Abstract
Arid salt marsh ecosystems endure chronic water scarcity and high salinity stress, with the stability of their functions inextricably linked to the pivotal role of the rhizosphere microenvironment of halophytes. This study focused on three typical halophytes (Kalidium cuspidatum, Nitraria tangutorum, Reaumuria soongarica) in the Jiantan wetland, and deeply explore how these halophytes differently regulate the soil microenvironment through the rhizosphere effect. The results showed that the rhizosphere soil of Kalidium cuspidatum had higher pH, Na+, and K+ contents, while the rhizosphere soil of R. soongarica had higher total carbon, soil organic carbon, alkali-hydrolyzable nitrogen, and microbial biomass. Microbial community analysis revealed that rhizosphere soil of fungal diversity was significantly higher in K. cuspidatum than in R. soongarica, with distinct differences in bacterial and fungal community structures. These differences were closely associated with factors such as Na+, Olsen phosphorus, microbial biomass carbon and alkali-hydrolyzable nitrogen. Among the dominant phyla, Proteobacteria and Ascomycota predominate, with Desulfobacterota and Mortierellomycota exhibiting the highest explanatory power (>48%) for physicochemical property variations. The microbial network of rhizosphere soil of R. soongarica has the highest complexity (with 633 nodes and 3300 edges), but the proportion of positive correlation edges was the lowest (21.58%). Structural equation modeling indicates that soil physical properties indirectly influence network complexity by negatively regulating chemical properties and microbial biomass, while microbial diversity had a direct positive effect on dominant phylum composition and network complexity. This study elucidated the differentiated adaptive strategies of rhizosphere microenvironment-microbe interactions in halophytes, providing a theoretical basis for wetland ecological restoration.
1. Introduction
In the salt marsh ecosystems of arid regions, halophytes are not evenly distributed but follow strict ecological gradients, presenting distinct band-like or patchy patterns [1]. From the core area of the salt marsh to the ecotone between water and land, with changes in soil salinity, moisture conditions, and groundwater level, there are plant species with different salt-tolerant levels distributed successively, such as Phragmites australis, Kalidium cuspidatum, Nitraria tangutorum, Reaumuria soongarica, and Tamarix chinensis. Together, they form a rare green living area in an arid region [2,3]. This distribution pattern is not only the result of plant adaptation and selection to environmental stress but also the core that maintains the stability of this ecosystem [4]. These halophytes fix quicksand and saline soil through their dense root systems, effectively mitigating soil wind and water erosion, and serve as a frontline barrier against desertification [5]. Its vast life forms can also fix carbon in the atmosphere, becoming an important carbon sink[4,5,6]. Therefore, the survival and distribution of halophytes in salt marshes in arid regions directly determine the structure, function, and stability of this ecosystem. In-depth research on these species is the scientific basis for ecological restoration and biodiversity conservation.
The rhizosphere microorganisms of plants serve as a link between plants and soil, and together they form a complex and dynamically balanced rhizosphere micro-ecosystem [7]. Compared with common plants, the rhizosphere of halophytes forms a highly specialized “stress screening environment”, thereby enriching a group of microbial groups with excellent salt tolerance and growth-promoting functions[8]. Bacterial communities, such as Bacillus and Pseudomonas, directly assist plants in resisting salt stress and improving their nutritional status through nitrogen fixation, phosphorus dissolution, secretion of osmotic regulatory substances, and ACC deaminase [9,10]. Fungal communities (especially arbuscular mycorrhizal fungi) play the role of “underground network engineers”, and their vast mycelial networks greatly expand the absorption range of the root system, helping plants obtain water and weakly diffused nutrients such as phosphorus more efficiently in the vast saline soil [11,12]. This rhizosphere micro-ecosystem, dominated by halophytes and featuring the precise collaboration of bacteria and fungi, is the core for its successful survival and ecological restoration functions in harsh saline environments and provides a crucial biological basis for the development of saline-alkali land management strategies based on plant-microbial consortia [13,14].
Bacteria and fungi in the plant rhizosphere do not exist in isolation. Through complex interactions, they form a sophisticated “bacterial–fungal co-occurrence network” [15]. This network was constructed based on high-throughput sequencing and bioinformatics analyses. By statistically analyzing the co-occurrence and mutual exclusion relationships among species, the internal organizational structure and ecological connections of microbial communities are visually revealed [16]. Microbial network complexity is directly related to the functional stability and environmental adaptability of the community [17]. In adverse environments, such as those inhabited by halophytes, a more connected and stable co-occurrence network usually indicates that the microbial community can collaborate more efficiently to help the host plant resist salt stress and obtain nutrients, thereby demonstrating stronger ecological resilience [18,19]. Therefore, analyzing this co-occurrence network is a core link in understanding the assembly rules of the rhizosphere microbial community, its functional synergy, and the mechanism of its impact on plant health.
This study selected three representative species, such as K. cuspidatum, N. tangutorum and R. soongarica. The aim of this study was to systematically reveal the composition, diversity, and structural characteristics of bacterial and fungal communities in the rhizosphere soil. This study will utilize high-throughput sequencing technology and combine the physicochemical parameters of the rhizosphere to explore the differences in the microbial community structure in the rhizosphere of different halophytes, with the aim of characterizing the associations between halophytes and rhizosphere microorganisms in salinized habitats by analyzing bacterial–fungal co-occurrence networks.
2. Materials and Methods
2.1. Overview of the Study Area and Collection of Rhizosphere Soil
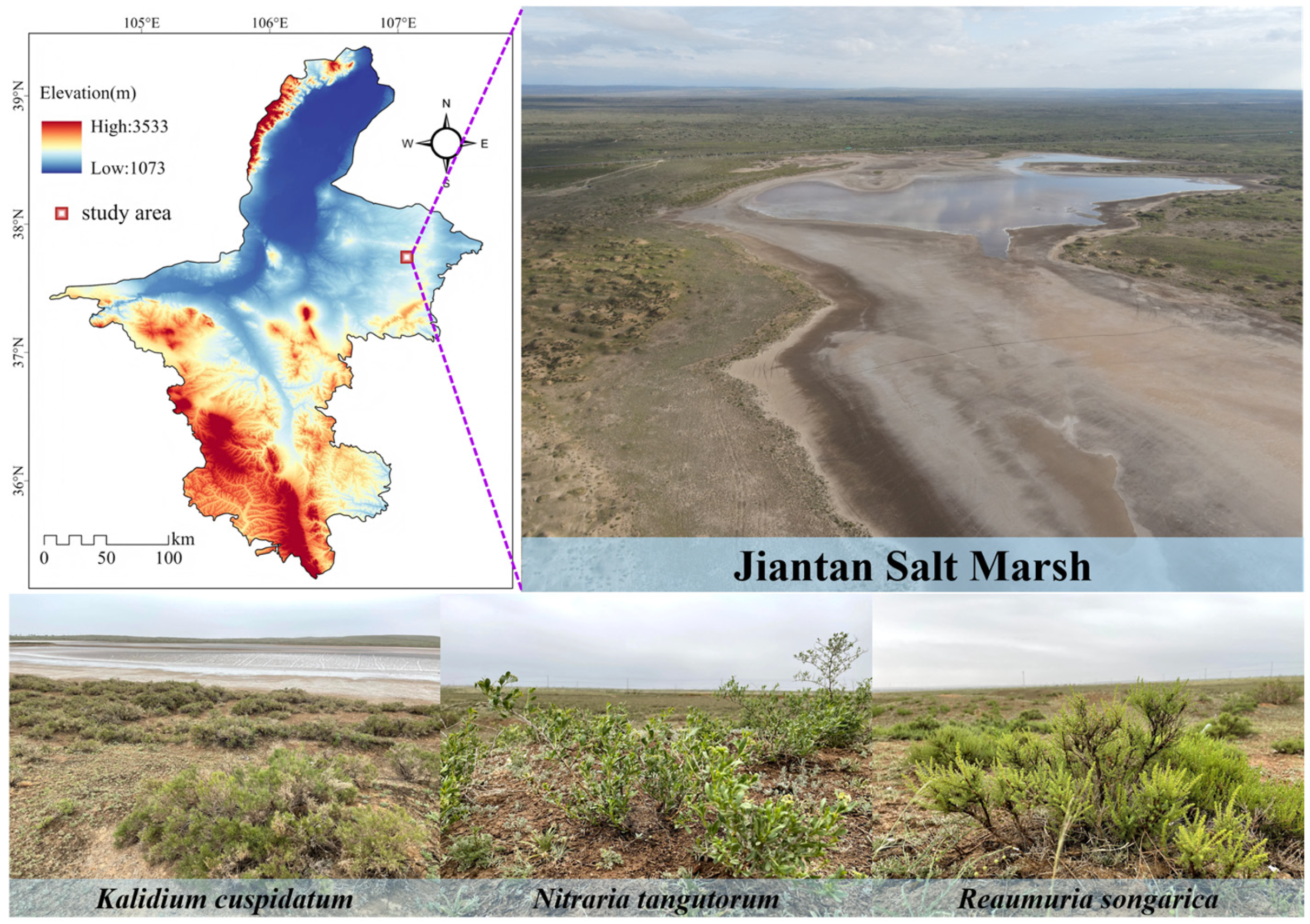
The study site was located in the Jiantan wetland in Yanchi County, central Ningxia. It belongs to a typical desert steppe salt marsh wetland ecosystem, with an altitude of 1398 m, an average annual temperature of 7.8 °C, an annual rainfall of approximately 280 mm, and an evaporation of over 2000 mm [20]. The wetland extends from the water-land ecotone to the desert steppe, mainly distributing plants such as Kalidium cuspidatum, Nitraria tangutorum, Reaumuria soongarica, Sophora alopecuroides, and Stipa caucasica. We investigated three halophytes, K. cuspidatum, N. tangutorum and R. soongarica (Figure 1). The soil attached to the roots of the plants was shaken and collected and defined as rhizosphere soil [21]. A total of 18 samples were collected from the rhizosphere soil of each halophytic plant, with three samples serving as one sample, totaling six samples. Soil samples were transported to the laboratory using a mobile refrigerator. After screening in the laboratory, the physical and chemical parameters and microbial biomass of the soil samples were determined, and soil DNA was extracted.
