Journal Description
Organoids
Organoids
is an international, peer-reviewed, open access journal on all aspects of organoids published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within ESCI (Web of Science), Scopus, and many other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 27.8 days after submission; acceptance to publication is undertaken in 3.2 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- Organoids is a companion journal of Cells.
Latest Articles
Dissecting PDE6-Associated Inherited Retinal Dystrophies Using Patient-Derived Retinal Models
Organoids 2026, 5(2), 13; https://doi.org/10.3390/organoids5020013 - 7 May 2026
Abstract
Inherited retinal dystrophies (IRDs) comprise a diverse group of genetic disorders that frequently result in irreversible vision loss due to photoreceptor dysfunction or degeneration. Among them, retinitis pigmentosa (RP) and achromatopsia (ACHM) are, in some cases, associated with pathogenic variants in PDE6A and
[...] Read more.
Inherited retinal dystrophies (IRDs) comprise a diverse group of genetic disorders that frequently result in irreversible vision loss due to photoreceptor dysfunction or degeneration. Among them, retinitis pigmentosa (RP) and achromatopsia (ACHM) are, in some cases, associated with pathogenic variants in PDE6A and PDE6C, respectively, which are key components of the phototransduction cascade. As most of IRDs still lack effective therapies, retinal organoids (ROs) provide a valuable in vitro model for the investigation of disease-associated mechanisms. Here, we generated induced pluripotent stem cell (iPSC)-derived ROs from an RP patient carrying compound heterozygous PDE6A mutations and from a patient with ACHM harboring a homozygous PDE6C mutation, along with their corresponding CRISPR/Cas9-corrected isogenic controls, which, to our knowledge, represent the first patient-derived RO models reported for the PDE6A and PDE6C genes. The mutant PDE6A line exhibited impaired neuroretinal vesicle formation and RO differentiation; however, a subset of RP-derived ROs matured appropriately and retained photoreceptor features. Moreover, the specific isoform expression pattern detected in retinal tissues reflected differences across developmental maturation stages that could influence disease severity. In contrast, the PDE6C_mutant ROs displayed normal structure and maturation, although cGMP hydrolysis within photoreceptors was likely compromised. In both models, CRISPR/Cas9-mediated correction restored the disease-associated phenotype resembling wild-type ROs. Collectively, these findings provide new insights into PDE6-associated pathogenesis, underscore the utility of patient-specific and gene-corrected ROs for elucidating IRD mechanisms, and support gene editing as a promising therapeutic strategy.
Full article
(This article belongs to the Special Issue Advances in Organoid Technology: Bridging the Gap between Research and Therapy)
►
Show Figures
Open AccessArticle
Advanced Glycation End Products Induce Microglial Activation and Impair Neurodevelopment in Human iPSC-Derived Brain Organoids
by
Rika Kumar, Grace Shinn, Jimmy Lin, Qingshun Q. Li and Yiling Hong
Organoids 2026, 5(2), 12; https://doi.org/10.3390/organoids5020012 - 20 Apr 2026
Abstract
►▼
Show Figures
Advanced Glycation End Products (AGEs) are reactive compounds formed through the non-enzymatic glycation of proteins, lipids, or nucleic acids due to exposure to reducing sugars. They accumulate through endogenous metabolic dysregulation and exogenous dietary intake, particularly high-fat and high-sugar foods prepared at high
[...] Read more.
Advanced Glycation End Products (AGEs) are reactive compounds formed through the non-enzymatic glycation of proteins, lipids, or nucleic acids due to exposure to reducing sugars. They accumulate through endogenous metabolic dysregulation and exogenous dietary intake, particularly high-fat and high-sugar foods prepared at high temperatures. The interaction between AGEs and their receptor, RAGE (receptor for Advanced Glycation End Products), has been implicated in a range of pathological conditions, including diabetes and metabolic syndrome. However, the impact of AGEs accumulation on neurodevelopment remains poorly understood. In this study, we investigated the effects of AGEs on human-induced pluripotent stem cell (iPSC)-derived cerebral organoids comprising neurons, astrocytes, and microglia. Our findings reveal that AGEs induce RAGE expression, leading to microglial activation, increased deposition of amyloid-beta (Aβ) aggregates, and impaired neurodevelopment. Additionally, elevated levels of AGE-modified proteins, along with altered microglial polarization, were observed in cerebral organoids modeling Western Pacific Amyotrophic Lateral Sclerosis and Parkinsonism–Dementia Complex (ALS-PDC). These findings demonstrate AGEs as active drivers of neurodevelopmental disruption and establish a mechanistic link between metabolic stress and increased susceptibility to neurodegenerative disease.
Full article

