Abstract
Background: Nano- and microplastics (N/MPs) are widespread pollutants in aquatic ecosystems and may interfere with the physiology of diatoms. However, the underlying molecular mechanisms remain poorly understood. Methods: Cultures of the diatom species Skeletonema marinoi, a key contributor to marine primary production and biogeochemical cycles, were exposed for 10 days to polyethylene (PE) N/MPs (25 and 75 µg/mL). Growth, morphology, and cell viability were monitored, while gene expression of stress, inflammation, programmed cell death (PCD), and extracellular polymeric substance (EPS) biosynthesis markers was assessed by RT-qPCR. Results: Growth was not significantly affected. Gene expression showed biphasic regulation of antioxidant enzymes, induction of HSP70/90, transient modulation of prostaglandin-related genes, and dose- and time-dependent activation of PCD markers. Selected CAZymes possibly involved in EPS synthesis were downregulated at early stages, suggesting impaired aggregation potential. Conclusions: Exposure to PE N/MPs elicited subtle yet multifaceted effects in S. marinoi, including oxidative stress, inflammatory-like responses, modulation of EPS pathways, and PCD activation. Despite apparent growth resilience, the molecular alterations observed may impact bloom dynamics and carbon export.
1. Introduction
Plastic pollution has become one of the most pressing environmental challenges of the 21st century, with microplastics (MPs, <5 mm) and nanoplastics (NPs, <1 μm) now recognized as ubiquitous contaminants in the world’s oceans [1]. Their persistence, small size, and ability to adsorb and transport other pollutants make them a complex threat to marine ecosystems. Once released into the water column, these particles can interact with a wide variety of organisms, ranging from bacteria and phytoplankton to higher trophic levels up to humans, potentially altering ecological processes and compromising ecosystem functioning [1]. The Mediterranean Sea, due to its semi-enclosed nature, intense anthropogenic activities, and limited water exchange with the Atlantic Ocean, is considered one of the global hotspots for plastic pollution [2]. Recent surveys have reported high concentrations of MPs and NPs in both surface waters and sediments, raising concerns about their impact on organisms’ physiology, marine biodiversity and the sustainability of this already vulnerable basin [3]. Among plastic polymers, polyethylene (PE) is one of the most persistent and widely produced types, accounting for a substantial fraction of global plastic output and being extensively used in packaging, consumer goods, and industrial applications [4]. Owing to its low density and high resistance to degradation, PE is particularly prone to dispersion and long-term persistence in marine environments. Its ubiquity across diverse habitats, including freshwater and terrestrial ecosystems, enhances its potential for interactions and bioaccumulation across multiple trophic levels, possibly posing risks also to human health [5,6]. Addressing the accumulation of PE in such ecosystems is therefore an urgent environmental priority. Diatoms (class Bacillariophyceae) are unicellular photosynthetic microalgae that represent one of the most important groups of marine phytoplankton. They are responsible for an estimated 20–25% of global primary production and play a pivotal role in the cycling of carbon and silicon [7]. Their distinctive silica-based cell wall, or frustule, provides structural protection and contributes to species-specific ecological interactions. Within this diverse group, the genus Skeletonema (order Thalassiosirales) is widely distributed in coastal waters and is characterized by cells forming chains through delicate intercalary processes [8]. This chain-forming morphology favors buoyancy and bloom formation, allowing Skeletonema species to dominate highly dynamic and nutrient-rich environments. Among them, Skeletonema marinoi is one of the most abundant and ecologically significant species in temperate and coastal regions, including the Mediterranean Sea, the Baltic Sea, and the North Atlantic [8]. It frequently forms seasonal blooms that strongly influence coastal food webs and carbon fluxes [9]. S. marinoi exhibits remarkable ecological plasticity, being able to tolerate fluctuations in light, temperature, and nutrient availability [10,11]. Moreover, it produces resting stages (benthic cysts) that ensure survival under unfavorable conditions and contribute to the long-term persistence and genetic diversity of populations [12]. Because of its dominance, ecological importance, and well-studied physiology and genomics, Skeletonema has become a model diatom for investigating the effects of environmental stressors, including emerging contaminants such as N/MPs at the primary producer level [13]. The present study investigates the responses of S. marinoi to PE N/MPs, focusing on growth performance, morphological alterations, stress-related gene expression, and extracellular polymeric substance (EPS) production, a microalgae component that is essential for chain integrity, ecological interactions, and global biogeochemical cycles, especially carbon sequestration. By combining physiological and molecular endpoints, this study seeks to explore potential effects of plastic exposure on diatoms and to contribute to understanding how plastic pollution might influence phytoplankton physiology and ecology. Beyond advancing our mechanistic understanding, the outcomes of this research may help refine ecological risk assessments of plastic pollution and support the development of more effective strategies for preserving the integrity and resilience of marine ecosystems.
2. Materials and Methods
2.1. Cell Culturing and Harvesting
Two replicate Skeletonema marinoi (CCMP2092) cultures (Bigelow NCMA, National Center for Marine Algae and Microbiota, East Boothbay, ME, USA), with initial cell concentrations of 5000 cells/mL, were grown in two-liter polycarbonate bottles filled with sterile filtered oligotrophic seawater at 36 ppt salinity amended with f/2 nutrients, vitamins and metals [14], under gentle bubbling with sterile-filtered ambient air and maintained at 18–20 °C, 12:12 h light:dark photoperiod under 100 μmol photons m2 s−1 irradiance. In total, 1 mL culture, fixed with Lugol’s iodine solution (final concentration ~2%), was used to follow the growth of each replicate by daily cell enumeration in a Sedgewick-Rafter counting chamber (Hausser Scientific, Horsham, PA, USA) under an Axioskop 2 light microscope at 20× magnification (Carl Zeiss GmbH, Jena, Germany). Microscopic images of a sample were acquired daily at 40× magnification using the EVOS XL Core Imaging System (Thermo Fisher Scientific, Waltham, MA, USA) to monitor cell morphology. At established timepoints (day 3, 5, 7, and 10), 250 mL from each culture was harvested by filtration onto 0.22 μm nitrocellulose membranes (RAWP047, Millipore, Merck KGaA, Darmstadt, Germany).
2.2. Nano/Microplastics and Reagents
PE N/MPs, with a size range of 200 nm to 9900 nm (Cospheric LLC, Santa Barbara, CA, USA), were suspended in 1% Tween20 solution (Dako, Agilent Technologies, Santa Clara, CA, USA) to prepare a 100 mg/mL stock, according to the manufacturer’s instructions. This stock suspension was directly added to S. marinoi cultures to achieve final concentrations of 25 μg/mL (PE 25) and 75 μg/mL (PE 75). The final concentration of Tween20 in the algal cultures was kept below 0.01% to minimize any potential surfactant-induced toxicity. The stock solution was thoroughly vortexed prior to each use. A vehicle (Veh) control containing an equivalent amount of Tween20 (as present in the highest PE N/MP treatment) was included to exclude any confounding effects of the dispersant. Cultures of S. marinoi were exposed to Veh, PE 25 and PE 75 for a continuous 10-day period, with treatments initiated on day 0. Each condition was tested in three biological replicates.
2.3. Selection of Candidate CAZyme Genes Implicated in EPS Biosynthesis
To identify candidate genes potentially involved in EPS production in S. marinoi, a comparative in silico strategy was adopted. As S. marinoi is not currently represented in the Carbohydrate Active enZymes (CAZy) database (www.cazy.org, accessed on 15/09/2025) [15], we selected Thalassiosira pseudonana—a phylogenetically related diatom species-as the reference organism. From the CAZy database, all amino acid motifs annotated for T. pseudonana were retrieved, and the corresponding mRNA sequences were identified. These sequences were subsequently queried against the S. marinoi transcriptome to search for homologous sequences [16]. Matches were identified for two CAZyme motifs assigned to the GT2 (glycosyltransferases) and GH31 (glycoside hydrolases) protein families. Further exploration of these CAZyme families within the CAZy database provided a list of associated protein functions and putative enzymes. This list was used to screen the functional annotations of the S. marinoi transcriptome, aiming to identify matches with protein functions characteristic of the GT2 and GH31 families. Given the high number of matches, only sequences plausibly involved in EPS biosynthesis were retained, as not all CAZymes are implicated in this process. A general literature review on EPS production in microalgae and diatoms highlighted fucosyltransferases and galactosyltransferases as the most frequently involved CAZymes in this context [17]. Most of the sequences identified by transcriptome annotation in S. marinoi corresponded to enzymes involved in N-glycosylation of proteins, a process not directly linked to EPS production [18]. Therefore, primers were designed and synthesized for only three selected sequences related to the functions of interest (Supplementary Material).
