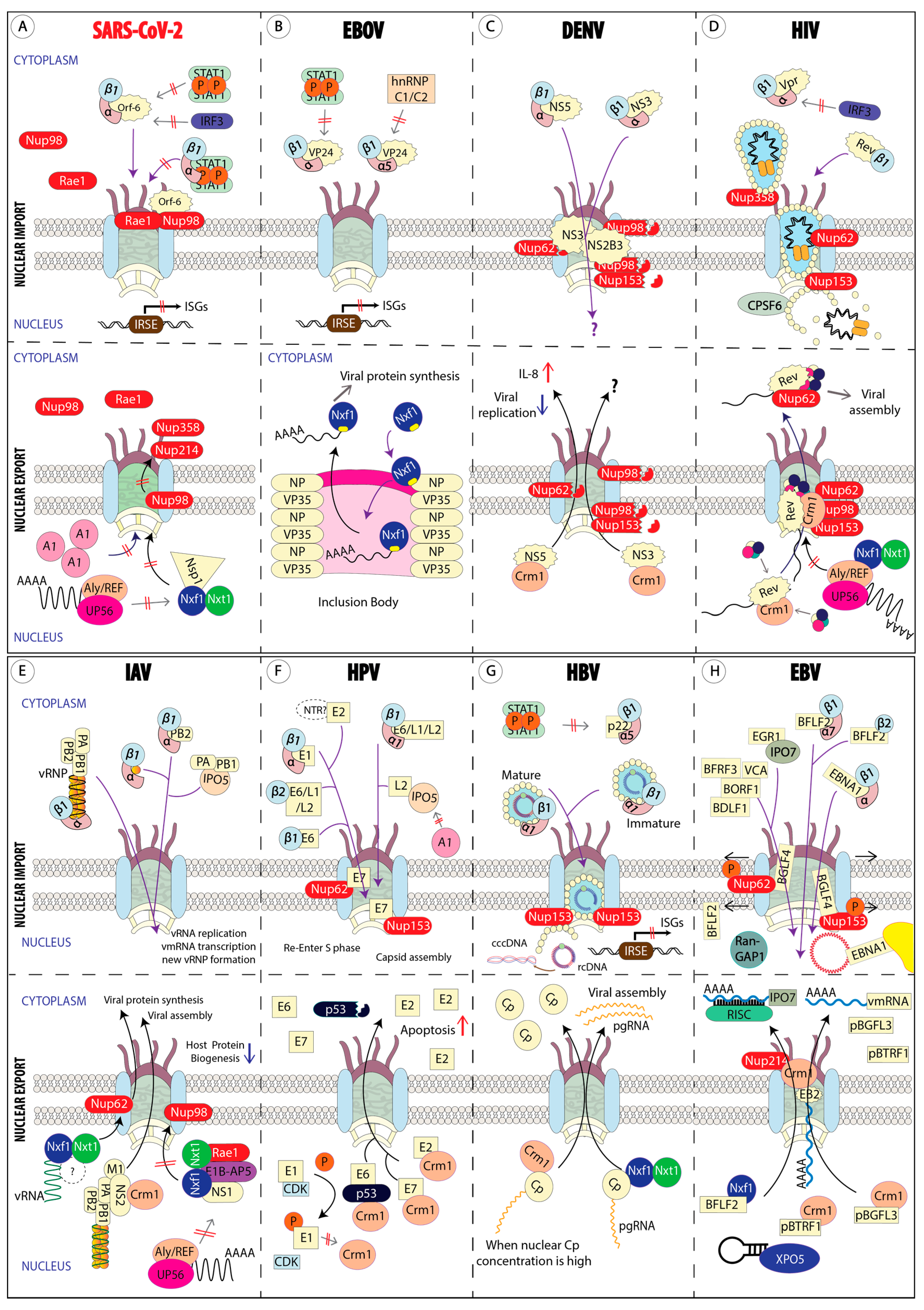
Abstract
The host nucleocytoplasmic trafficking system is often hijacked by viruses to accomplish their replication and to suppress the host immune response. Viruses encode many factors that interact with the host nuclear transport receptors (NTRs) and the nucleoporins of the nuclear pore complex (NPC) to access the host nucleus. In this review, we discuss the viral factors and the host factors involved in the nuclear import and export of viral components. As nucleocytoplasmic shuttling is vital for the replication of many viruses, we also review several drugs that target the host nuclear transport machinery and discuss their feasibility for use in antiviral treatment.
1. Introduction
Despite advancements in science and technology, humans are still plagued by communicable diseases, and the development and production of effective antiviral drugs and vaccines remains challenging. Pathogenic viruses, one major category of infectious agents, have caused not only substantial morbidity and mortality, but also devastating socioeconomic impacts. Within the past century, there have been two particularly severe pandemics, as follows: the 1918 influenza pandemic and the currently ongoing coronavirus disease 2019 (COVID-19) pandemic.
As part of their lifecycle, many viruses hijack the host transcription and translation machinery while evading the host immune responses [1]. The host nucleoplasm and cytoplasm are segregated by a nuclear envelope (NE), a lipid bilayer embedded with numerous nano-gates known as nuclear pore complexes (NPCs) (reviewed in [2]). Nuclear transport is important for mediating numerous cellular activities, such as cell division [3,4], cell metabolism [5,6], gene regulation (review in [7]), and innate immune response activation [8]. As exploitation of the host nuclear transport is important for viral replication, the mechanisms that are used by viruses to hijack the host nuclear trafficking are potential targets for antiviral drugs; such drugs might halt viral genome transcription, viral protein synthesis, and viral assembly. In this review, we focus on the various mechanisms used by viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), to hijack the host nucleocytoplasmic trafficking machinery. We describe various viral factors, and their target host factors, including importins and nucleoporins (Nups). Finally, we discuss the feasibility of using drugs that target the host nuclear transport machinery as antiviral therapies.
2. Fundamentals of Nucleocytoplasmic Trafficking
NPCs are mega-Dalton-sized protein complexes that consist of about thirty different types of Nup. Cryo-electron microscopy and tomography observations indicate that NPCs are formed by an eight-fold symmetric central scaffold, eight cytoplasmic filaments, and eight nucleoplasmic filaments (nuclear basket), with the pores presenting rotational symmetry [9,10,11]. Phenylalanine-glycine (FG) repeats are found in many Nups, and the dynamic hydrophobic interactions among the FG-repeats of Nups in the central scaffold create a cohesive meshwork that can make the NPC become a selective channel [12,13]. Several models to describe the FG-containing Nup interactions, including the hydrogel model [14], virtual gating or polymer brush model [13], reduction in dimensionality model (reviewed in [15]), and forest model [16], have been put forward. Recently, we successfully observed the dynamic behavior of Nup FGs by using high-speed atomic force microscopy (HS-AFM) and proposed a new model of FG-containing Nup interactions called the coweb FG-network model [17]. The functional NPC pore diameter was initially reported as being approximately 26 nm [18], but the value was later corrected to approximately 40 nm [19]. Small molecules (<40–60 kDa) pass freely through NPCs via passive diffusion. Conversely, large molecules, such as proteins, require receptor-mediated nuclear transport to move against a concentration gradient [19].
Receptor-mediated nuclear transport involves cargo proteins, nuclear transport receptors (NTRs), and the guanosine triphosphate-binding nuclear protein Ran (RanGTP) [2]. The types of NTRs required for transport differ depending on the traffic directionality. For nuclear import, selective cargo is recognized by specific NTRs via its nuclear localization signal (NLS) sites [2]. According to reviews authored by Pumroy et al. [20] and Mosammaparast et al. [21], several nuclear import pathways are available, depending on the cargo type. The best understood import pathway is the classical import pathway. It utilizes the heterodimeric importin-α/β1 transport receptor, in which importin-α works as an adaptor protein, bringing its cargo in concert with importin-β to translocate it into the nucleus. There are seven importin-α isoforms expressed in humans, and they can be grouped into the following three subfamilies: α1, α2, and α3 (see Table 1). The conserved N-terminal autoinhibitory importin-β-binding (IBB) domain and the C-terminal Armadillo (Arm) repeats (the NLS-binding sites) of α-importins are important for the nuclear import of the NLS-bearing cargo [22,23,24].

Table 1.
List of the nuclear transport receptors (NTRs) involved in nucleocytoplasmic shuttling.
Most of the identified NLS motifs consist of basic amino acids, such as lysine and arginine, and they are collectively named the classical NLSs (cNLSs). These NLSs are divided into the following two groups: monopartite cNLSs, which have a single stretch of amino acids (e.g., the NLS of the SV40 large T antigen), [25] and bipartite cNLSs, which have two stretches of amino acids separated by a linker region (e.g., the NLS of nucleoplasmin) [26]. There are many cargoes whose NLSs are still unknown. First, the cNLS-bearing cargo binds to the Arm repeats of importin-α [24], and then importin-β binds to the IBB domain of importin-α [27] to form a ternary complex. The IBB domain can be considered as an NLS for α-importins (i.e., a ubiquitous adaptor protein) as well as for other adaptor proteins that carry specific cargo (see Table 1). Once a ternary complex is formed by the cNLS-bearing cargo, importin-α, and importin-β1, the complex docks onto an NPC; subsequently, importin-β1–Nup FG interactions translocate the complex into the nucleus [28].
RanGTP is predominant in the nucleus, whereas the guanosine diphosphate-binding Ran (RanGDP) is abundant in the cytoplasm (reviewed in [29]). This RanGTP/RanGDP gradient is important for the dissociation of the cNLS-bearing cargo–importin-α/β1 ternary complex and the shuttling of importins back to the cytoplasm for the next functional cycle. RanGTP has a high affinity for importin-β1 [30], and the binding of RanGTP to importin-β1 dissociates the ternary complex, thus releasing the cargo in the nucleus [31,32,33]. The RanGTP–importin-β1 complex is then exported to the cytoplasmic site of an NPC. Similarly, the nuclear export of importin-α also requires RanGTP along with a soluble transport factor known as the CAS (Cellular Apoptosis Susceptibility gene) protein or exportin-2 (XPO2) [34]. The hydrolysis of RanGTP to RanGDP is mediated by the RanGTPase-activating protein (RanGAP1) [35,36] and the RanGTP-binding protein (RanBP1) [37,38] at the cytoplasmic site of an NPC, which helps to release importin-α and -β1 back to the cytoplasm. RanGDP is then reimported to the nucleus by nuclear transcription factor 2 (NTF2) [39]. RCC1, the major nucleotide exchange factor of Ran, converts RanGDP back to RanGTP [40].
The nuclear export of cargo requires NTRs (exportins) that read the nuclear export signals (NESs) within cargo [22,24,41]. Chromosomal region maintenance 1 (Crm1), more commonly known as exportin 1 (XPO1), is the major export receptor for roughly 1,000 different leucine-rich-NESs-containing cargoes in human cells [26,42]. For a list of other exportins, please refer to Table 1. A Crm1-mediated nuclear export starts with the formation of a Ran-binding protein 3 (RanBP3)–Crm1–RanGTP–NES-bearing cargo export complex. RanBP3 binds to Crm1 via its FG domains [43]. RanBP3 increases the affinity of the RanBP3–Crm1 complex for RanGTP and the NES-bearing cargo [43]. The quaternary export complex translocates through the NPCs by interacting with Nups and then docks at its terminal docking site, the cytoplasmic Nup214–Nup88 complex [44]. In most cases, RanGAP1 is not soluble but tethered to the NPCs via RanBP2. Cytoplasmic RanBP1 and RanGAP1 mediate RanGTP hydrolysis to disassemble the export complex [45]. A study reported that Crm1 interacts with NPCs in a Nup358/RanBP2-dependent manner [46]. In addition, in vitro experiments have shown that the isolated Ran-binding domain of Nup358 also induces the dissociation of Crm1-export complexes [47]. Collectively, these studies indicate that Crm1-export complexes dissociate after their interaction with soluble RanBP1 and/or Nup358, together with soluble and/or Nup358-associated RanGAP. Free Crm1 then interacts transiently with Nup358 to re-shuttle back to the nucleus [44].
Similar to cargo export, RNA export also requires specific receptors, which depend on the RNA type, in concert with adaptor proteins. These export receptors include the following: Nxf1 or TAP, the main mRNA export receptor [48]; Crm1, for ribosomal (r)RNA [49], small nuclear (sn)RNA [50], and some subsets of messenger (m)RNA [51]; Xpot, for transfer (t)RNA [52,53]; and exportin 5 (XPO5), for micro (mi)RNAs [54]. Several pathways mediate mRNA export, and the Nxf1-dependent pathway is the dominant pathway for exporting bulk mRNAs [55,56,57]. In an Nxf1-dependent mRNA export, mature mRNAs are packaged into messenger ribonucleoprotein particles (mRNPs) that then recruit mRNA export factors (e.g., an Nxf1–Nxt1 heterodimer) [58,59]. The interaction between an Nxf1–Nxt1 heterodimer, Rae1, and Nup98 enables mRNA to be delivered to the cytoplasm [60]. Nxf1 has two binding sites for Nxt1 (amino acid [aa] residues 372–445 [61] and 508–583 [60]) and Rae1 (aa residues 1–60 and 372–445) [60]. Only one Nxf1 binding site is necessary for Nxt1 binding (aa residues 372–445) [61], whereas the Nxf1–Rae1 interaction requires both binding sites [60]. The Nup98-binding site of Nxf1, which is located at the C-terminus of Nxf1 (aa residues 601–619), has the highest affinity for the GLFG-repeat domain of Nup98 [60]. Nup98 binds stably with Rae1, and the Rae1 that is pre-bound to Nup98 will not bind to Nxf1 [60]. Therefore, Nup98 provides a bridging site through which Nxf1 can bind to the adjacent site of Rae1 when both simultaneously interact with Nup98 [60]. This mechanism accomplishes mRNA export in a RanGTP-independent manner. Although Nxf1 has an RNA-binding domain at its C-terminus that can directly bind to the constitutive transport element (CTE) RNA of simian type D retroviruses [62], the Nxf1-Nxt1 heterodimer most frequently requires dedicated export adaptors, including Aly/REF and UAP56 (review in [63]). Likewise, Crm1 does not bind directly to mRNA; instead, it creates complexes with different types of mRNA-binding adaptor proteins, such as HuR [64], Nxf3 [65], and LRPPRC [66], to accomplish mRNA export in a RanGTP-dependent manner. In addition to interacting with mRNA, Crm1 also interacts with the adaptor protein Nmd3 to export the 60S ribosomal subunit [67].
3. Mechanisms of the Host Nuclear Transport Machinery Hijacking by Viruses
The dominant NTRs, including heterodimeric importin-α/β, Crm1, and Nxf1, together with the Nups involved in nuclear transport (Nup358, Nup214, Nup98, and Rae1), are the common targets that viruses hijack for shuttling viral factors between the cytoplasm and the nucleus. The viral replication site determines the purpose of the host nuclear transport subversion by the virus. For example, viruses that replicate in the cytoplasm tend to hijack the host nuclear transport for suppressing the interferon (IFN)-inducing antiviral responses. Conversely, viruses that replicate in the nucleus and assemble their products in the cytoplasm generally exploit the host nuclear transport for much more complicated activities, such as the nuclear import of a viral genome and the nuclear export of a newly synthesized viral genomic RNA to the cytoplasm. Below, we discuss the hijacking mechanisms employed by viruses in various viral families, grouped according to their site of replication, that is, cytoplasm (the Coronaviridae, Filoviridae, Flaviviridae, and Togaviridae families) or nucleus (the Orthomyxoviridae, Retroviridae, Papillomaviridae, Hepadnaviridae, and Herpesviridae families). We summarize these interactions in Table 2 and illustrate them in Figure 1.

Table 2.
Interactions between viral factors and host factors in the hijacking of the host nuclear transport machinery.
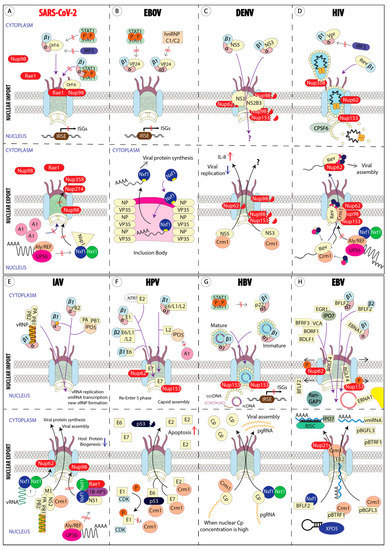
Figure 1.
Interaction between viral factors and host factors to hijack host nuclear transport machinery. (A) SARS-CoV-2 expresses Orf6 to suppress host antiviral response and Nsp1 to halt host protein biogenesis. (B) Ebola virus expresses VP24 to suppress host antiviral response. Ebola virus does not require nuclear export, but it needs the mRNA export factor, Nxf1, to mediate viral mRNA export from the inclusion body. (C) Dengue virus NS3 and NS2B3 degrade host Nups. The effect of the nucleocytoplasmic trafficking of both NS5 and NS3 in the host have yet to be elucidated. Nuclear export of Dengue NS5 is associated with elevated IL-8 production and reduced viral replication without a defined mechanism. (D) HIV capsids (yellowish balls) interact with host Nups (Nup358, Nup62, and Nup153) and CPSF6, then disassemble at an NPC basket to release PIC. HIV expresses Vpr to suppress host antiviral IFN production by blocking the nuclear translocation of activated IRF3. Nuclear import of HIV Rev is directly mediated by importin-β1. Nuclear Rev protein is needed to form a viral ribonucleoprotein (vRNP) transport complex to export vRNP to the cytoplasm via a Crm1-mediated pathway. (E) The Influenza-A virus nucleoprotein (NP, yellowish balls) interacts with importin-α/β1 to enter the nucleus for viral replication. Newly synthesized NPs and RdRp components (PA-PB1 and PB2) hijack host importins to enter the nucleus for new vRNP formation. IAV NS2 facilitates the nuclear export of new vRNP, whereas IAV NS1 blocks host mRNA export by forming an inhibitory complex with Nxf1. (F) HPV high-risk E6 and E7 nuclear imports induce carcinogenesis by reprogramming the cell cycle to an S-phase and promote degradation of p53. In addition, HPV viral capsid proteins (L1 and L2) exploit host importins to translocate to the nucleus for capsid assembly. Nuclear export of E2 of high-risk HPV triggers cellular apoptosis. (G) Mature HBV capsid interacts with importin-α/β1 and Nup153 to uncoat the capsid in the nuclear basket. Immature HBV capsid remains in the NPC channel after it interacts with Nup153. HBV p22 blocks the nuclear translocation of p-STAT1 to suppress host antiviral response. HBV pgRNA export is mediated either by an Nxf1-Nxt1-dependent or a Crm1-dependent pathway, depending on the nuclear Cp concentration. (H) EBV EBNA1 translocates to the nucleus to tether EBV episomal DNA on host chromosome (yellow region). EBV BGLF4 disrupts the NPC structure and promotes nuclear accumulation of RanGAP1 to promote nuclear ingress of viral proteins (VCA, BFRF3, BDLF1, and BORF1) and inhibits host classical importin-α/β1 nuclear import. EBV tegument proteins (pBTRF1 and pBGLF3) bind with Crm1 to translocate to the cytoplasm for viral tegumentation. On the other hand, EBV miRNA is carried to the cytoplasm by host Exportin-5 (XPO5) to suppress importin-7 (IPO7) translation.
3.1. Viruses That Replicate in the Cytoplasm
3.1.1. β-Coronaviruses (SARS-CoV-2, SARS-CoV-1, and MERS-CoV)
Coronaviruses (CoVs; members of the Coronaviridae family) are enveloped, positive-sense, single-stranded (+ss) RNA viruses. There are the following four genera in the Coronaviridae family: Alphacoronavirus (α-CoV), Betacoronavirus (β-CoV), Gammacoronavirus (γ-CoV), and Deltacoronavirus (δ-CoV) (reviewed in [68]). Although most human CoVs (HCoVs) are associated with mild upper respiratory diseases and enteric diseases (reviewed in [69]), the recently emerged zoonotic β-CoVs, such as severe respiratory acute syndrome CoV (SARS-CoV) and Middle East respiratory syndrome CoV (MERS-CoV), have caused severe lower respiratory diseases with high mortality (reviewed in [70]). Furthermore, a highly infectious novel β-CoV, SARS-CoV-2, that was first reported in Wuhan, China in 2019 has caused the greatest pandemic in the 21st century [71]. All CoVs, including SARS-CoV-2, replicate in cytosolic double-membrane vesicles (DMVs) [72,73].
