Abstract
Plasma-activated water (PAW) has gained attention across agricultural, medical, cosmetic, and sterilization fields due to its production of reactive oxygen and nitrogen species (ROS and RNS). Although PAW has been primarily explored for seed germination and sterilization in agriculture, its role as a nutrient source and physiological regulator remains less understood. In this study, PAW generated by a surface dielectric barrier discharge (SDBD) system contained approximately 1000 ppm nitrate (NO3−) and was designated as PAW1000. Diluted PAW solutions were applied to sprout crops—wheat (Triticum aestivum), barley (Hordeum vulgare), radish (Raphanus sativus), and broccoli (Brassica oleracea var. italica)—grown under hydroponic and soil-based conditions. PAW100 and PAW200 treatments enhanced growth, increasing fresh biomass by up to 26%, shoot length by 22%, and root length by 18%, depending on the species. In silico analysis identified nitrogen-responsive transcripts among several autophagy-related genes. Consistent with this, fluorescence microscopy of Arabidopsis thaliana GFP-StATG8 lines revealed increased autophagosome formation following PAW treatment. The growth-promoting effect of PAW was diminished in atg4 mutants, indicating that autophagy contributes to plant responses to PAW-derived ROS and RNS. Together, these findings demonstrate that diluted PAW generated by SDBD enhances biomass accumulation in sprout crops, and that autophagy plays a regulatory role in mediating PAW-induced physiological responses.
1. Introduction
Climate change has become a major global concern due to its effects on food security, the increasing frequency of extreme weather events, and the depletion of natural resources [1]. In response, indoor farming systems have gained attention, accompanied by the development of sustainable agricultural technologies [2,3,4,5]. Within smart farming environments, improving crop productivity through advanced materials such as nano-biofertilizers and bioactive agrochemicals has become an important objective [6,7]. Plant growth in enclosed systems is strongly influenced by environmental factors, including photosynthetically active radiation, atmospheric composition, and water availability, as well as temperature, soil properties, and nutrient status [8]. Excessive fertilizer use can lead to serious environmental issues, including groundwater contamination and nutrient runoff. Consequently, precision agriculture is being actively explored for its potential to enhance crop growth while reducing fertilizer-related pollution [9,10]. Meeting crop nutritional demands while limiting environmental impact has therefore become an urgent challenge. New strategies that improve both biotic stress resistance and abiotic stress tolerance—without relying on genetic modification—are increasingly needed.
Plasma occurs naturally in events such as thunderstorms and contributes reactive oxygen species (ROS) and reactive nitrogen species (RNS) to rainwater through atmospheric energy conversion [11]. When water is directly exposed to plasma, plasma-activated water (PAW) is produced, containing various ROS and RNS generated during this process [11]. PAW has attracted interest for its capacity to enhance plant growth by mitigating biological stress. Plasma treatment can effectively eliminate pathogens [12], sterilize seed surfaces, and induce controlled oxidative stress, ultimately improving seed germination and early growth [13].
Recent studies have shown that increased crop productivity can be achieved through the application of PAW alone, without direct plasma exposure on plant tissues [14]. PAW has been investigated in several crops, including pea, rice, and maize [15,16,17,18], and has been associated with increases in plant height—for example, approximately 11.9% in tomato and 5% in cabbage—although the effects vary by species [19]. However, most previous work has focused primarily on growth-related outcomes such as germination and biomass, with relatively limited attention to the underlying biological mechanisms.
ROS and RNS (RONS) play key roles in regulating autophagy in plant cells [20]. PAW irrigation in tomato seedlings has been reported to stimulate the expression of antioxidant enzymes such as superoxide dismutase, catalase, and glutathione S-transferase, as well as defense-related and MAPK signaling genes [21,22]. These responses indicate that PAW-derived RONS can act as signaling molecules that fine-tune cellular redox homeostasis and defense hormone pathways, depending on exposure duration and RONS concentration. Collectively, these findings suggest that PAW triggers controlled oxidative signaling events that activate plant defense and antioxidant mechanisms. However, the downstream regulatory processes linking PAW-induced redox signaling to specific cellular pathways, including autophagy, remain largely unexplored. Nitric oxide (NO) reacts with ROS to form secondary intermediates that modify proteins through mechanisms such as S-nitrosylation and tyrosine nitration, thereby altering protein function [23]. Recent findings suggest that ATG8ylation, the lipidation of ATG8, is closely linked to redox-dependent autophagy regulation, with ROS acting as an upstream signal that promotes ATG8 conjugation to single-membrane structures, including the tonoplast [24]. These redox-mediated modifications influence the activity of autophagy-related proteins such as ATG8 and its interacting partners, thereby modulating autophagic flux [25]. Additionally, NO has been reported to regulate the TOR and SnRK1 pathways—central components of nutrient and stress signaling—both of which influence autophagy under nutrient-limited or oxidative stress conditions [26]. Thus, appropriate concentrations of ROS and RNS in PAW may affect plant growth and development, at least in part, through autophagy-related regulatory pathways.
Sprout vegetables such as wheat, barley, radish, and broccoli are widely consumed and can be readily cultivated in smart-farm environments [27,28]. They contain high levels of bioactive compounds, including antioxidants and vitamins [27,29,30], and are characterized by rapid growth and high nutritional value. For these reasons, sprouts were selected in this study to evaluate the effects of diluted PAW under both hydroponic and soil-based cultivation systems.
In this work, PAW was generated using non-thermal plasma technology, and PAW1000—diluted based on nitrate concentration, the predominant reactive species—was applied to several sprout crops to assess its potential utility. The study also provides preliminary evidence that PAW may exert physiological effects through the activation of autophagy-related pathways. This foundational research contributes to the identification of environmentally sustainable agricultural inputs. Defining the physicochemical characteristics of PAW suitable for specific crops may offer a strategic direction for advancing eco-friendly agricultural practices in the future.
2. Materials and Methods
2.1. PAW Production and Anion Analysis
PAW production involved using a surface dielectric barrier discharge (SDBD) in closed containers [31]. In brief, the SDBD device was an SDBD reactor equipped with two electrodes on top, operating at an input power of 10 W and an actuation frequency of 17 kHz. The Ionized air generated by the plasma was dissolved in deionized water (D.W.) with the aid of a fan (Aone 15-LED 120). PAW1000 refers to plasma-activated water in which 2 L of deionized water (D.W.) was exposed to high-intensity plasma for approximately 230 min, resulting in a nitrate concentration of around 1000 ppm. The corresponding ionic composition, including nitrate levels, was analyzed using ion chromatography. Subsequently, the PAW1000 stock solution, containing 1000 ppm of nitrate, was diluted with triple-distilled water to prepare PAW solutions containing 1, 20, 100, 200, 400, and 500 ppm of nitrate. To analyze the chemical properties of PAW, the anion content was measured using ion chromatography. The optical emission spectrum (OES) of the plasma was recorded in the 200–600 nm range a Maya 2000 Pro spectrometer (Ocean Optics, Dunedin, FL, USA) with a 1 s integration time, averaged over 50 measurements. Plasma-activated water (PAW) was chemically characterized, and anion concentrations were determined using an ICS-2100 ion chromatography system (Thermo Fisher Scientific, Sunnyvale, CA, USA) following previously established procedures [32,33].
2.2. Plant Materials and Growth Conditions
Wheat, barley, radish, and broccoli sprout seeds were commercially purchased. All seeds were initially germinated in D.W. For hydroponic cultivation, the germinated seedlings were transferred to the respective treatment solutions (D.W. or PAW) and subsequently monitored for their growth responses. For soil-based cultivation, the seeds were likewise germinated in D.W. and then transplanted into soil, after which the experiments were conducted. All experiments were conducted using more than 40–100 sprout seeds per replicate, with three biological replicates for each treatment. A hydroponic system was established using handmade cultivation trays, consisting of plastic containers measuring 30 cm in diameter and 15 cm in height, designed to support optimal hydroponic growth conditions. The experimental setup ensured that the roots were submerged in approximately 0.5 cm of the respective solution. The plants were cultivated in a growth room and a growth chamber equipped with white LED lights providing a 16 h light/8 h dark photoperiod at an intensity of ~150 µmol m−2 s−1, at approximately 22 °C, and with a relative humidity of 60–70%. The sprout seeds were selected and grown in a hydroponic system. To prevent contamination, treated D.W. or PAW was replaced daily with an equal volume of fresh solution to maintain consistent treatment conditions. Measurements of the total length and weight of the sprouts were taken after 7 days.
Sixty sprouts were selected for soil cultivation. The sprouts were grown in a commercial potting substrate (Baroker, Seoul Bio Co., Ltd., Eumseong, Republic of Korea) composed of peat moss, perlite, and vermiculite mixed in a ratio of approximately 6:2:2 (v/v/v). Approximately 0.25 L of soil was uniformly filled into containers measuring 7 cm × 7 cm × 6 cm. The length and weight of the sprouts were measured after 14 days. We purchased Bio Garden Liquid Fertilizer II (Plant Pharmacy., Seoul, Republic of Korea). A Type IV compound fertilizer (N–P2O5–K2O = 6–8–5) was used, containing a total of 19% primary nutrients: 6% nitrogen (N), 8% available phosphate (P2O5), and 5% soluble potassium (K2O). This water-soluble formulation is commonly used in hydroponic and container-grown crop systems and may also include trace elements depending on the intended application. After seed germination, sprout crops in the 0.02% diluted liquid fertilizer, D.W., and PAW200 groups were irrigated every three days with 50 mL of their respective solutions per pot. The liquid fertilizer was applied using the same procedure as for PAW treatment.
2.3. Transgenic Plant Generation and Analysis
For the constitutive expression of the potato autophagy-related gene StATG8-1.1, the N-terminal GFP Gateway vector was used, and the 35S::GFP-StATG8.1-1 vector was obtained from Dr. Yasin Dagdas (Gregor Mendel Institute of Molecular Plant Biology) [34]. To generate Arabidopsis transgenic plants using modified floral dip method [35,36], Agrobacterium tumefaciens GV3101 carrying 35S::GFP-StATG8.1-1 were used, and selection of the T3 generation was performed using the antibiotic kanamycin. and qRT-PCR was performed using StATG8-1.1-specific primers: Forward (For): GACAATGTGCTACCGCCAAC and Reverse (Rev): TCGCAAATTATTCGGACACAG. The Arabidopsis transgenic plants were analyzed by confocal microscopy. Following a 1 h incubation with PAW20 or D.W, GFP signals were visualized using an LSM 910 confocal laser scanning microscope (Carl Zeiss, Jena, Germany) and a K1-Fluo confocal microscope (Nanoscope Systems, Daejeon, Republic of Korea).