2.4. RNA Extraction, Reverse Transcription, and RT-qPCR Analysis
Cells collected on filters (as previously described) were lysed using 1.5 mL of TRIsure reagent (Meridian Life Sciences, Memphis, TN, USA), supplemented with glass beads (Sigma-Aldrich, St. Louis, MO, USA) to facilitate mechanical disruption. The mixture was incubated on a thermo-shaker (Eppendorf, Hamburg, Germany) at 60 °C for 10 min at 1200 rpm. Following cell disruption, filters and beads were removed by a brief centrifugation step, and RNA extraction proceeded according to the TRIsure protocol. To eliminate residual genomic DNA, samples were treated with recombinant DNase I (Norgen Biotek Corp., Thorold, ON, Canada) following the manufacturer’s instructions. The efficiency of DNA removal was verified via conventional PCR using DNA-targeted primers; the absence of amplification confirmed successful digestion. PCR products were analyzed via agarose gel electrophoresis. Following DNase treatment, RNA was purified and concentrated using the RNA Clean-Up and concentration kit (Norgen Biotek Corp., Thorold, ON, Canada) and eluted in 30 μL of RNase-free water. RNA concentration was measured by absorbance at 260 nm using a NanoDrop 2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA), and integrity was assessed by agarose gel electrophoresis. One microgram of purified RNA per sample was reverse transcribed into complementary DNA (cDNA) using the SensiFAST™ cDNA Synthesis Kit (Meridian Life Sciences, Memphis, TN, USA), according to the manufacturer’s protocol. Gene expression was evaluated by SYBR Green-based real-time quantitative PCR (RT-qPCR), using specific primers (Table 1) [19,20]. Amplifications were performed on a QuantStudio 1 Real-Time PCR System (Thermo Fisher Scientific, Waltham, MA, USA). Gene expression levels were calculated using the 2−ΔΔCT method, and results are expressed as fold changes relative to untreated control cells for each time point. Each reaction was carried out in technical triplicate, with a minimum of three independent biological replicates.

Table 1.
Candidate reference genes and genes of interest used for RT-qPCR. Primer sequences and correlation factor (R2) are reported.
2.5. Data Analysis and Statistics
Gene expression levels were quantified relative to the control condition (Veh) for each timepoint using the Relative Expression Software Tool (REST, v2) [21]. For normalization, we tested Actin (ACT), Elongation Factor 1 Alpha (EF1α), Histone H4 (H4), Ribosomal Protein S (RPS), TATA-box Binding Protein (TBP), and Ubiquitin (UB) as reference genes. Three different software tools—BestKeeper (v1) [22], NormFinder (v21) [23], and geNorm (v3.4) [24]—were used to identify the most suitable reference genes in each experimental set, following the approach described by Orefice et al. [19]. As regards the growth curve experiment, the data obtained were plotted in a semi-logarithmic scatter plot (log10), where cell concentration (number of cells/mL) was reported as a function of time (days). For each experiment, at least three replicates were performed, and all results are expressed as the mean ± SEM. Statistical significance was calculated with GraphPad Prism 8.0.1 (GraphPad Software Inc., San Diego, CA, USA) using Student’s t-test, and p-values less than 0.05 were considered statistically significant.
3. Results
3.1. Effects of PE N/MP Exposure on the Growth and Morphology of S. marinoi
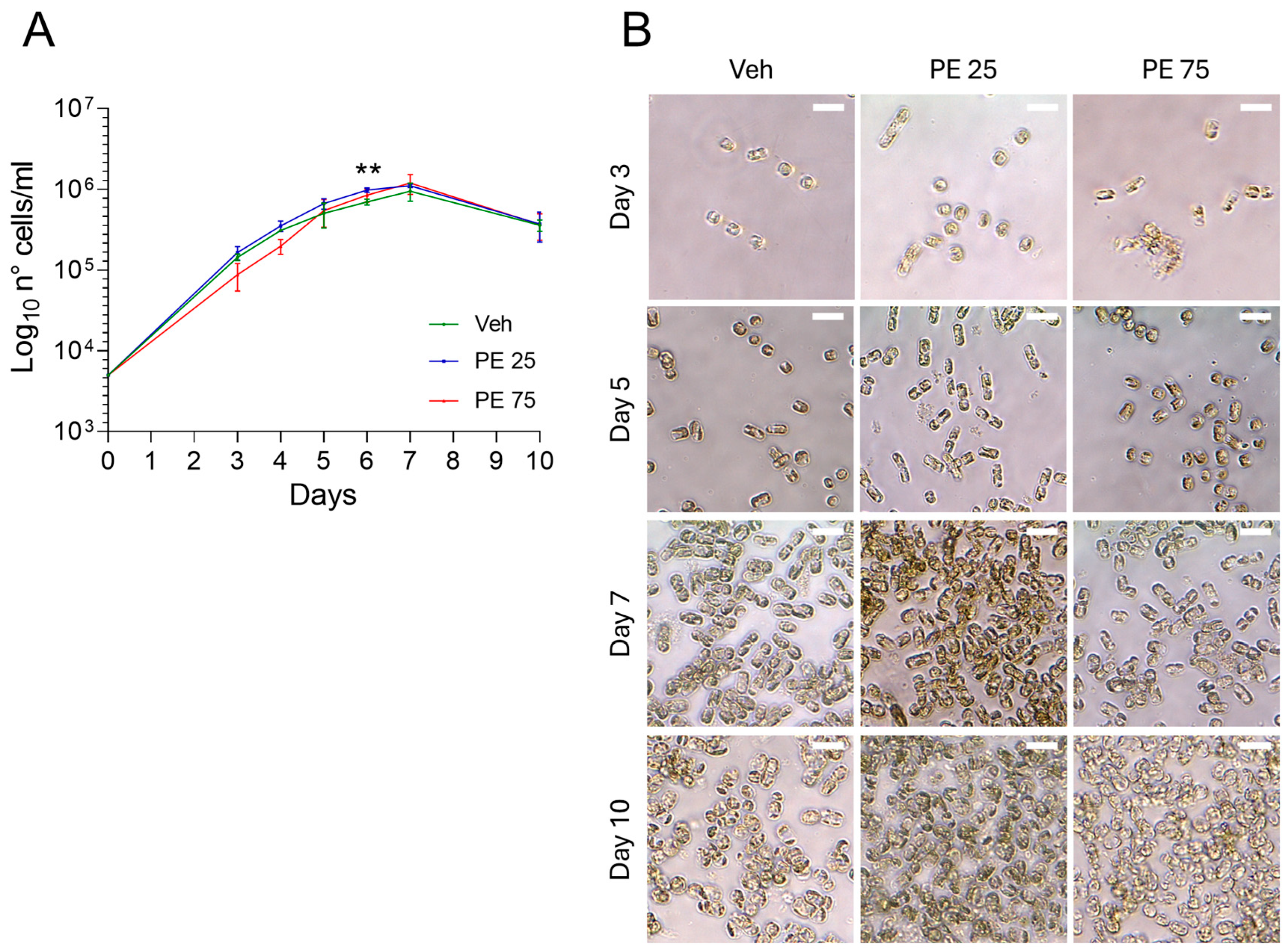
Both concentrations of PE, PE 25 and PE 75, did not impair the growth of S. marinoi, as all the cultures had comparable growth rates. The growth curves resulted in an exponential phase lasting 5 days, followed by 2 days of stationary phase until day 7, when cells started to exhibit a decline in viability. However, a statistically significant difference was observed at day 6, when PE 25-exposed cultures reached a higher cell concentration compared to both the Veh and PE 75 groups (Figure 1A). Microscopic inspection of S. marinoi cultures exposed to PE N/MPs did not reveal clear or consistent morphological alterations when compared to the Veh control (Figure 1B). Throughout the 10-day exposure period, cells largely maintained their typical chain organization and chloroplast morphology. While occasional variations in cell shape or chain length were observed, these appeared within the range of normal variability for the species. Overall, the optical microscopy observations suggest that the treatments did not induce significant morphological effects under the tested conditions.
