Abstract
Background: The signal peptide-CUB-EGF-like domain-containing protein 3 (SCUBE3) is a secretory protein that plays a role in cancer, cardiovascular, and immune disorders. SCUBE1, SCUBE2, and SCUBE3 belong to the SCUBE family. They contain multiple copies of EGF-like repeats at the amino acid terminal, a spacer region, three cysteine-rich motifs, and a CUB domain at the carboxyl terminus. The SCUBE family members are multifunctional proteins that act primarily as extracellular ligands or co-receptors in various cells. Methods: In this study, we examined the expression pattern and role of SCUBE3 in various cancers, as well as other diseases such as cardiovascular disease and immune disorders, and its impact on growth and development. Results: SCUBE3 expression is upregulated and secreted by the cells of lung cancer, hepatocellular carcinoma (HCC), melanoma, osteosarcoma, ovarian cancer, glioma, and breast cancer. Extracellular SCUBE3 protein often binds to TGFβRII or acts as a co-receptor for TGFβ and BMP2/BMP4 in regulating cellular signaling. Through the TGFβRII signaling, SCUBE3 activities promote tumor growth, metastasis, invasion, angiogenesis, and poor clinical outcomes. Conversely, in renal cell carcinoma, SCUBE3 expression suppresses growth. Altered SCUBE3 activity is associated with cardiovascular diseases, immune disorders, and hair growth. Conclusions: The review presents mechanistic evidence that SCUBE3 plays a crucial regulatory role in multiple cancers and other diseases. The evidence suggests the SCUBE3 protein could serve as a potential molecular target for various diseases and highlights its usefulness as a minimally invasive diagnostic marker, as it is a secreted protein.
Keywords:
SCUBE3; cancer; biomarker; ligand; co-receptor; TGFβRII; target; invasion; angiogenesis; metastasis; proliferation; clinical outcome 1. Introduction
Signal peptide-CUB-EGF-like domain-containing protein 3 (SCUBE3) is a 993 amino acid member of the SCUBE protein family. The SCUBE3 protein family consists of three members: SCUBE1, SCUBE2, and SCUBE3. The members are highly conserved in humans, mice, and zebrafish [1]. SCUBE3 protein shares about 65.6% homology with SCUBE1 and 58.5% with SCUBE2 in their amino acid compositions. SCUBE1 and SCUBE2 are 65.6% identical in their amino acid compositions [1]. These members are expressed differently in tissues of normal cells. The SCUBE1 protein is primarily expressed in platelets and endothelial cells. Outside the endothelial cells, SCUBE2 is broadly expressed in several cells and tissues. On the other hand, human SCUBE3 is ubiquitously expressed in osteoblasts and bone [2]. The association of SCUBE3 with cancer promotion is widely reported. Secreted SCUBE3 protein is crucial in inducing transforming growth factor-β receptor II (TGFβRII) activation and in increasing the expression of TGF-β. These events promote growth, metastasis, invasion, and angiogenesis. Interestingly, Scube3 is expressed in ectodermal, endodermal, and mesodermal-derivative in mice suggesting that it may be involved in development [3]. Embryonic depletion of Scube3 in mice has no physiological or developmental defect, suggesting that it is not essential for embryonic development or survival [4]. However, the developmental disorder observed due to defective SCUBE3 has been linked to the regulation of growth, morphogenesis, and teeth development through the Bone Morphogenetic Protein (BMP) signaling [5]. BMPs are among the TGF-β superfamily, and the family members are critical regulators of cell growth, differentiation, and apoptosis [6].
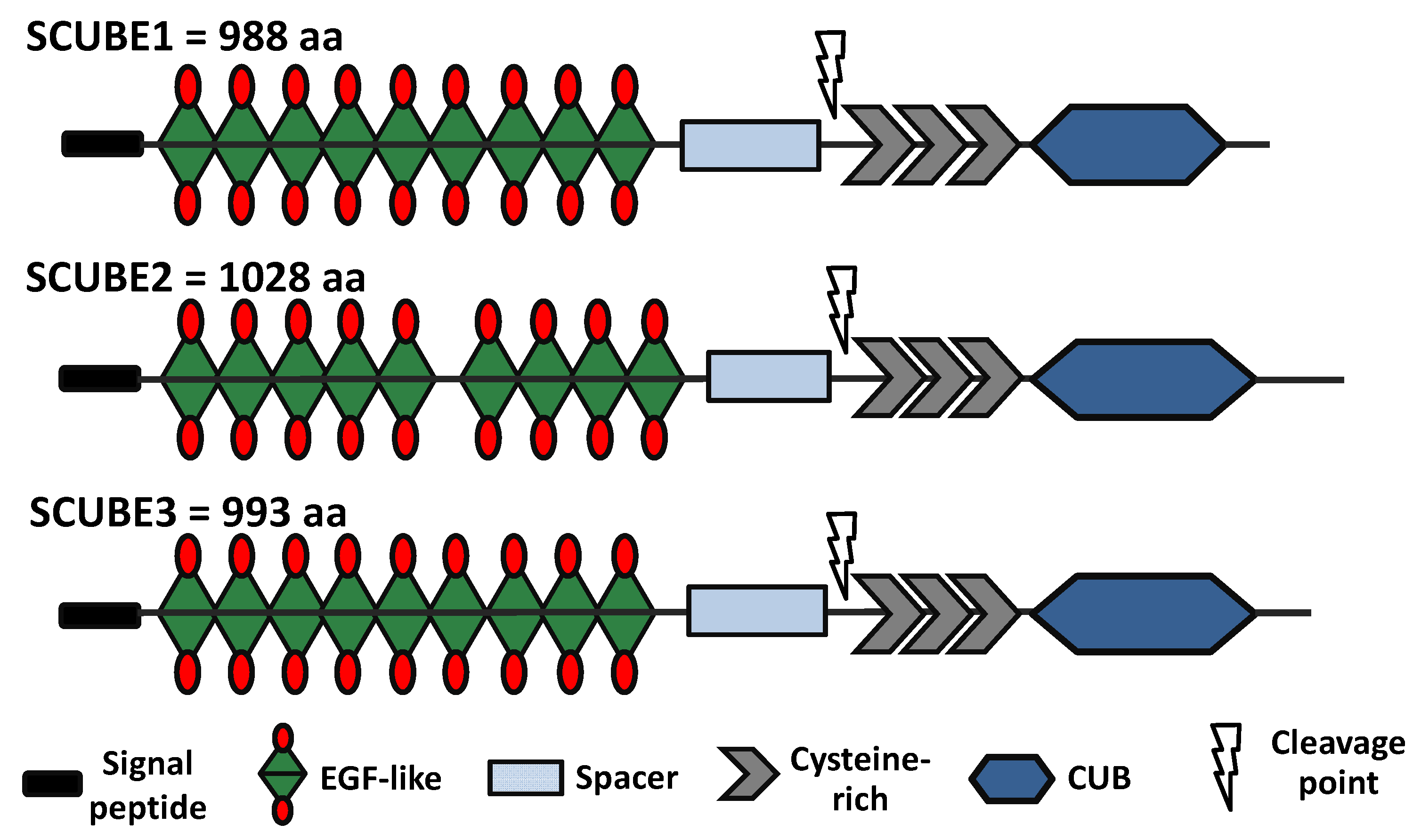
SCUBE family members are secreted glycosylated proteins containing five protein motifs: N-terminal signal peptide, epidermal growth factor (EGF)-like repeat domain, a spacer region, three cysteine-rich domains, and a complement protein C1r/C1s, Uegf, and Bmp1 (CUB) C-terminal domain (Figure 1). The highest homology among the members is found in the CUB domain (83%), followed by the EGF-like (73%) and Cys-rich (66%) domains. In contrast, the spacer also present in all members exhibits the lowest homology at about 34% [1]. This shows the functional importance of the regions to the members. Despite having the lowest amino acid homology among the members, the cysteine residues are instrumental for the diverse functions observed in the SCUBE members, as they can promote disulfide linkage with other proteins, promote protein stability, and serve as a proteolytic cleavage region (Figure 1). The proteolytic cleavage at the spacer region controls the modulatory function mediated by the C-terminal CUB domain and secretion of the N-terminal domain protein in the extracellular space as a soluble protein [7]. The overexpression of SCUBE3 has been observed in tissue specimens such as lung cancer, non-small cell cancer, breast cancer, renal carcinoma, glioma, ovarian, and osteosarcoma compared to the associated healthy tissues.
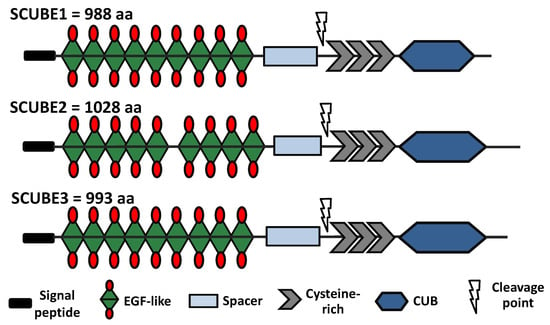
Figure 1.
Schematic structure of human SCUBE family members showing the different structural components: signal peptide, EGF-like, spacer, cysteine-rich, and CUB. The cysteine-rich region is prone to cleavage by MMP2 and MMP9.
Each SCUBE gene is located on a different chromosome and is differentially expressed. The gene for SCUBE3 is uniquely found on human chromosome 6, while the SCUBE1 and SCUBE2 genes are located on human chromosomes 22 and 11, respectively [1]. This shows that the SCUBE3 members are regulated differently. SCUBE3 protein was first isolated in the human umbilical vein endothelial cells (HUVECs) where it is significantly high [8]. Its 993 amino acid composition is heavily glycosylated, causing the molecular weight to be approximately 135 kDa. Report shows that SCUBE3 can be cleaved by MMP2 and MMP9 into the N-terminal containing the EGF-like repeat domain and C-terminal CUB domains, enabling it to interact with multiple proteins and be involved in different pathways [9]. Increasing evidence has linked SCUBE3 to cancers such as lung cancer, breast cancer, hepatocellular carcinoma, renal carcinoma, and osteosarcoma. Reports also show that elevated SCUBE3 expression is associated with poor clinical outcomes [10,11,12]. Also, differential alteration in the SCUBE3 gene, such as mutation, amplification, and deletion, could alter the normal function of the SCUBE3 protein in cells. However, the impacts of these alterations have yet to be fully deciphered. Our review is focused on the functional role of SCUBE3 in different cancer models, cardiovascular diseases, and immune disorders, and the potential benefit of developing new therapeutic agents for inhibiting SCUBE3 functions and as well making it a biomarker for cancer diagnosis. SCUBE3 overexpression is linked to various cancers, including lung cancer, hepatocellular carcinoma (HCC), ovarian cancer, osteosarcoma, breast cancer, melanoma, renal cell carcinoma, and glioma. It also regulates multiple signaling pathways in cancer, suggesting a need for pan-cancer targeting of SCUBE3.
Here, we provide a comprehensive review of the role of SCUBE3 in cancer development, cardiovascular disease, immune disorder, and fast muscle development.
2. SCUBE3 in Different Cancers
2.1. Lung Cancer
Lung cancer is the second most common type of cancer and the number one cause of death due to cancer in the United States [13]. Approximately 350 people die each day from lung cancer, with about 81% of the deaths connected to cigarette smoking and the rest linked to second-hand smoking [14,15]. Each year, approximately 226,650 individuals are diagnosed with lung cancer, and around 124,730 people succumb to the disease [13]. Exposure to radon, a naturally occurring radioactive element found in tobacco, increases the risk of lung cancer among tobacco smokers [16,17]. Radon is the second leading cause of lung cancer in non-smokers. It is odorless and colorless gas. When inhaled radon attack and damage lung cells. Approximately 85% of lung cancer cases are classified as non-small cell lung cancer (NSCLC) [18,19]. SCUBE3 is overexpressed in roughly 70.6% of NSCLC tissue specimens [10]. Elevated SCUBE3 protein level was observed in extremely invasive lung cancer. Another investigation demonstrated that SCUBE3 is significantly involved in the regulation of angiogenesis and increasing the metastatic potential in early lung cancer [10,20].