The Arabidopsis Col-0 ecotype T-DNA insertional mutant atg7-2 was obtained from Prof. Taijoon Chung (Pusan National University) [37] and Ws ecotype T-DNA insertion mutant atg7-1 was obtained from The European Arabidopsis Stock Centre (NASC). The qRT-PCR was performed to determine the transcript levels of ATG7 using the following primers: forward primer 5′-CGGAGTCCTACAACACCCAC-3′ and reverse primer 5′-AAGCGGTACAGCTGTTGGAA-3′.
To determine the effect of PAW on root growth of autophagy mutant, we conducted a root growth assay. After germination of atg7-1 or atg7-2 mutant seeds in 1/2MS medium, seedlings of the atg7-1 or atg7-2 mutant with consistent lengths were transferred to 0.8% agar medium made of PAW40 or D.W. The root growth was then observed and confirmed over a period of 5 days.
2.4. In Silco Autophagy-Related Gene Expression Analysis in Arabidopsis and Wheat
The Genevestigator program was used to analyze the in silico data. We selected the Affymetrix microarray dataset (ATH1-12) for ATG genes present in the data from Arabidopsis. To investigate nutritional responses in ATG genes present in wheat, data were acquired through the establishment of the mRNA sequencing dataset (WHEAT_GL-8). The response of the Arabidopsis and wheat ATG genes to nutrition-related stimuli was measured as Log2 ratio.
2.5. qRT-PCR Analysis Under PAW20 Treatment in Arabidopsis thaliana
For quantitative real-time PCR (qRT-PCR) analysis, Arabidopsis Col-0 seedlings were treated with PAW20 for 1 h to RNA extraction. Total RNA was isolated using a standard TRIzol-based method, and complementary DNA (cDNA) was synthesized from 1 µg of total RNA using the PrimeScript™ cDNA Synthesis Kit (Takara Bio, Kusatsu, Shiga, Japan) according to the manufacturer’s instructions. qRT-PCR was performed using a SYBR Green qPCR kit (Sigma-Aldrich, Darmstadt, Germany). Gene-specific primers were designed to amplify ATG family members, and ACT2 was used as an internal reference gene for normalization. The primer sequences were as follows: ATG3: forward 5′-GGAGGAGTTCGATGAGGCTG-3′, reverse 5′-CGATTGTGACCGTTTTACGAGC-3′. ATG5: forward 5′-AAGTTGTGCCTGAGACGGAG-3′, reverse 5′-TTACCACCCACGAAAACGGT-3′. ATG8C: forward 5′-TAGGGGAAACGAGAAGCCCT-3′, reverse 5′-CATCATTGCAGCAGTTGGAGG-3′. ATG10: forward 5′-AGGAAGGTTACTTGTCGCTGG-3′, reverse 5′-TTCTCCACCGCTGCAGTATC-3′. ACT2 (At3g18780): forward 5′-CCGCTCTTTCTTTCCAAGCTCAT-3′, reverse 5′-GGGTTAAGAGGAGCCTCGGT-3′. Relative expression levels of target genes were normalized to ACT2 expression, and fold changes were calculated [38].
2.6. Quantification of Autophagic Bodies in GFP–ATG8 Transgenic Lines Under Different PAW Concentrations
To evaluate the effect of PAW on autophagosome formation, GFP–ATG8-1.1 transgenic Arabidopsis thaliana seedlings were treated with D.W., PAW20, or PAW50 for 1 h. After treatment, root cells of 7-day-old seedlings were observed using a confocal laser scanning microscope (Carl Zeiss LSM910 and K1-Fluo). For GFP-tagged autophagy markers, excitation is typically achieved using a 488 nm argon laser, and fluorescence emission is collected within the 500–550 nm range. GFP-ATG8 puncta appear as distinct cytoplasmic fluorescent dots, representing autophagosome structures. Autophagic bodies displaying punctate GFP-ATG8 fluorescence were visualized and quantified. Image analysis was performed using ImageJ software (version 1.54p; National Institutes of Health, Bethesda, MD, USA) [39] to determine the number of GFP–ATG8 puncta per cell. For each treatment condition, ten independent seedlings (n = 10) were analyzed, and all experiments were conducted with three biological replicates.
2.7. Statistical Analysis
All experiments were repeated at least three times with similar results. Data are presented as means ± standard deviation (SD). Statistical significance was determined using unpaired two-tailed Student’s t-test (GraphPad Prism 9.0, San Diego, CA, USA). Significance levels were denoted as p < 0.05 (*), p < 0.01 (**), and p < 0.001 (***).
3. Results
3.1. Applicability of RNS-Rich Diluted PAW to Eco-Friendly Agriculture
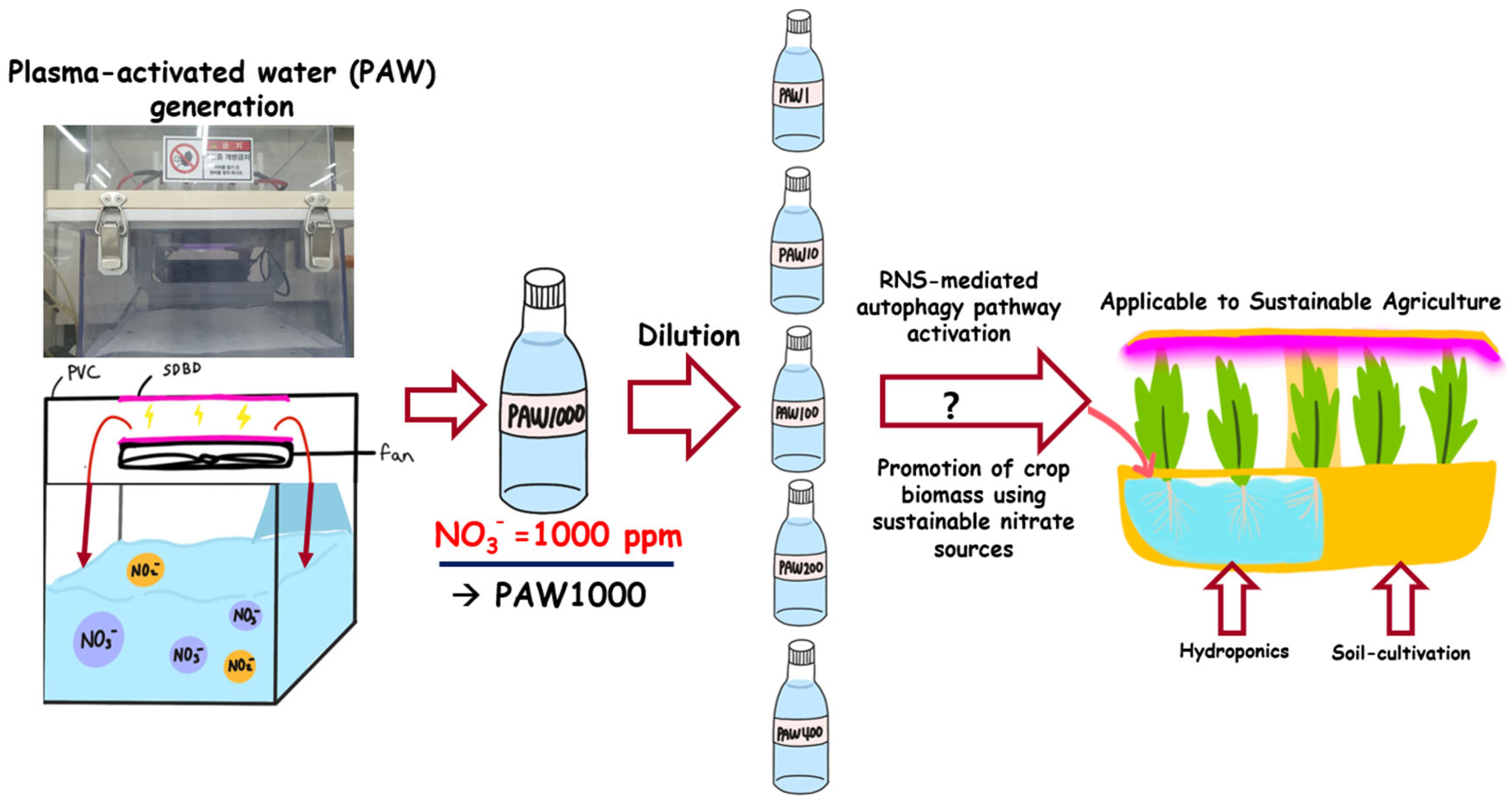
In previous studies, PAW application in agriculture has been extensively explored. The excessive presence of ROS and RNS depends on the plasma-generating device and processing time. In this study, we utilized a PAW-generating device that produces RNS-rich, ROS-excluded solutions [31]. Initially, plasma treatment was employed to generate conditions with 1000 ppm of RNS, particularly NO3−, resulting in a solution named PAW1000. PAW1000 was used as a stock PAW solution containing approximately 1000 ppm nitrate, and dilutions prepared on the basis of nitrate concentration were designated as PAW400, PAW200, and PAW100, as detailed in Table 1. In addition, when the pH of all PAW solutions was adjusted to 5.6 prior to plant treatment, no differences in the constituents present in the PAW were observed. According to ion analysis, nitrate was present at the highest concentration in PAW1000, reaching approximately 1045 ppm, while nitrite (NO2−) was detected at 4.5 ppm, chloride (Cl−) at 2.1 ppm, and fluoride (F−) at 1 ppm. Furthermore, other specific ROS-related factors are not present [31,40]. Our initial objective is to generate a raw PAW solution with a high concentration of RNS, as illustrated in Figure 1. We aim to determine the optimal PAW dilution ratio and subsequently apply it to small-scale home crop cultivation or smart farms [41]. Sprout vegetables have been selected as the most suitable crop for this purpose.

Table 1.
PAW ion analysis.
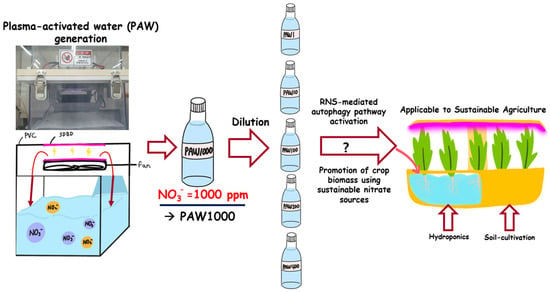
Figure 1.
