Regulatory and Non-Coding RNAs in Cancer Epigenetic Mechanisms
A topical collection in Cancers (ISSN 2072-6694). This collection belongs to the section "Molecular Cancer Biology".
Viewed by 213158Topical Collection Information
Dear Colleagues,
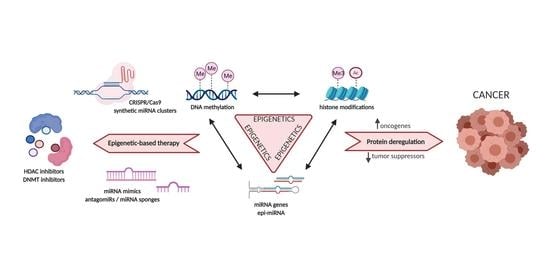
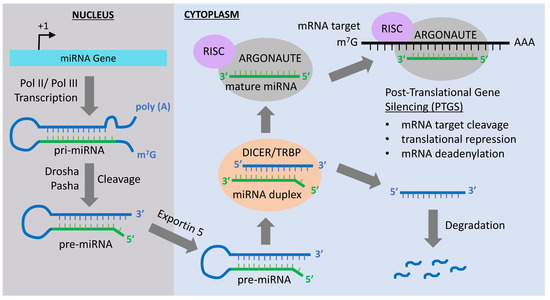
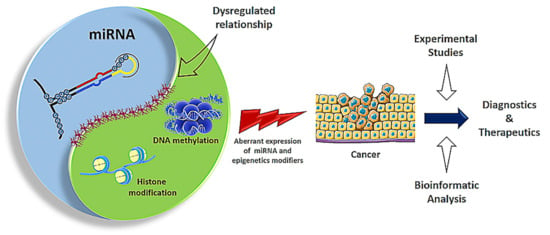
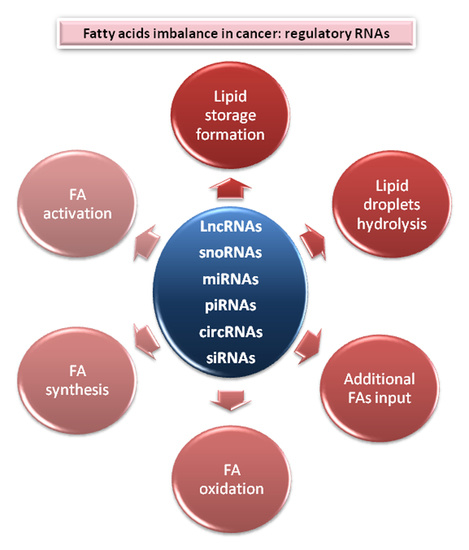
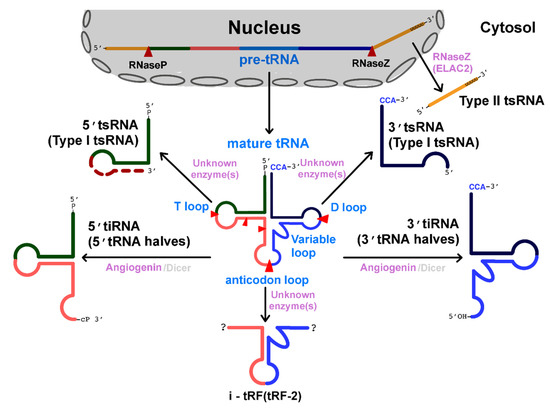
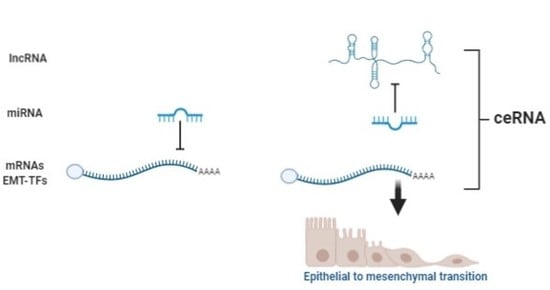
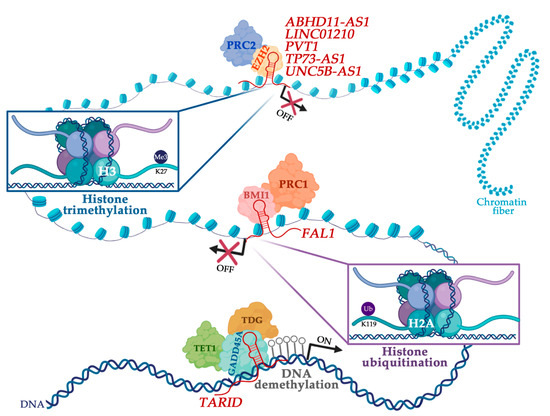
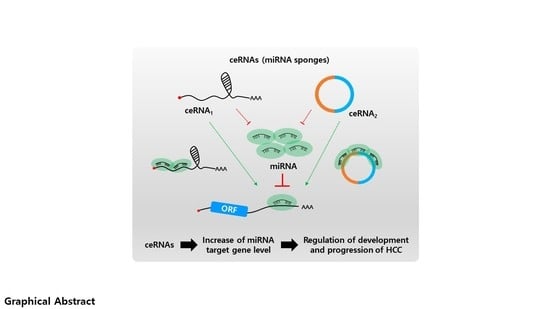
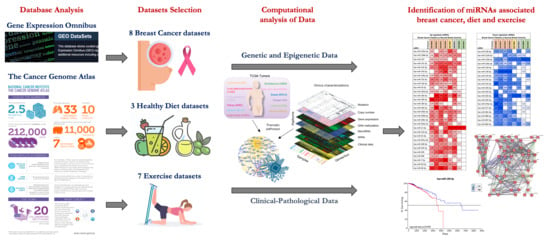
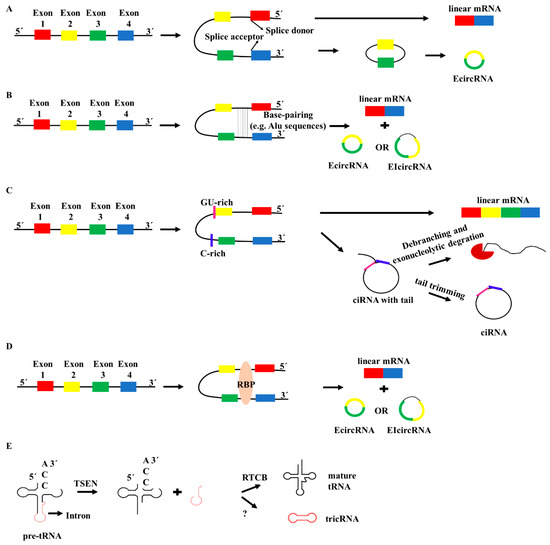
Over the years enormous progress has been made in our understanding of the way RNA controls and regulates processes in cells and organisms, and how these mechanisms—when they go awry—contribute to the progression of many diseases, including cancer. The regulatory and non-coding “RNA world” includes not only long non-coding RNAs (lncRNAs) and microRNAs (miRNAs) but also many more other types of RNAs (e.g., circular RNAs, enhancer RNAs) that are components of epigenetic regulatory mechanisms of development and cancer. The objective of this collection is to attract research studies and critical reviews on the central role of regulatory and non-coding RNAs in epigenetic mechanisms of cancer.
Prof. Dr. Nicoletta Sacchi
Guest Editor
Manuscript Submission Information
Manuscripts should be submitted online at www.mdpi.com by registering and logging in to this website. Once you are registered, click here to go to the submission form. All submissions that pass pre-check are peer-reviewed. Accepted papers will be published continuously in the journal (as soon as accepted) and will be listed together on the collection website. Research articles, review articles as well as communications are invited. For planned papers, a title and short abstract (about 250 words) can be sent to the Editorial Office for assessment.
Submitted manuscripts should not have been published previously, nor be under consideration for publication elsewhere (except conference proceedings papers). All manuscripts are thoroughly refereed through a single-blind peer-review process. A guide for authors and other relevant information for submission of manuscripts is available on the Instructions for Authors page. Cancers is an international peer-reviewed open access semimonthly journal published by MDPI.
Please visit the Instructions for Authors page before submitting a manuscript. The Article Processing Charge (APC) for publication in this open access journal is 2900 CHF (Swiss Francs). Submitted papers should be well formatted and use good English. Authors may use MDPI's English editing service prior to publication or during author revisions.
Keywords
- long non-coding RNAs
- microRNAs
- circular RNAs
- epigenetics
- development
- cancer