Figure 1
Open AccessArticle
Glucose Levels Impact the Morphology and Cell Type Composition of Human Cerebral Organoids
by
Gautami R. Kelkar, Balaji M. Rao and Albert J. Keung
Organoids 2026, 5(2), 11; https://doi.org/10.3390/organoids5020011 - 5 Apr 2026
Abstract
►▼
Show Figures
Human cerebral organoids, derived from pluripotent stem cells, are powerful models for studying human brain development. The understanding of how morphogens can be used to guide patterning and differentiation has matured rapidly; however, the influence of basal media components on organoid development remains
[...] Read more.
Human cerebral organoids, derived from pluripotent stem cells, are powerful models for studying human brain development. The understanding of how morphogens can be used to guide patterning and differentiation has matured rapidly; however, the influence of basal media components on organoid development remains unclear. Standard organoid media frequently contain non-physiological concentrations of nutrients, including glucose, a central regulator of cellular metabolism and signaling. Here, we examine how glucose availability shapes cerebral organoid growth, morphology, and cell type composition by comparing conventional hyperglycemic media to media with glucose levels more closely resembling normoglycemic conditions. We find that organoids derived from multiple human pluripotent stem cell lines can grow in low glucose, but they exhibit altered growth rates, structural features, and lineage distributions. In H9 embryonic stem cell-derived organoids, inhibition of the mammalian target of rapamycin pathway under low glucose restores neurodevelopmental cell types otherwise diminished in these conditions. These findings highlight glucose as a key determinant of organoid lineage specification and cellular signaling. Importantly, however, glucose modulation does not reduce variability across organoids or cell lines, underscoring the need to better understand and control sources of heterogeneity to improve organoid models.
Full article

Figure 1
Open AccessArticle
Optimized Large-Scale Longitudinal Biorepository of Gastroesophageal Adenocarcinoma Patient-Derived Organoids: High-Fidelity Models for Personalized Treatment to Overcome Resistance
by
Mingyang Kong, Sanjima Pal, Shuyuan Wang, Julie Bérubé, Ruoyu Ma, Yifei Yan, Wotan Zeng, France Bourdeau, Betty Giannias, Hong Zhao, Nathan Osman, Yehonatan Nevo, Kulsum Tai, Hellen Kuasne, James Tankel, Gertruda Evaristo, Pierre O. Fiset, Xin Su, Swneke Bailey, Morag Park, Nicholas Bertos, Veena Sangwan and Lorenzo Ferriadd
Show full author list
remove
Hide full author list
Organoids 2026, 5(2), 10; https://doi.org/10.3390/organoids5020010 - 30 Mar 2026
Abstract
►▼
Show Figures
A major limitation in studying gastroesophageal adenocarcinoma (GEA) has been the lack of reliable models that represent the disease’s complexity. We present lessons learned from a comprehensive large-scale biobanking effort combining traditional sample collection with several in vitro models, including 3-dimensional patient-derived organoids
[...] Read more.
A major limitation in studying gastroesophageal adenocarcinoma (GEA) has been the lack of reliable models that represent the disease’s complexity. We present lessons learned from a comprehensive large-scale biobanking effort combining traditional sample collection with several in vitro models, including 3-dimensional patient-derived organoids (PDOs), 2-dimensional cancer-associated fibroblasts (CAFs), tumor-infiltrating lymphocytes (TILs), and/or in vivo xenografts. This initiative started in 2018, integrating multiple advanced ex vivo models such as PDOs, patient-derived xenografts (PDXs), and organoids (PDXOs). This unique resource now includes tumor avatars from over 380 consented patients, making it the world’s largest living GEA biobank. We achieved a >90% success rate in creating per-patient models, including 227 tumor-derived and 203 neighboring normal PDOs. These organoids accurately mirror key features of the original tumors, such as their histology (e.g., microsatellite instability), mutations, and drug response across treatment points. Notably, PDOs can predict individual patient responses to chemotherapy within five weeks, underscoring their clinical relevance. Furthermore, high-throughput drug screening on PDO subsets with known genetic landscapes generates personalized chemosensitivity profiles for 22 drugs. Through a process of continued refinement of culture techniques and tumor sampling approach, our large-scale comprehensive collection of GEA avatars represents a unique and valuable preclinical experimental resource for precision oncology.
Full article

Figure 1
Open AccessReview
Organoid Models: Revolutionizing Disease Modeling and Personalized Therapeutics
by
Zhifeng Xue, Runze Yang, Yaling Liu and Han Luo
Organoids 2026, 5(1), 9; https://doi.org/10.3390/organoids5010009 - 13 Mar 2026
Abstract
As a three-dimensional in vitro model, organoid technology represents a revolutionary breakthrough in precision medicine. By harnessing the self-organizing capabilities of stem cells within biomimetic extracellular matrices, it enables the generation of miniature tissues that recapitulate key structural and functional characteristics of their
[...] Read more.
As a three-dimensional in vitro model, organoid technology represents a revolutionary breakthrough in precision medicine. By harnessing the self-organizing capabilities of stem cells within biomimetic extracellular matrices, it enables the generation of miniature tissues that recapitulate key structural and functional characteristics of their source organs. Conventional two-dimensional cell cultures lack tissue architecture and microenvironmental cues, whereas animal models are hindered by interspecies differences and inadequate representation of human pathological heterogeneity. By effectively addressing these limitations, organoids have emerged as powerful platforms that are highly representative of human physiology and disease processes in oncology, genetic disorders, and infectious diseases. They demonstrate significant potential for use in drug screening, toxicity assessment, and the development of personalized treatment strategies. Although challenges such as limited vascularization, lack of standardized culture protocols, and ethical considerations remain, the integration of multidisciplinary approaches such as AI-assisted analysis, organ-on-a-chip systems, and 3D bioprinting, together with increasing policy support and industrial advancement, is accelerating the clinical translation of organoid technology. In this review, the construction strategies for and applications of organoid models are systematically summarized, and their value and limitations in disease modeling, precision medicine, and preclinical research are highlighted. Finally, future development pathways driven by multidisciplinary collaboration and standardization are outlined.
Full article
(This article belongs to the Special Issue Advances in Organoid Technology: Bridging the Gap between Research and Therapy)
►▼
Show Figures