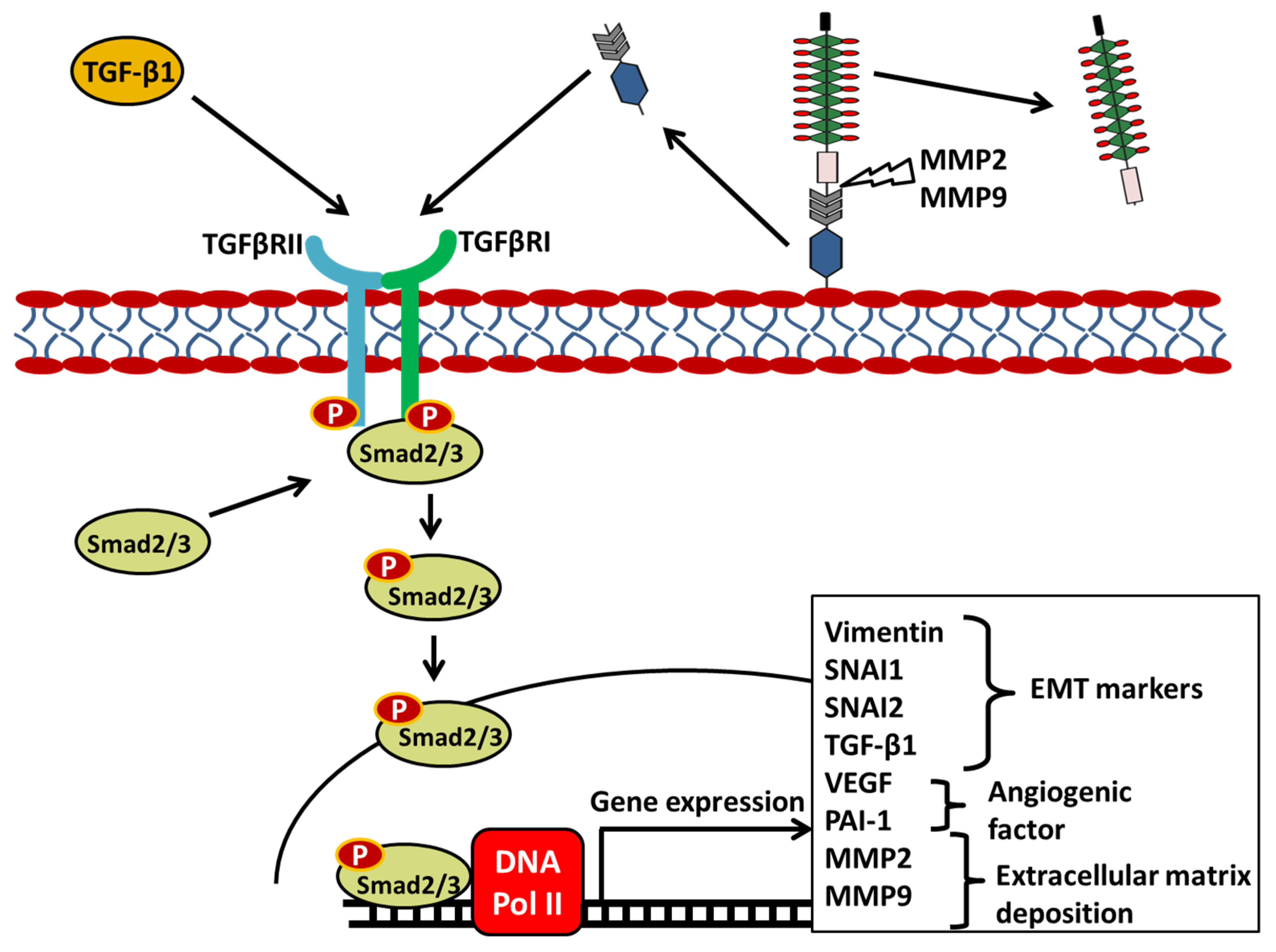
Upregulation of SCUBE3 expression alters the epithelial–mesenchymal transition (EMT) markers in non-small cell lung cancer. SCUBE3 involvement in EMT could result from its ability to bind and activate the transforming growth factor-β (TGF-β) type II receptor through the C-terminal CUB domain [9]. TGFβRII activation, in turn, stimulates the EMT processes, leading to the induction of receptor activated Smad2/3 by phosphorylation that triggers the localization of the Smad2/3 to the nucleus, where it acts as a transcriptional factor responsible for increasing the expression of matrix metalloproteinase-2 (MMP-2), matrix metalloproteinase-2 (MMP-9), plasminogen activator inhibitor type-1 (PAI-1), vascular endothelial growth factor (VEGF), and the transcription factors SNAI1 (Snail) and SNAI2 (Slug) involved in epithelial to mesenchymal transition (EMT) and cancer progression (Figure 2) [9,21].
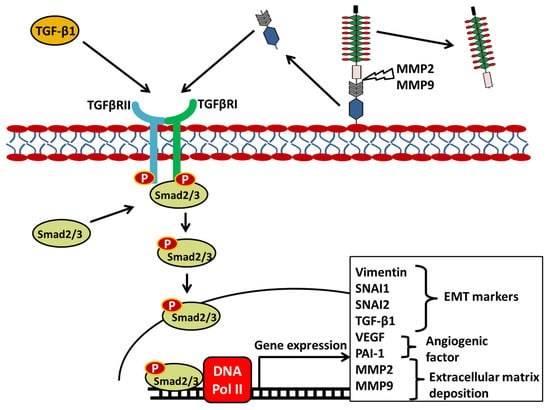
Figure 2.
SCUBE3 Regulates the TGFβRII Signaling Pathway in Lung Cancer. Secreted SCUBE3 is cleaved by MMP2 or MMP9 into the N-terminal and C-terminal domains. The C-terminal domain bearing the CUB domains binds to the TGFβRII triggering the phosphorylation and subsequent activation of the intracellular domain of the receptors. Smad2/3 binds to the intracellular domain and become activated, then translocate to the nucleus where they bind on the DNA to induce the expression of EMT markers, angiogenic factors, and extracellular matrix protein. Arrows indicate activation of signal transduction. Protein cleavage sites are indicated by a lightning bolt symbol.
Remarkably, the secreted SCUBE3 protein is cleaved by MMP2 and MMP9 into two fragments: one containing the EGF-like domain and spacer and the other containing the CUB domain and cysteine rich portion [9]. Silencing of SCUBE3 expression in NSCLC suppressed the proliferation and metastasis of tumor cells in an in vivo model [10]. Given the complexity of lung cancer, it is imperative to gain a comprehensive understanding of the molecular mechanisms involved in the development and progression of lung cancer in order to help in the development of effective therapies that focus on inhibiting SCUBE3 function.
2.2. Osteosarcoma
The SCUBE3 expression is associated with osteosarcoma, the most common primary bone cancer that affects children and young adults [22,23]. Osteosarcoma is characterized as an extremely rapid progressive and invasive cancer. SCUBE3 was identified among the heat shock protein family D (Hsp60) member 1 (HSPD1) and chemokine ligand 14 (CXCL14) as a signature gene that correlated with the prognosis of osteosarcoma [24]. The SCUBE3 protein level was significantly high compared to normal osteocytes [24].
The HSPD1 is encoded in the mitochondria and is essential for the folding and assembly of proteins imported into the mitochondria. Silencing of the HSPD1 suppressed the proliferation and progression of ovarian cancer and oral squamous cell carcinoma [25,26]. Similarly, aberrant expression of the CXCL14, which is involved in the innate immune system response, is linked to colorectal cancer, breast cancer, prostate cancer, and ovarian cancer [27,28,29]. Overexpression of SCUBE3 reduces the overall survival rate and improves poor treatment outcomes in osteosarcoma patients [24,30]. The suppression of SCUBE3, HSPD1, and CXCL14 expression significantly attenuated tumor cell proliferation. Additional evidence shows that SCUBE3 could be involved in regulating the calcium ion pathway in osteosarcoma [31] Dysregulation of the calcium ion pathway contributes to cancer. Together, these data suggest that treatment focusing on inhibition of SCUBE3, HSPD1, and CXCL14 genes or their proteins could improve treatment in osteosarcoma patients.
2.3. Hepatocellular Carcinoma
Hepatocellular carcinoma (HCC) is the fifth and ninth most frequent cancer in men and women, respectively, with a yearly incidence of 42,240 and 30,090 deaths [32]. HCC, cholangiocarcinoma, and angiosarcoma are the three main types of liver cancer, with HCC accounting for about 90% of all cases worldwide [33]. HCC prevalence is very low in the United States and is prevalent in Asia [34]. Globally, HCC is the second leading cause of death arising from cancer [35]. The Wnt signaling pathway, MAPK signaling pathway, TGF-β signaling pathway, and NF-κB signaling pathway are amongst the most common dysregulated signaling pathways associated with the development, growth, and metastasis of HCC [36,37]. Specific targeting of any pathways could inhibit tumor cell proliferation and progression.
The TGF-β is essential in cell differentiation and normal liver cell regeneration. However, its elevated activity in the liver is highly associated with the development and progression of the HCC. SCUBE3 expression is up-regulated in hepatocellular carcinoma tissue specimens and cell lines [12]. SCUBE3 overexpression promotes cell proliferation by regulating the expression of cyclin E1 (CCNE1) in HCC [12]. CCNE1 promotes cell survival and poor treatment outcomes in HCC and several cancers (ovarian cancer, breast cancer, and gastric cancer) [38,39,40,41,42]. Evidence demonstrated that inhibition of SCUBE3 expression in the HCC cells significantly attenuates cell proliferation and promotes apoptosis [12]. Also, a pre-clinical mouse model study demonstrated that SCUBE3 knockdown significantly inhibited xenograft tumor cell growth, indicating that SCUBE3 knockdown decreased tumor cell proliferation in vivo.
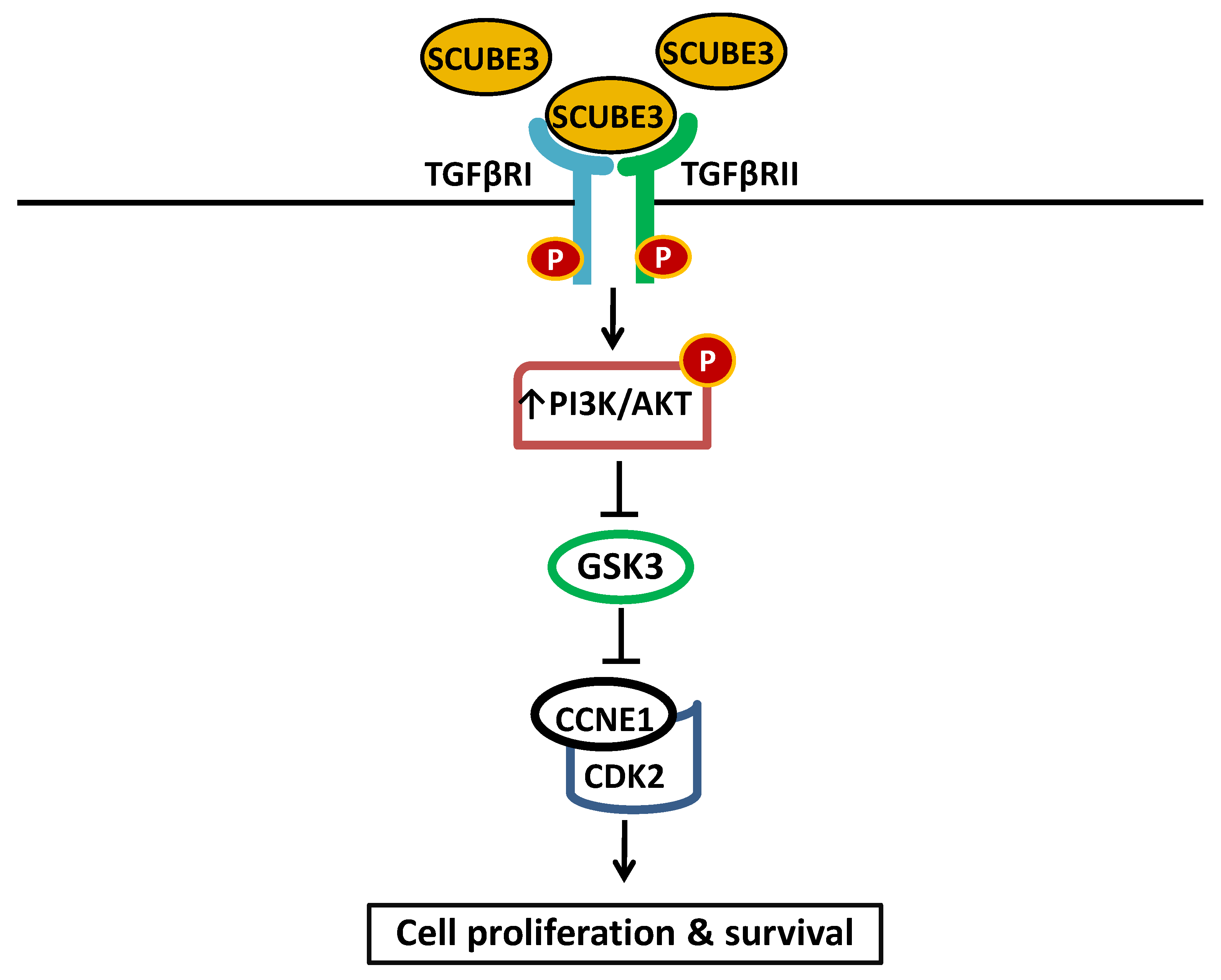
To determine the role of SCUBE3 in cell proliferation and apoptosis, SCUBE3 gene was knockdown using lentivirus particles carrying three SCUBE3 shRNAs in Bel7404 cells. Interestingly, the SCUBE3 knockdown cells have an increased number of Annexin V-APC/PI-positive cells and an increased number of cells in the G1 phase but a reduced number of cells in the S-and G2/M phases. These suggest that SCUBE3 knockdown suppressed proliferation and promoted apoptosis in HCC cells. Also, the knockdown of SCUBE3 significantly attenuated the oncogenic PI3K/AKT signaling pathway and GSK3β phosphorylation [12]. Using immunoprecipitation, the study showed that SCUBE3 protein interacts with TGFβRII. Further analysis indicated that the SCUBE3 protein interacts with the TGFβRII receptor and suggested that regulation of the PI3K/AKT signaling pathway occurs through the TGFβRII. Constitutive activation of PI3K/AKT signaling is known to promote processes associated with tumorigenesis and features that fuel cancer progression [43]. The result demonstrated the importance of the TGFβRII receptor signaling to the SCUBE3 oncogenic function in HCC. It also indicates that in HCC, SCUBE3 promotes cancer by activating the TGFβRII receptor, which then regulates the PI3K/AKT and GSK3β, causing the accumulation of CCNE1 (Figure 3). The CCNE1 accumulation due to the reduced activities of GSK3β promotes the progression of the HCC cells from the G1 phase to the S phase, thus promoting cell proliferation. This shows that SCUBE3 inhibitor-based therapy could be helpful in treating patients with HCC.
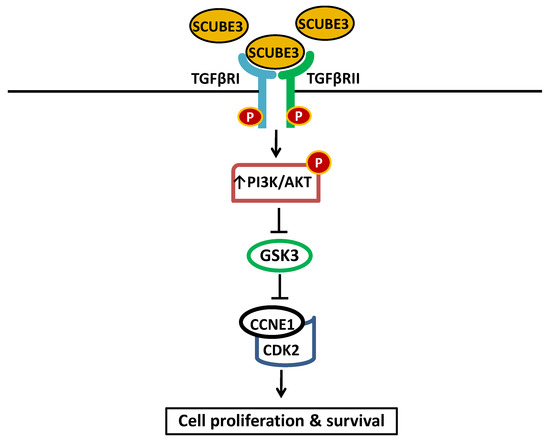
Figure 3.