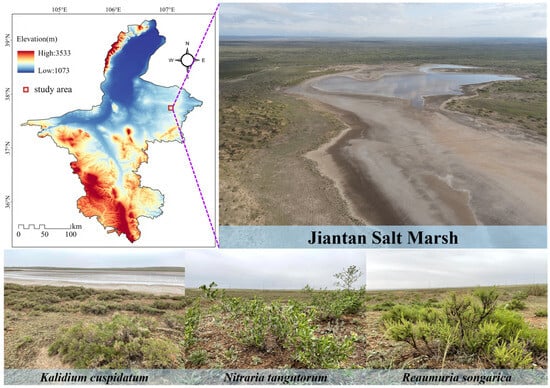
Figure 1.
Sampling sites in the Jiantan wetland in Yanchi County, Ningxia.
2.2. Determination of Physical and Chemical Properties of Soil
The soil water content (SWC) was determined using the weighing method. After the samples were air-dried, deionized water (soil/water = 1/2.5) was added and stirred evenly, and the pH value and electrical conductivity (EC) of the soil were measured using a pH meter and a conductivity meter [22]. The soil Na+ and K+ contents were determined using a flame photometer (FP6410, Shanghai INESA Analytical Instrument Co., Ltd., Shanghai, China) [23]. After the air-dried soil sample was sieved (0.15 mm), the total soil carbon (TC) and inorganic carbon (TIC) contents were determined using the TOC-VSeries SSM-5000A determination component [24]. The value of TC minus TIC is soil total organic carbon (TOC). Similarly, the total nitrogen content (TN) in the soil was determined using an elemental analyzer, and the total phosphorus (TP) content in the soil was determined using the sulfuric acid-molybdenum-dysprosium anticolorimetric method [25]. Soil alkali-hydrolyzable nitrogen (AN) was determined using the alkali-hydrolyzed diffusion method, and available phosphorus was extracted and determined using the sodium bicarbonate method [22]. After the soil samples were extracted with NaHCO3, the Olsen phosphorus content (OP) was determined using molybdenum blue colorimetry [26].
2.3. Determination of Soil Microbial Biomass
Fresh soil samples were fumigated with chloroform (CHCl3), and the fumigated and unfumigated soils were extracted with K2SO4 to determine the dissolved organic carbon (DOC) and total nitrogen in the soil, respectively [27,28]. The phosphorus content of soil microorganisms was determined using the molybdenum blue colorimetric method [29].
Among them, 0.45, 0.54, and 0.40 are the efficiency coefficients of fumigation for extracting carbon, nitrogen, and phosphorus from microbial biomass, respectively.
2.4. Soil DNA Extraction and High-Throughput Sequencing
Weigh fresh soil samples (1.0 g), add the extract (100 mmol·L−1 Tris-HCl, 100 mmol·L−1 EDTA, 200 mmol·L−1 NaCl, 1.0% PVP, 2.0% CTAB, pH 8.0), boil, cool and then centrifuge. The supernatant was extracted and added to (100 mmol·L−1 Tris-HCl, 200 mmol·L−1 NaCl, 2.0% SDS, pH8.0). After centrifugation, the supernatant was used for gel purification. Subsequently, the V5–V7 region of the bacterial 16S rRNA gene was amplified using primers 799F (5′-AACM GGAT TAGA TACC CKG-3′) and 1193R (5′-ACGT CATC CCCA CCTT CC-3′) [30]. PCR amplification of fungal ITS was performed using the primers ITS1F (5 ‘-CTTG GTCA TTTA GAGG AAGT AA-3′) and ITS2 (5 ‘-GCTG CGTT CTTC ATCG ATGC-3′) [31]. After PCR amplification, the samples were sequenced using the Illumina MiSeq PE300 platform. The original data were spliced using FLASH (V1.2.11) software [32]. Cluster analysis of operational taxonomic units (OTUs) was conducted using Usearch software (similarity = 97%) [33]. Subsequently, the α-diversity indices (Shannon and ACE) of bacterial and fungal communities were obtained using QIIME (Version 1.9) software [34]. The original data of bacteria and fungi in the soil samples of this study were submitted to the NCBI database (PRJNA1199940 and PRJNA1200139).
2.5. Data Analysis
All data were statistically analyzed using analysis of variance (ANOVA). Mantel tests for microbial diversity and physicochemical parameters were conducted using the “linkET” package in R software (V4.1) [35], and the data were visualized through the “ggplot2” package [36]. To further reveal the differences in rhizosphere microbial communities among different halophytes, non-metric multidimensional scaling analysis (NMDS) was used for visual display [37]. The “pheatmap” package was used to conduct Spearman analysis on the correlations between the data. The relative abundance (>1%) of the dominant phylum in the rhizosphere of halophytes was visualized using the “ggalluvial” package. Random forest regression analysis was conducted using the “randomForest” package to evaluate the significance of the dominant phyla in each microbial community to physicochemical properties and microbial biomass [38]. This study further identified the dominant bacterial genera in the rhizosphere of halophytes (top 10 in relative abundance) and conducted a variation partitioning analysis of the effects of soil physical behavior, chemical properties, and microbial biomass on the dominant bacterial genera using the “vegan” package [39].
The bacterial–fungal co-occurrence network in the rhizosphere provides direct evidence of the strength of the relationships between different taxa. In this study, Spearman analysis was conducted on the correlation of bacterial–fungal OTUs in the rhizosphere soil of three halophytes (|r| > 0.9, p < 0.01), and a matrix was constructed. The data was visualized using Cytoscape software (3.7.1). The “igraph” package was also used to extract relevant data from the subnetwork [40]. PCA was conducted on nodes, edges, average density, Transitivity, Diameter, and Average path length, and PCA1 was listed as the network complexity index (NCI) [41,42]. The linear regression relationship between dominant phyla and network complexity was demonstrated using the “ggplot2” package, and using “plspm” package to construct the partial least squares path model (PLS-PM) [43].
3. Results
3.1. Differences in Physical and Chemical Properties of Rhizosphere Soil of Halophytes
In the Jiantan wetland, the pH, SWC, Na+, and K+ contents in the K. cuspidatum rhizosphere soil were all higher than those in the N. tangutorum and R. soongarica rhizosphere soils. The Na+ content was 3.14 times and 68.65% higher than that of the N. tangutorum and R. soongarica rhizosphere soils, and the K+ content was 8.87 times and 1.06 times higher (p < 0.05). R. soongarica rhizosphere soil has a relatively high content of TC, TOC, and AN, which are 93.11%, 1.17 times, and 88.89% higher than those of K. cuspidatum, respectively. In addition, the MBC, MBN, and MBP content were higher in the R. soongarica rhizosphere soil, which were 31.45%, 25.66%, and 25.46% higher than those of K. cuspidatum, respectively (p < 0.05) (Table 1).

Table 1.
Physicochemical properties of rhizosphere soil of halophytes.
3.2. The Diversity of Rhizosphere Microorganisms of Halophytes
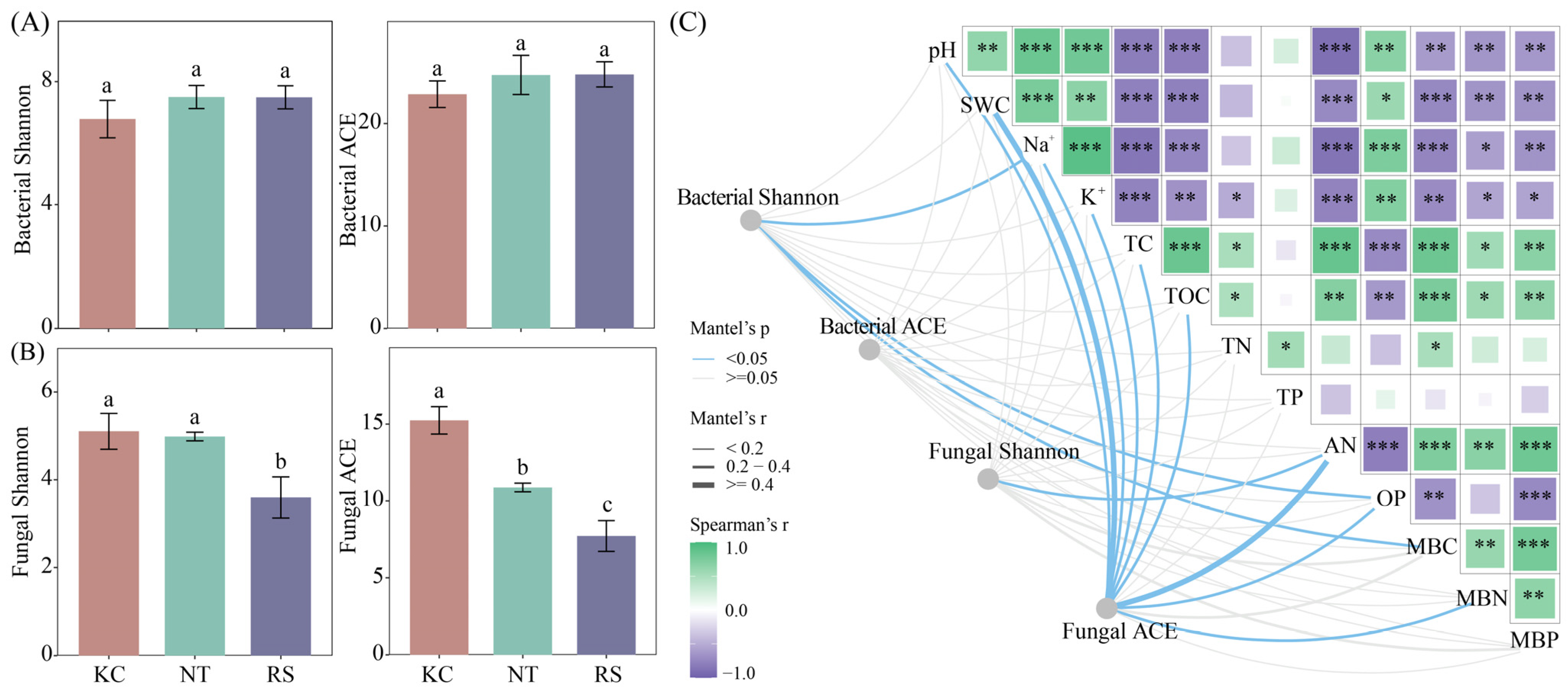
In the Jiantan wetland, there was no significant difference in the Shannon and ACE of the rhizosphere bacterial communities of K. cuspidatum, N. tangutorum, and R. soongarica. The Shannon and ACE fungal communities in the rhizosphere of K. cuspidatum were significantly higher than those in the rhizosphere of R. soongarica (p < 0.05) (Figure 2A,B). Mantel’s test showed that Shannon’s bacterial diversity was significantly correlated with Na+, OP, and MBC. Fungal Shannon and AN had a significant correlation. Fungal ACE was strongly correlated with pH, SWC, TOC, AN, OP and MBN (p < 0.05) (Figure 2C).
