Abstract
Artificial Intelligence (AI) and Precision Medicine are increasingly influencing pediatric pharmacotherapy, where age-dependent pharmacokinetic variability demands highly individualized therapeutic strategies. This review examines current applications of AI in pediatric precision medicine and evaluates their clinical relevance and translational challenges. Recent evidence shows substantial progress across multiple domains. In pharmacogenomics, predictive models have reached R2 = 0.95 for drug exposure. Tools for adverse drug reaction detection report sensitivities of 81.5% and specificities of 79.5%. Clinical decision support systems for pediatric epilepsy have achieved diagnostic accuracies of 93.4%. Real-world implementations have been associated with a 75% reduction in prescription distribution errors and a 65% improvement in adverse drug reaction detection. Despite these advances, clinical translation remains limited: only 0.38% of pediatric AI models progress to testing in real patients, and 77% of published studies carry a high risk of bias. These gaps highlight the need for rigorous validation, improved data quality, and careful consideration of ethical and algorithmic constraints. Overall, AI has the potential to shift pediatric pharmacotherapy from empirically driven decisions toward predictive, precision-based approaches. Achieving this goal will require well-designed pediatric studies and sustained interdisciplinary collaboration to ensure safe and effective integration into clinical practice.
1. Introduction
The landscape of pharmacological therapy continues to evolve, driven by technological innovations and an expanding understanding of human biological variability. In this context, Artificial Intelligence (AI) and Precision Medicine emerge as foundational pillars for redefining healthcare approaches, particularly in pediatric populations where challenges are both unique and complex. The World Health Organization has emphasized AI’s transformative potential in healthcare, highlighting how big data utilization and analytical techniques can optimize diagnostic and therapeutic processes, including the exploration of new therapies and drugs, identification of optimal treatments based on predicted patient responses, and improvement of public health monitoring [1].
Children cannot be considered simply “small adults,” as substantial age-related variations exist across all pharmacokinetic phases—absorption, distribution, metabolism, and elimination. In neonates, reduced gastric acid secretion can increase the bioavailability of acid-sensitive drugs while decreasing that of weakly acidic compounds. Similarly, reduced bile salt formation can diminish the bioavailability of lipophilic drugs. Renal plasma flow and glomerular filtration rate are significantly lower at birth and gradually increase, reaching levels comparable to those of adults by around one year of age [2]. These profound physiological differences necessitate highly personalized dosing and therapeutic approaches for pediatric patients [3].
The recognition that children exhibit distinct disease symptoms, drug responses, and recovery patterns compared to adults underscores the critical importance of precision medicine in pediatrics [4]. Currently, there persists a significant unmet need for safe and effective drugs specifically developed for preventing and treating diseases affecting children. This gap between pediatric physiological complexity and available therapeutic solutions creates an imperative for AI implementation, not merely as a technological advancement, but as an ethical and clinical necessity to ensure safety and efficacy in one of the most vulnerable patient populations. Moreover, persistent limitations in pediatric pharmacology—such as the scarcity of age-specific trials, the widespread reliance on off-label prescribing, and the difficulty of predicting drug response across rapidly changing developmental stages—raise key questions that motivate this review. Specifically, how AI can support pediatric drug development, improve the prediction of pharmacokinetic and pharmacodynamic parameters, enhance early detection of adverse drug events, and address the ethical and regulatory challenges of clinical implementation. These considerations define the conceptual framework for the present work.
2. Methods
We conducted a narrative review of the literature, drawing on recent publications that explore the application of artificial intelligence in pediatric pharmacotherapy. Searches were performed primarily in PubMed using broad combinations of terms related to artificial intelligence (“artificial intelligence,” “machine learning,” “deep learning”) and pediatric pharmacological domains (“pediatric,” “children,” “pharmacotherapy,” “precision medicine,” “pharmacogenomics,” “adverse drug reaction,” “clinical decision support”).
Eligible works included peer-reviewed articles and high-quality preprints presenting original research or comprehensive reviews relevant to AI in pediatric drug therapy. Adult-only studies and articles unrelated to pharmacotherapy were not considered, in keeping with the narrative scope of this review. Evidence was synthesized thematically according to major application areas—such as pharmacogenomics, adverse drug reaction prediction, drug discovery support, and clinical decision-making—and contextualized across key pediatric disease domains.
3. Artificial Intelligence and Precision Medicine: Theoretical Foundations and Clinical Convergence
AI in healthcare refers to the application of advanced algorithms and systems capable of processing complex information to support every phase of patient management, from diagnosis to treatment [5]. The technology’s core strength lies in analyzing vast and complex datasets to identify subtle, non-obvious correlations—a fundamental process for developing personalized therapies and refining treatment strategies and clinical outcomes. Neural networks have been successfully employed to process histopathological images, discriminating disease presence, including rare conditions, with high accuracy [6].
Machine-Learning (ML) and Deep-Learning (DL) models constitute the backbone of these applications, excelling in complex data analysis, automated feature extraction, and multi-task learning, thereby improving predictive accuracy in large biomedical datasets [7]. The medical sector is inundated with growing volumes of heterogeneous data, ranging from genomic and proteomic information to diagnostic images and electronic health records (EHR). Traditional methods struggle to manage this volume and identify meaningful patterns, while AI’s capacity to process “complex big data” and discover “subtle correlations” positions it not merely as a processing tool, but as a lens that renders visible and interpretable patterns and relationships that would otherwise be inaccessible or too complex for human analysis or simpler statistical methods.
Precision Medicine represents an approach that aims to adapt diagnoses and treatments to individual patient characteristics, considering genetic and metabolic aspects [8]. This paradigm is based on individual variability to optimize prevention and treatment strategies, marking a crucial transition from generic to “tailored” therapies. Despite its immense theoretical potential and promises of superior clinical outcomes, the practical application of precision medicine in real-world settings has not yet reached its full potential. One of the primary limitations lies in the ability to interpret complex genomic anomalies and effectively match them with the most appropriate molecular drugs or immunotherapies [9].
The integration of AI into precision medicine has already demonstrated revolutionary potential in improving disease diagnosis, treatment, and prevention, inaugurating a new era in medical practice. AI facilitates the identification of highly personalized treatments, considering patients’ genetic characteristics and lifestyle factors [6]. The convergence of these two disciplines promises to revolutionize healthcare by enabling the identification of patient phenotypes with less common therapeutic responses or unique healthcare needs [10]. However, to maximize the value of AI-mediated healthcare, it is imperative to ensure the quality and accuracy of utilized data, alongside transparency and ethics in implementing algorithms and predictive models [11] (Figure 1).
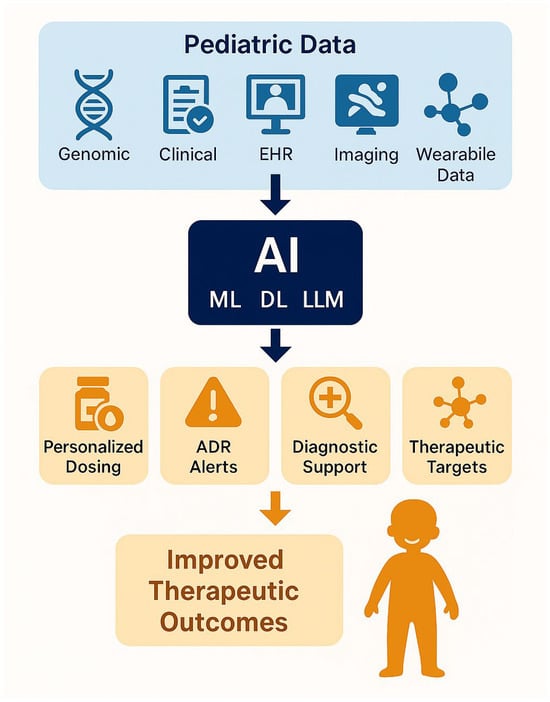
Figure 1.
Conceptual workflow for AI-supported precision pharmacotherapy in pediatrics, depicting the integration of genomic data, electronic health records (EHR), wearable sensors, and clinical information into AI decision-support systems. This diagram illustrates the theoretical ideal; actual real-world implementation faces substantial barriers: (1) Data quality limitations—pediatric EHR data completeness is significantly lower than adult systems, with studies reporting 30–50% missing data in critical clinical fields across even specialized pediatric centers; (2) Limited genomic integration—routine pharmacogenomic testing remains available in fewer than 10% of US pediatric hospitals, and international adoption is substantially lower due to cost, infrastructure, and regulatory barriers; (3) Wearable device availability—FDA-approved wearable sensors for pediatric use are extremely limited compared to the adult market, restricting real-time physiological data collection; (4) Infrastructure requirements—implementation requires substantial investment in interoperable EHR systems, computational infrastructure for real-time AI inference, clinical decision support integration, and healthcare provider training; (5) Geographic variation—feasibility of implementing this workflow varies dramatically between well-resourced pediatric academic medical centers and community hospitals, rural clinics, and healthcare systems in low/middle-income countries. Successful implementation of this workflow requires concurrent advances in healthcare information technology, data standardization, clinical decision support integration, and workforce training, not merely algorithmic sophistication. (ML: Machine Learning, DL: Deep Learning, LLM: Large Language Models).
4. Current Applications in Pediatric Precision Pharmacotherapy
4.1. Pharmacogenomics and Dosage Optimization
AI is revolutionizing pharmacogenomics (PGx) through its capacity to analyze complex datasets and develop advanced predictive models. The integration of these technologies with PGx offers potential for obtaining more precise, data-driven insights useful for identifying new drug targets, predicting drug efficacy, optimal therapy selection, and estimating adverse event risk. Pharmacogenomic variations in genes regulating drug disposition and therapeutic targets are recognized as primary determinants of interindividual differences in drug response and toxicity.