A schematic model illustrating how SCUBE3 is regulating the TGFβRII and PI3K/AKT in HCC. SCUBE3/TGFβRII signaling function is mediated through the PI3K/AKT. Binding to the TGFβRII by SCUBE3 leads to the activation and PI3K/AKT pathways and subsequent inhibition of GSK3β. This also led to the inhibition of the ubiquitination of CCNE1, causing its accumulation and promoting interaction with CDK2 and the subsequent promotion of the proliferation of HCC cells. Solid pointed arrows indicate the activation of the pathway, while flathead (or blunt) arrows indicate inhibition of the pathway.
Interestingly, in another study examining the effect of Triphenyl phosphate (TPP) on the proliferation, invasion, and migration of the HCC cell line, SCUBE3 was identified as one of the genes significantly upregulated by TPP treatment [44]. TPP is one of the organophosphate flame retardants (OPFR) found in the environment. The OPFRs are metabolized in the liver. The result showed that at low concentrations, TPP promoted Hep3B cell invasion and migration. In a tumor-bearing mouse model, HCC promotes tumor growth.
2.4. Melanoma
Just like normal cells, cancer cells require access to circulating blood. The growth and survival of cancer cells in the body also depend on their ability to increase blood vessels to the tumor peripheries. Blood carries nutrients, oxygen, and many soluble factors important for cellular life solutions. In a mouse model study, the data demonstrated that SCUBE3 expression promoted angiogenesis in melanoma [45]. Although melanoma accounts for approximately 1% of skin cancer cases, it is among the most aggressive type of cancer and the number one cause of death resulting from skin cancer. About 80% of all skin cancer deaths are due to melanoma [46]. Mutations of B-Raf (BRAF), NRAS, and CDKN2A genes are common in melanoma and mostly result in constitutive activation of the MAPK pathway, which are very common in melanoma [47,48].
The most common mutation of B-RAF is BRAFV600E, which is a missense mutation that resulted from substitution of valine with glutamic acid at the 600th amino acid. This change led to an abnormal flip of the catalytic domain that generates a constitutively active conformation with a kinase activity that is 500-fold higher than wild-type BRAF kinase [49,50]. The mutation of the BRAFV600E plus a dysregulation of any oncogenic genes such as NRAS Q61K, often results in an aggressive form. BRAFV600E mutation promotes cell proliferation, survival, and metastasis. The neuroblastoma RAS (NRAS) mutation is the second most common mutation, accounting for about 15–30% of the cases. NRAS is a viral oncogene homology and was first discovered in neuroblastoma cells by Robin Weiss led team at Institute of Cancer Research in London [51]. The majority of these forms of mutation are missense mutations at codons 12, 16, or 61. This mutation causes a prolonged NRAS binding with active GTP, resulting in increased NRAS signaling through MAPK kinase and PI3K pathways that promote the proliferation and survival of cells [52,53,54]. These mutations lead to constitutive activation of the MAPK kinase pathway, which promotes the proliferation and survival of cells.
The CDKN2A gene is a tumor suppressor that plays a vital role in regulating cell division. It encodes two important proteins: p16 (also known as p16INK4A) and p14 (ARF) [48]. These proteins are crucial in controlling the progression of the cell cycle from the G1 phase to the S phase. The p16(INK4A) protein specifically interacts with CDK4 and CDK6, which are proteins that promote cell cycle advancement and lead to cell division. By binding to these cyclin-dependent kinases, p16(INK4A) inhibits their activity, effectively preventing cell division. In contrast, the p14(ARF) protein functions to block the degradation of p53 proteins, which are also essential tumor suppressors that regulate the cell cycle. The loss of CDKN2A function promotes cell proliferation.
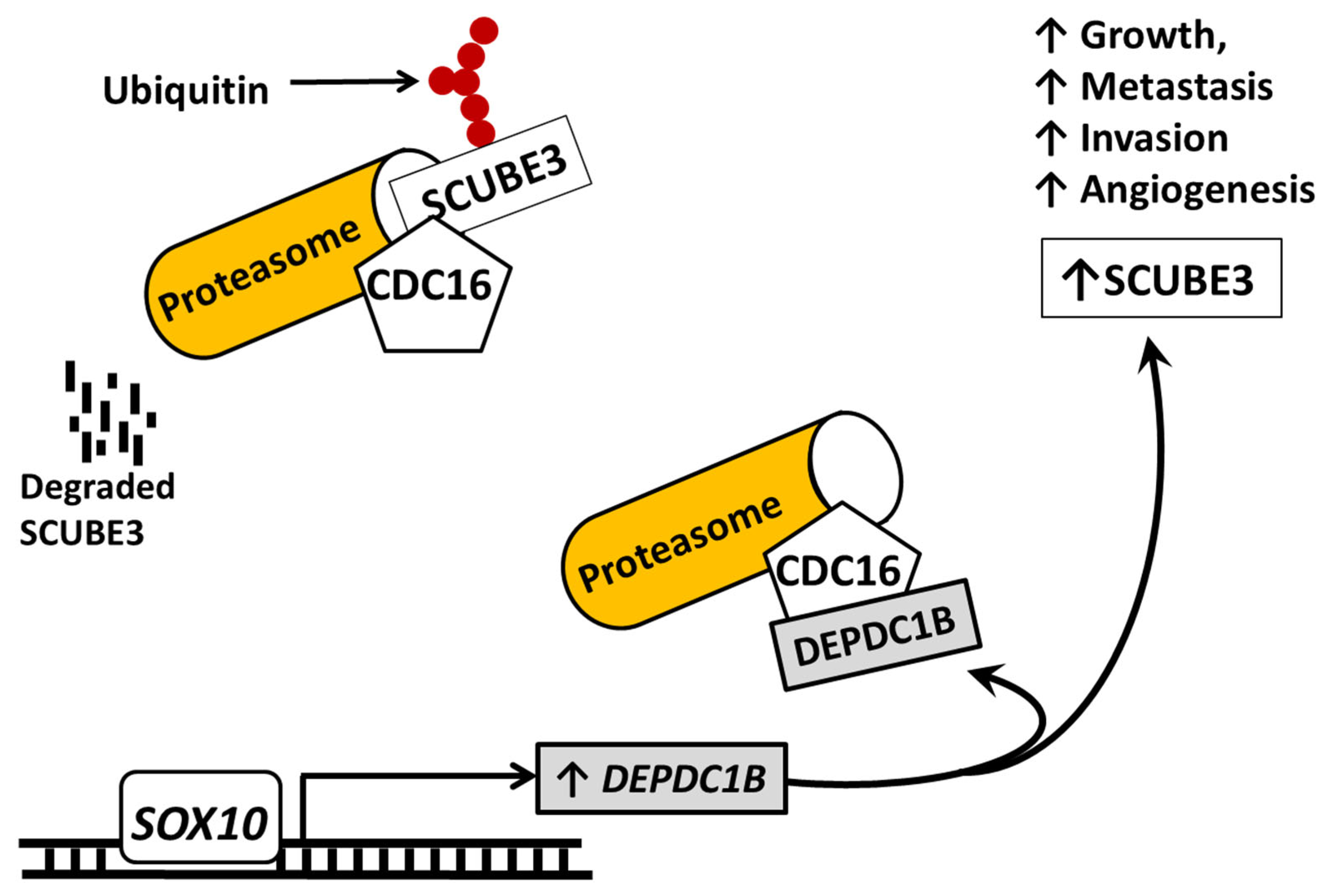
The overexpression and role of SCUBE3 in melanoma is not well reported. Recently published report has linked SCUBE3 to melanoma [45]. SCUBE3 protein is stabilized by the presence of the DEP domain containing 1B (DEPDC1B), which competitively binds to ubiquitin ligase cell division cycle 16 (CDC16) to prevent SCUBE3 from undergoing degradation through the ubiquitin-proteasome pathway [45]. DEPDC1B overexpression has been associated with cell proliferation, metastasis, and a poor treatment outcome [55]. Like DEPDC1B, SOX10 is known to promote proliferation, tumor formation, and growth [56]. The expression of SCUBE3 is directly correlated with the transcription factor SOX10 and DEPDC1B. DEPDC1B is downstream of SOX10, which regulates its expression. In turn, DEPDC1B mediates SOX10 functions such as the promotion of growth, metastasis, and angiogenesis.
DEPDC1B plays an important role in cell cycle progression allowing cells to lose focal adhesion long enough for cells to progress through mitosis where they become rounded prior to cytokinesis. The process competes with RhoA for interaction with focal adhesion-associated receptor protein tyrosine phosphatase, receptor type F (PTPRF) and also promotes the degradation of PTPRF resulting in de-attachment of the cells [57]. Similarly, the expression of SCUBE3 correlated to the DEPDC1B expression because of DEPDC1B involvement in protecting SCUBE3 from ubiquitination and subsequent degradation by the proteasome complex [45]. In the absence of DEPDC1B, CDC16 binds to SCUBE3 and initiates the ubiquitination of SCUBE3. Thus, DEPDC1B binding to the CDC16, inhibit the CDC16 from binding to SCUBE3. The CDC16 is a component of the anaphase promoting complex/cyclosome (APC/C), a cell cycle-regulating E3 ubiquitin ligase that control cycle progression through mitosis and the G1 phase [58]. The SCUBE3 protein level was significantly decreased when DEPDC1B was knocked down. The stabilization of SCUBE3 by DEPDC1B promotes increased SCUBE3 protein, enabling it to promote growth and angiogenesis in melanoma (Figure 4). Knockdown of SCUBE3 alone inhibits angiogenesis in DEPDC1B overexpressed melanoma cells. Likewise, the overexpression of SCUBE3 restored DEPDC1B expression and angiogenesis. These findings highlight the importance of SCUBE3 in cell division, suggesting it could be a potential target for melanoma treatment.
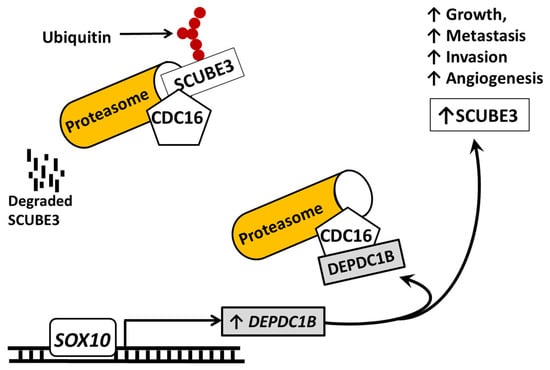
Figure 4.
The schematic diagram of illustration of events that alter SCUBE3 expression and function in melanoma. In the absence of DEPDC1B, the CDC16 binds to SCUBE3 in the cytoplasm to cause SCUBE3 ubiquitination and degradation by the proteasome complex. However, this action is inhibited by the expression of DEPDC1B by SOX10 because DEPDC1B will competitively bind to the site on the same site the CDC16 uses to attach to SCUBE3 and competitively prevent it from binding to SCUBE3. This binding by DEPDC1B prevents the binding by CDC16 and the degradation of SCUBE3. Overall, the action causes an increase in SCUBE3 protein and its tumor promotion action in melanoma. Solid pointed arrows indicate the activation of the pathway or process, while flathead (or blunt) arrows indicate inhibition of the pathway.
2.5. Breast Cancer
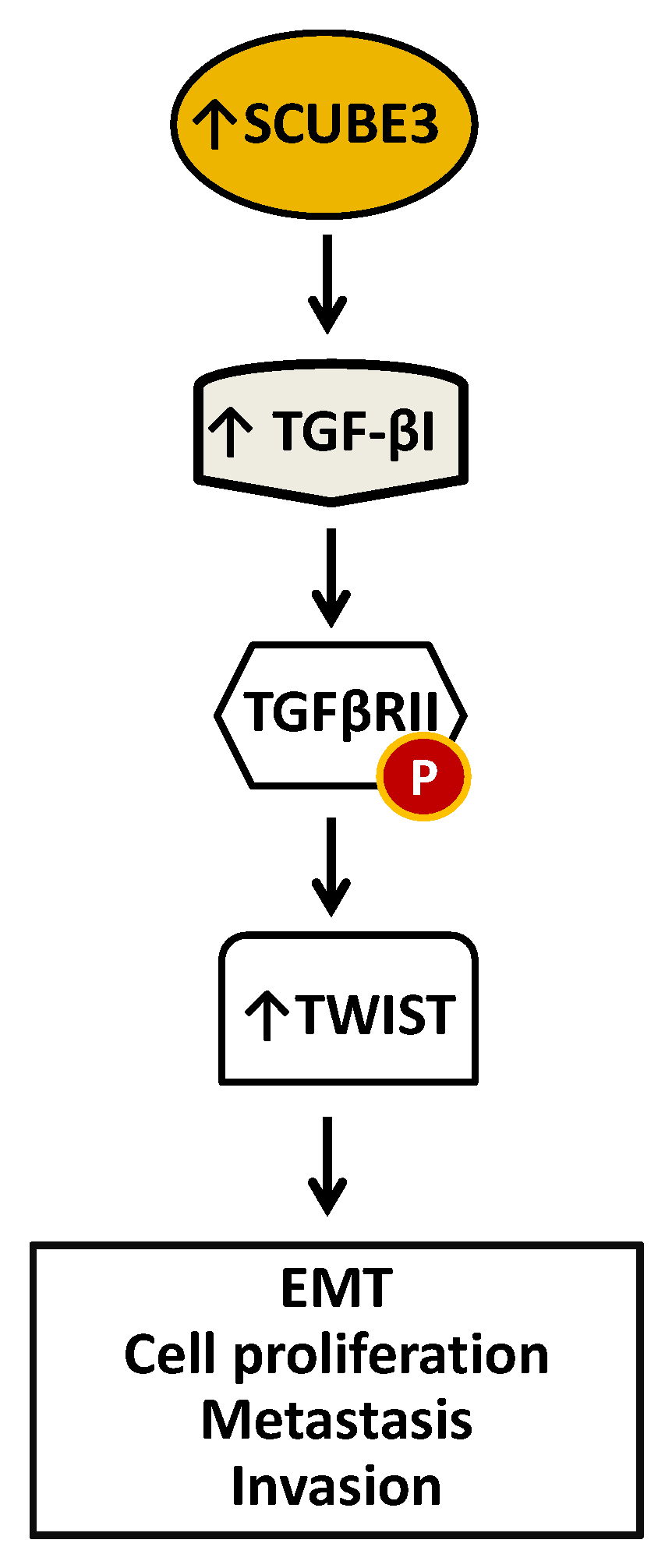
Arguably, breast cancer is the most frequent type of cancer and the second cause of cancer-related death in women [13]. Accumulating evidence has linked SCUBE3 overexpression to the development of several cancers, including breast cancer [11,59]. Like in several other cancers discussed, the elevation of SCUBE3 expression promotes the growth and progression of breast cancer cells. SCUBE3 expression was absent in normal breast mucosa but elevated in breast cancer tissue-invasive ductal carcinoma and invasive lobular carcinoma [11,59]. Remarkably, like in other cancers, SCUBE3 also constitutively activates the TGFβRII signaling in breast cancer cells, causing the cells to become metastatic and invasive. There is a direct correlation between the level of SCUBE3 expression and high histologic grade and negative E-cadherin expression tumors, with the highest SCUBE3 expression observed in late-stage cancer [11]. Also, there is a correlation in the expression of SCUBE3 and TGF-β1. Increased SCUBE3 expression leads to an increase in TGF-β1, a ligand of the TGFβRII, resulting in elevated TWIST1 expression (Figure 5). TWIST1 is a member of the TWIST family of basic helix-loop-helix (bHLH) factors and is an evolutionarily conserved family of proteins that play diverse roles in both embryonic development and pathological disease [60,61]. TWIST1 protects cancer cells from apoptosis and promotes cancer cell stemness, and drug resistance [60]. TWIST protein members have been the focus of much research for many years.