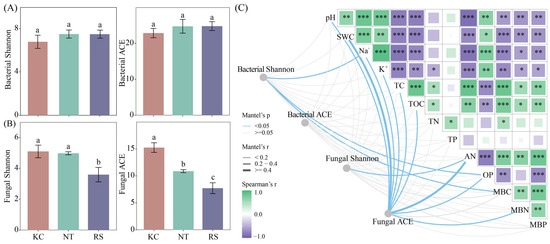
Figure 2.
The alpha diversity (A,B) of bacteria and fungi in the rhizosphere soil of halophytes and its correlation with physical and chemical properties (C). Different lowercase letters indicate significant differences. (*** p < 0.001, ** p < 0.01, * p < 0.05).
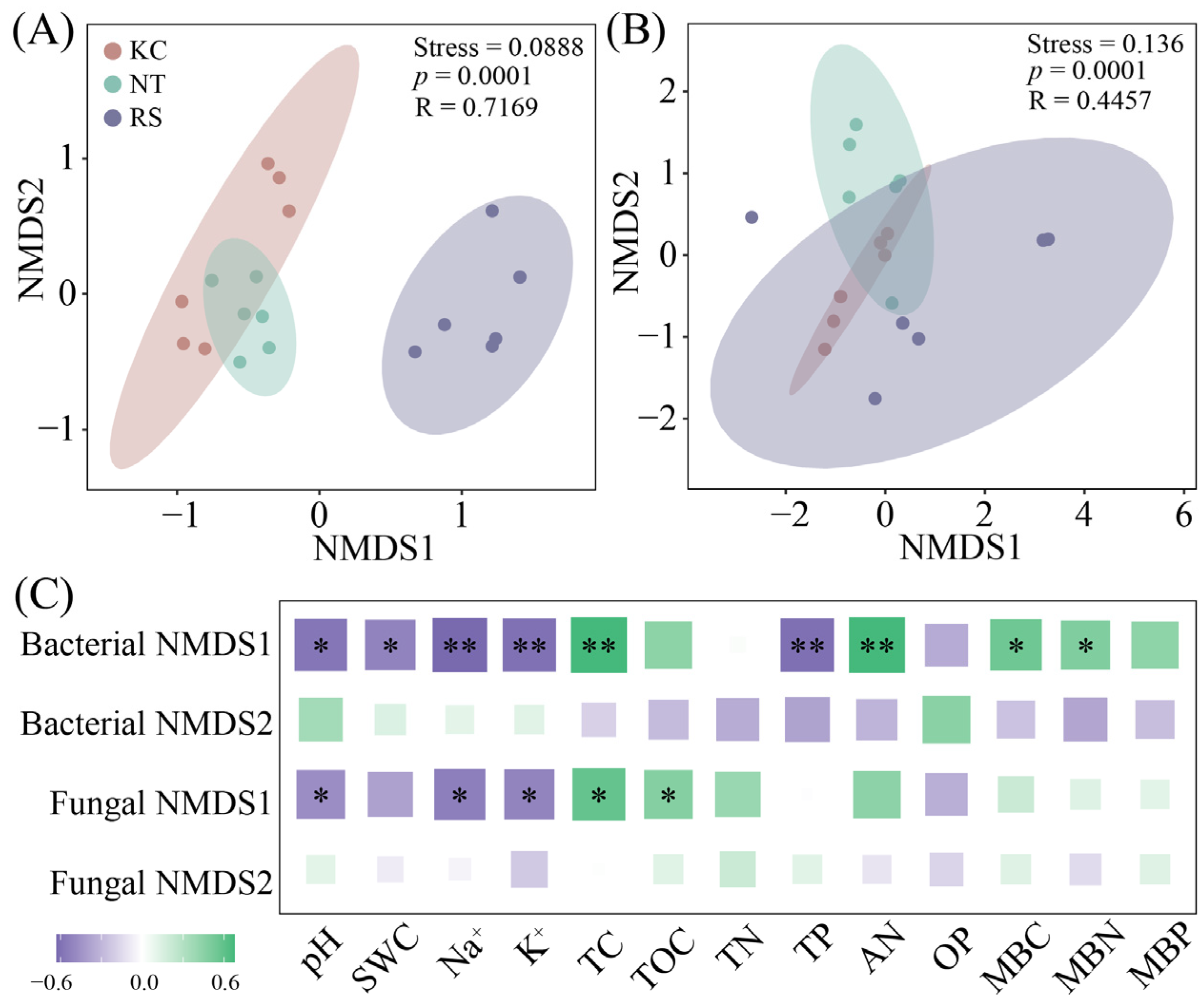
Further analysis revealed that both the rhizosphere bacterial (Stress = 0.0888, p = 0.001) and fungal (Stress = 0.136, p = 0.001) communities of halophytes showed certain differences (Figure 3A,B). Based on the analysis of physicochemical properties, bacterial NMDS1 showed a significant positive correlation with pH, SWC, Na+, K+, and TP, and a significant negative correlation with TC, AN, MBC, and MBN. Fungal NMDS1 was positively correlated with pH, Na+, and K+, and negatively correlated with TC and TOC (p < 0.05) (Figure 3C).
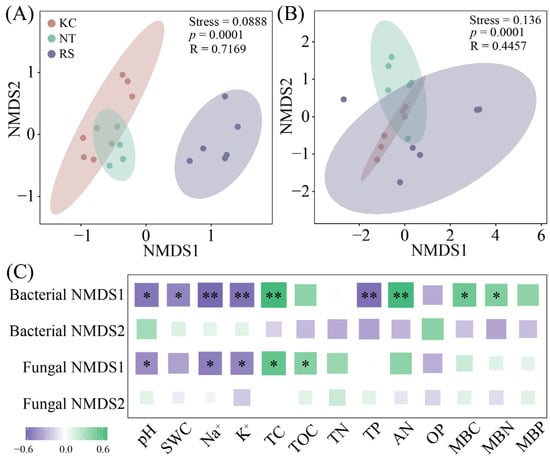
Figure 3.
The beta diversity (A,B) of bacteria and fungi in the rhizosphere soil of halophytes and its correlation with physical and chemical properties (C). (** p < 0.01, * p < 0.05).
3.3. The Community Structure of Rhizosphere Microorganisms of Halophytes
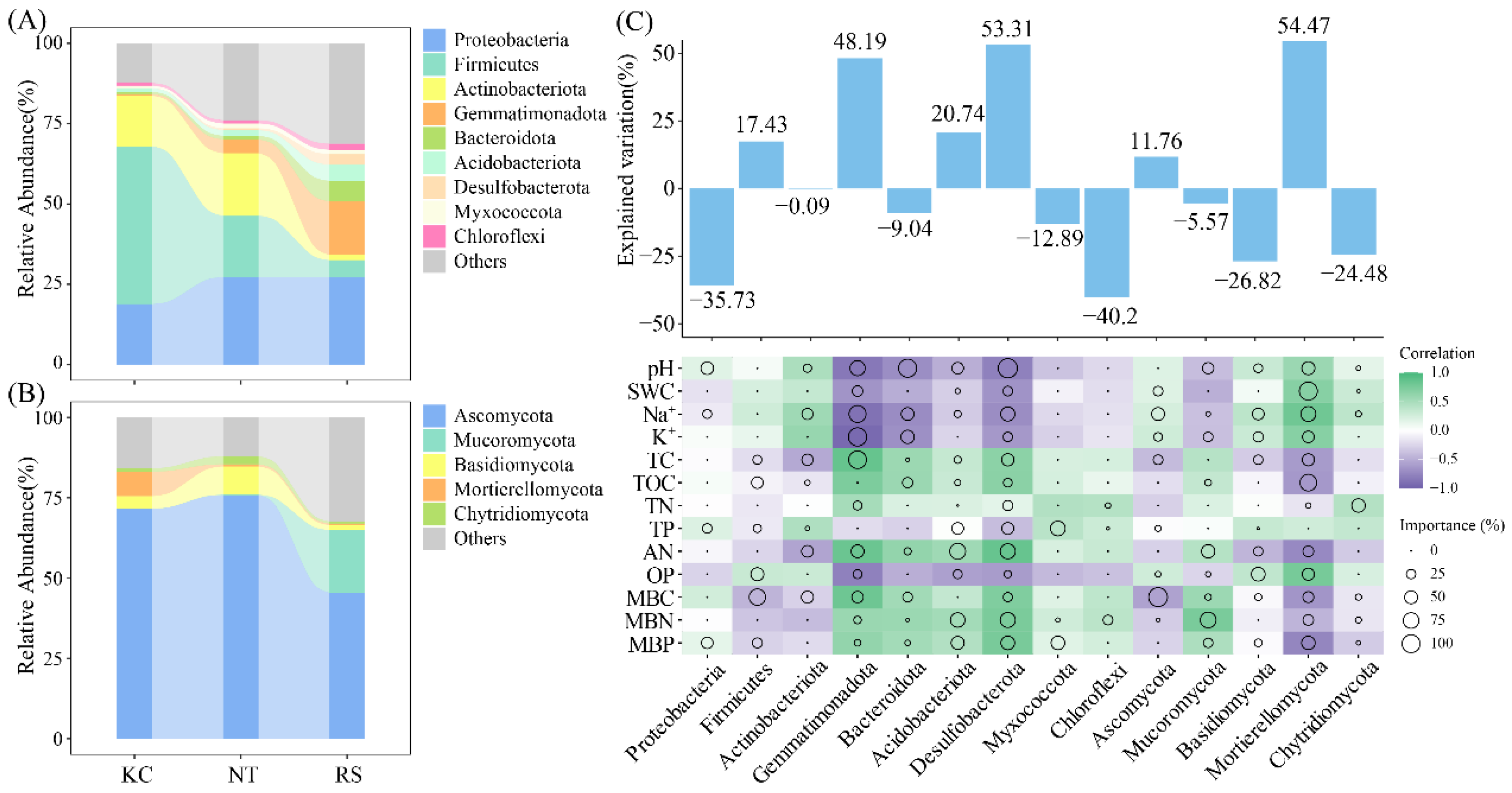
Among the rhizosphere bacterial communities of the three halophytes, Proteobacteria, Firmicutes, Actinobacteriota, and Gemmatimonadota were the dominant phyla (Figure 4A), whereas Ascomycota, Mucoromycota, and Basidiomycota were the dominant phyla of the rhizosphere fungal community (Figure 4B). Further random forest analysis revealed that among the rhizosphere bacteria, Desulfobacterota (53.31%) and Gemmatimonadota (48.19%), And Mortierellomycota (54.47%) among rhizosphere fungi had a higher explanatory rate for the changes in rhizosphere physicochemical properties (p < 0.05) (Figure 4C).