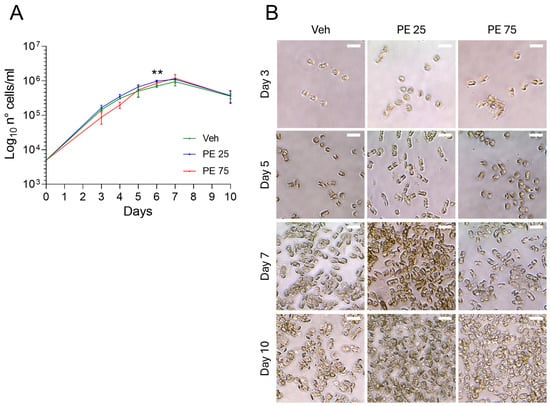
Figure 1.
Growth and Morphology of S. marinoi under PE N/MP Exposure: (A) Growth curve of S. marinoi cultures exposed to 25 μg/mL (PE 25; blue) and 75 μg/mL (PE 75; red) of PE N/MPs, or control Vehicle (Veh; green). Data are shown on a log10 scale. Statistical analyses were conducted by unpaired two-tailed Student’s t-test (n = 3; mean ± SEM) with ** p < 0.05. (B) Representative 40× micrographs of S. marinoi collected at four key time points. Scale bar = 10 μm.
Despite these findings, gene expression analyses revealed clear trends of molecular responses to PE N/MP exposure, although statistical significance was reached only for a subset of data points.
3.2. Stress and Inflammation Gene Responses in S. marinoi Exposed to PE N/MPs
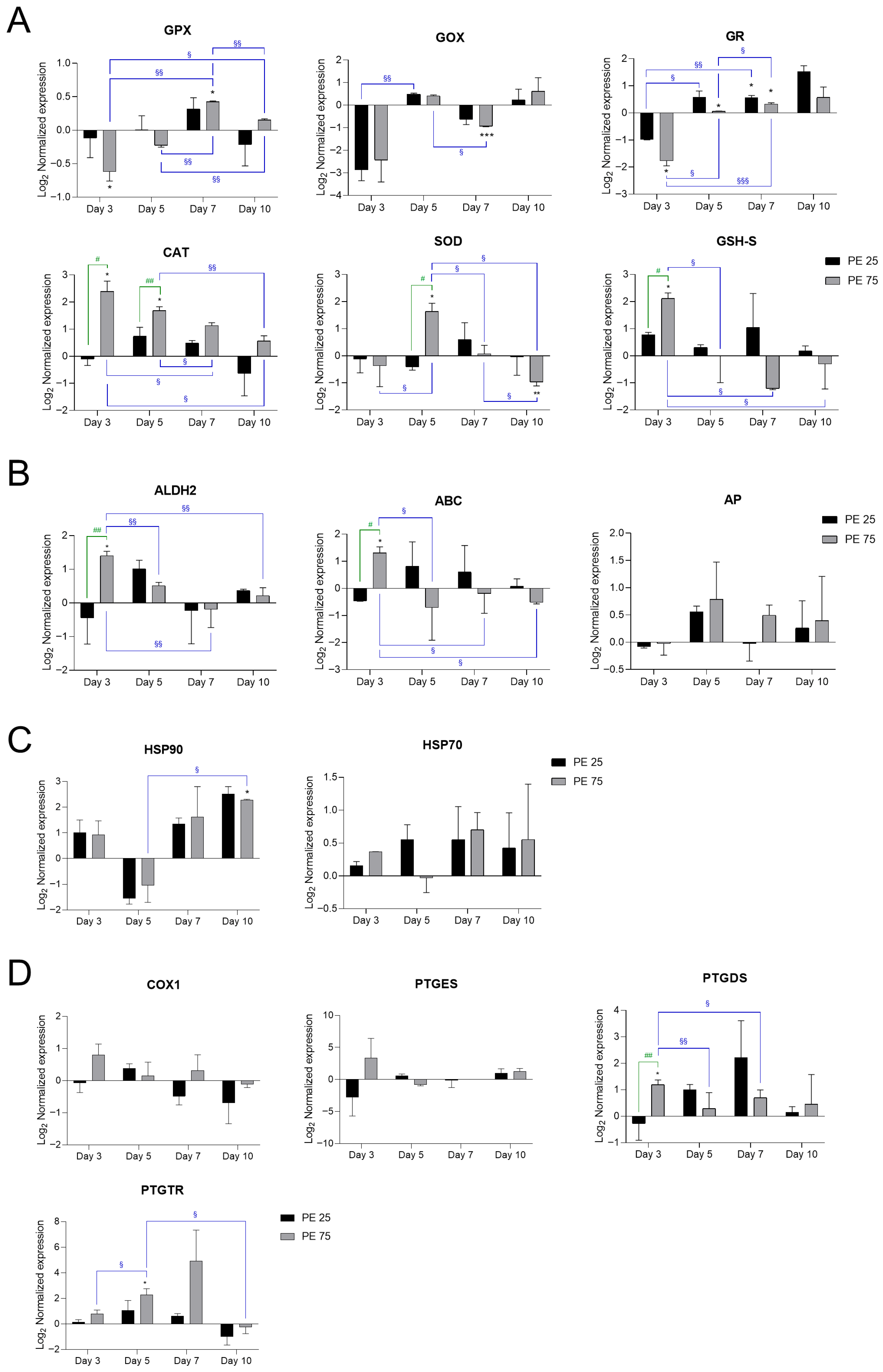
Antioxidant and stress-related genes, including glutathione peroxidase (GPX), glutathione oxidase (GOX), and glutathione reductase (GR), exhibited dynamic regulation in response to PE N/MP exposure (Figure 2A).
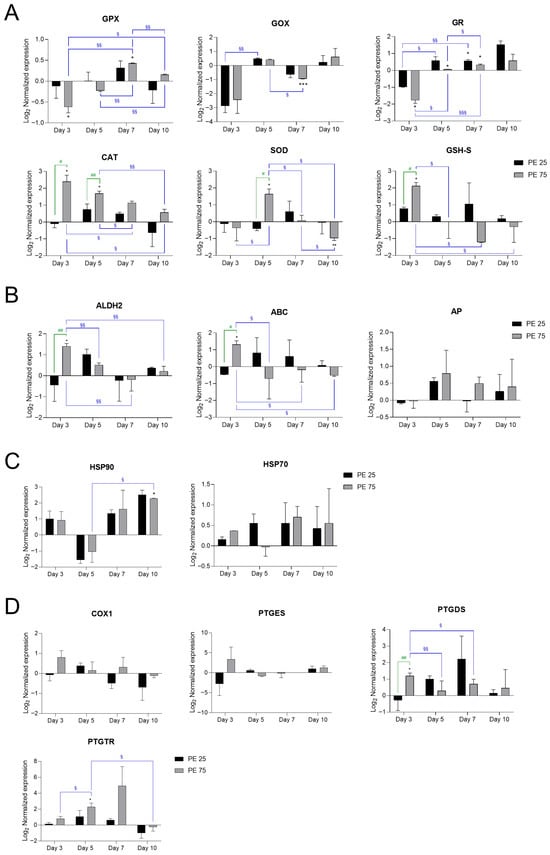
Figure 2.
