Abstract
Serine proteases are a class of enzymes that orchestrate an immune response. These proteases can be hijacked by viruses to facilitate entry and spread, while simultaneously supporting the innate immune system in neutralizing pathogens. This review highlights the dual roles of exogenous serine proteases, emphasizing neutrophil serine proteases (NSPs) that facilitate viral entry and promote disease progression while also contributing to antiviral defense by degrading viral glycoproteins. Additionally, the potential to modulate serine protease activity to boost host defenses will be discussed, offering both significant challenges and new opportunities for therapeutic intervention.
1. Introduction
Serine proteases constitute a class of enzymes that serve as essential orchestrators of the immune response, maintaining a delicate balance within the host–pathogen interface. These proteases are characterized by a paradoxical duality: on the one hand, viruses subvert serine proteases to facilitate cellular entry and propagate infection, while on the other hand, they enable the innate immune system to mount effective defense mechanisms [1,2,3,4,5]. We will explore the complex and dichotomous capacity of exogenous serine proteases in host defense, with a particular focus on neutrophil-derived serine proteases. The discussion concludes with an analysis of the therapeutic potential of controlling serine protease activity, aiming to augment host defenses.
1.1. Biochemical Nature of Serine Proteases
Biochemically, serine proteases hydrolyze peptide bonds by employing a catalytic triad that generally consists of a serine, histidine, and aspartate residues. The serine amino acid in the catalytic center acts as a nucleophile, attacking the carbonyl carbon of the scissile peptide bond, while the histidine residue is responsible for the general base catalysis that facilitates the transfer of a proton from the catalytically active serine residue, and the aspartate moiety stabilizes the positively charged histidine [6]. Serine proteases typically exhibit a specific cleavage site, recognizing distinct amino acid sequences adjacent to the peptide bond, which is hydrolyzed. These catalytic enzymes are also categorized into families based on substrate specificity, such as trypsin-like (which cleaves at lysine or arginine residues) and chymotrypsin-like (which preferentially cleaves after aromatic amino acids) [6,7,8].
1.2. The Role of Neutrophils and Their Serine Proteases in Immunity
During acute inflammation, particularly in an acute inflammatory response, activated neutrophils release granules packed with serine proteases known as neutrophil serine proteases (NSPs) [9]. NSPs encompass neutrophil elastase (NE), cathepsin G (CatG), proteinase 3 (PR3), and neutrophil serine protease 4 (NSP4), which are tightly regulated by endogenous serine protease inhibitors, commonly known as serpins. Serpins play a crucial role in maintaining homeostasis in order to prevent excessive tissue damage and ensure a balanced inflammatory response [10,11,12]. Generally, serine proteases are integral in activating immune responses by processing precursor proteins; however, viruses can hijack serine proteases to maintain productive infection [3,5,13].
Once released, NSPs can transiently bind to the cell surface of a panel of different cells, enabling them to modulate an immune response in the local microenvironment [2,7]. For instance, CatG, NE, and to a lesser extent PR3 are basic proteins, positively charged amino acid residues, which allow them to bind electrostatically to the anionic sulfate groups of heparan sulfate and chondroitin sulfate proteoglycans, as shown for neutrophils [14]. Additionally, PR3 can utilize general charge-dependent electrostatic interactions with negatively charged proteoglycans; PR3 binds to the cell surface through a cluster of differentiation 177 (CD177) and phosphatidylserine [15,16,17]. When a virus approaches the cellular surface where NSPs are proteolytically active, the resulting interactions are complex and can be either deleterious to the host cell or protective, depending on the virus type, the timing of the interaction, the microenvironment, the specific protease involved, or the local protease concentration. For instance, low concentrations of NSPs can cleave host or viral surface molecules, thereby promoting viral attachment, fusion, internalization, and effectively increasing infectivity. In contrast, high protease activity or a pro-inflammatory environment can cause proteolytic degradation of viral particles, leading to shedding of viral receptors and activation of innate immune signaling pathways that have antiviral effects [2,3,10].
Recent studies have elucidated the critical involvement of cell surface serine proteases in viral pathogenesis and host antiviral defense mechanisms, as summarized in the following sections.
2. Proteases That Prime Viral Proteins to Facilitate Infection of the Host Cell
For many enveloped viruses, entry into a host cell requires a priming step where the proteolytic cleavage of a viral surface glycoprotein triggers membrane fusion. The localization of specific host proteases can significantly influence tissue tropism and viral spread. For example, some strains harbor a polybasic furin cleavage site within the fusion protein, which is cleaved by the ubiquitously expressed serine protease furin, allowing the virus to spread systemically and become more virulent [18].
Furin-mediated cleavage of the fusion protein has been observed across a wide range of enveloped virus families, including Orthomyxoviridae, such as the influenza A virus, which uses host furin to cleave viral glycoproteins, aiding entry into target cells by enabling fusion with the cell membrane [19]. Similarly, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which causes coronavirus disease 2019 (COVID-19), relies on host cell serine proteases, such as furin that cleaves the spike protein (S protein) at the polybasic S1/S2 boundary during viral assembly, and membrane-anchored serine proteases, including the cell surface transmembrane protease serine subtype 2 (TMPRSS2) which catalyzes the hydrolysis of the peptide bond at the S2′ site, generating the fusion peptide (FP), thereby facilitating cellular entry, and subsequent replication [3,20].
The next section will explore the specific molecular mechanisms that enable viral fusion and entry into host cells for selected viruses.
2.1. Influenza Virus
Priming the hemagglutinin (HA) protein through cleavage by host serine proteases is an essential step for influenza viral productive infection. Without this proteolytic cleavage, the HA protein stays in a non-functional precursor trimer (HA0), which cannot trigger the viral membrane fusion needed for the virus to enter a host cell. The influenza virus relies on host serine proteases to cleave the HA0 precursor into two subunits, HA1 and HA2, at the single cleavage site (CS). The cleavage separates the receptor-binding HA1 from the hydrophobic FP at the N-terminus of the HA2 subunit [21].
The membrane-anchored TMPRSS2, along with other proteases, is the main HA activator in human respiratory epithelium [22]. It has been demonstrated that genetic deletion of TMPRSS2 in mice provides resistance to lethal H1N1 and H7N9 influenza A strains, emphasizing its critical role in vivo [23]. Other types of TTSPs, such as TMPRSS4 and TMPRSS11D, also cleave HA, which depends on viral subtype and tissue distribution [24]. For seasonal influenza viruses and low-pathogenic avian influenza (LPAI) viruses, the HA0 protein is usually cleaved extracellularly by TMPRSS2 and TMPRSS11D. Controlled severity of influenza A virus disease and limited viral spread were seen in mice lacking TMPRSS2. Other TTSPs, such as TMPRSS4 and TMPRSS11D, can partially compensate in certain subtypes. Matriptase (ST14), another TTSP, also cleaves HA, alongside TMPRSS2 and TMPRSS4 [24]. On the other hand, highly pathogenic avian influenza (HPAI) viruses have an HA0 precursor with a polybasic cleavage site, which is cleaved by the ubiquitously expressed furin protease, allowing the viruses to infect a broader range of tissues and contribute to higher pathogenicity [25]. In conclusion, the LPAI virus, which has a monobasic HA0 cleavage site for cell surface TMPRSS2 and TMPRSS11D, is restricted to infecting respiratory and intestinal tissues, while HPAI viruses can spread to many organs due to the intracellular polybasic cleavage site for furin, causing severe, widespread disease.
2.2. HIV
The pro-viral effect of CatG, with a similar but less significant outcome by NE, promotes the multiplication of HIV-1 within macrophages. These immune cells exhibit increased susceptibility to acute HIV infection following pretreatment with CatG, whereas CD4+ T cells remain unaffected. It was suggested that CatG interacts with cell surface receptors or activates specific signaling pathways that prime macrophages for more efficient viral infection. The mechanism relies on Gi-protein-dependent signaling mechanisms. Furthermore, CatG functions as a chemoattractant, recruiting monocytes and macrophages to sites of inflammation, thereby augmenting the pool of potential target cells for HIV. In summary, while CatG engages with host cellular factors, CatG does not directly interact with the viral particles [26]. Notably, furin controls viral activation by directly maturing the HIV-1 envelope glycoprotein 160 (gp160). Unlike the cell-surface triggers needed by influenza, this proteolytic cleavage into the receptor-binding (gp120) and transmembrane (gp41) subunits happens inside the producer cell within the secretory pathway [27] and renders the virus independent from further proteolytic activation upon entry into a new host cell.
2.3. The Hepatitis C Virus (HCV)
HCV infection of hepatocytes is a tightly regulated process that depends on coordinated interactions with attachment factors and essential entry factors, such as CD81, scavenger receptor class B type I, claudin-1, and occludin, followed by clathrin-mediated endocytosis [28]. Unlike influenza HA or coronavirus S protein, HCV entry is primed by receptor engagement at the cell surface for subsequent low-pH-dependent fusion in endosomes, usually without a single required protease cleavage step at the cell membrane [29,30]. However, HCV can use host cell surface serine proteases to assist viral entry and modulate the immune response. Among these, the activity of a trypsin-like serine protease (TMPRSS2 in this case) activates HCV infection at the post-binding and entry stages through proteolytic activation of the HCV envelope glycoprotein E2, which promotes fusion and entry into hepatocytes [31]. Additionally, HCV infection can modify the protease-protease inhibitor balance in infected cells. Notably, hepatitis B and hepatitis C virus replication increases the expression of serine protease inhibitor Kazal-type 1, leading to cellular resistance to serine protease-dependent apoptosis [32]. While the general HCV invasion process is mainly driven by host cell pathways, the activity of cell surface serine proteases can still impact infection efficiency at the initial stage, influencing viral spread and disease progression [31,32].