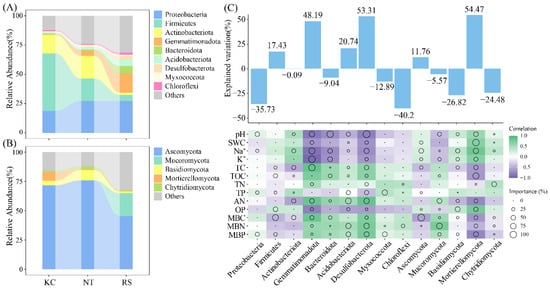
Figure 4.
Relative abundance of dominant bacterial and fungal phyla (A,B) in the rhizosphere soil of halophytes and the random forest analysis with physicochemical properties (C).
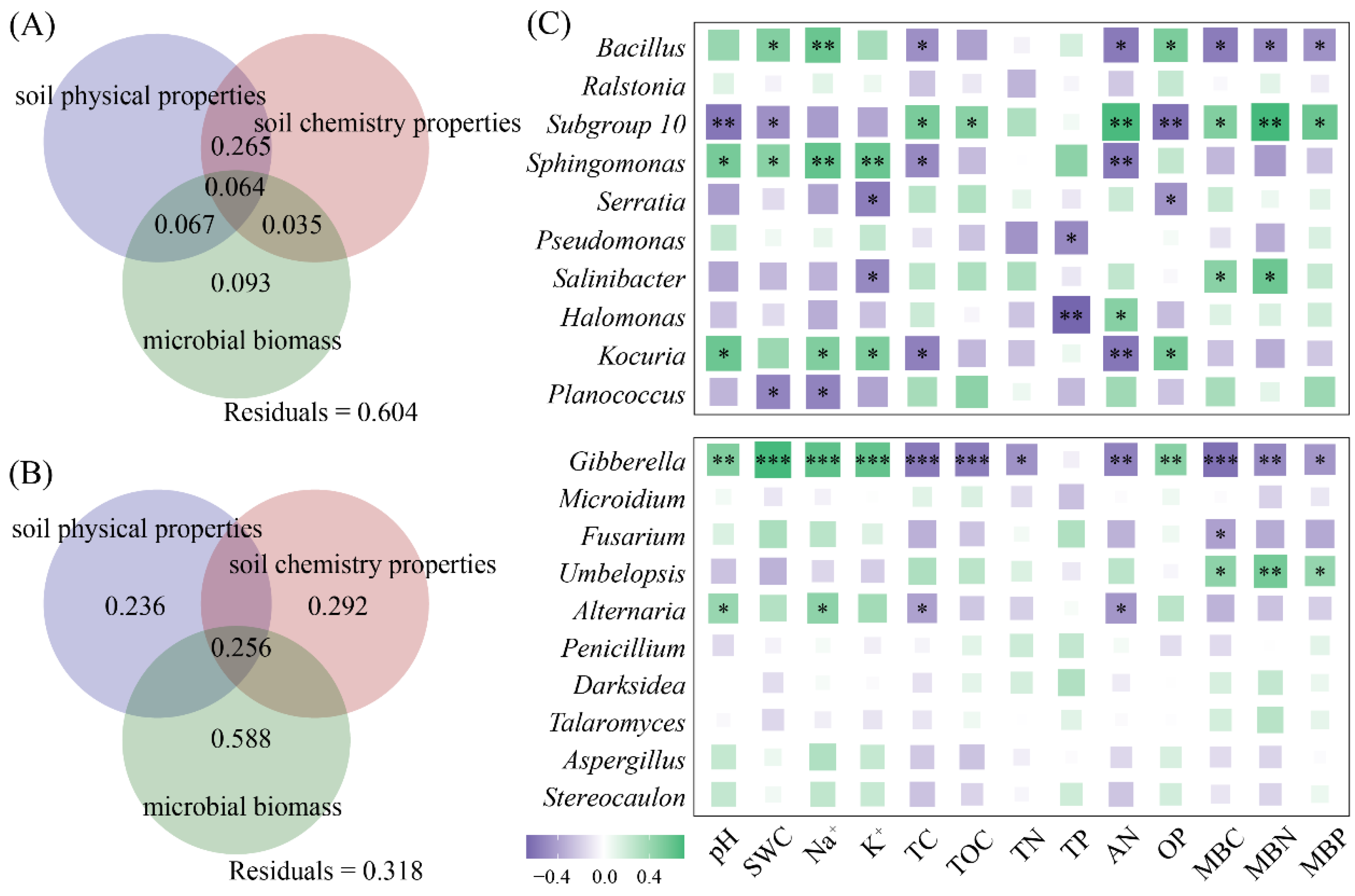
We further selected the top 10 dominant bacterial and fungal genera based on their relative abundance (Figures S1 and S2). Variation decomposition revealed that the physical properties, chemical characteristics, and microbial biomass of the halophyte rhizosphere soil jointly explained 39.6% of the changes in the dominant genera of rhizosphere bacteria (Figure 5A) and had an explanation rate of 68.2% for the changes in the dominant genera of rhizosphere fungi (Figure 5B). The genera and physicochemical properties of rhizosphere bacteria, such as Bacillus, Subgroup 10, Sphingomonas, and Kocuria, were strongly correlated, and the rhizosphere fungi Gibberella and Alternaria were strongly correlated with changes in pH, Na+, TC, and AN (Figure 5C).
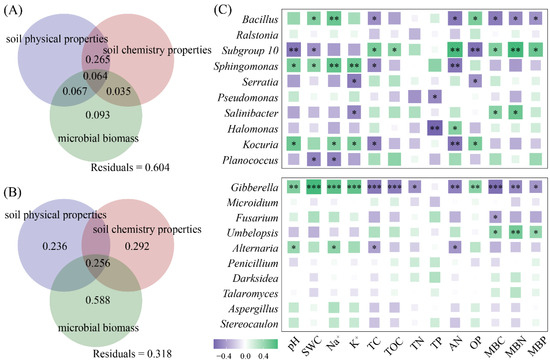
Figure 5.
Variations in physical properties, chemical properties, and microbial biomass in the rhizosphere soil of halophytes (A,B), as well as the correlation between the top 10 genera and the physicochemical properties (C). (*** p < 0.001, ** p < 0.01, * p < 0.05).
3.4. The Bacterial–Fungal Co-Occurrence Network in the Rhizosphere of Halophytes
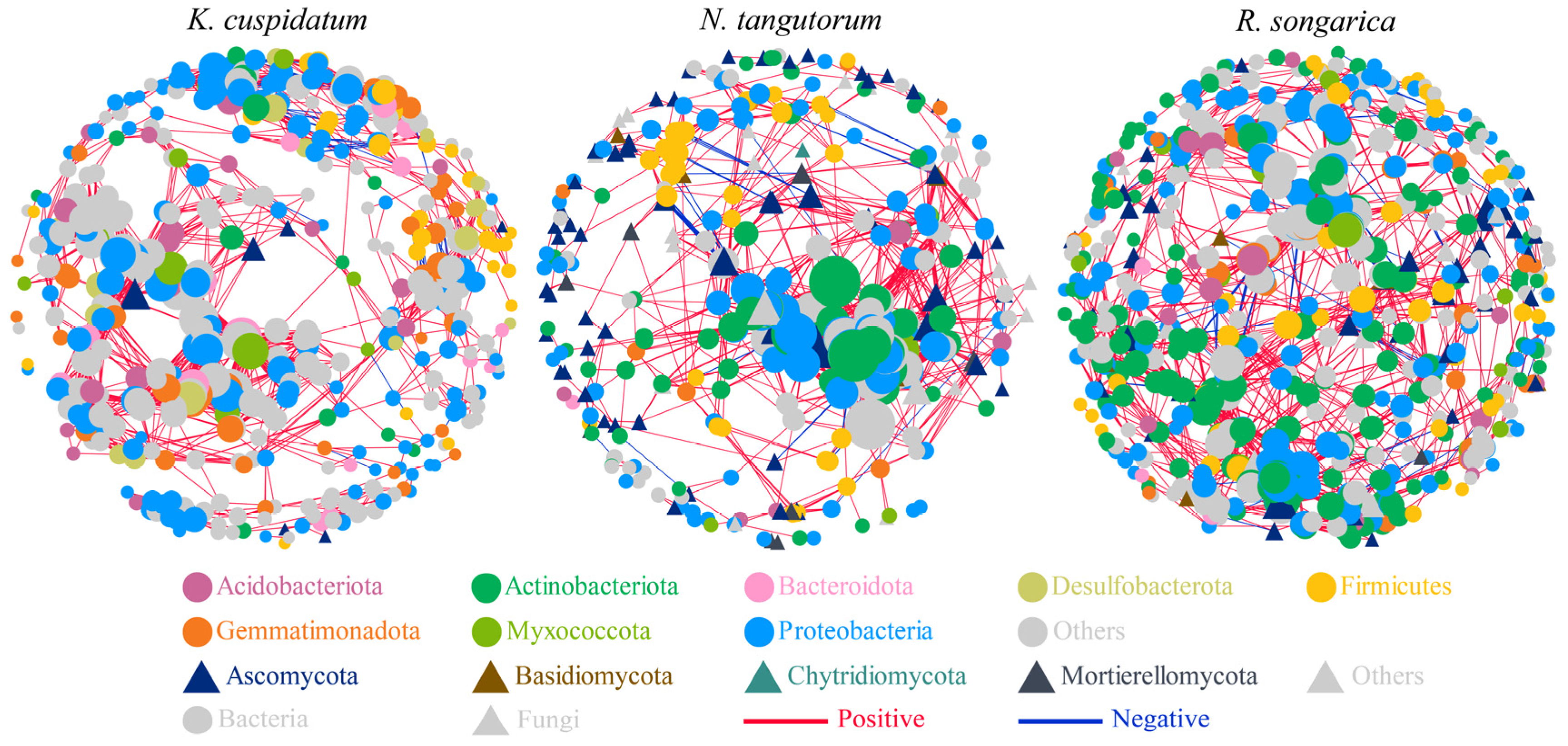
The rhizosphere bacterial–fungal co-occurrence network showed that the OTUs in the nodes were mainly composed of bacteria, such as Acidobacteriota, Actinobacteriota, and Bacteroidota, as well as fungi, including Ascomycota and Basidiomycota. Among them, the R. soongarica rhizosphere network had higher connectivity and complexity (nodes = 633, edges = 3300), but the positive/negative correlation of edges was the lowest (21.58%) (Figure 6, Table 2).