Figure 1
Open AccessReview
Understanding Alzheimer’s Disease Through Neurodevelopment: Insights from Human Cerebral Organoids
by
Patricia Mateos-Martínez, Deanira Patrone, Milagros González-Flores, Cristina Soriano-Amador, Rosa González-Sastre, Sabela Martín-Benito, Andreea Rosca, Raquel Coronel, Victoria López-Alonso and Isabel Liste
Organoids 2026, 5(1), 8; https://doi.org/10.3390/organoids5010008 - 10 Mar 2026
Abstract
►▼
Show Figures
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder and the leading cause of dementia, for which there is currently no cure. The causes of AD are still not well understood, although 5% of cases are known to have a genetic origin, associated with
[...] Read more.
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder and the leading cause of dementia, for which there is currently no cure. The causes of AD are still not well understood, although 5% of cases are known to have a genetic origin, associated with pathogenic genetic variants of the APP and PSEN1/2 genes. There is growing evidence that both APP and PSEN1/2 are also essential for proper human brain development and neural/neuronal function. This implies that abnormalities in early brain development could increase neuronal vulnerability to AD later in life. Human cerebral organoids (hCOs), generated from induced pluripotent stem cells (iPSCs) from AD patients, provide an exceptional model for better understanding the cellular and molecular mechanisms involved in human brain development, as well as early neurological alterations in the evolution of AD. This review compiles the main studies in which hCOs are used as a model for studying AD and for the discovery of new biomarkers. We also discuss the advantages and applications of these hCOs for studying the early stages of AD from a neurodevelopmental perspective. Finally, we mention the main current challenges in the use of hCOs for future research into AD.
Full article

Figure 1
Open AccessArticle
A Spheroid-Based In Vitro Model to Generate the Zonal Organisation of the Tendon-to-Bone Enthesis
by
Vinothini Prabhakaran and Jennifer Z. Paxton
Organoids 2026, 5(1), 7; https://doi.org/10.3390/organoids5010007 - 10 Feb 2026
Cited by 1
Abstract
►▼
Show Figures
The tendon-to-bone enthesis is a multiphasic structure with four structurally continuous and compositionally distinct regions: tendon, uncalcified fibrocartilage, calcified fibrocartilage and bone. Our study aimed to develop 3D scaffold-free in vitro spheroids and macro-tissues of the enthesis for applications as experimental tools to
[...] Read more.
The tendon-to-bone enthesis is a multiphasic structure with four structurally continuous and compositionally distinct regions: tendon, uncalcified fibrocartilage, calcified fibrocartilage and bone. Our study aimed to develop 3D scaffold-free in vitro spheroids and macro-tissues of the enthesis for applications as experimental tools to understand the development and repair of enthesis injury. This study hypothesises that integrating tendon and bone cell spheroids with bone marrow mesenchymal stem cell spheroids will facilitate the production of a fibrocartilaginous interface. 3D Spheroids: The biphasic (tendon–bone) and triphasic co-culture (tendon–stem cell–bone) of spheroids in growth media and chondrogenic media were investigated to establish fusion kinetics, and the cellular and ECM components produced via histology and immunohistochemistry. Complete fusion between spheroids occurred within 6-to-8 days in biphasic co-culture, and 15-to-20 days in triphasic co-culture. Compared to biphasic, the triphasic co-culture in chondrogenic media showed a continuous interface connecting the tendon and bone regions. The presence of collagen I, sulphated proteoglycans and collagen type II in the interface region of triphasic co-culture indicates fibrochondrogenic differentiation. 3D macro-tissues: The modular tissue engineering strategy was used in this study to produce enthesis macro-tissues using spheroids as building blocks. Spheroids were bio-assembled in the triphasic manner (12 tendon spheroids, 12 stem cell spheroids and 8 bone spheroids) in the custom-designed and 3D-printed temporary supports (Formlabs Clear Resin®) using a customised spheroid bio-assembly system. The fusion of spheroids occurred by day 8 after bio-assembly, and they were removed from temporary supports and cultured in scaffold-free conditions. Although the bio-assembly methodology was successful in producing fused scaffold-free macro-tissues, the histological analysis revealed the presence of an extensive necrotic core due to the large-sized constructs. To conclude, the findings support the hypothesis that a triphasic co-culture has the potential to produce a structurally continuous fibrocartilaginous interface but requires further optimisation to produce macro-tissues with anatomical morphologies and reduced necrotic cores.
Full article

