of antiretroviral therapy. However, aging with HIV is associated with increased risk of multimorbidity, neurocognitive impairment, frailty, psychosocial stress, and functional decline. Multidomain geriatric screening framed within an
[...] Read more.
Background: People living with HIV (PLWH) are increasingly reaching older ages due to the success of antiretroviral therapy. However, aging with HIV is associated with increased risk of multimorbidity, neurocognitive impairment, frailty, psychosocial stress, and functional decline. Multidomain geriatric screening framed within an Age-Friendly 4Ms Framework (Mentation, Medication, Mobility, What Matters Most) and consideration of multi-complexity may help identify aging-related vulnerabilities and guide multidisciplinary care with greater impact on patient outcomes. However, real-world implementation of such programs within HIV clinical settings remains limited. Methods: We conducted a retrospective analysis of adults aged ≥50 years enrolled in a multidisciplinary Healthy Aging Program within a large, integrated HIV care system. Multidomain screening assessments included cognitive evaluation (Montreal Cognitive Assessment), mental health screening (PHQ-2, GAD-2), functional assessment (Katz ADL, Lawton IADL), frailty screening (Edmonton Frail Scale), and intrinsic capacity domains using the WHO Integrated Care for Older People (ICOPE) framework. Screening results, referrals, clinical interventions, and cardiometabolic risk management measures were extracted from clinical program databases and electronic medical records. Results: A total of 317 adults aged ≥50 years completed multidomain screening. Participants had well-controlled HIV infection, with viral suppression in 96.2% and a median CD4 count of 660 cells/mm3. Despite this, aging-related vulnerabilities were common. Overall, 78.4% of participants had at least one abnormal screening domain. Cognitive impairment was identified in nearly half of individuals screened, including mild impairment in 39.8% and moderate impairment in 8.7%. Functional limitations were identified in 10.1% of participants, while anxiety symptoms were present in 9.5%. Sensory impairments were common, including vision impairment in 36.5% of participants. Polypharmacy was prevalent, with 33.2% of participants prescribed five or more chronic medications. Screening frequently generated multidisciplinary referrals, including behavioral health services (42.3%), social work support (42.9%), and pharmacist-led cardiometabolic risk review (56.8%). Age-stratified analyses demonstrated similar prevalence of screening abnormalities across age groups, including individuals aged 50–59 years. Modest improvements in cardiometabolic preventive care were observed during follow-up. Statin utilization increased from 65.6% at baseline to 70.0% at 12 months, and LDL cholesterol declined modestly during the observation period. Conclusions: Multidomain screening integrated into routine HIV care identified a high prevalence of aging-related vulnerabilities among PLWH aged ≥50 years despite excellent virologic control. These findings suggest that aging-related risk in HIV is not adequately captured by chronological age alone and support early, universal implementation of multidomain screening within HIV care models.
Full article
(This article belongs to the Special Issue HIV and Aging)
►▼
Show Figures

Figure 1
Open AccessReview
Living on the Edge: The Goldilocks Zone of Polyomavirus Replication and Persistence
by
Wenqing Yuan, Sheila A. Haley, Michael J. Imperiale and Walter J. Atwood
Viruses 2026, 18(5), 571; https://doi.org/10.3390/v18050571 - 19 May 2026
Abstract
BK and JC Polyomaviruses (BKPyV and JCPyV) are ubiquitous human pathogens capable of establishing lifelong, asymptomatic persistence in the majority of the global population. While decades of research have focused on their lytic replication cycles and the development of severe diseases, such as
[...] Read more.