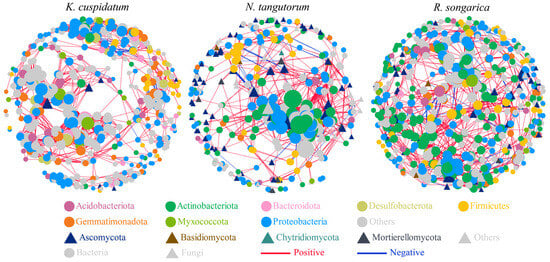
Figure 6.
Co-occurrence network analysis of bacteria and fungi in the rhizosphere soil of different halophytes.

Table 2.
Network parameters in the rhizosphere soil of halophytes.
3.5. The Complexity of Bacterial–Fungal Co-Occurrence Network in the Rhizosphere of Halophytes
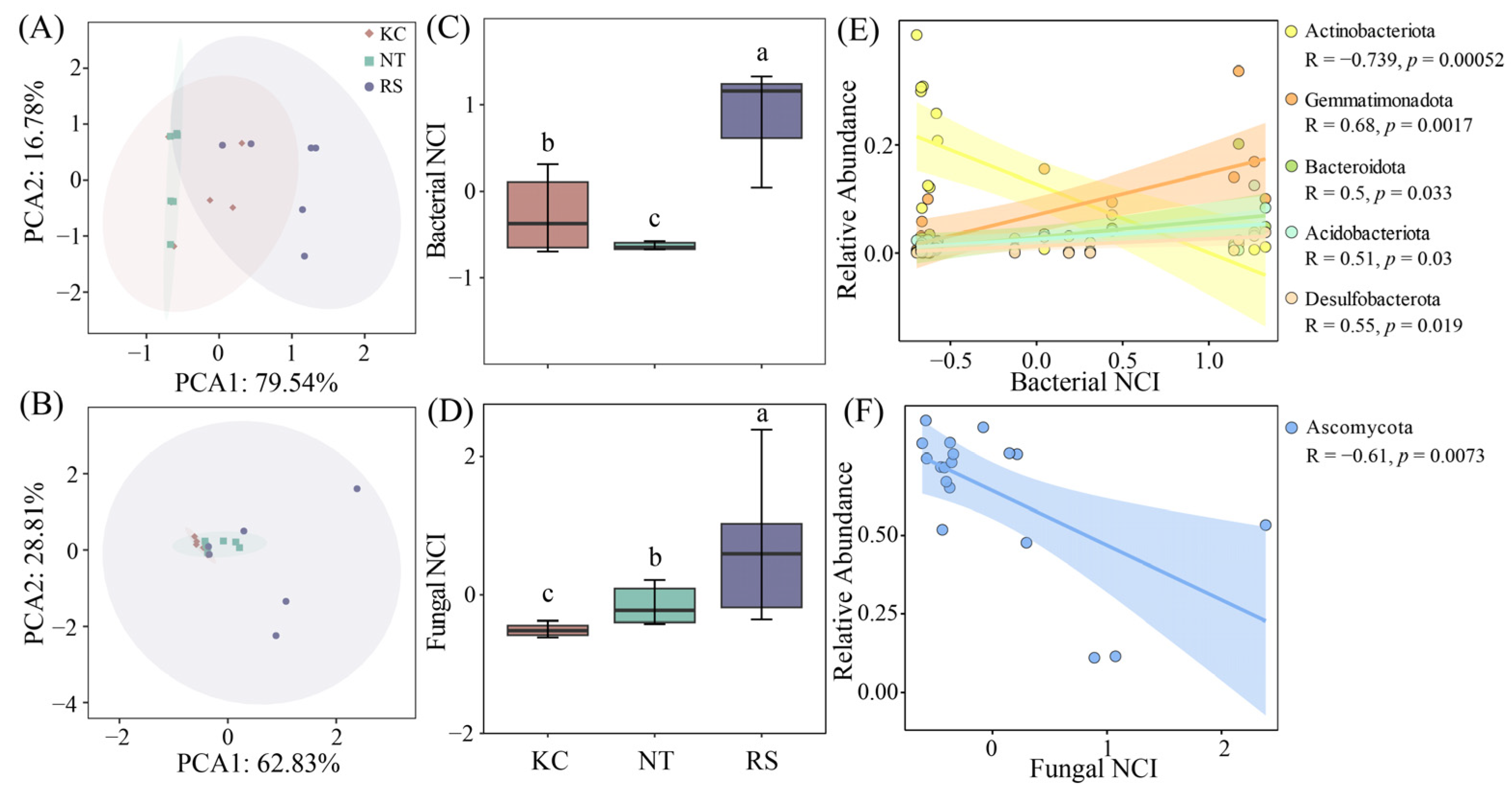
The complexity of the R. soongarica rhizosphere bacterial and fungal networks was higher than that of K. cuspidatum and N. tangutorum (Figure 7A–D). Actinobacteriota, Gemmatimonadota, Bacteroidota, Acidobacteriota, and Desulfobacterota had a significant linear relationship with the bacterial NCI index, and Actinobacteriota showed a negative correlation (Figure 7E). Among fungi, Ascomycota showed a negative correlation with the fungal NCI index (Figure 7F). In the subnetwork, the network nodes, edges, and average density all showed significant correlations with Na+ and TC. The edges, average density, Transitivity, and Average path length also had a relatively obvious correlation with the changes in TP and AN (Figure S3).
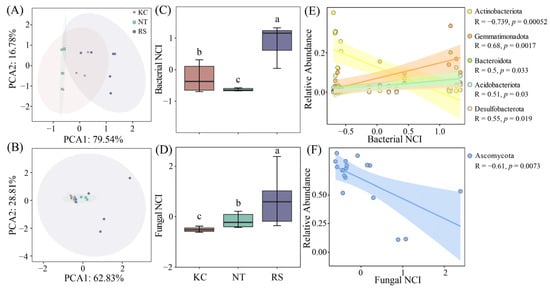
Figure 7.
PCA analysis of bacterial and fungal network parameters in the rhizosphere soil of halophytes (A,B), network complexity indices (C,D) and linear fitting of dominant phyla (E,F). Different lowercase letters indicate significant differences.