Figure 1
Open AccessArticle
Xenograft-Derived Human Breast Cancer Organoids Can Form Chimeras with Host Mouse Mammary Epithelial Cells Which Promote Tumor Cell Proliferation
by
Hiroyuki Uematsu, Chieko Saito, Jumpei Kondo, Kunishige Onuma, Roberto Coppo, Hiroko Endo, Takahiro Nakayama, Katsuhide Yoshidome, Taisei Nomura, Arihiro Kohara and Masahiro Inoue
Organoids 2026, 5(1), 6; https://doi.org/10.3390/organoids5010006 - 6 Feb 2026
Abstract
►▼
Show Figures
Breast cancer progression and treatment responsiveness are significantly influenced by the tumor microenvironment. Therefore, transplantation into the mammary fat pad is widely employed to establish a mouse xenograft model of breast cancer. This study reports chimeric organoids derived from breast cancer xenografts composed
[...] Read more.
Breast cancer progression and treatment responsiveness are significantly influenced by the tumor microenvironment. Therefore, transplantation into the mammary fat pad is widely employed to establish a mouse xenograft model of breast cancer. This study reports chimeric organoids derived from breast cancer xenografts composed of human and mouse cells. During passaging of an organoid line derived from breast cancer xenografts, characteristic cell clusters composed of smaller cells were observed. Immunostaining with a mouse-specific antibody revealed that the smaller cells were mouse cells composed of luminal- and basal-like cells. Chimeric organoids were observed in four of the six xenograft-derived organoid lines. Organoids composed solely of human cells rapidly diminished after passaging, with chimeric and mouse-cell-only organoids becoming predominant. When human breast cancer cells were co-cultured with mouse mammary epithelial cells, chimeras were frequently observed. The PCNA positivity rate in breast cancer cells within chimeras was higher than that in breast cancer cells within organoids composed solely of human cells. These findings indicate that xenograft-derived breast cancer organoids frequently contain mouse cells and that mouse mammary epithelial cells promote the proliferation of human breast cancer cells.
Full article

Figure 1
Open AccessArticle
High-Throughput Analysis of 3D Cell Culture Oxygen Consumption Using Sensor Arrays: A Novel Platform for Hypoxia/Normoxia Research
by
Christoph Grün, Cordula Nies, Magdalena Klesen, Enja Schwarz, Jonah ter Haseborg, Cornelius Dettmer, Christian Beyer, Larissa Funk and Eric Gottwald
Organoids 2026, 5(1), 5; https://doi.org/10.3390/organoids5010005 - 6 Feb 2026
Abstract
►▼
Show Figures
Precise control and measurement of the cellular microenvironment, particularly oxygen concentration, are crucial for developing physiologically relevant in vitro models. However, current methods often lack the spatial resolution and throughput needed to investigate complex, oxygen-dependent biological mechanisms in 3D cell cultures. Here, we
[...] Read more.
Precise control and measurement of the cellular microenvironment, particularly oxygen concentration, are crucial for developing physiologically relevant in vitro models. However, current methods often lack the spatial resolution and throughput needed to investigate complex, oxygen-dependent biological mechanisms in 3D cell cultures. Here, we present an advanced platform based on microcavity arrays featuring integrated, ratiometric oxygen sensors, so-called SensoSpheres. A unique bevel design at the cavity entrance enables the non-invasive, real-time measurement of pericellular oxygen concentration and oxygen gradients. We established protocols for generating spheroids from various cell lines (e.g., HepG2, HeLa) and characterized their metabolic responses under precisely controlled hypoxic, normoxic, and hyperoxic conditions. Using a dose–response assay, we demonstrate the platform’s sensitivity in capturing distinct metabolic shifts in response to acetaminophen and cisplatin. Furthermore, we introduce the Oxygen Consumption Recovery Rate (OCRR) as a novel parameter to quantify cellular resilience after exposure to toxic compounds such as cisplatin and acetaminophen. This high-throughput-compatible platform represents a significant methodological advancement, enabling detailed studies of oxygen-dependent cellular processes, drug toxicity, and metabolic adaptation. Its potential for integration into microfluidic systems paves the way for more sophisticated organ-on-chip models, ultimately improving the predictive power of preclinical research.
Full article

Figure 1
Open AccessArticle
A Fully Annotated Hepatoblastoma Tumoroid Biobank Details Treatment-Induced Evolution and Clonal Dynamics in Paediatric Cancer Patients
by
Gijs J. F. van Son, Femke C. A. S. Ringnalda, Markus J. van Roosmalen, Thomas A. Kluiver, Quinty Hansen, Evelien Duiker, Marius C. van den Heuvel, Vincent E. de Meijer, Ruben H. de Kleine, Ronald R. de Krijger, József Zsiros, Weng Chuan Peng, Ruben van Boxtel, Marc van de Wetering, Karin Sanders and Hans Clevers
Organoids 2026, 5(1), 4; https://doi.org/10.3390/organoids5010004 - 18 Jan 2026
Abstract
►▼
Show Figures
Hepatoblastoma (HB) is a paediatric liver malignancy arising from hepatic precursor cells, with >90% of cases harbouring a mutation in exon 3 of CTNNB1. We present a fully genetically characterised HB tumour organoid (tumoroid) biobank, which allows for in vitro studies of
[...] Read more.
Hepatoblastoma (HB) is a paediatric liver malignancy arising from hepatic precursor cells, with >90% of cases harbouring a mutation in exon 3 of CTNNB1. We present a fully genetically characterised HB tumour organoid (tumoroid) biobank, which allows for in vitro studies of disease progression and clonal dynamics in vitro. We established a biobank of 14 tumoroid lines from 9 different patients. Tumours and tumoroids were characterised by whole genome sequencing (WGS) and histology, revealing strong concordance in cell morphology and β-catenin staining. In tumour—tumoroid pairs, identical pathogenic CTNNB1 variants were found, alongside shared copy number alterations (CNAs) and mutations. Variant allele frequency (VAF) was consistently higher in tumoroids, indicating increased tumour purity in vitro. In addition to CTNNB1, we frequently observed ARID1A alterations (single-nucleotide variants [SNVs] or CNAs in 56% of patients), and MYC gains as described previously. In paired pre- and post-treatment samples, we observed a clear increase in mutational load, attributed to a chemotherapy signature. Notably, from one patient, we analysed 4 tumour samples (3 post-treatment) with 4 matching tumoroid lines, all carrying a novel BCL6 mutation and loss of ARID1A. Mutational profiles varied across samples from different locations, suggesting intratumoral heterogeneity and clonal selection during tumoroid derivation. Taken together, this biobank allows detailed analysis of HB tumour biology, including treatment-induced progression and clonal dynamics across temporally and spatially distinct samples.
Full article