2.4. SARS-CoV-2 S Protein Facilitates Entry into the Host Cell
SARS-CoV-2 relies on host cell surface receptor angiotensin-converting enzyme 2 (ACE2) and serine proteases for entry into target cells, which is mediated by the S protein. The S protein consists of two functional sites, the S1 for binding to ACE2 and the S2′ for fusion of the viral envelope with the host cell membrane. The S1/S2 interface at 685RS686 (furin cleavage motive is RRAR/S) is hydrolyzed by the serine protease furin, and the resulting S1 subunit is non-covalently associated with the remaining S2 part, which was generated in the infected producer cells before the virions were released [33,34,35]. In contrast, the S2′ part at 815RS816 is cleaved by TMPRSS2, generating the hydrophobic FP that mediates SARS-CoV-2 fusion with the host cell membrane [36]. Beyond the canonical S protein priming by furin and TMPRSS2, several other host proteases have been reported to be co-opted, depending on the tissue and viral variant, including TMPRSS1, TMPRSS4, TMPRSS11D, TMPRSS13, and matriptase [37,38,39,40,41]. In an alternative entry mechanism, SARS-CoV-2 can access the target cell through a TMPRSS2-independent endocytosis pathway, where the cysteine protease cathepsin L (CatL) catalyzes the cleavage of the S2′ site, thereby facilitating the fusion of SARS-CoV-2 with the endosomal membrane and the subsequent release of its genetic material into the cytosol [3,42,43].
Besides TMPRSS2 and furin, which regulate coronavirus entry into host cells, NE has also been shown to modulate viral entry and the immune response at the airway surface. Originally, it was identified that porcine elastase can activate the SARS-CoV-1 S protein, playing a critical role in fusion activation [44,45], and a recent study by Yamamoto et al. demonstrated that porcine elastase is capable of cleaving the S protein (SARS-CoV-2) of bat-derived sarbecoviruses RaTG13 and Khosta-2, promoting syncytium formation and enhancing TMPRSS-2-dependent entry into human cells [46]. These findings demonstrate that neutrophil-rich inflamed airways and NE deposition at the epithelial surface can create a proteolytic microenvironment that favors S protein activation for viral fusion with the host cell membrane.
2.5. Furin Also Cleaves at S2′ and Generates the Fusion Peptide
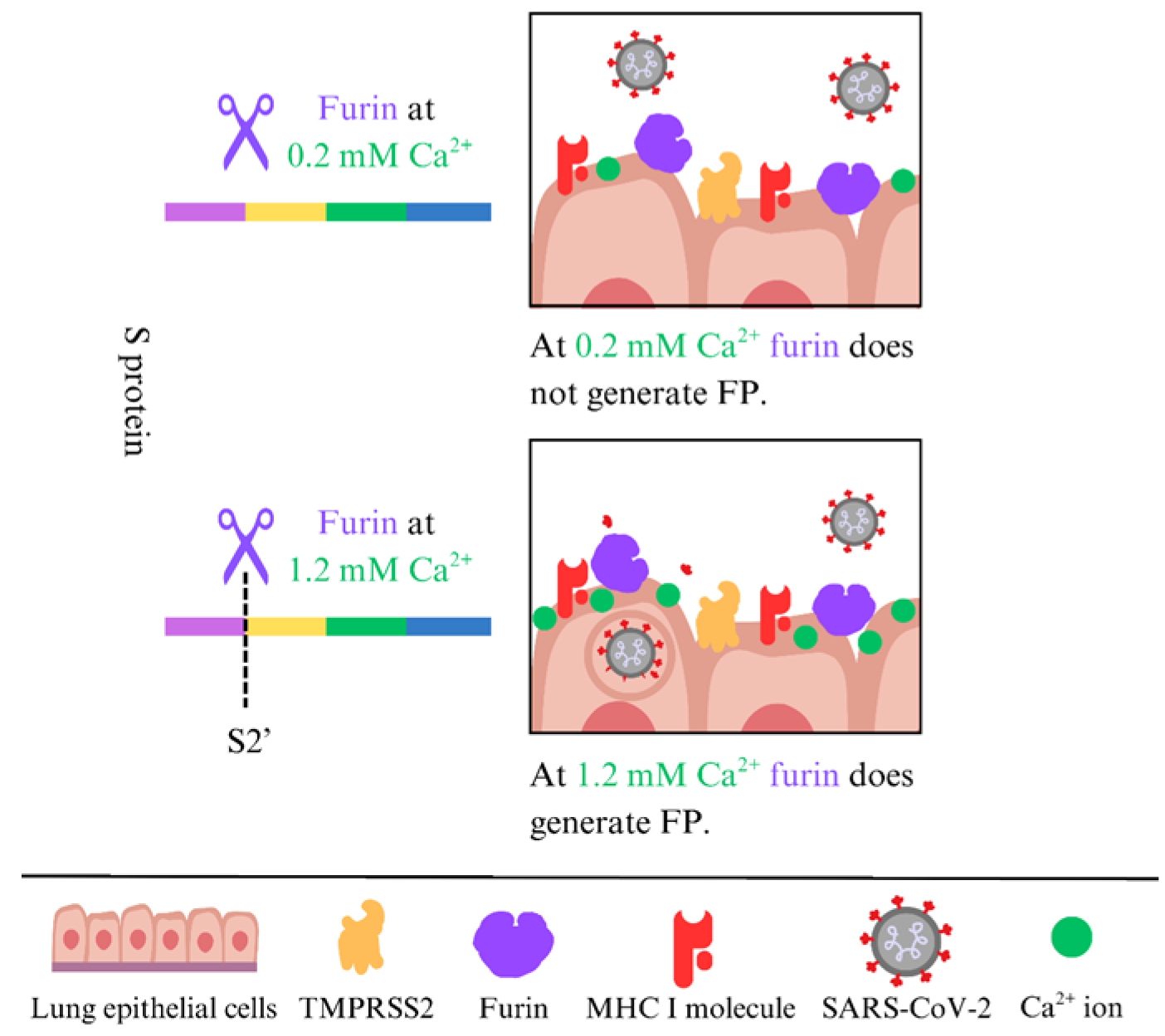
The peptide bond at the S1/S2 interface 685RS686 is hydrolyzed by furin [47]. Recently, we demonstrated that furin, with a final concentration of 0.2 mM Ca2+, cleaves at the S1/S2 interface at 685RS686. However, the S2′ site at 815RS816, which generates the hydrophobic FP, is not cleaved by furin at 0.2 mM Ca2+; higher concentrations are needed (1.2 mM) [48] (Figure 1). Generally, the level of free Ca2+ in the extracellular fluid is about 1 mM, compared to the resting intracellular level of roughly 100 nM [49]. These data indicate that only the S1/S2 interface at 685RS686 is hydrolyzed by furin, and the resulting S1 subunit remains non-covalently attached to the remaining S2 part, which is generated in the infected producer cells before the virions are released [33,34,35]. From the trans-Golgi network, where furin resides, furin can traffic through endosomal compartments to the cell surface and is tethered by actin-binding protein-280 [50,51]. At the cell surface, TMPRSS2 is the primary protease that generates the fusion peptide, but furin can also cleave at the S2′ site, as can TMPRSS2 [52].
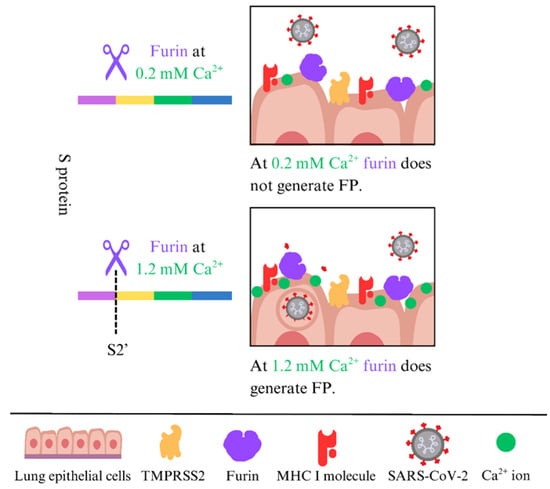
Figure 1.
Cleavage of the S2′ site by furin depends on the Ca2+ concentration.
3. Innate Immunity Involves Proteases Targeting and Neutralizing the Virus, Thereby Safeguarding the Host Cell
Conversely, the innate immune system, as the first line of defense, has developed strategies to counter pathogen invasion. For example, neutrophils secrete serine proteases that proteolytically degrade viral glycoproteins, neutralizing pathogens and preventing them from spreading within the body [53]. The principal defensive function of cell surface NSPs against viral pathogens is the capacity to facilitate the proteolytic degradation of viral constituents, thus providing protection against infection [54]. The mechanism will be further elucidated in the following section.
3.1. Influenza Virus
During influenza infection, neutrophils are rapidly recruited to the infected airway and contribute to antiviral defense through degranulation and the formation of neutrophil extracellular traps (NETs). NETs are reticular extracellular structures made of DNA, histones, and antimicrobial proteins, including NE, CatG, and myeloperoxidase (MPO), and they are released in response to viral infection [55]. NE contributes to antiviral defense primarily by driving NET formation. When activated, NE is released from azurophilic granules and translocates to the nucleus, where it cleaves histones and causes chromatin decondensation, with MPO assisting in this process [56]. Following NET release, NE and MPO are externalized within the NET scaffold at the epithelial surface, creating a localized protease-rich microenvironment that traps viral particles and can concentrate antimicrobial factors, thereby limiting local spread and subsequent entry into neighboring target cells [55]. However, a protease-rich NET response can become harmful when excessive or sustained. In influenza pneumonitis models, marked neutrophil influx and NET accumulation colocalize with tissue injury and associate with the disruption of the alveolar-capillary barrier, indicating that NET-associated enzymes and oxidant systems can aggravate lung pathology during dysregulated inflammation [57]. Thus, localized proteolysis may restrict viral spread early on, but prolonged or excessive activity can cause tissue damage and worsen disease severity beyond direct viral cytopathicity [57].
3.2. HIV
The V3 loop of HIV gp120 inhibits membrane-associated CatG on U-937 cells [58,59], and CatG has been shown to modulate HIV infection by processing the co-receptor for HIV RANTES/CCL5 at the N-terminus, generating a truncated 4-68 variant with decreased chemotactic activity and the ability to block viral binding, thereby preventing HIV from entering host cells [60], serving as an immune evasion mechanism. Prolonged or continuous exposure to CatG subsequent to the initial infection suppresses HIV replication within macrophages, indicating that CatG can function as an inhibitor of HIV infection. However, as discussed in Section 2.2, the role of CatG seems to be contingent upon specific contextual factors, notably the precise timing and duration of exposure. In fact, extended exposure to CatG underscores the critical significance of regulating CatG’s impact on HIV replication [26].