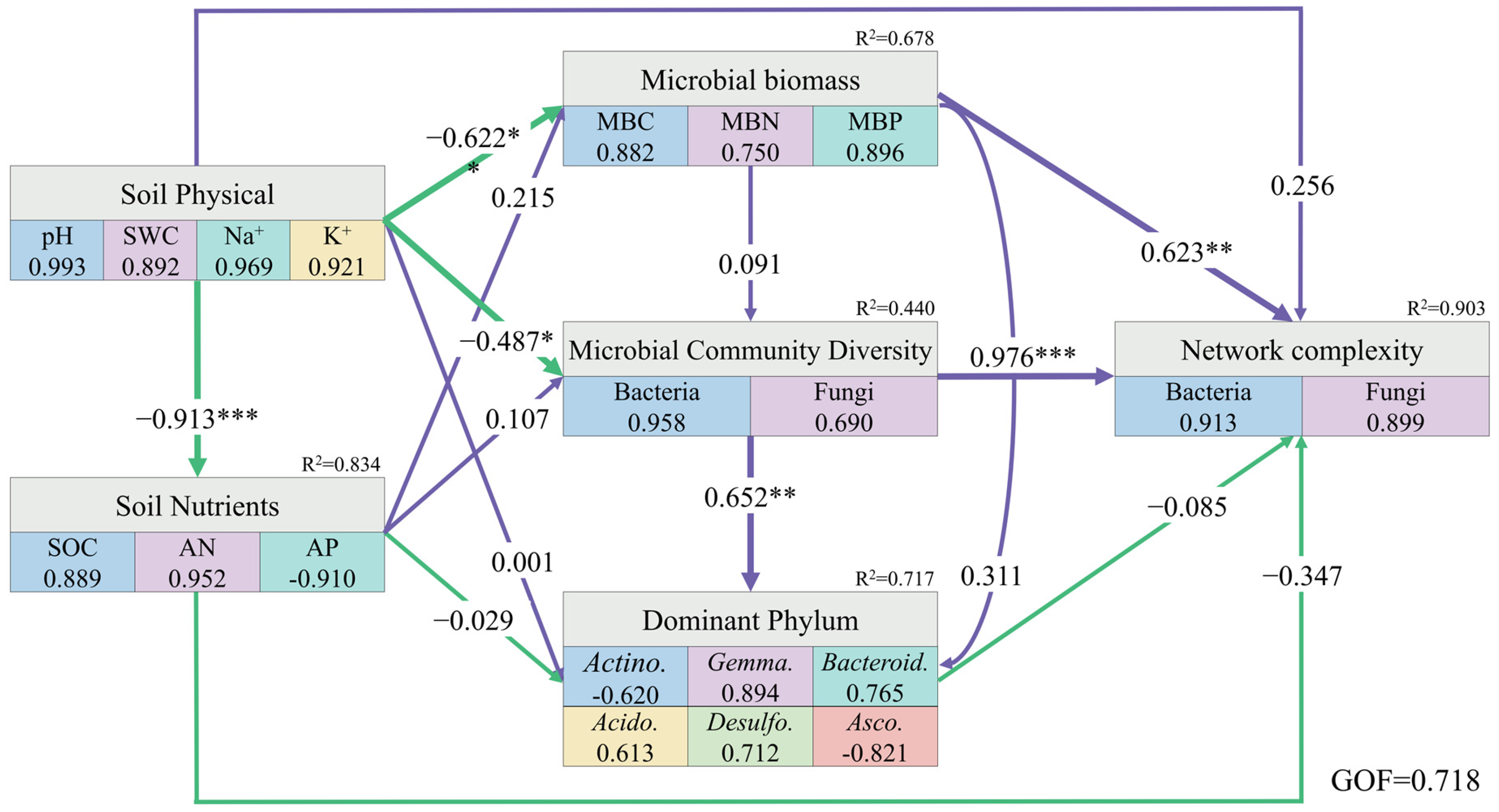
By constructing PLS-PM, the physical properties of the rhizosphere soil in halophytes had a direct negative effect on the chemical properties, while simultaneously negatively regulating microbial biomass and microbial community diversity. The biomass of rhizosphere microorganisms had a significantly positive effect on the network complexity. Moreover, microbial community diversity not only had a direct positive impact on the composition of the dominant phyla but also significantly regulated microbial network complexity (Figure 8).
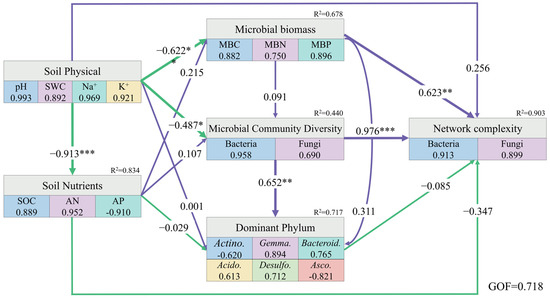
Figure 8.
Partial least squares path model of the rhizosphere soil of halophytes. Purple and green arrows indicate positive and negative causality, respectively, the numbers on the arrows indicate normalized path coefficient, R2 indicates the variance explained variable explained by the model. (*** p < 0.001, ** p < 0.01, * p < 0.05).
4. Discussion
4.1. Differential Rhizosphere Strategies of Halophytes in Desert-Grassland Salt Marsh
The differences in physicochemical properties and microbial biomass among the rhizosphere microenvironments of the three halophytes in the Jiantan wetland reflect their unique ecological adaptation strategies. The rhizosphere of K. cuspidatum exhibited the highest pH, SWC, and Na+ and K+ contents, suggesting that K. cuspidatum may have a stronger ion enrichment capacity or selectively retain high concentrations of Na+ and K+ in the rhizosphere region to maintain osmotic balance for physiological regulation [3]. In contrast, although the Na+ and K+ content in the rhizosphere of R. soongarica is relatively low, it has the highest TC, TOC, and AN content, which provides a more abundant carbon and nitrogen source for its rhizosphere microorganisms [18]. More importantly, the MBC, MBN, and MBP contents in the rhizosphere of R. soongarica were significantly higher than those of K. cuspidatum and N. tangutorum, indicating that its rhizosphere microbial community has a higher biomass and activity. This high microbial biomass may be closely related to the relatively high organic matter content and nutrient availability in the rhizosphere of R. soongarica, providing more favorable conditions for the growth and reproduction of microorganisms [44].
4.2. Typical Composition of Halophyte Rhizosphere Microbial Communities
In this study, the Shannon and ACE indices of the rhizosphere fungi of K. cuspidatum were significantly higher than those of R. soongarica, combined with higher salt (Na+ and K+) and SWC in the rhizosphere. On the one hand, it can be inferred that the rhizosphere fungal community of halophytes is more sensitive to environmental factors [45]; on the other hand, the high-salt environment in the rhizosphere of K. cuspidatum may have maintained a relatively high fungal diversity by inhibiting some sensitive fungal groups and screening out rare fungal species with stronger salt tolerance [3,4,5,6,7,8]. In the NMDS analysis, the differentiation of bacterial community structure (NMDS1 axis) was mainly positively driven by salinization factors (pH, Na+, K+, and TP), whereas organic matter and nitrogen (TC, AN, and MBN) had inhibitory effects. The structure of the fungal community was positively correlated with salt ions (Na+ and K+) and pH and negatively correlated with carbon sources (TC and TOC). These results suggest that the assembly of bacterial communities may be more directly regulated by salt stress and phosphorus dynamics, whereas fungal communities tend to form differentiated distributions in the trade-off between salt enrichment and the availability of carbon and nitrogen resources, which has been confirmed in previous studies [14,21,46].
The rhizosphere bacterial community of halophytes is mainly composed of Proteobacteria, Firmicutes, Actinobacteriota, and Gemmatimonadota. The fungal community was dominated by Ascomycota, Mucoromycota, and Basidiomycota, and this composition pattern is consistent with the characteristics of a typical halophytic environment microbial community [21,47]. More importantly, random forest analysis identified the microbial groups that were most sensitive to changes in the rhizosphere microenvironment of different plants. Their high interpretation rates for changes in physicochemical properties (all exceeding 48%) indicate that these groups are potential key species driving the functional differentiation of rhizosphere microorganisms [48]. Desulfobacterota participate in sulfur cycling, and abundance variation may be associated with rhizosphere redox potential and sulfur metabolism [49]. Mortierellomycota, an important rhizosphere fungus, its distribution may be strongly influenced by the heterogeneity of rhizosphere carbon and nitrogen resources dominated by plant types [50].
At the level of rhizosphere bacteria in halophytes, the close association between specific groups and environmental factors provides concrete evidence for this rule. For instance, the abundance of genera such as Bacillus and Pseudomonas, which are common multifunctional bacterial genera in the rhizosphere, is widely related to various environmental factors, demonstrating the flexibility and functional diversity of bacterial communities in response to environmental changes [51,52]. In contrast, fungi such as Gibberella and Alternaria show specific associations with key salt and nutrient factors such as pH, Na+, TC, and AN, which once again confirms the pattern that the fungal community structure is strongly screened by a few key environmental factors [53,54]. Further research indicated that soil pH profoundly influences plant root development and nutrient uptake efficiency by regulating the chemical form and bioavailability of mineral elements, thereby affecting the composition of the rhizosphere microbial community [55]. The TC content in the plant rhizosphere reflected that the allocation of photosynthetic products is the key factor driving spatial heterogeneity in the rhizosphere microbiome [56]. The nitrogen availability can effectively alleviate the N competition in the plant rhizosphere and enhance the decomposition of SOC by the microbial community [57].
4.3. Structure, Stability, and Environmental Drivers of the Microbial Interaction Network
This study conducted an in-depth analysis of the rhizosphere bacterial–fungal co-occurrence network of halophytes and found that the rhizosphere of R. soongarica has the largest number of nodes and edges and the highest connectivity co-occurrence network, indicating that it supports the most complex and close bacterial–fungal interaction relationship [58]. This high level of network complexity is usually associated with greater functional redundancy and ecosystem stability, which is consistent with the previously observed higher TC, MBC, and MBN in the rhizosphere of R. soongarica. Abundant resources provide a basis for the coexistence and interaction of a large number of microbial groups [59]. However, the proportion of negatively correlated connections in the R. soongarica network was the lowest, indicating that its complex network structure is mainly driven by a large number of positively correlated interactions (competition or antagonism), forming a more coordinated microbial community [60].
Subnetwork analysis provides clues for understanding the driving factors of this complexity [41]. Actinobacteriota and Ascomycota, key phyla of bacteria and fungi, respectively, had relative abundances that were negatively correlated with the NCI index. This indicates that they may play the role of “core competitors” or “dominant players” in the community, and the strengthening of their dominant position may simplify the interaction network [17]. In contrast, the positive correlation between groups such as Gemmatimonadota, Bacteroidota, and NCI suggests that they may be “bridge” groups that enhance network connectivity and complexity [61,62]. In particular, the correlation analysis between network topological attributes and environmental factors clearly outlines the two dominant forces driving the construction of interactive networks, namely, salt stress (Na+) and carbon source basis (TC), which jointly affect the scale (nodes, edges) and connection tightness (average density) of the network [46]. The availability of phosphorus (TP) and nitrogen (AN) more directly regulates the advanced organizational characteristics of the network, such as transitivity and average path length, which are closely related to the efficiency and stability of the network [63].
5. Conclusions
Overall, the differences in the environment and plant types in the Jiantan wetland jointly shape a unique rhizosphere physicochemical microenvironment, which in turn drives the differentiation of bacterial and fungal communities along different ecological gradients. Although the rhizosphere of halophytes shares a similar microbial phyla background, due to the differences in salt and nutrients, different intensities of ecological screening have been carried out for rhizosphere bacterial and fungal communities. The halophyte R. soongarica constructs a cooperative, structurally complex, and functionally highly integrated bacterial–fungal interaction network by creating a relatively low-salt and high-organic rhizosphere environment, which may be an important microbiological mechanism for maintaining the stability of its rhizosphere microecosystem.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microorganisms14030635/s1, Figure S1. The top 10 bacterial genera in the rhizosphere soil of halophytes. Different lowercase letters indicate significant differences. Figure S2. The top 10 fungal genera in the rhizosphere soil of halophytes. Different lowercase letters indicate significant differences. Figure S3. The correlation between the network parameters of bacteria and fungi in the rhizosphere soil of halophytes and their physicochemical properties. (*** p < 0.001, ** p < 0.01, * p < 0.05).