A protein–protein interaction (PPI) analysis of the viral factors and the host factors of SARS-CoV-2 revealed that the viral factors NSP9, NSP15, and Orf6 can interact with the host nuclear transport machinery [74]. The associations suggested by these PPI analyses need to be validated by further experiments. To date, only Orf6 has been shown to interrupt the host nucleocytoplasmic trafficking [75,76,77,78] and to induce an aberrant distribution of Nups (Nup98 and Rae1) [76]. Xia et al. found that Orf6 binds to importin-α1 to block the nuclear translocation of IRF3, resulting in an impaired type I IFN production in the HEK293T cell line [78]. Miorin and colleagues reported that type Ⅰ and type Ⅱ IFNs failed to induce the transcription of IFN-stimulated genes (ISGs) in SARS-CoV-2-infected Vero E6 cells owing to an impediment of the STAT1 and STAT2 nuclear translocation by Orf6 [75]. SARS-CoV-1 Orf6 plays a similar role in antagonizing IFN; it binds to importin-α1 and tethers importin-β1 to the endoplasmic reticulum (ER)/Golgi membrane, sequestrating the NTRs from activated STAT1 (pY-STAT1) [79], thus blocking the nuclear translocation of a pY-STAT1 [79,80]. Miorin and colleagues proposed two mechanisms to explain this blockade. First, Orf6 binds to Nup98 via its C-terminal domain to interrupt the docking of cargo–importin-α5/β1 ternary complexes at the NPCs. The Orf6–Nup98 interaction does not affect the Nup98–Rae1 interaction, which suggests that Orf6 binding to Nup98 targets only the nuclear import pathway. Second, Orf6 competes with STAT1 and STAT2 for α-importins (α5 and α1) to block the STAT1/2 nuclear ingress. Nevertheless, these α-importins may not be the direct key players because their overexpression failed to rescue an Orf6-dependent blockade of green fluorescent protein (GFP)-tagged STAT1 nuclear import. Interestingly, Addetia et al. reported that the SARS-CoV-2 Orf6 interactions with Nup98 and Rae1 were much stronger than those of SARS-CoV-1 Orf6, suggesting the strong IFN antagonism triggered by SARS-CoV-2 contributes to the high prevalence of asymptomatic cases of SARS-CoV-2 infection [77].
We found that, along with being able to impair nuclear import, SARS-CoV-2 Orf6 can also block nuclear export [76]. Our results show mislocalizations of Nup98 and Rae1 in Orf6-overexpressing cells [76]. In addition, nuclear accumulation of the nuclear RNA-binding protein hnRNPA1 was observed in Orf6-overexpressing cells. For mRNA export, hnRNPA1 is needed [81,82], and the saturation of hnRNPA1 in the nucleus inhibits mRNA export [81]. These findings, that is, the aberrant localization of Nup98 and Rae1 and the nuclear saturation of hnRNPA1, suggest that the Orf6–Nup98–Rae1 interaction can block mRNA export. Addetia et al. later found that SARS-CoV-2-infected cells exhibited a nuclear mRNA accumulation that could be mediated by Orf6 [77]. Recently, Zhang et al. reported that SARS-CoV-2 non-structural protein 1 (Nsp1) halted the host gene expression by blocking an Nxf1–Nxt1-mediated mRNA export [83]. Mechanistically, Nsp1 did not impair the RNA-binding ability of Nxf1; rather, it interrupted Nxf1 interacting with mRNA export adaptors, including Aly/REF and UAP56. Furthermore, Nsp1 reduced the interactions between Nxf1 and key Nups for nuclear export (Nup358, Nup214, and Nup98). SARS-CoV-1 Nsp1 was previously found to disrupt the localization of Nup93 (from the NE to the nucleoplasm) without affecting the NPC structure [84]. Furthermore, SARS-CoV-1 Nsp1 specifically promotes the cytoplasmic accumulation of the nuclear RNA-binding protein nucleolin with unknown consequences. These findings may indicate a role for SARS-CoV-1 Nsp1 in disrupting the host mRNA export [84].
SARS-CoV-1 Orf9b lacks an NLS and thus is localized mainly in the cytoplasm [85]. However, a small amount of Orf9b enters the nucleus passively, and nuclear Orf9b activates the host caspase-3-mediated apoptosis. As Orf9b degradation occurs in the cytoplasm, the nuclear export of Orf9b can protect the host and sustain viral replication [85]. Orf9b possesses an NES at aa residues 46–54 (LRLGSQLSL) that allows it to bind to Crm1 [85]. MERS-CoV Orf4b possesses an NLS (aa residues 2–38), and its nuclear localization is important for inhibiting IRF3- and IRF7-induced IFN-β production [86]. Canton et al. later demonstrated that MERS-CoV Orf4b has a strong affinity for importin-α3, which blocks the NF-κB p65 subunit from entering the nucleus [87].
3.1.2. Ebola Virus (Zaire Ebolavirus)
Ebola virus (EBOV), a member of the Filoviridae family, is a filamentous, enveloped, single-stranded, negative-sense (−ss) RNA virus [88]. At present, there are six recognized EBOV species. The Zaire ebolavirus species has caused two outbreaks in Africa with 40 and 66% fatality, respectively, [89]. As EBOVs replicate in the cytoplasm, they hijack the host nuclear transport to suppress IFN-mediated antiviral responses. The EBOV VP24 protein interacts with importin-α5, -α6, and -α7 [90], but does not interact with importin-α1, -α3, or -α4 [91]. A more recent study found that EBOV VP24 recognizes a non-classical NLS-binding site in importin-α6 [92]. These interactions allow EBOV to suppress the host IFN signaling by preventing the nuclear translocation of pY-STAT1 [91]. The VP24–importin-α interaction also inhibits IFN-λ production [93]. VP24 can directly bind to the inactive form of STAT1, unphosphorylated STAT1 (U-STAT1) [94]. The nuclear import of U-STAT1 is mediated by Nup153 and Nup214, independently of importins [95]. Within the nucleus, U-STAT1 activates and prolongs the expression of a set of IFN-induced genes that are distinct from those mediated by pY-STAT1 [96]. The VP24–importin-α5 interaction causes a cytoplasmic accumulation of the nuclear protein hnRNP C1/C2 [97]. During mitosis, hnRNP C1/C2 is exported to the cytoplasm for internal ribosomal entry site (IRES)-dependent c-myc translation [98]. Several viruses similarly re-localize hnRNP C1/C2 to the cytoplasm, either for viral replication or IRES-dependent viral protein translation [99,100,101,102], which suggests that VP24-dependent hnRNP C1/C2 cytoplasmic accumulation is essential for EBOV replication [97]. Together, nucleoproteins (NPs) and VP35 form EBOV inclusion bodies (IBs) for viral replication [103]. A study conducted by Gabriel et al. showed that importin-α7 is also involved in IB formation [104]. In addition to creating new copies of EBOV genomic RNA, sub-genomic EBOV RNAs (viral mRNAs) are also produced for viral protein synthesis. The EBOV NP recruits Nxf1 by interacting with the RNA-binding domain of Nxf1. In the presence of mRNA, the binding preference of the Nxf1 RNA-binding domain shifts from NPs to mRNA, and then it delivers mRNA to the cytoplasm for translation [105].
3.1.3. Dengue Virus (DENV) and Zika Virus (ZIKV)
Flaviviruses (members of the Flaviviridae family) are small, enveloped +ssRNA viruses that replicate in vesicle packets located at the ER [106,107]. The flavivirus Dengue virus (DENV) causes mosquito-borne dengue fever in tropical and sub-tropical countries. Two nonstructural proteins (NSs), the DENV NS3 helicase and the DENV NS5 RNA-dependent RNA polymerase (RdRp), form a complex to mediate the viral replication [108,109]. NS5 promotes STAT2 degradation to suppress the host IFN-mediated antiviral responses [110]. An early study reported that NS5 has a bipartite NLS composed of an importin-β1-binding site (βNLS) [111] and a heterodimeric importin-α/β1-binding site (cNLS) [112]. NS5 nuclear import is predominantly mediated by importin-α/β1 receptors [112]. However, later studies suggested that these mapped NLSs are not accessible to the host importins [113]. The subcellular localization of NS5 varies among the four DENV serotypes (DENV 1, 2, 3, and 4); the NS5 proteins of DENV2 and DENV3 reside in the nucleus, but the NS5 proteins of DENV1 and DENV4 are located in the cytoplasm [114,115]. A new monopartite NLS of NS5 has been identified at its C-terminus [115]. An in vitro assay revealed that the NS5 proteins of DENV2 and DENV3 have similar and strong affinities for importin-α2, whereas those of DENV1 and DENV4 have weak affinities for importin-α2 [115]. The NES in NS5 allows for Crm1-dependent NS5 nuclear egress, and this egression correlates with elevated IL-8 production and impaired viral replication, the mechanism of which has not yet been defined [116]. An NE abnormality together with deregulated NPC components were found in DENV-infected cells [117]. NS3 and its cofactor, NS2B3, disrupt the NPC integrity by inducing the proteolytic degradation of FG-Nups, including Nup62, Nup153, and Nup98 [117]. Palacios et al. reported that NS3 has a putative NLS and a putative NES. NS3 was localized within the nucleus during the early stages of infection, whereas it remained predominantly in the cytoplasm during the later stages of infection [118].
Another member of the Flaviviridae family, Zika virus (ZIKV), has posed serious health concerns owing to the rapid increase in cases of neonatal microcephaly linked to ZIKV-infected mothers in Brazil [119,120]. Similar to DENV, ZIKV also exploits the host NTRs and Nups during infection via its NS3 helicase and NS5 RdRp. A study conducted by De Jesús-González et al. showed that ZIKV NS3, in concert with NS2B3, promotes the proteolytic degradation of several FG-containing Nups, including TPR, Nup153, and Nup98, and also disrupts the NE structure [117]. ZIKV NS5, similar to DENV NS5, has a bipartite NLS (a βNLS and a cNLS), which is recognized by importin-α7 [121]. The nuclear import of ZIKV NS5 protects it from cytoplasmic degradation, and this strategy helps to sustain viral replication in the host cells [122]. The cNLS was initially thought to be the primary site of NTR binding [123], but a later study revealed that both the βNLS and the cNLS are required for NS5 nuclear import [122]. Nuclear NS5 sequesters various α-importins (α1, α3, and α4) in nuclear bodies [123]. Intriguingly, the NS5 accumulated in nuclear bodies was incorporated with STAT1 in a glioblastoma cell line (LN229) but not in a hepatocellular carcinoma cell line (Huh-7), suggesting a role for NS5 in the tissue-specific activation of inflammatory responses [123]. Unlike the NS5 protein in Japanese encephalitis virus (JEV), which competes with the host IRF3 and NF-κB for α-importins (α1, α3, and α4) [124], ZIKV NS5 inhibits the activation of TANK-binding kinase 1 to prevent IRF3 activation [125], a strategy used to inhibit the IFN production by infected cells. Furthermore, ZIKV NS2A promotes the chaperone-mediated autophagy (CMA) of importin-α1 [126], possibly in an effort to suppress the host antiviral response.
3.1.4. Chikungunya Virus (CHIKV)
Chikungunya virus (CHIKV), a member of the Togaviridae family, is an enveloped +ssRNA virus that replicates in the host cytoplasm [127]. A CHIKV infection is associated with chronic inflammatory arthritis and other musculoskeletal diseases [128]. The CHIKV capsid protein (CP) has been reported to have two NESs and one NLS [129,130]. Thomas and colleagues demonstrated that the CP bound specifically to the C-terminal NLS-binding site of importin-α3 for its nuclear translocation [129]. The CP binds to Crm1 via its Crm1-mediated NES, which was mapped to a leucine-rich region between aa residues 143 and 155 [129]. Thus, importin-α3 and Crm1 are the host factors targeted by CHIKV in its disruption of the host nucleocytoplasmic trafficking [129]. A mutation of the CHIKV CP’s NES near the N-terminus (aa residues 44–53) caused the retention of viral CPs in the nucleus and also blocked the host nuclear import system for unknown reasons [130]. CHIKV nsP2 inhibits the host IFN-induced antiviral response [131,132,133,134]. Although nsP2 lacks an NLS (reviewed in [135]), the nuclear import of nsP2 is necessary for suppressing the host antiviral response [134]. Interestingly, similar to that of DENV NS3, the nuclear localization of CHIKV nsP2 also occurs temporarily during early infection, after which this protein resides in the cytoplasm [136]. IFN antagonism by nsP2 can be achieved by several mechanisms, including via a reduction in the cGAS level by a global translational inhibition [131,136], an inhibition of STAT1 activation and/or block of pY-STAT1 nuclear import [133], and a promotion of STAT1 nuclear export [132].
3.2. Viruses That Replicate in the Nucleus
3.2.1. Human Immunodeficiency Virus (HIV)
Human immunodeficiency virus (HIV), a member of the Retroviridae family, is an enveloped +ssRNA virus that was first recognized in 1981 as the causative agent of a new disease affecting T lymphocytes [137]. HIV-1 is more virulent and infectious compared to HIV-2; thus, HIV-1 is the leading cause of acquired immunodeficiency syndrome (AIDS) in the ongoing AIDS pandemic [138]. Upon viral entry, the viral replication complex undergoes reverse transcription followed by integration to form a pre-integration complex (PIC) before entering the nucleus (reviewed in [139]). Mutations in HIV proteins and the silencing of some NPCs can change the HIV integration pattern, which suggests that nuclear import is closely associated with the selection of a viral cDNA integration site for integration into the host genome (review in [140]).
The HIV-1 viral capsid was initially postulated to rapidly disassemble upon viral entry, releasing PICs into the cytosol, from which they subsequently translocate to the nucleus through NPCs [141]. A group of RNA interference (RNAi) screening studies have identified several Nups that are required for a HIV-1 infection, which include Nup98, Nup85, Nup133, Nup107, Nup160, Nup153, Nup214, Nup358, Nup155, Crm1, and the nuclear import receptor transportin 3 (TNPO3) [142,143,144]. In addition, a HIV-1 infection induces a downregulation in Nup50 and an upregulation in Nup62 for currently unknown reasons [145]. Some HIV-1 proteins, such as integrase (IN) [146], the viral protein R (Vpr) [147], and the matrix protein (MA) [148], contain NLSs. Notably, the width of the broad end of the HIV-1 capsid is approximately 60 nm [149], whereas the NPC pore size has long been thought to be only approximately 40 nm wide [19]. Collectively, these findings suggested that capsid disassembly is a prerequisite for PIC nuclear import.
More recent findings showing that the HIV-1 capsid remained assembled for at least the reverse transcription process [150] and that capsid uncoating can occur in the nucleus [151,152,153] have challenged the model described above. With a combination of different imaging tools, including cryo-electron tomography (cryo-ET) and subtomogram averaging, Zila and colleagues have visualized the entry of a whole HIV-1 capsid into a host nucleus through an NPC [154]. The diameter of an NPC measured in intact human cells (i.e., in a transporting state) was in fact larger than the diameter of an NPC measured in an isolated nuclear envelope (i.e., in a constricted state). The diameter of an NPC in the transporting state is larger than the broad end of the HIV-1 capsid; therefore, it can accommodate the import of a HIV-1 capsid into the nucleus. The team proposed a three-step process for the nuclear import of a HIV-1 capsid, during which the intact capsid interacts with different Nups in each stage. First, the HIV-1 capsids travel along microtubules to dock onto NPCs and interact with the FG-repeats and cyclophilin (Cyp) domain of Nup358 at the NPC cytoplasmic sites. Second, the intact capsids move deep into the central channels of the NPCs, which contain high local concentrations of FG-containing Nups within the Nup62 complex. Last, the capsids bind to Nup153 and cleavage and polyadenylation specificity factor subunit 6 (CPSF6). The HIV capsid–Nup153–CPSF6 interaction triggers capsid disassembly, which releases PICs into the nucleoplasm.
HIV Vpr suppresses the host antiviral response by interacting with α-importins (preferentially α5, but also, to a lesser extent, α1 and α4) to inhibit the IRF3 activation and to block the nuclear import of IRF3 and NfκB [155]. HIV Vpr has an N-terminal NLS and a C-terminal NLS [156]. Its C-terminal NLS was initially thought to be non-functional [156], but later studies have shown that it binds α-importins and mediates nuclear import without importin-β1 [157,158].
HIV-1 Rev has an arginine-rich NLS site in its N-terminal domain and a leucine-rich NES sequence in its C-terminal domain (reviewed in [159]). HIV-1 Rev enters the nucleus without binding RNA because only the RNA-free form of Rev can bind to importin-β1 for nuclear import [160]. The Rev–importin-β1 interaction is highly specific and can be blocked by importin-α5 [160]. The Rev NES is required for the nuclear export of unspliced viral RNA (vRNA). Rev binds to vRNA via the Rev response element (RRE) (reviewed in [161]). Rev contains an NES that allows it to bind to Crm1 and subsequently bring vRNA, in the form of a viral ribonucleoprotein (vRNP) transport complex, out into the cytoplasm in a RanGTP-dependent manner [162]. Several studies have suggested that Nup214, Nup153, Nup98, and Nup62 interact indirectly with vRNP through different Rev co-factors [163,164,165]. For example, the Rev–vRNP complex disturbs NPC integrity and causes a Nup62 cytoplasmic localization and subsequent encapsidation into a progeny virus [166]. A downregulation in Nup62 induced a vRNA nuclear accumulation, suggesting the importance of Nup62 in vRNA export [166]. The host Nxf1-dependent RNA export always involves active pre-mRNA splicing to prevent aberrant protein translation. Rev suppresses Nxf1-mediated vRNA export to secure a viral gene expression in the host [167]. McCauley et al. reported that an HIV-1 provirus used Crm1 and Rev to export its unspliced HIV for translation, which triggered chronic inflammation in patients with HIV who were receiving anti-retroviral therapy [168].
3.2.2. Influenza A Virus (IAV)
Influenza A virus (IAV), a member of the Orthomyxoviridae family, is an enveloped, segmented −ssRNA virus that causes an epidemic respiratory disease (reviewed in [169]). IAV genomic RNAs are packed together with a viral nucleoprotein (NP) and a heterotrimeric RNA-dependent RNA polymerase (RdRp) complex (composed of PA, PB1, and PB2) into a rod-like vRNP (reviewed in [169]). Upon viral entry, free vRNPs translocate to the nucleus for viral RNA transcription and replication. The IAV NP and three components of the RdRp complex possess at least one NLS [170]. An IAV NP has two NLSs, a non-classical NLS in the N-terminus [171] and a classical bipartite NLS in the middle [172], and these are sufficient for vRNP nuclear import [170]. A mutation of the bipartite cNLS of the NP did not block its nuclear import, which suggests that the non-classical NLS of the NP is the predominant site [173] for its interaction with importins α5 and α7 [169]. Therefore, the non-classical NLS of an NP is essential for IAV replication [173]. Donchet et al. reported the binding affinity of an NP to different α-importins; an NP had the highest affinity for importin-α7 and the lowest affinity for importin-α1 [174].
IAV PB2 has a classical bipartite NLS [175], enabling its binding to the host α-importins (α1, α5, and α7) [176,177], preferentially importin-α7 [178]. Interestingly, both importins α1 and α7 act as positive regulators for PB2 polymerase activity, whereas importin-α3 acts as a negative regulator [177]. Highly pathogenic avian IAVs (HPAIVs) with human-like PB2 downregulate the expression of importin-α3, the main NTR for NF-κB, to suppress the host antiviral response [179]. The importin-β family member importin-5 (IPO5), also known as RanBP5, not only forms an import complex with PA and PB1 but also regulates the PA–PB1 complex for vRNA binding [180,181].
The Crm1-mediated nuclear export of newly synthesized IAV vRNPs is crucial for new viral synthesis. IAV vRNP, M1, and NES-bearing NS2 form a complex for vRNP export [182,183]. IAV NS1 exerts a negative regulatory effect on the host Nxf1-dependent mRNA export by downregulating Nup98 expression and forms an inhibitory complex with several of the host mRNA export factors (Nxf1, Nxt1, Rae1, and E1B-AP5) [184]. IAV NS1 binds at the FG-repeat binding site of Nxf1, where it interacts with the FG-domain of Nup98 during mRNA export [185]. In addition to disrupting the host mRNA export, IAV NS1 also interrupts the host mRNA processing, which it achieves by interacting with CPSF30 [186] and PABⅡ [187]. Collectively, these activities suppress the host protein biogenesis, particularly the factors required for an effective antiviral response [185]. Interestingly, in severe cases of influenza, a reduction in the protectin D1 (PD1) level enables IAV vRNA to bind directly to Nxf1, which then interacts with Nup62 for nuclear export. This mechanism does not require other mRNA export factors (e.g., Nxt1 or Crm1) or FG-containing Nups (e.g., Nup98 or Nup214) [188].
3.2.3. Human Papilloma Virus (HPV)
Human papilloma virus (HPV), a member of the Papillomaviridae family, is a small, icosahedral, non-enveloped, double-stranded (ds)DNA virus [189]. In 1995, the International Agency for Research (IARC) classified HPV16 and HPV18 as human carcinogens because infection with these viruses increases the risk of developing cervical cancer [190]. HPV replicates in the host nucleus, and the entry of its viral genome into the host nucleus requires NE breakdown (during mitosis) rather than passage through an NPC [191,192].
The nucleocytoplasmic shuttling of E6 and E7, two oncoproteins found in high-risk HPVs, promotes carcinogenesis in HPV-infected cells. Only high-risk HPV E6 (E6high) can translocate to the nucleus [193] because its C-terminal contains three NLSs [193,194,195] that interact with importin-α1/β1, importin-β1, and importin-β2 [194]. Low-risk HPV E6 (E6low) predominantly resides in the cytoplasm, but E6low acquires nuclear import activity when it is conjugated with an E6high NLS [193]. The nuclear import of E6high requires RanGDP, but GTP hydrolysis is not necessary [194]. Nuclear E6high is critical for E6-mediated p53 degradation, independent of MDM2 [196]. E6high mediates the polyubiquitination of p53, making it susceptible to proteosome degradation [197]. The E6high–p53 interaction promotes p53 export, and the p53 NES is required for a Crm1-mediated nuclear export [197]. The nuclear import of E6high is compulsory for E6high-mediated p53 degradation; however, the p53 degradation can occur in both the nucleus and the cytoplasm [197].