3.3. Proteases That Degrade the S Protein of SARS-CoV-2 to Block Cell Entry
Exogenous expression of TMPRSS11D enhances the SARS-CoV-2 life cycle [38], whereas soluble mature TMPRSS11D in the extracellular environment is linked to antiviral activity; however, the emergence of new variants (Delta and Omicron) has shown resistance to the anti-SARS-CoV-2 capacity of TMPRSS11D [61]. TMPRSS11D can serve as both a key factor in viral spread and an innate defense molecule, depending on its form, location, and the specific viral strain.
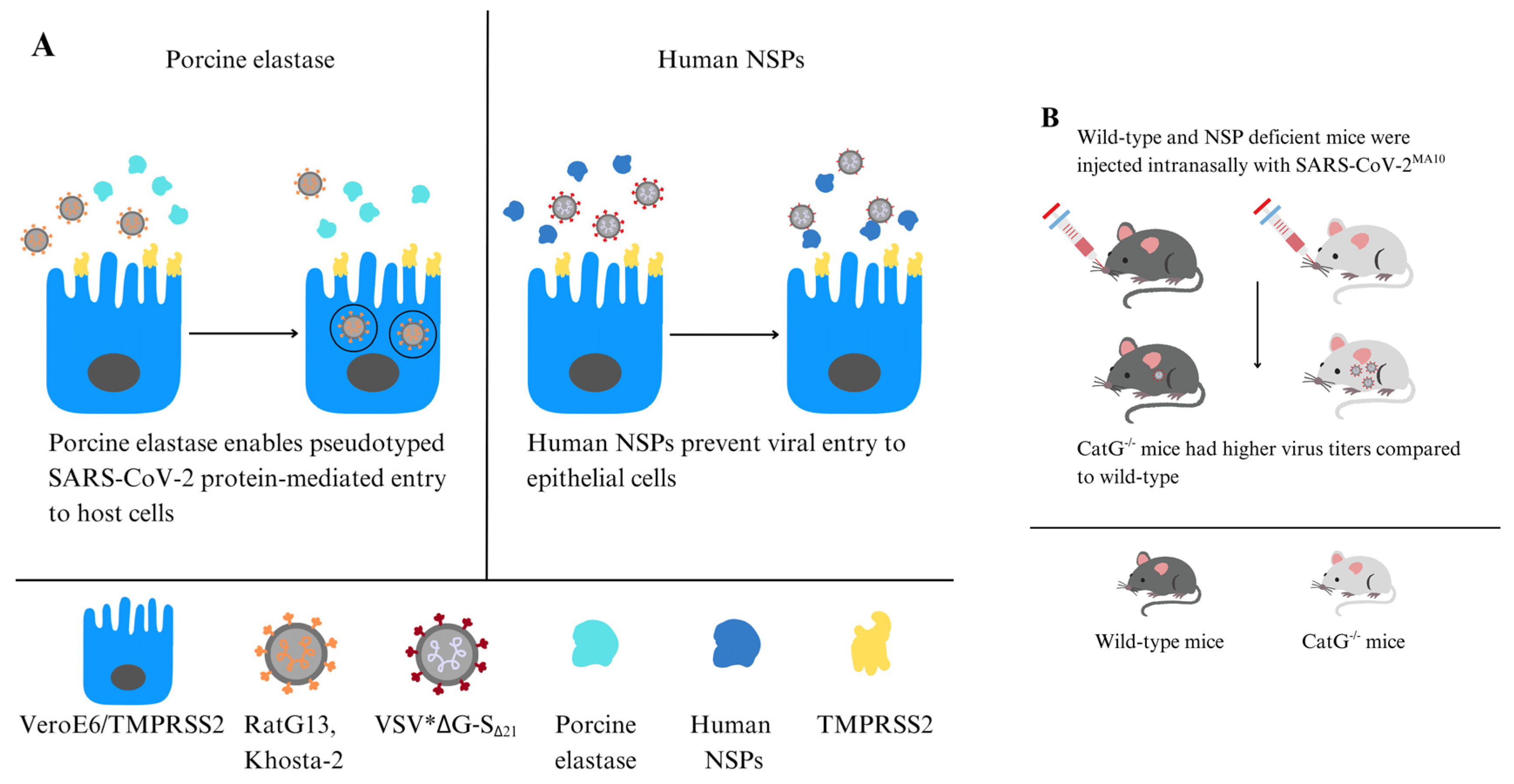
In the innate immune system, the mechanism by which serine proteases degrade the FP underscores their potential role in antiviral defense. NE and CatG can directly interact with the viral glycoprotein by cleaving the S protein, effectively rendering the virus inactive, as enzymatic activity disrupts the viral fusion process. Indeed, surface proteases assist in defending against viral infections, as the variability in SARS-CoV-2 infection outcomes between individuals may be attributed to differences in proteases responsible for viral degradation [62]. Benarafa and colleagues showed that NSPs degrade the SARS-CoV-2 S protein, reducing its stability, and inhibit viral entry into host cells, such as Vero cells expressing TMPRSS2 (Figure 2A). The absence of NE and PR3 directly affected the signs associated with SARS-CoV-2MA10 infection, as NE-/- and PR3-/- mice experienced significantly greater weight loss compared to WT mice after infection. CatG-/- mice had notably higher viral titers in the lungs than WT mice. Double knockout mice, NE-/- and CatG-/-, were the most severely affected, showing the greatest weight loss, increased inflammation, and lung tissue damage, indicating antiviral and anti-inflammatory roles of NSPs during SARS-CoV-2 infection. NE can also cleave other cell surface proteins, such as the ACE2 receptor used by SARS-CoV-2 to enter the host cell. This cleavage reduces the availability of receptors for viral binding, further hindering viral entry. In vivo studies showed that NE knockout (-/-) and CatG-/- mice increased pulmonary inflammation and more severe lung pathology than wild-type controls [54] (Figure 2B).
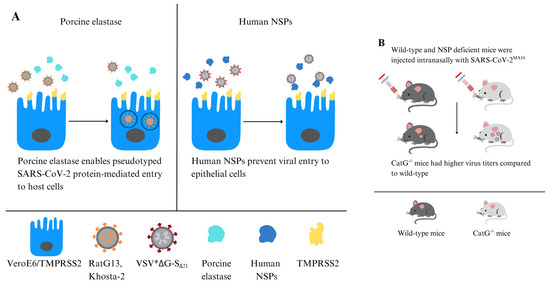
Figure 2.
NE can enhance or inhibit SARS-CoV-2 infection. (A) Porcine elastase facilitates host cell entry mediated by pseudo-typed SARS-CoV-2 S protein (left panel) [46]. NSPs digest the SARS-CoV-2 S protein, while CatG prevents the virus from entering host cells in Vero cells expressing TMPRSS2 (right panel). (B) NE-/- and PR3-/- mice experienced significant weight loss compared to WT mice after CoV-2MA10 infection. CatG-/- mice showed increased viral titers in the lungs in contrast to controls [54].
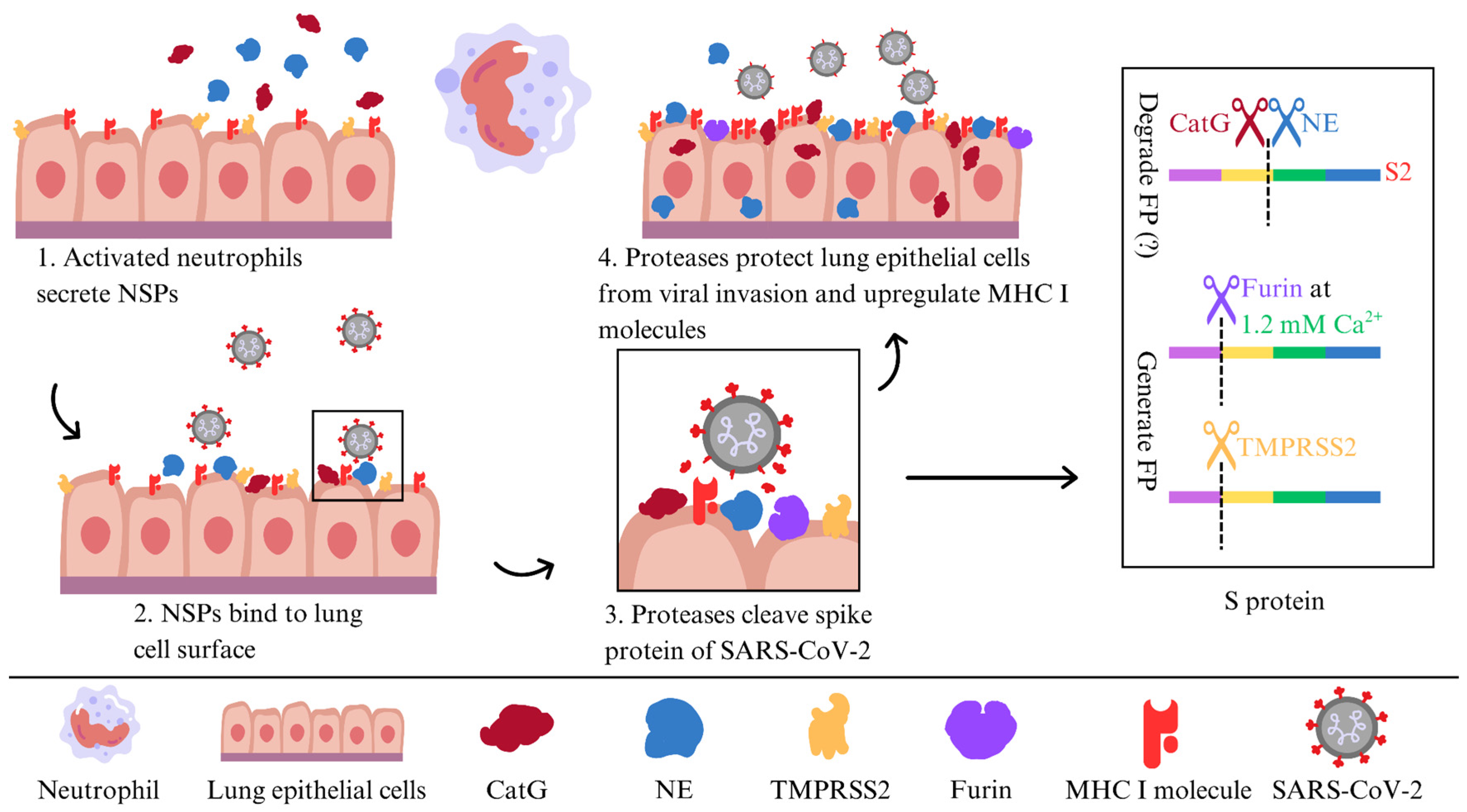
Furthermore, we revealed the pivotal role of NE and CatG in fortifying target cells, possibly against viral entry, through their binding capacity and being proteolytically active on the surface of the human alveolar basal epithelial cell line (A549) and the human lung epithelial-like cell line (H1299). Additionally, NE and CatG increase cell-surface levels of major histocompatibility complex class I (MHC I) molecules, which are crucial for presenting the endogenous antigenic repertoire to T cells. Moreover, our findings indicate that NE and CatG degrade the FP of SARS-CoV-2, as demonstrated by using a peptide spanning the S2′ site [48] (Figure 3). The augmentation of MHC I surface expression potentially arises from distinct mechanisms: NE is internalized via clathrin-mediated endocytosis [63] and might slow down MHC I turnover, leading to increased cell surface MHC I molecules [64,65], whereas CatG probably interacts with the protease-activated receptor 1 (PAR1) to promote recycling of MHC I back to the cell surface [66]. Another possibility is that serine proteases can modulate host cell receptors that viruses use for entry. By cleaving these receptors, serine proteases can hinder viral binding and decrease susceptibility to infection. Some serine proteases also regulate apoptotic pathways in response to viral infections. By regulating the apoptotic process, these enzymes can affect the viability of infected cells and constrain viral dissemination [53].