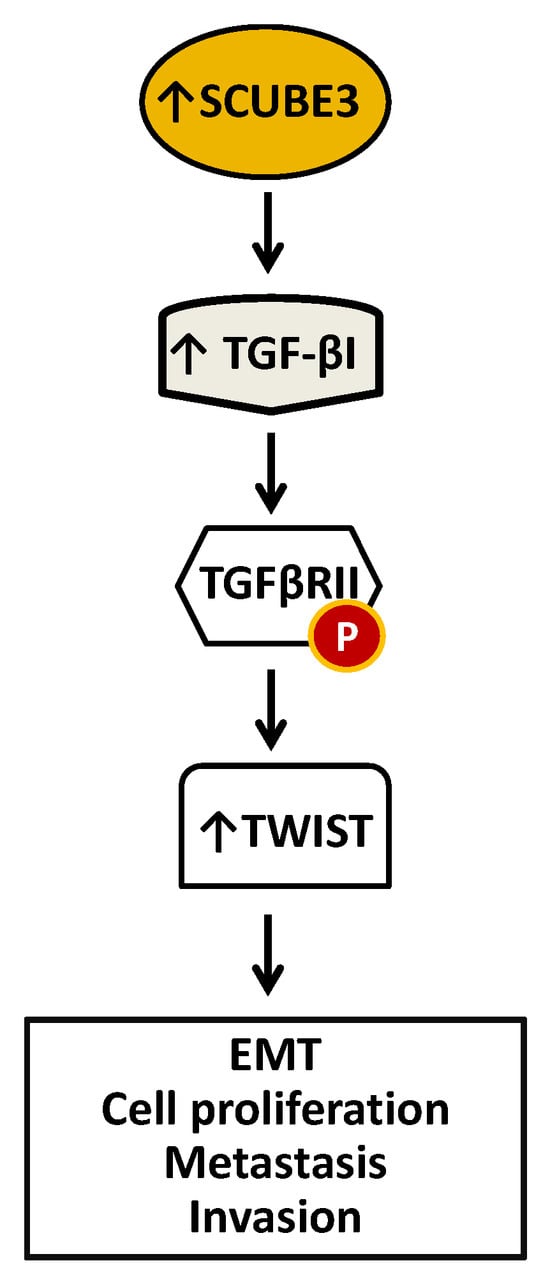
Figure 5.
A diagrammatic model of SCUBE3 mediated events in breast cancer. Overexpression of SCUBE3 caused an increase in TGF-β1, which binds and activates the TGFβRII by phosphorylation. Activated TGFβRII signals the expression of TWIST critical for EMT, proliferation, metastasis, and invasion.
A report has also shown that SCUBE3 is a ligand of TGFβRII and interacts with TGFβRII with its CUB domain [9]. Therefore, the elevation of both SCUBE3 and TGF-β1 would likely increase the activation of TGFβRII. The knockdown of SCUBE3 or TGF-β1 significantly suppressed growth, metastasis, invasion, and angiogenesis. The detection of SCUBE3 in the early stages of breast cancer suggests that SCUBE3 may be involved in tumor development and may be a marker to differentiate tumor lesions with a high risk of malignancy [11]. SCUBE3 expression is linked with decreased patient overall survival in breast cancer patients. Analysis using Kaplan–Meier demonstrated that increased expression of SCUBE3 was associated with poor prognosis in TMN3, lymph node (+), ER−, PR-, and HER2 +. Altogether, these findings suggest that high SCUBE3 expression is associated with poor treatment outcomes and that it could serve as an independent poor prognostic marker and a potential therapeutic target for breast cancer.
A recent study based on a prognostic risk scoring system identified the upregulation of SCUBE3 mRNA among three other mRNAs (RDH16, SPC24, and SPC25) that are differentially associated with HER2-positive breast cancer [62]. The aim of the study is to help improve the screening of high-risk patients with HER2-positive breast cancer. The result suggests that SCUBE3, RDH16, SPC24, and SPC25 are essential for the progression of HER2-positive breast cancer.
2.6. Ovarian Cancer
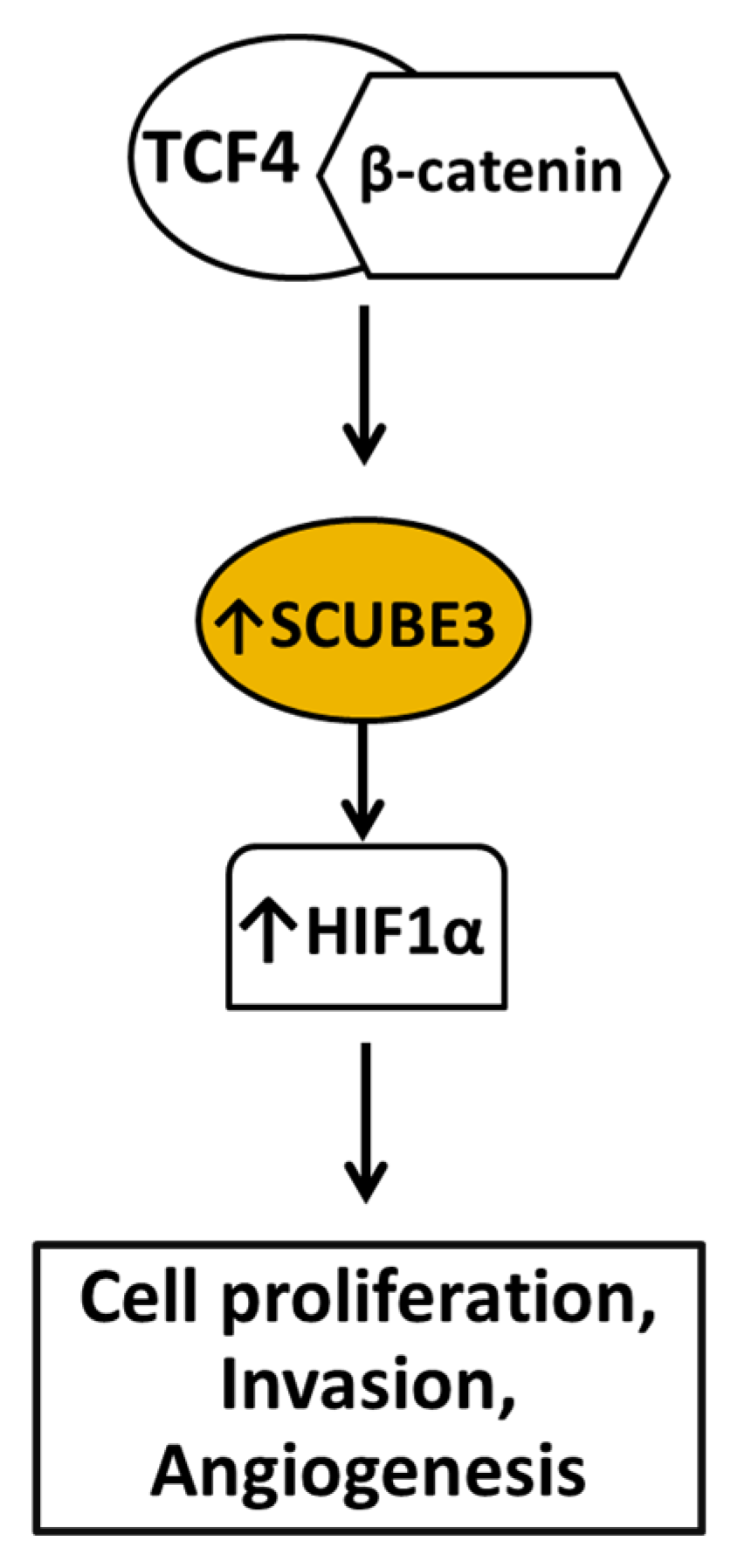
Ovarian cancer is the fifth cause of death due to cancer in women and the deadliest gynecologic cancer in the United States [63]. An estimated 20,890 people are diagnosed with ovarian cancer yearly, and about 12,730 people die of ovarian cancer annually globally [13]. Late diagnosis is the number one leading factor affecting treatment failure because most ovarian cancer cases are diagnosed at their late stages. Ovarian cancer is classified into subtypes based on histological markers, cells of origin, molecular compositions, clinical features, and treatments. There are four histological types of ovarian cancer: serous carcinoma, endometrioid, clear-cell carcinoma, and mucinous carcinoma. Serous ovarian cancer is the most common subtype of ovarian cancer based on histological classification and accounts for about 75% of all ovarian cancer cases [64]. SCUBE3 expression was elevated in ovarian cancer, and its overexpression is linked to poor prognosis [65]. The knockdown of SCUBE3 suppressed ovarian cancer cell growth. Also, in a preclinical study, the knockdown of SCUBE3 inhibited tumor growth in a xenograft mouse model [65]. Additional investigation demonstrated that SCUBE3 promotion of cell proliferation is mediated via hypoxia-inducible factor-1α (HIF1α). HIF1α is a transcription factor vital for synthesizing genes essential for enabling cells to adapt to hypoxic conditions, and it is overexpressed in tumors [66]. Accumulating evidence has linked HIF1α to cancer growth and metastasis because of its role in initiating angiogenesis [66]. Recent evidence demonstrated that the TCF4/β-catenin transcription factor complex regulates the expression of SCUBE3 at the transcriptional level (Figure 6). The expressions of SCUBE3 and HIF1α are directly correlated in ovarian cancer cells. The level of HIF1α protein was significantly decreased in SCUBE3 knockdown cells. Equally, overexpression of SCUBE3 resulted in increased HIF1α in the cells, showing a direct correlation in their expressions. This suggests that SCUBE3 is a regulator of HIF1α expression in ovarian cancer. Altogether, the evidence demonstrated the significant role of SCUBE3 in promoting ovarian cancer and showed that it is a potential therapeutic target in ovarian cancer.
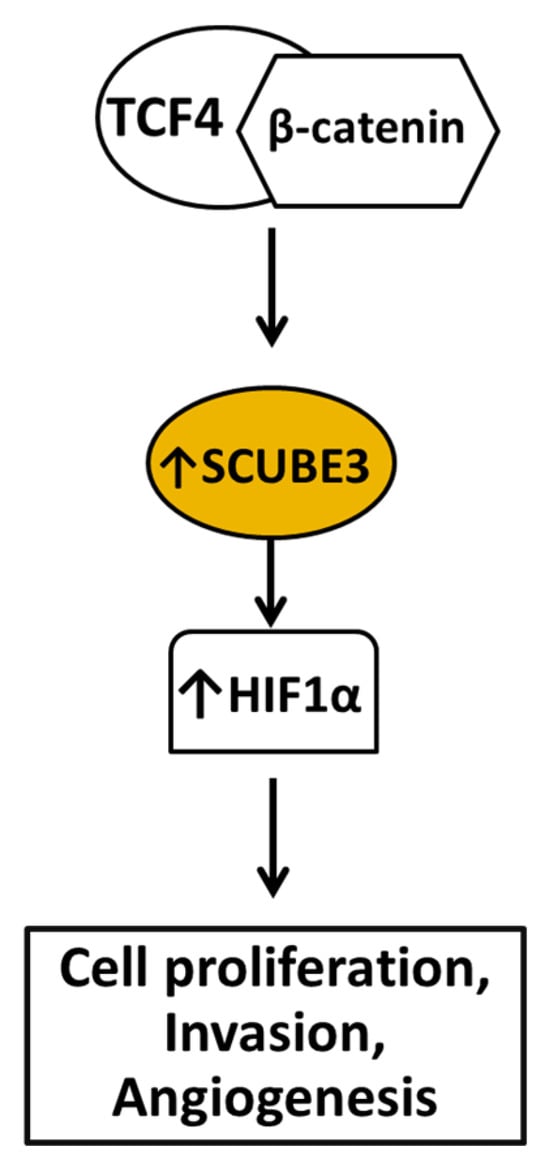
Figure 6.