To support viral DNA amplification, high-risk HPV E7 (E7high) must travel to the nucleus to hijack the host cell-cycle machinery, driving the host cell to re-enter the S-phase (reviewed in [198]). HPV16 E7 was initially thought to lack an NLS, and its nuclear import is Ran-dependent but independent of classical importin-α/β and importin-β2 receptors [199]. Knapp et al. later identified two NLSs in the N-terminal domain and CR3 domain of the HPV16 E7 protein [200]. Eberhard et al. found an explanation for the ability of the HPV16 E7 protein to undergo importin-independent nuclear import, despite having NLSs [201]. The zinc-binding domain within the E7 CR3 domain contains a hydrophobic patch (65LRLCV69) that enables E7 to accomplish its nuclear import by interacting with the FG-domain of Nup62 via a hydrophobic interaction [201]. Furthermore, this hydrophobic patch also facilitates the interaction of HPV16 E7 with Nup153 [202]. The hydrophobic interaction between HPV E7 and FG-containing Nups is conserved in other HPV serotypes (HPV8 and HPV11) [202,203]. A functional NES has been identified in HPV16 E7, which suggests that E7 nuclear egress is Crm1-dependent [200].
HPV encodes the E1 DNA helicase and the E2 original recognition protein and uses them to hijack the host DNA replication machinery for viral replication [204]. Therefore, the nuclear localization of E1 and E2 are important for efficient viral replication. The E1 proteins of several HPV serotypes (HPV11, HPV31, and HPV16) have bipartite NLSs [205], and these NLSs behave in a similar way to the bipartite NLS of the bovine HPV E1 protein [206], which can interact with multiple importins α (α1, α3, and α5) [207]. In addition, the phosphorylation of HPV E1 by the host ERK and/or JNK is needed for its nuclear import [205,206]. Regarding the HPV E1 nuclear export, this protein has a functional NES site, suggesting it undergoes a Crm1-dependent export [204,208]. This NES has a cyclin-dependent kinase (CDK) phosphorylation site, and NES phosphorylation by CDK causes HPV E1 to be retained in the nucleus, which is essential for viral replication [206,208].
The subcellular localizations of E2 proteins from low-risk (HPV11 and HPV6) and high-risk (HPV16 and HPV18) HPVs differ; low-risk HPV E2 is predominantly localized in the nucleus, whereas high-risk HPV E2 can be found in both the cytoplasm and the nucleus [209]. The NLS of the HPV11 E2 protein, but not those of the HPV16 and HPV18 E2 proteins, was found to have a dominant function [209]. Interestingly, cNLSs are found in high-risk HPV E2 but not in low-risk HPV E2; however, the NTRs for high-risk HPV E2 are still unknown [210]. The NES in high-risk HPV E2 enables the nucleocytoplasmic shuttling of this protein. Cytoplasmic accumulation of the high-risk HPV E2 proteins promotes caspase-8-mediated cell apoptosis [209].
The HPV L1 major capsid protein (L1) and L2 minor capsid protein (L2) are delivered to the nucleus for virion production. HPV16 L1 possesses both a monopartite and a bipartite NLS, and its nuclear import requires importin-α1/β1, RanGDP, and free GTP, but occurs independently of GTP hydrolysis [211]. The L1 protein also binds importin-β2, but RanGTP is unable to dissociate the complex [211]. This interaction inhibits the importin-β2-mediated nuclear import of hnRNP A1 [211]. An L2 protein nuclear import is required for HPV viral assembly [212]. For nuclear import, the HPV16 L2 protein can interact with importin-β2, importin-5, or the importin-α1/importin-β1 heterodimer. An NES has been identified in the L2 protein, but its function remains unclear.
3.2.4. Hepatitis B Virus (HBV)
Hepatitis B virus (HBV), a member of the Hepadnaviridae family, is a small, enveloped dsDNA virus that propagates in hepatocytes [213]. Acute infection with HBV causes acute hepatitis, whereas chronic infection with HBV increases the risk of hepatocellular carcinoma (HCC) (reviewed in [214]). The HBV capsid disassembles in the nucleus, releasing relaxed circular DNA (rc)DNA, which is subsequently repaired by the host DNA repair machinery to form a covalently closed circular DNA, (ccc)DNA. The HBV cccDNA serves as a template for viral RNA replication (reviewed in [215]). Phosphorylation of the C-terminal HBV core or the capsid protein (Cp) exposes the Cp NLS, allowing the Cp to recruit classical importin-α/β1 receptors to mediate the nuclear import of the HBV capsid [216,217]. In addition to phosphorylation, the completion of the (+) DNA and removal of viral DNA polymerase from rcDNA triggers a conformational change of the Cp, exposing the C-terminal NLS site [218]. During its translocation through an NPC, RanGTP triggers the dissociation of the Cp-importin-α/β1 ternary complex, which allows for an interaction between the Cp and Nup153 [219]. Notably, Nup153 is the sole FG-containing Nup that interacts with the Cp, which suggests that this interaction is not mediated by a hydrophobic interaction [219]. Finally, mature HBV capsids disintegrate in a Ran-independent manner [217], releasing viral rcDNA; capsid disassembly is halted in immature capsids [219].
The Cp has the following four arginine-rich domains (ARDs): ARD Ⅰ and Ⅲ each behave in a similar way to an NLS, and ARD Ⅱ and Ⅳ each behave in a similar way to an NES [220]. The Cp NES is needed for an Nxf1-dependent viral pre-genomic RNA (pgRNA) export [220]. The nuclear export of pgRNA is a critical step for new virion synthesis. The subcellular localization of the Cp is rather complex, depending on both the intrinsic factors (NLS and NES) and the extrinsic factors (importins, Nxf1, and cellular kinase) [220]. A recent study showed that increasing the nuclear Cp concentration induces Cp export in a Crm1-dependent manner, which suggests that the Cp concentration may manipulate different export signals [221]. An empty Cp can interact with importin-β1 independent of importin-α via its IBB domain, but the exact biological function of this interaction is yet to be elucidated [222]. Mitra and colleagues reported that the cytosolic HBV e antigen (HBeAg), also known as the precore protein intermediate (p22), interacts with importin-α5 through its C-terminal ARD. This interaction blocks the pY-STAT1 nuclear import and subsequently suppresses the host IFN response [223].
3.2.5. Herpes Simplex Virus Type-1, Human Cytomegalovirus, and Epstein–Barr Virus
Herpes simplex virus type-1 (HSV-1) is a dsDNA α-herpesvirus that causes cold sores in human beings [224]. After internalization, the viral capsid travels along microtubules to reach the nucleus to release a viral genome into the nucleoplasm for transcription and translation [225]. The tegument proteins VP1/2 [226] and pUL25 [225] interact with Nup358 and Nup214 to dock the viral capsid on an NPC. VP1/2 has an efficient NLS at its N-terminal, but the protein is mainly located in the cytoplasm [227]. The NLS of VP1/2 is specific to importin-β1 rather than importins α [227,228], and VP1/2-importin-β1 interaction is important for directing the viral capsid to dock on an NPC [227]. Using mouse embryonic fibroblast (MEF) cells, Döhner et al. comprehensively demonstrated the distinct importin-α isoforms (α1, α3 and α4) in mediating HSV-1 gene expression, nuclear localization of viral proteins, capsid assembly, and capsid egress [229]. Importins are not needed for the NPC docking of an HSV-1 capsid, for the nuclear import of incoming HSV-1 genomes, or for the nuclear translocation of the HSV-1 VP16 protein. Importins α indirectly regulate the HSV-1 protein expression, and within this regulation importin-α1 is facilitative but importin-α4 is restrictive. Importin-α1 and Importin-α3 orchestrate the nuclear localization of immediate-early (ICP4 and ICP0) and early (ICP8 and pUL42) HSV-1 proteins. The HSV-1 DNA polymerase subunits pUL30 and pUL42 utilize several nuclear import mechanisms. HSV-1 pUL30 has a non-canonical and a classical bipartite NLS, and interacts with importin-α5, but interactions with other importin-α isoforms remain unknown [230,231,232]. HSV-1 pUL42 has a bipartite NLS, and the NLS binds mainly to importin-α7 and, to a lesser extent, to importin-α1, but not to importin-α3 [233]. Interestingly, the mutation of the NLS in either pUL30 or pUL42 does not impede the nuclear import of both proteins under the holoenzyme form [233]. The holoenzyme only resides in cytosol if the NLSs in both pUL30 and pUL42 are mutated [233]. Importin-α1 is crucial for an HSV-1 infection because importin-α1 is required for efficient capsid assembly and egress to make new virions in cytosol. The team also illustrated that silencing either importin-α1 or importin-α3 is sufficient to suppress HSV-1 gene expression in terminally differentiated cells, neurons for example, but HSV-1 gene expression remains unperturbed in MEF cells [229]. MEF is not a terminally differentiated cell type [234]. Such a discrepancy indicates that importins α repertoire in MEF is sufficient to compensate for the absence of importin-α1 or importin-α3 to sustain HSV-1 gene expression.
HSV-1 ICP27 is an immediate-early protein that is required for enhancing HSV-1 gene expression and for exporting intronless HSV-1 mRNAs (review in [235,236]). Different conformations of ICP27 confer different specificities/preferences for nuclear export. Viral mRNA-bound ICP27 interacts with Aly/REF to recruit Nxf1 to export intronless HSV-1 mRNA for translation [237]. Nonetheless, Aly/REF knockdown does not significantly dampen viral mRNA export [238], suggesting that ICP27 could utilize other host export factors to compensate for Aly/REF. Free ICP27 does not require both Nxf1 and Crm1 for its export [239]. Instead, its N and C termini interact with Nup62 to enable ICP27 shuttles between the cytoplasm and the nucleus. The nuclear import of ICP27 inhibits both classical, importin-α/β1-dependent, and transportin-dependent nuclear import [240]. Beside the main export receptor (Nxf1), a recent finding reveals that the HSV-1 integral protein, glycoprotein M (gM), binds to exportin-6 (XPO6) for nuclear export to the trans-Golgi network (TGN) [241].
Human cytomegalovirus (HCMV) is a ds-DNA β-herpesvirus that is associated with mononucleosis syndrome (review in [242]), venous thromboembolism (VTE) (review in [242]), and cytomegalovirus encephalitis in immunocompromised patients (review in [243]). HCMV pUL97 is a multifunctional protein kinase that phosphorylates CDK1 and modifies the host G2/M cell cycle checkpoint regulators to promote viral replication [244]. There are two isoforms of pUL97, large and small isoforms. The large isoform has two bipartite NLSs (NLS1 and NLS2), whereas the small isoform only has NLS2, located at their N-terminals [245]. Therefore, the large isoform has a higher nuclear localization efficiency compared to the small isoform [245]. Only importin-α1 has been shown to interact with pUL97, and other importin-α isoforms have not been tested [245]. HCMV pUL79 is an elongation factor of RNA polymerase II for viral gene transcription during the late stages of HCMV infection [246]. HCMV pUL79 possesses a hydrophobic PY-NLS, which enables it to translocate to the nucleus through an importin-β2-mediated pathway [247]. The multifunctional protein, HMCV pUL84, is believed to initiate lytic viral DNA synthesis [248]. Lischka et al. performed in vitro transport assays and found that the nuclear import of pUL84 relies on the classical importin-mediated import pathway [249]. Although pUL84 has a putative NLS, the NLS does not have the NLS activity [249]. Instead, a large domain with 282 amino acids is needed for an importin-α–pUL84 interaction [249]. A team has shown that pUL84 interacted with several importin-α isoforms, including α1, α 3, α 4, and α5 [249]. Interestingly, the pUL84–importin-α interaction domain also contains two leucin-rich NESs, and this region allows the nuclear export of pUL84 via the Crm1-dependent pathway [250]. Gao and his coworkers conducted RNA pulldown assays to reveal pUL84′s viral mRNA export activity [248]. The team reported that pUL84 interacted with a viral-encoded transcript known as IRS1 and mediated the cytoplasmic localization of an IRS1 transcript [248].
Similar to HSV-1 ICP27, HCMV encodes the pUL69 regulatory protein to mediate a viral mRNA export. The arginine rich RNA-binding domain and the DExD/H-box helicase UAP56-binding motif in pUL69 [251] allows pUL69 to interact with the mRNA export factor UAP56/URH49 [252], transcription elongation factor hSPT6, and intronless viral mRNA to form a messenger ribonucleoprotein (mRNP) for an Nxf1-Nxt1-dependent mRNA export (reviewed in [253]). An NES is required for the nuclear export of free pUL69 but independent of Crm1, and the exact export mechanism remains elusive [254]. Likewise, in silico analysis failed to determine a classical NLS within pUL69, the NTRs associated with pUL69 nuclear import are also unknown [252]. CDK9 phosphorylation of pUL69 is crucial for a pUL69-mediated viral mRNA export because CDK inhibition triggers the nuclear accumulation of pUL69 and suppresses the mRNA export activity of pUL69 [255].
The Herpesviridae family member Epstein–Barr virus (EBV), also known as human herpesvirus 4, is a dsDNA γ-herpesvirus that is associated with the development of three types of B-cell lymphoma (Burkitt’s lymphoma, Hodgkin’s lymphoma, and diffuse large B-cell lymphoma) and of nasopharyngeal carcinoma (NPC) (reviewed in [256]). Viral replication, which occurs in the host nucleus, has two different replication patterns, latent replication (during B-cell proliferation) and lytic replication (virion production) (reviewed in [256]). In latent replication, EBV nuclear antigen 1 (EBNA1) is the only viral protein needed in the host nucleus for viral DNA retention [257]. EBNA1 has an NLS (379KRPRSPSS386) that binds to importin-α1 and importin-α5 for its nuclear import [257]. In lytic replication, several viral proteins, including the DNA-replicating enzymes BSLF1, BBLF2/3, BBLF4, and a major capsid protein (VCA), are translocated to the nucleus for viral DNA replication and nucleocapsid assembly [258]. However, these viral proteins do not possess canonical NLSs. To circumvent this limitation, EBV expresses the BGLF4 protein kinase to facilitate nuclear translocation, probably by inducing the nuclear accumulation of RanGAP1 to inhibit the nuclear import of cNLS-bearing cargoes, phosphorylating FG-containing Nups (Nup62 and Nup153) to dilate the NPC, and inducing microtubule reorganization to change the nuclear shape [258]. The BGLF4 homolog proteins Herpes Simplex 1 (HSV-1) UL13 (α-herpesvirus), human Cytomegalovirus (HCMV) UL97 (β-herpesvirus), Kaposi’s sarcoma-associated herpesvirus (KSHV) ORF36 (γ-herpesvirus), and Murine Gammaherpesvirus 68 (MHV68) ORF36 (γ-herpesvirus) promote nuclear lamina disassembly via a BGL4-like mechanism [259,260]. Intriguingly, only γ-herpesvirus BGL4 homolog proteins (KSHV ORF36 and MHV68 ORF36) can mediate the nuclear import of VCA [258].
The nuclear translocation of EBV EGR1 is mediated by importin-7 (IPO7) [261], and the accumulation of EGR1 is correlated with the viral lytic phase [262]. EGR1 can negatively regulate IPO7 expression using EBV miRNAs (mirBART3 and mirBART16) to maintain a level that is optimal for the growth of EBV-transformed cells [261]. EBV EB2 (also called M or SM) interacts with Crm1 to boost the EB2-mediated gene expression and also to mediate the export of unspliced lytic EBV mRNA [229]. EB2 was shown to be associated with the GTPase Ran and Nup214 during the export of EBV mRNA to the cytoplasm [263]. Herpesviridae family members express nuclear egress proteins 1 (BFLF1) and 2 (BFRF2) to form a nuclear egress complex (NEC) at the host inner NE, which allows the export of new viral capsids containing viral DNA [264,265,266]. EBV BFLF2 interacts with EBV BFRF1 and lamin B at the nuclear rim, making a nuclear lamina for new EBV maturation and budding through the inner nuclear membrane [265]. The homolog proteins of BFLF2, HSV-1 UL31 (α-herpesvirus) [264] and HCMV UL53 (β-herpesvirus) [266], also possess NLSs to mediate their nuclear import. HSV-1 UL31 has a functional bipartite NLS [264], whereas both HCMV UL53 [266] and EBV BFLF2 [267] have a functional monopartite NLS. BFLF2’s nuclear import requires Ran, importin-α7, importin-β1, and importin-β2, but not importins α1, α3, or α5 [267]. On the other hand, HSV-1 UL31 interacts with Ran, importin-α1, and importin-β2 for its nuclear import [268,269].
Non-functional NESs have been found in EBV BFLF2 [267] and HSV-1 UL31 [264,269], and no NESs have been found in HCMV UL53 [270]; these findings suggest that these proteins do not require Crm1 for their export. EBV BFLF2 interacts with Nxf1 in the absence of RNA for its export. The nuclear export receptors for HSV-1 UL31 and HCMV UL53 are yet to be determined [267]. Funk et al. conducted a comprehensive analysis on the EBV and HSV-1 tegument proteins using an in vitro assay called Nuclear EXport Trapped by RAPamycin (NEX-TRAP) [271] and found that two EBV (pBTRF1 and pBGFL3) and nine HSV-1 tegument proteins showed nuclear export activity. The group compared the nuclear export activity between the EBV and HSV-1 tegument orthologs; EBV pBTRF1 behaved in a similar way to its HSV-1 orthologue pUL21, exhibiting active export activity. Conversely, EBV pBGFL3 (exported) displayed the opposite behavior compared to its HSV-1 orthologue pUL14 (non-exported). The NES of pBTRF1 matches the Rev NES consensus, and that of pBGFL3 matches the PKI NES consensus. The NESs of both Rev and PKI are recognized by Crm1, which suggests that EBV pBTRF1’s and pBGFL3’s export activities are Crm1-dependent. A leptomycin B (LMB) assay revealed six HSV-1 tegument proteins (pUL4, pUL11, pUL13, pUL21, pUL37d11, and pUL48) that are exported in a Crm1-dependent manner.
5. Conclusions
Viral proteins masquerade as host factors to exploit the host nuclear transport machinery for viral replication, and they reprogram the host cellular environment to enhance their replication and evade the host immunity. Nuclear transport inhibitors can not only interrupt viral replication but also disrupt the viral assembly and restore the host immunity. Targeting host factors to block nuclear transport could exert a broad-spectrum antiviral effect, as reported for Ivermectin. Nonetheless, the feasibility of using inhibitors of host factors as antiviral drugs could be limited when they target important NTRs, such as importin-β1 and Crm1, owing to likely adverse effects. Furthermore, many infectious diseases, especially COVID-19, have a complicated clinical spectrum. The administration timing of the host-specific nuclear transport inhibitors needs to be carefully evaluated to maximize clinical benefits. In silico drug screening and in vitro and in vivo experiments could accelerate the identification of potent viral-specific nuclear transport inhibitors. As certain viral factors not only exploit the host NTRs but also have their own biological functions, viral-specific nuclear transport inhibitors could “kill two birds with one stone”, that is, not only improve the viral clearance but also preserve/restore the host nucleocytoplasmic trafficking. Lastly, on the basis of our experience applying HS-AFM imaging to human NPCs [17,336], DNA–DNA-binding protein interactions [337], and viral proteins [338,339], we believe it will be worthwhile to observe the interaction between the viral factors and the host nuclear transport factors in real-time.
Author Contributions
Conceptualization, K.L.; writing, E.S.S. and K.L.; overview and supervision, R.W.W. All authors have read and agreed to the published version of the manuscript.
Funding
This project was funded by grants from the MEXT/JSPS KAKENHI (19K23841, 20K16262 (to K.L.), 17H05874, 17K08655 (to R.W.W.)) from MEXT Japan, the NanoLSI Grant for Transdisciplinary Research Promotion (to K.L.), Kanazawa University against COVID-19 (to R.W.W.), the Kobayashi International Scholarship Foundation (to R.W.W.), and the Shimadzu Science Foundation (to R.W.W.).