RT-qPCR analysis of stress- and inflammation-related genes in S. marinoi exposed to PE N/MPs. Gene expression changes are presented in panels grouped according to their main biological functions: (A) Antioxidant defense: Glutathione peroxidase (GPX), Glutathione oxidase (GOX), Glutathione reductase (GR), Catalase (CAT), Superoxide dismutase (SOD), Glutathione synthetase (GHS-S); (B) Detoxification and transport: Aldehyde dehydrogenase 2 (ALDH2), ATP-binding cassette (ABC), Alkaline phosphatase (AP); (C) Protein stress response/chaperones: Heat shock protein 70 (HSP70), Heat shock protein 90 (HSP90); (D) Prostaglandin metabolism and signaling: Cyclooxygenase-1 (COX1), Prostaglandin D synthetase (PTGDS), Prostaglandin E synthetase (PTGES), Prostaglandin transporter (PTGTR). In the histograms, black bars represent PE 25 and gray bars represent PE 75. Values are reported on a log2 scale, with the zero-line set as the Veh reference. All the statistical analyses were conducted by unpaired, two-tailed Student’s t-test (n ≥ 3; mean ± SEM) with * p < 0.05, ** p < 0.005 and *** p < 0.001 for Veh vs. condition (black); § p < 0.05 and §§ p < 0.005 and §§§ p < 0.001 for timepoint vs. timepoint (blue); # p < 0.05 and ## p < 0.005 for PE 25 vs. PE 75 (green).
Initially, these genes were suppressed at day 3 with respect to Veh (zero-line), followed by a significant induction during the transition to the stationary phase (days 5–7), and a partial return to baseline levels by day 10. Notably, exposure to PE 75 led to more pronounced and significant changes in GPX, GOX, and GR activities, suggesting that the higher concentration imposed a stronger oxidative challenge. In contrast, PE 25 induced only limited or less consistent modulation, indicative of a milder stress response.
Among anti-oxidative stress genes, catalase (CAT) transcripts were strongly induced at days 3 and 5, with the highest expression observed under PE 75 exposure, followed by a gradual decline toward or below Veh levels by day 10. Superoxide dismutase (SOD) exhibited a similar pattern, with pronounced upregulation at day 5 under PE 75, decreasing markedly at day 7, and reaching significant downregulation by day 10. Glutathione synthetase (GSH-S) showed early and strong induction under PE 75 exposure, decreasing below basal levels at days 5–7, and partially recovering by day 10 (Figure 2A). Physiologically, SOD serves as the first line of defense by converting superoxide radicals into hydrogen peroxide (H2O2), which is then detoxified by both CAT and the glutathione system [25]. In our PE N/MP-exposed samples, CAT was induced earlier than SOD. This suggests that PE N/MP exposure may generate hydrogen peroxide or other ROS independently of superoxide, prompting an immediate activation of CAT to neutralize these reactive species. While H2O2 could theoretically also arise from abiotic/photo-oxidation of PE or interactions with the culture medium, under the experimental conditions used (10-day exposure under standard artificial light), such abiotic contributions are expected to be minimal. Therefore, the early CAT induction most likely reflects a cellular response to ROS generated by the algal metabolism in response to PE N/MP exposure.
Detoxification-related genes, such as Aldehyde dehydrogenase 2 (ALDH2), showed an early induction at day 3 under PE 75, suppression at days 5 and 7, and partial recovery at day 10 (Figure 2B). Moreover, ATP-binding cassette (ABC) transporter and alkaline phosphatase (AP) mirrored the temporal patterns observed in other gene categories. ABC transporter was strongly induced at day 3 under PE 75 exposure, with PE 25 showing expression levels close to Veh, followed by a steady decline over days 5–7, indicating downregulation. AP expression remained largely unchanged, with only minor and variable increases under PE 75 exposure (Figure 2B).
Heat shock protein 90 (HSP90) and Heat shock protein 70 (HSP70) are molecular chaperones that assist in protein folding and protect cells from stress-induced damage. HSP90 displayed modest increases from day 5 onward, becoming significantly different from Veh only at day 10. HSP70 showed a consistent trend of upregulation over time, although without attaining statistical significance (Figure 2C).
Prostaglandin-related genes, involved in inflammation and immune responses, including cyclooxygenase-1 (COX1), prostaglandin E synthase (PTGES), prostaglandin D synthase (PTGDS), and prostaglandin transporter (PTGTR), also displayed dynamic regulation (Figure 2D). COX1 remained stable across all treatments and time points. PTGES was transiently induced at day 3 under PE 75 exposure but showed high variability and no consistent modulation at later time points. PTGDS was strongly upregulated at day 3 in PE 75-exposed cultures compared to both Veh and PE 25 groups, followed by a progressive decline to near baseline levels by day 10. PTGTR exhibited a delayed response, peaking at day 7, particularly under PE 75 exposure, before sharply decreasing by day 10.
Overall, these findings suggest that PE N/MPs elicit a concentration-dependent, transient stress response in S. marinoi, occurring alongside the diatom’s natural growth cycle. High-dose exposure triggers robust but temporally limited defense mechanisms, likely limited by the diatom’s intrinsic physiological trajectory.
3.3. Differential Expression Dynamics of Programmed Cell Death-Related Genes in Response to PE N/MP Exposure
Expression analyses of programmed cell death (PCD)-related genes highlighted distinct molecular trajectories in response to PE N/MP exposure (Figure 3).
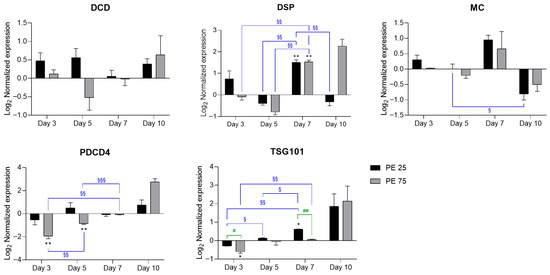
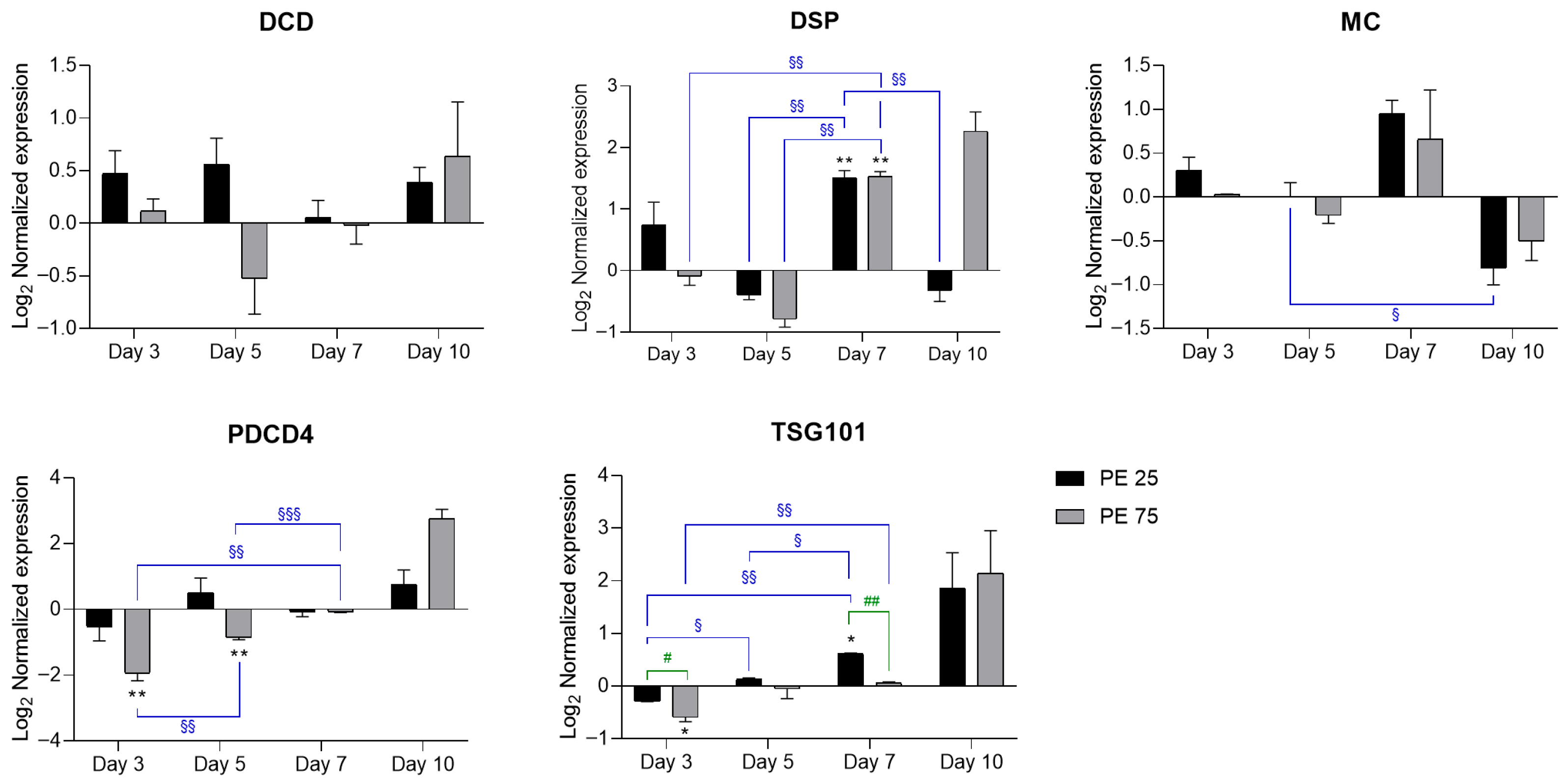
Figure 3.
