Journal Description
Biology
Biology
is an international, peer-reviewed, open access journal of biological sciences published semimonthly online by MDPI. The Spanish Society for Nitrogen Fixation (SEFIN) and Federation of European Laboratory Animal Science Associations (FELASA) are affiliated with Biology and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Biology) / CiteScore - Q1 (General Agricultural and Biological Sciences)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.8 days after submission; acceptance to publication is undertaken in 2.9 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.5 (2024);
5-Year Impact Factor:
4.0 (2024)
Latest Articles
Phytoremediation Potential of the Invasive Plant Datura stramonium (Solanaceae) for Toxic Metal Removal from Soil in the Qinghai–Tibet Plateau
Biology 2026, 15(10), 807; https://doi.org/10.3390/biology15100807 (registering DOI) - 19 May 2026
Abstract
The invasive plant Datura stramonium L. possesses strong reproductive capacity and ecological adaptability, showing a tendency to spread rapidly, especially in highly human-disturbed habitats. To explore its resource utilization pathway—turning waste into wealth—and to address toxic metal pollution in strongly human-disturbed areas (such
[...] Read more.
The invasive plant Datura stramonium L. possesses strong reproductive capacity and ecological adaptability, showing a tendency to spread rapidly, especially in highly human-disturbed habitats. To explore its resource utilization pathway—turning waste into wealth—and to address toxic metal pollution in strongly human-disturbed areas (such as mining regions), this study evaluates its phytoremediation potential in contaminated soils on the Qinghai–Tibet Plateau. We established a non-planted control and three planting density treatments to compare the removal rates of Pb, Cd, Cr, and As. To our knowledge, this is the first study to assess how planting density influences the multi-metal phytoremediation performance of this invasive species in a high-altitude plateau environment. The results showed that planting significantly increased toxic metal removal rates, with overall efficiency generally improving at higher densities, particularly for Cr. Analysis of bioconcentration and translocation factors revealed distinct element-specific accumulation patterns. Pb and As were primarily enriched and retained in the roots. Interestingly, while Cd exhibited a strong localized tendency to accumulate in the leaves, its overall root-to-shoot translocation remained relatively restricted at the whole-plant level, similar to Cr. Overall, D. stramonium functions primarily through root stabilization for Pb, As, and Cr, alongside partial aboveground accumulation for Cd. However, given its toxic and invasive nature, any practical phytoremediation application requires strict post-harvest biomass management and ecological monitoring to prevent secondary spread.
Full article
(This article belongs to the Section Ecology)
Open AccessArticle
Immune Evasion in Prostate Cancer: Resolving the Cold Tumour Paradox via a Hybrid Discrete–Continuum Computational Framework
by
Andile Kenneth Ntlokwana, Edinah Mudimu and Monde McMillan Ntwasa
Biology 2026, 15(10), 806; https://doi.org/10.3390/biology15100806 (registering DOI) - 19 May 2026
Abstract
Prostate cancer (PCa) is immunologically “cold” and resistant to immune checkpoint blockade (ICB), yet bulk analyses show low, non-prognostic PD-L1 expression. We hypothesised that this paradox reflects two overlooked dimensions: basal heterogeneity (static engine) and IFN-
Prostate cancer (PCa) is immunologically “cold” and resistant to immune checkpoint blockade (ICB), yet bulk analyses show low, non-prognostic PD-L1 expression. We hypothesised that this paradox reflects two overlooked dimensions: basal heterogeneity (static engine) and IFN-
(This article belongs to the Section Bioinformatics)
Open AccessReview
Cellular Stress and Immune Activation in Celiac Disease: Is the Chaperone System a Key Player?
by
Giuseppe Vergilio, Giusy Vultaggio, Rosalia Gagliardo, Letizia Paladino and Francesca Rappa
Biology 2026, 15(10), 805; https://doi.org/10.3390/biology15100805 (registering DOI) - 19 May 2026
Abstract
Celiac disease (CD) is a chronic immune-mediated enteropathy triggered by the ingestion of gluten in genetically predisposed individuals. While the adaptive immune response to deamidated gliadin peptides represents a central pathogenic mechanism, growing evidence suggests that epithelial stress and innate immune activation play
[...] Read more.
Celiac disease (CD) is a chronic immune-mediated enteropathy triggered by the ingestion of gluten in genetically predisposed individuals. While the adaptive immune response to deamidated gliadin peptides represents a central pathogenic mechanism, growing evidence suggests that epithelial stress and innate immune activation play a fundamental role in the onset and persistence of the disease. Heat shock proteins (Hsps), central regulators of cellular proteostasis, have emerged as potential mediators at the interface between epithelial distress and immune signaling. This review discusses the involvement of major Hsp families, including Hsp27, Hsp60, Hsp70, and Hsp90, in the pathophysiology of CD. The altered expression of Hsp27 and Hsp70 in the intestinal mucosa reflects a persistent state of epithelial stress that often persists despite a strict gluten-free diet (GFD). We focus specifically on Hsp60, whose extracellular release under stress conditions may allow it to function as a damage-associated molecular pattern (DAMP), engaging Toll-like receptors and promoting NF-κB- and inflammasome-dependent inflammatory pathways. Although direct mechanistic evidence linking Hsp60 to CD remains limited, the convergence of epithelial stress signs, Toll-like receptor (TLR) upregulation, and prolonged innate immune activation supports the hypothesis of a stress-induced inflammatory amplification circuit in the coeliac mucosa. Further studies are essential to clarify the pathogenic relevance and potential therapeutic implications of this proposed axis.