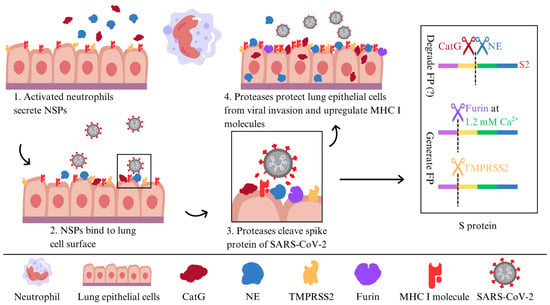
Figure 3.
A model of NE and CatG binding to the surface of lung epithelial cells to defend against SARS-CoV-2 infection.
3.4. NSPs Can Either Promote or Prevent Infection by SARS-CoV-2
Contradictory evidence has been reported regarding NE. On one hand, porcine elastase facilitates host cell entry mediated by pseudo-typed SARS-CoV-2 S protein and exhibits proviral effects [46]. Interestingly, porcine elastase also supports TMPRSS2 priming of bat RaTG13 and Khosta-2 (pseudo-typed bat SARS-like CoV) for viral fusion with the membrane, especially since low TMPRSS2 expression does not show this effect. Consequently, porcine elastase and TMPRSS2 synergistically facilitate cell entry mediated by the S proteins of bat RaTG13 and Khosta-2 [46] (Figure 2A). Moreover, neutrophil–epithelial interactions induce a pro-inflammatory state, enhancing SARS-CoV-2 infectivity by compromising epithelial barriers and exposing basal cells. Neutrophils can indirectly promote viral spread by weakening tissue defenses [67].
On the other hand, NE and CatG are defenders against viral infection as discussed in Section 3.3 [54]. Additionally, it has been shown that NSPs modulate the initial attachment phase of SARS-CoV-2 by targeting the viral receptor rather than directly interacting with the S protein, due to NE cleavage of the ACE2 ectodomain on human bronchial epithelial cells, thereby decreasing S protein binding affinity. Consequently, this reduction hampers viral attachment and entry [68]. This indicates a protective mechanism in which increased levels of NE in airway secretions trigger receptor shedding, thereby reducing viral infectivity.
NE and CatG have multiple cleavage sites within the S protein [54,69,70]. Besides the importance of the cleavage site itself, the three-dimensional structure of the S protein trimer (accessible by proteases), the timing of protease action, the source of proteases (which should be of human origin when using human host cells), species of host cells, the precise cleavage site 815RS816 to generate the FP (which is not the case for NE or CatG), and the specific SARS-CoV strain used are also significant. Even though the virus undergoes mutations that result in amino acid substitutions, eliminating all cleavage sites that could reduce the destructive capacity of NE or CatG might be unlikely. Thus, it is highly probable that NE and CatG are responsible for degrading the SARS-CoV-2 S protein, thereby impeding viral entry into host cells. This is consistent with the innate immune response, wherein neutrophils release NSPs and can bind to the surface of host cells, serving as a protective mechanism against SARS-CoV-2 infection.
During an innate immune response, administering proteases as prospective therapeutic agents or components that boost the catalytic efficiency of NSPs, provoking a transiently elevated proteolytic activity of NE and CatG, may be advantageous in neutralizing the virus; however, excessive enzymatic activity can precipitate pathological conditions in severe cases. For example, high levels of CatG are associated with detrimental inflammation and thrombogenesis, which are pivotal features of severe COVID-19 complications [71,72].
3.5. T Cells Are Armed with CatG, Possibly to Defend Themselves Against Viruses at the Site of Infection
CatG is present on the surface of immune cells, including T cells [73,74]. CatG bound to the surface of T cells may act as a protective barrier, potentially reducing the cells’ susceptibility to viral infections. Moreover, NE and CatG significantly increase the expression of MHC I molecules on a CD4+ T cell line (Jurkat cells), suggesting possible differences in how antigenic peptides are presented compared to other cell types, and imply that immune cells in transit may need enhanced surveillance to prevent viral spread [48]. Nevertheless, studies have shown that SARS-CoV-2 can directly infect human CD4+ T helper cells and cause their depletion in COVID-19, but not CD8+ T cells, indicating that viral entry is mediated by the CD4 molecule [75]. It is noteworthy that T cells do not express the cell surface protease TMPRSS2 [76], but T cells do express furin [18,77], resulting in the generation of the FP. One plausible explanation for infection is the imbalance of serpin concentrations in the microenvironment, which might inhibit cell-bound NSPs on T cells. Levels of serpins during inflammation and their inhibitory capacity will also be discussed in the subsequent section.
4. Long COVID, Potentially Regulated by NSPs and Serpins
Long COVID refers to the lingering symptoms experienced by individuals after their initial recovery from COVID-19, which has raised concerns about the long-term impact of SARS-CoV-2. Long COVID is characterized by symptoms that can persist for weeks or months after the initial infection. These symptoms include conditions such as cardiovascular, thrombotic, and cerebrovascular disease, as well as myalgic encephalomyelitis/chronic fatigue syndrome and postural orthostatic tachycardia syndrome [78,79]. The severity and duration of these symptoms can vary greatly between individuals, making it a complex and challenging condition to manage in daily life. The pathogenesis of Long COVID has been linked to hypotheses including persistent SARS-CoV-2 in tissues, reactivation of underlying pathogens (Epstein–Barr virus and human herpesvirus 6), autoimmunity, and immune dysregulation [79]. Notably, pharmacological inhibition of NE with sivelestat significantly reduces chronic lung inflammation [80], and NE, S100B, and related inflammatory markers are strongly linked to long-term respiratory dysfunction in patients months after SARS-CoV-2 infection [81]. Elevated levels of CatG and uncontrolled release of CatG by neutrophils contribute to detrimental inflammation and thrombogenesis, which are characteristic outcomes of severe COVID-19 [71,72].
A critical aspect of the immune response to SARS-CoV-2 involves protease-mediated protein degradation, with neutrophils initiating a cellular immune response against pathogens. Of particular interest is the role of NE and CatG, which can bind to the cell surface and exhibit proteolytic activity to fortify target cells [48] against viral invasion [54]. The protective function of the serine protease inhibitor SerpinA1 (also called alpha1-antitrypsin) in COVID-19 has been discussed, although this remains a topic of controversy [82,83]. In general, SerpinA1 is known to inhibit CatG, NE, PR3, and TMPRSS2, and is typically upregulated in response to managing inflammation and tissue damage, which is key after the immune system returns to homeostasis. Given the controversy surrounding the role of SerpinA1 in COVID-19, it is important to delve deeper into the regulation of serpin protease inhibitors, especially SerpinA3, which is a more potent CatG inhibitor and has been found to be upregulated in critical COVID-19 patients [84,85]. The dysregulation of these protease inhibitors, particularly during viral infections, raises intriguing questions about their potential impact on disease severity. One scenario to examine more closely is a potential decrease in SerpinA3 levels during viral infections, particularly in asymptomatic individuals, which could lead to reduced inhibition of CatG to enhance CatG-mediated viral clearance [48].
Besides proteolytic functions, genetic polymorphisms in serine protease genes influence host susceptibility to severe viral infections. A cross-sectional investigation involving over 1500 COVID-19 patients identified several single-nucleotide polymorphisms (SNPs) within the TMPRSS2 and SERPINE1 genes. The TMPRSS2 rs75603675 AA genotype has been correlated with increased mortality risk, whereas specific SERPINE1 variants, such as rs2227667, appeared to exert a protective effect. In contrast, the SERPINE1 rs2227692 T allele and TT genotype increased the risk of mortality and elevated D-dimer levels, implicating a role in hypercoagulability. These findings highlight that host genetic architecture influences viral entry mechanisms mediated by surface protease activity and subsequent inflammatory and thrombotic pathways. A more nuanced understanding of such genetic determinants could guide the development of therapies targeting serine protease pathways [86].