RT-qPCR analysis of PCD-related genes in response to PE N/MPs. In the histograms, black bars represent PE 25 and gray bars represent PE 75. Values are reported on a log2 scale, with the zero-line set as the Veh reference. All the statistical analyses were conducted by unpaired, two-tailed Student’s t-test (n ≥ 3; mean ± SEM) with * p < 0.05 and ** p < 0.005 for Veh vs. condition (black); § p < 0.05 and §§ p < 0.005 and §§§ p < 0.001 for timepoint vs. timepoint (blue); # p < 0.05 and ## p < 0.005 for PE 25 vs. PE 75 (green).
Development and Cell Death Domain protein (DCD) expression remained largely stable, with the only notable change being a transient downregulation at day 5 in PE 75. At later time points (days 7–10), both PE 25 and PE 75 showed a trend toward upregulation, although differences did not reach statistical significance. Death-specific protease (DSP) showed a significant induction at day 7, reaching comparable levels in both PE 25 and PE 75 relative to Veh. By day 10, DSP remained elevated in PE 75 (even if not statistically significant), whereas PE 25 tended toward the Veh baseline. Metacaspase (MC) expression showed modest fluctuations, with a slight induction at day 7 in both treatments, followed by repression at day 10, consistent with the observed culture decline. Programmed cell death 4 (PDCD4) followed a distinct biphasic trend, characterized by downregulation at day 3—particularly under PE 75—followed by a progressive increase that culminated in a moderate—though not statistically significant—upregulation by day 10, coinciding with higher levels of aging-related stress. Finally, Tumor Susceptibility Gene 101 (TSG101) was initially repressed at day 3 in both treatments and significantly upregulated at day 7 under PE 25, with respect to Veh. Notably, by day 10, TSG101 exhibited a further increase, although not statistically significant, possibly reflecting persistent late-stage activation. Collectively, these profiles suggest that moderate exposure (PE 25) supports a more coordinated and transient activation of PCD-related pathways, whereas high exposure (PE 75) elicits an unbalanced response, with delayed and sustained activation of pro-death signals during the terminal growth phase.
3.4. Expression Analysis of Candidate CAZyme Genes Implicated in EPS Biosynthesis
RT-qPCR analysis revealed that exposure of S. marinoi cultures to PE N/MPs significantly affected the expression of putative genes possibly involved in EPS secretion pathways (Figure 4).
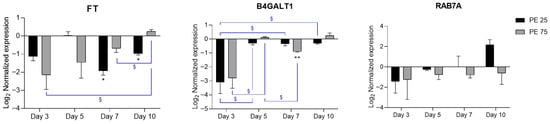

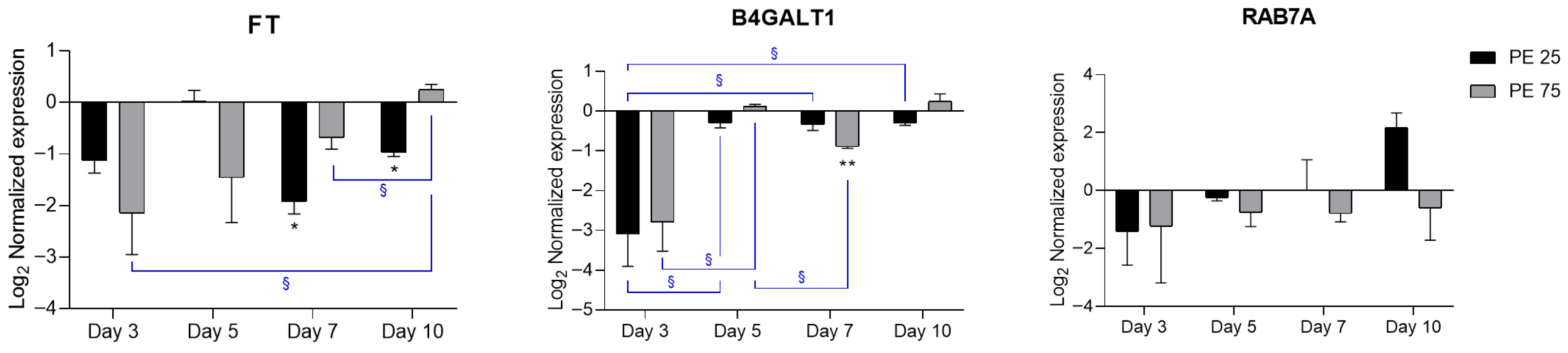
Figure 4.
RT-qPCR expression profiles of candidate CAZyme genes related to EPS biosynthesis. In the histograms, black bars represent PE 25 and gray bars represent PE 75. Values are reported on a log2 scale, with the zero-line set as the Veh reference. All the statistical analyses were conducted by unpaired, two-tailed Student’s t-test (n ≥ 3; mean ± SEM) with * p < 0.05 and ** p < 0.005 for Veh vs. condition (black); § p < 0.05 for timepoint vs. timepoint (blue).
Fucosyltransferase (FT) showed a downregulation trend at early time points, with statistical significance at Days 7 and 10 for PE 25, whereas Beta-1,4-galactosyltransferase 1 (B4GALT1) exhibited a decrease at Day 3, which reached significance at Day 7 for PE 75. Overall, both genes showed a recovery trend toward baseline expression by Day 10. Given their role in glycosylation and elongation of complex carbohydrates, this repression suggests an initial impairment in the biosynthetic machinery for EPS, which in diatoms is crucial for aggregation, biofilm formation, and particle interactions. The observed downregulation may thus reflect a stress response leading to reduced EPS production in the early stages of PE N/MP exposure. In contrast, Ras-related protein Rab-7a (RAB7A), a regulator of vesicular trafficking, displayed high variability without clear significant changes, yet its role in endosomal transport suggests that subtle modulation of trafficking processes cannot be excluded, potentially affecting the delivery of EPS-related enzymes or polysaccharides to the cell surface. It should be noted that, among the genes analyzed, only FT and B4GALT1 can be classified as bona fide CAZymes directly implicated in glycosylation. In contrast, RAB7A, while not a glycosyltransferase, may indirectly affect EPS production by modulating vesicular transport and secretion of EPS-related enzymes or polysaccharides. Interestingly, RAB7A displayed a deregulation pattern similar to PDCD4 and TSG101 genes, which are involved in vesicular trafficking and cellular stress responses, suggesting a possible activation of membrane remodeling and autophagy-related pathways at later exposure stages. Together, these data support the view that PE N/MP exposure might transiently disrupt glycosylation and secretion pathways, potentially impairing EPS production and remodeling in S. marinoi.