Full article
(This article belongs to the Special Issue Advances in Immunomodulation for Inflammatory Diseases)
►▼
Show Figures

Graphical abstract
Open AccessReview
Synergistic Integration of Enzyme and Microbial Platforms for Sustainable Management of Pharmaceutical Pollutants: Towards a Greener Pharmaceutical Lifecycle
by
Zhongshan Sun, Peitao Chen, Xiangyang Ge, Weiguo Zhang and Huanmin Liu
Biology 2026, 15(10), 804; https://doi.org/10.3390/biology15100804 (registering DOI) - 19 May 2026
Abstract
Purpose: This review aims to provide a theoretical basis and scientific reference for constructing environmentally friendly and economically feasible sustainable management systems for pharmaceutical pollution. Methods: This review discusses three synergistic mechanisms—“cascade degradation”, “symbiotic protection”, and “functional complementarity”—along with construction strategies including co-immobilization
[...] Read more.
Purpose: This review aims to provide a theoretical basis and scientific reference for constructing environmentally friendly and economically feasible sustainable management systems for pharmaceutical pollution. Methods: This review discusses three synergistic mechanisms—“cascade degradation”, “symbiotic protection”, and “functional complementarity”—along with construction strategies including co-immobilization technology, engineered biofilms, and engineered bacteria modified via synthetic biology. Result: Synergistic platforms have achieved significant progress in treating various types of pharmaceutical pollutants, including antibiotics, anti-inflammatories and hormones, antiviral drugs and pesticides. Conclusions: The synergistic integration of enzymes and microorganisms achieves the unification of efficient catalysis and deep mineralization, opening up a new pathway for the remediation of pharmaceutical pollution. It also transforms theoretically existing concepts into operable treatment technologies.
Full article
(This article belongs to the Section Biotechnology)
►▼
Show Figures

Figure 1
Open AccessArticle
Histological Evaluation of Radiotherapy-Induced Changes in Periodontal Tissues in a Rat Model of Experimental Periodontitis
by
Batuhan Hazar Ayşeşek, Buse Başak Feyizoğlu, Vakur Olgaç, İlknur Bingül, Nazlı Ayşeşek, Ülkü Başer and Ayşen Gülden Işık
Biology 2026, 15(10), 803; https://doi.org/10.3390/biology15100803 (registering DOI) - 19 May 2026
Abstract
Background/Objectives: Radiotherapy is a cornerstone of head and neck cancer treatment, but it can adversely affect periodontal tissues by impairing vascularity, cellularity, and healing capacity. The present study aimed to histologically and immunohistochemically evaluate radiotherapy-induced changes in periodontal tissues in a rat model
[...] Read more.
Background/Objectives: Radiotherapy is a cornerstone of head and neck cancer treatment, but it can adversely affect periodontal tissues by impairing vascularity, cellularity, and healing capacity. The present study aimed to histologically and immunohistochemically evaluate radiotherapy-induced changes in periodontal tissues in a rat model of experimental periodontitis, with particular focus on periodontal ligament width (PerioW) and the RANK/RANKL/OPG axis at different healing stages. Methods: Seventy-two male Sprague–Dawley rats were allocated to three groups: irradiation only (Rt), ligature-induced periodontitis only (Pt), and ligature-induced periodontitis followed by irradiation (PtRt). Experimental periodontitis was induced by placing ligatures around the maxillary first molars for two weeks. On the day of ligature removal, the irradiated groups received a single 20 Gy dose to the head and neck region. Animals were euthanized on days 1, 15, and 30 after ligature removal (n = 8/group/time point). Histomorphometric analysis of PerioW was performed on H&E-stained sections, and immunohistochemical staining was used to quantify RANK, RANKL, and OPG expression. Results: On day 1, PerioW did not differ significantly among groups, although the PtRt group had the highest mean. A significant intergroup difference was observed for RANKL, with higher expression in PtRt than in Pt. On day 15, PerioW differed significantly among groups, with the lowest value in Rt and the highest in PtRt; the Rt–PtRt comparison was significant. At this time point, RANK, RANKL, OPG, and the RANKL/OPG ratio showed no significant intergroup differences. On day 30, no significant intergroup differences were found for PerioW or immunohistochemical parameters; however, PtRt continued to show the highest PerioW and OPG values and the lowest RANKL/OPG ratio. Conclusions: Radiotherapy superimposed on periodontitis enhanced early pro-resorptive signaling and delayed structural normalization of periodontal tissues. Although late increases in OPG suggested a compensatory response, this appeared insufficient to fully reverse radiation-associated periodontal alterations. These findings support the importance of controlling periodontal inflammation before radiotherapy to reduce subsequent periodontal tissue damage.
Full article
(This article belongs to the Special Issue Head and Neck Cancer: Current Advances and Future Perspectives)
►▼
Show Figures

Figure 1
Open AccessArticle
Association Between Oxidative Stress Biomarkers and Sperm DNA Damage in Idiopathic and Unexplained Male Infertility
by
Kenza Berrada, Asmaa Serbouti, Abderrahmane Sadek, Yasmine Touhamia, Mohieddine Moumni, Noureddine Louanjli and Rachid Aboutaieb
Biology 2026, 15(10), 802; https://doi.org/10.3390/biology15100802 (registering DOI) - 19 May 2026
Abstract
(1) Background: Oxidative stress (OS) has been extensively associated with male infertility, contributing to its pathophysiology in approximately 30–80% of affected individuals. Excessive reactive oxygen species (ROS) levels trigger a cascade of interconnected processes, including lipid peroxidation, DNA damage, and mitochondrial dysfunction, ultimately
[...] Read more.