Diagram depicting steps leading to SCUBE3 overexpression and its roles in promoting ovarian cancer. The TCF4/β-catenin transcription complex plays a significant role in the expression of SCUBE3. Following its elevation, SCUBE3 promotes increased HIF1α expression to regulate cell proliferation, invasion, and angiogenesis.
2.7. Glioma
Glioma is a rare, aggressive neoplasm affecting the glial tissues of the brain. Although it accounts for approximately 80% of the malignant brain tumor, it remains one of the least understood human cancers and has poor treatment outcomes [67,68]. The survival rate among patients with glioma is low, with a median survival of about 14 months [69]. The three main subtypes of glioma are astrocytoma, oligodendroglioma, and glioblastoma. Amongst these forms, glioblastoma is the most malignant subtype. Despite the standard treatment strategy of surgical resection followed by radiotherapy and chemotherapy, the overall patients’ survival has not improved [70,71]. Interestingly, SCUBE3 expression is elevated in Glioma cells compared to normal brain tissue [72].
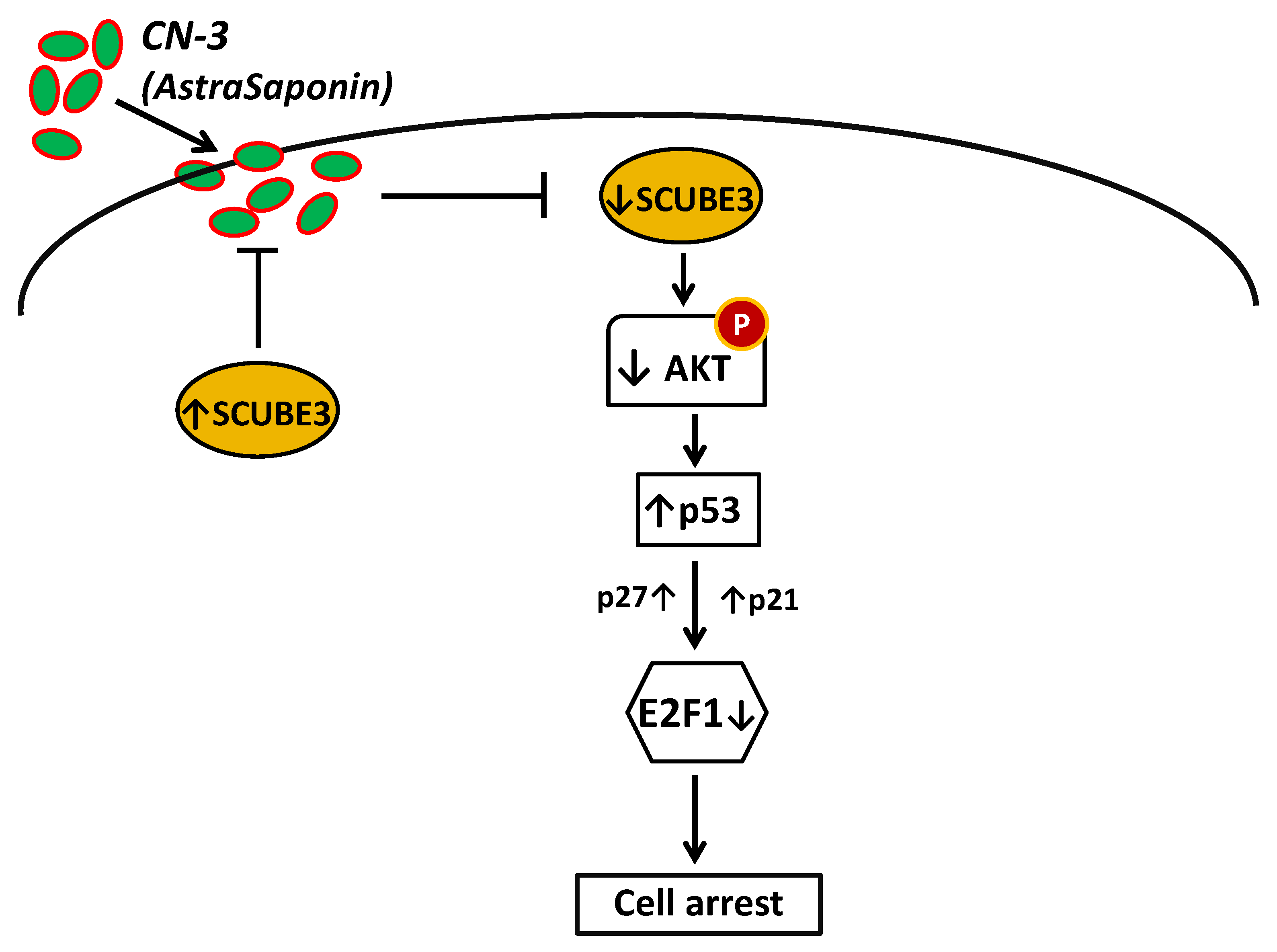
Ectopic SCUBE3 promotes cell proliferation by activating the AKT signaling pathway (Figure 7) [72]. The AKT signaling pathway is an essential pathway that promotes drug resistance in various types of cancers, and its constitutive activation promotes poor treatment outcomes in patients [73,74]. Furthermore, the data analysis showed the elevation of SCUBE3 protein levels in glioma specimens compared to normal brain tissue specimens. The U251 and U87 cells were treated with CN-3, a derivative of asterosaponin belonging to a steroid oligoglycosides class, isolated from the starfish Culcita novaeguineae. The CN-3 treatment at IC50 significantly suppressed U251 and U87 glioma cell growth and suppressed phosphorylated AKT, which indicated inhibition of AKT signaling. At IC40, CN-3 treatment significantly reduced SCUBE3 RNA level compared to eight other genes and control. The CN-3 treatment at IC40 also significantly suppressed the cell viability of U251 and U87 cells between 24 h to 72 h. Knockdown of SCUBE3 significantly inhibited cell proliferation of the glioma cells. SCUBE3 knockdown significantly suppressed AKT phosphorylation and increased p53, p21, and p27 levels, known as the mediators of cell cycle arrest (Figure 7). However, overexpression of SCUBE3 restores AKT signaling and cell proliferation. Also, the overexpression of SCUBE3 significantly damaged the effect of CN-3 treatment on the glioma cells. The study demonstrated that SCUBE3 plays a crucial role in promoting AKT signaling and suppressing the cell cycle mediators, leading to the proliferation of glioma cells. It also showed that treatment strategies targeting the SCUBE3 expression could improve patient treatment outcomes. Finally, it demonstrated that CN-3 is a promising inhibitor of the SCUBE3 oncogenic function in glioma.
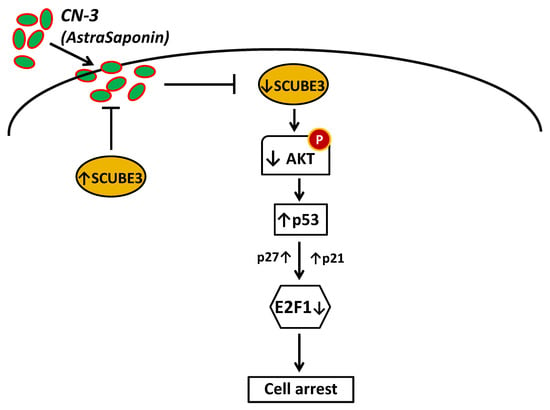
Figure 7.
Schematic representation of SCUBE3 role and the effects of CN-3 treatment in glioma cells. CN-3 inhibited SCUBE3/AKT signaling and induced cell arrest in glioma cells. SCUBE3 protein regulates AKT to mediate cell proliferation, survival, and CN-3 resistance. The treatment with CN-3, which interacts with SCUBE3 protein, significantly decreases SCUBE3 level, suppresses AKT phosphorylation, and subsequently induces cell arrest.
2.8. Renal Cell Carcinoma
A genome-wide methylation analysis study identified SCUBE3, among other genes, as epigenetically inactivated in renal cell carcinoma (RCC) [75]. RCC is the most common form of cancer affecting the kidney, accounting for about 85% of kidney tumors [76,77,78]. The data from the study found hypermethylation of SCUBE3’s promoter region in both RCC tumor specimens and cell lines [75]. The study utilized methylated DNA immunoprecipitation (MeDIP) and whole-genome array analysis techniques in combination with high-density expression array analysis in the investigation. Hypermethylation of CpG islands at the promoter regions is associated with gene silencing by multiple mechanisms involving chromatin modification and inhibition of the binding of the transcription machinery on the promoter. The loss of expression analysis using RNAi knockdown demonstrated that SCUBE3 knockdown promotes growth [75]. Additionally, Kaplan–Meier analyses revealed that the methylation of SCUBE3 was associated with a significantly increased risk of death and cancer death or relapse. The results demonstrated that SCUBE3 could function as a tumor suppressor in RCC. It also shows that SCUBE3 may have varying tissue-specific functions.
3. SCUBE3 in Other Diseases
3.1. Cardiovascular Disease
Cardiovascular disease is the prevalent cause of death in the US and accounts for approximately one-third of all causes of death [79]. SCUBE3 has been identified as a gene that is involved in cardiovascular diseases [80]. The overexpression of SCUBE3 promoted pathological cardiac hypertrophy in transgenic mice [80]. Cardiac hypertrophy is the abnormal enlargement or thickening of the heart, which makes it hard for the heart to pump blood. Evidence has shown that SCUBE3 alterations have no developmental defect [3,80,81]. SCUBE3 transgenic mice show no skeletal structure defects, but some abnormalities in bone morphology may arise when subjected to a pathological state such as stress. SCUBE3 was selectively expressed in the ventricular myocardium and may be necessary for maintaining myocyte integrity/growth or the compensatory response to myocardial stress. Observation with echocardiography and histopathology both showed cardiac hypertrophy in transgenic mice as they aged and also at eight months. The left-ventricular hypertrophy occurred more rapidly under pressure overload in the mice compared with age-matched wild-type littermates. Taken together, SCUBE3 might represent a new target for cardiac hypertrophy.
3.2. Systemic Lupus Erythematosus (SLE)
The SCUBE3 variant rs1888822 has been linked to systemic lupus erythematosus, the most common form of lupus [82]. The goal of the study was to determine the role of single nucleotide polymorphisms (SNPs) in SCUBE3 with systemic lupus erythematosus (SLE). SNP is a genomic variant at a single base position in the DNA of a gene or regulatory region. The change in the gene code may cause SNPs to alter the role of the gene in a disease. SLE is an autoimmune disease characterized by inflammation of the connective tissues and affects many organs [82,83,84]. Environmental and genetic factors have been linked as factors that cause SLE. The study has found that individuals with the rs1888822 SCUBE3 variant are predisposed to SLE. Further analysis showed that SCUBE3 mRNA expressions were significantly reduced in SLE patient’s whole blood, CD3 T-cells, and B cells compared with healthy controls [82]. However, more investigations are required to determine if the decreased SCUBE3 expression is linked with the disease condition. The study demonstrated that the alteration or/and expression of normal SCUBE3 could be associated with SLE.
3.3. Other Immune Disorder: Psoriasis and Rheumatoid Arthritis (RA)
SCUBE3 is implicated in both psoriasis and rheumatoid arthritis (RA). The SCUBE3 protein level was significantly higher in serum samples of patients with psoriasis than in the control [85]. Psoriasis is a chronic inflammatory disease that affects mainly the skin and joints. It is characterized by excessive growth and aberrant keratinocyte differentiation [86]. SCUBE3 level was also elevated in RA [87]. These findings suggest that SCUBE3 may play a significant role in the immune system and synovial angiogenesis and is an ideal target for treating psoriasis and RA.
3.4. Alopecia
A recently published study demonstrated that SCUBE3 is a mesenchymal niche factor that activates hair growth [88]. The hair dermal papilla cells are specialized mesenchymal fibroblasts found at the base of hair follicles [89]. They secrete extracellular molecules, for example, collagen, and are crucial for hair formation, growth, and cycling. The dermal papilla is connected to the dermal sheath, which surrounds the outside of the dermal papilla [90]. The dermal sheath contains progenitor cells that maintain and regenerate the dermal papilla. Together, the dermal papilla and sheath are crucial components for hair growth [91]. The SCUBE3 expression was broadly observed in the dermal papillae and some subsets of dermal sheath cells during the anagen phase, which is the hair growth phase and is mostly absent in the telogen phase, where the follicle is dormant [88]. Scattered expression of SCUBE3 was observed in the hair matrix, mainly during anagen. Microinjection of recombinant human SCUBE3 protein induced significant hair growth in mice back skin compared to bovine serum albumin control.
Aging mice experience miniaturization of hair follicles and elevated extra-follicular hair-growth inhibitor levels [88]. Microinjection of recombinant human SCUBE3 protein in aged 20-month-old skin mice stimulated hair growth. Further analysis showed that SCUBE3-deleted mice display mild anagen initiation delay, suggesting the importance of SCUBE3 in stimulating anagen effects. Recombinant human SCUBE3 protein treatment induced hair growth on the mouse xenograft. Additional evidence showed that the hair-stimulating effect is exerted mainly by exogenous SCUBE3 rather than endogenous SCUBE3 protein and via the TGF-β signaling [88]. All of these demonstrated that SCUBE3 is a hair-growth activator and further explained the importance of the TGF-β signaling in mediating SCUBE3-induced growth.