4. Discussion
Diatoms of the genus Skeletonema, including S. marinoi, play a pivotal role in primary carbon fixation and in regulating oceanic biogeochemical cycles. Increasing evidence indicates that these diatoms are sensitive to perturbations induced by N/MPs [26,27]. Similar stress responses occur under environmental drivers like nutrient or silicate limitation [28,29]. These observations highlight the interplay between plastic stress and natural environmental constraints in shaping diatom responses. Within this context, our study investigated the physiological and molecular effects of exposure to PE N/MPs in S. marinoi. Continuous exposure for 10 days did not significantly alter overall growth trends and morphology, yet transcriptional signatures emerged. Growth curves showed no persistent inhibition, but a transient increase in cell number was observed on day 6 at the lower concentration (PE 25). This finding is consistent with reports that moderate N/MP exposure often spares growth in microalgae [30]. A plausible explanation is that a mild oxidative challenge triggered a timely activation of antioxidant defenses, transiently enhancing resilience in a hormetic-like manner-a phenomenon also described in other microalgae under sublethal stress. However, this short-lived “physiological boost” was followed by a decline, consistent with the natural transition toward the stationary and death phases. Microscopy has previously revealed shorter chains, irregular chloroplasts, and cell shrinkage in microalgae exposed to N/MPs [31,32]. In contrast, in our S. marinoi cultures, no significant morphological alterations were detected under optical microscopy. However, optical microscopy provides limited resolution and may not capture fine-scale structural changes, such as alterations in frustule ultrastructure or subcellular features. More detailed imaging techniques, such as electron microscopy, would be required to fully assess subtle morphological responses to PE N/MP exposure.
Interestingly, at the molecular level, PE N/MP exposure interfered with stress- and antioxidant-related pathways in a concentration-dependent fashion. Key enzymes such as CAT, SOD, and glutathione-related proteins (GPX, GOX, GR, GSH-S) displayed biphasic expression patterns, with early induction followed by attenuation or downregulation. This dynamic mirrors responses reported in Thalassiosira pseudonana exposed to PS MPs [33], supporting the idea of a conserved oxidative stress program across diatoms. Heat shock proteins (HSP70, HSP90) were progressively induced, consistent with proteostatic adjustments. The timing of these responses, coinciding with the transition from exponential to stationary growth, suggests that PE exposure amplifies the physiological stress already imposed by nutrient depletion and population density.
Beyond antioxidant defenses, the regulation of prostaglandin-related genes (COX1, PTGES, PTGDS, PTGTR) points to an inflammatory-like signaling dimension in S. marinoi. Their transient upregulation strengthens the emerging view that diatoms employ oxylipin-based pathways as part of stress and defense responses [34] and that plastics may act as bioactive stressors rather than inert pollutants. Furthermore, oxidative stress in diatoms can drive the accumulation of lipid peroxidation products such as isoprostanoids, as reported in Phaeodactylum tricornutum. Even in species lacking canonical prostaglandin synthesis pathways, these non-enzymatic compounds accumulate, underscoring the chemical complexity of diatom stress responses [35]. Interestingly, exposure to PE N/MPs resulted in specific upregulation of PTGDS, whereas COX1 expression remained unchanged. This pattern suggests that the regulation of prostaglandin biosynthesis in this case may occur primarily at the level of the terminal synthase rather than the upstream cyclooxygenase. COX1, often constitutively expressed, may provide sufficient substrate (PGH2) for PTGDS activity, while the selective induction of PTGDS could represent a rapid mechanism to increase PGD2 production in response to PE N/MP-induced stress, without the need to modulate COX1 transcription.
An interesting outcome of this study is the transcriptional modulation of programmed cell death (PCD)-related genes, which followed distinct temporal trajectories. At high exposure (PE 75), late upregulation of PDCD4 and TSG101 was observed, coinciding with culture decline. Although these increases were not statistically significant, they suggest a trend whereby PE N/MPs may accelerate senescence and promote maladaptive activation of cell death programs. Conversely, the moderate exposure (PE 25) produced a more transient and coordinated activation of PCD markers, resembling an adaptive strategy to cope with stress. Similar dysregulation of PCD has been reported in other microalgae under plastic exposure [36], highlighting the ecological relevance of PCD in bloom termination and carbon export.
A novel important aspect of our study is the downregulation of candidate CAZyme genes, particularly FT and B4GALT1, during the initial phases of PE N/MP exposure (days 3–7), suggesting impaired EPS biosynthesis. EPS underpins diatom ecology by mediating aggregation, biofilm stability, and particle interactions [37,38]. Their repression under early PE N/MP exposure could collectively reduce aggregation capacity and hinder vertical fluxes, thereby weakening the biological carbon pump. From an ecological perspective, reduced EPS production may also alter colony morphology and chain formation, potentially influencing grazing susceptibility and predator-prey interactions. Previous work has shown that plastics can modulate EPS quantity and composition, altering hetero aggregation with particles and the sinking flux of organic carbon [39,40]. It should also be noted that abiotic factors can enhance EPS productivity in several microalgal species, since EPS secretion can function as a protective response to environmental stress [41]. Based on this framework, one might have expected to observe an increase in EPS production under PE N/MP exposure as well. This apparent discrepancy highlights the complexity of EPS regulation in diatoms. Moreover, only three CAZyme genes were analyzed in our study; while they are very likely to be directly involved in key steps of glycosylation and EPS biosynthesis and provide insight into core molecular responses, they do not represent the full EPS biosynthetic network. Importantly, CAZymes have never been functionally characterized in S. marinoi, nor, to our knowledge, in diatoms in general. This study, therefore, represents the first targeted analysis of EPS-related CAZymes in S. marinoi. These three genes were among the few with reliable annotation and robust expression in the transcriptome available at the time of the study. With the recent sequencing of the S. marinoi genome, more comprehensive and accurate annotation will soon allow a deeper exploration of the full EPS biosynthetic network. Additionally, the study relied exclusively on molecular analyses to infer EPS production, as direct biochemical quantification in S. marinoi is technically challenging due to its small size, dynamic EPS secretion, and the lack of established protocols. Consequently, extrapolation of gene expression patterns to actual EPS output should be made with caution. Broader transcriptomic or enzymatic analyses will be addressed to describe the complete complexity of EPS production in diatoms and to clarify the impact of PE N/MP exposure on S. marinoi.
Taken together, our findings indicate that PE N/MPs elicit subtle yet multifaceted effects in S. marinoi. Although population growth was not strongly inhibited, plastic pollution modulated stress and detoxification pathways, inflammatory signaling, PCD activation, and EPS biosynthesis. These responses were concentration-dependent and transient, reflecting both the plasticity of diatom stress responses and the risk of maladaptation under sustained or high-intensity exposure. Future work should integrate transcriptomic, proteomic, metabolomic, and live-cell imaging approaches to validate the functional and biological impact of the transcriptional changes and to assess whether similar responses are conserved across diatom taxa. Finally, it is important to acknowledge the main limitations of this study. Firstly, the concentrations of PE N/MPs we employed might be higher than those typically occurring in the marine environment; however, given the current lack of precise quantification of N/MP levels in the Mediterranean Sea and their expected variability between more and less polluted areas, we deliberately adopted elevated concentrations to highlight the potential effects on the diatom. Moreover, we focused on spherical, well-characterized PE particles to ensure reproducibility and mechanistic clarity. However, plastics in the environment occur in diverse and irregular forms, such as fibers and fragments, which may interact differently with diatom frustules and EPS matrices. These heterogeneous shapes, common in real-world matrices, could elicit distinct or complementary responses, and future research should address their ecological relevance. Moreover, investigating the combined effects of plastics with other environmental stressors—such as nutrient limitation, metal contamination, or ocean acidification—will be essential to capture the complexity of diatom responses in natural systems. Finally, short-term exposure limits our understanding of potential long-term or evolutionary effects, and findings from S. marinoi may not be directly generalizable to other diatom species or phytoplankton groups. Future studies integrating biochemical, physiological, and ecological endpoints over longer time scales and under more realistic environmental conditions are needed to fully assess the implications of PE N/MP exposure for diatoms.