ML models have significantly improved the prediction of genetic variant effects by incorporating DNA and protein sequence analysis, evolutionary conservation, and haplotype structures. Emerging DL models employ sophisticated techniques to capture evolutionary and biophysical properties, while ensemble approaches integrating multiple predictive models demonstrate greater accuracy, robustness, and interpretability. The combined application of these tools improves functional effect prediction of drug-related variants and offers a concrete strategy for translating comprehensive genomic information into clinically applicable pharmacogenetic recommendations in the near future [12].
In pediatrics, AI enables the creation of personalized treatment plans considering age, weight, genetic markers, and patient clinical history, ensuring safer and more effective therapies. AI-guided precision medicine can minimize side effects and improve treatment adherence, crucial factors in this vulnerable population [3]. For instance, a recent study on tacrolimus pharmacokinetics in 139 children with refractory nephrotic syndrome combined population pharmacokinetic modeling with six machine-learning algorithms, identifying the Lasso model as the most accurate (R2 = 0.42) for predicting drug clearance based on genetic and clinical variables—supporting the feasibility of AI-driven individualized therapy in pediatric pharmacogenomics [13]. However, it is important to acknowledge that an R2 of 0.42 indicates that more than half of the interindividual variability in tacrolimus clearance remains unexplained. For a drug with a narrow therapeutic index, this level of predictive accuracy is insufficient for clinical decision-making and requires careful interpretation. Moreover, the model was evaluated only through internal resampling strategies and was not externally validated on independent cohorts, which limits its generalizability and raises the possibility of overfitting. Recent AI-based approaches, such as interpretable neural networks coupled with adversarial training, have successfully identified genetic variants in CERS6 and TLR4 associated with cisplatin-induced ototoxicity in pediatric cancer patients, offering new insights for precision prevention strategies [14]. In a similar direction, a study on mycophenolate mofetil in 171 Chinese pediatric patients with immune-mediated renal diseases identified random forest as the best-performing model for predicting mycophenolic acid exposure (AUC0–12 h > 30 mg·h/L), with SHAP analysis highlighting albumin and hepato-renal function markers as key predictors—enabling exposure-guided, individualized dosing strategies to reduce proteinuria and improve outcomes [15]. Complementing these efforts, another investigation applied ten different AI models to 614 MPA samples from 209 pediatric patients with autoimmune diseases, demonstrating that the Wide&Deep model could accurately predict drug exposure with only three blood samples (R2 = 0.95), significantly reducing patient discomfort without compromising clinical precision. It is important to clarify, however, that these performances derive from an internal validation on a single-center dataset, as the original study did not include external validation on independent cohorts. The authors themselves highlight that the generalizability of the model remains to be confirmed in broader and more heterogeneous pediatric populations, and that future studies will focus on external validation. For this reason, the reported accuracy should be interpreted as a methodological proof-of-concept rather than evidence already ready for clinical implementation [16]. Along the same lines, a recent systematic review analyzing 59 studies on chemotherapy-induced toxicities in children with solid tumors found consistent pharmacogenomic associations—such as ABCC2, MTHFR, and SXR with methotrexate-induced hepatotoxicity and myelosuppression, and SLC28A3, CELF4, and RARG with anthracycline-related cardiotoxicity—despite inter-study variability, underscoring the potential of AI-supported PGx to refine pediatric oncology treatments [17].
Reinforcing this trajectory, a novel AI assistant powered by GPT-4 and enhanced through retrieval-augmented generation (RAG) demonstrated high efficacy in interpreting pharmacogenomic test results using CPIC-based knowledge. Compared to GPT-3.5, the assistant provided more accurate and context-relevant responses, particularly in provider-focused queries, highlighting the promise of generative AI in clinical decision-making and equitable access to personalized medicine [18]. The vast expanse of “millions of rare variations” makes their functional characterization and practical precision medicine implementation extremely challenging. AI, particularly ML and DL, can analyze these complex datasets while integrating information such as evolutionary conservation and biophysical properties, suggesting that AI not only identifies simple correlations but can also infer more complex functional effects and potentially causal relationships between genetic profiles and drug outcomes. Table 1 presents a summary of the various applications of artificial intelligence in the context of drug dosing.

Table 1.
Applications of Artificial Intelligence in Pediatric Precision Pharmacotherapy.
4.2. Adverse Drug Reaction Prediction
AI-assisted prediction of Adverse Drug Reactions (ADRs) has significant potential for improving drug safety and reducing healthcare costs. While initial studies relied on limited dimensions such as drug molecular structure or biomolecule interactions, integration of multiple characteristics now provides valuable insights from diverse perspectives, improving the completeness and accuracy of predictive models. A fusion deep-learning model developed for ADR prediction utilized four modules to extract information on drug molecular structure, drug-protein interaction data, and drug similarity. This model demonstrated remarkable performance on reference datasets, with ROC-AUC of 0.7002, AUPR of 0.6619, and F1 score of 0.6330.
The model’s AUPR showed significant improvement over conventional multi-label classifiers (from 64.02% to 66.19%) and surpassed State-of-the-Art methods on the LIU dataset (from 34.65% to 68.82%), highlighting its greater accuracy and robustness [19]. A systematic review of ML algorithms in pharmacovigilance for ADR prediction in hospitalized patients analyzed 13 studies and found that, for models in the development phase, the meta-analysis showed aggregated sensitivity of 78.1% and specificity of 70.6%. Studies that included external validation achieved superior performance, with an aggregated sensitivity of 81.5% and specificity of 79.5% [20]. Furthermore, AI implementation in hospital pharmacies led to reductions in prescription distribution errors of up to 75% and improvements in adverse drug reaction detection of up to 65% [21].
Recent advances in ML-based ADR prediction have demonstrated substantial progress across diverse clinical applications (predominantly in adult populations). A comprehensive meta-analysis of 59 studies covering 15 drugs and 15 ADEs revealed strong overall performance of ML models, with an average AUC of 76.68 ± 10.73%, accuracy of 76.00 ± 11.26%, sensitivity of 62.35 ± 20.19%, and specificity of 75.13 ± 16.60%. Random forest emerged as the most frequently used algorithm, followed by support vector machine, XGBoost, decision tree, and LightGBM [22]. Machine-learning approaches utilizing in vitro target-based pharmacology have identified 221 target-ADR associations, including established relations such as hERG binding with cardiac arrhythmias and novel associations like PDE3 with 40 different ADRs [23].
Notably, a study applying machine-learning techniques to predict adverse drug events in Chinese pediatric inpatients (n = 1746) with a median age of 3.84 years, demonstrated that the Gradient Boosting Decision Tree (GBDT) model significantly outperformed traditional Global Trigger Tools (44% vs. 13.3% in precision), while identifying key risk factors such as BMI, number of doses and drugs, and hospital stay length. This work highlights the feasibility and clinical value of ML-based ADR prediction in pediatric populations. However, it should be emphasized that the model was developed and evaluated exclusively within a single Chinese tertiary hospital, reflecting local prescribing practices, clinical workflows, and demographic characteristics. As also acknowledged by the original authors, the lack of multicenter or international validation limits the generalizability of the model, underscoring the need for external testing across diverse healthcare systems to mitigate algorithmic bias and ensure true clinical transferability [24]. In the neonatal setting, a prospective cohort study applied a machine-learning-based risk score to predict ADRs among 412 critically ill neonates, achieving a high classification accuracy (c-index: 0.914) based on key variables such as drug class, underlying conditions, and treatment characteristics—demonstrating the concrete potential of AI for early risk stratification and preventive pharmacovigilance in this highly vulnerable group [25].
Similarly, a study of 3152 ADR reports in Malaysian children tested four digital signal detection algorithms and showed that Multi-item Gamma Poisson Shrinker (MGPS) had the highest sensitivity (20%) and negative predictive value (23%), while all methods achieved 100% specificity and positive predictive value, confirming the effectiveness of digital pharmacovigilance systems for early identification of serious pediatric ADRs [26].
In pediatric tuberculosis, an AutoML-based gradient boosting model achieved high predictive performance for drug-induced liver injury (AUC = 0.838 on training set, 0.784 on testing set), identifying rifampicin Cmax and BMI as the most influential risk factors, and offering a promising tool for early clinical intervention [27]. Generative AI applications in pharmacovigilance show particular promise for enhanced adverse event detection, data-driven risk prediction, and optimized drug development, with the potential to streamline pharmacovigilance procedures through automation and big data processing [28].
These findings demonstrate AI’s capacity to transform pharmacovigilance from a reactive monitoring system to a proactive safety mechanism. This paradigm shift can lead to earlier interventions, reduced patient harm, and significant cost savings in healthcare systems, particularly beneficial in pediatric settings where adverse events can have long-term consequences on development and quality of life. Table 2 summarizes the artificial intelligence models developed to predict adverse drug reactions in pediatric populations.

Table 2.
AI Models for Predicting Adverse Drug Reactions (ADR) in Pediatric Populations.
4.3. Drug Discovery and Repositioning
AI can radically transform the drug discovery and early development process, addressing notable inefficiencies of traditional methods characterized by high costs, long timelines, and low success rates [29]. AI can improve every phase of this process, from molecular target identification to new molecule discovery and early clinical development. Through multiomics data analysis and network-based approaches, AI can identify new oncogenic vulnerabilities and key therapeutic targets. Advanced AI models like AlphaFold (Google) can predict protein structures with high accuracy, facilitating “druggability” assessments and structure-based drug design.