5. Application of Serine Protease Inhibitors or Activators
Understanding serine proteases on the cell surface has revealed new possibilities for treating viral infections. For example, inhibiting TMPRSS2, due to its role in facilitating the entry of several viruses into cells, could be a strategy to reduce infection rates by developing specific inhibitors for TMPRSS2 [22]. Therefore, selective TMPRSS2 inhibitors or the application of a broad-spectrum of protease inhibitors targeting the airway epithelium, where viral replication occurs, are implicated for drug development [87]. TMPRSS2 possesses the capacity to cleave the SARS-CoV-2 S protein at the S2′ site, a critical step that unveils the FP and facilitates membrane fusion at the cellular surface; however, inhibition of TMPRSS2 redirects the virus to alternative entry pathways, such as the cathepsin-mediated endosomal route [34]. Inhibition of TMPRSS2 by camostat or the more effective inhibitor nafamostat [88] was examined in a meta-analysis of randomized clinical trials. Camostat and nafamostat are well tolerated, although efficacy signals were inconsistent across different study populations. Some subgroup analyses favored nafamostat, but no definitive clinical benefit was established [89]. Randomized controlled trials of camostat, covering mild-to-severe COVID-19 cases, confirmed safety but did not demonstrate consistent therapeutic benefits [90], raising questions about the timing of the intervention limiting the effectiveness of camostat [91].
Aprotinin is a broad-spectrum inhibitor of serine proteases, some of which are involved in SARS-CoV-2 S protein activation and in the regulation of host cell-surface proteins. The ATAC Phase III Randomized Controlled Trial evaluated the effectiveness of inhaled aprotinin in patients with COVID-19 pneumonia [92]. Patients who received aprotinin in addition to the standard care experienced significant improvement. Aprotinin recipients had a 5-day reduction in hospital stay, a two-day shorter treatment duration, and were discharged earlier, with a hazard ratio for discharge of 2.19 (95% CI: 1.182–4.047; p = 0.013). In addition, the group showed reduced oxygen requirements and reported no significant adverse events [92]. A1-Antitrypsin (A1AT), an endogenous serpin that inhibits NSPs, is also known to be a direct inhibitor of TMPRSS2, which can prevent SARS-CoV-2 entry in airway epithelial models [93]. NE and CatG degrade the SARS-CoV-2 S protein, deactivate the virus, and reduce lung inflammation; broad protease inhibition by A1AT can negate NSP-mediated antiviral effect [54]. A1AT can reduce tissue damage during infection and also restrict the entry of viruses into cells by inhibiting TMPRSS2. However, inhibiting NSPs, which are part of the body’s antiviral defenses, risks removing an early antiviral barrier crucial in the initial stages of infection. This delicate balance emphasizes the importance of precise timing when considering the therapeutic use of aprotinin, A1AT, or NSP inhibitors to maximize benefits and reduce potential drawbacks.
Strikingly, eosinophil cationic protein (ECP) augments the proteolytic function of NE [94] or, as we demonstrated, lactoferrin (LF) enhances the proteolytic activity of CatG [95]. Furthermore, LF and its derivative peptides, such as lactoferricin, have direct antiviral effects by inhibiting TMPRSS2 activity, blocking S protein priming, and reducing SARS-CoV-2 infection in susceptible cell lines [96]. Moreover, a biophysical study demonstrated that full-length human LF limits viral infection by masking the host receptor, binding to ACE2 instead of the spike receptor-binding domain (RBD). Docking revealed three contact sites on ACE2, indicating multiple ACE2-centered mechanisms of entry inhibition [97]. Alves et al. showed that both apo- and holo-bovine LF exhibit nearly a 70% reduction in viral entry across ancestral and Omicron variants in vitro [98]. Interestingly, bovine LF was also shown to directly bind the RBD of the S protein and inhibit the viral RNA-dependent RNA polymerase (RdRp) in vitro, reducing lung viral loads in hamsters [99]. A clinical trial involving hospitalized patients with moderate-to-severe COVID-19 found LF to be safe and well tolerated; however, the trial did not show any significant improvement in clinical outcomes or inflammatory markers, suggesting that efficacy may depend on the disease stage, timing (patients took LF later in the disease, approximately 6 days after symptom onset), and means of LF delivery [100]. These findings suggest that LF and related peptides are potent natural adjuncts that can reduce SARS-CoV-2 infection and help decrease excessive inflammation but surprisingly did not lead to clinical improvement.
An initial enhancement of the proteolytic activity of NSPs combined with inhibition of TMPRSS2 and furin may offer certain advantages; however, prolonged proteolytic activity poses the potential for adverse effects.
6. Conclusions
Host cell-surface serine proteases have complex, often dual, roles in viral infections. Viruses can hijack these proteases to assist entry and spread, or they can be deployed by the innate immune system to neutralize pathogens. Consequently, this intricate interaction reflects the evolutionary arms race between the host and the virus, with the potential to modulate serine protease activity, initially boosting host defenses and offering insights into innovative antiviral strategies. To counteract immune evasion, components that enhance the catalytic efficiency of NSPs are indicated. For instance, ECP increases the proteolytic activity of NE [94], and LF for CatG [95]. On the other hand, the potential for off-target effects when targeting serine proteases therapeutically requires careful study. Depending on the individual’s medical condition or infection status, exploring the potential of combination therapies that include serine protease inhibitors and/or activators alongside traditional antiviral drugs may improve treatment efficacy. Finally, further research is needed to fully understand the dual roles of serine proteases in immunity and the potential functions that proteases may have in preventing viral infections. Exploring how serine protease activity can be modulated may lead to innovative therapeutic approaches in immunotherapy.
Author Contributions
A.S., M.A., A.M., A.B. and T.B. wrote, edited, and corrected the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Nazarbayev University Faculty-Development Competitive Research Grants Program, reference: 20122022FD4123, awarded to T.B.
Data Availability Statement
No new data were created or analyzed in this study.
Acknowledgments
The School of Sciences and Humanities, Nazarbayev University, paid the publication fee.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| NSPs | Neutrophil serine proteases |
| CatG | Cathepsin G |
| CS | Cleavage site |
| NE | Neutrophil elastase |
| PR3 | Proteinase 3 |
| NSP4 | Neutrophil serine protease 4 |
| CD177 | Cluster of differentiation 177 |
| SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 |
| COVID-19 | Coronavirus disease 2019 |
| S protein | Spike protein |
| TMPRSS2 | Transmembrane protease serine 2 |
| FP | Fusion peptide |
| HA | Hemagglutinin |
| TTSPs | Type II transmembrane serine proteases |
| TMPRSS4 | Transmembrane protease serine 4 |
| TMPRSS11D | Transmembrane protease serine 11D |
| LPAI | Low-pathogenic avian influenza |
| ST14 | Suppressor of tumorigenicity 14 protein (matriptase) |
| HPAI | Highly pathogenic avian influenza |
| HIV-1 | Human immunodeficiency virus type 1 |
| CD4+ | CD4 positive T helper cell |
| RANTES | Regulated on activation, normal T cell expressed and secreted |
| CCL5 | Chemokine (C-C motif) ligand 5 |
| HCV | Hepatitis C virus |
| ACE2 | Angiotensin-converting enzyme 2 |
| CatL | Cathepsin L |
| MOP | Myeloperoxidase |
| MHC I | Major histocompatibility complex class I |
| NET | Neutrophil extracellular trap |
| PAR1 | Protease-activated receptor 1 |
| CD8+ | CD8 positive T cell |
| SerpinA1 | Serine protease inhibitor A1 |
| SerpinA3 | Serine protease inhibitor A3 |
| SNPs | Single-nucleotide polymorphisms |
| A1AT | A1-Antitrypsin |
| ECP | Eosinophil cationic protein |
| LF | Lactoferrin |
| RBD | Receptor-binding domain |
| RdRp | RNA-dependent RNA polymerase |
References