Author Contributions
Y.P. and P.K. conceived and designed the study. R.W., J.H. and J.L. did running the experiments and data management. Y.P., R.W., J.H. and P.K. did writing the original draft. J.H., R.W., Z.C., B.A., X.L., Y.P. and P.K. performed the data mining, statistical analysis, interpretation, figure and table preparation. Y.P. and P.K. did the manuscript revising. All authors contributed to the article and approved the submitted version. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported financially by Xinjiang Natural Science Foundation Project (2025D01B69) and Doctoral Scientific Research Foundation of Xinjiang Normal University (XJNUZBS2532).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data were created or analyzed in this study.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Zhao, K.F.; Song, J.; Feng, G.; Zhao, M.; Liu, J.P. Species, types, distribution, and economic potential of halophytes in China. Plant Soil 2011, 342, 495–509, Erratum in Plant Soil 2011, 342, 511–526. https://doi.org/10.1007/s11104-010-0668-8. [Google Scholar] [CrossRef]
- Pan, Y.Q.; Kang, P.; Hu, J.P.; Song, N.P. Bacterial community demonstrates stronger network connectivity than fungal community in desert-grassland salt marsh. Sci. Total Environ. 2021, 798, 14911. [Google Scholar] [CrossRef]
- Pan, Y.Q.; Kang, P.; Zhang, Y.Q.; Li, X.R. Kalidium cuspidatum colonization changes the structure and function of salt crust microbial communities. Environ. Sci. Pollut. Res. 2024, 31, 19764–19778. [Google Scholar] [CrossRef]
- Qiu, D.D.; Zhu, G.F.; Bhat, M.A.; Wang, L.; Liu, Y.W.; Sang, L.Y.; Lin, X.R.; Zhang, W.H.; Sun, N. Water use strategy of nitraria tangutorum shrubs in ecological water delivery area of the lower inland river: Based on stable isotope data. J. Hydrol. 2023, 624, 129918. [Google Scholar] [CrossRef]
- Zhao, S.; van der Heijden, M.G.A.; Banerjee, S.; Liu, J.J.; Gu, H.D.; Zhou, N.; Yin, C.H.; Peng, B.; Liu, X.; Wang, B.Z.; et al. The role of halophyte-induced saline fertile islands in soil microbial biogeochemical cycling across arid ecosystems. Commun. Biol. 2024, 7, 1061. [Google Scholar] [CrossRef]
- Cong, M.F.; Zhang, Z.H.; Zhao, G.X.; Dong, X.P.; Lu, Y.; Li, L.; Hu, Y.; Wang, W.Q.; Mu, Z.B.; Tariq, A.; et al. The importance of fungal necromass in soil organic carbon: Insights from elevation gradients in arid regions. Appl. Soil Ecol. 2025, 214, 106376. [Google Scholar] [CrossRef]
- Xiao, Y.; Li, S.Z.; Zhang, M.M.; Zhou, H.; Qu, Y.Y.; Zhan, J.J.; Zhang, X.W. Environmental factors and biotic interactions shape the regional-scale assembly of rhizosphere microbial communities associated with Suaeda salsa. Plant Soil 2025, 518, 1545–1560. [Google Scholar] [CrossRef]
- Li, L.; Cheng, K.X.; Du, Y.; Zhang, Y.W.; Zhou, Y.W.; Jin, Y.; He, X.Q. Rhizosphere microbes from Populus euphratica conferred salt stress resistance to Populus alba × Populus glandulosa. Plant Cell Environ. 2025, 48, 8743–8755. [Google Scholar] [CrossRef]
- Huang, Y.; Zhang, X.Y.; Ma, J.B.; Jiao, H.C.; Muhammad, M.; Abdugheni, R.; Shurigin, V.; Egamberdieva, D.; Li, L. Diversity and plant growth-promoting properties of culturable bacteria associated with three halophytes in an arid land, Northwest China. J. Arid Land 2025, 17, 696–713. [Google Scholar] [CrossRef]
- Colchado-López, J.; Rougon-Cardoso, A.; Vélez, P.; Rosas, U. Meta-analysis of community composition patterns of halophyte and xerophyte rhizosphere associated bacteria. Rhizosphere 2022, 24, 100588. [Google Scholar] [CrossRef]
- Meena, K.K.; Bitla, U.; Sorty, A.M.; Kumar, S.; Kumar, S.; Wakchaure, G.C.; Singh, D.P.; Stougaard, P.; Suprasanna, P. Understanding the interaction and potential of halophytes and associated microbiome for bio-saline agriculture. J. Plant Growth Regul. 2023, 42, 6601–6619. [Google Scholar] [CrossRef]
- Diao, F.W.; Jia, B.B.; Luo, J.Q.; Ding, S.L.; Liu, T.; Guo, W. Arbuscular mycorrhizal fungi drive bacterial community assembly in halophyte Suaeda salsa. Microbiol. Res. 2024, 282, 127657. [Google Scholar] [CrossRef]
- Wang, R.M.; Cui, L.J.; Li, J.; Li, W. Factors driving the halophyte rhizosphere bacterial communities in coastal salt marshes. Front. Microbiol. 2023, 14, 1127958. [Google Scholar] [CrossRef] [PubMed]
- Qu, X.; Pan, Y.Q.; Wang, P.Q.; Ran, L.L.; Qin, G.F.; Li, Q.F.; Kang, P. Response of phyllosphere and rhizosphere microbial communities to salt stress of tamarix chinensis. Plants 2024, 13, 1091. [Google Scholar] [CrossRef]
- Gao, C.; Xu, L.; Montoya, L.; Madera, M.; Hollingsworth, J.; Chen, L.; Purdom, E.; Singan, V.; Vogel, J.; Hutmacher, R.; et al. Co-occurrence networks reveal more complexity than community composition in resistance and resilience of microbial communities. Nat. Commun. 2022, 13, 3867. [Google Scholar] [CrossRef] [PubMed]
- Kang, P.; Pan, Y.Q.; Yang, P.; Hu, J.P.; Zhao, T.L.; Zhang, Y.Q.; Ding, X.D.; Yan, X.F. A comparison of microbial composition under three tree ecosystems using the stochastic process and network complexity approaches. Front. Microbiol. 2022, 13, 1018077. [Google Scholar] [CrossRef]
- Pan, Y.Q.; Kang, P.; Qu, X.; Zhang, H.X.; Li, X.R. Response of the soil bacterial community to seasonal variations and land reclamation in a desert grassland. Ecol. Indic. 2024, 165, 112227. [Google Scholar] [CrossRef]
- Hu, J.P.; He, Y.Y.; Li, J.H.; Lü, Z.L.; Zhang, Y.W.; Li, Y.H.; Li, J.L.; Zhang, M.X.; Cao, Y.H.; Zhang, J.L. Planting halophytes increases the rhizosphere ecosystem multifunctionality via reducing soil salinity. Environ. Res. 2024, 261, 119707. [Google Scholar] [CrossRef]
- Qiao, Y.Z.; Wang, T.T.; Huang, Q.W.; Guo, H.Y.; Zhang, H.; Xu, Q.C.; Shen, Q.R.; Ling, N. Core species impact plant health by enhancing soil microbial cooperation and network complexity during community coalescence. Soil Biol. Biochem. 2024, 188, 109231. [Google Scholar]
- Chen, X.H.; Duan, Z.H. Changes in soil physical and chemical properties during reversal of desertification in Yanchi County of Ningxia Hui autonomous region, China. Environ. Geol. 2009, 57, 975–985. [Google Scholar]
- Pan, Y.Q.; Kang, P.; Tan, M.; Hu, J.P.; Zhang, Y.Q.; Zhang, J.L.; Song, N.P.; Li, X.R. Root exudates and rhizosphere soil bacterial relationships of Nitraria tangutorum are linked to k-strategists bacterial community under salt stress. Front. Plant Sci. 2022, 13, 997292. [Google Scholar] [CrossRef]
- Bao, S.D. Soil Agricultural Chemical Analysis; China Agricultural Press: Beijing, China, 2000. [Google Scholar]
- Bettinelli, M.; Beone, G.M.; Spezia, S.; Baffi, C. Determination of heavy metals in soils and sediments by microwave-assisted digestion and inductively coupled plasma optical emission spectrometry analysis. Anal. Chim. Acta 2000, 424, 289–296. [Google Scholar] [CrossRef]
- Shimadzu, C. Instruction Manual for TOC-V Series and SSM-5000A Solid Sample Measurement Unit; Shimadzu Corporation: Kyoto, Japan, 2004. [Google Scholar]
- Luo, R.Y.; Kuzyakov, Y.; Zhu, B.; Qiang, W.; Zhang, Y.; Pang, X.Y. Phosphorus addition decreases plant lignin but increases microbial necromass contribution to soil organic carbon in a subalpine forest. Glob. Change Biol. 2022, 28, 4194–4210. [Google Scholar] [CrossRef] [PubMed]
- Olsen, S.R. Estimation of Available Phosphorus in Soils by Extraction with Sodium Bicarbonate; United States Department of Agriculture: Washington, DC, USA, 1954. [Google Scholar]
- Brookes, P.C.; Landman, A.; Pruden, G.; Jenkinson, D.S. Chloroform fumigation and the release of soil nitrogen: A rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol. Biochem. 1985, 17, 837–842. [Google Scholar] [CrossRef]
- Vance, E.D.; Brookes, P.C.; Jenkinson, D.S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 1987, 19, 703–707. [Google Scholar] [CrossRef]
- Brookes, P.C.; Powlson, D.S.; Jenkinson, D.S. Measurement of microbial biomass phosphorus in soil. Soil Biol. Biochem. 1982, 14, 319–329. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef]