AI not only accelerates virtual screening of compound libraries but also enables de novo drug design, creating molecular structures optimized for specific biological properties [30]. Repositioning of existing drugs, a process significantly facilitated by AI, can drastically reduce time, costs, and risks associated with new drug development, offering a valuable avenue for addressing unmet medical needs. AI excels in analyzing large-scale datasets, identifying complex patterns in drug responses, and formulating predictions for potential drug repositioning [31].
The traditional drug discovery process is notoriously slow, expensive, and has a high failure rate. AI’s capabilities in analyzing vast datasets, predicting protein structures, and conducting virtual screening radically transform this process, dramatically increasing efficiency and reducing time and costs. This acceleration is particularly critical for pediatric diseases, many of which are rare and lack approved treatments [32]. By making drug discovery more efficient, AI can potentially address significant unmet medical needs in pediatric pharmacotherapy, bringing new therapies to children more rapidly.
4.4. Clinical Decision Support Systems
AI holds significant potential in supporting physicians in formulating more accurate diagnoses [33] and improving clinical decision-making through augmented intelligence [10]. AI-enabled Clinical Decision Support Systems (CDSS) can reduce diagnostic errors, enhance clinical intelligence supporting decisions, and assist physicians in extracting data from electronic health records (EHR) and documentation activities. AI can provide physicians with instant access to updated scientific information and evidence, fundamental for supporting timely, evidence-based clinical decisions. Additionally, AI can facilitate continuous monitoring and management of patient treatment over time [11].
LLMs are gaining ground in healthcare environments, offering decision support and assisting in documentation and communications [34]. A concrete example is the application of LLaMA that accurately classified clinical notes containing side effect requests (sensitivity = 87.2, specificity = 86.3, AUC = 0.93) in a pediatric ADHD medication management context, demonstrating the ability to scalably measure care quality [35]. These findings position AI as a “cognitive co-pilot,” improving clinicians’ ability to make informed decisions while maintaining human oversight as a central element.
Broader clinical applications demonstrate substantial AI-CDSS effectiveness across diverse healthcare settings, predominantly adult populations. AI systems for clinical trial eligibility screening achieved accuracy, sensitivity, and specificity greater than 80% in determining patient eligibility for breast cancer clinical trials, potentially increasing screening efficiency while accurately excluding ineligible patients [36]. In cardiovascular care, AI demonstrated remarkable performance in arrhythmia detection, achieving sensitivity and specificity greater than 96% for atrial fibrillation identification [37]. Cardiovascular CDSS applications show promise in risk assessment, emergency department differential diagnosis, and imaging-based approaches for coronary artery disease management, with AI-guided risk stratification potentially reducing patient numbers needed for targeted screening programs [38].
Primary care implementations reveal varying outcomes, with AI-CDSS demonstrating potential for managing diverse patient populations while maintaining up-to-date medical knowledge [39]. However, challenges persist in the interpretability and transparency of AI models, particularly deep-learning systems characterized by a black-box nature, posing difficulties for clinicians seeking to understand AI-driven recommendations. The generalizability and robustness of AI models across diverse patient populations and clinical settings remain critical challenges, with algorithms potentially exhibiting performance disparities across demographic groups [40]. These predominantly adult-focused findings highlight both the transformative potential and implementation challenges of AI-CDSS, emphasizing the need for pediatric-specific validation and development.
5. Disease-Specific Applications
5.1. Pediatric Oncology
AI is fundamental in oncological drug discovery and development, facilitating target identification, virtual screening, and de novo drug design. AI-based predictive models can assess drug toxicity in silico and identify specific biomarkers for patient stratification and personalized treatment approaches [41]. Recent overviews confirm that AI is increasingly shaping both adult and pediatric oncology by supporting diagnostic accuracy, risk stratification, and treatment personalization, while also raising important ethical considerations for its responsible clinical deployment [42]. An innovative approach for predicting cancer drug response (CDR) integrates the scGPT model within the DeepCDR model, using gene expression data embeddings. This scGPT-based method demonstrated superiority over previous work, including the original DeepCDR model and the scFoundation-based model. The scGPT-based approach showed steady improvement during training, starting at approximately 0.80 Pearson Correlation Coefficient (PCC) and reaching above 0.90, while maintaining superior training stability compared to the scFoundation model, which exhibited significant fluctuations throughout the training process. In leave-one-drug-out testing, the scGPT model consistently achieved higher PCC gains across various drugs compared to both alternative methods, demonstrating enhanced predictive capability for unseen drugs. Notably, the approach achieved highly accurate predictions for specific cases, including Low-Grade Gliomas as the best-performing cancer type and Tubastatin A among individual drug predictions [43].
MADRIGAL, a multimodal AI model, can learn from structural, pathway, cell viability, and transcriptomic data to predict effects of drug combinations on 953 clinical outcomes and 21,842 compounds, surpassing single-modality methods and State-of-the-Art models in predicting adverse drug interactions. The model uses a transformer bottleneck module to unify preclinical drug data modalities while handling missing data during training and inference, addressing a major challenge in multimodal learning. MADRIGAL successfully predicted resmetirom, the first and only FDA-approved drug for metabolic dysfunction-associated steatohepatitis (MASH), among therapies with the most favorable safety profile. The system supports personalized cancer therapy by integrating genomic profiles from cancer patients and demonstrated efficacy prediction capabilities using primary acute myeloid leukemia samples and patient-derived xenograft models for personalized drug combinations [44]. For acute lymphoblastic leukemia (ALL), a single-cell proteomic approach accurately predicted future relapse with 38% higher precision than standard prediction methods [45]. Another study demonstrated that a Long Short-Term Memory (LSTM) model achieved 100% accuracy in classifying leukemia types, a crucial result for treatment stratification and reducing unnecessary toxicities [46]. Accurate classification enables selection of the most effective drugs for a specific leukemia subtype, reducing unnecessary toxicities and improving cure rates, representing a significant step toward more effective precision medicine in pediatric oncology.
Further strengthening the role of AI in supportive pediatric oncology, a predictive model based on the C5.0 decision tree combined with SMOTE reached an AUC of 0.807 in identifying children at risk for delayed high-dose methotrexate (HD-MTX) clearance—outperforming conventional nomogram-based approaches and supporting early clinical intervention [47].
In a related application, a model combining random forest with adaptive synthetic resampling achieved excellent performance in predicting neutropenia (AUC = 0.927, sensitivity = 93.5%, specificity = 92.0%) and fever (AUC = 0.870) following HD-MTX, offering a valuable tool to anticipate complications and improve therapeutic decision-making in pediatric leukemia [48].
Similarly, in pediatric acute myeloid leukemia (AML), machine-learning models trained on electronic medical records achieved an AUC of 0.748 for predicting bloodstream infections, with improved specificity over neutropenia by 37.5%, supporting early infection risk stratification and guiding prophylactic strategies such as levofloxacin-vancomycin administration [49].
Reinforcing the relevance of ML in pediatric oncology, a recent study developed an extreme gradient boosting (XGB) model to estimate the risk of chemotherapy-induced myelosuppression in children with Wilms’ tumor, achieving excellent predictive performance (AUROC = 0.981 in training, 0.896 in testing) and identifying key predictors such as hemoglobin, white blood cell count, and albumin—offering a clinically valuable tool to anticipate hematologic toxicity and guide early supportive interventions [50].
5.2. Pediatric Infectious Diseases
AI has the potential to improve antimicrobial resistance prediction and optimize antibiotic prescriptions, supporting treatment strategies and bridging critical gaps in healthcare. ML models such as random forests and gradient-boosted decision trees have been used to predict antimicrobial resistance and optimize antibiotic selection by analyzing large datasets to discover actionable patterns [51]. In the pediatric sepsis context, which causes over 72,000 annual hospitalizations in the United States with significant mortality and morbidity, a predictive AI model called “Sepsis on ED to PICU Disposition” (SEPD) was developed and validated to accelerate antibiotic therapy.
The SEPD model outperformed a vendor-developed sepsis model with an AUROC of 81.8% versus 57.5%, demonstrating better precision-recall performance and more balanced identification of true positives. During “silent” implementation, the SEPD model maintained similar sensitivity (85.29%) and specificity (60.45%) to those observed during model testing. This model can facilitate identification and timely treatment of sepsis in high-risk pediatric patients [52].
In neonatal care, an ML-based clinical decision support system using the CatBoost algorithm was shown to reliably guide β-lactam antibiotic dosing across five commonly used drugs. It achieved over 80% prediction accuracy in real-world validation and improved key performance metrics compared to population pharmacokinetic models, with a 58.2% average increase in the probability of achieving therapeutic targets when compared to guideline-based dosing [53].
Similarly, in efforts to monitor antibiotic appropriateness in outpatient pediatrics, a generative language model—based on Llama 3.1 and refined via few-shot learning and chain-of-thought prompting—was able to identify 94.7% of clinical notes that justified antibiotic use in pediatric sinusitis and 66.2% of those where it was inappropriate, offering a scalable tool for real-time stewardship interventions via EHR integration [54].