BK and JC Polyomaviruses (BKPyV and JCPyV) are ubiquitous human pathogens capable of establishing lifelong, asymptomatic persistence in the majority of the global population. While decades of research have focused on their lytic replication cycles and the development of severe diseases, such as polyomavirus-associated nephropathy (PVAN) caused by BKPyV and progressive multifocal leukoencephalopathy (PML) caused by JCPyV, their primary evolutionary strategy is one of persistence rather than pathogenesis. This review shifts the perspective from a replication-centric framework towards an evolutionary persistence model, detailing the multi-layered host and viral determinants that maintain the homeostatic balance. At the cellular level, viral genomes are restricted by chromatinization into minichromosomes and host S-phase licensing. These constraints are reinforced by innate immune sensing and adaptive T-cell and antibody responses that curtail systemic dissemination while permitting periodic, low-level urinary shedding, which is essential for horizontal transmission. In addition to these host barriers, the viruses utilize intrinsic regulatory mechanisms to prevent excessive replication and immune detection, including the stable archetype non-coding control region (NCCR), viral microRNAs that downregulate early gene expression, and the small t antigen (STAg). Finally, we address unresolved questions regarding the full spectrum of cellular reservoirs, the molecular triggers of reactivation, and the ecological factors shaping their transmission routes. Understanding these maintenance mechanisms is crucial for refining clinical interventions and managing the rare, devastating transitions from silent persistence to lytic disease.
Full article
(This article belongs to the Special Issue Polyomavirus)
►▼
Show Figures

Figure 1
Open AccessArticle
Relationship Between the Size–Frequency Distribution of Nucleopolyhedrovirus Occlusion Bodies and Their Insecticidal Characteristics on Spodoptera frugiperda (Lepidoptera: Noctuidae)
by
Cristian Ángel-García, Rodrigo Lasa, Joel E. López-Meza, Selene Ramos-Ortiz, Trevor Williams and Ana Mabel Martínez-Castillo
Viruses 2026, 18(5), 570; https://doi.org/10.3390/v18050570 - 19 May 2026
Abstract
The Spodoptera frugiperda multiple nucleopolyhedrovirus (SfMNPV) is an important pathogen of the fall armyworm and is used as the basis for biological insecticides. In this study, we examined the relationship between the size–frequency distribution of SfMNPV occlusion bodies (OBs) and their insecticidal characteristics
[...] Read more.
The Spodoptera frugiperda multiple nucleopolyhedrovirus (SfMNPV) is an important pathogen of the fall armyworm and is used as the basis for biological insecticides. In this study, we examined the relationship between the size–frequency distribution of SfMNPV occlusion bodies (OBs) and their insecticidal characteristics when collected at the end of the replication cycle. Exposure of OBs to 40%, 70%, and 90% (wt/wt) glycerol had no effect on OB pathogenicity. Glycerol density gradient (50–100%) centrifugation was used to separate OBs into two fractions. OBs recovered from the upper fraction of the gradient had a significantly smaller median cross-sectional area than those harvested from the lower fraction. These fractions also differed significantly in their size–frequency distributions. The OB concentration–mortality response of S. frugiperda second instars did not differ significantly between the two fractions or with non-centrifuged OBs. The median survival time was similar for insects inoculated with OBs from the upper and lower fractions but was significantly shorter in insects inoculated with non-centrifuged OBs. The proportion of mature OBs (67–71%) and the number of viral genome copies (1.33–1.40 × 108 copies/µL) did not differ significantly between the upper and lower OB fractions. These findings suggest that altering the size–frequency distribution by density gradient centrifugation is not a useful technique for selecting large OBs with high insecticidal activity as part of the baculovirus insecticide production process. Future studies should evaluate a range of OB size separation techniques to determine their effects on OB insecticidal characteristics.
Full article
(This article belongs to the Section Invertebrate Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
Wastewater-Based Surveillance of SARS-CoV-2 for Early Warning of COVID-19 Infection Dynamics
by
Qiuyan Zhao, Xinye Zhang, Jing Peng, Xiaoyan Ma, Yongxing Wang, Jun Luo, Xiaohan Su, Siyu Yang, Xiaona Yan, Yuan Wei and Jie Zhang
Viruses 2026, 18(5), 569; https://doi.org/10.3390/v18050569 - 18 May 2026
Abstract
Wastewater-based epidemiology has emerged as a valuable complementary tool for population-level monitoring. This study evaluated the early warning value of wastewater surveillance for monitoring SARS-CoV-2 and its correlation with COVID-19 infection trends. From May 2024 to December 2025, 526 wastewater samples were collected
[...] Read more.