Figure 1
Open AccessArticle
Synergistic Overexpression of Sox9, TGFβ1, and Col II Induces Functional Chondrogenesis in hUC-MSCs Using a 3D Culture Approach
by
Shumaila Khalid, Sobia Ekram, Faiza Ramzan, Asmat Salim and Irfan Khan
Organoids 2026, 5(1), 3; https://doi.org/10.3390/organoids5010003 - 14 Jan 2026
Abstract
►▼
Show Figures
Human umbilical cord-derived mesenchymal stem cells (hUC-MSCs) possess the potential for chondrogenic differentiation, offering a promising alternative source for cartilage regeneration. To address the limited availability and expansion capacity of autologous chondrocytes, we investigated the effect of co-overexpression of Sox9, TGFβ1, and type
[...] Read more.
Human umbilical cord-derived mesenchymal stem cells (hUC-MSCs) possess the potential for chondrogenic differentiation, offering a promising alternative source for cartilage regeneration. To address the limited availability and expansion capacity of autologous chondrocytes, we investigated the effect of co-overexpression of Sox9, TGFβ1, and type II collagen (Col II) on the chondrogenic differentiation of hUC-MSCs using both 2D and 3D pellet culture systems. Following transfection, the cells exhibited a chondrocyte-like morphology and a marked downregulation of the stemness marker Stro-1. After 21 days in a 3D pellet culture system, the cells formed cartilage-like tissue characterized by the strong expression of chondrocyte-specific genes (Sox9, TGFβ1, Col II, Aggrecan) along with the significant secretion of sulfated glycosaminoglycans (sGaGs). These effects were attributed to enhanced cell–cell contact and extracellular matrix interactions promoted by the 3D environment. Our findings suggest that genetically modified hUC-MSCs cultured in a 3D pellet system represent a robust in vitro model for cartilage regeneration, with potential applications in transplantation and drug toxicity screening.
Full article

Figure 1
Open AccessReview
Decellularized Extracellular Matrix for Organoids Development and 3D Bioprinting
by
Elena Gkantzou, Alexandro Rodríguez-Rojas, Aleksandra Chmielewska, Barbara Pratscher, Surina Surina, Patricia Freund and Iwan A. Burgener
Organoids 2026, 5(1), 2; https://doi.org/10.3390/organoids5010002 - 8 Jan 2026
Cited by 2
Abstract
►▼
Show Figures
Organoids are three-dimensional multicellular structures that mimic key aspects of native tissues consisting ideal tools to study organ development and pathophysiology when incorporated in customized bioscaffolds. In vivo, the extracellular matrix (ECM) maintains tissue integrity and regulates cell adhesion, migration, differentiation, and survival
[...] Read more.
Organoids are three-dimensional multicellular structures that mimic key aspects of native tissues consisting ideal tools to study organ development and pathophysiology when incorporated in customized bioscaffolds. In vivo, the extracellular matrix (ECM) maintains tissue integrity and regulates cell adhesion, migration, differentiation, and survival through biochemical and mechanical signals. Tissue-derived decellularized extracellular matrix (dECM) can preserve organ-specific biochemical signals and cell-adhesive motifs, creating a bioactive environment that supports physiologically relevant organoid growth. 3D bioprinting technology marks a transformative phase in organoid research by enhancing the structural and functional complexity of organoid models and expanding their application in pharmacology and regenerative medicine. These systems enhance tissue modeling and drug testing while adhering to the principles of animal replacement, reduction, and refining (3Rs) in research. Remaining challenges include donor variability, limited mechanical stability, and the lack of standardized decellularization protocols that can be addressed by adopting quality and safety metrics. The combination of dECM-based biomaterials and 3D bioprinting holds great potential for the development of human-relevant, customizable, and ethically sound in vitro models for regenerative medicine and personalized therapies. In this review, we discuss the latest (2021–2025) developments in applying extracellular matrix bioprinting techniques to organoid technology, presenting examples for the most commonly referenced organoid types.
Full article