(1) Background: Oxidative stress (OS) has been extensively associated with male infertility, contributing to its pathophysiology in approximately 30–80% of affected individuals. Excessive reactive oxygen species (ROS) levels trigger a cascade of interconnected processes, including lipid peroxidation, DNA damage, and mitochondrial dysfunction, ultimately impairing sperm quality. (2) Methods: The present study was designed to examine the relationship between seminal oxidative stress biomarkers such as superoxide dismutase (SOD), catalase (CAT), and malondialdehyde (MDA) and sperm DNA integrity in men with unexplained (UMI) and idiopathic (IMI) male infertility. The study population comprised 235 participants, who were divided into three groups: fertile controls (n = 78), UMI (n = 88), and IMI (n = 69). Semen analysis was performed according to WHO criteria. Sperm DNA fragmentation index (DFI) was assessed using the TUNEL assay, while sperm decondensation index (SDI) was evaluated by aniline blue staining. Seminal OS biomarkers were measured in seminal plasma using spectrophotometric methods. (3) Results: DFI and SDI were significantly increased in infertile groups compared with fertile controls (p < 0.001), exceeding the clinical thresholds of 30% and 15%, respectively. Antioxidant enzyme activities (SOD and CAT) were significantly reduced, whereas MDA levels were significantly elevated, particularly in the IMI group (p < 0.001). Correlation analysis revealed strong positive associations between MDA and both DFI (r = 0.76, p < 0.001) and SDI (r = 0.80, p < 0.001). Additionally, MDA, DFI, and SDI were negatively correlated with semen parameters, whereas SOD and CAT were positively correlated with sperm quality (p < 0.001). (4) Conclusions: Our findings emphasize the critical role of oxidative stress and sperm DNA integrity as potential biomarkers for the diagnosis and evaluation of UMI and IMI.
Full article
(This article belongs to the Section Developmental and Reproductive Biology)
►▼
Show Figures

Figure 1
Open AccessReview
PRDM Family Proteins in Immune Regulation: Epigenetic Control and Implications in Immune-Related Diseases
by
Shiqi Deng, Hui Li, Chengyan Guo, Jun Zou and Lingli Zhang
Biology 2026, 15(10), 801; https://doi.org/10.3390/biology15100801 (registering DOI) - 18 May 2026
Abstract
PRDM family proteins are a group of epigenetic regulators characterized by a conserved PR domain and multiple zinc-finger motifs. To date, eighteen PRDM family members have been identified in both mice and humans. Increasing evidence indicates that PRDM members play critical roles in
[...] Read more.
PRDM family proteins are a group of epigenetic regulators characterized by a conserved PR domain and multiple zinc-finger motifs. To date, eighteen PRDM family members have been identified in both mice and humans. Increasing evidence indicates that PRDM members play critical roles in the regulation of immune responses and immune-related diseases. They are involved in key processes such as hematopoiesis, immune cell differentiation, inflammatory signaling, and immunometabolic regulation. Dysregulation of PRDM proteins has been associated with a variety of pathological conditions, including hematological malignancies, autoimmune and inflammatory diseases, and metabolic disorders. Notably, functional studies of PRDM family members remain highly uneven, with a few proteins extensively characterized while many others are still poorly understood. This review summarizes recent advances in the structural features and biological functions of PRDM proteins, with a particular focus on their roles in immune regulation and endocrine–metabolic homeostasis. Furthermore, we discuss their involvement in disease pathogenesis and highlight their potential as therapeutic targets. Overall, this work provides a systematic overview of PRDM family functions and offers insights for future research in immune-mediated and metabolic diseases.
Full article
(This article belongs to the Section Immunology)
►▼
Show Figures

Figure 1
Open AccessArticle
DIAG: A Framework for Evaluating Whole-Genome Amplification Quality in Single-Cell SNV Analysis
by
Di Zhang, Mengdong Zhang, Ao Zhang, Siqi Yang, Wenfeng Huang, Tianqi Cao, Xuan Bu, Zhan Liu, Bingjie Chen and Shanjun Deng
Biology 2026, 15(10), 800; https://doi.org/10.3390/biology15100800 (registering DOI) - 18 May 2026
Abstract
Single-cell genomics offers novel insights into genomic heterogeneity within cell populations, reframing our understanding of human development, tumorigenesis, and aging. However, constrained by the picogram-scale DNA templates of individual cells, Whole-Genome Amplification (WGA) remains a necessary precondition. Current quality control frameworks primarily focus
[...] Read more.
Single-cell genomics offers novel insights into genomic heterogeneity within cell populations, reframing our understanding of human development, tumorigenesis, and aging. However, constrained by the picogram-scale DNA templates of individual cells, Whole-Genome Amplification (WGA) remains a necessary precondition. Current quality control frameworks primarily focus on amplification uniformity but fail to capture the molecular independence of DNA amplicons, leading to an overestimation of information content in redundant WGA libraries. Here, we propose the Depth of Independent Amplicons Gauge (DIAG) to accurately quantify the effective number of amplicons derived from the primary template. The robustness of the DIAG was first validated using in silico datasets, revealing that the Depth of Independent Amplicons (DIA) is directly coupled with the precision and specificity of mutation calling. Furthermore, we established an organoid-derived ground-truth to evaluate mutation fidelity in real biological contexts, confirming the practical utility of the DIAG. Our results demonstrate that the DIAG provides a high-fidelity assessment of an individual WGA library without the need for costly external experiments, especially in Single-Nucleotide Variant (SNV) calling. Furthermore, we revealed that traditional uniformity indices, like the Gini index or Kullback–Leibler (KL) divergence, exhibit incongruous fluctuations under down-sampling perturbations. In contrast, the DIA remains a robust and high-fidelity predictor of mutational accuracy, maintaining stability across varying sequencing strategies. Finally, we conducted a systematic comparison of current single-cell Whole-Genome Amplification (scWGA) strategies, providing a standardized benchmarking of diverse technologies for high-resolution single-cell mutation analysis.
Full article
(This article belongs to the Section Bioinformatics)
►▼
Show Figures

Figure 1
Open AccessArticle
A Novel Nitrogen Metabolism Pathway in Strain Gordonia sp. TD-46: Genomic and Enzymatic Evidence
by
Peiyang Zheng, Hao Li, Xiaojie Yan, Da Ao, Wenlong Yue, Shuai Zhang, Guanghua Yang and Zhiqiang Cai
Biology 2026, 15(10), 799; https://doi.org/10.3390/biology15100799 (registering DOI) - 17 May 2026
Abstract
With the acceleration of industrialization, water pollution caused by ammonia-nitrogen compounds has become a severe environmental challenge. In recent years, significant breakthroughs in biological nitrogen removal technology have been achieved alongside the discovery of novel ammonia-nitrogen-degrading bacterial strains. This study delves into the
[...] Read more.