Wastewater-based epidemiology has emerged as a valuable complementary tool for population-level monitoring. This study evaluated the early warning value of wastewater surveillance for monitoring SARS-CoV-2 and its correlation with COVID-19 infection trends. From May 2024 to December 2025, 526 wastewater samples were collected from five treatment plants. Spearman correlation and a quasi-Poisson generalized additive model (adjusting for wastewater temperature) were used to assess relationships between SARS-CoV-2 RNA concentration, the number of reported cases, and lag associations. Wastewater viral loads (copies/mL) significantly correlated with reported cases. Wastewater temperature was positively correlated with both viral concentrations and case numbers. A significant lagged association was observed for the N gene, with relative risk peaking at a 10-day lag. Although the ORF1ab gene was not significant for most lag periods, its temporal trend was consistent with that of the N gene. Wastewater surveillance of SARS-CoV-2, particularly targeting the N gene, can effectively predict COVID-19 infection dynamics with a 10-day lead time, thereby supporting wastewater surveillance as an early warning tool for public health monitoring.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures

Figure 1
Open AccessCorrection
Correction: Mofed et al. Construction of a Macrophage-Tropic Subtype C HIV-1-mGreenLantern Reporter Virus for Studies on HIV-1 Replication and the Impact of Methamphetamine. Viruses 2024, 16, 1859
by
Dina Mofed, Angelo Mandarino, Xuhong Wu, Yuekun Lang, Anjali Gowripalan, Ganjam V. Kalpana and Vinayaka R. Prasad
Viruses 2026, 18(5), 568; https://doi.org/10.3390/v18050568 - 18 May 2026
Abstract
►▼
Show Figures
Error in Figure [...]
Full article
Figure 4
Open AccessArticle
Porcine Reproductive and Respiratory Syndrome Virus NSP8 Suppresses NF-κB Signaling by Hijacking Host UBE2K and IKKα
by
Da Liu, Yan Yan, Xuezhen Fu, Linglong Qin, Jiayu Ma, Hui Zhou, Shiping Sun, Haimin Li, Weiren Dong and Jiyong Zhou
Viruses 2026, 18(5), 567; https://doi.org/10.3390/v18050567 - 18 May 2026
Abstract
The Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) has evolved sophisticated immune-evasion strategies to establish a productive infection in the host, primarily by counteracting the innate antiviral response. Here, we demonstrate for the first time that the PRRSV non-structural protein NSP8 suppresses NF-κB-dependent
[...] Read more.
The Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) has evolved sophisticated immune-evasion strategies to establish a productive infection in the host, primarily by counteracting the innate antiviral response. Here, we demonstrate for the first time that the PRRSV non-structural protein NSP8 suppresses NF-κB-dependent antiviral signalling by hijacking the host ubiquitin-conjugating enzyme UBE2K and inducing the degradation of IKKα, a pivotal kinase in the NF-κB pathway. PRRSV infection led to significant upregulation of host UBE2K, which in turn facilitated viral replication. Mechanistically, we found that NSP8 interacts directly with IKKα, triggering its degradation by the proteasome. Furthermore, we revealed that this process was facilitated by the host protein UBE2K, which acted as a crucial cofactor by directly interacting with NSP8 and thereby enhancing its activity against IKKα. This disruption blocked the activation of the NF-κB pathway and suppressed the expression of downstream antiviral factors, such as TNF-α, IL-6 and IFN-β, ultimately facilitating PRRSV replication. All of these findings showed that NSP8 is an important part of the process by which the host NF-κB pathway is blocked by viruses. This is a new way in which PRRSV avoids the immune system.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
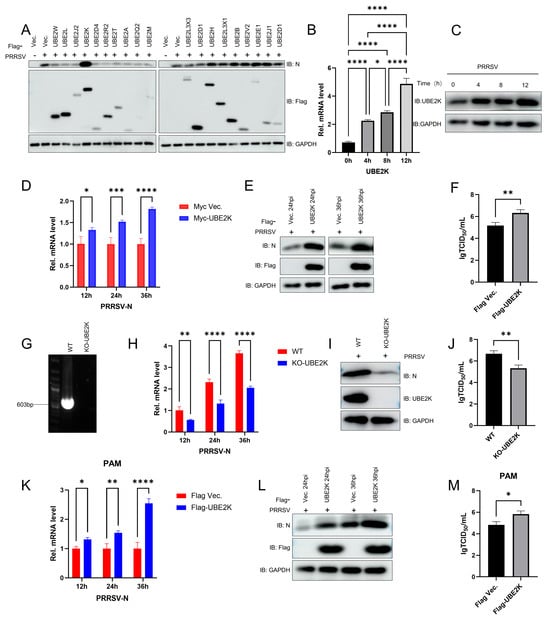
Figure 1
Open AccessArticle
The IFIT3 Protein of Porcine Induces Interferon Signaling and Inhibits the Early Gene Expression of African Swine Fever Virus
by
Wen-Li Wang, Deng-Wu Han, Xing Yang, Xi-Juan Shi, Ye-Sheng Shen, Shu-Yao Tian, Zhi-Hai Chang, Deng-Ji Zhang, Qiao-Ying Zeng, Shi-Jun Bao, Hai-Xue Zheng and Ruo-Qing Mao
Viruses 2026, 18(5), 566; https://doi.org/10.3390/v18050566 - 17 May 2026
Abstract
African swine fever virus (ASFV) is the causative agent of African swine fever (ASF), a fatal and highly contagious disease, resulting in enormous losses to the global swine industry. No licensed vaccines or effective therapeutics are currently available to control ASFV infection. Interferons
[...] Read more.
African swine fever virus (ASFV) is the causative agent of African swine fever (ASF), a fatal and highly contagious disease, resulting in enormous losses to the global swine industry. No licensed vaccines or effective therapeutics are currently available to control ASFV infection. Interferons (IFNs) serve as key mediators of host antiviral immunity by inducing interferon-stimulated genes (ISGs), but the specific mechanisms by which individual ISGs restrict ASFV replication remain unclear. Interferon-induced protein with tetratricopeptide repeats 3 (IFIT3, also called ISG60) has been shown to exhibit antiviral activity against various viruses, but its role in ASFV infection has not been previously studied. Here, we used porcine alveolar macrophages (PAMs), the primary target cells of ASFV, to investigate IFIT3’s function in ASFV replication. We found that overexpression of IFIT3 inhibited ASFV replication, while its knockdown enhanced viral propagation. Mechanistically, IFIT3 directly blocked ASFV adsorption to host cells, thereby suppressing all subsequent stages of the viral cycle. IFIT3 also specifically interacted with ASFV F334L, an early viral gene product that encodes the small subunit of ribonucleotide reductase, a key enzyme for viral DNA synthesis. Additionally, IFIT3 positively regulated the STAT1/TBK1/IRF3 signaling axis: its overexpression increased phosphorylation of TBK1 and IRF3, as well as the protein level of STAT1, while IFIT3 knockdown attenuated activation of these molecules. Transcriptomic analysis of IFIT3-knockout PAMs revealed significant suppression of innate immune pathways, including type I interferon, JAK-STAT, and RIG-I-like receptor pathways, along with downregulated expression of core antiviral molecules such as ISG15, MX1, and STAT1. Conversely, pathways related to viral adsorption, endocytosis, and cytoskeleton were activated, and pathways involved in protein translation initiation, endoplasmic reticulum stress, and autophagy were dysregulated, creating a favorable intracellular environment for ASFV replication. In conclusion, IFIT3 restricts ASFV replication possibly by inhibiting viral adsorption and promoting innate immune signaling, identifying it as a potential therapeutic target against ASFV. This study’s limitation is its in vitro PAM model; future work will validate IFIT3’s role in vivo and develop targeted inhibitors.