Graphical abstract
Open AccessArticle
Permeability of the Blood–Brain Barrier (BBB) to Nanoparticles, Bacteria and Phages Studied in BBB Organoids Under Normoxic and Hypoxic Conditions In Vitro
by
Kathrin Kostka-Wirtz, Nataniel Białas, Ivanna Kostina and Matthias Epple
Organoids 2026, 5(1), 1; https://doi.org/10.3390/organoids5010001 - 2 Jan 2026
Abstract
►▼
Show Figures
Organoids consisting of primary human cells, i.e., astrocytes, pericytes, and endothelial cells, form a functional blood–brain barrier (BBB) in vitro. The ability of FITC-dextran (70 kDa), calcium phosphate nanoparticles (100 nm), Escherichia coli bacteria (2 µm), and MS2 coliphages (27 nm, a model
[...] Read more.
Organoids consisting of primary human cells, i.e., astrocytes, pericytes, and endothelial cells, form a functional blood–brain barrier (BBB) in vitro. The ability of FITC-dextran (70 kDa), calcium phosphate nanoparticles (100 nm), Escherichia coli bacteria (2 µm), and MS2 coliphages (27 nm, a model for viruses) to penetrate the BBB under normoxic and hypoxic conditions (2.5% oxygen) for up to 12 days was assessed by fluorescence microscopy and confocal laser scanning microscopy. All agents were fluorescently labeled to trace them inside the organoids. Under normoxia, FITC-dextran, calcium phosphate nanoparticles, E. coli bacteria and MS2 coliphages did not penetrate the BBB. However, oxygen deficiency (hypoxia) triggered the penetration of the BBB by FITC-dextran and E. coli cells. This was underscored by a strong hypoxic center inside the organoids that developed in the presence of E. coli bacteria.
Full article

Graphical abstract
Open AccessEditorial
Organoids: Promoting Innovation in Organoid Technology for Basic Research and Therapeutic Applications
by
Süleyman Ergün and Philipp Wörsdörfer
Organoids 2025, 4(4), 33; https://doi.org/10.3390/organoids4040033 - 8 Dec 2025
Cited by 1
Abstract
►▼
Show Figures
Over the past decade, organoids representing a wide range of tissues have been developed, with increasing efforts to enhance their complexity, maturity, and resemblance to the corresponding native organs [...]
Full article
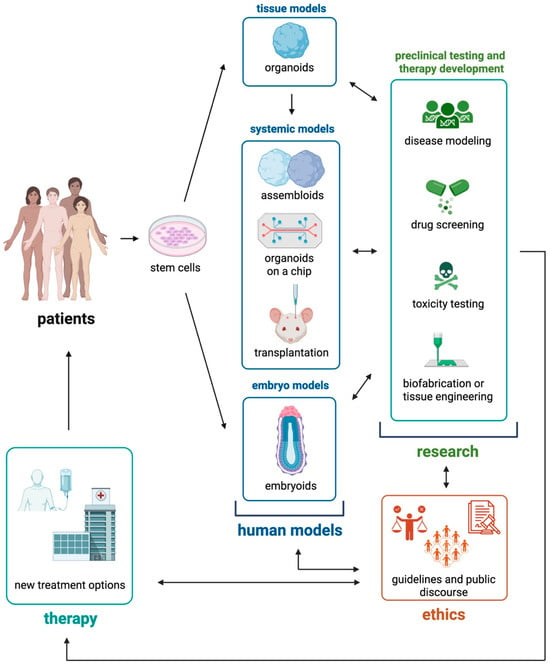
Figure 1
Open AccessArticle
Evaluation of a Gene Expression-Based Machine Learning Classifier to Discriminate Normal from Cancer Gastric Organoids
by
Daniel Skubleny, Hasnaien Ahmed, Sebastiao N. Martins-Filho, David Ross McLean, Daniel E. Schiller and Gina R. Rayat
Organoids 2025, 4(4), 32; https://doi.org/10.3390/organoids4040032 - 5 Dec 2025
Abstract
►▼
Show Figures
Three-dimensional cell model systems such as tumour organoids allow for in vitro modelling of self-organized tissue with functional and histologic similarity to in vivo tissue. However, there is a need for standard protocols and techniques to confirm the presence of cancer within organoids
[...] Read more.
Three-dimensional cell model systems such as tumour organoids allow for in vitro modelling of self-organized tissue with functional and histologic similarity to in vivo tissue. However, there is a need for standard protocols and techniques to confirm the presence of cancer within organoids derived from tumour tissue. The aim of this study was to assess the utility of a Nanostring gene expression-based machine learning classifier to determine the presence of cancer or normal organoids in cultures developed from both benign and cancerous stomach biopsies. A prospective cohort of normal and cancer stomach biopsies were collected from 2019 to 2022. Tissue specimens were processed for formalin-fixed paraffin-embedding (FFPE) and a subset of specimens were established in organoid cultures. Specimens were labelled as normal or cancer according to analysis of the FFPE tissue by two pathologists. The gene expression in FFPE and organoid tissue was measured using a 107 gene Nanostring codeset and normalized using the Removal of Unwanted Variation III algorithm. Our machine learning model was developed using five-fold nested cross-validation to classify normal or cancer gastric tissue from publicly available Asian Cancer Research Group (ACRG) gene expression data. The models were externally validated using the Cancer Genome Atlas (TCGA), as well as our own FFPE and organoid gene expression data. A total of 60 samples were collected, including 38 cancer FFPE specimens, 5 normal FFPE specimens, 12 cancer organoids, and 5 normal organoids. The optimal model design used a Least Absolute Shrinkage and Selection Operator model for feature selection and an ElasticNet model for classification, yielding area under the curve (AUC) values of 0.99 [95% CI: 0.99–1], 0.90 [95% CI: 0.87–0.93], and 0.79 [95% CI: 0.74–0.84] for ACRG (internal test), FFPE, and organoid (external test) data, respectively. The performance of our final model on external data achieved AUC values of 0.99 [95% CI: 0.98–1], 0.94 [95% CI: 0.86–1], and 0.85 [95% CI: 0.63–1] for TCGA, FFPE, and organoid specimens, respectively. Using a public database to create a machine learning model in combination with a Nanostring gene expression assay allows us to allocate organoids and their paired whole tissue samples. This platform yielded reasonable accuracy for FFPE and organoid specimens, with the former being more accurate. This study re-affirms that although organoids are a high-fidelity model, there are still limitations in validating the recapitulation of cancer in vitro.
Full article

