Journal Description
Epigenomes
Epigenomes
is an international, peer-reviewed, open access journal on epigenetics and epigenomics, published quarterly online by MDPI. The Epigenetics Society is affiliated with Epigenomes and its members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PMC, PubMed, Embase, PubAg, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q2 (Genetics and Heredity) / CiteScore - Q2 (Biochemistry, Genetics and Molecular Biology (miscellaneous))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 20.3 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the first half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.5 (2024);
5-Year Impact Factor:
3.1 (2024)
Latest Articles
Methylation of the Glucocorticoid Receptor Gene in Children with Somatic Symptom Disorder: A Case-Control Study
Epigenomes 2025, 9(2), 22; https://doi.org/10.3390/epigenomes9020022 - 13 Jun 2025
Abstract
►
Show Figures
Background: Somatic symptom disorder (SSD) in children may be influenced by stress reactivity and psychosocial factors. The glucocorticoid receptor (GR), encoded by NR3C1, is a key mediator of stress responses. However, the relationship between NR3C1 methylation and SSD remains unclear. Methods: We analyzed
[...] Read more.
Background: Somatic symptom disorder (SSD) in children may be influenced by stress reactivity and psychosocial factors. The glucocorticoid receptor (GR), encoded by NR3C1, is a key mediator of stress responses. However, the relationship between NR3C1 methylation and SSD remains unclear. Methods: We analyzed NR3C1 exon 1F methylation in cell-free DNA from saliva in 34 children with SSD and 29 age- and sex-matched controls using bisulfite amplicon sequencing. Psychological assessments included the Beck Depression Inventory-II (BDI-II) and KINDL questionnaires to evaluate associations with methylation patterns. Results: Methylation levels showed age-related differences. In children under 13, CpG sites displayed mixed methylation, and specific sites correlated with KINDL and BDI-II scores. KINDL physical and total well-being scores negatively correlated with CpG30 and positively with CpG35; BDI-II scores negatively correlated with CpG32 and CpG35. In children aged 13 or older, CpG sites showed uniformly high methylation with no correlation to psychological measures. The SSD group showed significantly higher average methylation across the exon 1F region than controls in the older age group. These children also had more cases of orthostatic dysregulation and longer illness duration. Conclusions: This study suggests age-dependent epigenetic regulation of NR3C1 in SSD. While younger children showed CpG-specific correlations with psychological symptoms, older children demonstrated uniformly high methylation and potentially reduced gene expression, potentially reflecting cumulative stress, autonomic dysfunction, and internalizing disorders such as anxiety and depression.
Full article
Open AccessReview
The Dynamic Interactions of m6A Modification and R-Loops: Implications for Genome Stability
by
Nicholas Kim and Hong Sun
Epigenomes 2025, 9(2), 21; https://doi.org/10.3390/epigenomes9020021 - 11 Jun 2025
Abstract
►▼
Show Figures
R-loops, three-stranded RNA-DNA hybrid nucleic acid structures, are recognized for their roles in both physiological and pathological processes. Regulation of R-loops is critical for genome stability as disruption of R-loop homeostasis can lead to aberrant gene expression, replication stress, and DNA damage. Recent
[...] Read more.
R-loops, three-stranded RNA-DNA hybrid nucleic acid structures, are recognized for their roles in both physiological and pathological processes. Regulation of R-loops is critical for genome stability as disruption of R-loop homeostasis can lead to aberrant gene expression, replication stress, and DNA damage. Recent studies suggest that the RNA modification, N6-methyladenosine (m6A), can modify R-loops and the writers, erasers, and readers of m6A are involved in the dynamic regulation of R-loops. Here, we discuss the reported functions of various m6A regulatory proteins in relation to R-loops, highlighting their distinct roles in recognizing and modulating the formation, stability, and resolution of these structures. We further examine the functional implications of m6A and R-loop interaction in human diseases, with a particular emphasis on their roles in cancer.
Full article

Figure 1
Open AccessReview
Epigenetic Insights into Tuberous Sclerosis Complex, Von Hippel–Lindau Syndrome, and Ataxia–Telangiectasia
by
Gavriel Hadjigavriel, Christina Stylianides, Evangelos Axarloglou, Maria Eleni Manthou, Efstratios Vakirlis, Paschalis Theotokis, Soultana Meditskou and Iasonas Dermitzakis
Epigenomes 2025, 9(2), 20; https://doi.org/10.3390/epigenomes9020020 - 9 Jun 2025
Abstract
Neurocutaneous syndromes represent a clinically and genetically heterogeneous group of disorders, with tuberous sclerosis complex (TSC), von Hippel–Lindau syndrome (VHL), and ataxia–telangiectasia (A-T) exemplifying some of the most complex entities within this category. These syndromes have traditionally been considered monogenic disorders, caused by
[...] Read more.
Neurocutaneous syndromes represent a clinically and genetically heterogeneous group of disorders, with tuberous sclerosis complex (TSC), von Hippel–Lindau syndrome (VHL), and ataxia–telangiectasia (A-T) exemplifying some of the most complex entities within this category. These syndromes have traditionally been considered monogenic disorders, caused by germline mutations in tumor suppressor or regulatory genes. However, they exhibit a striking degree of phenotypic variability and divergent clinical trajectories that cannot be fully explained by their underlying genetic alterations alone. Increasingly, epigenetic regulatory mechanisms, such as DNA methylation, histone modifications, chromatin remodeling, and non-coding RNA (ncRNA) activity, are recognized as key modulators of gene expression, cellular differentiation, and tissue-specific function. Disruption of these mechanisms has been implicated in disease pathogenesis, tumorigenesis, and neurodegeneration associated with TSC, VHL, and A-T. Aberrant epigenetic profiles may underlie the observed variability in clinical outcomes, even among individuals with identical mutations. This review consolidates current evidence on the epigenetic landscape of these syndromes, elucidating how these modifications may influence disease behavior and contribute to incomplete genotype–phenotype correlations. By integrating epigenetic insights with known molecular pathways, a more nuanced understanding of disease biology emerges, with potential implications for diagnostic stratification, prognostic assessment, and therapeutic innovation.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessArticle
Association of Model-Predicted Epigenetic Age and Female Infertility
by
Elena Pozdysheva, Vitaly Korchagin, Tatiana Rumyantseva, Daria Ogneva, Vera Zhivotova, Irina Gaponova, Konstantin Mironov and Vasily Akimkin
Epigenomes 2025, 9(2), 19; https://doi.org/10.3390/epigenomes9020019 - 5 Jun 2025
Abstract
►▼
Show Figures
Background: To date, there are no precise clinical and laboratory methods to accurately predict the onset of fertility decline in women, with chronological age being the ultimate predictor. This has led to increased interest in developing methods to determine biological age, as it
[...] Read more.
