Abstract
Non-Saccharomyces yeasts have the potential to ameliorate wine ethanol levels, but such fit-for-purpose yeast strains are still lacking. Seventy-one indigenous non-Saccharomyces yeasts isolated from spontaneous fermentations of four wine regions in China (Ningxia, Xinjiang, Gansu, and Shaanxi) were screened for ethanol formation and were characterized for major metabolite profiles in synthetic grape juice fermentation to obtain non-Saccharomyces yeasts with low ethanol yields. Four Hanseniaspora strains with less volatile acidity production were primarily selected, and their ethanol yield was reduced by 22–32% compared to S. cerevisiae. These strains were further evaluated for oenological properties, namely ethanol and temperature tolerance, H2S production, and killer activities against S. cerevisiae. Strain HuC-3-2 was then subjected to Atmospheric Room Temperature Plasma (ARTP) mutagenesis, and a mutant (HuC32-2-72) with rapid growth and optimized ethanol-reducing capability was obtained. The best-performing strains were further characterized in sequential fermentations with S. cerevisiae in Merlot juice, and resulted in a 1.4% v/v decrease in ethanol yield. Comprehensive analysis of yeast populations and the production of key metabolites highlighted important carbon sinks, as well as glycerol formation, partially accounting for the ethanol reduction. In addition to ethanol amelioration, the Hanseniaspora strains also led to alterations in many metabolites, including volatile compounds and some organic acids, which can further modulate wine aroma and flavor.
1. Introduction
Due to climate warming and advances in viticultural management, wine regions located in warm viticulture areas, including the northwest of China, are commonly related to elevated sugar concentrations in ripe grape berries [1]. Consequently, ethanol levels in commercial wines have been progressively increased [2]. Excessive ethanol content in wines exerts difficulties on both alcoholic and malolactic fermentations by inhibiting the growth of yeast and lactic acid bacteria, thereby resulting in stuck and sluggish fermentation [3]. Inferior sensory qualities of such wines can also be observed due to increased perceptions of hotness, bitterness, astringency, and roughness, as well as the masking of some important aroma compounds by ethanol [4]. Furthermore, many consumers are becoming more health-conscious and mindful of their alcohol intake [5]. According to the International Wine and Spirits Record (IWSR), globally, the growth rate for the wines—no- or low-alcohol (NOLO) category has been well above that of the total wine market. Between 2015 and 2020, the average annual growth rate was 25 percent. The growth rate forecast by the IWSR from 2021 to 2025 is 15% per annum on average, compared with < 1% per annum for total wine volume [6]. Low-alcohol wines have become an important trend in the wine industry, with most countries following the same broad trend of replacing alcohol with low-alcohol alternatives. Thus, there is a strong demand to seek strategies to efficiently reduce ethanol levels in these wines.
To manage the ethanol content in wines, different approaches targeting all stages of winemaking have been proposed. Previous studies have indicated that carbonic maceration processing can be used to obtain a wine fraction with lower alcoholic strength [7]. Pied de cuve (PdC) technology, which refers to a method of indirect inoculation through an inoculum made from must that is already fermenting, has been confirmed to effectively control the production of alcohol and the interaction between a low-ethanol environment and the population dynamics of non-Saccharomyces yeast [8]. The method of reducing the alcohol content in wine by industrially evaporating ethanol and then mixing it has a smaller impact on sensory characteristics [9]. Furthermore, the inhibitor MKS1 regulates the reverse reaction of yeast metabolism to increase glycerol and reduce ethanol [10]. Among these strategies, screening lower-ethanol-producing non-Saccharomyces yeasts has attracted more attention [11]. A few strains of Candida spp. [12], Hanseniaspora spp. [13], Lachancea spp. [14], Metschnikowia spp. [15], Zygosaccharomyces spp. [16], and Starmerella spp. [16], showed 0.3–3.8% v/v less ethanol formation compared to that of Saccharomyces cerevisiae during wine fermentation. Despite their potential, fit-for-purpose non-Saccharomyces strains are still lacking. These oenological characteristics can be diminished under vinification conditions and often require a combination with a more robust starter culture, such as commercial S. cerevisiae, to mitigate the risk of slow and sluggish fermentation [17]. During mixed fermentation, the rapid growth and strong fermentation capability of non-Saccharomyces are required upon fermentation onset. To improve the yeast growth rate, different strain optimization strategies, such as mutagenesis, hybridization, and adaptive laboratory evolution, have been widely used [18,19]. The technology of Atmospheric Room Temperature Plasma (ARTP) is a novel and powerful mutagenesis tool using radio-frequency atmospheric pressure glow discharge plasma jets. Compared to conventional mutagenesis techniques, ARTP offers a more efficient approach to inducing DNA damage and producing stable mutant strains [20]. Because of its operational simplicity, cost-effectiveness, and avoidance of hazardous chemicals, ARTP has been extensively utilized in the mutagenesis of bacteria, fungi, and microalgae, demonstrating significant potential for enhancing productivity and optimizing desirable traits [21,22]. It is, therefore, hypothesized that ARTP can be utilized to enhance the growth rate of lower-ethanol-producing yeasts.