Acknowledgments
We thank all Wong Lab members for their kind assistance. We also thank reviewers for their efforts and the constructive comments in our review.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Korobeinikov, A. Immune response and within-host viral evolution: Immune response can accelerate evolution. J. Theor. Biol. 2018, 456, 74–83. [Google Scholar] [CrossRef] [PubMed]
- Görlich, D.; Kutay, U. Transport between the cell nucleus and the cytoplasm. Ann. Rev. Cell Develop. Biol. 1999, 15, 607–660. [Google Scholar] [CrossRef]
- Makhnevych, T.; Lusk, C.P.; Anderson, A.M.; Aitchison, J.D.; Wozniak, R.W. Cell cycle regulated transport controlled by alterations in the nuclear pore complex. Cell 2003, 115, 813–823. [Google Scholar] [CrossRef]
- Nakano, H.; Funasaka, T.; Hashizume, C.; Wong, R.W. Nucleoporin translocated promoter region (Tpr) associates with dynein complex, preventing chromosome lagging formation during mitosis. J. Biol. Chem. 2010, 285, 10841–10849. [Google Scholar] [CrossRef]
- Perera, R.M.; Stoykova, S.; Nicolay, B.N.; Ross, K.N.; Fitamant, J.; Boukhali, M.; Lengrand, J.; Deshpande, V.; Selig, M.K.; Ferrone, C.R.; et al. Transcriptional control of autophagy-lysosome function drives pancreatic cancer metabolism. Nature 2015, 524, 361–365. [Google Scholar] [CrossRef]
- Dewi, F.R.P.; Jiapaer, S.; Kobayashi, A.; Hazawa, M.; Ikliptikawati, D.K.; Hartono; Sabit, H.; Nakada, M.; Wong, R.W. Nucleoporin TPR (translocated promoter region, nuclear basket protein) upregulation alters MTOR-HSF1 trails and suppresses autophagy induction in ependymoma. Autophagy 2021, 17, 1001–1012. [Google Scholar] [CrossRef]
- Sun, J.; Shi, Y.; Yildirim, E. The nuclear pore complex in cell type-specific chromatin structure and gene regulation. Trends Genet. TIG 2019, 35, 579–588. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Zebell, S.G.; Liang, Z.; Wang, S.; Kang, B.H.; Dong, X. Nuclear pore permeabilization is a convergent signaling event in effector-triggered immunity. Cell 2016, 166, 1526–1538. [Google Scholar] [CrossRef]
- Lezon, T.R.; Sali, A.; Bahar, I. Global motions of the nuclear pore complex: Insights from elastic network models. PLoS Comput. Biol. 2009, 5, e1000496. [Google Scholar] [CrossRef] [PubMed]
- Bui, K.H.; von Appen, A.; DiGuilio, A.L.; Ori, A.; Sparks, L.; Mackmull, M.T.; Bock, T.; Hagen, W.; Andrés-Pons, A.; Glavy, J.S.; et al. Integrated structural analysis of the human nuclear pore complex scaffold. Cell 2013, 155, 1233–1243. [Google Scholar] [CrossRef]
- Eibauer, M.; Pellanda, M.; Turgay, Y.; Dubrovsky, A.; Wild, A.; Medalia, O. Structure and gating of the nuclear pore complex. Nat. Commun. 2015, 6, 7532. [Google Scholar] [CrossRef]
- Hülsmann, B.B.; Labokha, A.A.; Görlich, D. The permeability of reconstituted nuclear pores provides direct evidence for the selective phase model. Cell 2012, 150, 738–751. [Google Scholar] [CrossRef]
- Lim, R.Y.; Huang, N.P.; Köser, J.; Deng, J.; Lau, K.H.; Schwarz-Herion, K.; Fahrenkrog, B.; Aebi, U. Flexible phenylalanine-glycine nucleoporins as entropic barriers to nucleocytoplasmic transport. Proc. Nat. Acad. Sci. USA 2006, 103, 9512–9517. [Google Scholar] [CrossRef]
- Frey, S.; Görlich, D. A saturated FG-repeat hydrogel can reproduce the permeability properties of nuclear pore complexes. Cell 2007, 130, 512–523. [Google Scholar] [CrossRef] [PubMed]
- Peters, R. Translocation through the nuclear pore complex: Selectivity and speed by reduction-of-dimensionality. Traffic 2005, 6, 421–427. [Google Scholar] [CrossRef]
- Patel, S.S.; Belmont, B.J.; Sante, J.M.; Rexach, M.F. Natively unfolded nucleoporins gate protein diffusion across the nuclear pore complex. Cell 2007, 129, 83–96. [Google Scholar] [CrossRef]
- Mohamed, M.S.; Hazawa, M.; Kobayashi, A.; Guillaud, L.; Watanabe-Nakayama, T.; Nakayama, M.; Wang, H.; Kodera, N.; Oshima, M.; Ando, T.; et al. Spatiotemporally tracking of nano-biofilaments inside the nuclear pore complex core. Biomaterials 2020, 256, 120198. [Google Scholar] [CrossRef]
- Dworetzky, S.I.; Lanford, R.E.; Feldherr, C.M. The effects of variations in the number and sequence of targeting signals on nuclear uptake. J. Cell Biol. 1988, 107, 1279–1287. [Google Scholar] [CrossRef]
- Panté, N.; Kann, M. Nuclear pore complex is able to transport macromolecules with diameters of about 39 nm. Mol. Biol. Cell 2002, 13, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Pumroy, R.A.; Cingolani, G. Diversification of importin-α isoforms in cellular trafficking and disease states. Biochem. J. 2015, 466, 13–28. [Google Scholar] [CrossRef] [PubMed]
- Mosammaparast, N.; Pemberton, L.F. Karyopherins: From nuclear-transport mediators to nuclear-function regulators. Trends Cell Biol. 2004, 14, 547–556. [Google Scholar] [CrossRef]
- Fontes, M.R.; Teh, T.; Kobe, B. Structural basis of recognition of monopartite and bipartite nuclear localization sequences by mammalian importin-alpha. J. Mol. Biol. 2000, 297, 1183–1194. [Google Scholar] [CrossRef] [PubMed]
- Herold, A.; Truant, R.; Wiegand, H.; Cullen, B.R. Determination of the functional domain organization of the importin alpha nuclear import factor. J. Cell Biol. 1998, 143, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Conti, E.; Uy, M.; Leighton, L.; Blobel, G.; Kuriyan, J. Crystallographic analysis of the recognition of a nuclear localization signal by the nuclear import factor karyopherin alpha. Cell 1998, 94, 193–204. [Google Scholar] [CrossRef]
- Kalderon, D.; Richardson, W.D.; Markham, A.F.; Smith, A.E. Sequence requirements for nuclear location of simian virus 40 large-T antigen. Nature 1984, 311, 33–38. [Google Scholar] [CrossRef]
- Robbins, J.; Dilworth, S.M.; Laskey, R.A.; Dingwall, C. Two interdependent basic domains in nucleoplasmin nuclear targeting sequence: Identification of a class of bipartite nuclear targeting sequence. Cell 1991, 64, 615–623. [Google Scholar] [CrossRef]
- Cingolani, G.; Petosa, C.; Weis, K.; Müller, C.W. Structure of importin-beta bound to the IBB domain of importin-alpha. Nature 1999, 399, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Bayliss, R.; Littlewood, T.; Stewart, M. Structural basis for the interaction between FxFG nucleoporin repeats and importin-beta in nuclear trafficking. Cell 2000, 102, 99–108. [Google Scholar] [CrossRef]
- Conti, E.; Izaurralde, E. Nucleocytoplasmic transport enters the atomic age. Curr. Opin. Cell Biol. 2001, 13, 310–319. [Google Scholar] [CrossRef]
- Lee, S.J.; Matsuura, Y.; Liu, S.M.; Stewart, M. Structural basis for nuclear import complex dissociation by RanGTP. Nature 2005, 435, 693–696. [Google Scholar] [CrossRef]
- Görlich, D.; Panté, N.; Kutay, U.; Aebi, U.; Bischoff, F.R. Identification of different roles for RanGDP and RanGTP in nuclear protein import. EMBO J. 1996, 15, 5584–5594. [Google Scholar] [CrossRef]
- Chi, N.C.; Adam, E.J.; Visser, G.D.; Adam, S.A. RanBP1 stabilizes the interaction of Ran with p97 nuclear protein import. J. Cell Biol. 1996, 135, 559–569. [Google Scholar] [CrossRef] [PubMed]
- Rexach, M.; Blobel, G. Protein import into nuclei: Association and dissociation reactions involving transport substrate, transport factors, and nucleoporins. Cell 1995, 83, 683–692. [Google Scholar] [CrossRef]
- Kutay, U.; Bischoff, F.R.; Kostka, S.; Kraft, R.; Görlich, D. Export of importin alpha from the nucleus is mediated by a specific nuclear transport factor. Cell 1997, 90, 1061–1071. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Klebe, C.; Kretschmer, J.; Wittinghofer, A.; Ponstingl, H. RanGAP1 induces GTPase activity of nuclear Ras-related Ran. Proc. Nat. Acad. Sci. USA 1994, 91, 2587–2591. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Krebber, H.; Kempf, T.; Hermes, I.; Ponstingl, H. Human RanGTPase-activating protein RanGAP1 is a homologue of yeast Rna1p involved in mRNA processing and transport. Proc. Nat. Acad. Sci. USA 1995, 92, 1749–1753. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Krebber, H.; Smirnova, E.; Dong, W.; Ponstingl, H. Co-activation of RanGTPase and inhibition of GTP dissociation by Ran-GTP binding protein RanBP1. EMBO J. 1995, 14, 705–715. [Google Scholar] [CrossRef]
- Coutavas, E.; Ren, M.; Oppenheim, J.D.; D’Eustachio, P.; Rush, M.G. Characterization of proteins that interact with the cell-cycle regulatory protein Ran/TC4. Nature 1993, 366, 585–587. [Google Scholar] [CrossRef] [PubMed]
- Bayliss, R.; Ribbeck, K.; Akin, D.; Kent, H.M.; Feldherr, C.M.; Görlich, D.; Stewart, M. Interaction between NTF2 and xFxFG-containing nucleoporins is required to mediate nuclear import of RanGDP. J. Mol. Biol. 1999, 293, 579–593. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Ponstingl, H. Catalysis of guanine nucleotide exchange on Ran by the mitotic regulator RCC1. Nature 1991, 354, 80–82. [Google Scholar] [CrossRef]
- Fischer, U.; Huber, J.; Boelens, W.C.; Mattaj, I.W.; Lührmann, R. The HIV-1 Rev activation domain is a nuclear export signal that accesses an export pathway used by specific cellular RNAs. Cell 1995, 82, 475–483. [Google Scholar] [CrossRef]
- Kırlı, K.; Karaca, S.; Dehne, H.J.; Samwer, M.; Pan, K.T.; Lenz, C.; Urlaub, H.; Görlich, D. A deep proteomics perspective on CRM1-mediated nuclear export and nucleocytoplasmic partitioning. eLife 2015, 4. [Google Scholar] [CrossRef]
- Lindsay, M.E.; Holaska, J.M.; Welch, K.; Paschal, B.M.; Macara, I.G. Ran-binding protein 3 is a cofactor for Crm1-mediated nuclear protein export. J. Cell Biol. 2001, 153, 1391–1402. [Google Scholar] [CrossRef]
- Hutten, S.; Kehlenbach, R.H. Nup214 is required for CRM1-dependent nuclear protein export in vivo. Mol. Cell. Biol. 2006, 26, 6772–6785. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, J.; Min, S.; Zhang, Y.; Zhang, Y.; Zhou, Q.; Shen, X.; Jia, D.; Han, J.; Sun, Q. Distinct RanBP1 nuclear export and cargo dissociation mechanisms between fungi and animals. eLife 2019, 8. [Google Scholar] [CrossRef]
- Engelsma, D.; Bernad, R.; Calafat, J.; Fornerod, M. Supraphysiological nuclear export signals bind CRM1 independently of RanGTP and arrest at Nup358. EMBO J. 2004, 23, 3643–3652. [Google Scholar] [CrossRef] [PubMed]
- Kehlenbach, R.H.; Dickmanns, A.; Kehlenbach, A.; Guan, T.; Gerace, L. A role for RanBP1 in the release of CRM1 from the nuclear pore complex in a terminal step of nuclear export. J. Cell Biol. 1999, 145, 645–657. [Google Scholar] [CrossRef]
- Herold, A.; Suyama, M.; Rodrigues, J.P.; Braun, I.C.; Kutay, U.; Carmo-Fonseca, M.; Bork, P.; Izaurralde, E. TAP (NXF1) belongs to a multigene family of putative RNA export factors with a conserved modular architecture. Mol. Cell. Biol. 2000, 20, 8996–9008. [Google Scholar] [CrossRef] [PubMed]
- Stage-Zimmermann, T.; Schmidt, U.; Silver, P.A. Factors affecting nuclear export of the 60S ribosomal subunit in vivo. Mol. Biol. Cell 2000, 11, 3777–3789. [Google Scholar] [CrossRef] [PubMed]
- Ohno, M.; Segref, A.; Bachi, A.; Wilm, M.; Mattaj, I.W. PHAX, a mediator of U snRNA nuclear export whose activity is regulated by phosphorylation. Cell 2000, 101, 187–198. [Google Scholar] [CrossRef]
- Stade, K.; Ford, C.S.; Guthrie, C.; Weis, K. Exportin 1 (Crm1p) is an essential nuclear export factor. Cell 1997, 90, 1041–1050. [Google Scholar] [CrossRef]
- Kutay, U.; Lipowsky, G.; Izaurralde, E.; Bischoff, F.R.; Schwarzmaier, P.; Hartmann, E.; Görlich, D. Identification of a tRNA-specific nuclear export receptor. Mol. Cell 1998, 1, 359–369. [Google Scholar] [CrossRef]
- Arts, G.J.; Fornerod, M.; Mattaj, I.W. Identification of a nuclear export receptor for tRNA. Curr. Biol. CB 1998, 8, 305–314. [Google Scholar] [CrossRef]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Hautbergue, G.M.; Hung, M.L.; Golovanov, A.P.; Lian, L.Y.; Wilson, S.A. Mutually exclusive interactions drive handover of mRNA from export adaptors to TAP. Proc. Nat. Acad. Sci. USA 2008, 105, 5154–5159. [Google Scholar] [CrossRef]
- Braun, I.C.; Herold, A.; Rode, M.; Izaurralde, E. Nuclear export of mRNA by TAP/NXF1 requires two nucleoporin-binding sites but not p15. Mol. Cell. Biol. 2002, 22, 5405–5418. [Google Scholar] [CrossRef] [PubMed]
- Farny, N.G.; Hurt, J.A.; Silver, P.A. Definition of global and transcript-specific mRNA export pathways in metazoans. Genes Dev. 2008, 22, 66–78. [Google Scholar] [CrossRef]
- Guzik, B.W.; Levesque, L.; Prasad, S.; Bor, Y.C.; Black, B.E.; Paschal, B.M.; Rekosh, D.; Hammarskjöld, M.L. NXT1 (p15) is a crucial cellular cofactor in TAP-dependent export of intron-containing RNA in mammalian cells. Mol. Cell. Biol. 2001, 21, 2545–2554. [Google Scholar] [CrossRef]
- Grüter, P.; Tabernero, C.; von Kobbe, C.; Schmitt, C.; Saavedra, C.; Bachi, A.; Wilm, M.; Felber, B.K.; Izaurralde, E. TAP, the human homolog of Mex67p, mediates CTE-dependent RNA export from the nucleus. Mol. Cell 1998, 1, 649–659. [Google Scholar] [CrossRef]
- Blevins, M.B.; Smith, A.M.; Phillips, E.M.; Powers, M.A. Complex formation among the RNA export proteins Nup98, Rae1/Gle2, and TAP. J. Biol. Chem. 2003, 278, 20979–20988. [Google Scholar] [CrossRef]
- Fribourg, S.; Braun, I.C.; Izaurralde, E.; Conti, E. Structural basis for the recognition of a nucleoporin FG repeat by the NTF2-like domain of the TAP/p15 mRNA nuclear export factor. Mol. Cell 2001, 8, 645–656. [Google Scholar] [CrossRef]
- Braun, I.C.; Rohrbach, E.; Schmitt, C.; Izaurralde, E. TAP binds to the constitutive transport element (CTE) through a novel RNA-binding motif that is sufficient to promote CTE-dependent RNA export from the nucleus. EMBO J. 1999, 18, 1953–1965. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Ren, Y. Mechanisms of nuclear mRNA export: A structural perspective. Traffic 2019, 20, 829–840. [Google Scholar] [CrossRef]
- Brennan, C.M.; Gallouzi, I.E.; Steitz, J.A. Protein ligands to HuR modulate its interaction with target mRNAs in vivo. J. Cell Biol. 2000, 151, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Bogerd, H.P.; Wang, P.J.; Page, D.C.; Cullen, B.R. Two closely related human nuclear export factors utilize entirely distinct export pathways. Mol. Cell 2001, 8, 397–406. [Google Scholar] [CrossRef]
- Volpon, L.; Culjkovic-Kraljacic, B.; Sohn, H.S.; Blanchet-Cohen, A.; Osborne, M.J.; Borden, K.L.B. A biochemical framework for eIF4E-dependent mRNA export and nuclear recycling of the export machinery. RNA 2017, 23, 927–937. [Google Scholar] [CrossRef] [PubMed]
- Trotta, C.R.; Lund, E.; Kahan, L.; Johnson, A.W.; Dahlberg, J.E. Coordinated nuclear export of 60S ribosomal subunits and NMD3 in vertebrates. EMBO J. 2003, 22, 2841–2851. [Google Scholar] [CrossRef]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Weiss, S.R.; Navas-Martin, S. Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol. Mol. Biol. Rev. MMBR 2005, 69, 635–664. [Google Scholar] [CrossRef]
- Bleibtreu, A.; Bertine, M.; Bertin, C.; Houhou-Fidouh, N.; Visseaux, B. Focus on Middle East respiratory syndrome coronavirus (MERS-CoV). Med. Mal. Infect. 2020, 50, 243–251. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Eng. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Snijder, E.J.; Limpens, R.; de Wilde, A.H.; de Jong, A.W.M.; Zevenhoven-Dobbe, J.C.; Maier, H.J.; Faas, F.; Koster, A.J.; Bárcena, M. A unifying structural and functional model of the coronavirus replication organelle: Tracking down RNA synthesis. PLoS Biol. 2020, 18, e3000715. [Google Scholar] [CrossRef]
- Wolff, G.; Limpens, R.; Zevenhoven-Dobbe, J.C.; Laugks, U.; Zheng, S.; de Jong, A.W.M.; Koning, R.I.; Agard, D.A.; Grünewald, K.; Koster, A.J.; et al. A molecular pore spans the double membrane of the coronavirus replication organelle. Science 2020, 369, 1395–1398. [Google Scholar] [CrossRef]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Miorin, L.; Kehrer, T.; Sanchez-Aparicio, M.T.; Zhang, K.; Cohen, P.; Patel, R.S.; Cupic, A.; Makio, T.; Mei, M.; Moreno, E.; et al. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Nat. Acad. Sci. USA 2020, 117, 28344–28354. [Google Scholar] [CrossRef]
- Kato, K.; Ikliptikawati, D.K.; Kobayashi, A.; Kondo, H.; Lim, K.; Hazawa, M.; Wong, R.W. Overexpression of SARS-CoV-2 protein ORF6 dislocates RAE1 and NUP98 from the nuclear pore complex. Biochem. Biophys. Res. Commun. 2021, 536, 59–66. [Google Scholar] [CrossRef]
- Addetia, A.; Lieberman, N.A.P.; Phung, Q.; Hsiang, T.Y.; Xie, H.; Roychoudhury, P.; Shrestha, L.; Loprieno, M.A.; Huang, M.L.; Gale, M., Jr.; et al. SARS-CoV-2 ORF6 Disrupts Bidirectional Nucleocytoplasmic Transport through Interactions with Rae1 and Nup98. mBio 2021, 12. [Google Scholar] [CrossRef]
- Xia, H.; Cao, Z.; Xie, X.; Zhang, X.; Chen, J.Y.; Wang, H.; Menachery, V.D.; Rajsbaum, R.; Shi, P.Y. Evasion of Type I Interferon by SARS-CoV-2. Cell Rep. 2020, 33, 108234. [Google Scholar] [CrossRef]
- Frieman, M.; Yount, B.; Heise, M.; Kopecky-Bromberg, S.A.; Palese, P.; Baric, R.S. Severe acute respiratory syndrome coronavirus ORF6 antagonizes STAT1 function by sequestering nuclear import factors on the rough endoplasmic reticulum/Golgi membrane. J. Virol. 2007, 81, 9812–9824. [Google Scholar] [CrossRef]
- Kopecky-Bromberg, S.A.; Martínez-Sobrido, L.; Frieman, M.; Baric, R.A.; Palese, P. Severe acute respiratory syndrome coronavirus open reading frame (ORF) 3b, ORF 6, and nucleocapsid proteins function as interferon antagonists. J. Virol. 2007, 81, 548–557. [Google Scholar] [CrossRef] [PubMed]