In the context of vancomycin management, which is particularly complex in younger children due to rapid physiological changes, an XGBoost model trained on pediatric patients under 4 years of age predicted trough concentrations with good accuracy (R2 = 0.59; MAE = 2.55), highlighting renal function as a key predictor [55]. A separate ensemble model, developed using data from 407 children, combined algorithms such as GBRT, Bagging eXtraTree, and decision trees, achieving even better predictive metrics (R2 = 0.614; prediction accuracy within ±30% of true values in 51.22% of cases), significantly outperforming traditional pharmacokinetic models [56].
The role of AI in improving time-sensitive conditions such as pediatric sepsis has also been demonstrated in a study that employed a graph neural network with deep forest modeling to predict sepsis onset in real time. The model reached AUCs of 93.63%, 96.73%, and 97.58% across successive diagnostic stages, with F1 scores progressively increasing to 86.48%, allowing for significantly earlier diagnosis and intervention [57].
More broadly, in febrile infants ≤ 60 days of age (n = 1470), a random forest model using just four clinical and laboratory features (urinalysis, white blood cell count, absolute neutrophil count, procalcitonin) achieved a remarkable sensitivity of 98.6% and specificity of 74.9%. This approach could reduce unnecessary lumbar punctures by nearly 70%, without compromising diagnostic safety [58].
In critically ill children, a random forest-based diagnostic model distinguished sepsis from non-infectious SIRS using eight routinely available variables, achieving an AUC of 0.78 in validation set and outperforming conventional biomarkers like CRP or procalcitonin alone—potentially reducing unnecessary antibiotic use by up to 30% in SIRS cases [59].
Lastly, in cases of pediatric pharyngitis—a frequent cause of inappropriate antibiotic use—an ML model based on support vector machines showed strong potential as a diagnostic aid, with a positive predictive value of 80.6% in distinguishing Group A streptococcal infections from viral etiologies, contributing to more targeted treatment strategies and reduced antimicrobial misuse [60].
Beyond pediatric-specific applications, AI has demonstrated significant advances in antimicrobial stewardship across clinical settings. The APEX Generative Optimization (APEX GO) framework, a generative AI system integrating transformer-based variational autoencoders with Bayesian optimization, achieved an outstanding 85% ground-truth experimental hit rate and 72% success rate in enhancing antimicrobial activity against clinically relevant Gram-negative pathogens [61]. AI-enabled clinical decision support systems have shown remarkable improvements over traditional rule-based approaches, with one study demonstrating enhancement of sensitivity from 26.44% to 80.84% and specificity from 99.23% to 99.95% in holistic healthcare prediction for inpatients [62].
Machine-learning models have achieved high accuracy rates in predicting antibiotic resistance without relying on traditional antibiogram methods. In a Turkish study analyzing clinical data from 103 patients with Escherichia coli infections, models demonstrated accuracy rates of 98% for fosfomycin, 98% for levofloxacin, and 96% for ertapenem predictions [63]. A Spanish study predicting antibiogram results for multidrug-resistant bacteria in intensive care units achieved the highest accuracy (77%) and specificity (82%) with random forest classifiers, while the highest sensitivity (69%) and ROC-AUC (76%) were obtained using Chi-square feature selection with logistic regression or XGBoost classifiers [64].
Telemedicine platforms leveraging AI algorithms have optimized antibiotic prescribing practices, with a study demonstrating slightly higher guideline-concordant antibiotic management in telemedicine visits (92.5%) compared to in-person visits (90.7%) for acute respiratory tract infections [65]. Furthermore, semi-supervised ensemble learning methods for urinary tract infection diagnosis have proven more accurate than traditional urinalysis or urine culture for UTI prediction, potentially reducing unnecessary antibiotic treatments and associated resistance development [66].
5.3. Pediatric Neurological Diseases
AI has demonstrated strong potential in pediatric epilepsy diagnosis and monitoring, often matching clinical accuracy. AI-based decision support achieved 93.4% accuracy in diagnosis, aligning closely with expert evaluations. Specific methods such as EEG-based AI for interictal discharge detection showed high specificity (93.33–96.67%) and sensitivity (76.67–93.33%). Neuroimaging approaches using resting-state functional MRI (rs-fMRI) and diffusion tensor imaging (DTI) achieved up to 97.5% accuracy in identifying microstructural abnormalities [67].
A systematic review and meta-analysis included 28 studies on ML and DL efficacy in pediatric epileptic seizure monitoring. For ML models in the training set, aggregated sensitivity was 0.77 (95% CI 0.73–0.80), specificity was 0.74 (95% CI 0.70–0.77), and accuracy was 0.75 (95% CI 0.72–0.77). In the validation set, ML aggregated sensitivity was 0.88 (95% CI 0.83–0.91), specificity was 0.83 (95% CI 0.71–0.90), and accuracy was 0.78 (95% CI 0.73–0.82). For DL models in the validation set, aggregated sensitivity was 0.89 (95% CI 0.85–0.91), specificity was 0.91 (95% CI 0.88–0.93), and accuracy was 0.89 (95% CI 0.86–0.92). The study concludes that DL algorithms appear to offer greater detection accuracy than ML algorithms [68].
5.4. Pediatric Rare Genetic Diseases
Rare diseases are often difficult to diagnose due to their low incidence and symptom overlap with more common conditions. AI, particularly LLMs, is improving the differential diagnosis of rare diseases. The RareScale system was designed to improve differential diagnoses of rare diseases directly from medical history dialogues. This approach improved Top-5 accuracy by over 17% compared to baseline “black-box” LLMs. Specifically, on dialogues generated by GPT-4o, Top-5 accuracy improved from 56.7% to 74.1%. On the Claude test sets, Top-5 accuracy improved from 56.69% to 71.41%. Top-1 accuracy also showed improvements, increasing from 28.65% to 33.12% for gpt-4o and from 30.65% to 33.23% for Claude [69].
By 2033, the drug development pipeline for pediatric-onset rare diseases is expected to result in approximately 45 new product approvals, alongside a 14% increase in the number of patients treated annually and a projected $10.7 billion rise in gross drug revenues, growing from $28.2 billion in 2023 to $38.9 billion in 2033. Despite these advances, the pace at which new rare diseases are being identified is likely to surpass the rate at which new treatments become available. It is anticipated that 95% of pediatric-onset rare diseases will still lack approved therapies over the next decade, and even among the remaining 5%, existing options will often be insufficient to meet clinical needs [32].
6. Discussion
The integration of AI into pediatric precision pharmacotherapy represents a paradigmatic transformation that addresses fundamental challenges inherent to pediatric medicine, yet its implementation reveals a complex landscape of promises and substantial barriers requiring critical examination (Figure 2).
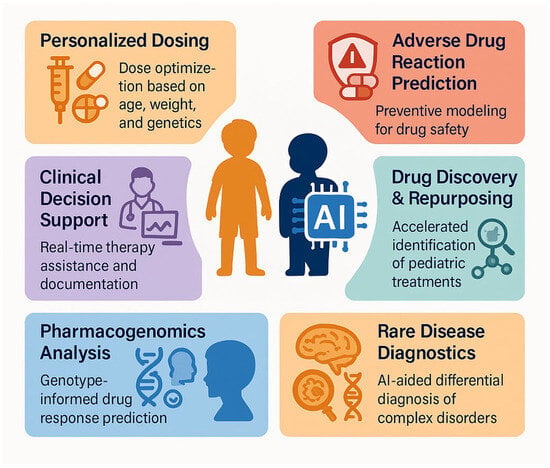
Figure 2.
Illustrative diagram highlighting key applications of AI in pediatric pharmacotherapy, including personalized dosing, ADR prediction, clinical decision support, drug repurposing, pharmacogenomics, and rare disease diagnostics—aimed at improving safety and efficacy of treatments in children.
Current evidence demonstrates remarkable clinical achievements: AI implementation in hospital pharmacies has achieved up to 75% reduction in prescription distribution errors and 65% improvement in adverse drug reaction detection [21]. The demonstrated capabilities of AI in analyzing complex, patient-specific data to personalize treatments represent an ideal solution for addressing the inherent complexity of pediatric populations, where physiological distinctiveness characterized by age-dependent pharmacokinetic and pharmacodynamic variations creates high uncertainty in drug response and increased risk of adverse effects. However, a critical implementation gap persists between research potential and clinical reality. Recent systematic analyses reveal that of 262 pediatric AI studies, only one model (0.38%) reached technology readiness level 8, indicating actual clinical testing [70]. Furthermore, 77% of pediatric AI studies demonstrate a high risk of bias, primarily due to insufficient sample sizes [71].
To reduce this gap and accelerate pediatric-specific model validation, a structured translational pathway is urgently needed. Such a framework may include multicenter pediatric data harmonization networks to overcome the limitations of fragmented datasets; age-stratified validation pipelines to ensure rigorous performance assessment across developmental stages; and prospective real-world pilot evaluations embedded in routine clinical workflows, allowing continuous recalibration and safety monitoring. These components together would establish a reproducible and scalable route for transitioning pediatric AI tools from research prototypes to clinically reliable systems.
The economic landscape of healthcare AI demonstrates substantial growth, with the global market reaching $26.57 billion in 2024 and projected to reach $187.69 billion by 2030, representing a compound annual growth rate of 38.5% [72]. Despite this growth, implementation costs remain significant, ranging from $50,000 to $300,000 for small clinics [73]. Organizations implementing ambient AI for clinical documentation report 15% improvements in charge capture and 60% better quality code capture [74]. However, economic barriers persist, with over one-third of healthcare technology leaders citing financial constraints as the main obstacle to AI adoption, and nearly 30% reporting doubts about the financial benefits of these investments [75].