Background: To date, there are no precise clinical and laboratory methods to accurately predict the onset of fertility decline in women, with chronological age being the ultimate predictor. This has led to increased interest in developing methods to determine biological age, as it provides a more accurate understanding of individual age-related physiological changes. Methods: In this study, we developed a model for estimating biological age based on DNA methylation levels in the ELOVL2, TRIM59, C1orf132, FHL2, and KLF14 genes using pyrosequencing. The model was tested in 64 Russian women, aged 25–39 years, to find an association between epigenetic age, infertility, low anti-Müllerian hormone (AMH) levels, and assisted reproductive technology (ART) failure. Results: The predictive performance of the model was evaluated. The mean absolute deviation of the model was 2.8 years; the mean absolute error was 2.6 years (R2 = 0.95). In the studied cohort, 33% of women exhibited epigenetic age acceleration (EAA), while 45% showed epigenetic age deceleration (EAD). All women with an EAA of ≥3 years (n = 6) had a history of infertility. Conclusions: In this study, no statistically significant associations were observed between EAA/EAD and AMH, body mass index, infertility, or ART failure in women.
Full article

Figure 1
Open AccessReview
DNA Methylation, Aging, and Cancer
by
Himani Vaidya, Jaroslav Jelinek and Jean-Pierre J. Issa
Epigenomes 2025, 9(2), 18; https://doi.org/10.3390/epigenomes9020018 - 3 Jun 2025
Abstract
►▼
Show Figures
Aging and cancer, though distinct biological processes, share overlapping molecular pathways, particularly in epigenetic regulation. Among these, DNA methylation is central to mediating gene expression, maintaining cellular identity, and regulating genome stability. This review explores how age-associated changes in DNA methylation, characterized by
[...] Read more.
Aging and cancer, though distinct biological processes, share overlapping molecular pathways, particularly in epigenetic regulation. Among these, DNA methylation is central to mediating gene expression, maintaining cellular identity, and regulating genome stability. This review explores how age-associated changes in DNA methylation, characterized by both global hypomethylation and focal hypermethylation, contribute to the emergence of cancer. We discuss mechanisms of DNA methylation drift, the development of epigenetic clocks, and the role of entropy and epigenetic mosaicism, in aging and tumorigenesis. Emphasis is placed on how stochastic methylation errors accumulate in aging cells and lead to epiallelic shifts and gene silencing, predisposing tissues to malignant transformation, even despite recently increased cancer incidences at younger ages. We also highlight the translational potential of DNA methylation-based biomarkers, and therapeutic targets, in age-related diseases. By framing cancer as a disease of accelerated epigenetic aging, this review offers a unifying perspective and calls for age-aware approaches to both basic research and clinical oncology.
Full article
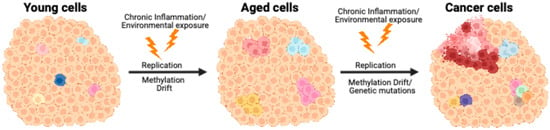
Figure 1
Open AccessOpinion
Epigenetic DNA Methylation Under the Influence of Low-Dose Ionizing Radiation, and Supplementation with Vitamin B12 and Folic Acid: Harmful or Beneficial for Professionals?
by
Borivoje Savic, Bozidar Savic and Svetlana Stanojlovic
Epigenomes 2025, 9(2), 17; https://doi.org/10.3390/epigenomes9020017 - 31 May 2025
Abstract
This review paper highlights the importance of educating current and future professionals about epigenetic mechanisms and recognizing epigenetics as a crucial model for protection against ionizing radiation. Two basic models for radiation-induced DNA damage are currently in use. The association between mutations and
[...] Read more.
This review paper highlights the importance of educating current and future professionals about epigenetic mechanisms and recognizing epigenetics as a crucial model for protection against ionizing radiation. Two basic models for radiation-induced DNA damage are currently in use. The association between mutations and chromosomal aberrations provides a framework for analyzing risks at low radiation doses and exposure to small doses. However, there is no monitoring of epigenetic changes in professionals exposed to low doses of ionizing radiation. Epigenetic events regulate gene activity and expression not only during cell development and differentiation but also in response to environmental stimuli, such as ionizing radiation. Furthermore, the potential occurrence of malignant and hereditary diseases at low doses of ionizing radiation is linearly correlated and is considered a scientifically accepted assumption, despite recognized scientific limitations associated with this assessment. The aim of this review is to integrate novel and intriguing radiobiological paradigms regarding the effects of ionizing radiation on DNA methylation and epigenetic regulation of the DNA molecule. Several hypothesized biological responses to ionizing radiation are examined, linking them to epigenetic mechanisms involved in health risk assessment for professionals. The second part of the review includes published research related to epigenetics, supplementation, and virus reactivation in the context of epigenetic modifications of the DNA molecule. We hypothesize that different cycles lead to changes in the epigenome, which may be associated with the reactivation of certain viruses and the deficiency of specific dietary elements. These findings are linked to minimal deficiencies in vitamin B12 and folic acid, which may contribute to epigenomic changes. This aspect is crucial for the immune status of individuals working in high-risk environments.