- Seth, S.; Batra, J.; Srinivasan, S. COVID-19: Targeting Proteases in Viral Invasion and Host Immune Response. Front. Mol. Biosci. 2020, 7, 215. [Google Scholar] [CrossRef]
- Pham, C.T. Neutrophil serine proteases fine-tune the inflammatory response. Int. J. Biochem. Cell Biol. 2008, 40, 1317–1333. [Google Scholar] [CrossRef]
- Xia, Q.; Liu, X.; Huang, H. Host proteases: Key regulators in viral infection and therapeutic targeting. Front. Immunol. 2025, 16, 1671173. [Google Scholar] [CrossRef] [PubMed]
- Lubinski, B.; Whittaker, G.R. Host Cell Proteases Involved in Human Respiratory Viral Infections and Their Inhibitors: A Review. Viruses 2024, 16, 984. [Google Scholar] [CrossRef] [PubMed]
- Majchrzak, M.; Poreba, M. The roles of cellular protease interactions in viral infections and programmed cell death: A lesson learned from the SARS-CoV-2 outbreak and COVID-19 pandemic. Pharmacol. Rep. 2022, 74, 1149–1165. [Google Scholar] [CrossRef]
- Kahler, J.P.; Vanhoutte, R.; Verhelst, S.H.L. Activity-Based Protein Profiling of Serine Proteases in Immune Cells. Arch. Immunol. Ther. Exp. 2020, 68, 23. [Google Scholar] [CrossRef]
- Burster, T.; Gartner, F.; Knippschild, U.; Zhanapiya, A. Activity-Based Probes to Utilize the Proteolytic Activity of Cathepsin G in Biological Samples. Front. Chem. 2021, 9, 628295. [Google Scholar] [CrossRef] [PubMed]
- Di Cera, E. Serine proteases. IUBMB Life 2009, 61, 510–515. [Google Scholar] [CrossRef]
- Korba-Mikolajczyk, A.; Sluzalska, K.D.; Kasperkiewicz, P. Exploring the involvement of serine proteases in neutrophil extracellular traps: A review of mechanisms and implications. Cell Death Dis. 2025, 16, 535. [Google Scholar] [CrossRef]
- Cheetham, C.J.; McKelvey, M.C.; McAuley, D.F.; Taggart, C.C. Neutrophil-Derived Proteases in Lung Inflammation: Old Players and New Prospects. Int. J. Mol. Sci. 2024, 25, 5492. [Google Scholar] [CrossRef]
- Burster, T.; Mustafa, Z.; Myrzakhmetova, D.; Zhanapiya, A.; Zimecki, M. Hindrance of the Proteolytic Activity of Neutrophil-Derived Serine Proteases by Serine Protease Inhibitors as a Management of Cardiovascular Diseases and Chronic Inflammation. Front. Chem. 2021, 9, 784003. [Google Scholar] [CrossRef]
- Benarafa, C.; Simon, H.U. Role of granule proteases in the life and death of neutrophils. Biochem. Biophys. Res. Commun. 2017, 482, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Rahbar Saadat, Y.; Hosseiniyan Khatibi, S.M.; Zununi Vahed, S.; Ardalan, M. Host Serine Proteases: A Potential Targeted Therapy for COVID-19 and Influenza. Front. Mol. Biosci. 2021, 8, 725528. [Google Scholar] [CrossRef]
- Campbell, E.J.; Owen, C.A. The sulfate groups of chondroitin sulfate- and heparan sulfate-containing proteoglycans in neutrophil plasma membranes are novel binding sites for human leukocyte elastase and cathepsin G. J. Biol. Chem. 2007, 282, 14645–14654. [Google Scholar] [CrossRef] [PubMed]
- Martin, K.R.; Kantari-Mimoun, C.; Yin, M.; Pederzoli-Ribeil, M.; Angelot-Delettre, F.; Ceroi, A.; Grauffel, C.; Benhamou, M.; Reuter, N.; Saas, P.; et al. Proteinase 3 is a Phosphatidylserine-binding Protein That Affects the Production and Function of Microvesicles. J. Biol. Chem. 2016, 291, 10476–10489. [Google Scholar] [CrossRef] [PubMed]
- Bauer, S.; Abdgawad, M.; Gunnarsson, L.; Segelmark, M.; Tapper, H.; Hellmark, T. Proteinase 3 and CD177 are expressed on the plasma membrane of the same subset of neutrophils. J. Leukoc. Biol. 2007, 81, 458–464. [Google Scholar] [CrossRef]
- von Vietinghoff, S.; Tunnemann, G.; Eulenberg, C.; Wellner, M.; Cristina Cardoso, M.; Luft, F.C.; Kettritz, R. NB1 mediates surface expression of the ANCA antigen proteinase 3 on human neutrophils. Blood 2007, 109, 4487–4493. [Google Scholar] [CrossRef]
- Braun, E.; Sauter, D. Furin-mediated protein processing in infectious diseases and cancer. Clin. Transl. Immunol. 2019, 8, e1073. [Google Scholar] [CrossRef]
- Kido, H.; Okumura, Y.; Takahashi, E.; Pan, H.Y.; Wang, S.; Chida, J.; Le, T.Q.; Yano, M. Host envelope glycoprotein processing proteases are indispensable for entry into human cells by seasonal and highly pathogenic avian influenza viruses. J. Mol. Genet. Med. 2008, 3, 167–175. [Google Scholar] [CrossRef]
- Bestle, D.; Heindl, M.R.; Limburg, H.; Van Lam van, T.; Pilgram, O.; Moulton, H.; Stein, D.A.; Hardes, K.; Eickmann, M.; Dolnik, O.; et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci. Alliance 2020, 3, 14. [Google Scholar] [CrossRef]
- Galloway, S.E.; Reed, M.L.; Russell, C.J.; Steinhauer, D.A. Influenza HA subtypes demonstrate divergent phenotypes for cleavage activation and pH of fusion: Implications for host range and adaptation. PLoS Pathog. 2013, 9, e1003151. [Google Scholar] [CrossRef]
- de Lima, G.B.; Nencioni, E.; Thimoteo, F.; Perea, C.; Pinto, R.F.A.; Sasaki, S.D. TMPRSS2 as a Key Player in Viral Pathogenesis: Influenza and Coronaviruses. Biomolecules 2025, 15, 75. [Google Scholar] [CrossRef]
- Tarnow, C.; Engels, G.; Arendt, A.; Schwalm, F.; Sediri, H.; Preuss, A.; Nelson, P.S.; Garten, W.; Klenk, H.D.; Gabriel, G.; et al. TMPRSS2 is a host factor that is essential for pneumotropism and pathogenicity of H7N9 influenza A virus in mice. J. Virol. 2014, 88, 4744–4751. [Google Scholar] [CrossRef]
- Limburg, H.; Harbig, A.; Bestle, D.; Stein, D.A.; Moulton, H.M.; Jaeger, J.; Janga, H.; Hardes, K.; Koepke, J.; Schulte, L.; et al. TMPRSS2 is the Major Activating Protease of Influenza A Virus in Primary Human Airway Cells and Influenza B Virus in Human Type II Pneumocytes. J. Virol. 2019, 93, e00649-19. [Google Scholar] [CrossRef] [PubMed]
- Tse, L.V.; Hamilton, A.M.; Friling, T.; Whittaker, G.R. A novel activation mechanism of avian influenza virus H9N2 by furin. J. Virol. 2014, 88, 1673–1683. [Google Scholar] [CrossRef]
- Moriuchi, H.; Moriuchi, M.; Fauci, A.S. Cathepsin G, a neutrophil-derived serine protease, increases susceptibility of macrophages to acute human immunodeficiency virus type 1 infection. J. Virol. 2000, 74, 6849–6855. [Google Scholar] [CrossRef]
- Hallenberger, S.; Bosch, V.; Angliker, H.; Shaw, E.; Klenk, H.D.; Garten, W. Inhibition of furin-mediated cleavage activation of HIV-1 glycoprotein gp160. Nature 1992, 360, 358–361. [Google Scholar] [CrossRef]
- Zeisel, M.B.; Fofana, I.; Fafi-Kremer, S.; Baumert, T.F. Hepatitis C virus entry into hepatocytes: Molecular mechanisms and targets for antiviral therapies. J. Hepatol. 2011, 54, 566–576. [Google Scholar] [CrossRef]
- Sharma, N.R.; Mateu, G.; Dreux, M.; Grakoui, A.; Cosset, F.L.; Melikyan, G.B. Hepatitis C virus is primed by CD81 protein for low pH-dependent fusion. J. Biol. Chem. 2011, 286, 30361–30376. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, B.S.; Whittaker, G.R.; Daniel, S. Influenza virus-mediated membrane fusion: Determinants of hemagglutinin fusogenic activity and experimental approaches for assessing virus fusion. Viruses 2012, 4, 1144–1168. [Google Scholar] [CrossRef] [PubMed]
- Esumi, M.; Ishibashi, M.; Yamaguchi, H.; Nakajima, S.; Tai, Y.; Kikuta, S.; Sugitani, M.; Takayama, T.; Tahara, M.; Takeda, M.; et al. Transmembrane serine protease TMPRSS2 activates hepatitis C virus infection. Hepatology 2015, 61, 437–446. [Google Scholar] [CrossRef]
- Lamontagne, J.; Pinkerton, M.; Block, T.M.; Lu, X. Hepatitis B and hepatitis C virus replication upregulates serine protease inhibitor Kazal, resulting in cellular resistance to serine protease-dependent apoptosis. J. Virol. 2010, 84, 907–917. [Google Scholar] [CrossRef]
- Wettstein, L.; Kirchhoff, F.; Munch, J. The Transmembrane Protease TMPRSS2 as a Therapeutic Target for COVID-19 Treatment. Int. J. Mol. Sci. 2022, 23, 1351. [Google Scholar] [CrossRef]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2022, 23, 3–20. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Pohlmann, S. A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 is Essential for Infection of Human Lung Cells. Mol. Cell 2020, 78, 779–784.e5. [Google Scholar] [CrossRef]
- Takeda, M. Proteolytic activation of SARS-CoV-2 spike protein. Microbiol. Immunol. 2022, 66, 15–23. [Google Scholar] [CrossRef]
- Zang, R.; Gomez Castro, M.F.; McCune, B.T.; Zeng, Q.; Rothlauf, P.W.; Sonnek, N.M.; Liu, Z.; Brulois, K.F.; Wang, X.; Greenberg, H.B.; et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 2020, 5, eabc3582. [Google Scholar] [CrossRef]