The use of diluted PAW for sprout cultivation. In a plasma device based on SDBD, distilled water (D.W.) was placed in a PVC chamber and subjected to plasma processing. The resulting plasma-activated water (PAW) contains approximately 1000 ppm of NO3− and does not contain factors involved in reactive oxygen species (ROS). After adjusting the pH to 5.6, which is suitable for plant growth, PAW1000 was diluted as a stock solution using D.W to make PAW500, PAW400, PAW200, PAW100, PAW20, and PAW1. The growth impact of each PAW was evaluated when sprouts were grown hydroponically or in soil. This positive effect on sprout growth can be exploited in smart farming.
3.2. Enhanced Growth of Wheat and Barley Sprouts Was Observed When Treated with PAW100 and PAW200
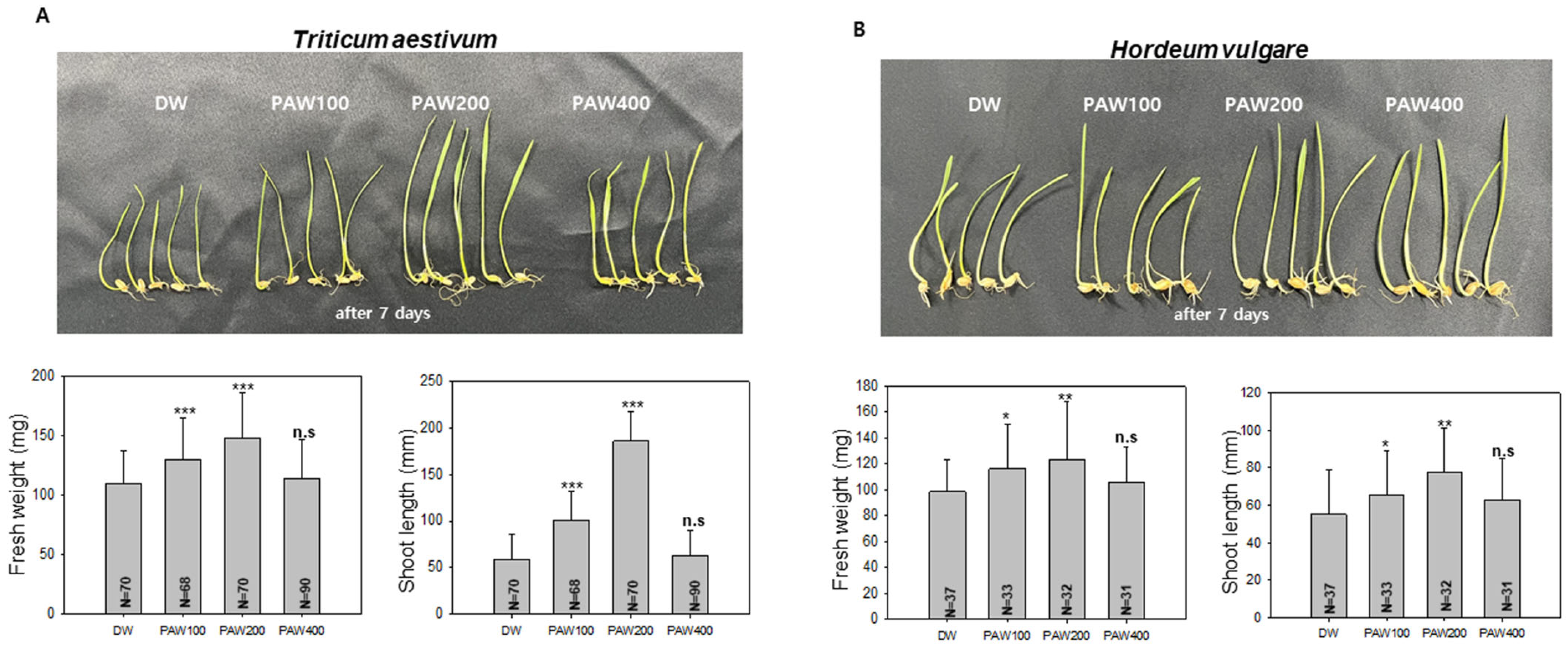
As shown in the preliminary experiment (Figure S1), treatment with the stock solution PAW1000 resulted in inhibited plant growth, due to excessive nitrogen content. Based on this observation, we subsequently investigated the effects of diluted PAW treatments (PAW400, PAW200, and PAW100) to identify an appropriate concentration for promoting plant growth. As shown in Figure 2A, the fresh weight of T. aestivum sprouts grown in PAW200 increased by about 40% and the sprout length increased by about 110% compared to sprouts grown in D.W. In PAW100, the fresh weight increased by approximately 18%, and the shoot length increased by about 30% compared with D.W. control. While there was a notable difference in the growth rate of wheat and barley, a similar trend was observed in barley irrigated with PAW200, showing a significant increase in both fresh weight and shoot length (Figure 2B). This suggests that the impact of diluted PAW is influenced by nitrogen source content and type of sprout crop.
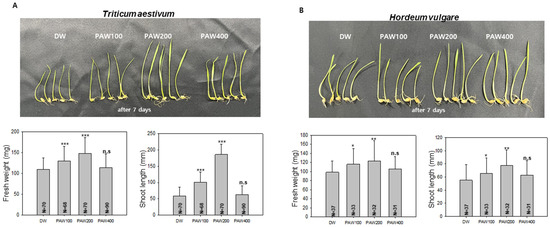
Figure 2.
Optimal growth of Triticum aestivum (wheat) and Hordeum vulgare (barley) sprouts using diluted PAW. (A). Wheat sprout growth in diluted PAW was assessed by cultivating 68 to 90 sprouts hydroponically for 7 days. Measurements of total fresh weight and shoot length were then taken. A t-test was conducted using the values measured in D.W. as a control group. The asterisk indicates significantly different (*** p < 0.001) from the control group and n.s. indicates no statistically significant difference. (B). Barley sprouts (31 to 37) were cultivated hydroponically, and measurements were taken after 7 days. A t-test was performed using the values measured in D.W. as a control group. Data are presented as means ± SD. The asterisk indicates statistical significance compared with the control group, with * p < 0.05 and ** p < 0.01. These experiments were conducted at least three times.
3.3. Irrigation with All Diluted PAWs Resulted in Improved Growth for Both Broccoli and Radish Sprouts
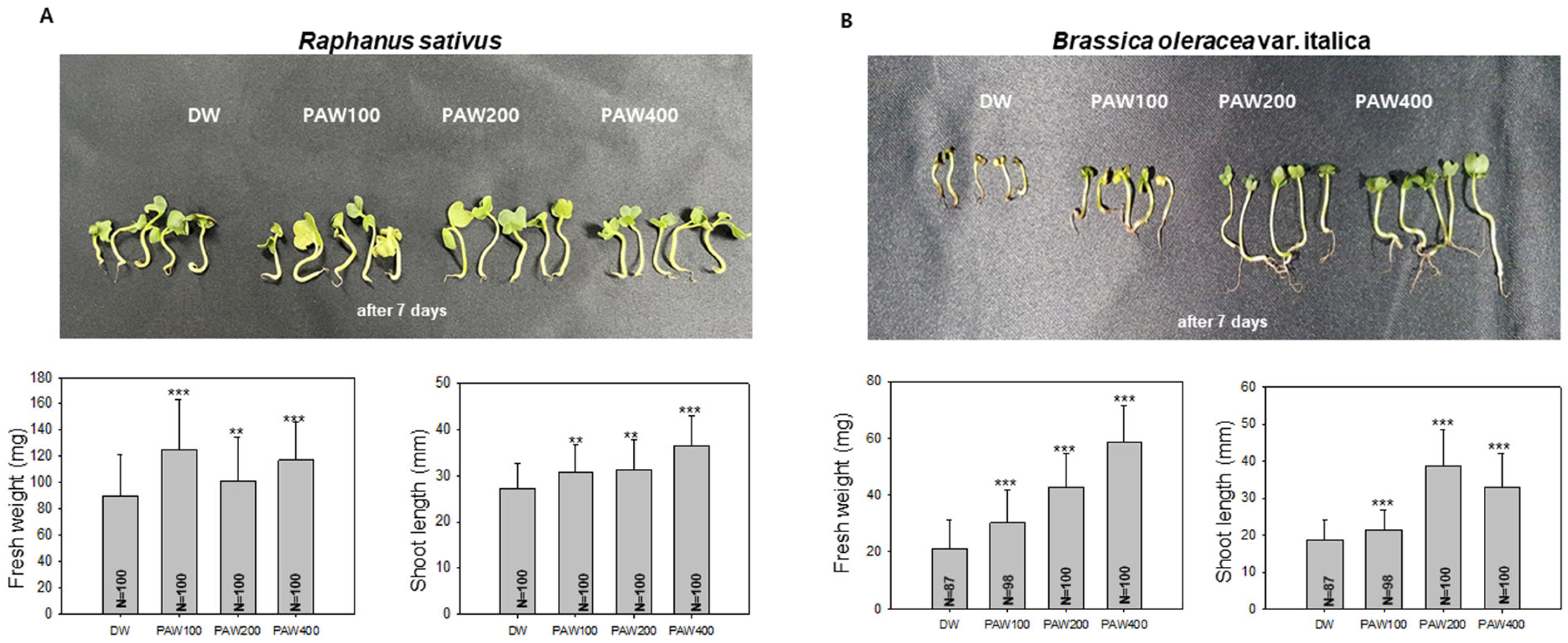
Radish exhibited a growth increase of approximately 30% in fresh weight and about 25% in shoot length with PAW400. Interestingly, significant growth enhancements were also observed with both PAW100 and PAW200 (Figure 3A). The fresh weight of broccoli sprouts increased significantly about 175% in PAW400, 110% in PAW200, and 50% in PAW100 (Figure 3B). These results correlated with the shoot length. This suggests that the growth of both radish and broccoli can be enhanced, even with PAW400. However, it is worth noting that the application of diluted PAW may vary, as seen in Figure 2 with barley and wheat.
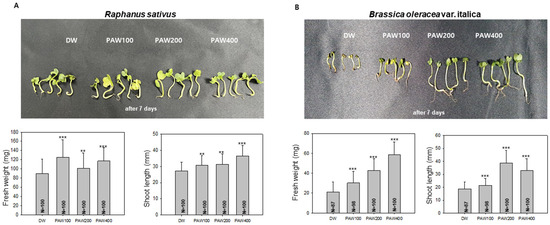
Figure 3.