Full article
(This article belongs to the Special Issue Features Papers in Epigenomes 2025)
►▼
Show Figures

Figure 1
Open AccessReview
Genetics and Epigenetics of Chemoinduced Oral Mucositis in Paediatric Patients with Haematological Malignancies—A Review
by
Juliana Ramalho Guimarães, José Maria Chagas Viana Filho and Naila Francis Paulo de Oliveira
Epigenomes 2025, 9(2), 16; https://doi.org/10.3390/epigenomes9020016 - 30 May 2025
Abstract
Background: Oral mucositis (OM) is a painful inflammation resulting from chemotherapy. It is dependent on factors such as age, gender, chemotherapy regimen, oral health, immunological and nutritional status, and genetics. Objectives: The aim of the study was to conduct a narrative review to
[...] Read more.
Background: Oral mucositis (OM) is a painful inflammation resulting from chemotherapy. It is dependent on factors such as age, gender, chemotherapy regimen, oral health, immunological and nutritional status, and genetics. Objectives: The aim of the study was to conduct a narrative review to compile studies on the contribution of genetic and epigenetic aspects to the pathogenesis of OM in children with haematological malignancies undergoing chemotherapy treatment. Methods: The literature search was performed in Pubmed, Scopus, Web of Science, Cochrane, Lilacs, and grey literature databases covering articles published since 2010. Results: Twenty-two studies investigating polymorphisms and four studies investigating DNA methylation were included. Polymorphisms in the MTHFR, ABCB1, ABCC2, ABCG2, SLCO1B, miR-1206, miR-3683, CAT, and VDR genes were associated as risk factors for OM and polymorphisms in the TYMS and miR-4268 genes were associated as protective factors. With regard to DNA methylation, associations such as protection or susceptibility to OM have not yet been proven. However, studies have shown that DNMT1 methylation and hypomethylation in total DNA and in the TNF-α gene are associated with recovery of the oral mucosa. Conclusions: Genetic variants are associated with OM in various biological pathways, such as folate metabolism, transport proteins, epigenetic machinery, oxidative stress, and vitamin D metabolism. The DNA methylation profile, which is still poorly understood in the pathogenesis of OM, is associated with mucosal recovery (inflammation and epigenetic machinery). Genetic and epigenetic markers may be tools to indicate a patient’s susceptibility to developing OM, and epigenetic markers may be a target for therapies.
Full article
(This article belongs to the Special Issue Epigenetic Mechanisms of Hematologic Malignancies)
►▼
Show Figures
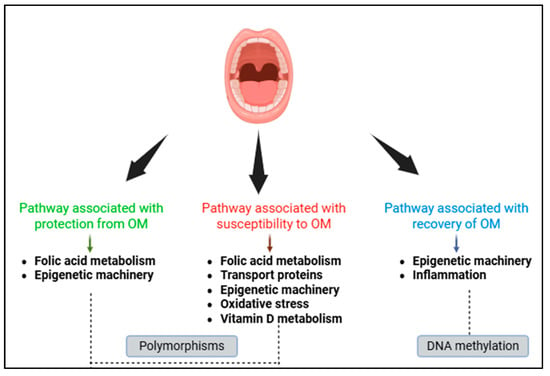
Figure 1
Open AccessArticle
Histone H3 Lysine 9 Acetylation Plays a Role in Adipogenesis of Periodontal Ligament-Derived Stem Cells
by
Julio A. Montero-Del-Toro, Angelica A. Serralta-Interian, Geovanny I. Nic-Can, Mónica Lamas, Rodrigo A. Rivera-Solís and Beatriz A. Rodas-Junco
Epigenomes 2025, 9(2), 15; https://doi.org/10.3390/epigenomes9020015 - 24 May 2025
Abstract
Background: The epigenetic regulation of adipogenic differentiation in dental stem cells (DSCs) remains poorly understood, as research has prioritized osteogenic differentiation for dental applications. However, elucidating these mechanisms could enable novel regenerative strategies for soft tissue engineering. Periodontal ligament stem cells (PDLSCs) exhibit
[...] Read more.
Background: The epigenetic regulation of adipogenic differentiation in dental stem cells (DSCs) remains poorly understood, as research has prioritized osteogenic differentiation for dental applications. However, elucidating these mechanisms could enable novel regenerative strategies for soft tissue engineering. Periodontal ligament stem cells (PDLSCs) exhibit notable adipogenic potential, possibly linked to histone 3 acetylation at lysine 9 (H3K9ac); however, the mechanistic role of this modification remains unclear. Methods: To address this gap, we investigated how histone deacetylase inhibitors (HDACis)—valproic acid (VPA, 8 mM) and trichostatin A (TSA, 100 nM)—modulate H3K9ac dynamics, adipogenic gene expression (C/EBPβ and PPARγ-2), and chromatin remodeling during PDLSCs differentiation. Techniques used included quantitative PCR (qPCR), lipid droplet analysis, and chromatin immunoprecipitation followed by qPCR (ChIP-qPCR). Results: TSA-treated cells exhibited increased lipid deposition with smaller lipid droplets compared to VPA-treated cells. Global H3K9ac levels correlated positively with adipogenic progression. VPA induced early upregulation of C/EBPβ and PPARγ-2 (day 7), whereas TSA triggered a delayed but stronger PPARγ-2 expression. ChIP-qPCR analysis revealed significant H3K9ac enrichment at the PPARγ-2 promoter in TSA-treated cells, indicating enhanced chromatin accessibility. Conclusions: These findings demonstrate that H3K9ac-mediated epigenetic remodeling plays a critical role in the adipogenic differentiation of PDLSCs and identifies TSA as a potential tool for modulating this process.