3.5. Fast Muscle Development
In mice, SCUBE3 is broadly expressed in various developing tissues such as somites, neural tubes, and limb buds [3]. Evidence shows that SCUBE3 is an essential regulator of fibroblast signaling, which plays a significant role in fast muscle development [92]. The knockdown of SCUBE3 in C2C12 myoblasts suppressed FGF receptor 4 (FGFR4) expression and fibroblast growth factor (FGF) signaling, thus resulting in poor myogenic differentiation. Also, the knockdown of SCUBE3 in zebra fish attenuated the expression of myogenic marker myod1 within the lateral fast muscle precursors. In contrast, its overexpression in the adaxial slow muscle precursors was mainly unaffected [92]. The data also demonstrated that SCUBE3 could act as an FGF co-receptor to boost FGF8 signaling during zebrafish embryogenesis. Overall, the study showed that SCUBE3 is an essential player in fast fiber development through the regulation of FGF signaling and can act alone as a multifunctional protein or together with other SCUBE family members in a context-dependent manner.
4. Clinical Utility Focusing on SCUBE3 and Future Expectations
The roles of SCUBE3 in the proliferation and progression of cancer and other diseases have been identified in many studies. The key characteristics of SCUBE3 in different physiological conditions are summaries in Table 1 and Table 2. Increasing evidence showed that SCUBE3 was involved in the growth and promotion of malignant phenotypes in lung cancer. All the reports on SCUBE3 underscored the interaction between SCUBE3 and TGFβRII in the extracellular as the leading promoter of cancer and the upregulation of TGF-β, which also activates the TGFβRII signaling to promote metastasis and invasion. Upon its release from the cytoplasm, SCUBE3 and TGF-β function as ligands, acting in an autocrine and paracrine manner by binding and activating TGFβRII signaling. Activating this signaling thus promotes tumor cell growth, metastasis, and invasion.

Table 1.
SCUBE3 expression and function in cancer.

Table 2.
SCUBE3 is associated with various other diseases.
Due to its expression pattern, SCUBE3 could serve as a biomarker for early detection of cancer and monitoring treatment outcomes. It could also serve as a biomarker for detection of cardiac stress in patients and also for other patients that are at risk of heart attack. A reliable circulating protein biomarker can be detected by inexpensive and noninvasive blood tests that can be used for early diagnosis and monitoring throughout the disease’s treatment and progression. As we have shown, SCUBE3 expression is significantly higher in cancer tissues than in normal tissue and is likely to lead to an increased level of secreted SCUBE3. That could be used as a predictive or prognostic biomarker in cancer. The glycosylation of secreted SCUBE3 protein makes it reasonably stable for clinical analysis and thus makes it a good candidate for biomarker. We have seen the success of other secreted biomarkers detected in blood tests, including prostate-specific antigen (PSA) for prostate cancer, CA-125 for ovarian cancer, and carcinoembryonic antigen (CEA) for colorectal cancers [93,94,95,96,97]. Other secreted proteins are being evaluated for their ability to serve as biomarkers including TGF-β and its’ signaling network.
Recent findings demonstrated that the aberrant SCUBE3 expression is associated with systemic lupus erythematosus (SLE) [82]. SLE is a complex auto-immune disorder caused when the immune system attacks its own tissue, resulting in tissue damage and inflammation. Low expression of SCUBE3 mRNA was observed in Systemic lupus erythematosus (SLE). In CD3+ T cells and B cells of patients with SLE, the SCUBE3 level was also decreased. In the embryonic mouse, SCUBE3 is expressed in the skin dermal papillae of developed hair follicles [98]. This report widened the involvement of SCUBE3 in human diseases such as immune disorder. However, the mechanism by which loss of SCUBE3 alters or contributes to the immune system is unknown.
Aging in mice is associated with shrinking of hair follicular because of increase expression of hair-growth inhibitors [99]. SCID mouse xenograft human occipital scalp hair follicles treated with recombinant human SCUBE3 protein increases hair growth, thus suggesting that SCUBE3 is a mesenchymal niche-derived hair-growth activator in mice, whose expression pattern and function are at least partially conserved in human scalp [88]. The SCUBE3/TGF-β signaling mediates the regulation of mesenchymal niche function in the hair follicle through Hedgehog. In general, because of its high expression, secretion into the extracellular matrix and mechanisms of action, extracellular SCUBE3 is a promising target for future strategies to treat cancer. This report has also validated the critical role of SCUBE3 in growth and demonstrated that SCUBE3 promotion of growth is extended to the reprogramming of the stem cell niche in hair follicles. There are still issues to resolve regarding extracellular SCUBE3. There are possibilities that extracellular SCUBE3 can act through receptors other than the TGFβRII to promote its functions in cancers because the composition of three cysteine-rich motifs amino acid residues indicates the possibility of proteolytic cleavage. In fast fiber myogenesis in zebra fish, SCUBE3 acted as a co-receptor for fibroblast growth factor 8 (FGF 8) signaling during embryogenesis [92]. The multifunctional character of SCUBE3, functioning intra and extracellularly, should not impede its suitability as a therapeutic target because SCUBE3 inside and outside the cells promotes cell growth, metastasis, and invasion.
Hypothetically, the up-regulation in intracellular SCUBE3 protein in cancer cells could lead to increased secretion of SCUBE3 into the extracellular space. To better understand these mechanisms, we need to know how SCUBE3 is released, whether SCUBE3 interacts with other receptors, and the manner of the interactions. It is also important to determine if SCUBE3 may be acting as a Damage-Associated Molecular Patterns (DAMPs) protein to indicate the presence of cancer cells in the tumor microenvironment. DAMP proteins are released in response to stress, apoptosis, necrosis, and acute DNA damage, and during response to cytotoxic anti-tumor treatment, their release may trigger inflammation [100]. Cells communicate directly and indirectly about their physiological states to other cells using chemical or mechanical signals [101]. There may be tumor-specific DAMPs that are upregulated in cancers to signal the pathophysiological states of the cells. Understanding these proteins can help in the diagnosis and treatment of cancer patients.
5. Concluding Remarks and Future Directions
There are several potential research focus areas for SCUBE3, such as the evaluation of secreted SCUBE3 in the bodily fluid as a marker in cancer and predictive responses and delineation of the functional overlaps and differences for extracellular SCUBE3 in tumorigenesis, neo-angiogenesis, and fibrotic diseases could help shed more information on the functions and mechanisms of SCUBE3. Furthermore, identifying and characterizing selective SCUBE3 inhibitors for therapy in pre-clinical and clinical settings could provide more information on the clinical value of inhibiting SCUBE3 protein in cancer. The role of SCUBE3 as a hair-growth activator highlights the importance of understanding SCUBE3’s functions and mechanisms in various conditions, including cardiac hypertrophy, systemic lupus erythematosus (SLE), rheumatoid arthritis, and psoriasis.
Targeted treatment of SCUBE3 for cancer may encounter several limitations due to its widespread expression and diverse roles in different tissues. SCUBE3 can have varying effects in cancer cells; for instance, it acts as a tumor suppressor in renal cell carcinoma (RCC), while serving as an oncogene in other cancers, such as lung and breast cancer. This also shows that SCUBE3 can be regulated through methylation. Identifying effective drugs to block SCUBE3 without harming normal cells is challenging because many compounds lack specificity or require high, potentially toxic concentrations. Additionally, using an antibody or small molecule inhibitor to target SCUBE3 may yield limited success, as research indicates that SCUBE3 has intracellular functions that promote cell proliferation. SCUBE3 might also function extracellularly by binding to TGFBRII and activating associated signaling pathways. Antibody treatment is good if the protein is secreted or expressed on the cell membrane. Cancer cells can increase SCUBE3 to bypass other targeted therapies or become resistant to antiestrogens, indicating SCUBE3 can be a resistance driver.
Quantifying SCUBE3 in small clinical samples poses challenges due to issues related to mRNA and protein stability, as well as variability in protein abundance. Therefore, understanding these factors is crucial for identifying the potential benefits and challenges of targeting SCUBE3 in medical treatments. A key focus should be on designing and developing high-affinity inhibitors that disrupt SCUBE3 interactions with target proteins, such as TGFBRII. Additionally, efforts should be directed towards quantifying SCUBE3 levels in tissue samples and patients’ bodily fluids to establish baseline levels in healthy individuals and compare them with cancer patients.
6. Conclusions
In this review, we have summarized the current knowledge on SCUBE3 with the addition of our recent observations of SCUBE3 expression in human cancer to demonstrate how SCUBE3 influences the pathogenesis of several cancers. Nearly all the reports have shown that SCUBE3 is a secreted protein with elevated expression in several cancers. The involvements of SCUBE3 in growth, metastasis, and invasion were also demonstrated in several reports. The TGFβRII is a critical player in mediating SCUBE3 functions. Future investigation could help determine if SCUBE3 is a diagnostic or prognostic biomarker in cancer and other diseases. In addition, the identification of selective SCUBE3 inhibitors could help determine whether targeting SCUBE3 is a promising strategy in cancer treatment. The role of SCUBE3 in cancer must be clear so that it gains significant attention as an essential protein that promotes cancers and can be utilized as a diagnostic or prognostic biomarker and a promising therapeutic target. Since its role spans beyond cancer, crossing to other areas such as cardiac hypertrophy, bone development, inflammation, and hair growth, research on SCUBE3 will thus continue to be essential and exciting in life sciences.
Author Contributions
Conceptualization: A.I., B.C.O., T.J.S. and D.C.; Methodology: B.C.O., T.J.S., D.C. and P.S.; Formal Analysis: A.I., B.C.O., D.C., C.S.-W., T.J.S., S.J., K.A., A.A. and P.S.; Resource: A.I., B.C.O., D.C., C.S.-W., T.J.S., S.J., K.A., A.A. and P.S.; Data curator: B.C.O. and T.J.S.; Writing-Original Draft Preparation: B.C.O. and T.J.S.; Writing-Review and Editing: A.I., B.C.O., D.C., C.S.-W., T.J.S., S.J., K.A., A.A. and P.S.; Visualization: A.I., B.C.O., D.C., C.S.-W., T.J.S., S.J., K.A., A.A. and P.S.; Supervision: B.C.O., D.C., T.J.S. and P.S.; Project Administration: B.C.O. and T.J.S.; Funding acquisition: B.C.O. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by MS-INBRE (NIH/MS-INBRE P20GM103476).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data supporting the finding of this study are available from the corresponding author upon request.