The growth of Raphanus sativus (radish) and Brassica oleracea var. Italica (broccoli) sprouts in diluted PAW. (A). Growth of radish in diluted PAW. One hundred radish sprouts were hydroponically cultivated for 7 days in D.W or diluted PAW. Subsequently, measurements of fresh weight and shoot length were taken, and differences from the control group raised with D.W were analyzed. Data are presented as means ± SD. Asterisks indicate statistical significance compared with the control group, with ** p < 0.01 and *** p < 0.001. (B). Growth of broccoli sprouts with D.W and diluted PAW. 87 to 100 broccoli sprouts were hydroponically cultivated. After 7 days, measurements of fresh weight and shoot length were taken, and a t-test was performed to analyze the results. Data are presented as means ± SD. The asterisk indicates statistical significance compared with the control group, with *** p < 0.001. These experiments were conducted at least three times.
3.4. The Growth of Radishes and Broccoli Showed a More Effective Increase with PAW200 Compared to Liquid Fertilizer Applied to the Soil
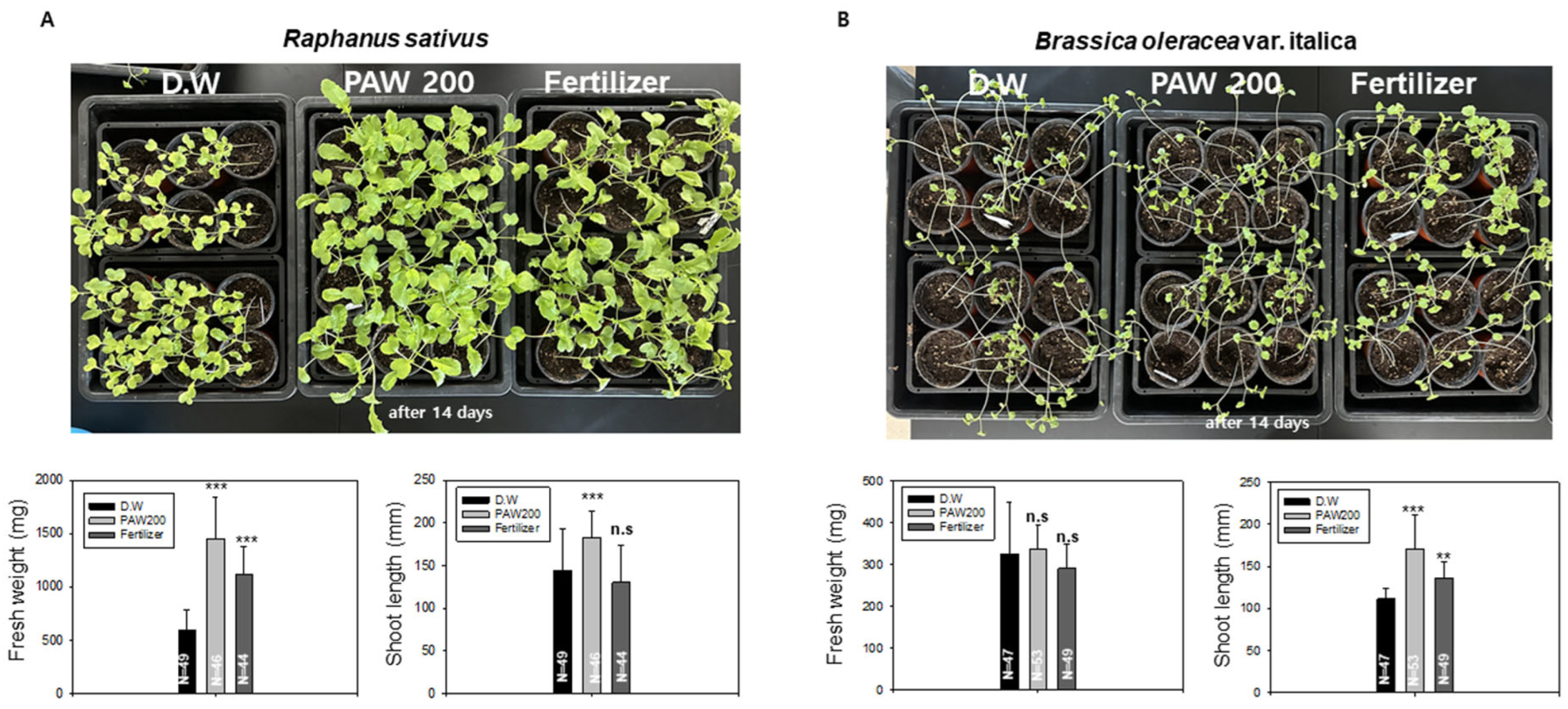
In the subsequent experiment, we conducted a comparative study on the growth of radish and broccoli cultivated in soil treated with a 0.02% liquid fertilizer, D.W., and PAW200. In Figure 4A, we compared the growth of radish cultivated in soil with D.W., PAW200, and 0.02% liquid fertilizer over a 14-day period. The fresh weight in the PAW200 treatment group exhibited an increase of about 130% compared to the D.W. treatment group, whereas the fertilizer treatment group showed an approximately 85% increase. While the effectiveness of the 0.02% liquid fertilizer used in this study could potentially impact radish growth with increased concentration, the research indirectly supports the idea that the availability of PAW, a nitrogen-rich source, may offer a practical means to lessen dependence on liquid fertilizers. Overall, the conditions for growing broccoli in soil are not optimally applied. In the PAW200 treatment group, there was an increase in total length, but there was no significant effect on live weight (Figure 4B).
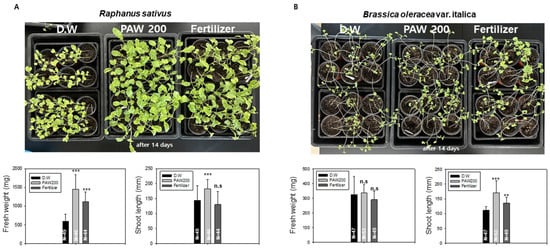
Figure 4.
The growth of radish and broccoli was assessed when irrigated with PAW200 and liquid fertilizer in soil. (A). Growth of radish sprouts in soil with PAW200 and 0.02% liquid fertilizer. 44 to 49 radish sprouts were cultivated on soil, the sprouts were irrigated with D.W, PAW200, and 0.02% liquid fertilizer every 3 days. Measurements of fresh weight and total length were taken after 14 days. The asterisk indicates statistical significance, with *** p < 0.001 from the control group. n.s. indicates no statistically significant difference. (B). Growth of broccoli sprouts in soil with PAW200 and 0.02% liquid fertilizer. 47 to 53 broccoli sprouts were irrigated with D.W, PAW200, and 0.02% liquid fertilizer every 3 days. Measurements of fresh weight and total length were taken after 14 days. Data are presented as means ± SD. Asterisks indicate statistical significance compared with the control group, with ** p < 0.01 and *** p < 0.001. These experiments were conducted at least three times.
In this study, the effects of PAW, which contains abundant RNS, were compared with those of simple nitrogen sources such as KNO3 and NaNO3 on plant growth. PAW200, containing approximately 200 mg/L of nitrate (NO3−), was evaluated for its efficacy as a nitrogen source. The performance of PAW200 was compared to treatments with KNO3 and NaNO3 at the same concentration (200 mg/L). For radish, PAW200 significantly enhanced fresh weight compared to both KNO3 and NaNO3 treatments, demonstrating its efficacy as a nitrogen source (Supplemental Figure S2A). The observed impact of PAW200 on plant growth may be attributed to its abundant RNS, which appears to exert greater effects than simple nitrogen sources. In contrast, no significant differences were observed in broccoli between PAW200 and the KNO3 or NaNO3 treatments (Supplemental Figure S2B). These results suggest that PAW has potential as an effective nitrogen source, though its impact may vary depending on the crop species. The observed variability in plant responses may also reflect differences in growth conditions, such as hydroponic versus soil-based cultivation systems.
3.5. Autophagy Responds to PAW
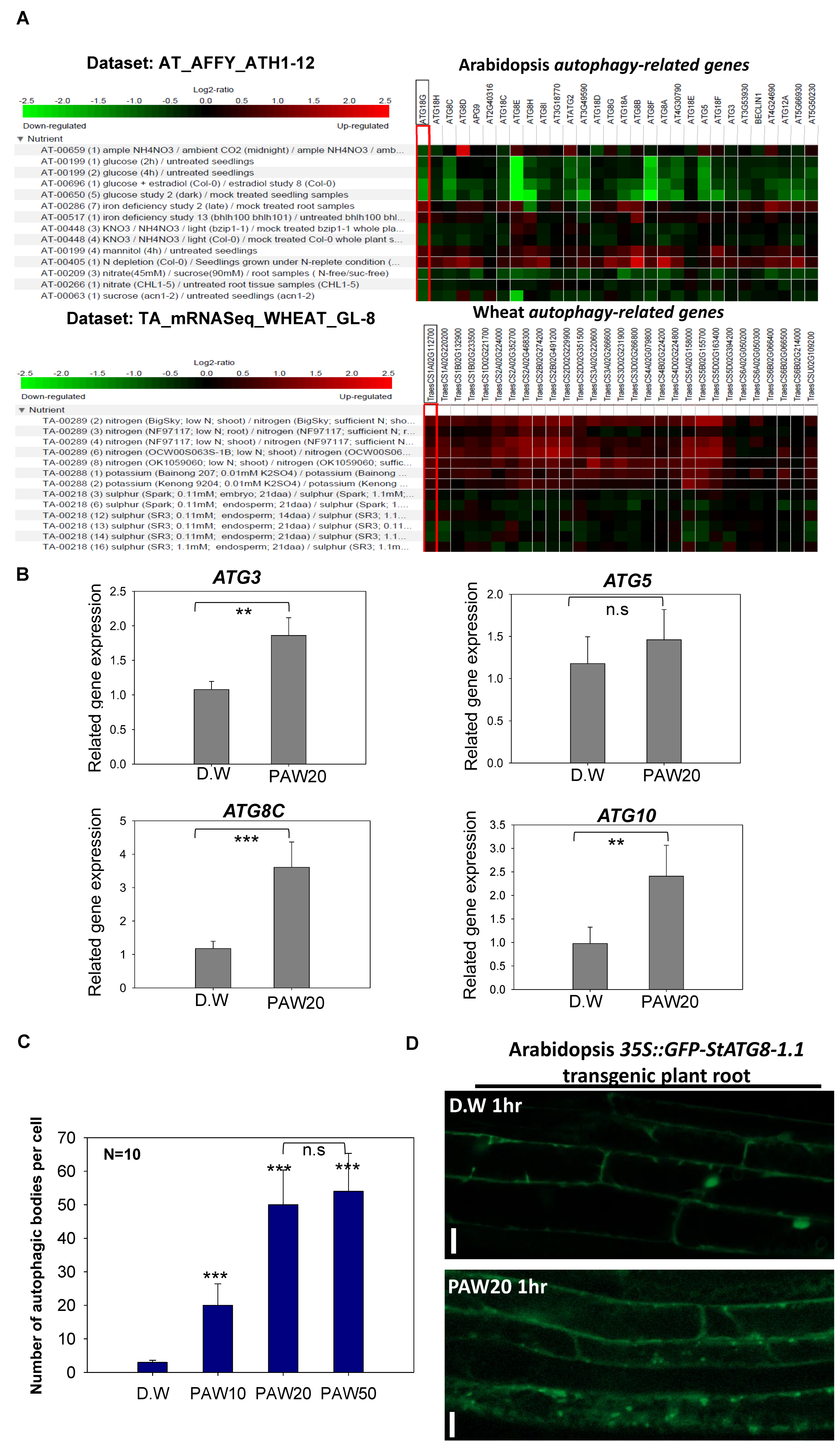
Autophagy, a cellular phenomenon central to the degradation and recycling of cellular components, is intricately linked to the presence of ROS and RNS [42]. Nitric oxide (NO), a molecule belonging to the RNS, is known to regulate intracellular autophagy [43]. Although NO and nitrate (NO3−) are separate molecules, both are involved in cell signaling, and NO has been recognized for its role in regulating the autophagy process. In our initial approach, we employed bioinformatics to assess the responsiveness of autophagy-related genes to nitrate. Analyzing Arabidopsis microarray and wheat RNA-seq data through the Geneinvestigator which is a web-based platform for gene expression [44], we observed that the treatment with nitrogen sources such as KNO3 induced the expression of autophagy-related genes (Figure 5A). Specifically, a strong response of autophagy-related genes was observed when there was an insufficiency in nitrogen, a crucial nutrient. To further validate the preceding in silico data, we performed qRT-PCR analysis using Col-0 seedlings treated with PAW20 for 1 h and gene-specific primers for four known Arabidopsis autophagy-related genes (ATG3, ATG5, ATG8C, and ATG10). As shown in Figure 5B, the expression levels of most of these selected ATG genes were upregulated in response to PAW20 treatment.