Full article
(This article belongs to the Collection Epigenetic Regulation of Cellular Differentiation)
►▼
Show Figures

Figure 1
Open AccessArticle
Discrimination, Coping, and DNAm Accelerated Aging Among African American Mothers of the InterGEN Study
by
Alexandria Nyembwe, Yihong Zhao, Billy A. Caceres, Daniel W. Belsky, Calen Patrick Ryan, Brittany Taylor, Morgan T. Morrison, Laura Prescott, Stephanie Potts-Thompson, Arezo Aziz, Fisola Aruleba, Erica Matute-Arcos, Olajide Williams, Cindy Crusto and Jacquelyn Y. Taylor
Epigenomes 2025, 9(2), 14; https://doi.org/10.3390/epigenomes9020014 - 4 May 2025
Abstract
Background: Racial discrimination experiences are associated with the activation of stress biology pathways and signs of accelerated biological aging, including alterations in DNA methylation (DNAm). Coping strategies may mitigate stress from racial discrimination and protect against long-term adverse health outcomes. Methods: We conducted
[...] Read more.
Background: Racial discrimination experiences are associated with the activation of stress biology pathways and signs of accelerated biological aging, including alterations in DNA methylation (DNAm). Coping strategies may mitigate stress from racial discrimination and protect against long-term adverse health outcomes. Methods: We conducted a secondary analysis of data from the Intergenerational Impact of Genetic and Psychological Factors on Blood Pressure cohort, an all-African-American sample, to test the hypothesis that social support can protect against accelerated biological aging associated with experiences of racial discrimination. We measured biological aging from saliva DNAm using six epigenetic clocks. Clock values were residualized on participant age and the estimated proportion of epithelial cells contributing to the DNA sample and standardized to M = 0, SD = 1 within the analysis sample. The primary analysis was focused on the second-generation PhenoAge and GrimAge clocks and the third-generation DunedinPACE “speedometer,” which previous studies have linked with racial discrimination. Results: In our sample (n = 234; mean age = 31.9 years; SD = 5.80), we found evidence consistent with our hypothesis in the case of the PhenoAge clock, but not the other clocks. Among mothers who did not seek social support, experiences of racial discrimination were associated with an older PhenoAge (b = 0.26, 95% CI = 0.02–0.50, p = 0.03). However, social-support seeking mitigated this risk; at the highest levels of social support, no adverse consequences of discrimination were observed (interaction b = −0.01, 95% CI = −0.02–−0.00, p = 0.03). Conclusions: The replication of results is needed. Future research should also investigate additional adaptive and maladaptive coping strategies utilized by African American women and mothers to identify protective measures that influence health outcomes.
Full article
(This article belongs to the Special Issue Features Papers in Epigenomes 2025)
►▼
Show Figures

Figure 1
Open AccessArticle
Arabidopsis thaliana Roots Exposed to Extracellular Self-DNA: Evidence of Epigenetic Effects
by
Alessia Ronchi, Guido Incerti, Emanuele De Paoli, Speranza Claudia Panico, Giovanni Luca Sciabbarrasi, Pasquale Termolino, Fabrizio Cartenì, Mariachiara Langella, Maria Luisa Chiusano and Stefano Mazzoleni
Epigenomes 2025, 9(2), 13; https://doi.org/10.3390/epigenomes9020013 - 30 Apr 2025
Abstract
Background: Previous evidence demonstrated DNA methylation changes in response to stress in plants, showing rapid changes within a limited time frame. Exposure to self-DNA inhibits seedling root elongation, and it was shown that it causes changes in CG DNA methylation in Lactuca sativa
[...] Read more.
Background: Previous evidence demonstrated DNA methylation changes in response to stress in plants, showing rapid changes within a limited time frame. Exposure to self-DNA inhibits seedling root elongation, and it was shown that it causes changes in CG DNA methylation in Lactuca sativa. We assessed cytosine methylation changes and associated gene expression patterns in roots of Arabidopsis thaliana Col-0 seedlings exposed to self-DNA for 6 and 24 h. Methods: We used whole genome bisulfite sequencing (WGBS) and RNA-seq analyses to assess genomic cytosine methylation and corresponding gene expression, respectively, on DNA and RNA extracted with commercial kits from roots exposed to self-DNA by an original setup. Fifteen hundred roots replicates, including the control in distilled water, were collected after exposure. Sequencing was performed on a NovaSeq 6000 platform and Ultralow Methyl-Seq System for RNA and DNA WGBS, respectively. Results: Gene expression in roots exposed to self-DNA differed from that of untreated controls, with a total of 305 genes differentially expressed and 87 ontologies enriched in at least one treatment vs. control comparison, and particularly after 24 h of exposure. DNA methylation, particularly in CHG and CHH contexts, was also different, with hyper- and hypomethylation prevailing in treatments vs. controls at 6 h and 24 h, respectively. Differentially expressed genes (DEGs) analysis, Gene Ontology (GO) enrichment analysis, and differentially methylated regions (DMRs) analysis, provided an integrated understanding of the changes associated with self-DNA exposure. Our results suggest differential gene expression associated with DNA methylation in response to self-DNA exposure in A. thaliana roots, enhanced after prolonged exposure. Conclusions: Main functional indications of association between DNA methylation and gene expression involved hypomethylation and downregulation of genes related to nucleotide/nucleoside metabolism (ATP synthase subunit) and cell wall structure (XyG synthase), consistent with previous observations from metabolomics and physiological studies. Further confirmation of these findings will contribute to improving our understanding of the plant molecular response to self-DNA and its implications in stress responses.