Acknowledgments
The authors express gratitude to Ugoeze I. Onyeagucha and Oluchukwu I. Onyeagucha for their help in creating the figures.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Lin, Y.C.; Sahoo, B.K.; Gau, S.S.; Yang, R.B. The biology of SCUBE. J. Biomed. Sci. 2023, 30, 33. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.T.; Su, Y.H.; Tsai, M.T.; Wasserman, S.M.; Topper, J.N.; Yang, R.B. A novel secreted, cell-surface glycoprotein containing multiple epidermal growth factor-like repeats and one CUB domain is highly expressed in primary osteoblasts and bones. J. Biol. Chem. 2004, 279, 37485–37490. [Google Scholar] [CrossRef] [PubMed]
- Haworth, K.; Smith, F.; Zoupa, M.; Seppala, M.; Sharpe, P.T.; Cobourne, M.T. Expression of the Scube3 epidermal growth factor-related gene during early embryonic development in the mouse. Gene. Expr. Patterns 2007, 7, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Xavier, G.M.; Economou, A.; Senna Guimaraes, A.L.; Sharpe, P.T.; Cobourne, M.T. Characterization of a mouse Scube3 reporter line. Genesis 2010, 48, 684–692. [Google Scholar] [CrossRef]
- Lin, Y.C.; Niceta, M.; Muto, V.; Vona, B.; Pagnamenta, A.T.; Maroofian, R.; Beetz, C.; van Duyvenvoorde, H.; Dentici, M.L.; Lauffer, P.; et al. SCUBE3 loss-of-function causes a recognizable recessive developmental disorder due to defective bone morphogenetic protein signaling. Am. J. Hum. Genet. 2021, 108, 115–133. [Google Scholar] [CrossRef]
- Kobayashi, T.; Lyons, K.M.; McMahon, A.P.; Kronenberg, H.M. BMP signaling stimulates cellular differentiation at multiple steps during cartilage development. Proc. Natl. Acad. Sci. USA 2005, 102, 18023–18027. [Google Scholar] [CrossRef]
- Tsai, M.T.; Cheng, C.J.; Lin, Y.C.; Chen, C.C.; Wu, A.R.; Wu, M.T.; Hsu, C.C.; Yang, R.B. Isolation and characterization of a secreted, cell-surface glycoprotein SCUBE2 from humans. Biochem. J. 2009, 422, 119–128. [Google Scholar] [CrossRef]
- Lin, Y.C.; Chao, T.Y.; Yeh, C.T.; Roffler, S.R.; Kannagi, R.; Yang, R.B. Endothelial SCUBE2 Interacts With VEGFR2 and Regulates VEGF-Induced Angiogenesis. Arterioscler. Thromb. Vasc. Biol. 2017, 37, 144–155. [Google Scholar] [CrossRef]
- Wu, Y.Y.; Peck, K.; Chang, Y.L.; Pan, S.H.; Cheng, Y.F.; Lin, J.C.; Yang, R.B.; Hong, T.M.; Yang, P.C. SCUBE3 is an endogenous TGF-beta receptor ligand and regulates the epithelial-mesenchymal transition in lung cancer. Oncogene 2011, 30, 3682–3693. [Google Scholar] [CrossRef]
- Zhao, C.; Qin, Q.; Wang, Q.; Zhang, J.; Xu, Y.; Li, W.; Gu, M.; Chen, S.; Deng, A. SCUBE3 overexpression predicts poor prognosis in non-small cell lung cancer. Biosci. Trends 2013, 7, 264–269. [Google Scholar]
- Huo, Q.; He, X.; Li, Z.; Yang, F.; He, S.; Shao, L.; Hu, Y.; Chen, S.; Xie, N. SCUBE3 serves as an independent poor prognostic factor in breast cancer. Cancer Cell Int. 2021, 21, 268. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Luo, A.; Xiong, C.; Ren, H.; Yan, L.; Luo, Q. SCUBE3 downregulation modulates hepatocellular carcinoma by inhibiting CCNE1 via TGFbeta/PI3K/AKT/GSK3beta pathway. Cancer Cell Int. 2022, 22, 1. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Islami, F.; Goding Sauer, A.; Miller, K.D.; Siegel, R.L.; Fedewa, S.A.; Jacobs, E.J.; McCullough, M.L.; Patel, A.V.; Ma, J.; Soerjomataram, I.; et al. Proportion and number of cancer cases and deaths attributable to potentially modifiable risk factors in the United States. CA Cancer J. Clin. 2018, 68, 31–54. [Google Scholar] [CrossRef]
- Field, R.W. Radon: A Leading Environmental Cause of Lung Cancer. Am. Fam. Physician 2018, 98, 280–282. [Google Scholar]
- Liu, Y.; Xu, Y.; Xu, W.; He, Z.; Fu, C.; Du, F. Radon and lung cancer: Current status and future prospects. Crit. Rev. Oncol. Hematol. 2024, 198, 104363. [Google Scholar] [CrossRef]
- Molina, J.R.; Yang, P.; Cassivi, S.D.; Schild, S.E.; Adjei, A.A. Non-small cell lung cancer: Epidemiology, risk factors, treatment, and survivorship. Mayo Clin. Proc. 2008, 83, 584–594. [Google Scholar] [CrossRef]
- Sher, T.; Dy, G.K.; Adjei, A.A. Small cell lung cancer. Mayo Clin. Proc. 2008, 83, 355–367. [Google Scholar] [CrossRef]
- Chou, C.H.; Cheng, Y.F.; Siow, T.Y.; Kumar, A.; Peck, K.; Chang, C. SCUBE3 regulation of early lung cancer angiogenesis and metastatic progression. Clin. Exp. Metastasis 2013, 30, 741–752. [Google Scholar] [CrossRef]
- Simms, J.A.; Pearson, D.D.; Cholowsky, N.L.; Irvine, J.L.; Nielsen, M.E.; Jacques, W.R.; Taron, J.M.; Peters, C.E.; Carlson, L.E.; Goodarzi, A.A. Younger North Americans are exposed to more radon gas due to occupancy biases within the residential built environment. Sci. Rep. 2021, 11, 6724. [Google Scholar] [CrossRef]
- Mirabello, L.; Troisi, R.J.; Savage, S.A. Osteosarcoma incidence and survival rates from 1973 to 2004: Data from the Surveillance, Epidemiology, and End Results Program. Cancer 2009, 115, 1531–1543. [Google Scholar] [CrossRef]
- Khandekar, S.; Dive, A.; Munde, P.; Fande, P.Z. Chondroblastic osteosarcoma of the left zygomatic bone: Rare case report and review of the literature. J. Oral. Maxillofac. Pathol. 2014, 18, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Yang, C.; Peng, J.; Qian, Y.; Wang, Z. The Expression of HSPD1, SCUBE3, CXCL14 and Its Relations with the Prognosis in Osteosarcoma. Cell Biochem. Biophys. 2015, 73, 763–768. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.H.; Shu, C.W.; Chao, J.K.; Lee, C.H.; Fu, T.Y.; Liou, H.H.; Ger, L.P.; Liu, P.F. HSPD1 repressed E-cadherin expression to promote cell invasion and migration for poor prognosis in oral squamous cell carcinoma. Sci. Rep. 2019, 9, 8932. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Yu, J.; Chen, M.; Lu, Q.; Ning, F.; Gan, X.; Liu, H.; Ye, Y.; Lu, S.; Lash, G.E. Knockdown of heat shock protein family D member 1 (HSPD1) promotes proliferation and migration of ovarian cancer cells via disrupting the stability of mitochondrial 3-oxoacyl-ACP synthase (OXSM). J. Ovarian Res. 2023, 16, 81. [Google Scholar] [CrossRef]
- Pan, Q.; Liu, R.; Zhang, X.; Cai, L.; Li, Y.; Dong, P.; Gao, J.; Liu, Y.; He, L. CXCL14 as a potential marker for immunotherapy response prediction in renal cell carcinoma. Ther. Adv. Med. Oncol. 2023, 15, 17588359231217966. [Google Scholar] [CrossRef]
- Schwarze, S.R.; Luo, J.; Isaacs, W.B.; Jarrard, D.F. Modulation of CXCL14 (BRAK) expression in prostate cancer. Prostate 2005, 64, 67–74. [Google Scholar] [CrossRef]
- Gibbs, C.; So, J.Y.; Ahad, A.; Michalowski, A.M.; Son, D.S.; Li, Y. CXCL14 Attenuates Triple-Negative Breast Cancer Progression by Regulating Immune Profiles of the Tumor Microenvironment in a T Cell-Dependent Manner. Int. J. Mol. Sci. 2022, 23, 9314. [Google Scholar] [CrossRef]
- Song, G.; Wang, J.; Lu, J.; Xu, H.; Zhao, Z.; Tang, Q.; Zou, C.; Yin, J.; Xie, X.; Shen, J. Role of SCUBE3 in promoting osteosarcoma cell growth and its association with prognosis. Nan Fang Yi Ke Da Xue Xue Bao 2014, 34, 617–621. [Google Scholar]
- Chen, H.; Wu, X.; Lan, Y.; Zhou, X.; Zhang, Y.; Long, L.; Zhong, Y.; Hao, Z.; Zhang, W.; Xue, D. SCUBE3 promotes osteogenic differentiation and mitophagy in human bone marrow mesenchymal stem cells through the BMP2/TGF-beta signaling pathway. FASEB J. 2024, 38, e70011. [Google Scholar] [CrossRef]
- Villanueva, A. Hepatocellular Carcinoma. N. Engl. J. Med. 2019, 380, 1450–1462. [Google Scholar] [CrossRef]
- Bruix, J.; Takayama, T.; Mazzaferro, V.; Chau, G.Y.; Yang, J.; Kudo, M.; Cai, J.; Poon, R.T.; Han, K.H.; Tak, W.Y.; et al. Adjuvant sorafenib for hepatocellular carcinoma after resection or ablation (STORM): A phase 3, randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2015, 16, 1344–1354. [Google Scholar] [CrossRef]
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Pineros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2019, 144, 1941–1953. [Google Scholar] [CrossRef] [PubMed]
- Balogh, J.; Victor, D., 3rd; Asham, E.H.; Burroughs, S.G.; Boktour, M.; Saharia, A.; Li, X.; Ghobrial, R.M.; Monsour, H.P., Jr. Hepatocellular carcinoma: A review. J. Hepatocell. Carcinoma 2016, 3, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Swamy, S.G.; Kameshwar, V.H.; Shubha, P.B.; Looi, C.Y.; Shanmugam, M.K.; Arfuso, F.; Dharmarajan, A.; Sethi, G.; Shivananju, N.S.; Bishayee, A. Targeting multiple oncogenic pathways for the treatment of hepatocellular carcinoma. Target. Oncol. 2017, 12, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Alqahtani, A.; Khan, Z.; Alloghbi, A.; Said Ahmed, T.S.; Ashraf, M.; Hammouda, D.M. Hepatocellular Carcinoma: Molecular Mechanisms and Targeted Therapies. Medicina 2019, 55, 526. [Google Scholar] [CrossRef]
- Margolis, B.; Dao, F.; Licciardi, M.; Misirlioglu, S.; Olvera, N.; Ramaswami, S.; Levine, D.A. CCNE1 amplification among metastatic sites in patients with gynecologic high-grade serous carcinoma. Gynecol. Oncol. Rep. 2021, 37, 100850. [Google Scholar] [CrossRef]
- Ayhan, A.; Kuhn, E.; Wu, R.C.; Ogawa, H.; Bahadirli-Talbott, A.; Mao, T.L.; Sugimura, H.; Shih, I.M.; Wang, T.L. CCNE1 copy-number gain and overexpression identify ovarian clear cell carcinoma with a poor prognosis. Mod. Pathol. 2017, 30, 297–303. [Google Scholar] [CrossRef]
- Guerrero-Zotano, A.; Belli, S.; Zielinski, C.; Gil-Gil, M.; Fernandez-Serra, A.; Ruiz-Borrego, M.; Ciruelos Gil, E.M.; Pascual, J.; Munoz-Mateu, M.; Bermejo, B.; et al. CCNE1 and PLK1 Mediate Resistance to Palbociclib in HR+/HER2- Metastatic Breast Cancer. Clin. Cancer Res. 2023, 29, 1557–1568. [Google Scholar] [CrossRef]
- Turner, N.C.; Liu, Y.; Zhu, Z.; Loi, S.; Colleoni, M.; Loibl, S.; DeMichele, A.; Harbeck, N.; Andre, F.; Bayar, M.A.; et al. Cyclin E1 Expression and Palbociclib Efficacy in Previously Treated Hormone Receptor-Positive Metastatic Breast Cancer. J. Clin. Oncol. 2019, 37, 1169–1178. [Google Scholar] [CrossRef]
- Ma, G.; Yang, L.; Dong, J.; Zhang, L. CCNE1 Promotes Progression and is Associated with Poor Prognosis in Lung Adenocarcinoma. Curr. Pharm. Biotechnol. 2022, 23, 1168–1178. [Google Scholar] [CrossRef] [PubMed]
- Fresno Vara, J.A.; Casado, E.; de Castro, J.; Cejas, P.; Belda-Iniesta, C.; Gonzalez-Baron, M. PI3K/Akt signalling pathway and cancer. Cancer Treat. Rev. 2004, 30, 193–204. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Zhang, X.; Wang, P.; Zhang, Y.; He, S.; Li, Y.; Li, S.; Liang, K.; Liao, S.; Gao, Y.; et al. Low concentration triphenyl phosphate fuels proliferation and migration of hepatocellular carcinoma cells. Environ. Toxicol. 2022, 37, 2445–2459. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Fong, K.O.; Cheung, M.P.L.; Liu, J.A.; Liang, R.; Li, T.W.; Sharma, R.; Ip, P.P.; Yang, X.; Cheung, M. DEPDC1B Promotes Melanoma Angiogenesis and Metastasis through Sequestration of Ubiquitin Ligase CDC16 to Stabilize Secreted SCUBE3. Adv. Sci. 2022, 9, e2105226. [Google Scholar] [CrossRef]
- Saginala, K.; Barsouk, A.; Aluru, J.S.; Rawla, P. Epidemiology of Melanoma. Med. Sci. 2021, 9, 63. [Google Scholar] [CrossRef]
- Castellani, G.; Buccarelli, M.; Arasi, M.B.; Rossi, S.; Pisanu, M.E.; Bellenghi, M.; Lintas, C.; Tabolacci, C. BRAF Mutations in Melanoma: Biological Aspects, Therapeutic Implications, and Circulating Biomarkers. Cancers 2023, 15, 4026. [Google Scholar] [CrossRef]
- Kreuger, I.Z.M.; Slieker, R.C.; van Groningen, T.; van Doorn, R. Therapeutic Strategies for Targeting CDKN2A Loss in Melanoma. J. Investig. Dermatol. 2023, 143, 18–25 e11. [Google Scholar] [CrossRef]
- Wan, P.T.; Garnett, M.J.; Roe, S.M.; Lee, S.; Niculescu-Duvaz, D.; Good, V.M.; Jones, C.M.; Marshall, C.J.; Springer, C.J.; Barford, D.; et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 2004, 116, 855–867. [Google Scholar] [CrossRef]
- Richtig, G.; Hoeller, C.; Kashofer, K.; Aigelsreiter, A.; Heinemann, A.; Kwong, L.N.; Pichler, M.; Richtig, E. Beyond the BRAF(V)(600E) hotspot: Biology and clinical implications of rare BRAF gene mutations in melanoma patients. Br. J. Dermatol. 2017, 177, 936–944. [Google Scholar] [CrossRef]