- Izaurralde, E.; Jarmolowski, A.; Beisel, C.; Mattaj, I.W.; Dreyfuss, G.; Fischer, U. A role for the M9 transport signal of hnRNP A1 in mRNA nuclear export. J. Cell Biol. 1997, 137, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Durie, D.; Li, H.; Liu, B.Q.; Skehel, J.M.; Mauri, F.; Cuorvo, L.V.; Barbareschi, M.; Guo, L.; Holcik, M.; et al. hnRNPA1 couples nuclear export and translation of specific mRNAs downstream of FGF-2/S6K2 signalling. Nucl. Acids Res. 2014, 42, 12483–12497. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Miorin, L.; Makio, T.; Dehghan, I.; Gao, S.; Xie, Y.; Zhong, H.; Esparza, M.; Kehrer, T.; Kumar, A.; et al. Nsp1 protein of SARS-CoV-2 disrupts the mRNA export machinery to inhibit host gene expression. Sci. Adv. 2021, 7. [Google Scholar] [CrossRef]
- Gomez, G.N.; Abrar, F.; Dodhia, M.P.; Gonzalez, F.G.; Nag, A. SARS coronavirus protein nsp1 disrupts localization of Nup93 from the nuclear pore complex. Biochem. Cell Biol. Biochim. Biol. Cell. 2019, 97, 758–766. [Google Scholar] [CrossRef]
- Sharma, K.; Åkerström, S.; Sharma, A.K.; Chow, V.T.; Teow, S.; Abrenica, B.; Booth, S.A.; Booth, T.F.; Mirazimi, A.; Lal, S.K. SARS-CoV 9b protein diffuses into nucleus, undergoes active Crm1 mediated nucleocytoplasmic export and triggers apoptosis when retained in the nucleus. PLoS ONE 2011, 6, e19436. [Google Scholar] [CrossRef]
- Yang, Y.; Ye, F.; Zhu, N.; Wang, W.; Deng, Y.; Zhao, Z.; Tan, W. Middle East respiratory syndrome coronavirus ORF4b protein inhibits type I interferon production through both cytoplasmic and nuclear targets. Sci. Rep. 2015, 5, 17554. [Google Scholar] [CrossRef] [PubMed]
- Canton, J.; Fehr, A.R.; Fernandez-Delgado, R.; Gutierrez-Alvarez, F.J.; Sanchez-Aparicio, M.T.; García-Sastre, A.; Perlman, S.; Enjuanes, L.; Sola, I. MERS-CoV 4b protein interferes with the NF-κB-dependent innate immune response during infection. PLoS Pathog. 2018, 14, e1006838. [Google Scholar] [CrossRef]
- Schmidt, M.L.; Hoenen, T. Characterization of the catalytic center of the Ebola virus L polymerase. PLoS Negl. Trop. Dis. 2017, 11, e0005996. [Google Scholar] [CrossRef]
- Bodmer, B.S.; Greßler, J.; Schmidt, M.L.; Holzerland, J.; Brandt, J.; Braun, S.; Groseth, A.; Hoenen, T. Differences in Viral RNA Synthesis but Not Budding or Entry Contribute to the In Vitro Attenuation of Reston Virus Compared to Ebola Virus. Microorganisms 2020, 8, 1215. [Google Scholar] [CrossRef]
- Reid, S.P.; Valmas, C.; Martinez, O.; Sanchez, F.M.; Basler, C.F. Ebola virus VP24 proteins inhibit the interaction of NPI-1 subfamily karyopherin alpha proteins with activated STAT1. J. Virol. 2007, 81, 13469–13477. [Google Scholar] [CrossRef]
- Reid, S.P.; Leung, L.W.; Hartman, A.L.; Martinez, O.; Shaw, M.L.; Carbonnelle, C.; Volchkov, V.E.; Nichol, S.T.; Basler, C.F. Ebola virus VP24 binds karyopherin alpha1 and blocks STAT1 nuclear accumulation. J. Virol. 2006, 80, 5156–5167. [Google Scholar] [CrossRef]
- Xu, W.; Edwards, M.R.; Borek, D.M.; Feagins, A.R.; Mittal, A.; Alinger, J.B.; Berry, K.N.; Yen, B.; Hamilton, J.; Brett, T.J.; et al. Ebola virus VP24 targets a unique NLS binding site on karyopherin alpha 5 to selectively compete with nuclear import of phosphorylated STAT1. Cell Host Microbe 2014, 16, 187–200. [Google Scholar] [CrossRef]
- He, F.; Melén, K.; Maljanen, S.; Lundberg, R.; Jiang, M.; Österlund, P.; Kakkola, L.; Julkunen, I. Ebolavirus protein VP24 interferes with innate immune responses by inhibiting interferon-λ1 gene expression. Virology 2017, 509, 23–34. [Google Scholar] [CrossRef]
- Zhang, A.P.; Bornholdt, Z.A.; Liu, T.; Abelson, D.M.; Lee, D.E.; Li, S.; Woods, V.L., Jr.; Saphire, E.O. The ebola virus interferon antagonist VP24 directly binds STAT1 and has a novel, pyramidal fold. PLoS Pathog. 2012, 8, e1002550. [Google Scholar] [CrossRef]
- Marg, A.; Shan, Y.; Meyer, T.; Meissner, T.; Brandenburg, M.; Vinkemeier, U. Nucleocytoplasmic shuttling by nucleoporins Nup153 and Nup214 and CRM1-dependent nuclear export control the subcellular distribution of latent Stat1. J. Cell Biol. 2004, 165, 823–833. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee-Kishore, M.; Wright, K.L.; Ting, J.P.; Stark, G.R. How Stat1 mediates constitutive gene expression: A complex of unphosphorylated Stat1 and IRF1 supports transcription of the LMP2 gene. EMBO J. 2000, 19, 4111–4122. [Google Scholar] [CrossRef]
- Shabman, R.S.; Gulcicek, E.E.; Stone, K.L.; Basler, C.F. The Ebola virus VP24 protein prevents hnRNP C1/C2 binding to karyopherin α1 and partially alters its nuclear import. J. Infect. Dis. 2011, 204, S904–S910. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Paek, K.Y.; Choi, K.; Kim, T.D.; Hahm, B.; Kim, K.T.; Jang, S.K. Heterogeneous nuclear ribonucleoprotein C modulates translation of c-myc mRNA in a cell cycle phase-dependent manner. Mol. Cell. Biol. 2003, 23, 708–720. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, I.; Elsby, R.; Fernandez, M.; Faria, P.A.; Nussenzveig, D.R.; Lossos, I.S.; Fontoura, B.M.; Martin, W.D.; Barber, G.N. NFAR-1 and -2 modulate translation and are required for efficient host defense. Proc. Nat. Acad. Sci. USA 2008, 105, 4173–4178. [Google Scholar] [CrossRef]
- Brunner, J.E.; Nguyen, J.H.; Roehl, H.H.; Ho, T.V.; Swiderek, K.M.; Semler, B.L. Functional interaction of heterogeneous nuclear ribonucleoprotein C with poliovirus RNA synthesis initiation complexes. J. Virol. 2005, 79, 3254–3266. [Google Scholar] [CrossRef]
- Gontarek, R.R.; Gutshall, L.L.; Herold, K.M.; Tsai, J.; Sathe, G.M.; Mao, J.; Prescott, C.; Del Vecchio, A.M. hnRNP C and polypyrimidine tract-binding protein specifically interact with the pyrimidine-rich region within the 3′NTR of the HCV RNA genome. Nucl. Acids Res. 1999, 27, 1457–1463. [Google Scholar] [CrossRef] [PubMed]
- Sokolowski, M.; Schwartz, S. Heterogeneous nuclear ribonucleoprotein C binds exclusively to the functionally important UUUUU-motifs in the human papillomavirus type-1 AU-rich inhibitory element. Virus Res. 2001, 73, 163–175. [Google Scholar] [CrossRef]
- Miyake, T.; Farley, C.M.; Neubauer, B.E.; Beddow, T.P.; Hoenen, T.; Engel, D.A. ebola virus inclusion body formation and RNA synthesis are controlled by a novel domain of nucleoprotein interacting with VP35. J. Virol. 2020, 94. [Google Scholar] [CrossRef]
- Gabriel, G.; Feldmann, F.; Reimer, R.; Thiele, S.; Fischer, M.; Hartmann, E.; Bader, M.; Ebihara, H.; Hoenen, T.; Feldmann, H. Importin-α7 Is Involved in the Formation of Ebola Virus Inclusion Bodies but Is Not Essential for Pathogenicity in Mice. J. Infect. Dis. 2015, 212, S316–S321. [Google Scholar] [CrossRef][Green Version]
- Wendt, L.; Brandt, J.; Bodmer, B.S.; Reiche, S.; Schmidt, M.L.; Traeger, S.; Hoenen, T. The ebola virus nucleoprotein recruits the nuclear RNA Export Factor NXF1 into inclusion bodies to facilitate viral protein expression. Cells 2020, 9, 187. [Google Scholar] [CrossRef] [PubMed]
- Hertzog, J.; Dias Junior, A.G.; Rigby, R.E.; Donald, C.L.; Mayer, A.; Sezgin, E.; Song, C.; Jin, B.; Hublitz, P.; Eggeling, C.; et al. Infection with a Brazilian isolate of Zika virus generates RIG-I stimulatory RNA and the viral NS5 protein blocks type I IFN induction and signaling. Eur. J. Immunol. 2018, 48, 1120–1136. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, L.K.; Hoenen, A.; Morgan, G.; Mackenzie, J.M. The endoplasmic reticulum provides the membrane platform for biogenesis of the flavivirus replication complex. J. Virol. 2010, 84, 10438–10447. [Google Scholar] [CrossRef] [PubMed]
- Welsch, S.; Miller, S.; Romero-Brey, I.; Merz, A.; Bleck, C.K.; Walther, P.; Fuller, S.D.; Antony, C.; Krijnse-Locker, J.; Bartenschlager, R. Composition and three-dimensional architecture of the dengue virus replication and assembly sites. Cell Host Microbe 2009, 5, 365–375. [Google Scholar] [CrossRef]
- Yu, L.; Takeda, K.; Markoff, L. Protein-protein interactions among West Nile non-structural proteins and transmembrane complex formation in mammalian cells. Virology 2013, 446, 365–377. [Google Scholar] [CrossRef]
- Ashour, J.; Laurent-Rolle, M.; Shi, P.Y.; García-Sastre, A. NS5 of dengue virus mediates STAT2 binding and degradation. J. Virol. 2009, 83, 5408–5418. [Google Scholar] [CrossRef]
- Brooks, A.J.; Johansson, M.; John, A.V.; Xu, Y.; Jans, D.A.; Vasudevan, S.G. The interdomain region of dengue NS5 protein that binds to the viral helicase NS3 contains independently functional importin beta 1 and importin alpha/beta-recognized nuclear localization signals. J. Biol. Chem. 2002, 277, 36399–36407. [Google Scholar] [CrossRef]
- Pryor, M.J.; Rawlinson, S.M.; Butcher, R.E.; Barton, C.L.; Waterhouse, T.A.; Vasudevan, S.G.; Bardin, P.G.; Wright, P.J.; Jans, D.A.; Davidson, A.D. Nuclear localization of dengue virus nonstructural protein 5 through its importin alpha/beta-recognized nuclear localization sequences is integral to viral infection. Traffic 2007, 8, 795–807. [Google Scholar] [CrossRef]
- Potisopon, S.; Priet, S.; Collet, A.; Decroly, E.; Canard, B.; Selisko, B. The methyltransferase domain of dengue virus protein NS5 ensures efficient RNA synthesis initiation and elongation by the polymerase domain. Nucl. Acids Res. 2014, 42, 11642–11656. [Google Scholar] [CrossRef]
- Hannemann, H.; Sung, P.Y.; Chiu, H.C.; Yousuf, A.; Bird, J.; Lim, S.P.; Davidson, A.D. Serotype-specific differences in dengue virus non-structural protein 5 nuclear localization. J. Biol. Chem. 2013, 288, 22621–22635. [Google Scholar] [CrossRef]
- Tay, M.Y.; Smith, K.; Ng, I.H.; Chan, K.W.; Zhao, Y.; Ooi, E.E.; Lescar, J.; Luo, D.; Jans, D.A.; Forwood, J.K.; et al. The C-terminal 18 Amino Acid Region of Dengue Virus NS5 Regulates its Subcellular Localization and Contains a Conserved Arginine Residue Essential for Infectious Virus Production. PLoS Pathog. 2016, 12, e1005886. [Google Scholar] [CrossRef]
- Rawlinson, S.M.; Pryor, M.J.; Wright, P.J.; Jans, D.A. CRM1-mediated nuclear export of dengue virus RNA polymerase NS5 modulates interleukin-8 induction and virus production. J. Biol. Chem. 2009, 284, 15589–15597. [Google Scholar] [CrossRef]
- De Jesús-González, L.A.; Cervantes-Salazar, M.; Reyes-Ruiz, J.M.; Osuna-Ramos, J.F.; Farfán-Morales, C.N.; Palacios-Rápalo, S.N.; Pérez-Olais, J.H.; Cordero-Rivera, C.D.; Hurtado-Monzón, A.M.; Ruíz-Jiménez, F.; et al. The nuclear pore complex: A Target for NS3 protease of dengue and zika viruses. Viruses 2020, 12, 583. [Google Scholar] [CrossRef] [PubMed]
- Palacios-Rápalo, S.N.; De Jesús-González, L.A.; Reyes-Ruiz, J.M.; Osuna-Ramos, J.F.; Farfan-Morales, C.N.; Gutiérrez-Escolano, A.L.; Del Ángel, R.M. Nuclear localization of non-structural protein 3 (NS3) during dengue virus infection. Arch. Virol. 2021, 166, 1439–1446. [Google Scholar] [CrossRef] [PubMed]
- Gulland, A. Zika virus is a global public health emergency, declares WHO. BMJ 2016, 352, i657. [Google Scholar] [CrossRef] [PubMed]
- Kleber de Oliveira, W.; Cortez-Escalante, J.; De Oliveira, W.T.; do Carmo, G.M.; Henriques, C.M.; Coelho, G.E.; Araújo de França, G.V. Increase in reported prevalence of microcephaly in infants born to women living in areas with confirmed zika virus transmission during the first trimester of pregnancy-brazil, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 242–247. [Google Scholar] [CrossRef]
- Yang, L.; Wang, R.; Yang, S.; Ma, Z.; Lin, S.; Nan, Y.; Li, Q.; Tang, Q.; Zhang, Y.J. Karyopherin alpha 6 is required for replication of porcine reproductive and respiratory syndrome virus and zika virus. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [PubMed]
- Ji, W.; Luo, G. Zika virus NS5 nuclear accumulation is protective of protein degradation and is required for viral RNA replication. Virology 2020, 541, 124–135. [Google Scholar] [CrossRef] [PubMed]
- Ng, I.H.W.; Chan, K.W.; Tan, M.J.A.; Gwee, C.P.; Smith, K.M.; Jeffress, S.J.; Saw, W.G.; Swarbrick, C.M.D.; Watanabe, S.; Jans, D.A.; et al. Zika Virus NS5 forms supramolecular nuclear bodies that sequester importin-α and modulate the host immune and pro-inflammatory response in neuronal cells. ACS Infect. Dis. 2019, 5, 932–948. [Google Scholar] [CrossRef]
- Ye, J.; Chen, Z.; Li, Y.; Zhao, Z.; He, W.; Zohaib, A.; Song, Y.; Deng, C.; Zhang, B.; Chen, H.; et al. Japanese encephalitis virus NS5 Inhibits Type I Interferon (IFN) production by blocking the nuclear translocation of ifn regulatory factor 3 and NF-κB. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Lin, S.; Yang, S.; He, J.; Guest, J.D.; Ma, Z.; Yang, L.; Pierce, B.G.; Tang, Q.; Zhang, Y.J. Zika virus NS5 protein antagonizes type I interferon production via blocking TBK1 activation. Virology 2019, 527, 180–187. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Yang, L.; Chang, P.; Yang, S.; Lin, S.; Tang, Q.; Wang, X.; Zhang, Y.J. Zika virus NS2A protein induces the degradation of KPNA2 (karyopherin subunit alpha 2) via chaperone-mediated autophagy. Autophagy 2020, 16, 2238–2251. [Google Scholar] [CrossRef]
- Remenyi, R.; Gao, Y.; Hughes, R.E.; Curd, A.; Zothner, C.; Peckham, M.; Merits, A.; Harris, M. Persistent replication of a chikungunya virus replicon in human cells is associated with presence of stable cytoplasmic granules containing nonstructural protein 3. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [PubMed]
- Hawman, D.W.; Stoermer, K.A.; Montgomery, S.A.; Pal, P.; Oko, L.; Diamond, M.S.; Morrison, T.E. Chronic joint disease caused by persistent Chikungunya virus infection is controlled by the adaptive immune response. J. Virol. 2013, 87, 13878–13888. [Google Scholar] [CrossRef]
- Thomas, S.; Rai, J.; John, L.; Schaefer, S.; Pützer, B.M.; Herchenröder, O. Chikungunya virus capsid protein contains nuclear import and export signals. Virol. J. 2013, 10, 269. [Google Scholar] [CrossRef]
- Jacobs, S.C.; Taylor, A.; Herrero, L.J.; Mahalingam, S.; Fazakerley, J.K. Mutation of a conserved nuclear export sequence in chikungunya virus capsid protein disrupts host cell nuclear import. Viruses 2017, 9, 306. [Google Scholar] [CrossRef]
- Webb, L.G.; Veloz, J.; Pintado-Silva, J.; Zhu, T.; Rangel, M.V.; Mutetwa, T.; Zhang, L.; Bernal-Rubio, D.; Figueroa, D.; Carrau, L.; et al. Chikungunya virus antagonizes cGAS-STING mediated type-I interferon responses by degrading cGAS. PLoS Pathog. 2020, 16, e1008999. [Google Scholar] [CrossRef]
- Göertz, G.P.; McNally, K.L.; Robertson, S.J.; Best, S.M.; Pijlman, G.P.; Fros, J.J. The methyltransferase-like domain of chikungunya virus nsP2 inhibits the interferon response by promoting the nuclear export of STAT1. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Fros, J.J.; Liu, W.J.; Prow, N.A.; Geertsema, C.; Ligtenberg, M.; Vanlandingham, D.L.; Schnettler, E.; Vlak, J.M.; Suhrbier, A.; Khromykh, A.A.; et al. Chikungunya virus nonstructural protein 2 inhibits type I/II interferon-stimulated JAK-STAT signaling. J. Virol. 2010, 84, 10877–10887. [Google Scholar] [CrossRef] [PubMed]
- Fros, J.J.; van der Maten, E.; Vlak, J.M.; Pijlman, G.P. The C-terminal domain of chikungunya virus nsP2 independently governs viral RNA replication, cytopathicity, and inhibition of interferon signaling. J. Virol. 2013, 87, 10394–10400. [Google Scholar] [CrossRef]
- Wong, K.Z.; Chu, J.J.H. The interplay of viral and host factors in chikungunya virus infection: Targets for antiviral strategies. Viruses 2018, 10, 294. [Google Scholar] [CrossRef]
- Fros, J.J.; Major, L.D.; Scholte, F.E.M.; Gardner, J.; van Hemert, M.J.; Suhrbier, A.; Pijlman, G.P. Chikungunya virus non-structural protein 2-mediated host shut-off disables the unfolded protein response. J. Gen. Virol. 2015, 96, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Poiesz, B.J.; Ruscetti, F.W.; Gazdar, A.F.; Bunn, P.A.; Minna, J.D.; Gallo, R.C. Detection and isolation of type C retrovirus particles from fresh and cultured lymphocytes of a patient with cutaneous T-cell lymphoma. Proc. Nat. Acad. Sci. USA 1980, 77, 7415–7419. [Google Scholar] [CrossRef]
- Barré-Sinoussi, F.; Chermann, J.C.; Rey, F.; Nugeyre, M.T.; Chamaret, S.; Gruest, J.; Dauguet, C.; Axler-Blin, C.; Vézinet-Brun, F.; Rouzioux, C.; et al. Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science 1983, 220, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Engelman, A.N.; Singh, P.K. Cellular and molecular mechanisms of HIV-1 integration targeting. Cell. Mol. Life Sci. 2018, 75, 2491–2507. [Google Scholar] [CrossRef]
- Wong, R.W.; Mamede, J.I.; Hope, T.J. Impact of nucleoporin-mediated chromatin localization and nuclear architecture on HIV integration site selection. J. Virol. 2015, 89, 9702–9705. [Google Scholar] [CrossRef][Green Version]
- Popov, S.; Rexach, M.; Ratner, L.; Blobel, G.; Bukrinsky, M. Viral protein R regulates docking of the HIV-1 preintegration complex to the nuclear pore complex. J. Biol. Chem. 1998, 273, 13347–13352. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Xu, M.; Huang, Q.; Gates, A.T.; Zhang, X.D.; Castle, J.C.; Stec, E.; Ferrer, M.; Strulovici, B.; Hazuda, D.J.; et al. Genome-scale RNAi screen for host factors required for HIV replication. Cell Host Microbe 2008, 4, 495–504. [Google Scholar] [CrossRef]
- König, R.; Zhou, Y.; Elleder, D.; Diamond, T.L.; Bonamy, G.M.; Irelan, J.T.; Chiang, C.Y.; Tu, B.P.; De Jesus, P.D.; Lilley, C.E.; et al. Global analysis of host-pathogen interactions that regulate early-stage HIV-1 replication. Cell 2008, 135, 49–60. [Google Scholar] [CrossRef]