The present work aimed to generate and characterize lower-ethanol-producing yeast strains with rapid growth. Four low-ethanol-producing Hanseniaspora strains were first screened from 71 indigenous non-Saccharomyces strains based on ethanol yield and fermentation traits. One of the strains, HuC-3-2, was then subjected to ARTP mutagenesis to allow the generation of a novel low-ethanol-formation strain that is capable of rapidly initiating alcoholic fermentation. The best-performing strains were further characterized in a sequential culture with S. cerevisiae in Merlot juice, to determine their fermentation performance, with a focus on ethanol levels and the alterations of the primary and secondary metabolites.
2. Materials and Methods
2.1. Yeast Strains and Culture Conditions
The 71 indigenous non-Saccharomyces yeast strains used in this study were isolated from spontaneous fermentations using grapes sourced from four wine-producing regions of China, namely Ningxia, Xinjiang, Gansu, and Shaanxi. These yeast isolates were preserved in 20% glycerol stocks at −80 °C, and are listed in Table 1. The cryogenically preserved strains were revived for 2 days at 28 °C on YPD plates (2% glucose, 2% peptone, 1% yeast extract, 2% agar). Yeast starter cultures were prepared by inoculating approximately 5 × 106 cells/mL into 250 mL Erlenmeyer flasks containing 100 mL YPD liquid medium, and incubated overnight at 28 °C with agitation at 150 rpm.

Table 1.
Yeast strains used in this study.
The commercial yeast S. cerevisiae CECA (Angel Yeast, Yichang, China) was used as the reference strain.
2.2. Screening of Strains with Low Ethanol Production
Lab-scale alcoholic fermentation trials (150 mL) were performed to screen for indigenous yeast strains with low ethanol yields. Fermentations were conducted in triplicate in a chemically defined medium (‘Triple M’ synthetic grape juice, 100 g/L glucose, 100 g/L fructose) at 20 °C [23]. Overnight yeast cultures were prepared in YPD liquid medium at 28 °C prior to being inoculated into Triple M at a rate of 1 × 106 cells/mL. All tested strains were characterized for their ethanol yield and the change in ethanol yield, which were calculated by the following formula.
where A refers to the ethanol yield of the tested non-Saccharomyces strain, whilst B is the ethanol yield of CECA at the same sugar consumption.
Ethanol yield (g/g) = ethanol production (g/L)/sugar consumption (g/L)
Reduction in ethanol (%) = (A − B)/B × 100%
Strains with sugar consumption above 170 g/L and ethanol yield below 0.32 g/g (approximate 20% ethanol reduction compared with CECA) were selected from all tested indigenous non-Saccharomyces strains.
2.3. Evaluation of Enological Performance of the Candidate Strains
Candidate strains with low ethanol yields from the above screening assay were subjected to characterization for their oenological properties. Based on previous research findings, we conducted an ethanol tolerance assay within a 150 mL liquid YPD medium, with ethanol concentrations ranging from 2% to 8% (v/v), incremented by 2% (v/v), the initial yeast cell concentration at 1 × 106 cells/mL, and shaking at 150 rpm. Yeast culture density was monitored 48 h after inoculation using a Cary 60 UV-Vis spectrophotometer (Agilent Technologies, Santa, Clara, CA, USA) at 600 nm. Similarly, yeast growth was characterized in normal YPD at four different temperatures (10, 20, 30, 40 °C). H2S production was evaluated using the BiGGY agar approach according to a previous study [24]. Killer toxin activity of the indigenous strains against the sensitive S. cerevisiae strain 1296, at 28 °C, was assessed using the agar diffusion well method, as previously described [25].
2.4. ARTP Mutagenesis to Generate Novel Yeast Strains with Lower Ethanol Yield
Atmospheric and Room Temperature Plasma (ARTP) mutagenesis was performed using the ARTP biological mutagenesis system (Wuxi Yuanqing Tianmu Biological Technology Co., Ltd., Wuxi, China). Mid-log phase non-Saccharomyces yeast cells were collected, washed twice with 5% PBS, and diluted to a suspension with an OD600 (the optical density value measured at a wavelength of 600 nm) of 0.6–0.8. A dose of 10 μL yeast culture was then evenly spread on a sterile metal plate, and was exposed to 100 W power of radio frequency for 0 -150 s. The culture was sampled at various time points during the ARTP treatment to determine the lethality rate, which was calculated as the ratio of the total number of dead spores after the ARTP treatment to that of the live spores before the treatment. Simultaneously, samples were collected and eluted with 1 mL sterile PBS (5%). Cells were then harvested after centrifugation (5000 rpm, 10 min), and the medium was replaced with YPD and sterile glycerol (added to 20% v/v) for cryo-storage.