- Weiss, R.A. A perspective on the early days of RAS research. Cancer Metastasis Rev. 2020, 39, 1023–1028. [Google Scholar] [CrossRef]
- Hodis, E.; Watson, I.R.; Kryukov, G.V.; Arold, S.T.; Imielinski, M.; Theurillat, J.P.; Nickerson, E.; Auclair, D.; Li, L.; Place, C.; et al. A landscape of driver mutations in melanoma. Cell 2012, 150, 251–263. [Google Scholar] [CrossRef]
- Krauthammer, M.; Kong, Y.; Ha, B.H.; Evans, P.; Bacchiocchi, A.; McCusker, J.P.; Cheng, E.; Davis, M.J.; Goh, G.; Choi, M.; et al. Exome sequencing identifies recurrent somatic RAC1 mutations in melanoma. Nat. Genet. 2012, 44, 1006–1014. [Google Scholar] [CrossRef] [PubMed]
- Chappell, W.H.; Steelman, L.S.; Long, J.M.; Kempf, R.C.; Abrams, S.L.; Franklin, R.A.; Basecke, J.; Stivala, F.; Donia, M.; Fagone, P.; et al. Ras/Raf/MEK/ERK and PI3K/PTEN/Akt/mTOR inhibitors: Rationale and importance to inhibiting these pathways in human health. Oncotarget 2011, 2, 135–164. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Shi, W.; Hu, W.; Ma, D.; Yan, D.; Yu, K.; Zhang, G.; Cao, Y.; Wu, J.; Jiang, C.; et al. DEP Domain-Containing Protein 1B (DEPDC1B) Promotes Migration and Invasion in Pancreatic Cancer Through the Rac1/PAK1-LIMK1-Cofilin1 Signaling Pathway. Onco Targets Ther. 2020, 13, 1481–1496. [Google Scholar] [CrossRef]
- Cronin, J.C.; Watkins-Chow, D.E.; Incao, A.; Hasskamp, J.H.; Schonewolf, N.; Aoude, L.G.; Hayward, N.K.; Bastian, B.C.; Dummer, R.; Loftus, S.K.; et al. SOX10 ablation arrests cell cycle, induces senescence, and suppresses melanomagenesis. Cancer Res. 2013, 73, 5709–5718. [Google Scholar] [CrossRef]
- Marchesi, S.; Montani, F.; Deflorian, G.; D’Antuono, R.; Cuomo, A.; Bologna, S.; Mazzoccoli, C.; Bonaldi, T.; Di Fiore, P.P.; Nicassio, F. DEPDC1B coordinates de-adhesion events and cell-cycle progression at mitosis. Dev. Cell 2014, 31, 420–433. [Google Scholar] [CrossRef]
- Heichman, K.A.; Roberts, J.M. CDC16 controls initiation at chromosome replication origins. Mol. Cell 1998, 1, 457–463. [Google Scholar] [CrossRef]
- Yang, X.; Hu, J.; Shi, C.; Dai, J. Activation of TGF-beta1 Pathway by SCUBE3 Regulates TWIST1 Expression and Promotes Breast Cancer Progression. Cancer Biother. Radiopharm. 2020, 35, 120–128. [Google Scholar] [CrossRef]
- Khan, M.A.; Chen, H.C.; Zhang, D.; Fu, J. Twist: A molecular target in cancer therapeutics. Tumour. Biol. 2013, 34, 2497–2506. [Google Scholar] [CrossRef]
- Barnes, R.M.; Firulli, A.B. A twist of insight—The role of Twist-family bHLH factors in development. Int. J. Dev. Biol. 2009, 53, 909–924. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Zhuang, J.; Li, H.; Liu, C.; Zhou, C.; Liu, L.; Feng, F.; Sun, C.; Wu, J. Development of a risk scoring system for evaluating the prognosis of patients with Her2-positive breast cancer. Cancer Cell Int. 2020, 20, 121. [Google Scholar] [CrossRef]
- Webb, P.M.; Jordan, S.J. Epidemiology of epithelial ovarian cancer. Best. Pract. Res. Clin. Obstet. Gynaecol. 2017, 41, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Torre, L.A.; Trabert, B.; DeSantis, C.E.; Miller, K.D.; Samimi, G.; Runowicz, C.D.; Gaudet, M.M.; Jemal, A.; Siegel, R.L. Ovarian cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 284–296. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Ma, X.; Wei, Z.; Qian, Q.; Jing, A.; Ding, Y.; Geng, T.; Qin, J.; Ma, L.; Chen, Y.; et al. beta-catenin/TCF4-induced SCUBE3 upregulation promotes ovarian cancer development via HIF-1 signaling pathway. Mol. Cell. Endocrinol. 2023, 582, 112127. [Google Scholar] [CrossRef]
- Jun, J.C.; Rathore, A.; Younas, H.; Gilkes, D.; Polotsky, V.Y. Hypoxia-Inducible Factors and Cancer. Curr. Sleep Med. Rep. 2017, 3, 1–10. [Google Scholar] [CrossRef]
- Huang, Y.T.; Zhang, Y.; Wu, Z.; Michaud, D.S. Genotype-based gene signature of glioma risk. Neuro. Oncol. 2017, 19, 940–950. [Google Scholar] [CrossRef]
- Zhou, Y.; Wu, S.; Liang, C.; Lin, Y.; Zou, Y.; Li, K.; Lu, B.; Shu, M.; Huang, Y.; Zhu, W.; et al. Transcriptional upregulation of microtubule-associated protein 2 is involved in the protein kinase A-induced decrease in the invasiveness of glioma cells. Neuro. Oncol. 2015, 17, 1578–1588. [Google Scholar] [CrossRef]
- Prados, M.D.; Chang, S.M.; Butowski, N.; DeBoer, R.; Parvataneni, R.; Carliner, H.; Kabuubi, P.; Ayers-Ringler, J.; Rabbitt, J.; Page, M.; et al. Phase II study of erlotinib plus temozolomide during and after radiation therapy in patients with newly diagnosed glioblastoma multiforme or gliosarcoma. J. Clin. Oncol. 2009, 27, 579–584. [Google Scholar] [CrossRef]
- Lee, J.H.; Wee, C.W. Treatment of Adult Gliomas: A Current Update. Brain Neurorehabil. 2022, 15, e24. [Google Scholar] [CrossRef]
- Zhao, J.; Wang, L.; Hu, G.; Wei, B. A 6-Gene Risk Signature Predicts Survival of Glioblastoma Multiforme. Biomed. Res. Int. 2019, 2019, 1649423. [Google Scholar] [CrossRef] [PubMed]
- Qiu, P.C.; Lu, Y.Y.; Zhang, S.; Li, H.; Bao, H.; Ji, Y.Q.; Fang, F.; Tang, H.F.; Cheng, G. Reduction of SCUBE3 by a new marine-derived asterosaponin leads to arrest of glioma cells in G1/S. Oncogenesis 2020, 9, 71. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Chen, Y.; Liu, G.; Li, C.; Song, Y.; Cao, Z.; Li, W.; Hu, J.; Lu, C.; Liu, Y. PI3K/AKT pathway as a key link modulates the multidrug resistance of cancers. Cell Death Dis. 2020, 11, 797. [Google Scholar] [CrossRef] [PubMed]
- Rascio, F.; Spadaccino, F.; Rocchetti, M.T.; Castellano, G.; Stallone, G.; Netti, G.S.; Ranieri, E. The Pathogenic Role of PI3K/AKT Pathway in Cancer Onset and Drug Resistance: An Updated Review. Cancers 2021, 13, 3949. [Google Scholar] [CrossRef]
- Joosten, S.C.; Deckers, I.A.; Aarts, M.J.; Hoeben, A.; van Roermund, J.G.; Smits, K.M.; Melotte, V.; van Engeland, M.; Tjan-Heijnen, V.C. Prognostic DNA methylation markers for renal cell carcinoma: A systematic review. Epigenomics 2017, 9, 1243–1257. [Google Scholar] [CrossRef]
- Moch, H.; Gasser, T.; Amin, M.B.; Torhorst, J.; Sauter, G.; Mihatsch, M.J. Prognostic utility of the recently recommended histologic classification and revised TNM staging system of renal cell carcinoma: A Swiss experience with 588 tumors. Cancer 2000, 89, 604–614. [Google Scholar] [CrossRef]
- Lipworth, L.; Morgans, A.K.; Edwards, T.L.; Barocas, D.A.; Chang, S.S.; Herrell, S.D.; Penson, D.F.; Resnick, M.J.; Smith, J.A.; Clark, P.E. Renal cell cancer histological subtype distribution differs by race and sex. BJU Int. 2016, 117, 260–265. [Google Scholar] [CrossRef]
- Leibovich, B.C.; Lohse, C.M.; Crispen, P.L.; Boorjian, S.A.; Thompson, R.H.; Blute, M.L.; Cheville, J.C. Histological subtype is an independent predictor of outcome for patients with renal cell carcinoma. J. Urol. 2010, 183, 1309–1315. [Google Scholar] [CrossRef]
- Leopold, J.A.; Loscalzo, J. Emerging Role of Precision Medicine in Cardiovascular Disease. Circ. Res. 2018, 122, 1302–1315. [Google Scholar] [CrossRef]
- Yang, H.Y.; Cheng, C.F.; Djoko, B.; Lian, W.S.; Tu, C.F.; Tsai, M.T.; Chen, Y.H.; Chen, C.C.; Cheng, C.J.; Yang, R.B. Transgenic overexpression of the secreted, extracellular EGF-CUB domain-containing protein SCUBE3 induces cardiac hypertrophy in mice. Cardiovasc. Res. 2007, 75, 139–147. [Google Scholar] [CrossRef]
- Xavier, G.M.; Panousopoulos, L.; Cobourne, M.T. Scube3 is expressed in multiple tissues during development but is dispensable for embryonic survival in the mouse. PLoS ONE 2013, 8, e55274. [Google Scholar] [CrossRef]
- Qi, Y.Y.; Zhao, Y.F.; Zhai, Y.L.; Zhang, X.X.; Wang, X.Y.; Liu, X.R.; Cui, Y.; Ning, X.H.; Zhao, Z.Z. SCUBE3 Is Likely a Susceptibility Gene for Systemic Lupus Erythematosus for Chinese Populations. J. Immunol. Res. 2020, 2020, 8897936. [Google Scholar] [CrossRef] [PubMed]
- Fortuna, G.; Brennan, M.T. Systemic lupus erythematosus: Epidemiology, pathophysiology, manifestations, and management. Dent. Clin. N. Am. 2013, 57, 631–655. [Google Scholar] [CrossRef] [PubMed]
- Caielli, S.; Wan, Z.; Pascual, V. Systemic Lupus Erythematosus Pathogenesis: Interferon and Beyond. Annu. Rev. Immunol. 2023, 41, 533–560. [Google Scholar] [CrossRef] [PubMed]
- Capkin, A.A.; Demir, S.; Mentese, A.; Bulut, C.; Ayar, A. Can signal peptide-CUB-EGF domain-containing protein (SCUBE) levels be a marker of angiogenesis in patients with psoriasis? Arch. Dermatol. Res. 2017, 309, 203–207. [Google Scholar] [CrossRef]
- Lowes, M.A.; Bowcock, A.M.; Krueger, J.G. Pathogenesis and therapy of psoriasis. Nature 2007, 445, 866–873. [Google Scholar] [CrossRef]
- Yang, M.; Guo, M.; Hu, Y.; Jiang, Y. Scube regulates synovial angiogenesis-related signaling. Med. Hypotheses 2013, 81, 948–953. [Google Scholar] [CrossRef]
- Liu, Y.; Guerrero-Juarez, C.F.; Xiao, F.; Shettigar, N.U.; Ramos, R.; Kuan, C.H.; Lin, Y.C.; de Jesus Martinez Lomeli, L.; Park, J.M.; Oh, J.W.; et al. Hedgehog signaling reprograms hair follicle niche fibroblasts to a hyper-activated state. Dev. Cell 2022, 57, 1758–1775 e7. [Google Scholar] [CrossRef]
- Driskell, R.R.; Clavel, C.; Rendl, M.; Watt, F.M. Hair follicle dermal papilla cells at a glance. J. Cell Sci. 2011, 124, 1179–1182. [Google Scholar] [CrossRef]
- Biggs, L.C.; Makela, O.J.; Myllymaki, S.M.; Das Roy, R.; Narhi, K.; Pispa, J.; Mustonen, T.; Mikkola, M.L. Hair follicle dermal condensation forms via Fgf20 primed cell cycle exit, cell motility, and aggregation. eLife 2018, 7, e36468. [Google Scholar] [CrossRef]
- Sennett, R.; Wang, Z.; Rezza, A.; Grisanti, L.; Roitershtein, N.; Sicchio, C.; Mok, K.W.; Heitman, N.J.; Clavel, C.; Ma’ayan, A.; et al. An Integrated Transcriptome Atlas of Embryonic Hair Follicle Progenitors, Their Niche, and the Developing Skin. Dev. Cell 2015, 34, 577–591. [Google Scholar] [CrossRef]
- Tu, C.F.; Tsao, K.C.; Lee, S.J.; Yang, R.B. SCUBE3 (signal peptide-CUB-EGF domain-containing protein 3) modulates fibroblast growth factor signaling during fast muscle development. J. Biol. Chem. 2014, 289, 18928–18942. [Google Scholar] [CrossRef]
- Xue, H.; Lu, B.; Lai, M. The cancer secretome: A reservoir of biomarkers. J. Transl. Med. 2008, 6, 52. [Google Scholar] [CrossRef] [PubMed]
- Veyssiere, H.; Bidet, Y.; Penault-Llorca, F.; Radosevic-Robin, N.; Durando, X. Circulating proteins as predictive and prognostic biomarkers in breast cancer. Clin. Proteom. 2022, 19, 25. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Agarwal, R.; Dituri, F.; Lupo, L.; Trerotoli, P.; Mancarella, S.; Winter, P.; Giannelli, G. NGS-based transcriptome profiling reveals biomarkers for companion diagnostics of the TGF-beta receptor blocker galunisertib in HCC. Cell Death Dis. 2017, 8, e2634. [Google Scholar] [CrossRef] [PubMed]
- Javle, M.; Li, Y.; Tan, D.; Dong, X.; Chang, P.; Kar, S.; Li, D. Biomarkers of TGF-beta signaling pathway and prognosis of pancreatic cancer. PLoS ONE 2014, 9, e85942. [Google Scholar] [CrossRef]
- Hadid, R.; Spinedi, E.; Chautard, T.; Giacomini, M.; Gaillard, R.C. Role of several mediators of inflammation on the mouse hypothalamo-pituitary-adrenal axis response during acute endotoxemia. Neuroimmunomodulation 1999, 6, 336–343. [Google Scholar] [CrossRef]
- Matsumura, H.; Mohri, Y.; Binh, N.T.; Morinaga, H.; Fukuda, M.; Ito, M.; Kurata, S.; Hoeijmakers, J.; Nishimura, E.K. Hair follicle aging is driven by transepidermal elimination of stem cells via COL17A1 proteolysis. Science 2016, 351, aad4395. [Google Scholar] [CrossRef]
- Chen, C.C.; Murray, P.J.; Jiang, T.X.; Plikus, M.V.; Chang, Y.T.; Lee, O.K.; Widelitz, R.B.; Chuong, C.M. Regenerative hair waves in aging mice and extra-follicular modulators follistatin, dkk1, and sfrp4. J. Investig. Dermatol. 2014, 134, 2086–2096. [Google Scholar] [CrossRef]
- Hernandez, C.; Huebener, P.; Schwabe, R.F. Damage-associated molecular patterns in cancer: A double-edged sword. Oncogene 2016, 35, 5931–5941. [Google Scholar] [CrossRef]
- Xu, R.; Zhou, X.; Wang, S.; Trinkle, C. Tumor organoid models in precision medicine and investigating cancer-stromal interactions. Pharmacol. Ther. 2021, 218, 107668. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.