With the acceleration of industrialization, water pollution caused by ammonia-nitrogen compounds has become a severe environmental challenge. In recent years, significant breakthroughs in biological nitrogen removal technology have been achieved alongside the discovery of novel ammonia-nitrogen-degrading bacterial strains. This study delves into the metabolic pathways and molecular mechanisms of ammonia-nitrogen degradation by Gordonia sp. TD-46. A comprehensive understanding of the strain’s nitrogen metabolic pathways and the functions of key genes was achieved by optimizing its ammonia-nitrogen degradation conditions, analyzing its whole-genome sequence, and conducting heterologous expression of crucial genes. The results demonstrated that when ammonium chloride served as the nitrogen source, sodium acetate as the carbon source, with a C/N ratio of 25, pH of 7, and an inoculum size of 15%, the strain achieved an ammonia-nitrogen degradation rate exceeding 80% under these conditions. Whole-genome sequencing analysis identified genes involved in nitrogen metabolism, including glnA, gdhA, narB, narGHI, nirBD, and nasBDE. These genes indicate that the nitrogen metabolism pathway of strain TD 46 follows the assimilatory nitrate reduction pathway (NO3− → NO2− → NH4+) and the ammonia assimilation pathway (NH4+ → Gln → Glu). Thus, strain TD-46 is capable of efficient nitrogen assimilation.
Full article
(This article belongs to the Section Microbiology)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Conservation Status, Plastome Diversity, and Evolutionary Diversification of Three Arabian Desmidorchis Endemics (Apocynaceae)
by
Samah A. Alharbi and Othman S. S. Al-Hawshabi
Biology 2026, 15(10), 798; https://doi.org/10.3390/biology15100798 (registering DOI) - 17 May 2026
Abstract
The genus Desmidorchis Ehrenb. (Apocynaceae) is a characteristic component of the succulent flora of the Arabian Peninsula, where high levels of endemism and increasing environmental pressures highlight the need for integrated genomic and conservation research. This study assessed the conservation status of three
[...] Read more.
The genus Desmidorchis Ehrenb. (Apocynaceae) is a characteristic component of the succulent flora of the Arabian Peninsula, where high levels of endemism and increasing environmental pressures highlight the need for integrated genomic and conservation research. This study assessed the conservation status of three ethnomedicinally important endemics—D. adenensis, D. arabica, and D. awdeliana—and characterizes their complete plastomes to resolve their evolutionary and temporal history. Conservation assessments were conducted following IUCN Red List criteria, and complete plastomes were sequenced and compared within a dataset of 15 subtribe Stapeliinae taxa. Comparative analyses examined the genome structure, divergence hotspots, repetitive sequences, codon usage bias, and selection pressure, while divergence times were estimated using fossil-calibrated molecular clock analyses. All three species were classified as Near Threatened (NT), primarily due to anthropogenic and environmental pressures. Plastome analyses revealed a highly conserved genome structure; however, hypervariable regions, particularly ycf1 and clpP1, exhibited elevated sequence divergence and phylogenetic informativeness. Simple sequence repeats (SSRs) were also identified as potentially informative features at the genus level. Codon usage and Ka/Ks analyses further indicated that most plastid protein-coding genes are under strong purifying selection, whereas only a few loci, particularly clpP1, showed comparatively elevated evolutionary rates. Phylogenomic analyses supported the monophyly of Desmidorchis, with molecular dating indicating recent Pleistocene diversification (~0.34–1.51 Ma), potentially associated with Quaternary climatic oscillations. Overall, this study provides an important genomic foundation for future taxonomic, evolutionary, and conservation studies of rare Arabian taxa.
Full article
Open AccessArticle
One-Pot LAMP-Coupled CRISPR/Cas12b Assay Enables Sensitive Detection of Helicobacter pylori
by
Ziyan Tang, Wentao Bai, Shuting Yan, Gaoming Luo, Yanheng Zheng, Zhuojun Bai and Zhu Chen
Biology 2026, 15(10), 797; https://doi.org/10.3390/biology15100797 (registering DOI) - 16 May 2026
Abstract
Helicobacter pylori (H. pylori) infection is closely associated with the development of chronic gastritis, peptic ulcers, and gastric cancer, highlighting the importance of rapid and accurate detection for disease prevention and clinical management. In this study, a one-pot LAMP-CRISPR/Cas12b assay targeting
[...] Read more.
Helicobacter pylori (H. pylori) infection is closely associated with the development of chronic gastritis, peptic ulcers, and gastric cancer, highlighting the importance of rapid and accurate detection for disease prevention and clinical management. In this study, a one-pot LAMP-CRISPR/Cas12b assay targeting the CagA gene was developed for H. pylori detection. First, the LAMP system was optimized by systematically screening key reaction components. Subsequently, a one-step LAMP-CRISPR/Cas12b detection platform was established through optimization of the ratio between the LAMP premix and CRISPR buffer, reaction temperature, Cas12b concentration, and ssDNA reporter concentration. Under optimal conditions, the assay achieved a detection limit of 3.14 × 101 copies/µL, representing a tenfold improvement in sensitivity compared with conventional LAMP and PCR assays (3.14 × 102 copies/µL). In addition, the entire detection process could be completed within 1 h. Validation using 17 culture-positive and 17 culture-negative samples demonstrated complete concordance with culture-based results, with no false-positive or false-negative detections observed. These findings indicate that the proposed platform possesses high sensitivity, excellent specificity, rapid turnaround, and operational simplicity, demonstrating strong potential for point-of-care testing and applications in resource-limited settings.
Full article
(This article belongs to the Section Biotechnology)
►▼
Show Figures

Figure 1
Open AccessArticle
Potential Role of Captive Environments in Reshaping the Compositions of Pathogenic Gut Bacteria in Equus Species
by
Haotian Li, Xinyuan Hou, Shile Han, Songtao Xie, Yuchun Li and Xibao Wang
Biology 2026, 15(10), 796; https://doi.org/10.3390/biology15100796 (registering DOI) - 16 May 2026
Abstract
Captive environments can have detrimental effects on the health of Equus species. However, due to the limitations of available databases, the types and abundance of pathogenic bacteria in Equus species remain largely unknown. Therefore, this study aimed to explore pathogenic gut bacteria in
[...] Read more.