Full article
(This article belongs to the Special Issue Features Papers in Epigenomes 2025)
►▼
Show Figures

Figure 1
Open AccessArticle
The Effect of Clinical Factors on the Reversion of Cg05575921 Methylation in Smoking Cessation
by
Robert Philibert, Steven R. H. Beach, Michelle R. vanDellen, James A. Mills and Jeffrey D. Long
Epigenomes 2025, 9(2), 12; https://doi.org/10.3390/epigenomes9020012 - 28 Apr 2025
Abstract
Background: Financial Incentive Treatments (FIT) can be effective in the treatment of smoking. However, weaknesses in current biochemical approaches for assessing smoking cessation may hinder its implementation, particularly for management of long-term smoking cessation. The use of cg05575921 methylation assessments could address some
[...] Read more.
Background: Financial Incentive Treatments (FIT) can be effective in the treatment of smoking. However, weaknesses in current biochemical approaches for assessing smoking cessation may hinder its implementation, particularly for management of long-term smoking cessation. The use of cg05575921 methylation assessments could address some of the shortcomings of current self-report and non-self-report methods, but additional information is needed about the speed of methylation reversion as a function of key clinical and demographic variables. Methods: To better understand those relationships, we analyzed data from 3040 subjects from the National Lung Screening Trial (NLST), including 1552 self-reported quitters. Results: Plotting of the data as a function of time since quitting shows that methylation increases approximately 14%, on average, after at least one full year of cessation with a subsequent slow non-linear increase in methylation over the next 14 years. Least Squares Regression modeling shows strong effects of quit time and a modest, yet significant, effect of body mass index (BMI) on the rate of reversion. Prior cigarette consumption characteristics and sex made modest contributions as well, with the latter largely offset by pre-cessation methylation levels. Race and age were not significant factors in the models. Conclusions: When combined with data from prior studies, these analyses of the long-term reversion of cg05575921 methylation will be informative to those considering FIT approaches to incentivizing reversion of cg05575921 as an index of short- and long-term smoking cessation.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessReview
Induction of DNA Demethylation: Strategies and Consequences
by
Pietro Salvatore Carollo and Viviana Barra
Epigenomes 2025, 9(2), 11; https://doi.org/10.3390/epigenomes9020011 - 12 Apr 2025
Abstract
DNA methylation is an important epigenetic modification with a plethora of effects on cells, ranging from the regulation of gene transcription to shaping chromatin structure. Notably, DNA methylation occurs thanks to the activity of DNA methyltransferases (DNMTs), which covalently add a methyl group
[...] Read more.
DNA methylation is an important epigenetic modification with a plethora of effects on cells, ranging from the regulation of gene transcription to shaping chromatin structure. Notably, DNA methylation occurs thanks to the activity of DNA methyltransferases (DNMTs), which covalently add a methyl group to the cytosine in position 5′ in CpG dinucleotides. Different strategies have been developed to study the effects of DNA methylation in cells, involving either DNMTs inhibition (passive DNA demethylation) or the use of Ten-eleven translocation protein (TET) family enzymes, which directly demethylate DNA (active DNA demethylation). In this manuscript, we will briefly cover the most commonly used strategies in the last two decades to achieve DNA demethylation, along with their effects on cells. We will also discuss some of the newest inducible ways to inhibit DNMTs without remarkable side effects, as well as the effect of non-coding RNAs on DNA methylation. Lastly, we will briefly examine the use of DNA methylation inhibition in biomedical research.
Full article
(This article belongs to the Special Issue Features Papers in Epigenomes 2025)
►▼
Show Figures

Graphical abstract
Open AccessReview
The Good, the Bad, and the Epigenetic: Stress-Induced Metabolite Regulation and Transgenerational Effects
by
Saida Ibragić, Sabina Dahija and Erna Karalija
Epigenomes 2025, 9(2), 10; https://doi.org/10.3390/epigenomes9020010 - 29 Mar 2025
Cited by 4
Abstract
Background: Plants face a wide range of environmental stresses that disrupt growth and productivity. To survive and adapt, they undergo complex metabolic reprogramming by redirecting carbon and nitrogen fluxes toward the biosynthesis of protective secondary metabolites such as phenylpropanoids, flavonoids, and lignin. Recent
[...] Read more.
Background: Plants face a wide range of environmental stresses that disrupt growth and productivity. To survive and adapt, they undergo complex metabolic reprogramming by redirecting carbon and nitrogen fluxes toward the biosynthesis of protective secondary metabolites such as phenylpropanoids, flavonoids, and lignin. Recent research has revealed that these stress-induced metabolic processes are tightly regulated by epigenetic mechanisms, including DNA methylation, histone modifications, chromatin remodeling, and non-coding RNAs. Methods: This review synthesizes current findings from studies on both model and crop plants, examining the roles of key epigenetic regulators in controlling secondary metabolism under stress. Special focus is placed on dynamic changes in DNA methylation, histone acetylation, and the action of small RNAs such as siRNAs and miRNAs in transcriptional and post-transcriptional regulation. Results: Evidence indicates that stress triggers rapid and reversible epigenetic modifications that modulate gene expression linked to secondary metabolic pathways. These modifications not only facilitate immediate metabolic responses but can also contribute to stress memory. In some cases, this memory is retained and transmitted to the next generation, influencing progeny stress responses. However, critical knowledge gaps remain, particularly concerning the temporal dynamics, tissue specificity, and long-term stability of these epigenetic marks in crops. Conclusions: Understanding how epigenetic regulation governs secondary metabolite production offers promising avenues to enhance crop resilience and productivity in the context of climate change. Future research should prioritize dissecting the stability and heritability of these modifications to support the development of epigenetically informed breeding strategies.
Full article
(This article belongs to the Collection Epigenetic Control in Plants)
►▼
Show Figures

Figure 1
Open AccessArticle
The Association of Childhood Allergic Diseases with Prenatal Exposure to Pollen Grains Through At-Birth DNA Methylation
by
Rajesh Melaram, Hongmei Zhang, James Adefisoye and Hasan Arshad
Epigenomes 2025, 9(1), 9; https://doi.org/10.3390/epigenomes9010009 - 11 Mar 2025
Abstract
Background: Pollen exposure in early life is shown to be associated with allergy and asthma. DNA methylation (DNAm), an epigenetic marker, potentially reacts to pollen. However, the role of at-birth DNAm between prenatal pollen grain (PPG) exposure and childhood asthma and allergic rhinitis
[...] Read more.