For smaller pediatric institutions, cost-conscious and incremental adoption strategies are essential. Practical approaches may include leveraging cloud-based AI infrastructures to minimize upfront hardware expenses, deploying modular decision-support tools that can be integrated stepwise, and prioritizing applications with immediate demonstrated benefits, such as ADR detection or medication error reduction. These strategies make AI deployment feasible even in resource-limited pediatric settings, while maintaining adherence to safety, regulatory, and ethical standards.
The concept of age-related algorithmic bias emerges as a critical consideration unique to pediatric AI implementation. Sources of algorithmic bias affecting children and young people may arise from a lack of transparency in participant age reporting, inadequate clinically and developmentally appropriate representation of children, and inappropriate generalizations made to the pediatric population from adult data [76]. Recent evidence demonstrates that some LLMs show substantial performance decreases when addressing pediatric versus adult medical questions, with models like Gemini 2.0 Flash showing about a 10.3 percentage point decline (81.24% vs. 70.96%) in pediatric questions [77]. Healthcare AI systems are particularly susceptible to bias because they reflect the training data, and if models are trained predominantly on data from certain demographic groups, they may underperform when applied to underrepresented populations [78]. The ACCEPT-AI framework has emerged as a comprehensive approach to mitigate these challenges, incorporating six key sections: age, communication, consent and assent, equity, protection of data, and technological considerations [76]. This framework provides actionable recommendations for researchers, regulators, and clinicians to ensure equitable, ethical, and appropriate representation of children and young people in AI/ML research.
The availability of high-quality, age-specific pediatric data represents one of the primary limitations for developing robust AI models [79]. Pediatric datasets are typically smaller compared to adult ones, limiting ML model effectiveness and generalizability. Privacy concerns further limit data sharing between institutions, leading to restricted datasets for model training. Recent publication trends demonstrate exponential growth in pediatric AI research, with projections suggesting 2024 could cross 300 original studies integrating pediatrics and machine learning [80]. This pediatric-specific growth mirrors the broader expansion of AI research in healthcare, which has increased from 158 articles (3.54%) in 2014 to 731 articles (16.33%) by 2024 [81]. However, this quantitative growth has not translated proportionally into clinically validated applications. A comprehensive analysis of AI implementation in pediatric clinical settings found that among 126 full-text articles examined, only 17 met the inclusion criteria for AI implementation in pediatric clinical settings, with 30% showing no differences in clinical outcomes. Among pediatric AI implementations, only 47% reported both clinical outcomes and process measures, while 35% reported only process measures [82], highlighting the need for standardized reporting of outcomes, care processes, and human performance measures in pediatric AI research.
Although research environments frequently report high predictive performance, translation into real-world pediatric care remains challenging. Data heterogeneity, inconsistent documentation practices, and varying levels of EHR interoperability can significantly diminish model accuracy outside controlled settings. These issues are even more prominent in low-resource pediatric environments, where dataset sparsity and infrastructural constraints limit reliable deployment. Real-world testing and adaptive recalibration should therefore be integrated early in the development pipeline to ensure robustness, equity, and clinical safety across diverse healthcare contexts.
Current applications demonstrate tangible clinical impact across multiple domains. In pharmacogenomics, AI’s ability to integrate evolutionary conservation and biophysical properties suggests movement beyond simple correlations toward inferring complex functional effects between genetic profiles and drug outcomes [12]. This capability is fundamental for truly personalized dosing, especially for drugs with narrow therapeutic indices commonly used in pediatric oncology, where accurate dosing based on individual metabolism can prevent severe toxicities or ensure therapeutic efficacy. The transformation of pharmacovigilance from reactive monitoring to proactive safety mechanisms through AI-predicted adverse drug reactions represents a paradigm shift with particular significance in pediatrics, where adverse events can have long-term developmental consequences. However, implementation requires addressing significant ethical considerations. The ACCEPT-AI framework specifically addresses the ethical use of pediatric data in AI research, emphasizing age-appropriate consent processes, data protection, and prevention of algorithmic discrimination [76]. Healthcare professionals require urgent training and orientation to AI applications, as training algorithms often require large datasets that pose challenges when building unbiased AI algorithms in pediatric populations with small sample sizes [83].
Despite the abundance of published pediatric AI models, the number of AI implementations in clinical practice is minimal, and standardized reporting of outcomes, care processes, or human performance measures is lacking. The clinical decision support capabilities demonstrate promise, with AI-based systems achieving 93.4% accuracy in pediatric epilepsy diagnosis [67] and AI-enabled clinical decision support systems showing enhancement of sensitivity from 26.44% to 80.84% in healthcare prediction for inpatients [62]. However, challenges persist in the interpretability and transparency of AI models, particularly deep-learning systems characterized by a black-box nature, posing difficulties for clinicians seeking to understand AI-driven recommendations. The “black box” nature of AI algorithms is particularly problematic in pediatric contexts, where transparency and clinician trust are paramount for ensuring safe decision-making in this vulnerable population [84]. As noted by Shortliffe, the reasoning behind AI decisions should be transparent for clinicians to comprehend the rationale, as black box algorithms lack the interpretability required for clinical contexts where understanding the decision-making process is essential for patient safety [85]. This challenge is amplified in pediatrics, where the consequences of misdiagnosis or inappropriate treatment can have profound long-term developmental impacts. The development of explainable AI (XAI) techniques specifically tailored for pediatric applications represents a critical research priority to bridge the gap between AI performance and clinical acceptability [86]. Terminology standardization represents an important, often overlooked aspect requiring agreed definitions within respective healthcare systems. Many different terms are used in precision medicine literature, including personalized, individualized, stratified medicine, and precision dosing [87].
A critical gap in the current literature is the scarcity of studies specifically designed to evaluate AI applications in pediatric pharmacotherapy. While numerous AI models demonstrate promising results in adult populations or mixed-age cohorts, dedicated pediatric-focused investigations remain limited. This represents a significant limitation given the unique physiological, pharmacokinetic, and safety considerations that distinguish pediatric from adult pharmacotherapy. The absence of age-specific validation studies raises concerns about the direct applicability of current AI models to pediatric populations and underscores the urgent need for pediatric-centered research initiatives that can validate AI-driven therapeutic decisions in this vulnerable population.
Addressing these challenges will require coordinated action: harmonized multicenter pediatric datasets, rigorous age-stratified evaluation, early real-world testing, and implementation models tailored to institutions with varying resources. Such an approach directly answers the need highlighted by recent reviews and aligns with the ethical obligations emphasized in ACCEPT-AI.
Future research must prioritize real-world validation through rigorous clinical studies and standardized outcome reporting. Emerging techniques, including increasingly sophisticated LLMs and multimodal data integration, promise to further refine AI precision and applicability [88]. However, successful implementation will require sustained commitment to addressing current limitations while maintaining focus on optimizing therapeutic outcomes for this vulnerable population.
7. Conclusions
The convergence of Artificial Intelligence and Precision Medicine creates a promising paradigm for pediatric therapeutic decision-making. Children’s unique physiological characteristics make AI an indispensable tool for transforming pediatric vulnerability into an innovation catalyst.
Current AI applications demonstrate tangible clinical impact across pharmacogenomics, adverse drug reaction prediction, drug discovery, and clinical decision support. AI shifts understanding from correlation to causality, enabling personalized dosing and transforming pharmacovigilance into proactive safety mechanisms.
Despite promising advances, large-scale pediatric AI implementation faces significant challenges, including limited data quality, algorithmic bias, generalizability issues, and the “black-box” nature that undermines clinical trust. Economic barriers and ethical considerations regarding data privacy and informed consent require rigorous attention.
A critical gap exists in pediatric pharmacotherapy-specific AI evaluation studies. The absence of age-specific validation raises concerns about current AI model applicability to pediatric populations, underscoring the need for dedicated pediatric-centered research initiatives.
Future development of sophisticated AI models and multimodal data integration promises enhanced precision. However, real-world validation through rigorous clinical studies and interdisciplinary collaboration remains imperative for building robust pediatric AI ecosystems.
AI represents a transformative element in reshaping pediatric therapeutic decision-making. Successful implementation requires sustained commitment to addressing current limitations while maintaining focus on optimizing therapeutic outcomes for our most vulnerable patient populations.
Author Contributions
Conceptualization, G.M., M.M. and F.G.A.; resources, S.C.; data curation, V.F.; writing—original draft preparation, G.M.; writing—review and editing, G.M., A.P., S.C., V.F., M.M. and F.G.A. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data was created or analyzed in this study. Data is contained within the article.
Acknowledgments
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| Abbreviation | Definition |
| ACCEPT-AI | Age, Communication, Consent and Assent, Equity, Protection of Data, Technology |
| ADE | Adverse Drug Event |
| ADRs | Adverse Drug Reactions |
| ALL | Acute Lymphoblastic Leukemia |
| AML | Acute Myeloid Leukemia |
| APEX GO | APEX Generative Optimization |
| AUC | Area Under the Curve |
| AUPR | Area Under the Precision-Recall Curve |
| CDSS | Clinical Decision Support System |
| CPIC | Clinical Pharmacogenetics Implementation Consortium |
| DL | Deep Learning |
| DTI | Diffusion Tensor Imaging |
| EHR | Electronic Health Record |
| FDA | Food and Drug Administration |
| GBDT | Gradient Boosting Decision Tree |
| GBRT | Gradient Boosted Regression Trees |
| HD-MTX | High-Dose Methotrexate |
| LIU | Labeled Independent Users dataset |
| LLM | Large Language Model |
| LSTM | Long Short-Term Memory |
| MAE | Mean Absolute Error |
| MGPS | Multi-item Gamma Poisson Shrinker |
| ML | Machine Learning |
| PCC | Pearson Correlation Coefficient |
| PGx | Pharmacogenomics |
| RAG | Retrieval-Augmented Generation |
| ROC | Receiver Operating Characteristic |
| rs-fMRI | Resting-State Functional Magnetic Resonance Imaging |
| SEPD | Sepsis on ED to PICU Disposition |
| SHAP | SHapley Additive exPlanations |
| SIRS | Systemic Inflammatory Response Syndrome |
| SMOTE | Synthetic Minority Over-sampling Technique |
| UTI | Urinary Tract Infection |
| XGB | Extreme Gradient Boosting |
| XGBoost | Extreme Gradient Boosting |
References
- World Health Organization. Artificial Intelligence for Health. 2023. Available online: https://www.who.int/publications/m/item/artificial-intelligence-for-health (accessed on 22 June 2025).