Captive environments can have detrimental effects on the health of Equus species. However, due to the limitations of available databases, the types and abundance of pathogenic bacteria in Equus species remain largely unknown. Therefore, this study aimed to explore pathogenic gut bacteria in wild and captive Equus species. Using 16S rRNA gene sequencing and a comprehensive multiple bacterial pathogen detection database, we compared the pathogenic gut bacterial profiles of three Equus species (Equus ferus przewalskii, E. hemionus, and E. kiang) across different captive and wild environments. The pheatmap revealed that the three Equus species living in different captive environments showed convergence in their pathogenic gut bacterial composition. Captive Equus species had significantly enriched zoonotic bacteria, whereas wild Equus species had significantly enriched animal pathogenic bacteria. These findings suggest that a captive environment may increase the risk of zoonotic bacterial transmission between Equus species and humans. We hope that this study can provide a reasonable scientific basis for the management and protection of captive Equus species.
Full article
(This article belongs to the Section Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Habitat Associations Shape Phlebotomine Sand Fly Assemblages at the Andes–Amazon Interface in Southeastern Peru
by
Sergio Méndez-Cardona, Juliana A. Morales-Monje, Alejandro Lopera-Toro, Adrian Forsyth, Alexandra J. Bauer, Olivia R. Magaletta, Panpim Thongsripong and Olga L. Cabrera-Quintero
Biology 2026, 15(10), 795; https://doi.org/10.3390/biology15100795 (registering DOI) - 16 May 2026
Abstract
Phlebotomine sand flies remain poorly studied in southeastern Peru, a region with a high burden of cutaneous leishmaniasis (CL). Using modified ultraviolet (UV) light traps, we surveyed sand fly assemblages at Manu Biological Station during the wet season within secondary forest, Guadua bamboo
[...] Read more.
Phlebotomine sand flies remain poorly studied in southeastern Peru, a region with a high burden of cutaneous leishmaniasis (CL). Using modified ultraviolet (UV) light traps, we surveyed sand fly assemblages at Manu Biological Station during the wet season within secondary forest, Guadua bamboo forest, fruit crop plots, and peridomicile habitats. We collected 2641 sand flies representing 32 species. Habitat type was the primary driver of assemblage composition, with minimum nightly temperature as the strongest environmental correlate. Sand fly abundance was highest in secondary forest (n = 921) and peridomicile habitats (n = 836), where assemblages were dominated by Nyssomyia shawi, a generalist species. Although Guadua bamboo forests harbored lower abundance (n = 386), potential vector species comprised 92% of the assemblage compared to 42–86% in other habitats, suggesting that expanding bamboo forests may favor a higher proportion of potential CL vectors. Peridomicile assemblages consisted largely of generalist species that overlapped with adjacent forested habitats, indicating potential pathways for sylvatic-to-peridomestic spillover. Although the study’s limited scope (i.e., limited to a single season and locality) does not allow broad generalization, our findings suggest the importance of habitat in structuring patterns of potential vector exposure.
Full article
(This article belongs to the Special Issue Ecological Dynamics of Vector-Borne Pathogens: From Hosts to Vectors)
►▼
Show Figures

Figure 1
Open AccessReview
Immunomodulatory Mechanisms of Mesenchymal Stromal Cells: Cytokine Networks and Therapeutic Potential Across Immune-Mediated, Inflammatory, and Regenerative Disorders
by
Tamerlan Nurlybek, Nursulu Altaeva, Baglan Kazhiyakhmetova, Zhansaya Seitkumarova, Yerkezhan Baidildina, Anastassiya Vizigina and Yerlan Kashkinbayev
Biology 2026, 15(10), 794; https://doi.org/10.3390/biology15100794 (registering DOI) - 16 May 2026
Abstract
Mesenchymal stromal cells (MSCs) are multipotent cells characterized by their regenerative capacity and strong immunomodulatory properties. In recent years, MSC-based therapy has attracted significant attention as a potential treatment for a wide range of immune-mediated and degenerative diseases. The therapeutic effects of MSCs
[...] Read more.
Mesenchymal stromal cells (MSCs) are multipotent cells characterized by their regenerative capacity and strong immunomodulatory properties. In recent years, MSC-based therapy has attracted significant attention as a potential treatment for a wide range of immune-mediated and degenerative diseases. The therapeutic effects of MSCs are primarily mediated through paracrine signaling and secretion of cytokines that regulate immune responses and promote tissue repair. This review focuses on five key cytokines involved in MSC immunomodulation: interleukin-6 (IL-6), interleukin-10 (IL-10), transforming growth factor-beta (TGF-β), tumor necrosis factor-alpha (TNF-α), and interleukin-1 beta (IL-1β). These cytokines interact within a complex signaling network that allows MSCs to suppress excessive inflammation and restore immune balance. The role of MSC therapy is examined in several clinically relevant conditions, including systemic lupus erythematosus, systemic sclerosis, ischemic stroke, spinal cord injury, diabetes mellitus, and female infertility. Across these diseases, MSCs demonstrate the ability to inhibit pro-inflammatory immune cell activity, promote regulatory immune phenotypes, reduce oxidative stress, and stimulate regeneration through the secretion of growth factors and extracellular vesicles. Despite promising experimental and early clinical findings, several limitations remain, including variability in MSC sources, limited cell survival after transplantation, and the need for optimized dosing strategies. Overall, MSC therapy represents a multifunctional therapeutic approach combining immunomodulation, anti-inflammatory activity, and regenerative support. Further research is required to better understand cytokine interactions, improve standardization of MSC-based treatments, and enhance clinical efficacy across diverse pathological conditions.