Full article
(This article belongs to the Special Issue Virus–Host Protein Interactions)
►▼
Show Figures
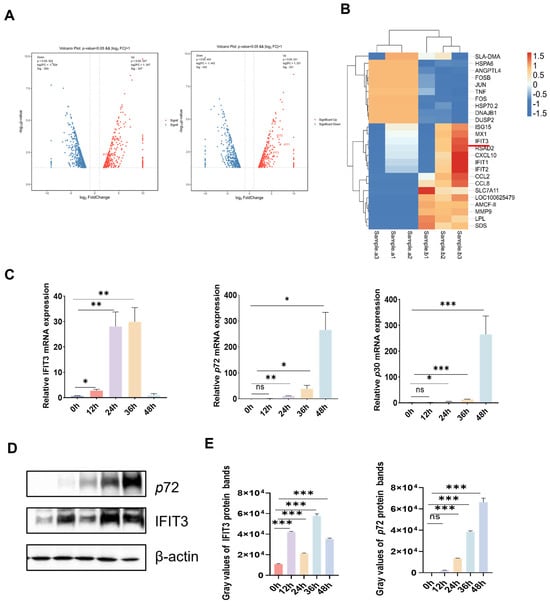
Figure 1

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
20 May 2026
Meet Us at the 10th National Congress of the Italian Society for Virology, 14–16 June 2026, Palermo, Italy
Meet Us at the 10th National Congress of the Italian Society for Virology, 14–16 June 2026, Palermo, Italy

15 May 2026
Meet Us at the 1st South-East Transnational Congress of Microbial Sciences (SETCoMS), 10–13 June 2026, Belgrade, Serbia
Meet Us at the 1st South-East Transnational Congress of Microbial Sciences (SETCoMS), 10–13 June 2026, Belgrade, Serbia

Topics
Topic in
Biomedicines, Cancers, IJMS, Vaccines, Viruses, JMP
Advances in HPV-Driven Head and Neck Cancer: Pathogenesis, Treatment, and Prevention
Topic Editors: Shilpi Gupta, Bhudev Chandra Das, Prabhat KumarDeadline: 20 April 2027
Topic in
Antibiotics, Diseases, Vaccines, Viruses, Epidemiologia
HIV Prevention and Treatment: A Guide to Health and Safety
Topic Editors: Olanrewaju Oladimeji, Martha ChadyiwaDeadline: 31 December 2027

Conferences
Special Issues
Special Issue in
Viruses
Influenza B Virus: From Infection to Prevention
Guest Editor: Saira HussainDeadline: 30 May 2026
Special Issue in
Viruses
Citrus Viral Diseases: Advances in Knowledge and Technologies and Their Containment Measures
Guest Editors: Antonino F. Catara, Salvatore Davino, Moshe Bar-JosephDeadline: 31 May 2026
Special Issue in
Viruses
Roles of Macrophages in Viral Infections, 2nd Edition
Guest Editor: Stefano AquaroDeadline: 31 May 2026
Special Issue in
Viruses
Crimean-Congo Hemorrhagic Fever Virus (CCHFV)
Guest Editor: Remi N. CharrelDeadline: 31 May 2026
Topical Collections
Topical Collection in
Viruses
Efficacy and Safety of Antiviral Therapy
Collection Editors: Giordano Madeddu, Andrea De Vito, Agnese Colpani
Topical Collection in
Viruses
Phage Therapy
Collection Editors: Nina Chanishvili, Jean-Paul Pirnay, Mikael Skurnik