5. Conclusions
This study demonstrates that S. marinoi exhibits a multifaceted response to continuous exposure to PE N/MPs. While overall growth and morphology were not markedly affected, PE N/MP exposure induced modulation of antioxidant and stress-related pathways, transient impairment of EPS biosynthesis, and activation of PCD markers. These effects were concentration-dependent and largely dynamic, highlighting both the resilience and the vulnerability of diatoms when coping with plastic-induced stress. Given the ecological importance of S. marinoi in primary production, aggregation processes, and carbon export, such responses may have broader implications for bloom dynamics and biogeochemical cycles in plastic-polluted oceans. Future studies should address environmentally realistic plastic morphologies and their interactions with additional stressors to better predict the fate of marine diatom populations under global change scenarios.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/microplastics5010033/s1. File S1: “Comparative in silico retrieval and selection of putative EPS-related CAZymes in S. marinoi: sequences of interest”.
Author Contributions
Conceptualization, F.M. and S.C.; methodology, P.P., V.D.D. and C.L.; validation, F.C. and E.R.; formal analysis, P.P.; investigation, P.P., V.D.D., C.L., F.C. and E.R.; resources, S.C.; data curation, P.P.; writing—original draft preparation, P.P.; writing—review and editing, V.D.D., C.L., A.A., C.M. and F.M.; visualization, P.P., F.C. and E.R.; supervision, S.C. and F.M.; project administration, A.A., C.M. and F.M.; funding acquisition, A.A., C.M. and F.M. All authors have read and agreed to the published version of the manuscript.
Funding
This study was conducted under the funding of the Italian Ministry of University and Research (MUR) through the PON “Ricerca e Innovazione” 2014–2020 program for the implementation of “Action IV.6–Research contracts on Green topics,” contract no. B85F21004490001; Sapienza University of Rome—“Ricerca di Ateneo” 2023 (Prot. RM123188F1F65F5C); Ministero dell’Università e della Ricerca-PRIN 2022 (Prot. 2022KCP7YP).
Data Availability Statement
The data presented in this study are available upon reasonable request from the corresponding authors.
Acknowledgments
The authors thank Mariano Amoroso and Arianna Smerilli from Stazione Zoologica Anton Dohrn for their technical support.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| ABC | ATP-binding cassette |
| ACT | Actin |
| ALDH2 | Aldehyde dehydrogenase 2 |
| AP | Alkaline phosphatase |
| B4GALT1 | Beta-1,4-galactosyltransferase 1 |
| CAT | Catalase |
| CAZy | Carbohydrate Active enZymes |
| COX1 | Cyclooxygenase-1 |
| DCD | Cell Death Domain protein |
| DSP | Death-specific protease |
| EF1α | Elongation Factor 1 Alpha |
| EPS | Extracellular polymeric substance |
| FT | Fucosyltransferase |
| GOX | Glutathione oxidase |
| GPX | Glutathione peroxidase |
| GR | Glutathione reductase |
| GSH-S | Glutathione synthetase |
| H4 | Histone H4 |
| HSP70 | Heat shock protein 70 |
| HSP90 | Heat shock protein 90 |
| MC | Metacaspase |
| MPs | Microplastics |
| NPs | Nanoplastics |
| PDCD4 | Programmed cell death 4 |
| PE | Polyethylene |
| PS | Polystyrene |
| PTGDS | Prostaglandin D synthase |
| PTGES | Prostaglandin E synthase |
| PTGTR | Prostaglandin transporter |
| RAB7A | Ras-related protein Rab-7a |
| ROS | Reactive oxygen species |
| RPS | Ribosomal Protein S |
| RT-qPCR | Real-time quantitative PCR |
| SOD | Superoxide dismutase |
| TBP | TATA-box Binding Protein |
| TSG101 | Tumor Susceptibility Gene 101 |
| UB | Ubiquitin |
| Veh | Vehicle |
References
- Hossain, M.; Engelhardt, I. Global plastic footprint: Unveiling property trends, environmental fate, and emerging threats of microplastic and nanoplastics pollution across ecosystems. Energy Ecol. Environ. 2025, 10, 637–674. [Google Scholar] [CrossRef]
- Sharma, S.; Sharma, V.; Chatterjee, S. Microplastics in the Mediterranean Sea: Sources, Pollution Intensity, Sea Health, and Regulatory Policies. Front. Mar. Sci. 2021, 8, 634934. [Google Scholar] [CrossRef]
- Chatziparaskeva, G.; Papamichael, I.; Zorpas, A.A. Microplastics in the coastal environment of Mediterranean and the impact on sustainability level. Sustain. Chem. Pharm. 2022, 29, 100768. [Google Scholar] [CrossRef]
- Tiago, G.A.O.; Mariquito, A.; Martins-Dias, S.; Marques, A.C. The problem of polyethylene waste—Recent attempts for its mitigation. Sci. Total Environ. 2023, 892, 164629. [Google Scholar] [CrossRef]
- Pontecorvi, P.; Ceccarelli, S.; Cece, F.; Camero, S.; Lotti, L.V.; Niccolai, E.; Nannini, G.; Gerini, G.; Anastasiadou, E.; Scialis, E.S.; et al. Assessing the Impact of Polyethylene Nano/Microplastic Exposure on Human Vaginal Keratinocytes. Int. J. Mol. Sci. 2023, 24, 11379. [Google Scholar] [CrossRef] [PubMed]
- Traversa, A.; Mari, E.; Pontecorvi, P.; Gerini, G.; Romano, E.; Megiorni, F.; Amedei, A.; Marchese, C.; Ranieri, D.; Ceccarelli, S. Polyethylene Micro/Nanoplastics Exposure Induces Epithelial-Mesenchymal Transition in Human Bronchial and Alveolar Epithelial Cells. Int. J. Mol. Sci. 2024, 25, 10168. [Google Scholar] [CrossRef]
- Leblanc, K.; Quéguiner, B.; Diaz, F.; Cornet, V.; Michel-Rodriguez, M.; de Madron, X.D.; Bowler, C.; Malviya, S.; Thyssen, M.; Grégori, G.; et al. Nanoplanktonic diatoms are globally overlooked but play a role in spring blooms and carbon export. Nat. Commun. 2018, 9, 953. [Google Scholar] [CrossRef]
- Sarno, D.; Kooistra, W.H.C.F.; Medlin, L.K.; Percopo, I.; Zingone, A. Diversity in the genus Skeletonema (bacillariophyceae). Ii. An assessment of the taxonomy of s. Costatum-like species with the description of four new species. J. Phycol. 2005, 41, 151–176. [Google Scholar] [CrossRef]
- Lauritano, C.; Romano, G.; Roncalli, V.; Amoresano, A.; Fontanarosa, C.; Bastianini, M.; Braga, F.; Carotenuto, Y.; Ianora, A. New oxylipins produced at the end of a diatom bloom and their effects on copepod reproductive success and gene expression levels. Harmful Algae 2016, 55, 221–229. [Google Scholar] [CrossRef]
- Armbrust, E.V. The life of diatoms in the world’s oceans. Nature 2009, 459, 185–192. [Google Scholar] [CrossRef]
- Chandrasekaran, R.; Barra, L.; Carillo, S.; Caruso, T.; Corsaro, M.M.; Piaz, F.D.; Graziani, G.; Corato, F.; Pepe, D.; Manfredonia, A.; et al. Light modulation of biomass and macromolecular composition of the diatom Skeletonema marinoi. J. Biotechnol. 2014, 192, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Godhe, A.; Härnström, K. Linking the planktonic and benthic habitat: Genetic structure of the marine diatom Skeletonema marinoi. Mol. Ecol. 2010, 19, 4478–4490. [Google Scholar] [CrossRef] [PubMed]