- Brass, A.L.; Dykxhoorn, D.M.; Benita, Y.; Yan, N.; Engelman, A.; Xavier, R.J.; Lieberman, J.; Elledge, S.J. Identification of host proteins required for HIV infection through a functional genomic screen. Science 2008, 319, 921–926. [Google Scholar] [CrossRef]
- Imbeault, M.; Ouellet, M.; Tremblay, M.J. Microarray study reveals that HIV-1 induces rapid type-I interferon-dependent p53 mRNA up-regulation in human primary CD4+ T cells. Retrovirology 2009, 6, 5. [Google Scholar] [CrossRef]
- Tsurutani, N.; Kubo, M.; Maeda, Y.; Ohashi, T.; Yamamoto, N.; Kannagi, M.; Masuda, T. Identification of critical amino acid residues in human immunodeficiency virus type 1 IN required for efficient proviral DNA formation at steps prior to integration in dividing and nondividing cells. J. Virol. 2000, 74, 4795–4806. [Google Scholar] [CrossRef] [PubMed]
- Gallay, P.; Stitt, V.; Mundy, C.; Oettinger, M.; Trono, D. Role of the karyopherin pathway in human immunodeficiency virus type 1 nuclear import. J. Virol. 1996, 70, 1027–1032. [Google Scholar] [CrossRef] [PubMed]
- Fouchier, R.A.; Meyer, B.E.; Simon, J.H.; Fischer, U.; Malim, M.H. HIV-1 infection of non-dividing cells: Evidence that the amino-terminal basic region of the viral matrix protein is important for Gag processing but not for post-entry nuclear import. EMBO J. 1997, 16, 4531–4539. [Google Scholar] [CrossRef]
- Ganser-Pornillos, B.K.; Cheng, A.; Yeager, M. Structure of full-length HIV-1 CA: A model for the mature capsid lattice. Cell 2007, 131, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Arhel, N.J.; Souquere-Besse, S.; Munier, S.; Souque, P.; Guadagnini, S.; Rutherford, S.; Prévost, M.C.; Allen, T.D.; Charneau, P. HIV-1 DNA Flap formation promotes uncoating of the pre-integration complex at the nuclear pore. EMBO J. 2007, 26, 3025–3037. [Google Scholar] [CrossRef]
- Li, C.; Burdick, R.C.; Nagashima, K.; Hu, W.S.; Pathak, V.K. HIV-1 cores retain their integrity until minutes before uncoating in the nucleus. Proc. Nat. Acad. Sci. USA 2021, 118, e2019467118. [Google Scholar] [CrossRef]
- Francis, A.C.; Marin, M.; Prellberg, M.J.; Palermino-Rowland, K.; Melikyan, G.B. HIV-1 Uncoating and Nuclear Import Precede the Completion of Reverse Transcription in Cell Lines and in Primary Macrophages. Viruses 2020, 12, 1234. [Google Scholar] [CrossRef]
- Dharan, A.; Bachmann, N.; Talley, S.; Zwikelmaier, V.; Campbell, E.M. Nuclear pore blockade reveals that HIV-1 completes reverse transcription and uncoating in the nucleus. Nat. Microbiol. 2020, 5, 1088–1095. [Google Scholar] [CrossRef]
- Zila, V.; Margiotta, E.; Turoňová, B.; Müller, T.G.; Zimmerli, C.E.; Mattei, S.; Allegretti, M.; Börner, K.; Rada, J.; Müller, B.; et al. Cone-shaped HIV-1 capsids are transported through intact nuclear pores. Cell 2021, 184, 1032–1046.e18. [Google Scholar] [CrossRef] [PubMed]
- Khan, H.; Sumner, R.P.; Rasaiyaah, J.; Tan, C.P.; Rodriguez-Plata, M.T.; Van Tulleken, C.; Fink, D.; Zuliani-Alvarez, L.; Thorne, L.; Stirling, D.; et al. HIV-1 Vpr antagonizes innate immune activation by targeting karyopherin-mediated NF-κB/IRF3 nuclear transport. eLife 2020, 9, e60821. [Google Scholar] [CrossRef]
- Karni, O.; Friedler, A.; Zakai, N.; Gilon, C.; Loyter, A. A peptide derived from the N-terminal region of HIV-1 Vpr promotes nuclear import in permeabilized cells: Elucidation of the NLS region of the Vpr. FEBS Lett. 1998, 429, 421–425. [Google Scholar] [CrossRef]
- Nitahara-Kasahara, Y.; Kamata, M.; Yamamoto, T.; Zhang, X.; Miyamoto, Y.; Muneta, K.; Iijima, S.; Yoneda, Y.; Tsunetsugu-Yokota, Y.; Aida, Y. Novel nuclear import of Vpr promoted by importin alpha is crucial for human immunodeficiency virus type 1 replication in macrophages. J. Virol. 2007, 81, 5284–5293. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, H.; Sanjoh, A.; Murakami, T.; Murakami, H.; Matsuda, G.; Hagiwara, K.; Yokoyama, M.; Sato, H.; Miyamoto, Y.; Dohmae, N.; et al. Molecular Mechanism of HIV-1 Vpr for Binding to Importin-α. J. Mol. Biol. 2016, 428, 2744–2757. [Google Scholar] [CrossRef]
- Pollard, V.W.; Malim, M.H. The HIV-1 Rev protein. Ann. Rev. Microbiol. 1998, 52, 491–532. [Google Scholar] [CrossRef] [PubMed]
- Henderson, B.R.; Percipalle, P. Interactions between HIV Rev and nuclear import and export factors: The Rev nuclear localisation signal mediates specific binding to human importin-beta. J. Mol. Biol. 1997, 274, 693–707. [Google Scholar] [CrossRef] [PubMed]
- Emerman, M.; Malim, M.H. HIV-1 regulatory/accessory genes: Keys to unraveling viral and host cell biology. Science 1998, 280, 1880–1884. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Echarri, A.; Ross, T.M.; Cullen, B.R. Inhibition of human immunodeficiency virus Rev and human T-cell leukemia virus Rex function, but not Mason-Pfizer monkey virus constitutive transport element activity, by a mutant human nucleoporin targeted to Crm1. J. Virol. 1998, 72, 8627–8635. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, W.; Reichart, B.; Ewald, A.; Müller, E.; Schmitt, I.; Stauber, R.H.; Lottspeich, F.; Jockusch, B.M.; Scheer, U.; Hauber, J.; et al. Cofactor requirements for nuclear export of Rev response element (RRE)- and constitutive transport element (CTE)-containing retroviral RNAs. An unexpected role for actin. J. Cell Biol. 2001, 152, 895–910. [Google Scholar] [CrossRef] [PubMed]
- Ullman, K.S.; Shah, S.; Powers, M.A.; Forbes, D.J. The nucleoporin nup153 plays a critical role in multiple types of nuclear export. Mol. Biol. Cell 1999, 10, 649–664. [Google Scholar] [CrossRef]
- Zolotukhin, A.S.; Felber, B.K. Nucleoporins nup98 and nup214 participate in nuclear export of human immunodeficiency virus type 1 Rev. J. Virol. 1999, 73, 120–127. [Google Scholar] [CrossRef]
- Monette, A.; Panté, N.; Mouland, A.J. HIV-1 remodels the nuclear pore complex. J. Cell Biol. 2011, 193, 619–631. [Google Scholar] [CrossRef]
- Taniguchi, I.; Mabuchi, N.; Ohno, M. HIV-1 Rev protein specifies the viral RNA export pathway by suppressing TAP/NXF1 recruitment. Nucl. Acids Res. 2014, 42, 6645–6658. [Google Scholar] [CrossRef] [PubMed]
- McCauley, S.M.; Kim, K.; Nowosielska, A.; Dauphin, A.; Yurkovetskiy, L.; Diehl, W.E.; Luban, J. Intron-containing RNA from the HIV-1 provirus activates type I interferon and inflammatory cytokines. Nat. Commun. 2018, 9, 5305. [Google Scholar] [CrossRef]
- Te Velthuis, A.J.W.; Grimes, J.M.; Fodor, E. Structural insights into RNA polymerases of negative-sense RNA viruses. Nat. Rev. Microbiol. 2021, 19, 303–318. [Google Scholar] [CrossRef]
- O’Neill, R.E.; Jaskunas, R.; Blobel, G.; Palese, P.; Moroianu, J. Nuclear import of influenza virus RNA can be mediated by viral nucleoprotein and transport factors required for protein import. J. Biol. Chem. 1995, 270, 22701–22704. [Google Scholar] [CrossRef]
- Wang, P.; Palese, P.; O’Neill, R.E. The NPI-1/NPI-3 (karyopherin alpha) binding site on the influenza a virus nucleoprotein NP is a nonconventional nuclear localization signal. J. Virol. 1997, 71, 1850–1856. [Google Scholar] [CrossRef]
- Weber, F.; Kochs, G.; Gruber, S.; Haller, O. A classical bipartite nuclear localization signal on Thogoto and influenza A virus nucleoproteins. Virology 1998, 250, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Cros, J.F.; García-Sastre, A.; Palese, P. An unconventional NLS is critical for the nuclear import of the influenza A virus nucleoprotein and ribonucleoprotein. Traffic 2005, 6, 205–213. [Google Scholar] [CrossRef] [PubMed]
- Donchet, A.; Vassal-Stermann, E.; Gérard, F.C.A.; Ruigrok, R.W.H.; Crépin, T. Differential behaviours and preferential bindings of influenza nucleoproteins on importins-α. Viruses 2020, 12, 834. [Google Scholar] [CrossRef] [PubMed]
- Tarendeau, F.; Boudet, J.; Guilligay, D.; Mas, P.J.; Bougault, C.M.; Boulo, S.; Baudin, F.; Ruigrok, R.W.; Daigle, N.; Ellenberg, J.; et al. Structure and nuclear import function of the C-terminal domain of influenza virus polymerase PB2 subunit. Nat. Struct. Mol. Biol. 2007, 14, 229–233. [Google Scholar] [CrossRef]
- Gabriel, G.; Herwig, A.; Klenk, H.D. Interaction of polymerase subunit PB2 and NP with importin alpha1 is a determinant of host range of influenza A virus. PLoS Pathog. 2008, 4, e11. [Google Scholar] [CrossRef] [PubMed]
- Hudjetz, B.; Gabriel, G. Human-like PB2 627K influenza virus polymerase activity is regulated by importin-α1 and -α7. PLoS Pathog. 2012, 8, e1002488. [Google Scholar] [CrossRef]
- Pumroy, R.A.; Ke, S.; Hart, D.J.; Zachariae, U.; Cingolani, G. Molecular determinants for nuclear import of influenza A PB2 by importin α isoforms 3 and 7. Structure 2015, 23, 374–384. [Google Scholar] [CrossRef] [PubMed]
- Thiele, S.; Stanelle-Bertram, S.; Beck, S.; Kouassi, N.M.; Zickler, M.; Müller, M.; Tuku, B.; Resa-Infante, P.; van Riel, D.; Alawi, M.; et al. Cellular Importin-α3 Expression Dynamics in the Lung Regulate Antiviral Response Pathways against Influenza A Virus Infection. Cell Rep. 2020, 31, 107549. [Google Scholar] [CrossRef]
- Swale, C.; Monod, A.; Tengo, L.; Labaronne, A.; Garzoni, F.; Bourhis, J.M.; Cusack, S.; Schoehn, G.; Berger, I.; Ruigrok, R.W.; et al. Structural characterization of recombinant IAV polymerase reveals a stable complex between viral PA-PB1 heterodimer and host RanBP5. Sci. Rep. 2016, 6, 24727. [Google Scholar] [CrossRef]
- Swale, C.; Da Costa, B.; Sedano, L.; Garzoni, F.; McCarthy, A.A.; Berger, I.; Bieniossek, C.; Ruigrok, R.W.H.; Delmas, B.; Crépin, T. X-ray structure of the human karyopherin RanBP5, an essential factor for influenza polymerase nuclear trafficking. J. Mol. Biol. 2020, 432, 3353–3359. [Google Scholar] [CrossRef]
- Brunotte, L.; Flies, J.; Bolte, H.; Reuther, P.; Vreede, F.; Schwemmle, M. The nuclear export protein of H5N1 influenza A viruses recruits Matrix 1 (M1) protein to the viral ribonucleoprotein to mediate nuclear export. J. Biol. Chem. 2014, 289, 20067–20077. [Google Scholar] [CrossRef]
- Gao, S.; Wang, S.; Cao, S.; Sun, L.; Li, J.; Bi, Y.; Gao, G.F.; Liu, W. Characteristics of nucleocytoplasmic transport of H1N1 influenza A virus nuclear export protein. J. Virol. 2014, 88, 7455–7463. [Google Scholar] [CrossRef]
- Satterly, N.; Tsai, P.L.; van Deursen, J.; Nussenzveig, D.R.; Wang, Y.; Faria, P.A.; Levay, A.; Levy, D.E.; Fontoura, B.M. Influenza virus targets the mRNA export machinery and the nuclear pore complex. Proc. Nat. Acad. Sci. USA 2007, 104, 1853–1858. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Xie, Y.; Muñoz-Moreno, R.; Wang, J.; Zhang, L.; Esparza, M.; García-Sastre, A.; Fontoura, B.M.A.; Ren, Y. Structural basis for influenza virus NS1 protein block of mRNA nuclear export. Nat. Microb. 2019, 4, 1671–1679. [Google Scholar] [CrossRef]
- Nemeroff, M.E.; Barabino, S.M.; Li, Y.; Keller, W.; Krug, R.M. Influenza virus NS1 protein interacts with the cellular 30 kDa subunit of CPSF and inhibits 3′end formation of cellular pre-mRNAs. Mol. Cell 1998, 1, 991–1000. [Google Scholar] [CrossRef]
- Chen, Z.; Li, Y.; Krug, R.M. Influenza A virus NS1 protein targets poly(A)-binding protein II of the cellular 3′-end processing machinery. EMBO J. 1999, 18, 2273–2283. [Google Scholar] [CrossRef]
- Morita, M.; Kuba, K.; Ichikawa, A.; Nakayama, M.; Katahira, J.; Iwamoto, R.; Watanebe, T.; Sakabe, S.; Daidoji, T.; Nakamura, S.; et al. The lipid mediator protectin D1 inhibits influenza virus replication and improves severe influenza. Cell 2013, 153, 112–125. [Google Scholar] [CrossRef]
- Van Doorslaer, K.; Chen, Z.; Bernard, H.U.; Chan, P.K.S.; DeSalle, R.; Dillner, J.; Forslund, O.; Haga, T.; McBride, A.A.; Villa, L.L.; et al. ICTV Virus Taxonomy Profile: Papillomaviridae. J. Gen. Virol. 2018, 99, 989–990. [Google Scholar] [CrossRef]
- Muñoz, N.; Bosch, F.X.; de Sanjosé, S.; Herrero, R.; Castellsagué, X.; Shah, K.V.; Snijders, P.J.; Meijer, C.J. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N. Eng. J. Med. 2003, 348, 518–527. [Google Scholar] [CrossRef]
- Aydin, I.; Weber, S.; Snijder, B.; Samperio Ventayol, P.; Kühbacher, A.; Becker, M.; Day, P.M.; Schiller, J.T.; Kann, M.; Pelkmans, L.; et al. Large scale RNAi reveals the requirement of nuclear envelope breakdown for nuclear import of human papillomaviruses. PLoS Pathog. 2014, 10, e1004162. [Google Scholar] [CrossRef]
- Pyeon, D.; Pearce, S.M.; Lank, S.M.; Ahlquist, P.; Lambert, P.F. Establishment of human papillomavirus infection requires cell cycle progression. PLoS Pathog. 2009, 5, e1000318. [Google Scholar] [CrossRef]
- Tao, M.; Kruhlak, M.; Xia, S.; Androphy, E.; Zheng, Z.M. Signals that dictate nuclear localization of human papillomavirus type 16 oncoprotein E6 in living cells. J. Virol. 2003, 77, 13232–13247. [Google Scholar] [CrossRef]
- Le Roux, L.G.; Moroianu, J. Nuclear entry of high-risk human papillomavirus type 16 E6 oncoprotein occurs via several pathways. J. Virol. 2003, 77, 2330–2337. [Google Scholar] [CrossRef]
- Masson, M.; Hindelang, C.; Sibler, A.P.; Schwalbach, G.; Travé, G.; Weiss, E. Preferential nuclear localization of the human papillomavirus type 16 E6 oncoprotein in cervical carcinoma cells. J. Gen. Virol. 2003, 84, 2099–2104. [Google Scholar] [CrossRef]
- Freedman, D.A.; Levine, A.J. Nuclear export is required for degradation of endogenous p53 by MDM2 and human papillomavirus E6. Mol. Cell. Biol. 1998, 18, 7288–7293. [Google Scholar] [CrossRef]
- Stewart, D.; Ghosh, A.; Matlashewski, G. Involvement of nuclear export in human papillomavirus type 18 E6-mediated ubiquitination and degradation of p53. J. Virol. 2005, 79, 8773–8783. [Google Scholar] [CrossRef][Green Version]
- McLaughlin-Drubin, M.E.; Münger, K. The human papillomavirus E7 oncoprotein. Virology 2009, 384, 335–344. [Google Scholar] [CrossRef]
- Angeline, M.; Merle, E.; Moroianu, J. The E7 oncoprotein of high-risk human papillomavirus type 16 enters the nucleus via a nonclassical Ran-dependent pathway. Virology 2003, 317, 13–23. [Google Scholar] [CrossRef]
- Knapp, A.A.; McManus, P.M.; Bockstall, K.; Moroianu, J. Identification of the nuclear localization and export signals of high risk HPV16 E7 oncoprotein. Virology 2009, 383, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Eberhard, J.; Onder, Z.; Moroianu, J. Nuclear import of high risk HPV16 E7 oncoprotein is mediated by its zinc-binding domain via hydrophobic interactions with Nup62. Virology 2013, 446, 334–345. [Google Scholar] [CrossRef]
- Onder, Z.; Moroianu, J. Nuclear import of cutaneous beta genus HPV8 E7 oncoprotein is mediated by hydrophobic interactions between its zinc-binding domain and FG nucleoporins. Virology 2014, 449, 150–162. [Google Scholar] [CrossRef][Green Version]
- McKee, C.H.; Onder, Z.; Ashok, A.; Cardoso, R.; Moroianu, J. Characterization of the transport signals that mediate the nucleocytoplasmic traffic of low risk HPV11 E7. Virology 2013, 443, 113–122. [Google Scholar] [CrossRef]
- Deng, W.; Lin, B.Y.; Jin, G.; Wheeler, C.G.; Ma, T.; Harper, J.W.; Broker, T.R.; Chow, L.T. Cyclin/CDK regulates the nucleocytoplasmic localization of the human papillomavirus E1 DNA helicase. J. Virol. 2004, 78, 13954–13965. [Google Scholar] [CrossRef] [PubMed]
- Egawa, N.; Wang, Q.; Griffin, H.M.; Murakami, I.; Jackson, D.; Mahmood, R.; Doorbar, J. HPV16 and 18 genome amplification show different E4-dependence, with 16E4 enhancing E1 nuclear accumulation and replicative efficiency via its cell cycle arrest and kinase activation functions. PLoS Pathog. 2017, 13, e1006282. [Google Scholar] [CrossRef]
- Yu, J.H.; Lin, B.Y.; Deng, W.; Broker, T.R.; Chow, L.T. Mitogen-activated protein kinases activate the nuclear localization sequence of human papillomavirus type 11 E1 DNA helicase to promote efficient nuclear import. J. Virol. 2007, 81, 5066–5078. [Google Scholar] [CrossRef] [PubMed]
- Bian, X.L.; Rosas-Acosta, G.; Wu, Y.C.; Wilson, V.G. Nuclear import of bovine papillomavirus type 1 E1 protein is mediated by multiple alpha importins and is negatively regulated by phosphorylation near a nuclear localization signal. J. Virol. 2007, 81, 2899–2908. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fradet-Turcotte, A.; Moody, C.; Laimins, L.A.; Archambault, J. Nuclear export of human papillomavirus type 31 E1 is regulated by Cdk2 phosphorylation and required for viral genome maintenance. J. Virol. 2010, 84, 11747–11760. [Google Scholar] [CrossRef] [PubMed]
- Blachon, S.; Bellanger, S.; Demeret, C.; Thierry, F. Nucleo-cytoplasmic shuttling of high risk human Papillomavirus E2 proteins induces apoptosis. J. Biol. Chem. 2005, 280, 36088–36098. [Google Scholar] [CrossRef] [PubMed]
- Klucevsek, K.; Wertz, M.; Lucchi, J.; Leszczynski, A.; Moroianu, J. Characterization of the nuclear localization signal of high risk HPV16 E2 protein. Virology 2007, 360, 191–198. [Google Scholar] [CrossRef]
- Nelson, L.M.; Rose, R.C.; Moroianu, J. Nuclear import strategies of high risk HPV16 L1 major capsid protein. J. Biol. Chem. 2002, 277, 23958–23964. [Google Scholar] [CrossRef] [PubMed]
- Darshan, M.S.; Lucchi, J.; Harding, E.; Moroianu, J. The l2 minor capsid protein of human papillomavirus type 16 interacts with a network of nuclear import receptors. J. Virol. 2004, 78, 12179–12188. [Google Scholar] [CrossRef] [PubMed]
