- Article
Epigenome-Wide DNA Methylation Levels During Pregnancy: Associations with Parity Across Diverse Populations
- Su Chen,
- Yunsung Lee and
- Susan L. Ewart
- + 9 authors
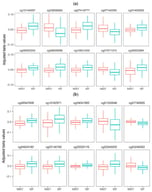
Background/Objectives: Parity, the number of times a woman carries a pregnancy to viability, has been linked to long-term maternal health outcomes. The mechanisms linking parity to health outcomes are poorly understood but may reflect influences of pregnancy on the maternal epigenome. Methods: This study examines the relationship between parity and DNA methylation (DNAm) during pregnancy using data from three cohorts: the Norwegian Mother, Father and Child Cohort Study (MoBa), the Atlanta African American Maternal-Child (AAAMC) cohort, and the Isle of Wight (IOW) Birth Cohort. Results: An epigenome-wide association study (EWAS) in MoBa identified 5374 cytosine–phosphate–guanine sites (CpGs) that were statistically significantly associated with parity, of which 69% were positively and 31% negatively correlated. Replication analyses confirmed 3491 CpGs in at least one cohort, and 93 CpGs in both AAAMC and IOW. Gene enrichment analysis revealed significant involvement of developmental and signaling pathways, including calcium signaling and neuroactive ligand–receptor interaction. Additionally, 584 differentially methylated regions (DMRs) were detected, with 90% overlapping individual parity-related CpGs. Conclusions: These findings suggest that parity influences epigenetic patterns, potentially affecting biological processes and molecular functions relevant to maternal health later in life.
5 May 2026





![HA-related gene and protein expression across species. Expression of hyaluronan synthase (HAS) and hyaluronidase (HYAL) isoform RNA in human (A), pig (B) and mouse (C) brains by region and expressed as normalised transcripts per million (nTPM) according to the Human Protein Atlas (HPA) database Version: 25.0, updated 11 November 2025, accessed 29 January 2026 [54]. Abbreviations: cerebral cortex (CC), olfactory bulb (OB), hippocampal formation (HPC), amygdala (AMY), basal ganglia (BG), thalamus (THA), hypothalamus (HYP), midbrain (MB), cerebellum (CER), medulla oblongata (MO), spinal cord (SC), white matter (WM), choroid plexus (CP). (D) Characterisation of protein expression of HA-related enzymes in human brain samples from HPA confirmed the expression of HAS1 and HYAL2; however, expression of HAS2 and HAS3 was not detected by this resource, despite evidence of brain expression from the literature. Expression of HYAL1, HYAL3 and HYAL4 was not explored as part of the HPA project so no data on the expression of these enzymes in the HPA database exists.](https://mdpi-res.com/cdn-cgi/image/width=281%2Cheight=192/https://mdpi-res.com/epigenomes/epigenomes-10-00028/article_deploy/html/images/epigenomes-10-00028-g001-550.jpg)