- Review
Russian Dolls of Heme Metabolism in Malaria-Infected Red Blood Cells: Nested Vulnerabilities and Therapeutic Opportunities
- Swamy R. Adapa,
- Faiza A. Siddiqui and
- Rays H. Y. Jiang
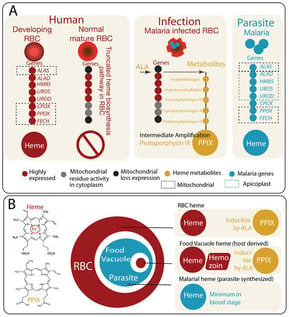
Heme metabolism is central to the biology of malaria parasites and to the mechanism of action of artemisinin-based therapies. Within malaria-infected red blood cells (RBCs), heme-related chemistry arises from multiple nested metabolic sources that function as “Russian dolls”: the truncated heme biosynthetic capacity of the host erythrocyte, the parasite’s own heme synthesis pathway, and host heme released through hemoglobin digestion in the parasite food vacuole. These overlapping metabolic layers create distinct pools of heme that can influence redox balance and drug activation. Recent studies highlight that exogenous 5-aminolevulinic acid (5-ALA) can perturb host heme biosynthesis in infected erythrocytes, potentially increasing intracellular levels of the heme intermediate protoporphyrin IX and sensitizing parasites to oxidative stress. However, the extent to which such metabolic perturbations affect artemisinin susceptibility depends strongly on parasite stage and exposure duration. Here we review the compartmentalized architecture of heme metabolism in malaria-infected RBCs and discuss how these nested vulnerabilities may be exploited for therapeutic intervention.
29 April 2026