Figure 1
Open AccessReview
Organoids as a Tool for Assessing Drinking Water Safety and Guidelines Relevance
by
Roberto Coppo and Edoardo Bertone
Organoids 2025, 4(4), 31; https://doi.org/10.3390/organoids4040031 - 4 Dec 2025
Abstract
►▼
Show Figures
Ensuring access to safe drinking water is a fundamental public health priority, yet the growing diversity of contaminants demands more human-relevant toxicity assessment frameworks. Conventional models based on immortalized cell lines or sentinel species, while informative, lack the tissue complexity and inter-individual variability
[...] Read more.
Ensuring access to safe drinking water is a fundamental public health priority, yet the growing diversity of contaminants demands more human-relevant toxicity assessment frameworks. Conventional models based on immortalized cell lines or sentinel species, while informative, lack the tissue complexity and inter-individual variability required to capture realistic human responses. Organoids, three-dimensional epithelial structures derived from adult or pluripotent stem cells, retain the genomic, histological, and functional characteristics of their original tissue, enabling assessment of contaminant-induced toxicity, short-term peak exposures, and inter-donor variability within a single system. This study examined whether current international drinking water guidelines remain protective or if recent organoid-based findings reveal toxicity at differing concentrations. Comparative synthesis indicates that per- and polyfluoroalkyl substances (PFAS) often display organoid toxicity at concentrations above current thresholds, suggesting conservative guidelines, whereas most metals are properly regulated. However, some metals exhibit toxicity at concentrations that include levels below guideline values, highlighting the need for further investigation. Emerging contaminants, including pesticides, nanoparticles, microplastics, and endocrine disruptors, induce adverse effects at environmentally relevant concentrations, despite limited or absent regulatory limits. Integrating organoid-based toxicology with high-frequency monitoring and dynamic exposure modeling could refine water quality guidelines and support adaptive regulatory frameworks that better reflect real-world exposure patterns and human diversity.
Full article

Figure 1
Open AccessReview
AI-Enhanced Patient-Derived Cancer Organoids: Integrating Machine Learning for Precision Oncology
by
Elisa Heinzelmann and Francesco Piraino
Organoids 2025, 4(4), 30; https://doi.org/10.3390/organoids4040030 - 3 Dec 2025
Cited by 3
Abstract
►▼
Show Figures
Cancer remains a leading cause of mortality worldwide. Patient-derived organoids (PDOs) are three-dimensional (3D) cultures that recapitulate tumor histology, genetics, and cellular heterogeneity, providing physiologically relevant preclinical models. Integrating PDOs with artificial intelligence (AI) and machine learning (ML) enables scalable analysis of high-dimensional
[...] Read more.
Cancer remains a leading cause of mortality worldwide. Patient-derived organoids (PDOs) are three-dimensional (3D) cultures that recapitulate tumor histology, genetics, and cellular heterogeneity, providing physiologically relevant preclinical models. Integrating PDOs with artificial intelligence (AI) and machine learning (ML) enables scalable analysis of high-dimensional datasets, including imaging, transcriptomics, proteomics, and pharmacological readouts. These approaches support prediction of drug sensitivity, biomarker discovery, and patient stratification. Recent advances—such as deep learning (DL), transfer learning, federated learning, and self-supervised learning—enhance phenotypic profiling, cross-institutional model training, and translational prediction. In this review, we summarize the current state of AI-driven PDO research, highlighting methodological approaches, preclinical and clinical applications, challenges, and emerging trends. We also propose strategies for standardization, validation, and multi-modal integration to accelerate patient-specific therapeutic strategies.
Full article