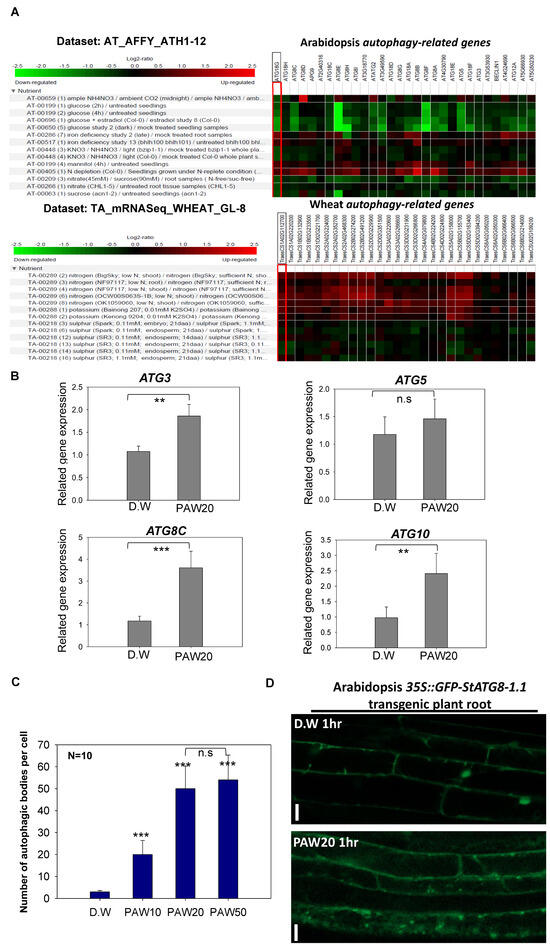
Figure 5.
Autophagy responds to PAW containing nitrate (NO3−). (A). In silico analysis of Arabidopsis and wheat ATG gene expressions using the Genevestigator. In silico gene expression analysis was conducted using microarray data for Arabidopsis and RNA-seq data for wheat. The data were classified to reveal the response of autophagy genes to nitrogen sources, and the results were exhibited on a log2 scale. (B). Expression patterns of autophagy-related genes in Arabidopsis under PAW treatment. Seven-day-old Col-0 seedlings were treated with PAW20 for 1 h, and relative transcript levels of autophagy-related genes were quantified by qRT-PCR. Each value represents the mean ± SD of three biological replicates (n = 3). Statistical significance was determined using a t-test: ** p < 0.01 and *** p < 0.001. (C). Differences in autophagic body formation under varying concentrations of PAW. Seven-day-old GFP–StATG8-1.1 transgenic Arabidopsis thaliana seedlings were treated with different concentrations of PAW. After 1 h of treatment, autophagic bodies appearing as GFP puncta were visualized in root cells using a confocal laser scanning microscope and quantified with ImageJ software. For each treatment, ten seedlings (n = 10) were analyzed in each of three independent biological replicates. Asterisks indicate statistically significant differences compared with the control (D.W.), where *** represents p < 0.001, while the lack of significant difference between PAW20 and PAW50 is indicated as n.s. The n.s. indicates no statistically significant difference. (D). The formation of autophagic bodies in potato GFP-StATG8.1-1 transformed Arabidopsis roots was investigated following PAW20 treatment. The roots of transgenic plants were treated with either D.W or PAW20 for 1 h. The autophagic body formation was observed using a confocal microscope. A round spot formed by GFP represents an autophagic body. The scale bar indicates a length of 20 µm.
For additional experiments to determine whether PAW is involved in the activation of autophagy, we examined autophagosome formation using the 35S::GFP-StATG8 transgenic line (Supplemental Figure S3). The quantification of autophagic bodies under different PAW concentrations revealed that autophagic activity was maximally induced at PAW20, with no further increase observed at higher concentrations such as PAW50 (Figure 5C,D). This result indicates that PAW-induced autophagy depends on an optimal range of ROS/RNS levels, beyond which additional activation is not promoted and PAW is capable of sufficiently activating autophagy, including RNS. In conclusion, our findings confirm that PAW has the capacity to activate genes associated with autophagy, influencing the formation of autophagosomes.
3.6. Root Growth of Autophagy Mutants Is Partially Affected by PAW
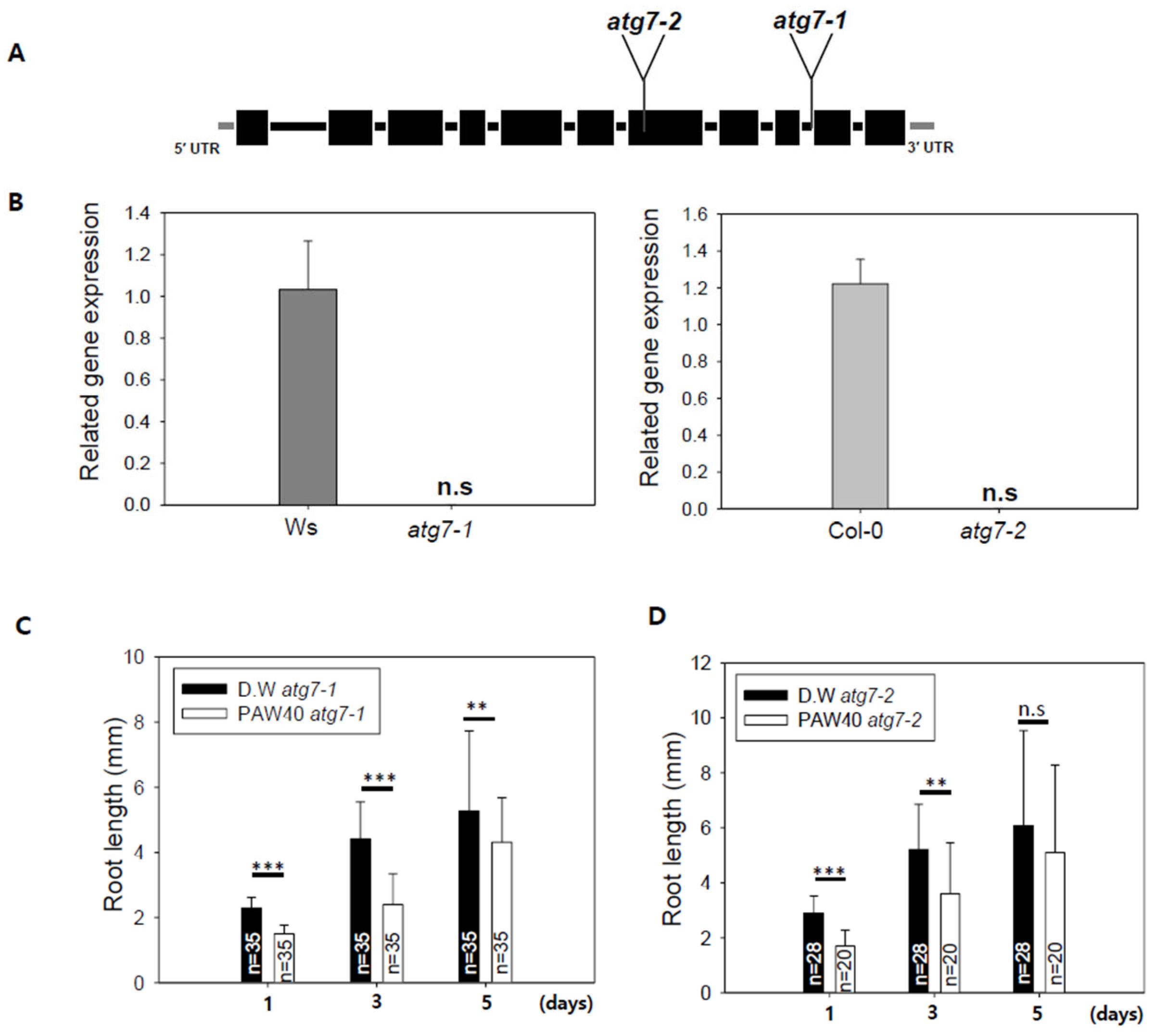
Based on previous findings, we genetically investigated the effect of PAW on autophagy. The atg7-1 and atg7-2 mutant alleles (Figure 6A) are known to be impaired in autophagy [37,45]. The presence or absence of ATG7 mRNA transcripts was confirmed by qRT-PCR (Figure 6B). Seeds of the Arabidopsis atg7-1 and atg7-2 mutants were cultivated on 0.8% agar medium prepared with either D.W. or PAW40. PAW treatment promoted root growth in Arabidopsis [46]. In our study, PAW40 significantly increased root growth compared with the D.W.-treated control in both Col-0 and the Ws-2 ecotypes (Supplemental Figure S4). In Figure 6C,D, the growth of both atg7 mutant alleles was significantly inhibited under PAW40 treatment compared to D.W. This suggests a potential relationship between the effects of PAW and the function of autophagy.