- Kearns, G.L.; Abdel-Rahman, S.M.; Alander, S.W.; Blowey, D.L.; Leeder, J.S.; Kauffman, R.E. Developmental Pharmacology—Drug Disposition, Action, and Therapy in Infants and Children. N. Engl. J. Med. 2003, 349, 1157–1167. [Google Scholar] [CrossRef]
- Precision for Medicine. Precision Medicine in Pediatrics: Biomarkers and Assay Development. Available online: https://www.precisionformedicine.com/blog/pediatric-studies-precision-medicine-approach (accessed on 22 June 2025).
- Matellio. AI in Pediatric Healthcare: How Custom AI Solutions Improve Child-Centric Medical Services. Available online: https://www.matellio.com/blog/ai-in-pediatric-healthcare-services/ (accessed on 22 June 2025).
- Tekkeşin, A.İ. Artificial Intelligence in Healthcare: Past, Present and Future. Anatol. J. Cardiol. 2019, 22, 8–9. [Google Scholar] [CrossRef]
- Johnson, K.B.; Wei, W.; Weeraratne, D.; Frisse, M.E.; Misulis, K.; Rhee, K.; Zhao, J.; Snowdon, J.L. Precision Medicine, AI, and the Future of Personalized Health Care. Clin. Transl. Sci. 2021, 14, 86–93. [Google Scholar] [CrossRef]
- Sartori, F.; Codicè, F.; Caranzano, I.; Rollo, C.; Birolo, G.; Fariselli, P.; Pancotti, C. A Comprehensive Review of Deep Learning Applications with Multi-Omics Data in Cancer Research. Genes 2025, 16, 648. [Google Scholar] [CrossRef]
- Guo, C.; Chen, J. Big Data Analytics in Healthcare. In Knowledge Technology and Systems: Toward Establishing Knowledge Systems Science; Springer: Singapore, 2023. [Google Scholar] [CrossRef]
- Schork, N.J. Artificial Intelligence and Personalized Medicine. Cancer Treat Res. 2019, 178, 265–283. [Google Scholar] [CrossRef]
- Alsaedi, S.; Ogasawara, M.; Alarawi, M.; Gao, X.; Gojobori, T. AI-powered precision medicine: Utilizing genetic risk factor optimization to revolutionize healthcare. NAR Genom. Bioinform. 2025, 7, lqaf038. [Google Scholar] [CrossRef]
- Lorkowski, J.; Kolaszyńska, O.; Pokorski, M. Artificial Intelligence and Precision Medicine: A Perspective. In Integrative Clinical Research: Advances in Experimental Medicine and Biology; Springer: Cham, Switzerland, 2022; Volume 1375, pp. 1–11. [Google Scholar] [CrossRef]
- Tremmel, R.; Honore, A.; Park, Y.; Zhou, Y.; Xiao, M.; Lauschke, V.M. Machine learning models for pharmacogenomic variant effect predictions—Recent developments and future frontiers. Pharmacogenomics 2025, 26, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Lin, X.; Wang, Y.; Chen, X.; Zheng, W.; Zhong, X.; Shang, D.; Huang, M.; Gao, X.; Deng, H.; et al. Tacrolimus pharmacokinetics in pediatric nephrotic syndrome: A combination of population pharmacokinetic modelling and machine learning approaches to improve individual prediction. Front. Pharmacol. 2022, 13, 942129. [Google Scholar] [CrossRef] [PubMed]
- Arab, A.; Kashani, B.; Cordova-Delgado, M.; Scott, E.N.; Alemi, K.; Trueman, J.; Groeneweg, G.; Chang, W.-C.; Loucks, C.M.; Ross, C.J.D.; et al. Machine learning model identifies genetic predictors of cisplatin-induced ototoxicity in CERS6 and TLR4. Comput. Biol. Med. 2024, 183, 109324. [Google Scholar] [CrossRef]
- Han, L.-Y.; Chen, X.; Liu, T.-S.; Zhang, Z.-L.; Chen, F.; Zhan, D.-C.; Yu, Y.; Yu, G. Applying exposure-response analysis to enhance Mycophenolate Mofetil dosing precision in pediatric patients with immune-mediated renal diseases by machine learning models. Eur. J. Pharm. Sci. 2025, 211, 107146. [Google Scholar] [CrossRef] [PubMed]
- Zheng, P.; Pan, T.; Gao, Y.; Chen, J.; Li, L.; Chen, Y.; Fang, D.; Li, X.; Gao, F.; Li, Y. Predicting the exposure of mycophenolic acid in children with autoimmune diseases using a limited sampling strategy: A retrospective study. Clin. Transl. Sci. 2025, 18, e70092. [Google Scholar] [CrossRef]
- Hansson, P.; Blacker, C.; Uvdal, H.; Wadelius, M.; Green, H.; Ljungman, G. Pharmacogenomics in pediatric oncology patients with solid tumors related to chemotherapy-induced toxicity: A systematic review. Crit. Rev. Oncol./Hematol. 2025, 211, 104720. [Google Scholar] [CrossRef]
- Murugan, M.; Yuan, B.; Venner, E.; Ballantyne, C.M.; Robinson, K.M.; Coons, J.C.; Wang, L.; Empey, P.E.; Gibbs, R.A. Empowering personalized pharmacogenomics with generative AI solutions. J. Am. Med. Inform. Assoc. 2024, 31, 1356–1366. [Google Scholar] [CrossRef]
- Ou, Q.; Jiang, X.; Guo, Z.; Jiang, J.; Gan, Z.; Han, F.; Cai, Y. A Fusion Deep Learning Model for Predicting Adverse Drug Reactions Based on Multiple Drug Characteristics. Life 2025, 15, 436. [Google Scholar] [CrossRef]
- Dsouza, V.S.; Leyens, L.; Kurian, J.R.; Brand, A.; Brand, H. Artificial intelligence (AI) in pharmacovigilance: A systematic review on predicting adverse drug reactions (ADR) in hospitalized patients. Res. Soc. Adm. Pharm. 2025, 21, 453–462. [Google Scholar] [CrossRef]
- Simpson, M.D.; Qasim, H.S. Clinical and Operational Applications of Artificial Intelligence and Machine Learning in Pharmacy: A Narrative Review of Real-World Applications. Pharmacy 2025, 13, 41. [Google Scholar] [CrossRef]
- Hu, Q.; Chen, Y.; Zou, D.; He, Z.; Xu, T. Predicting adverse drug event using machine learning based on electronic health records: A systematic review and meta-analysis. Front. Pharmacol. 2024, 15, 1497397. [Google Scholar] [CrossRef] [PubMed]
- Ietswaart, R.; Arat, S.; Chen, A.X.; Farahmand, S.; Kim, B.; DuMouchel, W.; Armstrong, D.; Fekete, A.; Sutherland, J.J.; Urban, L. Machine learning guided association of adverse drug reactions with in vitro target-based pharmacology. EBioMedicine 2020, 57, 102837. [Google Scholar] [CrossRef]
- Yu, Z.; Ji, H.; Xiao, J.; Wei, P.; Song, L.; Tang, T.; Hao, X.; Zhang, J.; Qi, Q.; Zhou, Y.; et al. Predicting Adverse Drug Events in Chinese Pediatric Inpatients With the Associated Risk Factors: A Machine Learning Study. Front. Pharmacol. 2021, 12, 659099. [Google Scholar] [CrossRef] [PubMed]
- Yalçın, N.; Kaşıkcı, M.; Çelik, H.T.; Allegaert, K.; Demirkan, K.; Yiğit, Ş.; Yurdakök, M. An Artificial Intelligence Approach to Support Detection of Neonatal Adverse Drug Reactions Based on Severity and Probability Scores: A New Risk Score as Web-Tool. Children 2022, 9, 1826. [Google Scholar] [CrossRef] [PubMed]
- Rekha, B.H.; Hisham, S.A.; Wahab, I.A.; Ali, N.M.; Goh, K.W.; Ming, L.C. Digital monitoring of medication safety in children: An investigation of ADR signalling techniques in Malaysia. BMC Med. Inform. Decis. Mak. 2024, 24, 395. [Google Scholar] [CrossRef]
- Zeng, Y.; Lu, H.; Li, S.; Shi, Q.-Z.; Liu, L.; Gong, Y.-Q.; Yan, P. Risk Prediction of Liver Injury in Pediatric Tuberculosis Treatment: Development of an Automated Machine Learning Model. Drug Des. Dev. Ther. 2025, 19, 239–250. [Google Scholar] [CrossRef]
- Mishra, H.P.; Gupta, R. Leveraging Generative AI for Drug Safety and Pharmacovigilance. Curr. Rev. Clin. Exp. Pharmacol. 2025, 20, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Niazi, S.K.; Mariam, Z. Artificial intelligence in drug development: Reshaping the therapeutic landscape. Ther. Adv. Drug Saf. 2025, 16, 20420986251321704. [Google Scholar] [CrossRef] [PubMed]
- Ocana, A.; Pandiella, A.; Privat, C.; Bravo, I.; Luengo-Oroz, M.; Amir, E.; Gyorffy, B. Integrating artificial intelligence in drug discovery and early drug development: A transformative approach. Biomark. Res. 2025, 13, 45. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Sun, X.; Li, Y.; Chu, T.; Hao, X.; Cao, Y.; Zhang, P. Applications of Artificial Intelligence in Drug Repurposing. Adv. Sci. 2025, 12, e2411325. [Google Scholar] [CrossRef]