Graphical abstract
Open AccessCommentary
Organoid Intelligence: Can We Separate Intelligent Behavior from an Intelligent Being?
by
Daniel Montoya
Organoids 2025, 4(4), 29; https://doi.org/10.3390/organoids4040029 - 18 Nov 2025
Abstract
As brain organoids and organoid-based computational models grow in complexity, they increasingly exhibit electrophysiological patterns consistent with plasticity and information processing. This article explores a central question at the intersection of neuroscience, synthetic biology, and philosophy of mind: Can intelligent behavior be meaningfully
[...] Read more.
As brain organoids and organoid-based computational models grow in complexity, they increasingly exhibit electrophysiological patterns consistent with plasticity and information processing. This article explores a central question at the intersection of neuroscience, synthetic biology, and philosophy of mind: Can intelligent behavior be meaningfully separated from an intelligent being? In other words, can adaptive, goal-directed behavior exist independently of subjective awareness—a question that challenges conventional definitions of cognition and consciousness. Drawing from neuroscience, artificial intelligence, and philosophy, I propose a tiered framework based on neural complexity and environmental responsiveness. This includes a simple level analysis and a context-sensitive benchmark for evaluating intelligence in organoid systems without presupposing sentience. Ethical and ontological implications are also addressed, particularly the risk of anthropomorphizing synthetic cognition and the importance of developing context-aware definitions of intelligence. By distinguishing functional sophistication from subjective experience, the framework aims to guide responsible scientific inquiry while clarifying the boundaries of synthetic cognition.
Full article
(This article belongs to the Special Issue Advances in Organoid Technology: Bridging the Gap between Research and Therapy)
Open AccessReview
Combining Proteomics and Organoid Research to Unravel the Multifunctional Complexity of Kidney Physiology Enhances the Need for Controlled Organoid Maturation
by
Kathrin Groeneveld and Ralf Mrowka
Organoids 2025, 4(4), 28; https://doi.org/10.3390/organoids4040028 - 14 Nov 2025
Cited by 1
Abstract
This review aims to highlight how the study of kidney organoids combined with proteomic analysis can deepen our understanding of renal physiology and disease. Proteomics quantifies proteins in a sample, allowing us to determine which proteins are present, how abundant they are, and
[...] Read more.
This review aims to highlight how the study of kidney organoids combined with proteomic analysis can deepen our understanding of renal physiology and disease. Proteomics quantifies proteins in a sample, allowing us to determine which proteins are present, how abundant they are, and how they are modified. These data may reveal the pathways that are active in the kidney organoids and how they change in disease, helping to pinpoint candidate biomarkers. Kidney organoids are three-dimensional structures derived from induced pluripotent stem cells (iPS) that recapitulate many architectural and functional features of the adult organ. Because they can be generated in large numbers under defined conditions, organoids provide a promising platform for testing how genetic mutations, environmental stresses, or drugs affect kidney development and pathology. When proteomic profiles are obtained from mature organoids, researchers can directly link protein-level changes to phenotypic outcomes observed in the model. This integration makes it possible to map disease-related networks at the molecular level and to assess the impact of therapeutic interventions in a system that more closely resembles human kidney tissue than traditional cell lines. A current limitation is that many kidney organoids do not reach the full maturation seen in vivo; they often lack complete segmental differentiation and the functional robustness of adult nephrons. Improving the maturation state of organoids will be essential for accurately modeling chronic kidney diseases and for translating findings into clinically relevant therapies.
Full article
(This article belongs to the Special Issue Advances in Organoid Technology: Bridging the Gap between Research and Therapy)
►▼
Show Figures

Figure 1
Open AccessArticle
Improved Differentiation of Human Retinal Organoids Producing Mature Photoreceptors with Budding Calyceal Process-like Structure and Usher Protein Expression
by
Tokiyoshi Matsushita, Takahiro Matsuyama, Takayuki Kawasaki and Fumiaki Uchiumi
Organoids 2025, 4(4), 27; https://doi.org/10.3390/organoids4040027 - 6 Nov 2025
Abstract
Human retinal organoids derived from pluripotent stem cells represent a robust in vitro model for investigating retinal development and disease mechanisms of retinal disorders. However, achieving structural maturation that faithfully recapitulates the intricate architecture of photoreceptors within a feasible and cost-efficient culture timeframe
[...] Read more.
Human retinal organoids derived from pluripotent stem cells represent a robust in vitro model for investigating retinal development and disease mechanisms of retinal disorders. However, achieving structural maturation that faithfully recapitulates the intricate architecture of photoreceptors within a feasible and cost-efficient culture timeframe remains a significant challenge. Here, we present an optimized differentiation protocol that enables the generation of retinal organoids exhibiting advanced photoreceptor maturation within 140 days. Photoreceptors in the retinal organoids displayed compartmentalized architecture, including distinct inner and outer segments and connecting cilia. Notably, we observed the emergence of budding calyceal process-like structures—a feature not previously emphasized in photoreceptors derived from pluripotent stem cells. These results suggest that our protocol may promote advanced photoreceptor maturation within a relatively shortened culture period. Thus, this method could serve as a useful model for investigating retinal development and related pathologies, building upon previous protocols.
Full article
(This article belongs to the Special Issue The Current Applications and Potential of Stem Cell-Derived Organoids)
►▼
Show Figures

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Cells, JCM, Organoids, JMP
Novel Discoveries in Oncology 2nd Edition
Topic Editors: Michela Campolo, Giovanna Casili, Alessia Filippone, Marika LanzaDeadline: 20 June 2026
Topic in
CIMB, IJMS, JCDD, Organoids, Biomedicines
Molecular and Cellular Mechanisms of Heart Disease
Topic Editors: Pasi Tavi, Ebru Arioglu-InanDeadline: 31 December 2026

Special Issues
Special Issue in
Organoids
Advances in Organoid Technology: Bridging the Gap between Research and Therapy
Guest Editors: Elizabeth Vincan, Ramanuj DasGupta, Somponnat Sampattavanich, Joao FerreiraDeadline: 15 May 2026
Special Issue in
Organoids
Modeling Lung Regeneration and Injury Responses Using Organoids and Engineered Niches
Guest Editor: Tristan FrumDeadline: 31 May 2026
Special Issue in
Organoids
Organs-on-Chips and Organoids: From Disease Modeling to Advanced Therapeutics
Guest Editor: Gautam MahajanDeadline: 30 November 2026
Special Issue in
Organoids
From Development to Degeneration: Advances in Stem Cell and Organoid Models of the Nervous System
Guest Editor: Elena CocciaDeadline: 30 November 2026