Background: Pollen exposure in early life is shown to be associated with allergy and asthma. DNA methylation (DNAm), an epigenetic marker, potentially reacts to pollen. However, the role of at-birth DNAm between prenatal pollen grain (PPG) exposure and childhood asthma and allergic rhinitis is unknown. Methods: Data in a birth cohort study on the Isle of Wight, UK, were analyzed (n = 236). Newborn DNAm was measured in cord blood or blood spots on Guthrie cards and screened for potential association with PPG exposure using the R package ttScreening. CpGs that passed screening were further assessed for such associations via linear regressions with adjusting covariates included. Finally, DNAm at PPG-associated CpGs were evaluated for their association with asthma and allergic rhinitis using logistic regressions, adjusting for covariates. The impact of cell heterogeneity on the findings was assessed. Statistical significance was set at p < 0.05. Results: In total, 42 CpGs passed screening, with 41 remaining statistically significant after adjusting for covariates and cell types (p < 0.05). High PPG exposure was associated with lower DNAm at cg12318501 (ZNF99, β = −0.029, p = 0.032) and cg00929606 (ADM2, β = −0.023, p = 0.008), which subsequently was associated with decreased odds of asthma (OR = 0.11, 95% CI 0.02–0.53, p = 0.006; OR = 0.14, 95% CI 0.02–1.00, p = 0.049). For rhinitis, cg15790214 (HCG11) was shown to play such a role as a mediator (β = −0.027, p ≤ 0.0001; OR = 0.22, 95% CI 0.07–0.72, p = 0.01). Conclusions: The association of PPG exposure with childhood asthma and allergic rhinitis incidence is potentially mediated by DNAm at birth.
Full article
(This article belongs to the Special Issue Longitudinal Studies on Prenatal or Early Life Exposures, Their Associated Epigenetics, and Population Health)
►▼
Show Figures
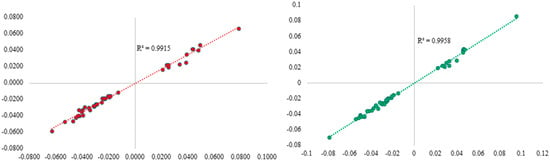
Figure 1
Open AccessArticle
Investigating Single-Molecule Molecular Inversion Probes for Medium-Scale Targeted DNA Methylation Analysis
by
Roy B. Simons, Hieab H. H. Adams, Manfred Kayser and Athina Vidaki
Epigenomes 2025, 9(1), 8; https://doi.org/10.3390/epigenomes9010008 - 2 Mar 2025
Abstract
Background: Epigenetic biomarkers, particularly CpG methylation, are increasingly employed in clinical and forensic settings. However, we still lack a cost-effective, sensitive, medium-scale method for the analysis of hundreds to thousands of user-defined CpGs suitable for minute DNA input amounts (<10 ng). In this
[...] Read more.
Background: Epigenetic biomarkers, particularly CpG methylation, are increasingly employed in clinical and forensic settings. However, we still lack a cost-effective, sensitive, medium-scale method for the analysis of hundreds to thousands of user-defined CpGs suitable for minute DNA input amounts (<10 ng). In this study, motivated by promising results in the genetics field, we investigated single-molecule molecular inversion probes (smMIPs) for simultaneous analysis of hundreds of CpGs by using an example set of 514 age-associated CpGs (Zhang model). Methods: First, we developed a novel smMIP design tool to suit bisulfite-converted DNA (Locksmith). Then, to optimize the capture process, we performed single-probe capture for ten selected, representative smMIPs. Based on this pilot, the full smMIP panel was tested under varying capture conditions, including hybridization and elongation temperature, smMIP and template DNA amounts, dNTP concentration and elongation time. Results: Overall, we found that the capture efficiency was highly probe-(and hence, sequence-) dependent, with a heterogeneous coverage distribution across CpGs higher than the 1000-fold range. Considering CpGs with at least 20X coverage, we yielded robust methylation detection with levels comparable to those obtained from the gold standard EPIC microarray analysis (Pearsons’s r: 0.96). Conclusions: The observed low specificity and uniformity indicate that smMIPs in their current form are not compatible with the lowered complexity of bisulfite-converted DNA.
Full article
(This article belongs to the Special Issue A Commemorative Issue in Honor of the 30th Anniversary of the Epigenetics Society)
►▼
Show Figures

Figure 1
Open AccessArticle
Isothiocyanates Enhance the Anti-Melanoma Effect of Zebularine Through Modulation of Apoptosis and Regulation of DNMTs’ Expression, Chromatin Configuration and Histone Posttranslational Modifications Associated with Altered Gene Expression Patterns
by
Ioannis Anestopoulos, Ioannis Paraskevaidis, Sotiris Kyriakou, Louiza Potamiti, Dimitrios T. Trafalis, Sotiris Botaitis, Rodrigo Franco, Aglaia Pappa and Mihalis I. Panayiotidis
Epigenomes 2025, 9(1), 7; https://doi.org/10.3390/epigenomes9010007 - 25 Feb 2025
Abstract
Background: In the present study, we aimed to characterize the cytotoxic efficacy of Zebularine either as a single agent or in combination with various isothiocyanates in an in vitro model consisting of human melanoma (A375, Colo-679) as well as non-tumorigenic immortalized keratinocyte (HaCaT)
[...] Read more.