- Op De Beeck, M.; Lievens, B.; Busschaert, P.; Declerck, S.; Vangronsveld, J.; Colpaert, J.V. Comparison and validation of some ITS primer pairs useful for fungal metabarcoding studies. PLoS ONE 2014, 9, e97629. [Google Scholar] [CrossRef]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Haas, B.J.; Gevers, D.; Earl, A.M.; Feldgarden, M.; Ward, D.V.; Giannoukos, G.; Ciulla, D.; Tabbaa, D.; Highlander, S.K.; Sodergren, E.; et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 2011, 21, 494–504. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Sunagawa, S.; Coelho, L.P.; Chaffron, S.; Kultima, J.R.; Labadie, K.; Salazar, G.; Djahanschiri, B.; Zeller, G.; Mende, D.R.; Alberti, A.A. Structure and function of the global ocean microbiome. Science 2015, 348, 1261359. [Google Scholar] [CrossRef]
- Racine, J.S. RStudio:A platform-independent IDE for R and Sweave. J. Appl. Economet. 2012, 27, 167–172. [Google Scholar] [CrossRef]
- Ramette, A. Multivariate analyses in microbial ecology. FEMS Microbiol. Ecol. 2007, 62, 142–160. [Google Scholar] [CrossRef]
- Jiao, S.; Chen, W.M.; Wang, J.L.; Du, N.N.; Li, Q.P.; Wei, G.H. Soil microbiomes with distinct assemblies through vertical soil profiles drive the cycling of multiple nutrients in reforested ecosystems. Microbiome 2018, 6, 146. [Google Scholar] [CrossRef]
- Blanchet, F.G.; Legendre, P.; Borcard, D. Forward selection of explanatory variables. Ecology 2008, 89, 2623–2632. [Google Scholar] [CrossRef]
- Csardi, G.; Nepusz, T. The igraph software. Complex Syst. 2006, 1695, 1–9. [Google Scholar]
- Ma, B.; Wang, H.Z.; Dsouza, M.; Lou, J.; He, Y.; Dai, Z.M.; Brookes, P.C.; Xu, J.; Gilbert, J.A. Geographic patterns of co-occurrence network topological features for soil microbiota at continental scale in eastern China. ISME J. 2016, 10, 1891–1901. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.P.; Zhang, Q.; Zhu, H.S.; Reich, P.B.; Banerjee, S.; van der Heijden, M.G.A.; Sadowsky, M.J.; Ishii, S.S.; Jia, X.X.; Shao, M.G.; et al. Erosion reduces soil microbial diversity, network complexity and multifunctionality. ISME J. 2021, 15, 2474–2489. [Google Scholar] [CrossRef] [PubMed]
- Rigdon, E.E. Choosing PLS path modeling as analytical method in European management research: A realist perspective. Eur. Manag. J. 2016, 34, 598–605. [Google Scholar] [CrossRef]
- Kang, P.; Pan, Y.Q.; Hu, J.P.; Qu, X.; Ji, Q.B.; Zhuang, C.Y.; Ren, Y.F.; Zhou, J.; Wei, T.J. Straw mulch and orchard grass mediate soil microbial nutrient acquisition and microbial community composition in Ziziphus Jujuba orchard. Plant Soil 2025, 512, 1203–1219. [Google Scholar] [CrossRef]
- Abrego, N.; Roslin, T.; Huotari, T.; Tack, A.J.M.; Lindahl, B.D.; Tikhonov, G.; Somervuo, P.; Schmidt, N.M.; Ovaskainen, O. Accounting for environmental variation in co-occurrence modelling reveals the importance of positive interactions in root-associated fungal communities. Mol. Ecol. 2020, 29, 2736–2746. [Google Scholar] [CrossRef]
- Zhang, X.C.; Zhai, H.L.; Xu, H.Y.; Song, W.Z.; Liu, M.; Dong, X.F.; Sun, W.; Ma, J.Y. Different salt stress types regulated rhizosphere rare bacterial communities through root exudates and soil physicochemical properties. Plant Soil 2025, 516, 797–811. [Google Scholar] [CrossRef]
- Li, W.J.; Li, Y.; Lv, J.; He, X.M.; Wang, J.L.; Teng, D.X.; Jiang, L.M.; Wang, H.F.; Lv, G.H. Rhizosphere effect alters the soil microbiome composition and C, N transformation in an arid ecosystem. Appl. Soil Ecol. 2022, 170, 104296. [Google Scholar] [CrossRef]
- Kang, P.; Pan, Y.Q.; Ran, Y.C.; Li, W.N.; Shao, M.X.; Zhang, Y.Q.; Ji, Q.B.; Ding, X.D. Soil saprophytic fungi could be used as an important ecological indicator for land management in desert steppe. Ecol. Indic. 2023, 150, 110224. [Google Scholar] [CrossRef]
- Fang, Y.; Liu, J.; Yang, J.; Wu, G.; Hua, Z.S.; Dong, H.L.; Hedlund, B.P.; Baker, B.J.; Jiang, H.C. Compositional and metabolic responses of autotrophic microbial community to salinity in lacustrine environments. mSystems 2022, 7, e0033522. [Google Scholar] [CrossRef]
- Liu, Y.P.; Ise, Y.; Takami, H.; Urakawa, R.; Tateno, R.; Toyoda, A.; Ohte, N.; Shi, W.; Jiang, L.F.; Isobe, K. Soil pH modulates microbial nitrogen allocation in soil via compositional and metabolic shifts across forests in Japan. iMetaOmics 2025, 2, e70054. [Google Scholar] [CrossRef]
- Sierra-García, I.N.; Ferreira, M.J.; Torres-Ballesteros, A.; Louvado, A.; Gomes, N.; Cunha, A. Brevibacterium EB3 inoculation enhances rhizobacterial community interactions leading to improved growth of Salicornia europaea. Appl. Soil Ecol. 2024, 196, 105306. [Google Scholar] [CrossRef]
- Slatni, T.; Zorrig, W.; Razzegui, A.; Antonio Hernández, J.; Barba-Espín, G.; Ben Hamed, K.; Díaz-Vivancos, P. Halophilic Bacillus improve barley growth on calcareous soil via enhanced photosynthetic performance and metabolomic re-programing. J. Plant Physiol. 2025, 309, 154495. [Google Scholar] [CrossRef]
- Pinho, D.; Barroso, C.; Froufe, H.; Brown, N.; Vanguelova, E.; Egas, C.; Denman, S. Linking tree health, rhizosphere physicochemical properties, and microbiome in acute oak decline. Forests 2020, 11, 1153. [Google Scholar] [CrossRef]
- Qi, X.Y.; Chen, T.; Ding, C.; Chen, X.M.; He, B.; Hu, W.G. Diversity of fungal communities in the rhizosphere soil of Tamarix chinensis in saline–alkaline wetland. Environ. Dev. Sustain. 2025, 27, 8693–8709. [Google Scholar] [CrossRef]
- Liu, L.; Song, W.; Huang, S.J.; Jiang, K.; Moriwaki, Y.; Wang, Y.C.; Men, Y.F.; Zhang, D.; Wen, X.; Han, Z.F.; et al. Extracellular pH sensing by plant cell-surface peptide-receptor complexes. Cell 2022, 185, 3341–3355.e13. [Google Scholar] [CrossRef]
- Schultes, S.R.; Rüger, L.; Niedeggen, D.; Freudenthal, J.; Frindte, K.; Becker, M.F.; Metzner, R.; Pflugfelder, D.; Chlubek, A.; Hinz, C.; et al. Photosynthate distribution determines spatial patterns in the rhizosphere microbiota of the maize root system. Nat. Commun. 2025, 16, 7286. [Google Scholar] [CrossRef]
- Bai, T.S.; Qiu, Y.P.; Hu, S.J. Nitrogen availability mediates the effects of roots and mycorrhizal fungi on soil organic carbon decomposition: A meta-analysis. Pedosphere 2024, 34, 289–296, Erratum in Pedosphere 2024, in press. Available online: https://www.sciencedirect.com/science/article/pii/S1002016024001139?utm_source=chatgpt.com (accessed on 5 December 2024). [Google Scholar] [CrossRef]
- Chen, W.Q.; Wang, J.Y.; Chen, X.; Meng, Z.X.; Xu, R.; Duoji, D.Z.; Zhang, J.H.; He, J.; Wang, Z.G.; Chen, J.; et al. Soil microbial network complexity predicts ecosystem function along elevation gradients on the Tibetan Plateau. Soil Biol. Biochem. 2022, 172, 108766. [Google Scholar] [CrossRef]
- Liu, Q.; Tang, L.; Sun, H.R.; Kong, X.Q.; Jiao, Y.; Wu, W.Q.; Li, S.; Shen, Y.F. Responses of the fungal-bacterial community and network to surface mulching and nitrogen fertilization in the Loess Plateau. Plant Soil 2024, 494, 111–126. [Google Scholar] [CrossRef]
- Liu, B.; Li, X.; Cao, W.; Li, Q. Negative interactions between bacteria and fungi modulate life history strategies of soil bacterial communities in temperate shrublands under precipitation gradients. Funct. Ecol. 2025, 39, 3459–3473. [Google Scholar] [CrossRef]
- Hu, P.L.; Zhang, W.; Wanek, W.; Chen, J.; Abalos, D.; Zhao, J.; Xiao, D.; Hou, X.Y.; Li, J.; Chen, H.S.; et al. Calcium-rich parent materials enhance multiple soil functions and bacterial network complexity. Commun. Earth Environ. 2025, 6, 797. [Google Scholar] [CrossRef]
- Shi, J.W.; Yang, L.; Liao, Y.; Li, J.W.; Jiao, S.; Shangguan, Z.P.; Deng, L. Soil labile organic carbon fractions mediate microbial community assembly processes during long-term vegetation succession in a semiarid region. Imeta 2023, 2, e142. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Guo, X.N.; Luo, K.Y.; Xiao, X.Z.; Zhong, H.H.; Sun, J.; Xia, J.X.; Chen, Q. Long-term N and NPK fertilizations induce significant alterations in microbial compositions and co-occurrence patterns in saturated soil layers. J. Clean. Prod. 2025, 506, 145531. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.