Full article
(This article belongs to the Section Immunology)
Open AccessArticle
The Ascosphaera apis Invasion of Apis cerana Worker Larvae: Long Non-Coding RNA-Mediated Regulation
by
Yunzhen Yang, Kaiyao Zhang, Genchao Gan, Shuai Zhou, Qingwei Tan, Jianfeng Qiu, Dafu Chen, Zhongmin Fu and Rui Guo
Biology 2026, 15(10), 793; https://doi.org/10.3390/biology15100793 (registering DOI) - 15 May 2026
Abstract
Ascosphaera apis, an obligate lethal fungal pathogen that infects bee larvae, and causes chalkbrood disease, poses a significant threat to the global beekeeping industry. Long non-coding RNAs (lncRNAs) are employed by pathogens to enhance infectivity and evade host immunity. Here, lncRNAs in
[...] Read more.
Ascosphaera apis, an obligate lethal fungal pathogen that infects bee larvae, and causes chalkbrood disease, poses a significant threat to the global beekeeping industry. Long non-coding RNAs (lncRNAs) are employed by pathogens to enhance infectivity and evade host immunity. Here, lncRNAs in A. apis spores (AaCK group) and the guts of 4-, 5-, and 6-day-old Apis cerana cerana worker larvae inoculated with A. apis spores (AaT1, AaT2, and AaT3 groups) were identified, characterized, and validated. Additionally, the expression pattern of fungal lncRNAs during infection was analyzed, followed by an investigation of the regulatory manners and roles of differentially expressed lncRNAs (DElncRNAs). A total of 1379 lncRNAs were identified in AaCK, AaT1, AaT2, and AaT3 groups using bioinformatics, involving various types such as sense lncRNAs, antisense lncRNAs, bidirectional lncRNAs, intergenic lncRNAs, and intronic lncRNAs. Additionally, 4, 9, and 75 up-regulated lncRNAs as well as 2, 1, and 15 down-regulated ones were identified in the 4-, 5-, and 6-day-old larval guts following A. apis inoculation. Fifteen DElncRNAs as potential antisense lncRNAs may interact with 15 sense-strand mRNAs in the AaCK vs. AaT3 comparison group. Cis-acting analysis identified 10, 16, and 136 upstream and downstream genes of DElncRNAs in the aforementioned comparison groups, involving a series of GO terms and KEGG pathways like metabolic process and biosynthesis of secondary metabolites. Following the trans-acting investigation, 752, 821, and 1327 co-transcribed genes with DElncRNAs were discovered, spanning an array of functional terms and pathways such as biological processes and glycerophospholipid metabolism. Analysis of a competing endogenous RNA (ceRNA) network indicated that 1 and 5 DElncRNAs in the AaCK vs. AaT1 and AaCK vs. AaT3 comparison groups potentially targeted 1 and 2 miRNAs, further targeting 208 and 286 mRNAs, respectively. Further analysis identified one ceRNA axis relevant to the MAPK signaling pathway and several ceRNA networks associated with the biosynthesis of secondary metabolites. Finally, RT-qPCR results confirmed that the expression trends of six randomly selected DElncRNAs were consistent with those in the transcriptome data. These findings not only offer a foundation for elucidating the mechanisms underlying DElncRNA-mediated A. apis infection but also enrich our understanding of honeybee host–fungal pathogen interactions.
Full article
(This article belongs to the Section Infection Biology)
►▼
Show Figures
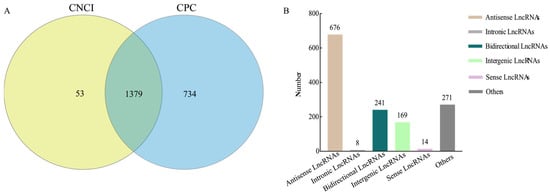
Figure 1
Open AccessReview
Deconvolution of Red Blood Cells Thermal Fluid Biopsy Following Systematic Cyclophosphamide or Cilostazol Drug Therapies
by
Andrea Ferencz and Dénes Lőrinczy
Biology 2026, 15(10), 792; https://doi.org/10.3390/biology15100792 (registering DOI) - 15 May 2026
Abstract
Beyond gas transport, red blood cells (RBCs) have emerging roles regarding innate immunity, regulating blood flow, and participating in nutrient transport, which can be engineered as drug delivery systems since they contribute to maintaining water homeostasis. Following extensive thermoanalytical studies of human blood
[...] Read more.
Beyond gas transport, red blood cells (RBCs) have emerging roles regarding innate immunity, regulating blood flow, and participating in nutrient transport, which can be engineered as drug delivery systems since they contribute to maintaining water homeostasis. Following extensive thermoanalytical studies of human blood plasma, our working group investigated the properties of RBCs, examining their role in healthy and in different disease states by using differential scanning calorimetry (DSC) and the deconvolution of the resulting thermal curve. In the first study, guinea pigs were treated with intraperitoneal chemotherapy. Cyclophosphamide treatment showed a dose-dependent difference between the thermal parameters of control and treated samples, indicating that DSC can be used in this area. Following deconvolution of the DSC studies, the changes can be attributed to the damaged compounds. In the second part of our study, a method for the thermal analysis and deconvolution of RBCs in patients with lower limb ischemia during a three-month cilostazol treatment was developed. The control DSC curve showed 5-6 distinct thermal domains, and in contrast to other drug treatments, this remained stable throughout the entire study period. No effects of stiffness or compact were caused by the anticancer drug cyclophosphamide were observed in the structure of RBCs. These preliminary results highlight the uniqueness of thermodynamic studies of RBCs and provide a fingerprint-like identification of a given individual or disease state.