- Kishimoto, M.; Uemura, K.; Sanaki, T.; Sato, A.; Hall, W.W.; Kariwa, H.; Orba, Y.; Sawa, H.; Sasaki, M. TMPRSS11D and TMPRSS13 Activate the SARS-CoV-2 Spike Protein. Viruses 2021, 13, 384. [Google Scholar] [CrossRef] [PubMed]
- Jaimes, J.A.; Millet, J.K.; Whittaker, G.R. Proteolytic Cleavage of the SARS-CoV-2 Spike Protein and the Role of the Novel S1/S2 Site. iScience 2020, 23, 101212. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Cofas-Vargas, L.F.; Poma, A.B.; Pyrc, K. Evolution of the SARS-CoV-2 spike protein in utilizing host transmembrane serine proteases. iScience 2025, 28, 113318. [Google Scholar] [CrossRef]
- Hoffmann, M.; Hofmann-Winkler, H.; Smith, J.C.; Kruger, N.; Arora, P.; Sorensen, L.K.; Sogaard, O.S.; Hasselstrom, J.B.; Winkler, M.; Hempel, T.; et al. Camostat mesylate inhibits SARS-CoV-2 activation by TMPRSS2-related proteases and its metabolite GBPA exerts antiviral activity. EBioMedicine 2021, 65, 103255. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280 e278. [Google Scholar] [CrossRef]
- Ou, T.; Mou, H.; Zhang, L.; Ojha, A.; Choe, H.; Farzan, M. Hydroxychloroquine-mediated inhibition of SARS-CoV-2 entry is attenuated by TMPRSS2. PLoS Pathog. 2021, 17, e1009212. [Google Scholar] [CrossRef]
- Belouzard, S.; Madu, I.; Whittaker, G.R. Elastase-mediated activation of the severe acute respiratory syndrome coronavirus spike protein at discrete sites within the S2 domain. J. Biol. Chem. 2010, 285, 22758–22763. [Google Scholar] [CrossRef]
- Matsuyama, S.; Ujike, M.; Morikawa, S.; Tashiro, M.; Taguchi, F. Protease-mediated enhancement of severe acute respiratory syndrome coronavirus infection. Proc. Natl. Acad. Sci. USA 2005, 102, 12543–12547. [Google Scholar] [CrossRef]
- Yamamoto, Y.; Inoue, T.; Sugiyama, N.; Furukawa, M.; Sato, K.; Onodera, T.; Takahashi, Y.; Wakita, T.; Fukasawa, M.; Noguchi, K. Synergistic activation of bat SARS-like coronaviruses spike protein by elastase and TMPRSS2. Sci. Rep. 2025, 15, 26469. [Google Scholar] [CrossRef] [PubMed]
- Coutard, B.; Valle, C.; de Lamballerie, X.; Canard, B.; Seidah, N.G.; Decroly, E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 2020, 176, 104742. [Google Scholar] [CrossRef] [PubMed]
- Assylbekova, A.; Allayarova, M.; Konysbekova, M.; Bekturgan, A.; Makhanova, A.; Brown, S.; Grzegorzek, N.; Kalbacher, H.; Kalendar, R.; Burster, T. The Proteolytic Activity of Neutrophil-Derived Serine Proteases Bound to the Cell Surface Arming Lung Epithelial Cells for Viral Defense. Molecules 2024, 29, 4449. [Google Scholar] [CrossRef] [PubMed]
- Simons, T.J. Calcium and neuronal function. Neurosurg. Rev. 1988, 11, 119–129. [Google Scholar] [CrossRef]
- Molloy, S.S.; Thomas, L.; VanSlyke, J.K.; Stenberg, P.E.; Thomas, G. Intracellular trafficking and activation of the furin proprotein convertase: Localization to the TGN and recycling from the cell surface. EMBO J. 1994, 13, 18–33. [Google Scholar] [CrossRef]
- Thomas, G. Furin at the cutting edge: From protein traffic to embryogenesis and disease. Nat. Rev. Mol. Cell Biol. 2002, 3, 753–766. [Google Scholar] [CrossRef]
- Essalmani, R.; Jain, J.; Susan-Resiga, D.; Andreo, U.; Evagelidis, A.; Derbali, R.M.; Huynh, D.N.; Dallaire, F.; Laporte, M.; Delpal, A.; et al. Distinctive Roles of Furin and TMPRSS2 in SARS-CoV-2 Infectivity. J. Virol. 2022, 96, e0012822. [Google Scholar] [CrossRef]
- Korkmaz, B.; Moreau, T.; Gauthier, F. Neutrophil elastase, proteinase 3 and cathepsin G: Physicochemical properties, activity and physiopathological functions. Biochimie 2008, 90, 227–242. [Google Scholar] [CrossRef] [PubMed]
- Leborgne, N.G.; Devisme, C.; Kozarac, N.; Berenguer Veiga, I.; Ebert, N.; Godel, A.; Grau-Roma, L.; Scherer, M.; Plattet, P.; Thiel, V.; et al. Neutrophil proteases are protective against SARS-CoV-2 by degrading the spike protein and dampening virus-mediated inflammation. JCI Insight 2024, 9, e174133. [Google Scholar] [CrossRef]
- Zafarani, A.; Razizadeh, M.H.; Haghi, A. Neutrophil extracellular traps in influenza infection. Heliyon 2023, 9, e23306. [Google Scholar] [CrossRef]
- Papayannopoulos, V.; Metzler, K.D.; Hakkim, A.; Zychlinsky, A. Neutrophil elastase and myeloperoxidase regulate the formation of neutrophil extracellular traps. J. Cell Biol. 2010, 191, 677–691. [Google Scholar] [CrossRef]
- Narasaraju, T.; Yang, E.; Samy, R.P.; Ng, H.H.; Poh, W.P.; Liew, A.A.; Phoon, M.C.; van Rooijen, N.; Chow, V.T. Excessive neutrophils and neutrophil extracellular traps contribute to acute lung injury of influenza pneumonitis. Am. J. Pathol. 2011, 179, 199–210. [Google Scholar] [CrossRef]
- Avril, L.E.; di Martino-Ferrer, M.; Brillard-Bourdet, M.; Gauthier, F. Inhibition of U-937 membrane-associated cathepsin G by GP120 (IIIB) and V3 loop-derived peptides from several strains of HIV-1. FEBS Lett. 1995, 367, 251–256. [Google Scholar] [CrossRef]
- Avril, L.E.; Di Martino-Ferrer, M.; Pignede, G.; Seman, M.; Gauthier, F. Identification of the U-937 membrane-associated proteinase interacting with the V3 loop of HIV-1 gp120 as cathepsin G. FEBS Lett. 1994, 345, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.K.; Lu, W.; Hartley, O.; DeVico, A.L. N-terminal proteolytic processing by cathepsin G converts RANTES/CCL5 and related analogs into a truncated 4-68 variant. J. Leukoc. Biol. 2006, 80, 1395–1404. [Google Scholar] [CrossRef] [PubMed]
- Ren, W.; Hong, W.; Yang, J.; Zou, J.; Chen, L.; Zhou, Y.; Lei, H.; Alu, A.; Que, H.; Gong, Y.; et al. SARS-CoV-2 Delta and Omicron variants resist spike cleavage by human airway trypsin-like protease. J. Clin. Investig. 2024, 134, e174304. [Google Scholar] [CrossRef]
- Zhang, Y.; Bharathi, V.; Dokoshi, T.; de Anda, J.; Ursery, L.T.; Kulkarni, N.N.; Nakamura, Y.; Chen, J.; Luo, E.W.C.; Wang, L.; et al. Viral afterlife: SARS-CoV-2 as a reservoir of immunomimetic peptides that reassemble into proinflammatory supramolecular complexes. Proc. Natl. Acad. Sci. USA 2024, 121, e2300644120. [Google Scholar] [CrossRef]
- Kummarapurugu, A.B.; Zheng, S.; Ma, J.; Ghosh, S.; Hawkridge, A.; Voynow, J.A. Neutrophil Elastase Triggers the Release of Macrophage Extracellular Traps: Relevance to Cystic Fibrosis. Am. J. Respir. Cell Mol. Biol. 2022, 66, 76–85. [Google Scholar] [CrossRef]
- Eruslanov, E.B.; Singhal, S.; Albelda, S.M. Mouse versus Human Neutrophils in Cancer: A Major Knowledge Gap. Trends Cancer 2017, 3, 149–160. [Google Scholar] [CrossRef]
- Chawla, A.; Alatrash, G.; Philips, A.V.; Qiao, N.; Sukhumalchandra, P.; Kerros, C.; Diaconu, I.; Gall, V.; Neal, S.; Peters, H.L.; et al. Neutrophil elastase enhances antigen presentation by upregulating human leukocyte antigen class I expression on tumor cells. Cancer Immunol. Immunother. 2016, 65, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Giese, M.; Turiello, N.; Molenda, N.; Palesch, D.; Meid, A.; Schroeder, R.; Basilico, P.; Benarafa, C.; Halatsch, M.E.; Zimecki, M.; et al. Exogenous cathepsin G upregulates cell surface MHC class I molecules on immune and glioblastoma cells. Oncotarget 2016, 7, 74602–74611. [Google Scholar] [CrossRef] [PubMed]
- Calvert, B.A.; Quiroz, E.J.; Lorenzana, Z.; Doan, N.; Kim, S.; Senger, C.N.; Anders, J.J.; Wallace, W.D.; Salomon, M.P.; Henley, J.; et al. Neutrophilic inflammation promotes SARS-CoV-2 infectivity and augments the inflammatory responses in airway epithelial cells. Front. Immunol. 2023, 14, 1112870. [Google Scholar] [CrossRef] [PubMed]
- Kummarapurugu, A.B.; Hawkridge, A.M.; Ma, J.; Osei, S.; Martin, R.K.; Zheng, S.; Voynow, J.A. Neutrophil elastase decreases SARS-CoV-2 spike protein binding to human bronchial epithelia by clipping ACE-2 ectodomain from the epithelial surface. J. Biol. Chem. 2023, 299, 104820. [Google Scholar] [CrossRef]
- Mustafa, Z.; Zhanapiya, A.; Kalbacher, H.; Burster, T. Neutrophil Elastase and Proteinase 3 Cleavage Sites Are Adjacent to the Polybasic Sequence within the Proteolytic Sensitive Activation Loop of the SARS-CoV-2 Spike Protein. ACS Omega 2021, 6, 7181–7185. [Google Scholar] [CrossRef]
- Mustafa, Z.; Kalbacher, H.; Burster, T. Occurrence of a novel cleavage site for cathepsin G adjacent to the polybasic sequence within the proteolytically sensitive activation loop of the SARS-CoV-2 Omicron variant: The amino acid substitution N679K and P681H of the spike protein. PLoS ONE 2022, 17, e0264723. [Google Scholar] [CrossRef]
- Beloglazov, V.; Yatskov, I.; Nikolaeva, A.; Lavrenchuk, E.; DuBuske, L. Cathepsin G in Patients with SARS-CoV-2 Infection of Various Degrees of Severity. J. Allergy Clin. Immunol. 2022, 149, AB59. [Google Scholar] [CrossRef]