Background: In the present study, we aimed to characterize the cytotoxic efficacy of Zebularine either as a single agent or in combination with various isothiocyanates in an in vitro model consisting of human melanoma (A375, Colo-679) as well as non-tumorigenic immortalized keratinocyte (HaCaT) cells. Methods: In this model, we have evaluated the anti-melanoma effect of Zebularine (in single and combinatorial protocols) in terms of cell viability, apoptotic induction and alterations in ultrastructural chromatin configuration, protein expression levels of DNA methyltransferases (DNMTs) and associated histone epigenetic marks capable of mediating gene expression. Results: Exposure to Zebularine resulted in dose- and time-dependent cytotoxicity through apoptotic induction in malignant melanoma cells, while neighboring non-tumorigenic keratinocytes remained unaffected. A more profound response was observed in combinational protocols, as evidenced by a further decline in cell viability leading to an even more robust apoptotic induction followed by a differential response (i.e., activation/de-activation) of various apoptotic genes. Furthermore, combined exposure protocols caused a significant decrease of DNMT1, DNMT3A and DNMT3B protein expression levels together with alterations in ultrastructural chromatin configuration and protein expression levels of specific histone modification marks capable of modulating gene expression. Conclusions: Overall, we have developed a novel experimental approach capable of potentiating the cytotoxic efficacy of Zebularine against human malignant melanoma cells while at the same time maintaining a non-cytotoxic profile against neighboring non-tumorigenic keratinocyte (HaCaT) cells.
Full article
(This article belongs to the Collection Epigenetics of Melanoma)
►▼
Show Figures

Figure 1
Open AccessArticle
Novel Epigenetics Control (EpC) Nanocarrier for Cancer Therapy Through Dual-Targeting Approach to DNA Methyltransferase and Ten-Eleven Translocation Enzymes
by
Risa Mitsuhashi, Kiyoshi Sato and Hiroyoshi Kawakami
Epigenomes 2025, 9(1), 6; https://doi.org/10.3390/epigenomes9010006 - 11 Feb 2025
Cited by 1
Abstract
Background/Objectives: Aberrant hypermethylation in the promoter regions of tumor suppressor genes facilitates the pathogenesis and progression of cancer. Therefore, inhibitors targeting DNA methyltransferase (DNMT) have been tested in clinical studies. However, the current monotherapy of DNMT inhibitors shows limited efficacy. Furthermore, the mechanism
[...] Read more.
Background/Objectives: Aberrant hypermethylation in the promoter regions of tumor suppressor genes facilitates the pathogenesis and progression of cancer. Therefore, inhibitors targeting DNA methyltransferase (DNMT) have been tested in clinical studies. However, the current monotherapy of DNMT inhibitors shows limited efficacy. Furthermore, the mechanism of action of DNMT inhibitors is DNA replication-dependent. To address these limitations, we developed a novel core–shell-type “epigenetics control (EpC) nanocarrier” that encapsulated decitabine (5-aza-dC) in the PLGA core nanoparticle and hybridized TET1 gene-encoding pDNA on the lipid shell surface. This study aimed to evaluate whether the dual delivery of DNMT inhibitors and pDNA of TET1 could synergistically enhance tumor suppressor gene expression and induce cell cycle arrest and/or apoptosis in cancer cells. Herein, we demonstrate the potential of the EpC carrier in HCT116 human colon cancer cells to upregulate tumor suppressor gene expression and rapidly achieve cell cycle arrest. Methods: PLGA core nanoparticles were prepared by the W/O/W double emulsion method. The formation of core–shell nanoparticles and complexation with pDNA were investigated and optimized by dynamic light scattering, zeta potential measurement, and agarose gel electrophoresis. The cellular uptake and transfection efficiency were measured by confocal laser scanning microscopy and a luciferase assay, respectively. The expression of p53 protein was detected by Western blotting. The anti-tumor effects of the EpC nanocarrier were evaluated by cell cycle analysis and an apoptosis assay. Results: The EpC nanocarrier delivered the DNMT inhibitor and TET gene-encoding pDNA into HCT116 cells. It promoted the expression of the tumor suppressor protein p53 and induced rapid cell cycle arrest in the G2/M phase in HCT116 cells. Conclusions: Our findings suggest that the dual-targeting of DNMT and TET enzymes effectively repairs aberrant DNA methylation and induces growth arrest in cancer cells, and the dual-targeting strategy may contribute to the advancement of epigenetic cancer therapy.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessReview
Epigenomic Echoes—Decoding Genomic and Epigenetic Instability to Distinguish Lung Cancer Types and Predict Relapse
by
Alexandra A. Baumann, Zholdas Buribayev, Olaf Wolkenhauer, Amankeldi A. Salybekov and Markus Wolfien
Epigenomes 2025, 9(1), 5; https://doi.org/10.3390/epigenomes9010005 - 5 Feb 2025
Abstract
Genomic and epigenomic instability are defining features of cancer, driving tumor progression, heterogeneity, and therapeutic resistance. Central to this process are epigenetic echoes, persistent and dynamic modifications in DNA methylation, histone modifications, non-coding RNA regulation, and chromatin remodeling that mirror underlying genomic chaos
[...] Read more.
Genomic and epigenomic instability are defining features of cancer, driving tumor progression, heterogeneity, and therapeutic resistance. Central to this process are epigenetic echoes, persistent and dynamic modifications in DNA methylation, histone modifications, non-coding RNA regulation, and chromatin remodeling that mirror underlying genomic chaos and actively influence cancer cell behavior. This review delves into the complex relationship between genomic instability and these epigenetic echoes, illustrating how they collectively shape the cancer genome, affect DNA repair mechanisms, and contribute to tumor evolution. However, the dynamic, context-dependent nature of epigenetic changes presents scientific and ethical challenges, particularly concerning privacy and clinical applicability. Focusing on lung cancer, we examine how specific epigenetic patterns function as biomarkers for distinguishing cancer subtypes and monitoring disease progression and relapse.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessArticle
Single-Molecule Nanopore Sequencing of the CpG Island from the Promoter of O6-Methylguanine-DNA Methyltransferase Provides Insights into the Mechanism of De Novo Methylation of G/C-Rich Regions
by
Alexander V. Sergeev, Daniil P. Malyshev, Adelya I. Genatullina, Galina V. Pavlova, Elizaveta S. Gromova and Maria I. Zvereva
Epigenomes 2025, 9(1), 4; https://doi.org/10.3390/epigenomes9010004 - 26 Jan 2025
Cited by 1
Abstract
►▼
Show Figures
Background: The methylation of cytosine residues at CpG sites within the O6-methylguanine-DNA methyltransferase (MGMT) promoter is a key biomarker in glioblastoma therapy. The MGMT promoter (MGMTp) contains multiple guanine-rich sequences capable of folding into G-quadruplexes (G4s), but their relevance for MGMTp
[...] Read more.