Full article
(This article belongs to the Special Issue Erythrocytes in Human Life—Functions Beyond Oxygen Transport)
Open AccessArticle
Single-Cell Transcriptomic Analysis Reveals Early Transcriptional Heterogeneity of Cardiac-Associated Cell Populations During Zebrafish Embryogenesis
by
Samer N. Khalaf, Mundher Jabbar Al-Okhedi, Amal Saeed Alayed, Mariam M. Jaddah and Asra’a Adnan Abdul-Jalil
Biology 2026, 15(10), 791; https://doi.org/10.3390/biology15100791 (registering DOI) - 15 May 2026
Abstract
Understanding the development and differentiation of cardiac progenitor cells during the initial stages of embryogenesis is central to a complete understanding of vertebrate heart development. In zebrafish, cardiac specification begins during gastrulation; however, the single-cell transcriptional dynamics of initial cardiac lineage commitment remain
[...] Read more.
Understanding the development and differentiation of cardiac progenitor cells during the initial stages of embryogenesis is central to a complete understanding of vertebrate heart development. In zebrafish, cardiac specification begins during gastrulation; however, the single-cell transcriptional dynamics of initial cardiac lineage commitment remain not fully defined. In this case, we integrated single-cell RNA sequencing datasets of zebrafish embryos at 4 and 6 h post-fertilisation (hpf) to investigate early cardiac lineage specification. The unsupervised clustering of the integrated dataset identified 12 distinct cell clusters, which made it possible to identify a transcriptionally distinct population of cells characterised by the coordinated expression of transcription factors associated with cardiac development. A further subclustering of the cells expressing cardiac-associated transcription factors showed a significant level of early diversification of the cardiac progenitor group. A projection onto low-dimensional embedding revealed a structured transcriptional organisation of the cardiac subclusters, marked by the differential expression of key cardiac transcription factors, including Gata5, Gata6, Hand2, Nkx2.5, and Tbx5a. A pseudotemporal trajectory analysis uncovered a continuous developmental progression within the cardiac lineage and indicated the gene-specific dynamic regulation and temporal hierarchy of cardiac transcriptional programs. Collectively, these results indicate that zebrafish cardiac progenitors are transcriptionally diverse and acquire cardiac fate through a sustained, continuous regulatory process rather than an abrupt fate transition. This work provides an informative, high-resolution model of early cardiac lineage specification and highlights the power of single-cell transcriptomics for analysing dynamic events in vertebrate embryogenesis.
Full article
(This article belongs to the Section Bioinformatics)
Open AccessReview
Antimicrobial Peptides in Fish: Mechanisms of Action and Applications in Aquaculture
by
Fan Zhou, Leyi Zhou, Pengfei Wang, Mariano Elisio, Sally Salaah, Bakhtiyor Karimov and Quanquan Cao
Biology 2026, 15(10), 790; https://doi.org/10.3390/biology15100790 (registering DOI) - 15 May 2026
Abstract
With the rapid development of global aquaculture, the frequent occurrence of fish diseases has had a serious impact on the efficiency of aquaculture and the ecological environment. Antimicrobial peptides, as a kind of natural immune active substance existing in organisms, participate in innate
[...] Read more.
With the rapid development of global aquaculture, the frequent occurrence of fish diseases has had a serious impact on the efficiency of aquaculture and the ecological environment. Antimicrobial peptides, as a kind of natural immune active substance existing in organisms, participate in innate immunity and adaptive immunity. Due to their extensive antibacterial properties and low toxicity, they have gradually become a hot topic in scientific research. This article reviews the classification, tissue distribution, mechanism of action, extraction, and synthesis techniques of antimicrobial peptides (AMPs) derived from fish, as well as their applications in disease prevention in aquaculture, product preservation, and antibiotic substitution. Although antimicrobial peptides are expected to become alternatives to antibiotics, challenges such as environmental stability, production costs, and regulatory frameworks remain to be addressed. This article holds that antimicrobial peptides derived from fish are a feasible strategy for sustainable aquaculture. The future development direction lies in biotechnology-driven optimization, carrier innovation, and combined application with traditional antibiotics.
Full article
(This article belongs to the Special Issue Pathology and Physiology Insights in Animals)
Open AccessArticle
Effects of Alpha Particle Exposure on Genetic Stability and Morphogenesis in Drosophila melanogaster
by
Zarema Biyasheva, Yuliya Zaripova, Anna Lovinskaya, Vyacheslav Dyachkov and Alexandr Yushkov
Biology 2026, 15(10), 789; https://doi.org/10.3390/biology15100789 (registering DOI) - 15 May 2026
Abstract
The study of genetic effects induced by low-dose alpha radiation associated with radon and its decay progeny is critically important for assessing radiation risks in regions with elevated natural background levels. The aim of this study was to evaluate the mutagenic effects (in
[...] Read more.
The study of genetic effects induced by low-dose alpha radiation associated with radon and its decay progeny is critically important for assessing radiation risks in regions with elevated natural background levels. The aim of this study was to evaluate the mutagenic effects (in germline cells) and teratogenic effects (in somatic tissues) of alpha radiation using the D. melanogaster model. To differentiate between these effects, teratogenic outcomes were analyzed in directly exposed individuals (phenotypic analysis of adults that developed from irradiated larvae), whereas mutagenic effects were assessed in the progeny of irradiated flies. Larvae and adult flies were exposed to calibrated alpha-particle sources with energies ranging from 4.8 to 7.7 MeV and absorbed doses of 1.90–44.96 mGy. The results demonstrated a statistically significant increase in the frequency of morphological abnormalities in the exposed groups, including melanotic masses and deformities of the wings, thorax, and tergites. Under 72 h exposure, a strong correlation between absorbed dose and abnormality frequency was observed (r = 0.98). In the reporter system, induction of GFP expression was detected in imaginal discs at doses above 10 mGy, indicating threshold activation of the cellular stress response. The obtained data demonstrate that chronic low-dose α-irradiation leads to an increased frequency of morphological abnormalities (indirect phenotypic manifestations of compromised genetic stability) in D. melanogaster, with the most pronounced effects observed at the level of morphogenesis. The high sensitivity of the applied test systems was confirmed, supporting the use of D. melanogaster as a bioindicator for ecogenetic monitoring of radon-prone areas, including regions of Kazakhstan.