- Young, C.M.; Phares, S.E.; Kennedy, A.; Sullivan, J.; McGowan, B.; Trusheim, M.R. Pediatric-onset rare disease therapy pipeline yields hope for some and gaps for many: 10-year projection of approvals, treated patients, and list price revenues. J. Manag. Care Spéc. Pharm. 2025, 31, 491–498. [Google Scholar] [CrossRef]
- Choudhury, A.; Asan, O. Role of Artificial Intelligence in Patient Safety Outcomes: Systematic Literature Review. JMIR Med Inform. 2020, 8, e18599. [Google Scholar] [CrossRef]
- Abdalwahab Abdallah, A.B.A.; Hafez Sadaka, S.I.; Ali, E.I.; Mustafa Bilal, S.A.; Abdelrahman, M.O.; Fakiali Mohammed, F.B.; Nimir Ahmed, S.D.; Abdelrahim Saeed, N.E. The Role of Artificial Intelligence in Pediatric Intensive Care: A Systematic Review. Cureus 2025, 17, e80142. [Google Scholar] [CrossRef]
- Bannett, Y.; Gunturkun, F.; Pillai, M.; Herrmann, J.E.; Luo, I.; Huffman, L.C.; Feldman, H.M. Applying Large Language Models to Assess Quality of Care: Monitoring ADHD Medication Side Effects. Pediatrics 2024, 155, e2024067223. [Google Scholar] [CrossRef]
- Haddad, T.; Helgeson, J.M.; Pomerleau, K.E.; Preininger, A.M.; Roebuck, M.C.; Dankwa-Mullan, I.; Jackson, G.P.; Goetz, M.P. Accuracy of an Artificial Intelligence System for Cancer Clinical Trial Eligibility Screening: Retrospective Pilot Study. JMIR Med Inform. 2021, 9, e27767. [Google Scholar] [CrossRef]
- Parsi, A.; Glavin, M.; Jones, E.; Byrne, D. Prediction of paroxysmal atrial fibrillation using new heart rate variability features. Comput. Biol. Med. 2021, 133, 104367. [Google Scholar] [CrossRef]
- Bozyel, S.; Şimşek, E.; Koçyiğit Burunkaya, D.; Güler, A.; Korkmaz, Y.; Şeker, M.; Ertürk, M.; Keser, N. Artificial Intelligence-Based Clinical Decision Support Systems in Cardiovascular Diseases. Anatol. J. Cardiol. 2024, 28, 74–86. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Cabello, C.A.; Borna, S.; Pressman, S.; Haider, S.A.; Haider, C.R.; Forte, A.J. Artificial-Intelligence-Based Clinical Decision Support Systems in Primary Care: A Scoping Review of Current Clinical Implementations. Eur. J. Investig. Health Psychol. Educ. 2024, 14, 685–698. [Google Scholar] [CrossRef]
- Elhaddad, M.; Hamam, S. AI-Driven Clinical Decision Support Systems: An Ongoing Pursuit of Potential. Cureus 2024, 16, e57728. [Google Scholar] [CrossRef]
- Sarvepalli, S.; Vadarevu, S. Role of artificial intelligence in cancer drug discovery and development. Cancer Lett. 2025, 627, 217821. [Google Scholar] [CrossRef]
- Bongurala, A.R.; Save, D.; Virmani, A. Progressive role of artificial intelligence in treatment decision-making in the field of medical oncology. Front. Med. 2025, 12, 1533910. [Google Scholar] [CrossRef]
- Rossner, T.; Li, Z.; Balke, J.; Salehfard, N.; Seifert, T.; Tang, M. Integrating single-cell foundation models with graph neural networks for drug response prediction. arXiv 2025, arXiv:2504.14361. [Google Scholar] [CrossRef]
- Huang, Y.; Su, X.; Ullanat, V.; Liang, I.; Clegg, L.; Olabode, D.; Ho, N.; John, B.; Gibbs, M.; Zitnik, M. Multimodal AI predicts clinical outcomes of drug combinations from preclinical data. arXiv 2025, arXiv:2503.02781. [Google Scholar] [CrossRef]
- Predicting Relapse at the Time of Diagnosis in Acute Lymphoblastic Leukemia. NIH Reporter. Available online: https://reporter.nih.gov/project-details/11047678 (accessed on 22 June 2025).
- Ilyas, M.; Ramzan, M.; Deriche, M.; Mahmood, K.; Naz, A. An efficient leukemia prediction method using machine learning and deep learning with selected features. PLoS ONE 2025, 20, e0320669. [Google Scholar] [CrossRef]
- Zhan, M.; Chen, Z.; Ding, C.; Qu, Q.; Wang, G.; Liu, S.; Wen, F. Risk prediction for delayed clearance of high-dose methotrexate in pediatric hematological malignancies by machine learning. Int. J. Hematol. 2021, 114, 483–493. [Google Scholar] [CrossRef]
- Zhan, M.; Chen, Z.-B.; Ding, C.-C.; Qu, Q.; Wang, G.-Q.; Liu, S.; Wen, F.-Q. Machine learning to predict high-dose methotrexate-related neutropenia and fever in children with B-cell acute lymphoblastic leukemia. Leuk. Lymphoma 2021, 62, 2502–2513. [Google Scholar] [CrossRef]
- Chappell, T.L.; Pflaster, E.G.; Namata, R.; Bell, J.; Miller, L.H.; Pomputius, W.F.; Boutilier, J.J.; Messinger, Y.H. Bloodstream Infections in Childhood Acute Myeloid Leukemia and Machine Learning Models: A Single-institutional Analysis. J. Pediatr. Hematol. 2025, 47, e26–e33. [Google Scholar] [CrossRef]
- Li, M.; Wang, Q.; Lu, P.; Zhang, D.; Hua, Y.; Liu, F.; Liu, X.; Lin, T.; Wei, G.; He, D. Development of a Machine Learning-Based Prediction Model for Chemotherapy-Induced Myelosuppression in Children with Wilms’ Tumor. Cancers 2023, 15, 1078. [Google Scholar] [CrossRef] [PubMed]
- AlGain, S.; Marra, A.R.; Kobayashi, T.; Marra, P.S.; Celeghini, P.D.; Hsieh, M.K.; Shatari, M.A.; Althagafi, S.; Alayed, M.; Ranavaya, J.I.; et al. Can we rely on artificial intelligence to guide antimicrobial therapy? A systematic literature review. Antimicrob. Steward. Healthc. Epidemiol. 2025, 5, e90. [Google Scholar] [CrossRef] [PubMed]
- Cao, K.; Braykov, N.; McCarter, A.; Kandaswamy, S.; Orenstein, E.W.; Ray, E.; Carter, R.; Gleeson, M.B.; Iyer, S.; Muthu, N.; et al. De-velopment and Validation of an Artificial Intelligence Predictive Model to Accelerate Antibiotic Therapy for Critical Ill Children with Sepsis in the Pediatric ED with Pediatric ICU Disposition. medRxiv 2025. [Google Scholar] [CrossRef]
- Tang, B.-H.; Yao, B.-F.; Zhang, W.; Zhang, X.-F.; Fu, S.-M.; Hao, G.-X.; Zhou, Y.; Sun, D.-Q.; Liu, G.; van den Anker, J.; et al. Optimal use of β-lactams in neonates: Machine learning-based clinical decision support system. EBioMedicine 2024, 105, 105221. [Google Scholar] [CrossRef]
- Weissenbacher, D.; Dutcher, L.; Boustany, M.; Cressman, L.; O’Connor, K.; Hamilton, K.W.; Gerber, J.; Grundmeier, R.; Gonzalez-Hernandez, G. Automated Evaluation of Antibiotic Prescribing Guideline Concordance in Pediatric Sinusitis Clinical Notes. Biocomputing 2025, 30, 138–153. [Google Scholar] [CrossRef]
- Yin, M.; Jiang, Y.; Yuan, Y.; Li, C.; Gao, Q.; Lu, H.; Li, Z. Optimizing vancomycin dosing in pediatrics: A machine learning approach to predict trough concentrations in children under four years of age. Int. J. Clin. Pharm. 2024, 46, 1134–1142. [Google Scholar] [CrossRef]
- Huang, X.; Yu, Z.; Bu, S.; Lin, Z.; Hao, X.; He, W.; Yu, P.; Wang, Z.; Gao, F.; Zhang, J.; et al. An Ensemble Model for Prediction of Vancomycin Trough Concentrations in Pediatric Patients. Drug Des. Dev. Ther. 2021, 15, 1549–1559. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, R.; Tang, X.-Y. Towards real-time diagnosis for pediatric sepsis using graph neural network and ensemble methods. Eur. Rev. Med. Pharmacol. Sci. 2021, 25, 4693–4701. [Google Scholar] [CrossRef]
- Ramgopal, S.; Horvat, C.M.; Yanamala, N.; Alpern, E.R. Machine Learning To Predict Serious Bacterial Infections in Young Febrile Infants. Pediatrics 2020, 146, e20194096. [Google Scholar] [CrossRef]