- Papaevangelou, G.; Roumeliotou-Karayannis, A.; Tassopoulos, N.; Kolaitis, N.; Contoyannis, P.; Krugman, S. Post-exposure hepatitis B vaccination of sexual partners of acute viral hepatitis patients. J. Infect. 1983, 7, 63–67. [Google Scholar] [CrossRef]
- Chaturvedi, V.K.; Singh, A.; Dubey, S.K.; Hetta, H.F.; John, J.; Singh, M.P. Molecular mechanistic insight of hepatitis B virus mediated hepatocellular carcinoma. Microb. Pathog. 2019, 128, 184–194. [Google Scholar] [CrossRef]
- Nassal, M. HBV cccDNA: Viral persistence reservoir and key obstacle for a cure of chronic hepatitis B. Gut 2015, 64, 1972–1984. [Google Scholar] [CrossRef]
- Kann, M.; Sodeik, B.; Vlachou, A.; Gerlich, W.H.; Helenius, A. Phosphorylation-dependent binding of hepatitis B virus core particles to the nuclear pore complex. J. Cell Biol. 1999, 145, 45–55. [Google Scholar] [CrossRef]
- Rabe, B.; Vlachou, A.; Panté, N.; Helenius, A.; Kann, M. Nuclear import of hepatitis B virus capsids and release of the viral genome. Proc. Nat. Acad. Sci. USA 2003, 100, 9849–9854. [Google Scholar] [CrossRef]
- Guo, H.; Mao, R.; Block, T.M.; Guo, J.T. Production and function of the cytoplasmic deproteinized relaxed circular DNA of hepadnaviruses. J. Virol. 2010, 84, 387–396. [Google Scholar] [CrossRef]
- Schmitz, A.; Schwarz, A.; Foss, M.; Zhou, L.; Rabe, B.; Hoellenriegel, J.; Stoeber, M.; Panté, N.; Kann, M. Nucleoporin 153 arrests the nuclear import of hepatitis B virus capsids in the nuclear basket. PLoS Pathog. 2010, 6, e1000741. [Google Scholar] [CrossRef]
- Li, H.C.; Huang, E.Y.; Su, P.Y.; Wu, S.Y.; Yang, C.C.; Lin, Y.S.; Chang, W.C.; Shih, C. Nuclear export and import of human hepatitis B virus capsid protein and particles. PLoS Pathog. 2010, 6, e1001162. [Google Scholar] [CrossRef]
- Nair, S.; Zlotnick, A. HBV Core protein is in flux between cytoplasmic, nuclear, and nucleolar compartments. mBio 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Wang, J.C.; Pierson, E.E.; Keifer, D.Z.; Delaleau, M.; Gallucci, L.; Cazenave, C.; Kann, M.; Jarrold, M.F.; Zlotnick, A. Importin β can bind hepatitis b virus core protein and empty core-like particles and induce structural changes. PLoS Pathog. 2016, 12, e1005802. [Google Scholar] [CrossRef]
- Mitra, B.; Wang, J.; Kim, E.S.; Mao, R.; Dong, M.; Liu, Y.; Zhang, J.; Guo, H. Hepatitis B Virus Precore Protein p22 Inhibits Alpha Interferon Signaling by Blocking STAT Nuclear Translocation. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Dai, X.; Zhou, Z.H. Structure of the herpes simplex virus 1 capsid with associated tegument protein complexes. Science 2018, 360, eaao7298. [Google Scholar] [CrossRef]
- Pasdeloup, D.; Blondel, D.; Isidro, A.L.; Rixon, F.J. Herpesvirus capsid association with the nuclear pore complex and viral DNA release involve the nucleoporin CAN/Nup214 and the capsid protein pUL25. J. Virol. 2009, 83, 6610–6623. [Google Scholar] [CrossRef]
- Copeland, A.M.; Newcomb, W.W.; Brown, J.C. Herpes simplex virus replication: Roles of viral proteins and nucleoporins in capsid-nucleus attachment. J. Virol. 2009, 83, 1660–1668. [Google Scholar] [CrossRef]
- Abaitua, F.; Hollinshead, M.; Bolstad, M.; Crump, C.M.; O’Hare, P. A Nuclear localization signal in herpesvirus protein VP1-2 is essential for infection via capsid routing to the nuclear pore. J. Virol. 2012, 86, 8998–9014. [Google Scholar] [CrossRef] [PubMed]
- Ojala, P.M.; Sodeik, B.; Ebersold, M.W.; Kutay, U.; Helenius, A. Herpes simplex virus type 1 entry into host cells: Reconstitution of capsid binding and uncoating at the nuclear pore complex in vitro. Mol. Cell. Biol. 2000, 20, 4922–4931. [Google Scholar] [CrossRef] [PubMed]
- Döhner, K.; Ramos-Nascimento, A.; Bialy, D.; Anderson, F.; Hickford-Martinez, A.; Rother, F.; Koithan, T.; Rudolph, K.; Buch, A.; Prank, U.; et al. Importin α1 is required for nuclear import of herpes simplex virus proteins and capsid assembly in fibroblasts and neurons. PLoS Pathog. 2018, 14, e1006823. [Google Scholar] [CrossRef] [PubMed]
- Marsden, H.S.; Murphy, M.; McVey, G.L.; MacEachran, K.A.; Owsianka, A.M.; Stow, N.D. Role of the carboxy terminus of herpes simplex virus type 1 DNA polymerase in its interaction with UL42. J. Gen. Virol. 1994, 75, 3127–3135. [Google Scholar] [CrossRef]
- Loregian, A.; Piaia, E.; Cancellotti, E.; Papini, E.; Marsden, H.S.; Palù, G. The catalytic subunit of herpes simplex virus type 1 DNA polymerase contains a nuclear localization signal in the UL42-binding region. Virology 2000, 273, 139–148. [Google Scholar] [CrossRef]
- Alvisi, G.; Musiani, D.; Jans, D.A.; Ripalti, A. An importin alpha/beta-recognized bipartite nuclear localization signal mediates targeting of the human herpes simplex virus type 1 DNA polymerase catalytic subunit pUL30 to the nucleus. Biochemistry 2007, 46, 9155–9163. [Google Scholar] [CrossRef] [PubMed]
- Alvisi, G.; Avanzi, S.; Musiani, D.; Camozzi, D.; Leoni, V.; Ly-Huynh, J.D.; Ripalti, A. Nuclear import of HSV-1 DNA polymerase processivity factor UL42 is mediated by a C-terminally located bipartite nuclear localization signal. Biochemistry 2008, 47, 13764–13777. [Google Scholar] [CrossRef]
- Yusuf, B.; Gopurappilly, R.; Dadheech, N.; Gupta, S.; Bhonde, R.; Pal, R. Embryonic fibroblasts represent a connecting link between mesenchymal and embryonic stem cells. Dev. Growth Differ. 2013, 55, 330–340. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.W.; Malik, P.; Clements, J.B. The herpes simplex virus ICP27 protein: A multifunctional post-transcriptional regulator of gene expression. Biochem. Soc. Trans. 2005, 33, 499–501. [Google Scholar] [CrossRef]
- Sandri-Goldin, R.M. The many roles of the regulatory protein ICP27 during herpes simplex virus infection. Front. Biosci. J. Virtual Librar. 2008, 13, 5241–5256. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.H.; Li, L.; Silva, L.; Sandri-Goldin, R.M. ICP27 recruits Aly/REF but not TAP/NXF1 to herpes simplex virus type 1 transcription sites although TAP/NXF1 is required for ICP27 export. J. Virol. 2005, 79, 3949–3961. [Google Scholar] [CrossRef]
- Escudero-Paunetto, L.; Li, L.; Hernandez, F.P.; Sandri-Goldin, R.M. SR proteins SRp20 and 9G8 contribute to efficient export of herpes simplex virus 1 mRNAs. Virology 2010, 401, 155–164. [Google Scholar] [CrossRef]
- Koffa, M.D.; Clements, J.B.; Izaurralde, E.; Wadd, S.; Wilson, S.A.; Mattaj, I.W.; Kuersten, S. Herpes simplex virus ICP27 protein provides viral mRNAs with access to the cellular mRNA export pathway. EMBO J. 2001, 20, 5769–5778. [Google Scholar] [CrossRef] [PubMed]
- Malik, P.; Tabarraei, A.; Kehlenbach, R.H.; Korfali, N.; Iwasawa, R.; Graham, S.V.; Schirmer, E.C. Herpes simplex virus ICP27 protein directly interacts with the nuclear pore complex through Nup62, inhibiting host nucleocytoplasmic transport pathways. J. Biol. Chem. 2012, 287, 12277–12292. [Google Scholar] [CrossRef]
- Boruchowicz, H.; Hawkins, J.; Cruz-Palomar, K.; Lippé, R. The XPO6 Exportin Mediates Herpes Simplex Virus 1 gM Nuclear Release Late in Infection. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [PubMed]
- Walter, G.; Richert, Q.; Ponnampalam, A.; Sharma, A. Acute superior mesenteric vein thrombosis in the setting of cytomegalovirus mononucleosis: A case report and review of the literature. Lancet Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Bowen, L.N.; Smith, B.; Reich, D.; Quezado, M.; Nath, A. HIV-associated opportunistic CNS infections: Pathophysiology, diagnosis and treatment. Nat. Rev. Neurol. 2016, 12, 662–674. [Google Scholar] [CrossRef] [PubMed]
- Gill, R.B.; James, S.H.; Prichard, M.N. Human cytomegalovirus UL97 kinase alters the accumulation of CDK1. J. Gen. Virol. 2012, 93, 1743–1755. [Google Scholar] [CrossRef]
- Webel, R.; Solbak, S.; Held, C.; Milbradt, J.; Groß, A.; Eichler, J.; Wittenberg, T.; Jardin, C.; Sticht, H.; Fossen, T.; et al. Nuclear import of isoforms of the cytomegalovirus kinase pUL97 is mediated by differential activity of NLS1 and NLS2 both acting through classical importin-α binding. J. Gen. Virol. 2012, 93, 1756–1768. [Google Scholar] [CrossRef]
- Perng, Y.C.; Campbell, J.A.; Lenschow, D.J.; Yu, D. Human cytomegalovirus pUL79 is an elongation factor of RNA polymerase II for viral gene transcription. PLoS Pathog. 2014, 10, e1004350. [Google Scholar] [CrossRef]
- Wang, L.; Li, M.; Cai, M.; Xing, J.; Wang, S.; Zheng, C. A PY-nuclear localization signal is required for nuclear accumulation of HCMV UL79 protein. Med. Microbiol. Immunol. 2012, 201, 381–387. [Google Scholar] [CrossRef]
- Gao, Y.; Kagele, D.; Smallenberg, K.; Pari, G.S. Nucleocytoplasmic shuttling of human cytomegalovirus UL84 is essential for virus growth. J. Virol. 2010, 84, 8484–8494. [Google Scholar] [CrossRef]
- Lischka, P.; Sorg, G.; Kann, M.; Winkler, M.; Stamminger, T. A nonconventional nuclear localization signal within the UL84 protein of human cytomegalovirus mediates nuclear import via the importin alpha/beta pathway. J. Virol. 2003, 77, 3734–3748. [Google Scholar] [CrossRef]
- Lischka, P.; Rauh, C.; Mueller, R.; Stamminger, T. Human cytomegalovirus UL84 protein contains two nuclear export signals and shuttles between the nucleus and the cytoplasm. J. Virol. 2006, 80, 10274–10280. [Google Scholar] [CrossRef][Green Version]
- Toth, Z.; Lischka, P.; Stamminger, T. RNA-binding of the human cytomegalovirus transactivator protein UL69, mediated by arginine-rich motifs, is not required for nuclear export of unspliced RNA. Nucl. Acids Res. 2006, 34, 1237–1249. [Google Scholar] [CrossRef] [PubMed]
- Zielke, B.; Thomas, M.; Giede-Jeppe, A.; Müller, R.; Stamminger, T. Characterization of the betaherpesviral pUL69 protein family reveals binding of the cellular mRNA export factor UAP56 as a prerequisite for stimulation of nuclear mRNA export and for efficient viral replication. J. Virol. 2011, 85, 1804–1819. [Google Scholar] [CrossRef] [PubMed]
- Stamminger, T. Interactions of human cytomegalovirus proteins with the nuclear transport machinery. Curr. Top. Microbiol. Immunol. 2008, 325, 167–185. [Google Scholar] [CrossRef] [PubMed]
- Lischka, P.; Rosorius, O.; Trommer, E.; Stamminger, T. A novel transferable nuclear export signal mediates CRM1-independent nucleocytoplasmic shuttling of the human cytomegalovirus transactivator protein pUL69. EMBO J. 2001, 20, 7271–7283. [Google Scholar] [CrossRef][Green Version]
- Rechter, S.; Scott, G.M.; Eickhoff, J.; Zielke, K.; Auerochs, S.; Müller, R.; Stamminger, T.; Rawlinson, W.D.; Marschall, M. Cyclin-dependent Kinases Phosphorylate the Cytomegalovirus RNA Export Protein pUL69 and Modulate Its Nuclear Localization and Activity. J. Biol. Chem. 2009, 284, 8605–8613. [Google Scholar] [CrossRef] [PubMed]
- Münz, C. Latency and lytic replication in Epstein-Barr virus-associated oncogenesis. Nat. Rev. Microbiol. 2019, 17, 691–700. [Google Scholar] [CrossRef]
- Ito, S.; Ikeda, M.; Kato, N.; Matsumoto, A.; Ishikawa, Y.; Kumakubo, S.; Yanagi, K. Epstein-barr virus nuclear antigen-1 binds to nuclear transporter karyopherin alpha1/NPI-1 in addition to karyopherin alpha2/Rch1. Virology 2000, 266, 110–119. [Google Scholar] [CrossRef][Green Version]
- Chang, C.W.; Lee, C.P.; Su, M.T.; Tsai, C.H.; Chen, M.R. BGLF4 kinase modulates the structure and transport preference of the nuclear pore complex to facilitate nuclear import of Epstein-Barr virus lytic proteins. J. Virol. 2015, 89, 1703–1718. [Google Scholar] [CrossRef]
- Lee, C.P.; Huang, Y.H.; Lin, S.F.; Chang, Y.; Chang, Y.H.; Takada, K.; Chen, M.R. Epstein-Barr virus BGLF4 kinase induces disassembly of the nuclear lamina to facilitate virion production. J. Virol. 2008, 82, 11913–11926. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Kamil, J.P.; Coughlin, M.; Reim, N.I.; Coen, D.M. Human cytomegalovirus UL50 and UL53 recruit viral protein kinase UL97, not protein kinase C, for disruption of nuclear lamina and nuclear egress in infected cells. J. Virol. 2014, 88, 249–262. [Google Scholar] [CrossRef]
- Yang, Y.C.; Sugden, B. Epstein-Barr Virus Limits the Accumulation of IPO7, an Essential Gene Product. Front. Microbiol. 2021, 12, 643327. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Gradoville, L.; Miller, G. Cellular immediate-early gene expression occurs kinetically upstream of Epstein-Barr virus bzlf1 and brlf1 following cross-linking of the B cell antigen receptor in the Akata Burkitt lymphoma cell line. J. Virol. 2010, 84, 12405–12418. [Google Scholar] [CrossRef]
- Boyle, S.M.; Ruvolo, V.; Gupta, A.K.; Swaminathan, S. Association with the cellular export receptor CRM 1 mediates function and intracellular localization of Epstein-Barr virus SM protein, a regulator of gene expression. J. Virol. 1999, 73, 6872–6881. [Google Scholar] [CrossRef] [PubMed]
- Funk, C.; Ott, M.; Raschbichler, V.; Nagel, C.H.; Binz, A.; Sodeik, B.; Bauerfeind, R.; Bailer, S.M. The herpes simplex virus protein pUL31 escorts nucleocapsids to sites of nuclear egress, a process coordinated by its N-terminal domain. PLoS Pathog. 2015, 11, e1004957. [Google Scholar] [CrossRef]
- Gonnella, R.; Farina, A.; Santarelli, R.; Raffa, S.; Feederle, R.; Bei, R.; Granato, M.; Modesti, A.; Frati, L.; Delecluse, H.J.; et al. Characterization and intracellular localization of the Epstein-Barr virus protein BFLF2: Interactions with BFRF1 and with the nuclear lamina. J. Virol. 2005, 79, 3713–3727. [Google Scholar] [CrossRef] [PubMed]
- Schmeiser, C.; Borst, E.; Sticht, H.; Marschall, M.; Milbradt, J. The cytomegalovirus egress proteins pUL50 and pUL53 are translocated to the nuclear envelope through two distinct modes of nuclear import. J. Gen. Virol. 2013, 94, 2056–2069. [Google Scholar] [CrossRef]
- Li, M.; Chen, T.; Zou, X.; Xu, Z.; Wang, Y.; Wang, P.; Ou, X.; Li, Y.; Chen, D.; Peng, T.; et al. Characterization of the nucleocytoplasmic transport mechanisms of epstein-barr virus BFLF2. Cell. Physiol. Biochem. 2018, 51, 1500–1517. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Jiang, S.; Mo, C.; Zeng, Z.; Li, X.; Chen, C.; Yang, Y.; Wang, J.; Huang, J.; Chen, D.; et al. Identification of molecular determinants for the nuclear import of pseudorabies virus UL31. Arch. Biochem. Biophys. 2015, 587, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Cai, M.; Chen, D.; Zeng, Z.; Yang, H.; Jiang, S.; Li, X.; Mai, J.; Peng, T.; Li, M. Characterization of the nuclear import signal of herpes simplex virus 1 UL31. Arch. Virol. 2016, 161, 2379–2385. [Google Scholar] [CrossRef]
- Paßvogel, L.; Klupp, B.G.; Granzow, H.; Fuchs, W.; Mettenleiter, T.C. Functional characterization of nuclear trafficking signals in pseudorabies virus pUL31. J. Virol. 2015, 89, 2002–2012. [Google Scholar] [CrossRef]
- Funk, C.; Raschbichler, V.; Lieber, D.; Wetschky, J.; Arnold, E.K.; Leimser, J.; Biggel, M.; Friedel, C.C.; Ruzsics, Z.; Bailer, S.M. Comprehensive analysis of nuclear export of herpes simplex virus type 1 tegument proteins and their Epstein-Barr virus orthologs. Traffic 2019, 20, 152–167. [Google Scholar] [CrossRef]
- Gabriel, G.; Klingel, K.; Otte, A.; Thiele, S.; Hudjetz, B.; Arman-Kalcek, G.; Sauter, M.; Shmidt, T.; Rother, F.; Baumgarte, S.; et al. Differential use of importin-α isoforms governs cell tropism and host adaptation of influenza virus. Nat. Commun. 2011, 2, 156. [Google Scholar] [CrossRef]
- Higgs, E.S.; Gayedyu-Dennis, D.; Fisher, W.; Nason, M.; Reilly, C.; Beavogui, A.H.; Aboulhab, J.; Nordwall, J.; Lobbo, P.; Wachekwa, I.; et al. PREVAIL IV: A Randomized, Double-Blind, Two-Phase, Phase 2 Trial of Remdesivir versus Placebo for Reduction of Ebola Virus RNA in the Semen of Male Survivors. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Choopanya, K.; Martin, M.; Suntharasamai, P.; Sangkum, U.; Mock, P.A.; Leethochawalit, M.; Chiamwongpaet, S.; Kitisin, P.; Natrujirote, P.; Kittimunkong, S.; et al. Antiretroviral prophylaxis for HIV infection in injecting drug users in Bangkok, Thailand (the Bangkok Tenofovir Study): A randomised, double-blind, placebo-controlled phase 3 trial. Lancet 2013, 381, 2083–2090. [Google Scholar] [CrossRef]
- Abdool Karim, S.S.; Abdool Karim, Q.; Kharsany, A.B.; Baxter, C.; Grobler, A.C.; Werner, L.; Kashuba, A.; Mansoor, L.E.; Samsunder, N.; Mindel, A.; et al. Tenofovir Gel for the Prevention of Herpes Simplex Virus Type 2 Infection. N. Engl. J. Med. 2015, 373, 530–539. [Google Scholar] [CrossRef]
- Kosugi, S.; Hasebe, M.; Entani, T.; Takayama, S.; Tomita, M.; Yanagawa, H. Design of peptide inhibitors for the importin alpha/beta nuclear import pathway by activity-based profiling. Chem. Biol. 2008, 15, 940–949. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.Z.; Yao, S.Y.; Veach, R.A.; Torgerson, T.R.; Hawiger, J. Inhibition of nuclear translocation of transcription factor NF-kappa B by a synthetic peptide containing a cell membrane-permeable motif and nuclear localization sequence. J. Biol. Chem. 1995, 270, 14255–14258. [Google Scholar] [CrossRef]
- Zienkiewicz, J.; Armitage, A.; Hawiger, J. Targeting nuclear import shuttles, importins/karyopherins alpha by a peptide mimicking the NFκB1/p50 nuclear localization sequence. J. Am. Heart Assoc. 2013, 2, e000386. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Y.; Fraser, J.E.; Chan, W.K.; Moreland, N.J.; Rathore, A.P.; Wang, C.; Vasudevan, S.G.; Jans, D.A. Nuclear localization of dengue virus (DENV) 1-4 non-structural protein 5; protection against all 4 DENV serotypes by the inhibitor Ivermectin. Antivir. Res. 2013, 99, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Wagstaff, K.M.; Sivakumaran, H.; Heaton, S.M.; Harrich, D.; Jans, D.A. Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus. Biochem. J. 2012, 443, 851–856. [Google Scholar] [CrossRef]