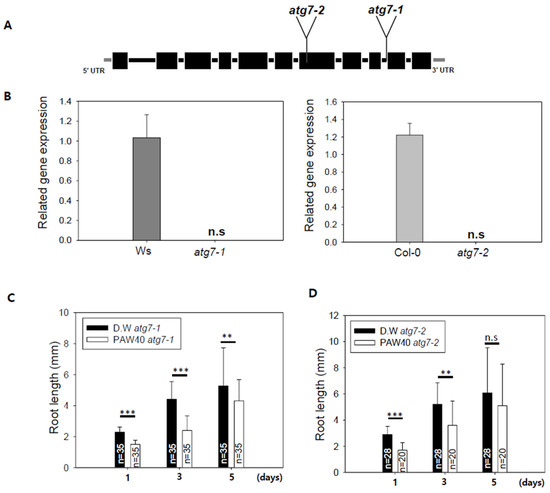
Figure 6.
Root growth of the Arabidopsis autophagy mutant atg7 alleles was evaluated on a medium containing PAW. (A). Characterization and expression analysis of T-DNA insertion mutant alleles in atg7. The atg7-1 allele originates from the Wassilewskija (Ws) background, while the atg7-2 allele is derived from the Columbia (Col) background. Introns are represented by lines, and exons are represented by boxes, with coding regions in black and untranslated regions in grey. The positions of T-DNA insertions were determined using the T-DNA Express (Arabidopsis Gene Mapping Tool). (B). The qRT-PCR analysis was conducted to assess ATG7 transcript levels in homozygous atg7 mutants. n.s. indicates no statistically significant difference. (C,D). Arabidopsis atg7-1 and atg7-2 mutant seeds were cultivated on 0.8% agar medium prepared with D.W or 0.8% agar medium prepared with PAW40. Root length was measured after 1, 3, and 5 days. Data are presented as means ± SD. Asterisks indicate statistical significance, with ** p < 0.01 and *** p < 0.001. This experiment was conducted at least three times. n.s. indicates no statistically significant difference.
4. Discussion
4.1. Overview of PAW Effects on Sprout Growth
In this study, we examined the impact of diluted PAW1000, which initially contained 1000 ppm of NO3−, on sprout cultivation. Diluted PAW was applied in a rapid and simple manner to evaluate its growth-promoting effects. Although responses varied across sprout species, the minimum effective concentration was approximately PAW200. Our findings also indicate that PAW can promote growth at levels comparable to commercially available nitrogen fertilizers. Furthermore, we identified a correlation between autophagy-related gene expression and nitrogen responsiveness, suggesting that PAW may activate autophagy in plants. This activation is likely influenced by reactive nitrogen species (RNS) present in PAW, which may indirectly influence plant growth.
4.2. Implications of PAW in Sustainable Agriculture
Agricultural practices are rapidly changing in response to climate change, with increasing emphasis on sustainable technologies that enhance production while minimizing environmental pollution [47,48]. Precision agriculture has emerged as an important approach to optimize nutrient use and reduce environmental burden [49,50]. Our findings indicate that crop responses to PAW are not determined solely by the presence of nitrogenous compounds such as NO3−. Plasma treatment also introduces trace elements, highlighting the need for further investigations into changes in PAW ion composition and their roles in both plant growth and autophagy activation. In addition, deeper understanding of plasma-derived ions beyond nitrate will be essential for evaluating the broader potential of PAW in agricultural systems. Nutrient mismanagement in smart farming increases costs and risks eutrophication. Therefore, PAW, which contains plasma-generated ROS and RNS, offers a sustainable complementary or alternative source of nitrogen.
4.3. Crop-Specific Responses and Physiological Mechanisms
Previous studies have shown that plant responses to PAW are highly species dependent. While PAW irrigation enhanced germination and biomass accumulation in crops such as bell pepper and tomato, excessive application has produced limited or nonsignificant effects [51]. These observations imply that plant perception of PAW-derived RONS and nitrogenous compounds varies among species, supporting the idea that PAW functions as a signaling agent rather than merely a nutrient source. In our study, Brassicaceae crops such as radish and broccoli exhibited subtle but distinct differences in their growth responses to PAW, despite phylogenetic similarity. These differences may arise from species-specific variations in root development, nutrient demand, and sensitivity to oxidative or nitrogen-related signals. The concentration-dependent growth responses observed here further indicate that PAW effects are both crop specific and dose regulated, while also suggesting that a generally applicable effective concentration range may exist.
4.4. Agronomic Potential and Limitations of PAW Application
PAW offers several advantages for agriculture, particularly because it can be generated on-site using only water [52,53]. By supplying low levels of nitrogen together with signaling-active ROS and RNS, PAW may promote growth through regulatory pathways rather than nutrient oversupply, underscoring its potential as a sustainable alternative to conventional fertilizers. However, limitations remain. The chemical composition of PAW varies depending on plasma generation conditions, including the gas type and treatment duration, resulting in variable plant responses. Therefore, systematic studies are needed to identify crop-specific optimal PAW concentrations and treatment strategies. Additionally, the long-term effects of repeated PAW application on soil properties, microbial communities, and plant performance require further investigation. Nevertheless, our results show that PAW extends beyond a laboratory-scale concept and shows strong potential for practical use in agricultural systems.
4.5. Autophagy Activation and Molecular Responses to PAW
Autophagy is essential for plant development, stress adaptation, and cellular homeostasis [54,55]. In this study, we highlight the potential applicability of PAW in promoting plant growth via activation of autophagy-related pathways. Nitrate produced as part of plasma-generated RNS has been widely recognized as a major contributor to plant growth enhancement [56,57]. Transcriptome analysis in PAW-treated mung bean revealed significant changes, including differential expression of 258 genes [58]. Among them, WRKY transcription factors—key regulators of immunity and development—were upregulated [59,60], similar to nitrate treatment [58]. These findings suggest that plant growth promotion involves not only nutrient uptake but also transcriptional regulation associated with nitrate-responsive pathways. Moreover, WRKY proteins have been reported to bind autophagy gene promoters, activate their transcription, and interact with ATG8 [61,62].
4.6. ROS/RNS-Mediated Regulation of Autophagy
ROS and RNS encompass diverse signaling molecules that regulate cellular pathways depending on intracellular redox balance, although additional studies are required to identify the specific molecules responsible for PAW-mediated signaling [63,64,65]. The ROS/RNS network constitutes a highly sophisticated redox regulatory system that fine-tunes the activation threshold of autophagy [42,66]. While balanced ROS/RNS levels promote survival by facilitating autophagy-mediated recycling of damaged components, excessive accumulation can trigger oxidative damage and cell death [66,67,68]. These findings underscore the importance of redox homeostasis in determining autophagy outcomes [66,67]. Thus, appropriate concentrations of ROS/RNS present in PAW may contribute to growth promotion and developmental regulation through autophagic modulation.
4.7. Agricultural Prospects for Autophagy-Modulating PAW
Autophagy regulates diverse processes in plants, including programmed cell death, development, homeostasis, and immunity [69,70,71,72,73,74]. Accordingly, agricultural application of PAW containing appropriate levels of ROS and RNS may enhance both plant growth and immune function through autophagy-associated signaling pathways. Optimizing PAW formulations to leverage autophagy modulation may offer a promising strategy for improving plant health and resilience in sustainable agricultural systems.
5. Conclusions
This study aimed to clarify plant responses to PAW treatment and to explore its potential applications in sprout crop cultivation. Our experimental results demonstrated that diluted PAW1000 significantly promotes the growth of various sprout crops. Additionally, PAW irrigation was found to partially induce autophagic activity, suggesting activation of autophagy pathways. These effects are mediated by RNS present in PAW. To deepen understanding, further experiments are needed to compare the impacts of PAW containing both ROS and RNS with those containing only RNS. Investigating how combined ROS and RNS influence plant growth, immunity, and autophagy will provide valuable insights into the mechanisms behind PAW’s effects. Overall, these findings contribute foundational knowledge to the molecular mechanisms of PAW action and support its application for enhancing crop yield in hydroponic smart farming systems.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agronomy16020207/s1, Supporting Information Figures S1–S4: Figure S1. Effects of PAW irrigation on total plant length of wheat and radish under hydroponic cultivation. After cultivating wheat and radish for 7 days under hydroponic cultivation conditions with PAW1000, PAW100, PAW10, and D.W irrigation, the total length, including roots, was measured. Data are presented as means ± SD. Asterisks indicate statistical significance, with * p < 0.05 and ** p < 0.001. This experiment was conducted at least three times. Figure S2. Growth responses of radish and broccoli sprouts irrigated with PAW200, KNO3, and NaNO3 treatments. A and B. Growth of radish and broccoli sprouts irrigated with PAW200, 200 mg/L KNO3, or 200 mg/L NaNO3. Fifty radish sprouts were cultivated in soil and irrigated with distilled water (D.W.), PAW200, or 200 mg/L KNO3 or NaNO3 every three days. Fresh weight was measured after 14 days. Data are presented as means ± SD. Asterisks indicate statistically significant differences compared with the control group (** p < 0.001). All experiments were performed at least three times. Figure S3. qRT-PCR analysis of gene expression in 35S::GFP-stATG8.1-1. Arabidopsis transgenic lines. qRT-PCR analysis was performed to evaluate gene expression in T3 transgenic lines of Arabidopsis carrying the 35S::GFP-StATG8.1-1 construct. Wild-type Arabidopsis Col-0 plants were used as a negative control. qRT-PCR analysis was performed using three independent biological samples per replicate. Figure S4. Effects of PAW40 on root growth in two Arabidopsis ecotypes, Col-0 and Ws2. Seedlings of the two ecotypes were transferred to D.W. or PAW40 immediately after germination, and root length was measured at 1, 3, and 5 days. A total of 50 seedlings were analyzed for each treatment. Data are presented as means ± SD. Asterisks indicate statistically significant differences compared with the control group. All experiments were performed at least three times.