Full article
(This article belongs to the Topic Disease Risks from Environmental Radiological Exposure)
►▼
Show Figures
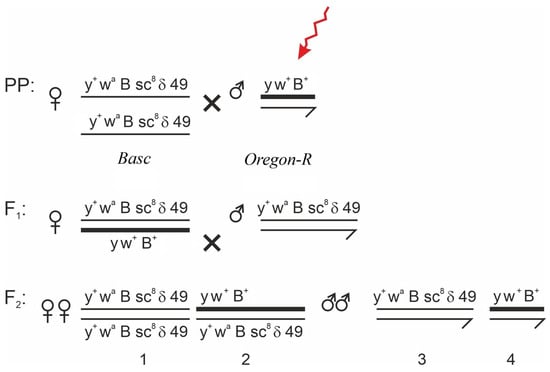
Figure 1
Open AccessArticle
Transcriptomic and Metabolomic Profiling Reveals the Antiproliferative Mechanism of Goose Serum and Plasma in SW1990 Cells
by
Xiaolong Zhou, Mihan Wu, Han Wang, Xiangchen Li, Songbai Yang and Ayong Zhao
Biology 2026, 15(10), 788; https://doi.org/10.3390/biology15100788 (registering DOI) - 15 May 2026
Abstract
►▼
Show Figures
Goose blood has anticancer properties and was recorded in ancient China, but the specific molecular mechanisms underlying this effect still require further exploration. In this study, SW1990 cells were treated with goose serum or plasma, and transcriptome analysis was performed to explore the
[...] Read more.
Goose blood has anticancer properties and was recorded in ancient China, but the specific molecular mechanisms underlying this effect still require further exploration. In this study, SW1990 cells were treated with goose serum or plasma, and transcriptome analysis was performed to explore the function of goose blood on cancer cells. Metabolomic profiling was also performed on goose serum, goose plasma, chicken serum, and chicken plasma to identify the bioactive substances responsible for the anticancer effect. The study examined the effects of goose plasma and serum on SW1990 cells and compared the metabolites between goose and chicken blood. Wound scratch, CCK-8, and Annexin V-PI assays showed that goose plasma and serum inhibited SW1990 cell proliferation at 24 and 48 h. Both treatments reduced cell viability, with serum inducing early and late apoptosis and plasma inducing late apoptosis. RNA sequencing (RNA-seq) identified 2259 (1418 upregulated, 841 downregulated) and 2731 (1844 upregulated, 887 downregulated) differentially expressed genes (DEGs) in the plasma and serum groups versus the negative control (NC), respectively, and 689 DEGs between the plasma and serum groups. Gene Ontology (GO) and KEGG pathway analyses revealed that the DEGs were enriched in processes such as lipid metabolism, JAK-STAT, and IL-17 pathways. Untargeted liquid chromatography–tandem mass spectrometry (LC-MS/MS) analysis identified distinct metabolites in goose and chicken blood, with unique metabolites and differential ones between groups. In SW1990 cells, four metabolite subclusters matched the plasma and serum effects. In summary, goose blood can suppress cancer cells by regulating gene expression to affect the key signaling pathways involved in cancer cell apoptosis and autophagy. Certain metabolites present at high concentrations in goose blood, such as cucurbitacin D and Oleoyl-L-carnitine, may also contribute to the inhibition of cancer cell proliferation and migration. These findings suggest that goose blood holds broad application prospects as a future auxiliary drug for cancer treatment, and this study provides a theoretical basis for the further application of goose products.
Full article

Figure 1

Journal Menu
► ▼ Journal Menu-
- Biology Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, JCM, Diagnostics, Dentistry Journal
Assessment of Craniofacial Morphology: Traditional Methods and Innovative Approaches
Topic Editors: Nikolaos Gkantidis, Carlalberta VernaDeadline: 1 June 2026
Topic in
Animals, Biology, Conservation, Diversity, Ecologies, Forests, Land
Conservation at the Crossroads: Forest Ecology, Wildlife Dynamics, and Emerging Challenges for Ecosystem Resilience
Topic Editors: Yiannis G. Zevgolis, Triantaphyllos Akriotis, Anastasia Christopoulou, Panayiotis G. Dimitrakopoulos, Dimitra-Lida Rammou, Dionisios YoulatosDeadline: 31 July 2026
Topic in
Biology, Biomolecules, Cancers, Cells, IJMS, Pharmaceuticals, Kinases and Phosphatases
Kinases in Cancer and Other Diseases, 2nd Edition
Topic Editors: Jonas Cicenas, Anna M. CzarneckaDeadline: 31 August 2026
Topic in
JoX, Sustainability, Water, Hydrology, Biology, Environments
Sustainable Water Resource Management: Controlling Nutrient and Contaminant Transport in Aquatic Systems
Topic Editors: Taotao Lu, Shuangcheng TangDeadline: 30 October 2026

Special Issues
Special Issue in
Biology
The Biology of Animal Reproduction
Guest Editor: Jingli TaoDeadline: 30 May 2026
Special Issue in
Biology
Studies on Antioxidant Activity of Natural Products from Plants
Guest Editors: Pornsiri Pitchakarn, Piya TemviriyanukulDeadline: 31 May 2026
Special Issue in
Biology
Recent Advances in Biological Research on Prostate Cancer
Guest Editors: Samson Mashele, Pakiso MakhoahleDeadline: 31 May 2026
Special Issue in
Biology
Artificial Intelligence Research for Complex Biological Systems
Guest Editors: Yong Chen, Milana Frenkel-MorgensternDeadline: 31 May 2026
Topical Collections
Topical Collection in
Biology
Crop Improvement Now and Beyond
Collection Editors: Pierre Devaux, Pierre Sourdille
Topical Collection in
Biology
Applied Physics in Cancer Cells
Collection Editors: Jagoba Iturri, José Toca-Herrera
Topical Collection in
Biology
Molecular Mechanisms of Aging
Collection Editors: Serena Dato, Giuseppina Rose, Paolina Crocco