- Parikh, H.S.; Dave, G.; Tiwari, A. Microplastic pollution in aquatic ecosystems: Impacts on diatom communities. Environ. Monit. Assess. 2025, 197, 206. [Google Scholar] [CrossRef]
- Guillard, R.R.L. Culture of Phytoplankton for Feeding Marine Invertebrates. In Culture of Marine Invertebrate Animals: Proceedings—1st Conference on Culture of Marine Invertebrate Animals Greenport; Smith, W.L., Chanley, M.H., Eds.; Springer: Boston, MA, USA, 1975; pp. 29–60. [Google Scholar] [CrossRef]
- Cantarel, B.L.; Coutinho, P.M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-Active EnZymes database (CAZy): An expert resource for Glycogenomics. Nucleic Acids Res. 2009, 37, D233–D238. [Google Scholar] [CrossRef]
- Keeling, P.J.; Burki, F.; Wilcox, H.M.; Allam, B.; Allen, E.E.; Amaral-Zettler, L.A.; Armbrust, E.V.; Archibald, J.M.; Bharti, A.K.; Bell, C.J.; et al. The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): Illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 2014, 12, e1001889. [Google Scholar] [CrossRef]
- Xiao, R.; Zheng, Y. Overview of microalgal extracellular polymeric substances (EPS) and their applications. Biotechnol. Adv. 2016, 34, 1225–1244. [Google Scholar] [CrossRef]
- van Bockstaele-Fuentes, J.; Mati-Baouche, N.; Lupette, J.; Gargouch, N.; Rivet, E.; Lerouge, P.; Bardor, M. An overview of protein N-glycosylation diversity in microalgae. Front. Plant Sci. 2025, 16, 1669918. [Google Scholar] [CrossRef] [PubMed]
- Orefice, I.; Lauritano, C.; Procaccini, G.; Ianora, A.; Romano, G. Insights into possible cell-death markers in the diatom Skeletonema marinoi in response to senescence and silica starvation. Mar. Genom. 2015, 24, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Lauritano, C.; Orefice, I.; Procaccini, G.; Romano, G.; Ianora, A. Key genes as stress indicators in the ubiquitous diatom Skeletonema marinoi. BMC Genom. 2015, 16, 411. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST©) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002, 30, e36. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper--Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034. [Google Scholar] [CrossRef]
- Koletti, A.; Skliros, D.; Dervisi, I.; Roussis, A.; Flemetakis, E. Oxidative Stress Responses in Microalgae: Modern Insights into an Old Topic. Appl. Microbiol. 2025, 5, 37. [Google Scholar] [CrossRef]
- Bellingeri, A.; Casabianca, S.; Capellacci, S.; Faleri, C.; Paccagnini, E.; Lupetti, P.; Koelmans, A.A.; Penna, A.; Corsi, I. Impact of polystyrene nanoparticles on marine diatom Skeletonema marinoi chain assemblages and consequences on their ecological role in marine ecosystems. Environ. Pollut. 2020, 262, 114268. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zheng, X.; Liu, X.; Zhang, L.; Zhang, S.; Li, Y.; Koelmans, A.A.; Penna, A.; Corsi, I. Effect and mechanism of microplastics exposure against microalgae: Photosynthesis and oxidative stress. Sci. Total Environ. 2023, 905, 167017. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Chen, F.; Mi, T.; Liu, Q.; Yu, Z.; Zhen, Y. Responses of Marine Diatom Skeletonema marinoi to Nutrient Deficiency: Programmed Cell Death. Appl. Environ. Microbiol. 2020, 86, e02460-19. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Ma, J.; Zhong, Z.; Liu, H.; Miao, A.; Zhu, X.; Pan, K. Silicon Limitation Impairs the Tolerance of Marine Diatoms to Pristine Microplastics. Environ. Sci. Technol. 2023, 57, 3291–3300. [Google Scholar] [CrossRef]
- Reichelt, S.; Gorokhova, E. Micro- and Nanoplastic Exposure Effects in Microalgae: A Meta-Analysis of Standard Growth Inhibition Tests. Front. Environ. Sci. 2020, 8, 131. [Google Scholar] [CrossRef]
- Sun, Z.; Zhang, S.; Zheng, T.; He, C.; Xu, J.; Lin, D.; Zhang, L. Nanoplastics inhibit carbon fixation in algae: The effect of aging. Heliyon 2024, 10, e29814. [Google Scholar] [CrossRef]
- Nguyen, M.K.; Moon, J.-Y.; Lee, Y.-C. Microalgal ecotoxicity of nanoparticles: An updated review. Ecotoxicol. Environ. Saf. 2020, 201, 110781. [Google Scholar] [CrossRef]
- Zhang, B.; Tang, X.; Liu, Q.; Li, L.; Zhao, Y.; Zhao, Y. Different effecting mechanisms of two sized polystyrene microplastics on microalgal oxidative stress and photosynthetic responses. Ecotoxicol. Environ. Saf. 2022, 244, 114072. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Albarano, L.; Esposito, R.; Zupo, V.; Costantini, M.; Ianora, A. Multiple Roles of Diatom-Derived Oxylipins within Marine Environments and Their Potential Biotechnological Applications. Mar. Drugs 2020, 18, 342. [Google Scholar] [CrossRef]
- Lupette, J.A.-O.X.; Jaussaud, A.A.-O.; Vigor, C.; Oger, C.; Galano, J.M.; Réversat, G.; Vercauteren, J.; Jouhet, J.; Durand, T.; Maréchal, E. Non-Enzymatic Synthesis of Bioactive Isoprostanoids in the Diatom Phaeodactylum following Oxidative Stress. Plant Physiol. 2018, 178, 1344–1357. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Tan, L.; Han, X.; Wang, X.; Zhang, Y.; Ma, X.; Lin, K.; Wang, R.; Ni, Z.; Wang, J.; et al. Microplastic-induced apoptosis and metabolism responses in marine Dinoflagellate, Karenia mikimotoi. Sci. Total Environ. 2022, 804, 150252. [Google Scholar] [CrossRef] [PubMed]
- Bahulikar, R.A.; Kroth, P.G. Localization of EPS components secreted by freshwater diatoms using differential staining with fluorophore-conjugated lectins and other fluorochromes. Eur. J. Phycol. 2007, 42, 199–208. [Google Scholar] [CrossRef]
- Ge, R.; Gong, Z.; Feng, Q.; Wang, S.; Jin, C.; Wei, J. Particle aggregation induced by microorganisms is a key mechanism for the biostabilization of coastal sediment. Sediment. Geol. 2024, 464, 106630. [Google Scholar] [CrossRef]
- Guo, M.; Noori, R.; Abolfathi, S. Microplastics in freshwater systems: Dynamic behaviour and transport processes. Resour. Conserv. Recycl. 2024, 205, 107578. [Google Scholar] [CrossRef]
- Demir-Yilmaz, I.; Yakovenko, N.; Roux, C.; Guiraud, P.; Collin, F.; Coudret, C.; ter Halle, A.; Formosa-Dague, C. The role of microplastics in microalgae cells aggregation: A study at the molecular scale using atomic force microscopy. Sci. Total Environ. 2022, 832, 155036. [Google Scholar] [CrossRef]
- Garza-Rodríguez, Z.B.; Hernández-Pérez, J.; Santacruz, A.; Jacobo-Velázquez, D.A.; Benavides, J. Prospective on the application of abiotic stresses to enhance the industrial production of exopolysaccharides from microalgae. Curr. Res. Biotechnol. 2022, 4, 439–444. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.