Author Contributions
Conceptualization, I.S., S.H., Y.K.L. and S.U.H.; Data curation, I.S., S.H., Y.J.N., S.Y.J., J.Y.J., Y.K.L. and S.U.H.; Formal analysis, S.U.H.; Investigation, S.U.H.; Methodology and sources, Y.K.L. and S.U.H.; Software, S.U.H.; Supervision, Y.K.L. and S.U.H.; Writing—original draft, S.U.H.; Writing—review and editing, Y.K.L. and S.U.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the X-Corps program of the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (No. NRF-2022H1D8A3038669), and the ‘Regional Innovation Strategy (RIS)’ program through the NRF, funded by the Ministry of Education (MOE) (2023RIS-008). Additional support for this work was provided by the R&D program ‘Plasma Advanced Technology for Agriculture and Food (Plasma Farming)’ (Nos. EN2325-10 and EN2425-11) through the Korea Institute of Fusion Energy (KFE), and by the NRF grant funded by the Korea government (MSIT) (No. NRF-2021R1A2C1005650) to Y.K.L.
Data Availability Statement
The original contributions presented in the study are included in the article; further inquiries can be directed to the corresponding authors.
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
References
- Campbell, E.; Uppalapati, S.S.; Kotcher, J.; Maibach, E. Communication research to improve engagement with climate change and human health: A review. Front. Public Health 2022, 10, 1086858. [Google Scholar] [CrossRef] [PubMed]
- Lashari, M.H.; Karim, S.; Alhussein, M.; Hoshu, A.A.; Aurangzeb, K.; Anwar, M.S. Internet of Things-based sustainable environment management for large indoor facilities. PeerJ Comput. Sci. 2023, 9, e1623. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Udayakumar, A.; Anbarasa Kumar, A.; Senthamarai Kannan, K.; Krishnan, N. Multiparameter optimization system with DCNN in precision agriculture for advanced irrigation planning and scheduling based on soil moisture estimation. Environ. Monit. Assess. 2022, 195, 13. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-de-Santos, P.; Fernández, R.; Sepúlveda, D.; Navas, E.; Emmi, L.; Armada, M. Field Robots for Intelligent Farms—Inhering Features from Industry. Agronomy 2020, 10, 1638. [Google Scholar] [CrossRef]
- Huang, W.T.; Su, T.Y.; Chuang, J.H.; Lu, K.M.; Hu, S.F.; Liu, R.S. Plant Growth Modeling and Response from Broadband Phosphor-Converted Lighting for Indoor Agriculture. ACS Appl. Mater. Interfaces 2023, 15, 32589–32596. [Google Scholar] [CrossRef]
- Balusamy, S.R.; Joshi, A.S.; Perumalsamy, H.; Mijakovic, I.; Singh, P. Advancing sustainable agriculture: A critical review of smart and eco-friendly nanomaterial applications. J. Nanobiotechnol. 2023, 21, 372. [Google Scholar] [CrossRef]
- Garg, D.; Sridhar, K.; Stephen Inbaraj, B.; Chawla, P.; Tripathi, M.; Sharma, M. Nano-Biofertilizer Formulations for Agriculture: A Systematic Review on Recent Advances and Prospective Applications. Bioengineering 2023, 10, 1010. [Google Scholar] [CrossRef]
- Klerkx, L.; Jakku, E.; Labarthe, P. A review of social science on digital agriculture, smart farming and agriculture 4.0: New contributions and a future research agenda. NJAS Wagening. J. Life Sci. 2019, 90–91, 100315. [Google Scholar] [CrossRef]
- Kumari, N.; Kumar, M.; Singh, A.; Pandey, A.K. Recent Innovation in Precision Agriculture and their Impact on Crop and Soil Health: A Compressive Review. J. Sci. Res. Rep. 2024, 30, 382–393. [Google Scholar] [CrossRef]
- Munz, J. What if precision agriculture is not profitable?: A comprehensive analysis of the right timing for exiting, taking into account different entry options. Precis. Agric. 2024, 25, 1284–1323. [Google Scholar] [CrossRef]
- Naïtali, M.; Kamgang-Youbi, G.; Herry, J.-M.; Bellon-Fontaine, M.-N.; Brisset, J.-L. Combined Effects of Long-Living Chemical Species during Microbial Inactivation Using Atmospheric Plasma-Treated Water. Appl. Environ. Microbiol. 2010, 76, 7662–7664. [Google Scholar] [CrossRef]
- Zhao, Y.M.; Patange, A.; Sun, D.W.; Tiwari, B. Plasma-activated water: Physicochemical properties, microbial inactivation mechanisms, factors influencing antimicrobial effectiveness, and applications in the food industry. Compr. Rev. Food Sci. Food Saf. 2020, 19, 3951–3979. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Cheng, J.-H.; Sun, D.-W. Enhancement of Wheat Seed Germination, Seedling Growth and Nutritional Properties of Wheat Plantlet Juice by Plasma Activated Water. J. Plant Growth Regul. 2023, 42, 2006–2022. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Francis, K.; Zhang, X. Review on formation of cold plasma activated water (PAW) and the applications in food and agriculture. Food Res. Int. 2022, 157, 111246. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Zhang, A.; Heroux, P.; Sand, W.; Sun, Z.; Zhan, J.; Wang, C.; Hao, S.; Li, Z.; Li, Z.; et al. Effect of Dielectric Barrier Discharge Cold Plasma on Pea Seed Growth. J. Agric. Food Chem. 2019, 67, 10813–10822. [Google Scholar] [CrossRef]
- Liu, Q.; Wu, H.; Luo, J.; Liu, J.; Zhao, S.; Hu, Q.; Ding, C. Effect of dielectric barrier discharge cold plasma treatments on flavor fingerprints of brown rice. Food Chem. 2021, 352, 129402. [Google Scholar] [CrossRef]
- Zheng, Z.; Niu, L.; Yang, W.; Chen, Y.; Huang, Y.; Li, C. Degradation of Zearalenone by Dielectric Barrier Discharge Cold Plasma and Its Effect on Maize Quality. Foods 2023, 12, 1129. [Google Scholar] [CrossRef]
- Loganathan, S.; Khacef, A. Enhanced seed germination and plant growth by atmospheric pressure cold air plasma: Combined effect of seed and water treatment. RSC Adv. 2017, 7, 1822–1832. [Google Scholar] [CrossRef]
- Locatelli, S.; Triolone, S.; De Bonis, M.; Zanin, G.; Nicoletto, C. Non-Thermal Plasma-Activated Water Enhances Nursery Production of Vegetables: A Species-Specific Study. Agronomy 2025, 15, 209. [Google Scholar] [CrossRef]
- Del Río, L.A. ROS and RNS in plant physiology: An overview. J. Exp. Bot. 2015, 66, 2827–2837. [Google Scholar] [CrossRef]
- Adhikari, B.; Adhikari, M.; Ghimire, B.; Park, G.; Choi, E.H. Cold Atmospheric Plasma-Activated Water Irrigation Induces Defense Hormone and Gene expression in Tomato seedlings. Sci. Rep. 2019, 9, 16080. [Google Scholar] [CrossRef] [PubMed]
- Zambon, Y.; Contaldo, N.; Laurita, R.; Várallyay, E.; Canel, A.; Gherardi, M.; Colombo, V.; Bertaccini, A. Plasma activated water triggers plant defence responses. Sci. Rep. 2020, 10, 19211. [Google Scholar] [CrossRef] [PubMed]
- Yun, B.W.; Skelly, M.J.; Yin, M.; Yu, M.; Mun, B.G.; Lee, S.U.; Hussain, A.; Spoel, S.H.; Loake, G.J. Nitric oxide and S-nitrosoglutathione function additively during plant immunity. New Phytol. 2016, 211, 516–526. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Ma, J.; Li, J.; Chen, S.; Luo, J.; Wu, J.; Zhang, K.; Peng, C.; Zeng, Y.; Li, F.; et al. ATG8ylation-mediated tonoplast invagination mitigates vacuole damage. Nat. Commun. 2025, 16, 6621. [Google Scholar] [CrossRef]
- Li, Y.; Xu, X.; Qi, G.; Cui, D.; Huang, C.; Sui, X.; Li, G.; Fan, Q. Mechanisms of autophagy function and regulation in plant growth, development, and response to abiotic stress. Crop J. 2023, 11, 1611–1625. [Google Scholar] [CrossRef]
- Haq, S.I.U.; Tariq, F.; Sama, N.U.; Jamal, H.; Mohamed, H.I. Role of autophagy in plant growth and adaptation to salt stress. Planta 2025, 261, 49. [Google Scholar] [CrossRef]
- Geng, J.; Li, J.; Zhu, F.; Chen, X.; Du, B.; Tian, H.; Li, J. Plant sprout foods: Biological activities, health benefits, and bioavailability. J. Food Biochem. 2022, 46, e13777. [Google Scholar] [CrossRef]
- Ebert, A.W. Sprouts and Microgreens-Novel Food Sources for Healthy Diets. Plants 2022, 11, 571. [Google Scholar] [CrossRef]
- Holman, J.; Hurd, M.; Moses, P.L.; Mawe, G.M.; Zhang, T.; Ishaq, S.L.; Li, Y. Interplay of broccoli/broccoli sprout bioactives with gut microbiota in reducing inflammation in inflammatory bowel diseases. J. Nutr. Biochem. 2023, 113, 109238. [Google Scholar] [CrossRef]
- Kehinde, B.A.; Majid, I.; Hussain, S. Isolation of bioactive peptides and multiple nutraceuticals of antidiabetic and antioxidant functionalities through sprouting: Recent advances. J. Food Biochem. 2022, 46, e14317. [Google Scholar] [CrossRef]
- Lee, Y.K.; Lim, J.; Hong, E.J.; Kim, S.B. Plasma-activated water regulates root hairs and cotyledon size dependent on cell elongation in Nicotiana tabacum L. Plant Biotechnol. Rep. 2020, 14, 663–672. [Google Scholar] [CrossRef]
- Everett, S.A.; Dennis, M.F.; Tozer, G.M.; Prise, V.E.; Wardman, P.; Stratford, M.R.L. Nitric oxide in biological fluids: Analysis of nitrite and nitrate by high-performance ion chromatography. J. Chromatogr. A 1995, 706, 437–442. [Google Scholar] [CrossRef]