- Lamping, F.; Jack, T.; Rübsamen, N.; Sasse, M.; Beerbaum, P.; Mikolajczyk, R.T.; Boehne, M.; Karch, A. Development and validation of a diagnostic model for early differentiation of sepsis and non-infectious SIRS in critically ill children—A data-driven approach using machine-learning algorithms. BMC Pediatr. 2018, 18, 112. [Google Scholar] [CrossRef]
- Hoffer, O.; Cohen, M.; Gerstein, M.; Shkalim Zemer, V.; Richenberg, Y.; Nathanson, S.; Avner Cohen, H. Machine Learning for Clinical Decision Support of Acute Streptococcal Pharyngitis: A Pilot Study. Isr Med. Assoc. J. 2024, 26, 299–303. [Google Scholar] [PubMed]
- Torres, M.D.T.; Zeng, Y.; Wan, F.; Maus, N.; Gardner, J.; de la Fuente-Nunez, C. A generative artificial intelligence approach for antibiotic optimization. bioRxiv 2024. bioRxiv:2024.11.27.625757. [Google Scholar] [CrossRef] [PubMed]
- Juang, W.-C.; Hsu, M.-H.; Cai, Z.-X.; Chen, C.-M. Developing an AI-assisted clinical decision support system to enhance in-patient holistic health care. PLoS ONE 2022, 17, e0276501. [Google Scholar] [CrossRef]
- Ayyıldız, H.; Tuncer, S.A. Is it possible to determine antibiotic resistance of E. coli by analyzing laboratory data with machine learning? Turk. J. Biochem. 2021, 46, 623–630. [Google Scholar] [CrossRef]
- Pascual-Sánchez, L.; Mora-Jiménez, I.; Martínez-Agüero, S.; Álvarez-Rodríguez, J.; Soguero-Ruiz, C. Predicting Multidrug Resistance Using Temporal Clinical Data and Machine Learning Methods. In Proceedings of the 2021 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Houston, TX, USA, 9–12 December 2021; pp. 2826–2833. [Google Scholar] [CrossRef]
- de la Lastra, J.M.P.; Wardell, S.J.T.; Pal, T.; de la Fuente-Nunez, C.; Pletzer, D. From Data to Decisions: Leveraging Artificial Intelligence and Machine Learning in Combating Antimicrobial Resistance—A Comprehensive Review. J. Med. Syst. 2024, 48, 71. [Google Scholar] [CrossRef]
- de Vries, S.; Doesschate, T.T.; Totté, J.E.E.; Heutz, J.W.; Loeffen, Y.G.T.; Oosterheert, J.J.; Thierens, D.; Boel, E. A semi-supervised decision support system to facilitate antibiotic stewardship for urinary tract infections. Comput. Biol. Med. 2022, 146, 105621. [Google Scholar] [CrossRef] [PubMed]
- Mourid, M.R.; Irfan, H.; Oduoye, M.O. Artificial Intelligence in Pediatric Epilepsy Detection: Balancing Effectiveness With Ethical Considerations for Welfare. Health Sci. Rep. 2025, 8, e70372. [Google Scholar] [CrossRef]
- Zou, Z.; Chen, B.; Xiao, D.; Tang, F.; Li, X. Accuracy of Machine Learning in Detecting Pediatric Epileptic Seizures: Systematic Review and Meta-Analysis. J. Med. Internet Res. 2024, 26, e55986. [Google Scholar] [CrossRef]
- Schumacher, E.; Naik, D.; Kannan, A. Rare disease differential diagnosis with large language models at scale: From abdominal actinomycosis to Wilson’s disease. arXiv 2025, arXiv:2502.15069. [Google Scholar] [CrossRef]
- Barry, J.S.; Beam, K.; McAdams, R.M. Artificial intelligence in pediatric medicine: A call for rigorous reporting standards. J. Perinatol. 2025, 45, 1031–1033. [Google Scholar] [CrossRef]
- Schouten, J.S.; Kalden, M.A.C.M.; van Twist, E.; Reiss, I.K.M.; Gommers, D.A.M.P.J.; van Genderen, M.E.; Taal, H.R. From bytes to bedside: A systematic review on the use and readiness of artificial intelligence in the neonatal and pediatric intensive care unit. Intensive Care Med. 2024, 50, 1767–1777. [Google Scholar] [CrossRef]
- Grand View Research. AI In Healthcare Market Size, Share|Industry Report, 2030. Available online: https://www.grandviewresearch.com/industry-analysis/artificial-intelligence-ai-healthcare-market (accessed on 24 June 2025).
- Aalpha. The Cost of Implementing AI in Healthcare in 2025. Available online: https://www.aalpha.net/blog/cost-of-implementing-ai-in-healthcare/ (accessed on 24 June 2025).
- Axis Intelligence. Healthcare AI Implementation: $2.4M ROI Blueprint for Medical Organizations in 2025. Available online: https://axis-intelligence.com/healthcare-ai-implementation-ai-health-2025/ (accessed on 24 June 2025).
- The AI Journal. The Good, the Bad: Behind the Scenes Economic Impact of AI in Healthcare. Available online: https://aijourn.com/economicimpacthealthcare/ (accessed on 24 June 2025).
- Muralidharan, V.; Burgart, A.; Daneshjou, R.; Rose, S. Recommendations for the use of pediatric data in artificial intelligence and machine learning ACCEPT-AI. npj Digit. Med. 2023, 6, 66. [Google Scholar] [CrossRef] [PubMed]
- Mondillo, G.; Colosimo, S.; Perrotta, A.; Frattolillo, V.; Masino, M. Are LLMs ready for pediatrics? A comparative evaluation of model accuracy across clinical domains. medRxiv 2025. [Google Scholar] [CrossRef]
- Cross, J.L.; Choma, M.A.; Onofrey, J.A. Bias in medical AI: Implications for clinical decision-making. PLoS Digit. Health 2024, 3, e0000651. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Mondillo, G.; Perrotta, A.; Colosimo, S.; Frattolillo, V.; Masino, M.; Pettoello-Mantovani, M. Artificial intelligence in pediatrics: An opportunity to lead, not to follow. J. Pediatr. 2025, 283, 114641. [Google Scholar] [CrossRef]
- Ganatra, H.A. Machine Learning in Pediatric Healthcare: Current Trends, Challenges, and Future Directions. J. Clin. Med. 2025, 14, 807. [Google Scholar] [CrossRef] [PubMed]
- Faiyazuddin, M.; Rahman, S.J.Q.; Anand, G.; Siddiqui, R.K.; Mehta, R.; Khatib, M.N.; Gaidhane, S.; Zahiruddin, Q.S.; Hussain, A.; Sah, R. The Impact of Artificial Intelligence on Healthcare: A Comprehensive Review of Advancements in Diagnostics, Treatment, and Operational Efficiency. Health Sci. Rep. 2025, 8, e70312. [Google Scholar] [CrossRef]
- Kandaswamy, S.; Knake, L.A.; Dziorny, A.C.; Hernandez, S.M.; McCoy, A.B.; Hess, L.M.; Orenstein, E.; White, M.S.; Kirkendall, E.S.; Molloy, M.J.; et al. Pediatric Predictive Artificial Intelligence Implemented in Clinical Practice from 2010 to 2021: A Systematic Review. Appl. Clin. Inform. 2025, 16, 477–487. [Google Scholar] [CrossRef]
- Bhargava, H.; Salomon, C.; Suresh, S.; Chang, A.; Kilian, R.; van Stijn, D.; Oriol, A.; Low, D.; Knebel, A.; Taraman, S. Promises, Pitfalls, and Clinical Applications of Artificial Intelligence in Pediatrics. J. Med. Internet Res. 2024, 26, e49022. [Google Scholar] [CrossRef]
- Ramgopal, S.; Sanchez-Pinto, L.N.; Horvat, C.M.; Carroll, M.S.; Luo, Y.; Florin, T.A. Artificial intelligence-based clinical decision support in pediatrics. Pediatr. Res. 2022, 93, 334–341. [Google Scholar] [CrossRef] [PubMed]
- Shortliffe, E.H.; Sepúlveda, M.J. Clinical decision support in the era of artificial intelligence. JAMA 2018, 320, 2199–2200. [Google Scholar] [CrossRef] [PubMed]
- Balla, Y.; Tirunagari, S.; Windridge, D. Pediatrics in Artificial Intelligence Era: A Systematic Review on Challenges, Opportunities, and Explainability. Indian Pediatr. 2023, 60, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Elzagallaai, A.; Barker, C.; Lewis, T.; Cohn, R.; Rieder, M. Advancing Precision Medicine in Paediatrics: Past, present and future. Camb. Prism. Precis. Med. 2023, 1, e11. [Google Scholar] [CrossRef]
- Shahin, M.H.; Desai, P.; Terranova, N.; Guan, Y.; Helikar, T.; Lobentanzer, S.; Liu, Q.; Lu, J.; Madhavan, S.; Mo, G.; et al. AI-Driven Applications in Clinical Pharmacology and Translational Science: Insights From the ASCPT 2024 AI Preconference. Clin. Transl. Sci. 2025, 18, e70203. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.