Background: The methylation of cytosine residues at CpG sites within the O6-methylguanine-DNA methyltransferase (MGMT) promoter is a key biomarker in glioblastoma therapy. The MGMT promoter (MGMTp) contains multiple guanine-rich sequences capable of folding into G-quadruplexes (G4s), but their relevance for MGMTp methylation is poorly understood. Objectives: Our study explores the impact of potential G-quadruplex-forming sequences (PQS) in the MGMT promoter CpG island on the activity of de novo DNA methyltransferase Dnmt3a. Additionally, we investigate their influence on the accuracy of methylation pattern detection using nanopore sequencing. Methods: Nanopore sequencing was employed to analyze the methylation of 94 clinically significant CpG sites in the human MGMTp using an in vitro de novo methylation system. Circular dichroism spectroscopy was used to identify G4 structures within the MGMTp CpG island. Interactions between the catalytic domain of Dnmt3a and the PQS from the MGMTp were examined by biolayer interferometry. Results: Guanine-rich DNA strands of the PQSs in the MGMTp were hypomethylated, while the complementary cytosine-rich strands were methylated by DNA methyltransferase Dnmt3a with higher efficiency. The accuracy of detecting modified bases in the PQS was significantly lower compared to surrounding sequences. Single-stranded guanine-rich DNA sequences from the MGMTp exhibited strong binding to Dnmt3a-CD, with an affinity approximately 10 times higher than their cytosine-rich complements (Kd = 3 × 10−8 M and 3 × 10−7 M, respectively). By binding to Dnmt3a, G4-forming oligonucleotides from MGMTp effectively inhibited the methylation reaction (IC50 6 × 10−7 M). Conclusions: The obtained data indicate the role of PQSs in establishing de novo methylation of the MGMT promoter. They also highlight the challenges of sequencing guanine-rich regions and the impact of specific de novo methylation patterns on clinical data interpretation.
Full article

Figure 1
Open AccessReview
Epigenetics in Skin Homeostasis and Ageing
by
Iasonas Dermitzakis, Stella Aikaterini Kyriakoudi, Sofia Chatzianagnosti, Despoina Chatzi, Efstratios Vakirlis, Soultana Meditskou, Maria Eleni Manthou and Paschalis Theotokis
Epigenomes 2025, 9(1), 3; https://doi.org/10.3390/epigenomes9010003 - 9 Jan 2025
Cited by 1
Abstract
►▼
Show Figures
The skin, the largest organ of the human body, plays numerous essential roles, including protection against environmental hazards and the regulation of body temperature. The processes of skin homeostasis and ageing are complex and influenced by many factors, with epigenetic mechanisms being particularly
[...] Read more.
The skin, the largest organ of the human body, plays numerous essential roles, including protection against environmental hazards and the regulation of body temperature. The processes of skin homeostasis and ageing are complex and influenced by many factors, with epigenetic mechanisms being particularly significant. Epigenetics refers to the regulation of gene expression without altering the underlying DNA sequence. The dynamic nature of the skin, characterized by constant cellular turnover and responsiveness to environmental stimuli, requires precise gene activity control. This control is largely mediated by epigenetic modifications such as DNA methylation, histone modification, and regulation by non-coding RNAs. The present review endeavours to provide a comprehensive exploration and elucidation of the role of epigenetic mechanisms in regulating skin homeostasis and ageing. By integrating our current knowledge of epigenetic modifications with the latest advancements in dermatological research, we can gain a deeper comprehension of the complex regulatory networks that govern skin biology. Understanding these mechanisms also presents promising avenues for therapeutic interventions aimed at improving skin health and mitigating age-related skin conditions.
Full article

Figure 1

Journal Menu
► ▼ Journal Menu-
- Epigenomes Home
- Aims & Scope
- Editorial Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Topical Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Brain Sciences, CIMB, Epigenomes, Genes, IJMS, DNA
Genetics and Epigenetics of Substance Use Disorders
Topic Editors: Aleksandra Suchanecka, Anna Maria Grzywacz, Kszysztof ChmielowiecDeadline: 15 November 2025

Conferences
Special Issues
Special Issue in
Epigenomes
The Interplay Between Epigenetics and Inflammation in Chronic Diseases
Guest Editor: Baihai JiaoDeadline: 31 August 2025
Special Issue in
Epigenomes
Longitudinal Studies on Prenatal or Early Life Exposures, Their Associated Epigenetics, and Population Health
Guest Editors: Hongmei Zhang, Yu (Joyce) JiangDeadline: 31 August 2025
Special Issue in
Epigenomes
Features Papers in Epigenomes 2025
Guest Editors: Ivana De la Serna, Che-Kun James ShenDeadline: 31 December 2025
Special Issue in
Epigenomes
DNA Methylation Markers in Health and Disease
Guest Editor: Osman El‐MaarriDeadline: 31 May 2026
Topical Collections
Topical Collection in
Epigenomes
Epigenetic Mechanisms in Diabetes Research
Collection Editor: Assam El-Osta
Topical Collection in
Epigenomes
Epigenetic Regulation of Cellular Differentiation
Collection Editor: Wiesława Leśniak