- Priatama, R.A.; Beak, H.K.; Park, S.; Song, I.; Park, S.J.; Kim, S.B.; Lee, Y.K. Tomato yield enhancement with plasma-activated water as an alternative nitrogen source. BMC Plant Biol. 2025, 25, 668. [Google Scholar] [CrossRef] [PubMed]
- Dagdas, Y.F.; Belhaj, K.; Maqbool, A.; Chaparro-Garcia, A.; Pandey, P.; Petre, B.; Tabassum, N.; Cruz-Mireles, N.; Hughes, R.K.; Sklenar, J.; et al. An effector of the Irish potato famine pathogen antagonizes a host autophagy cargo receptor. eLife 2016, 5, e10856. [Google Scholar] [CrossRef] [PubMed]
- Huh, S.U. Interaction of Potato Autophagy-Related StATG8 Family Proteins with Pathogen Effector and WRKY Transcription Factor in the Nucleus. Microorganisms 2025, 13, 1589. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Chung, T.; Phillips, A.R.; Vierstra, R.D. ATG8 lipidation and ATG8-mediated autophagy in Arabidopsis require ATG12 expressed from the differentially controlled ATG12A AND ATG12B loci. Plant J. 2010, 62, 483–493. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Schindelin, J.; Rueden, C.T.; Hiner, M.C.; Eliceiri, K.W. The ImageJ ecosystem: An open platform for biomedical image analysis. Mol. Reprod. Dev. 2015, 82, 518–529. [Google Scholar] [CrossRef]
- Kaneko, T.; Takashima, K.; Sasaki, S. Integrated Transport Model for Controlled Delivery of Short-Lived Reactive Species via Plasma-Activated Liquid with Practical Applications in Plant Disease Control. Plasma Chem. Plasma Process. 2024, 44, 1165–1201. [Google Scholar] [CrossRef]
- Rathore, V.; Nema, S.K. A Nitrogen Alternative: Use of Plasma Activated Water as Nitrogen Source in Hydroponic Solution for Radish Growth. arXiv 2024, arXiv:2404.16910. [Google Scholar] [CrossRef]
- Navarro-Yepes, J.; Burns, M.; Anandhan, A.; Khalimonchuk, O.; del Razo, L.M.; Quintanilla-Vega, B.; Pappa, A.; Panayiotidis, M.I.; Franco, R. Oxidative stress, redox signaling, and autophagy: Cell death versus survival. Antioxid. Redox Signal 2014, 21, 66–85. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, D.N.; Chowdhury, R.; Trudel, L.J.; Tee, A.R.; Slack, R.S.; Walker, C.L.; Wogan, G.N. Reactive nitrogen species regulate autophagy through ATM-AMPK-TSC2-mediated suppression of mTORC1. Proc. Natl. Acad. Sci. USA 2013, 110, E2950–E2957. [Google Scholar] [CrossRef] [PubMed]
- Hruz, T.; Laule, O.; Szabo, G.; Wessendorp, F.; Bleuler, S.; Oertle, L.; Widmayer, P.; Gruissem, W.; Zimmermann, P. Genevestigator v3: A reference expression database for the meta-analysis of transcriptomes. Adv. Bioinform. 2008, 2008, 420747. [Google Scholar] [CrossRef]
- Yan, H.; Lu, Z.; Du, X.; You, Z.; Yang, M.; Li, N.; Li, X.; Ni, Z.; Wu, H.; Wang, X.; et al. Autophagy modulates Arabidopsis male gametophyte fertility and controls actin organization. Nat. Commun. 2024, 15, 10071. [Google Scholar] [CrossRef]
- Ka, D.H.; Priatama, R.A.; Park, J.Y.; Park, S.J.; Kim, S.B.; Lee, I.A.; Lee, Y.K. Plasma-Activated Water Modulates Root Hair Cell Density via Root Developmental Genes in Arabidopsis thaliana L. Appl. Sci. 2021, 11, 2240. [Google Scholar] [CrossRef]
- Datta, S.; Hamim, I.; Jaiswal, D.K.; Sungthong, R. Sustainable agriculture. BMC Plant Biol. 2023, 23, 588. [Google Scholar] [CrossRef]
- Yang, Y.; Jin, Z.; Mueller, N.D.; Driscoll, A.W.; Hernandez, R.R.; Grodsky, S.M.; Sloat, L.L.; Chester, M.V.; Zhu, Y.G.; Lobell, D.B. Sustainable irrigation and climate feedbacks. Nat. Food 2023, 4, 654–663. [Google Scholar] [CrossRef]
- Zain, M.; Ma, H.; Ur Rahman, S.; Nuruzzaman, M.; Chaudhary, S.; Azeem, I.; Mehmood, F.; Duan, A.; Sun, C. Nanotechnology in precision agriculture: Advancing towards sustainable crop production. Plant Physiol. Biochem. 2023, 206, 108244. [Google Scholar] [CrossRef]
- Melash, A.A.; Bogale, A.A.; Bytyqi, B.; Nyandi, M.S.; Ábrahám, É.B. Nutrient management: As a panacea to improve the caryopsis quality and yield potential of durum wheat (Triticum turgidum L.) under the changing climatic conditions. Front. Plant Sci. 2023, 14, 1232675. [Google Scholar] [CrossRef]
- Ferreyra, M.G.; Caffaro, M.M.; Santamaría, B.; Zilli, C.; Hernández, A.; Fina, B.L.; Vélez, A.S.; Balestrasse, K.B.; Prevosto, L. Plasma-Activated Water Produced by a Moderately High Energy-Efficient 1-Liter Reactor: Effects on Germination and Growth of Tomato and Bell Pepper Plants. Plants 2025, 14, 722. [Google Scholar] [CrossRef]
- Andrade, P.E.; Savi, P.J.; Almeida, F.S.; Carciofi, B.A.; Pace, A.; Zou, Y.; Eylands, N.; Annor, G.; Mattson, N.; Nansen, C. Plasma-Activated Water as a Sustainable Nitrogen Source: Supporting the UN Sustainable Development Goals (SDGs) in Controlled Environment Agriculture. Crops 2025, 5, 35. [Google Scholar] [CrossRef]
- Chalise, R.; Rijal, O.S.; Oli, D.; Regmi, K.; Nepal, S.; Khanal, R. Physical properties of natural and processed water using dielectric barrier and gliding arc discharge. AIP Adv. 2025, 15, 105321. [Google Scholar] [CrossRef]
- Sertsuvalkul, N.; DeMell, A.; Dinesh-Kumar, S.P. The complex roles of autophagy in plant immunity. FEBS Lett. 2022, 596, 2163–2171. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of cells and tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef]
- Lamichhane, P.; Veerana, M.; Lim, J.S.; Mumtaz, S.; Shrestha, B.; Kaushik, N.K.; Park, G.; Choi, E.H. Low-Temperature Plasma-Assisted Nitrogen Fixation for Corn Plant Growth and Development. Int. J. Mol. Sci. 2021, 22, 5360. [Google Scholar] [CrossRef]
- Bafoil, M.; Jemmat, A.; Martinez, Y.; Merbahi, N.; Eichwald, O.; Dunand, C.; Yousfi, M. Effects of low temperature plasmas and plasma activated waters on Arabidopsis thaliana germination and growth. PLoS ONE 2018, 13, e0195512. [Google Scholar] [CrossRef]
- Chou, Y.-J.; Ting, Y. Plasma-activated water regulated transcriptome gene expression leading to high germination and growth of mung beans. Chem. Biol. Technol. Agric. 2023, 10, 146. [Google Scholar] [CrossRef]
- Liu, Q.; Li, X.; Yan, S.; Yu, T.; Yang, J.; Dong, J.; Zhang, S.; Zhao, J.; Yang, T.; Mao, X.; et al. OsWRKY67 positively regulates blast and bacteria blight resistance by direct activation of PR genes in rice. BMC Plant Biol. 2018, 18, 257. [Google Scholar] [CrossRef]
- Wang, H.; Chen, W.; Xu, Z.; Chen, M.; Yu, D. Functions of WRKYs in plant growth and development. Trends Plant Sci. 2023, 28, 630–645. [Google Scholar] [CrossRef]
- Song, I.; Hong, S.; Huh, S.U. Identification and Expression Analysis of the Solanum tuberosum StATG8 Family Associated with the WRKY Transcription Factor. Plants 2022, 11, 2858. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Nolan, T.M.; Yin, Y.; Bassham, D.C. Identification of transcription factors that regulate ATG8 expression and autophagy in Arabidopsis. Autophagy 2020, 16, 123–139. [Google Scholar] [CrossRef] [PubMed]
- Praveen, A. Nitric oxide mediated alleviation of abiotic challenges in plants. Nitric Oxide 2022, 128, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Qi, S.; Wang, Y. Nitrate signaling and use efficiency in crops. Plant Commun. 2022, 3, 100353. [Google Scholar] [CrossRef]
- Gupta, K.J.; Kaladhar, V.C.; Fitzpatrick, T.B.; Fernie, A.R.; Møller, I.M.; Loake, G.J. Nitric oxide regulation of plant metabolism. Mol. Plant 2022, 15, 228–242. [Google Scholar] [CrossRef]
- Robert, G.; Enet, A.; Saavedra, L.; Lascano, R. Redox regulation of autophagy in Arabidopsis: The different ROS effects. Plant Physiol. Biochem. 2025, 223, 109800. [Google Scholar] [CrossRef]
- Mittler, R. ROS Are Good. Trends Plant Sci. 2017, 22, 11–19. [Google Scholar] [CrossRef]
- Sakil, M.A.; Angon, P.B.; Polash, M.A.S.; Mondal, S.; Zannat, M.; Islam, M.S.; Norizuki, T.; Ueda, T. Autophagic cell death in plants. Plant Cell Physiol. 2025, 66, 1389–1396. [Google Scholar] [CrossRef]
- Zou, J.; Chen, X.; Liu, C.; Guo, M.; Kanwar, M.K.; Qi, Z.; Yang, P.; Wang, G.; Bao, Y.; Bassham, D.C.; et al. Autophagy promotes jasmonate-mediated defense against nematodes. Nat. Commun. 2023, 14, 4769. [Google Scholar] [CrossRef]
- Cao, J.; Zheng, X.; Xie, D.; Zhou, H.; Shao, S.; Zhou, J. Autophagic pathway contributes to low-nitrogen tolerance by optimizing nitrogen uptake and utilization in tomato. Hortic. Res. 2022, 9, uhac068. [Google Scholar] [CrossRef]
- Yagyu, M.; Yoshimoto, K. New Insights into Plant Autophagy: Molecular Mechanisms and Roles in Development and Stress Responses. J. Exp. Bot. 2023, 75, 1234–1251. [Google Scholar] [CrossRef]
- Feng, Y.; Chen, Y.; Wu, X.; Chen, J.; Zhou, Q.; Liu, B.; Zhang, L.; Yi, C. Interplay of energy metabolism and autophagy. Autophagy 2024, 20, 4–14. [Google Scholar] [CrossRef]
- Wleklik, K.; Borek, S. Vacuolar Processing Enzymes in Plant Programmed Cell Death and Autophagy. Int. J. Mol. Sci. 2023, 24, 1198. [Google Scholar] [CrossRef]
- Huh, S.U. Evolutionary Diversity and Function of Metacaspases in Plants: Similar to but Not Caspases. Int. J. Mol. Sci. 2022, 23, 4588. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.