Advances in Livestock Breeding: From DNA Sequencing to Selection Techniques
A special issue of Animals (ISSN 2076-2615). This special issue belongs to the section "Animal Genetics and Genomics".
Deadline for manuscript submissions: 31 August 2026 | Viewed by 961
Special Issue Editors
Interests: proteomics; gene expression; SNP; NGS; mass spectrometry
Interests: population genetics; species identification; SNP; STR
Special Issues, Collections and Topics in MDPI journals
Special Issue Information
Dear Colleagues,
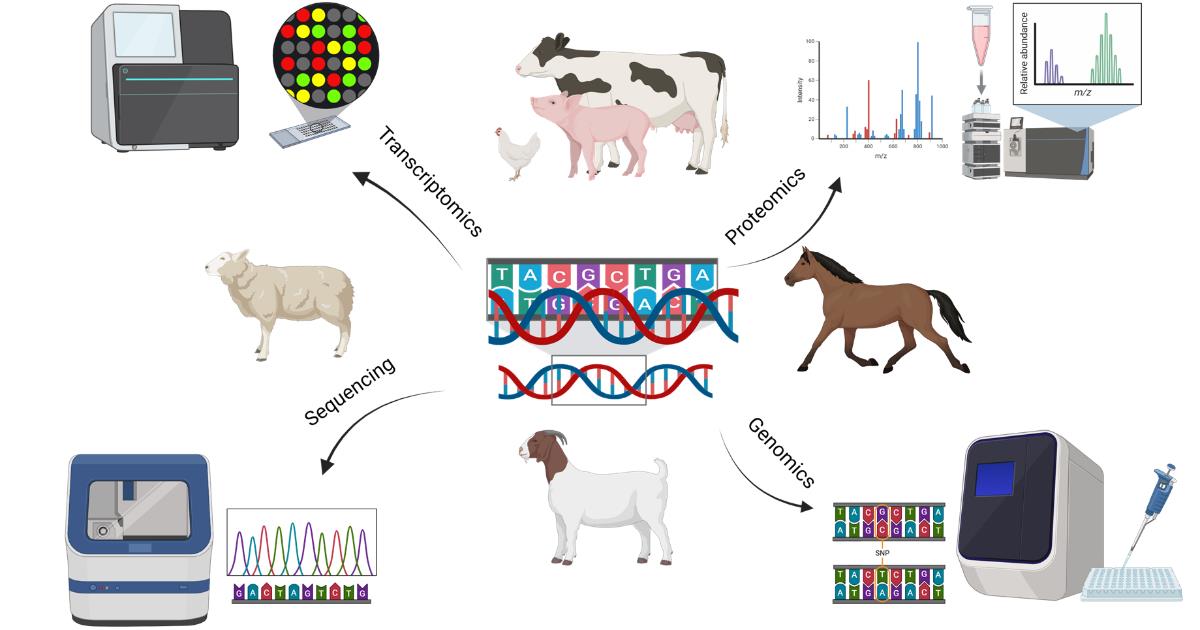
The field of livestock breeding has undergone a transformative evolution with the integration of high-throughput sequencing and advanced selection techniques. While traditional breeding methods have been pivotal in improving desirable traits, the advent of omics technologies—such as genomics, transcriptomics, proteomics, and metabolomics—has deepened our understanding of the complex biological networks that govern animal performance, health, and reproduction. This Special Issue aims to highlight the latest advancements in livestock breeding, from DNA sequencing and SNP discovery to the functional analysis of gene expression and protein dynamics. Particular emphasis is placed on the role of proteomics in bridging the gap between genotype and phenotype, enabling more accurate selection strategies. We invite contributions that explore innovative breeding programs, biomarker discovery, multi-omics data integration, and the application of cutting-edge technologies, such as mass spectrometry, in animal science. Studies focusing on sustainable breeding practices, disease resistance, fertility, and the improvement of production traits in any species (terrestrial, birds, and pollinators) are especially welcome.
Dr. Grzegorz Smołucha
Dr. Anna Koseniuk
Guest Editors
Manuscript Submission Information
Manuscripts should be submitted online at www.mdpi.com by registering and logging in to this website. Once you are registered, click here to go to the submission form. Manuscripts can be submitted until the deadline. All submissions that pass pre-check are peer-reviewed. Accepted papers will be published continuously in the journal (as soon as accepted) and will be listed together on the special issue website. Research articles, review articles as well as short communications are invited. For planned papers, a title and short abstract (about 250 words) can be sent to the Editorial Office for assessment.
Submitted manuscripts should not have been published previously, nor be under consideration for publication elsewhere (except conference proceedings papers). All manuscripts are thoroughly refereed through a single-blind peer-review process. A guide for authors and other relevant information for submission of manuscripts is available on the Instructions for Authors page. Animals is an international peer-reviewed open access semimonthly journal published by MDPI.
Please visit the Instructions for Authors page before submitting a manuscript. The Article Processing Charge (APC) for publication in this open access journal is 2400 CHF (Swiss Francs). Submitted papers should be well formatted and use good English. Authors may use MDPI's English editing service prior to publication or during author revisions.
Keywords
- livestock breeding
- DNA sequencing
- proteomics
- genomics
- mass spectrometry
- biomarkers
- genetic selection
- multi-omics integration
- functional genomic
- sustainable agriculture
Benefits of Publishing in a Special Issue
- Ease of navigation: Grouping papers by topic helps scholars navigate broad scope journals more efficiently.
- Greater discoverability: Special Issues support the reach and impact of scientific research. Articles in Special Issues are more discoverable and cited more frequently.
- Expansion of research network: Special Issues facilitate connections among authors, fostering scientific collaborations.
- External promotion: Articles in Special Issues are often promoted through the journal's social media, increasing their visibility.
- Reprint: MDPI Books provides the opportunity to republish successful Special Issues in book format, both online and in print.
Further information on MDPI's Special Issue policies can be found here.