- Yang, S.N.Y.; Atkinson, S.C.; Fraser, J.E.; Wang, C.; Maher, B.; Roman, N.; Forwood, J.K.; Wagstaff, K.M.; Borg, N.A.; Jans, D.A. Novel flavivirus antiviral that targets the host nuclear transport importin α/β1 heterodimer. Cells 2019, 8, 281. [Google Scholar] [CrossRef]
- Yang, S.N.Y.; Atkinson, S.C.; Wang, C.; Lee, A.; Bogoyevitch, M.A.; Borg, N.A.; Jans, D.A. The broad spectrum antiviral ivermectin targets the host nuclear transport importin α/β1 heterodimer. Antivir. Res. 2020, 177, 104760. [Google Scholar] [CrossRef] [PubMed]
- Soderholm, J.F.; Bird, S.L.; Kalab, P.; Sampathkumar, Y.; Hasegawa, K.; Uehara-Bingen, M.; Weis, K.; Heald, R. Importazole, a small molecule inhibitor of the transport receptor importin-β. ACS Chem. Biol. 2011, 6, 700–708. [Google Scholar] [CrossRef] [PubMed]
- Van der Watt, P.J.; Chi, A.; Stelma, T.; Stowell, C.; Strydom, E.; Carden, S.; Angus, L.; Hadley, K.; Lang, D.; Wei, W.; et al. Targeting the Nuclear Import Receptor Kpnβ1 as an Anticancer Therapeutic. Mol. Cancer Ther. 2016, 15, 560–573. [Google Scholar] [CrossRef] [PubMed]
- Hintersteiner, M.; Ambrus, G.; Bednenko, J.; Schmied, M.; Knox, A.J.; Meisner, N.C.; Gstach, H.; Seifert, J.M.; Singer, E.L.; Gerace, L.; et al. Identification of a small molecule inhibitor of importin β mediated nuclear import by confocal on-bead screening of tagged one-bead one-compound libraries. ACS Chem. Biol. 2010, 5, 967–979. [Google Scholar] [CrossRef] [PubMed]
- Cansizoglu, A.E.; Lee, B.J.; Zhang, Z.C.; Fontoura, B.M.; Chook, Y.M. Structure-based design of a pathway-specific nuclear import inhibitor. Nat. Struct. Mol. Biol. 2007, 14, 452–454. [Google Scholar] [CrossRef] [PubMed]
- Cai, M.; Ou, X.; Li, Y.; Zou, X.; Xu, Z.; Wang, Y.; Peng, H.; Deng, Y.; Guo, Y.; Lu, M.; et al. Molecular anatomy of the subcellular localization and nuclear import mechanism of herpes simplex virus 1 UL6. Aging 2020, 12, 5751–5763. [Google Scholar] [CrossRef] [PubMed]
- Nishi, K.; Yoshida, M.; Fujiwara, D.; Nishikawa, M.; Horinouchi, S.; Beppu, T. Leptomycin B targets a regulatory cascade of crm1, a fission yeast nuclear protein, involved in control of higher order chromosome structure and gene expression. J. Biol. Chem. 1994, 269, 6320–6324. [Google Scholar] [CrossRef]
- Kudo, N.; Matsumori, N.; Taoka, H.; Fujiwara, D.; Schreiner, E.P.; Wolff, B.; Yoshida, M.; Horinouchi, S. Leptomycin B inactivates CRM1/exportin 1 by covalent modification at a cysteine residue in the central conserved region. Proc. Nat. Acad. Sci. USA 1999, 96, 9112–9117. [Google Scholar] [CrossRef]
- Zhang, K.; Wang, M.; Tamayo, A.T.; Shacham, S.; Kauffman, M.; Lee, J.; Zhang, L.; Ou, Z.; Li, C.; Sun, L.; et al. Novel selective inhibitors of nuclear export CRM1 antagonists for therapy in mantle cell lymphoma. Exp. Hematol. 2013, 41, 67–78.e64. [Google Scholar] [CrossRef]
- Ranganathan, P.; Yu, X.; Na, C.; Santhanam, R.; Shacham, S.; Kauffman, M.; Walker, A.; Klisovic, R.; Blum, W.; Caligiuri, M.; et al. Preclinical activity of a novel CRM1 inhibitor in acute myeloid leukemia. Blood 2012, 120, 1765–1773. [Google Scholar] [CrossRef]
- Sun, Q.; Carrasco, Y.P.; Hu, Y.; Guo, X.; Mirzaei, H.; Macmillan, J.; Chook, Y.M. Nuclear export inhibition through covalent conjugation and hydrolysis of Leptomycin B by CRM1. Proc. Nat. Acad. Sci. USA 2013, 110, 1303–1308. [Google Scholar] [CrossRef]
- Lapalombella, R.; Sun, Q.; Williams, K.; Tangeman, L.; Jha, S.; Zhong, Y.; Goettl, V.; Mahoney, E.; Berglund, C.; Gupta, S.; et al. Selective inhibitors of nuclear export show that CRM1/XPO1 is a target in chronic lymphocytic leukemia. Blood 2012, 120, 4621–4634. [Google Scholar] [CrossRef]
- Haines, J.D.; Herbin, O.; de la Hera, B.; Vidaurre, O.G.; Moy, G.A.; Sun, Q.; Fung, H.Y.; Albrecht, S.; Alexandropoulos, K.; McCauley, D.; et al. Nuclear export inhibitors avert progression in preclinical models of inflammatory demyelination. Nat. Neurosci. 2015, 18, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Tai, Y.T.; Landesman, Y.; Acharya, C.; Calle, Y.; Zhong, M.Y.; Cea, M.; Tannenbaum, D.; Cagnetta, A.; Reagan, M.; Munshi, A.A.; et al. CRM1 inhibition induces tumor cell cytotoxicity and impairs osteoclastogenesis in multiple myeloma: Molecular mechanisms and therapeutic implications. Leukemia 2014, 28, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Newlands, E.S.; Rustin, G.J.; Brampton, M.H. Phase I trial of elactocin. Br. J. Cancer 1996, 74, 648–649. [Google Scholar] [CrossRef]
- Sun, Q.; Chen, X.; Zhou, Q.; Burstein, E.; Yang, S.; Jia, D. Inhibiting cancer cell hallmark features through nuclear export inhibition. Signal Transduct. Target. Ther. 2016, 1, 16010. [Google Scholar] [CrossRef]
- Wang, C.; Yang, S.N.Y.; Smith, K.; Forwood, J.K.; Jans, D.A. Nuclear import inhibitor N-(4-hydroxyphenyl) retinamide targets Zika virus (ZIKV) nonstructural protein 5 to inhibit ZIKV infection. Biochem. Biophys. Res. Commun. 2017, 493, 1555–1559. [Google Scholar] [CrossRef]
- Pitts, J.D.; Li, P.C.; de Wispelaere, M.; Yang, P.L. Antiviral activity of N-(4-hydroxyphenyl) retinamide (4-HPR) against Zika virus. Antivir. Res. 2017, 147, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.E.; Watanabe, S.; Wang, C.; Chan, W.K.; Maher, B.; Lopez-Denman, A.; Hick, C.; Wagstaff, K.M.; Mackenzie, J.M.; Sexton, P.M.; et al. A nuclear transport inhibitor that modulates the unfolded protein response and provides in vivo protection against lethal dengue virus infection. J. Infect. Dis. 2014, 210, 1780–1791. [Google Scholar] [CrossRef]
- Link, J.O.; Rhee, M.S.; Tse, W.C.; Zheng, J.; Somoza, J.R.; Rowe, W.; Begley, R.; Chiu, A.; Mulato, A.; Hansen, D.; et al. Clinical targeting of HIV capsid protein with a long-acting small molecule. Nature 2020, 584, 614–618. [Google Scholar] [CrossRef] [PubMed]
- Yant, S.R.; Mulato, A.; Hansen, D.; Tse, W.C.; Niedziela-Majka, A.; Zhang, J.R.; Stepan, G.J.; Jin, D.; Wong, M.H.; Perreira, J.M.; et al. A highly potent long-acting small-molecule HIV-1 capsid inhibitor with efficacy in a humanized mouse model. Nat. Med. 2019, 25, 1377–1384. [Google Scholar] [CrossRef]
- Mohl, G.; Liddle, N.; Nygaard, J.; Dorius, A.; Lyons, N.; Hodek, J.; Weber, J.; Michaelis, D.J.; Busath, D.D. Novel influenza inhibitors designed to target PB1 interactions with host importin RanBP5. Antivir. Res. 2019, 164, 81–90. [Google Scholar] [CrossRef]
- Tanaka, K.; Kasahara, Y.; Miyamoto, Y.; Takumi, O.; Kasai, T.; Onodera, K.; Kuwahara, M.; Oka, M.; Yoneda, Y.; Obika, S. Development of oligonucleotide-based antagonists of Ebola virus protein 24 inhibiting its interaction with karyopherin alpha 1. Org. Biomol. Chem. 2018, 16, 4456–4463. [Google Scholar] [CrossRef]
- Song, X.; Lu, L.Y.; Passioura, T.; Suga, H. Macrocyclic peptide inhibitors for the protein-protein interaction of Zaire Ebola virus protein 24 and karyopherin alpha 5. Org. Biomol. Chem. 2017, 15, 5155–5160. [Google Scholar] [CrossRef]
- Gonzalez-Sanchez, J.L.; Martinez-Chequer, J.C.; Hernandez-Celaya, M.E.; Barahona-Bustillos, E.; Andrade-Manzano, A.F. Randomized placebo-controlled evaluation of intramuscular interferon beta treatment of recurrent human papillomavirus. Obstet. Gynecol. 2001, 97, 621–624. [Google Scholar] [CrossRef] [PubMed]
- Terenzi, F.; Saikia, P.; Sen, G.C. Interferon-inducible protein, P56, inhibits HPV DNA replication by binding to the viral protein E1. EMBO J. 2008, 27, 3311–3321. [Google Scholar] [CrossRef]
- Wu, G.; Liu, B.; Zhang, Y.; Li, J.; Arzumanyan, A.; Clayton, M.M.; Schinazi, R.F.; Wang, Z.; Goldmann, S.; Ren, Q.; et al. Preclinical characterization of GLS4, an inhibitor of hepatitis B virus core particle assembly. Antimicrob. Agents Chemother. 2013, 57, 5344–5354. [Google Scholar] [CrossRef] [PubMed]
- Weber, O.; Schlemmer, K.H.; Hartmann, E.; Hagelschuer, I.; Paessens, A.; Graef, E.; Deres, K.; Goldmann, S.; Niewoehner, U.; Stoltefuss, J.; et al. Inhibition of human hepatitis B virus (HBV) by a novel non-nucleosidic compound in a transgenic mouse model. Antivir. Res. 2002, 54, 69–78. [Google Scholar] [CrossRef]
- Brezillon, N.; Brunelle, M.N.; Massinet, H.; Giang, E.; Lamant, C.; DaSilva, L.; Berissi, S.; Belghiti, J.; Hannoun, L.; Puerstinger, G.; et al. Antiviral activity of Bay 41-4109 on hepatitis B virus in humanized Alb-uPA/SCID mice. PLoS ONE 2011, 6, e25096. [Google Scholar] [CrossRef]
- Perni, R.B.; Conway, S.C.; Ladner, S.K.; Zaifert, K.; Otto, M.J.; King, R.W. Phenylpropenamide derivatives as inhibitors of hepatitis B virus replication. Bioorg. Med. Chem. Lett. 2000, 10, 2687–2690. [Google Scholar] [CrossRef]
- King, R.W.; Ladner, S.K.; Miller, T.J.; Zaifert, K.; Perni, R.B.; Conway, S.C.; Otto, M.J. Inhibition of human hepatitis B virus replication by AT-61, a phenylpropenamide derivative, alone and in combination with (-)beta-L-2′,3′-dideoxy-3′-thiacytidine. Antimicrob. Agents Chemother. 1998, 42, 3179–3186. [Google Scholar] [CrossRef] [PubMed]
- Delaney, W.E.t.; Edwards, R.; Colledge, D.; Shaw, T.; Furman, P.; Painter, G.; Locarnini, S. Phenylpropenamide derivatives AT-61 and AT-130 inhibit replication of wild-type and lamivudine-resistant strains of hepatitis B virus in vitro. Antimicrob. Agents Chemother. 2002, 46, 3057–3060. [Google Scholar] [CrossRef] [PubMed]
- Campagna, M.R.; Liu, F.; Mao, R.; Mills, C.; Cai, D.; Guo, F.; Zhao, X.; Ye, H.; Cuconati, A.; Guo, H.; et al. Sulfamoylbenzamide derivatives inhibit the assembly of hepatitis B virus nucleocapsids. J. Virol. 2013, 87, 6931–6942. [Google Scholar] [CrossRef]
- Berke, J.M.; Dehertogh, P.; Vergauwen, K.; Van Damme, E.; Mostmans, W.; Vandyck, K.; Pauwels, F. Capsid Assembly Modulators Have a Dual Mechanism of Action in Primary Human Hepatocytes Infected with Hepatitis B Virus. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef]
- Guo, F.; Zhao, Q.; Sheraz, M.; Cheng, J.; Qi, Y.; Su, Q.; Cuconati, A.; Wei, L.; Du, Y.; Li, W.; et al. HBV core protein allosteric modulators differentially alter cccDNA biosynthesis from de novo infection and intracellular amplification pathways. PLoS Pathog. 2017, 13, e1006658. [Google Scholar] [CrossRef] [PubMed]
- Ligat, G.; Goto, K.; Verrier, E.; Baumert, T.F. Targeting Viral cccDNA for Cure of Chronic Hepatitis, B. Curr. Hepatol. Rep. 2020, 19, 235–244. [Google Scholar] [CrossRef]
- Caly, L.; Druce, J.D.; Catton, M.G.; Jans, D.A.; Wagstaff, K.M. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antivir. Res. 2020, 178, 104787. [Google Scholar] [CrossRef]
- Varghese, F.S.; Kaukinen, P.; Gläsker, S.; Bespalov, M.; Hanski, L.; Wennerberg, K.; Kümmerer, B.M.; Ahola, T. Discovery of berberine, abamectin and ivermectin as antivirals against chikungunya and other alphaviruses. Antivir. Res. 2016, 126, 117–124. [Google Scholar] [CrossRef]
- King, C.R.; Tessier, T.M.; Dodge, M.J.; Weinberg, J.B.; Mymryk, J.S. Inhibition of Human Adenovirus Replication by the Importin α/β1 Nuclear Import Inhibitor Ivermectin. J. Virol. 2020, 94. [Google Scholar] [CrossRef]
- Bennett, S.M.; Zhao, L.; Bosard, C.; Imperiale, M.J. Role of a nuclear localization signal on the minor capsid proteins VP2 and VP3 in BKPyV nuclear entry. Virology 2015, 474, 110–116. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.G.; Huang, W.; Lee, H.; van de Leemput, J.; Kane, M.A.; Han, Z. Characterization of SARS-CoV-2 proteins reveals Orf6 pathogenicity, subcellular localization, host interactions and attenuation by Selinexor. Cell Biosci. 2021, 11, 58. [Google Scholar] [CrossRef] [PubMed]
- Perwitasari, O.; Johnson, S.; Yan, X.; Howerth, E.; Shacham, S.; Landesman, Y.; Baloglu, E.; McCauley, D.; Tamir, S.; Tompkins, S.M.; et al. Verdinexor, a novel selective inhibitor of nuclear export, reduces influenza a virus replication in vitro and in vivo. J. Virol. 2014, 88, 10228–10243. [Google Scholar] [CrossRef] [PubMed]
- Perwitasari, O.; Johnson, S.; Yan, X.; Register, E.; Crabtree, J.; Gabbard, J.; Howerth, E.; Shacham, S.; Carlson, R.; Tamir, S.; et al. Antiviral Efficacy of Verdinexor In Vivo in Two Animal Models of Influenza A Virus Infection. PLoS ONE 2016, 11, e0167221. [Google Scholar] [CrossRef] [PubMed]
- López-Medina, E.; López, P.; Hurtado, I.C.; Dávalos, D.M.; Ramirez, O.; Martínez, E.; Díazgranados, J.A.; Oñate, J.M.; Chavarriaga, H.; Herrera, S.; et al. Effect of ivermectin on time to resolution of symptoms among adults with mild COVID-19: A randomized clinical trial. JAMA 2021, 325, 1426–1435. [Google Scholar] [CrossRef] [PubMed]
- Elalfy, H.; Besheer, T.; El-Mesery, A.; El-Gilany, A.H.; Soliman, M.A.; Alhawarey, A.; Alegezy, M.; Elhadidy, T.; Hewidy, A.A.; Zaghloul, H.; et al. Effect of a combination of nitazoxanide, ribavirin, and ivermectin plus zinc supplement (MANS.NRIZ study) on the clearance of mild COVID-19. J. Med. Virol. 2021, 93, 3176–3183. [Google Scholar] [CrossRef]
- Ahmed, S.; Karim, M.M.; Ross, A.G.; Hossain, M.S.; Clemens, J.D.; Sumiya, M.K.; Phru, C.S.; Rahman, M.; Zaman, K.; Somani, J.; et al. A five-day course of ivermectin for the treatment of COVID-19 may reduce the duration of illness. Int. J. Infect. Dis. 2021, 103, 214–216. [Google Scholar] [CrossRef]
- Gandhi, R.T.; Lynch, J.B.; del Rio, C. Mild or Moderate Covid-19. N. Engl. J. Med. 2020, 383, 1757–1766. [Google Scholar] [CrossRef] [PubMed]
- Suputtamongkol, Y.; Avirutnan, P.; Mairiang, D.; Angkasekwinai, N.; Niwattayakul, K.; Yamasmith, E.; Saleh-Arong, F.A.; Songjaeng, A.; Prommool, T.; Tangthawornchaikul, N.; et al. Ivermectin accelerates circulating nonstructural protein 1 (NS1) clearance in adult dengue patients: A combined phase 2/3 randomized double-blinded placebo controlled trial. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Libraty, D.H.; Young, P.R.; Pickering, D.; Endy, T.P.; Kalayanarooj, S.; Green, S.; Vaughn, D.W.; Nisalak, A.; Ennis, F.A.; Rothman, A.L. High circulating levels of the dengue virus nonstructural protein NS1 early in dengue illness correlate with the development of dengue hemorrhagic fever. J. Infect. Dis. 2002, 186, 1165–1168. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Meng, Y.L.; Duan, S.M.; Zhan, S.B.; Guan, R.L.; Yue, T.F.; Kong, L.H.; Zhou, L.; Deng, L.H.; Huang, C.; et al. REBACIN® as a noninvasive clinical intervention for high-risk human papillomavirus persistent infection. Int. J. Cancer 2019, 145, 2712–2719. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Hu, T.; Ming, X.; Yang, E.; Min, W.; Li, Z. REBACIN® is an optional intervention for persistent high-risk human papillomavirus infection: A retrospective analysis of 364 patients. Int. J. Gynaecol. Obstet. 2021, 152, 82–87. [Google Scholar] [CrossRef]
- Yuen, M.F.; Gane, E.J.; Kim, D.J.; Weilert, F.; Yuen Chan, H.L.; Lalezari, J.; Hwang, S.G.; Nguyen, T.; Flores, O.; Hartman, G.; et al. Antiviral Activity, Safety, and Pharmacokinetics of Capsid Assembly Modulator NVR 3-778 in Patients with Chronic HBV Infection. Gastroenterology 2019, 156, 1392–1403. [Google Scholar] [CrossRef]
- Zhang, H.; Wang, F.; Zhu, X.; Chen, Y.; Chen, H.; Li, X.; Wu, M.; Li, C.; Liu, J.; Zhang, Y.; et al. Antiviral Activity and Pharmacokinetics of the HBV Capsid Assembly Modulator GLS4 in Patients with Chronic HBV Infection. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Chaccour, C.; Casellas, A.; Blanco-Di Matteo, A.; Pineda, I.; Fernandez-Montero, A.; Ruiz-Castillo, P.; Richardson, M.A.; Rodríguez-Mateos, M.; Jordán-Iborra, C.; Brew, J.; et al. The effect of early treatment with ivermectin on viral load, symptoms and humoral response in patients with non-severe COVID-19: A pilot, double-blind, placebo-controlled, randomized clinical trial. EClinicalMedicine 2021, 32, 100720. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M.S.; Kobayashi, A.; Taoka, A.; Watanabe-Nakayama, T.; Kikuchi, Y.; Hazawa, M.; Minamoto, T.; Fukumori, Y.; Kodera, N.; Uchihashi, T.; et al. High-speed atomic force microscopy reveals loss of nuclear pore resilience as a dying code in colorectal cancer cells. ACS Nano 2017, 11, 5567–5578. [Google Scholar] [CrossRef] [PubMed]
- Nishide, G.; Lim, K.; Mohamed, M.S.; Kobayashi, A.; Hazawa, M.; Watanabe-Nakayama, T.; Kodera, N.; Ando, T.; Wong, R.W. High-speed atomic force microscopy reveals spatiotemporal dynamics of histone protein H2A involution by DNA inchworming. J. Phys. Chem. Lett. 2021, 12, 3837–3846. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.; Kodera, N.; Wang, H.; Mohamed, M.S.; Hazawa, M.; Kobayashi, A.; Yoshida, T.; Hanayama, R.; Yano, S.; Ando, T.; et al. High-Speed AFM reveals molecular dynamics of human influenza A hemagglutinin and its interaction with exosomes. Nano Lett. 2020, 20, 6320–6328. [Google Scholar] [CrossRef]
- Lim, K.S.; Mohamed, M.S.; Wang, H.; Hazawa, M.; Kobayashi, A.; Voon, D.C.; Kodera, N.; Ando, T.; Wong, R.W. Direct visualization of avian influenza H5N1 hemagglutinin precursor and its conformational change by high-speed atomic force microscopy. Biochim. Biophys. Acta. Gen. Subj. 2020, 1864, 129313. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).