- Violi, F.; Bartimoccia, S.; Cangemi, R.; Nocella, C.; D’Amico, A.; Oliva, A.; Venditti, M.; Pugliese, F.; Mastroianni, C.M.; Ridola, L.; et al. Neutrophil Cathepsin G and Thrombosis in COVID-19. Circ. Res. 2024, 135, 350–352. [Google Scholar] [CrossRef]
- Delgado, M.B.; Clark-Lewis, I.; Loetscher, P.; Langen, H.; Thelen, M.; Baggiolini, M.; Wolf, M. Rapid inactivation of stromal cell-derived factor-1 by cathepsin G associated with lymphocytes. Eur. J. Immunol. 2001, 31, 699–707. [Google Scholar] [CrossRef]
- Yamazaki, T.; Aoki, Y. Cathepsin G binds to human lymphocytes. J. Leukoc. Biol. 1997, 61, 73–79. [Google Scholar] [CrossRef]
- Brunetti, N.S.; Davanzo, G.G.; de Moraes, D.; Ferrari, A.J.R.; Souza, G.F.; Muraro, S.P.; Knittel, T.L.; Boldrini, V.O.; Monteiro, L.B.; Virgilio-da-Silva, J.V.; et al. SARS-CoV-2 uses CD4 to infect T helper lymphocytes. Elife 2023, 12, e84790. [Google Scholar] [CrossRef]
- Bao, R.; Hernandez, K.; Huang, L.; Luke, J.J. ACE2 and TMPRSS2 expression by clinical, HLA, immune, and microbial correlates across 34 human cancers and matched normal tissues: Implications for SARS-CoV-2 COVID-19. J. Immunother. Cancer 2020, 8, e001020. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, A.; Aittomaki, S.; Jankovic, D.; Ortutay, Z.; Pulkkinen, K.; Hamalainen, S.; Rokka, A.; Corthals, G.L.; Watford, W.T.; Junttila, I.; et al. Proprotein convertase FURIN constrains Th2 differentiation and is critical for host resistance against Toxoplasma gondii. J. Immunol. 2014, 193, 5470–5479. [Google Scholar] [CrossRef] [PubMed]
- Castellano, B.; Castellano, C.; Sobczak, A.; Khanna, D. Long-Term Manifestations of COVID-19: A Review. Cureus 2025, 17, e91492. [Google Scholar] [CrossRef]
- Davis, H.E.; McCorkell, L.; Vogel, J.M.; Topol, E.J. Long COVID: Major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 2023, 21, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Hwang, S.; Jang, H.; Park, D.; Kim, J.; Gil, J.; Choi, J.H.; Jang, S.G.; Choi, I.; Jung, Y.; et al. Prolonged immune activation in post-acute sequelae of SARS-CoV-2: Neutrophil dynamics and therapeutic insights. Exp. Mol. Med. 2025, 57, 2067–2082. [Google Scholar] [CrossRef]
- Rabczynski, M.; Chwalek, S.; Adamiec-Mroczek, J.; Lewandowski, L.; Trocha, M.; Nowak, B.; Misiuk-Hojlo, M.; Bednarska-Chabowska, D.; Kuznik, E.; Lubieniecki, P.; et al. Neutrophil Elastase, Neuron-Specific Enolase, and S100B Protein as Potential Markers of Long-Term Complications Caused by COVID-19 in Patients with Type 2 Diabetes Mellitus (T2DM) and Advanced Stage of Diabetic Nephropathy (NfT2DM)-Observational Studies. Int. J. Mol. Sci. 2024, 25, 11791. [Google Scholar] [CrossRef] [PubMed]
- Lotke, R.; Petersen, M.; Sauter, D. Restriction of Viral Glycoprotein Maturation by Cellular Protease Inhibitors. Viruses 2024, 16, 332. [Google Scholar] [CrossRef]
- Janciauskiene, S.; Wrenger, S.; Immenschuh, S.; Olejnicka, B.; Greulich, T.; Welte, T.; Chorostowska-Wynimko, J. The Multifaceted Effects of Alpha1-Antitrypsin on Neutrophil Functions. Front. Pharmacol. 2018, 9, 341. [Google Scholar] [CrossRef]
- Carapito, R.; Li, R.; Helms, J.; Carapito, C.; Gujja, S.; Rolli, V.; Guimaraes, R.; Malagon-Lopez, J.; Spinnhirny, P.; Lederle, A.; et al. Identification of driver genes for critical forms of COVID-19 in a deeply phenotyped young patient cohort. Sci. Transl. Med. 2022, 14, eabj7521. [Google Scholar] [CrossRef]
- Beatty, K.; Bieth, J.; Travis, J. Kinetics of association of serine proteinases with native and oxidized alpha-1-proteinase inhibitor and alpha-1-antichymotrypsin. J. Biol. Chem. 1980, 255, 3931–3934. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Gomez, L.E.; Martinez-Armenta, C.; Tusie-Luna, T.; Vazquez-Cardenas, P.; Vidal-Vazquez, R.P.; Ramirez-Hinojosa, J.P.; Gomez-Martin, D.; Vargas-Alarcon, G.; Posadas-Sanchez, R.; Fragoso, J.M.; et al. The fatal contribution of serine protease-related genetic variants to COVID-19 outcomes. Front. Immunol. 2024, 15, 1335963. [Google Scholar] [CrossRef]
- Millet, J.K.; Whittaker, G.R. Host cell proteases: Critical determinants of coronavirus tropism and pathogenesis. Virus Res. 2015, 202, 120–134. [Google Scholar] [CrossRef]
- Hempel, T.; Raich, L.; Olsson, S.; Azouz, N.P.; Klingler, A.M.; Hoffmann, M.; Pohlmann, S.; Rothenberg, M.E.; Noe, F. Molecular mechanism of inhibiting the SARS-CoV-2 cell entry facilitator TMPRSS2 with camostat and nafamostat. Chem. Sci. 2021, 12, 983–992. [Google Scholar] [CrossRef]
- Hernandez-Mitre, M.P.; Morpeth, S.C.; Venkatesh, B.; Hills, T.E.; Davis, J.; Mahar, R.K.; McPhee, G.; Jones, M.; Totterdell, J.; Tong, S.Y.C.; et al. TMPRSS2 inhibitors for the treatment of COVID-19 in adults: A systematic review and meta-analysis of randomized clinical trials of nafamostat and camostat mesylate. Clin. Microbiol. Infect. 2024, 30, 743–754. [Google Scholar] [CrossRef]
- Khan, U.; Mubariz, M.; Khlidj, Y.; Nasir, M.M.; Ramadan, S.; Saeed, F.; Muhammad, A.; Abuelazm, M. Safety and Efficacy of Camostat Mesylate for COVID-19: A systematic review and Meta-analysis of Randomized controlled trials. BMC Infect. Dis. 2024, 24, 709. [Google Scholar] [CrossRef] [PubMed]
- Kosinsky, Y.; Peskov, K.; Stanski, D.R.; Wetmore, D.; Vinetz, J. Semi-Mechanistic Pharmacokinetic-Pharmacodynamic Model of Camostat Mesylate-Predicted Efficacy against SARS-CoV-2 in COVID-19. Microbiol. Spectr. 2022, 10, e0216721. [Google Scholar] [CrossRef]
- Redondo-Calvo, F.J.; Padin, J.F.; Munoz-Rodriguez, J.R.; Serrano-Oviedo, L.; Lopez-Juarez, P.; Porras Leal, M.L.; Gonzalez Gasca, F.J.; Rodriguez Martinez, M.; Perez Serrano, R.; Sanchez Cadena, A.; et al. Aprotinin treatment against SARS-CoV-2: A randomized phase III study to evaluate the safety and efficacy of a pan-protease inhibitor for moderate COVID-19. Eur. J. Clin. Investig. 2022, 52, e13776. [Google Scholar] [CrossRef]
- Wettstein, L.; Weil, T.; Conzelmann, C.; Muller, J.A.; Gross, R.; Hirschenberger, M.; Seidel, A.; Klute, S.; Zech, F.; Prelli Bozzo, C.; et al. Alpha-1 antitrypsin inhibits TMPRSS2 protease activity and SARS-CoV-2 infection. Nat. Commun. 2021, 12, 1726. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Chakraborty, K.; Tang, X.A.; Zhou, G.; Schoenfelt, K.Q.; Becker, K.M.; Hoffman, A.; Chang, Y.F.; Blank, A.; Reardon, C.A.; et al. Neutrophil elastase selectively kills cancer cells and attenuates tumorigenesis. Cell 2021, 184, 3163–3177 e3121. [Google Scholar] [CrossRef]
- Eipper, S.; Steiner, R.; Lesner, A.; Sienczyk, M.; Palesch, D.; Halatsch, M.E.; Zaczynska, E.; Heim, C.; Hartmann, M.D.; Zimecki, M.; et al. Lactoferrin is an Allosteric Enhancer of the Proteolytic Activity of Cathepsin G. PLoS ONE 2016, 11, e0151509. [Google Scholar] [CrossRef] [PubMed]
- Ohradanova-Repic, A.; Skrabana, R.; Gebetsberger, L.; Tajti, G.; Barath, P.; Ondrovicova, G.; Prazenicova, R.; Jantova, N.; Hrasnova, P.; Stockinger, H.; et al. Blockade of TMPRSS2-mediated priming of SARS-CoV-2 by lactoferricin. Front. Immunol. 2022, 13, 958581. [Google Scholar] [CrossRef]
- Piacentini, R.; Centi, L.; Miotto, M.; Milanetti, E.; Di Rienzo, L.; Pitea, M.; Piazza, P.; Ruocco, G.; Boffi, A.; Parisi, G. Lactoferrin Inhibition of the Complex Formation between ACE2 Receptor and SARS-CoV-2 Recognition Binding Domain. Int. J. Mol. Sci. 2022, 23, 5436. [Google Scholar] [CrossRef] [PubMed]
- Alves, N.S.; Azevedo, A.S.; Dias, B.M.; Horbach, I.S.; Setatino, B.P.; Denani, C.B.; Schwarcz, W.D.; Lima, S.M.B.; Missailidis, S.; Ano Bom, A.P.D.; et al. Inhibition of SARS-CoV-2 Infection in Vero Cells by Bovine Lactoferrin under Different Iron-Saturation States. Pharmaceuticals 2023, 16, 1352. [Google Scholar] [CrossRef]
- He, S.T.; Qin, H.; Guan, L.; Liu, K.; Hong, B.; Zhang, X.; Lou, F.; Li, M.; Lin, W.; Chen, Y.; et al. Bovine lactoferrin inhibits SARS-CoV-2 and SARS-CoV-1 by targeting the RdRp complex and alleviates viral infection in the hamster model. J. Med. Virol. 2023, 95, e28281. [Google Scholar] [CrossRef]
- Matino, E.; Tavella, E.; Rizzi, M.; Avanzi, G.C.; Azzolina, D.; Battaglia, A.; Becco, P.; Bellan, M.; Bertinieri, G.; Bertoletti, M.; et al. Effect of Lactoferrin on Clinical Outcomes of Hospitalized Patients with COVID-19: The LAC Randomized Clinical Trial. Nutrients 2023, 15, 1285. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.