2.5. Screening for the ARTP Mutagenesis Isolates with Lower Ethanol Yield via Micro- and Lab-Scale Fermentations
Glycerol stocks of the mixed culture collected at the mutagenic lethality of 80–90% were streaked onto YPD agar medium for isolation of single colonies. Preliminary characterization of these single colonies was performed in micro-scale fermentations in 24-well micro-titer plates. Each well containing 1.8 mL YPD was inoculated with approximately 1 × 106 cells /mL yeast isolates. Plates sealed with breathable titer tops (Coad: SF-200) were incubated at 20 °C for 72 h, after which the cell density was measured using a microplate spectra-photometer (BioTek, ELx800, Winooski, VT, USA) at 600 nm. Mutants with higher biomass than the parent strain were selected for further evaluation by lab-scale sequential fermentations.
The filter-sterilized Merlot juice (2021, Yuma winery, Ningxia, China) with 254 g/L sugar was used for the lab-scale fermentation experiment. Sequential fermentations were performed in triplicate in 500 mL Erlenmeyer flasks equipped with air locks and containing 300 mL of grape juice at 25 °C. The Merlot juice was first inoculated with overnight cultures of the selected non-Saccharomyces at 1 × 107 cells/mL, followed by inoculation of the commercial wine yeast CECA at 1 × 106 cells/mL after 48 h. Fermentation samples were collected every 48 h to determine yeast viability and sugar consumption. The final sample was collected at the end of fermentation for downstream analyses of metabolites and volatile compounds.
2.6. Chemical Analysis
Residual sugar and volatile acidity were measured according to a previously described process [26]. Glycerol, ethanol, and organic acids were quantified using HPLC equipment (Agilent 1100, Agilent Technologies, Santa, Clara, CA, USA) equipped with an HPX-87H Aminex ionexchange column (300 mm × 7.8 mm, 9 µm particle size, 8% cross-linkage; Bio-Rad, Hercules, CA, USA). Specifically, citric, tartaric, pyruvic, malic, succinic, fumaric, and acetic acids were assayed by a diode array detector (Agilent DAD G7115A, Agilent Technologies, Santa, Clara, CA, USA) at 210 nm with the column temperature maintained at 60 °C whilst an extra cation H+ micro-guard cartridge (BioRad, Hercules, CA, USA) was included for the determination of glycerol and ethanol at 40°C. Prior to the HPLC analysis, samples were diluted 1 in 5 using ultra-pure water, and filtered through 0.22 μM nylon syringe filters. A volume of 20 μL of diluted samples was finally injected into the HPLC equipment. Analytes were eluted with 5 mM H2SO4 at a flow rate of 0.6 mL/min. Quantitative analusis of the analytes was performed using the standard curves.
Volatile compounds were analyzed using head space–solid phase microextraction–gas chromatography with mass spectrometry (HS-SPME-GC-MS) following Chen et al. [26]. In brief, an Agilent 6890 GC coupled with an Agilent 5975B MSD (Agilent Technologies, Santa, Clara, CA, USA) and an HP-INNOWAX (J&W Scientific, Folsom, CA, USA; 60 m × 0.25 mm, 0.25 μm film thickness) column were used for volatile profiling. The carrier gas was helium at 1 mL/min; the solid phase microextraction was manually injected by placing the SPME fiber at the GC inlet for 25 min in a non-split mode. The temperature of the GC system was adjusted as follows: 40 °C for 5 min, then increased to 200 °C at a rate of 3 °C/min and held for 2 min. The ion source temperature was kept at 230 °C. Spectra were acquired on electron ionization (EI) at 70 eV, and the mass scan range was 29–350 m/z, with a scan interval of 0.2 s.
Standard calibration curves were obtained using volatile compound standards in a synthetic wine medium (14% v/v ethanol, 5 g/L tartaric acid, pH 3.8). The standard mixture was blended with a 10 µL internal standard (4-methyl-2-pentanol, 20 mg/L), and analyzed according to the HS-SPME-GC-MS protocol described above. Agilent ChemStation was used to qualify and quantify the volatile compounds. The concentrations of the compounds were calculated using calibration curves following [26].
2.7. Statistical Analysis
Data were preliminarily processed using SPSS 26.0 (SPSS Inc., Chicago, IL, USA), and are expressed as mean values ± standard deviations. The same software was used to perform One-way ANOVA to allow the determination of significant differences among the tested strains. A confidence interval for One-way ANOVA was set at 95%. Volatile compounds with means that are significantly different (p < 0.05) across the treatments were further analyzed by principal component analysis (PCA) using Origin 2024 (Originlab Inc., Northampton, MA, USA). The rest of the figures were plotted using GraphPad Prism 9 (GraphPad Software, San Diego, CA, USA).
3. Results
3.1. Evaluation of Non-Saccharomyces Yeasts with Low Ethanol Yield
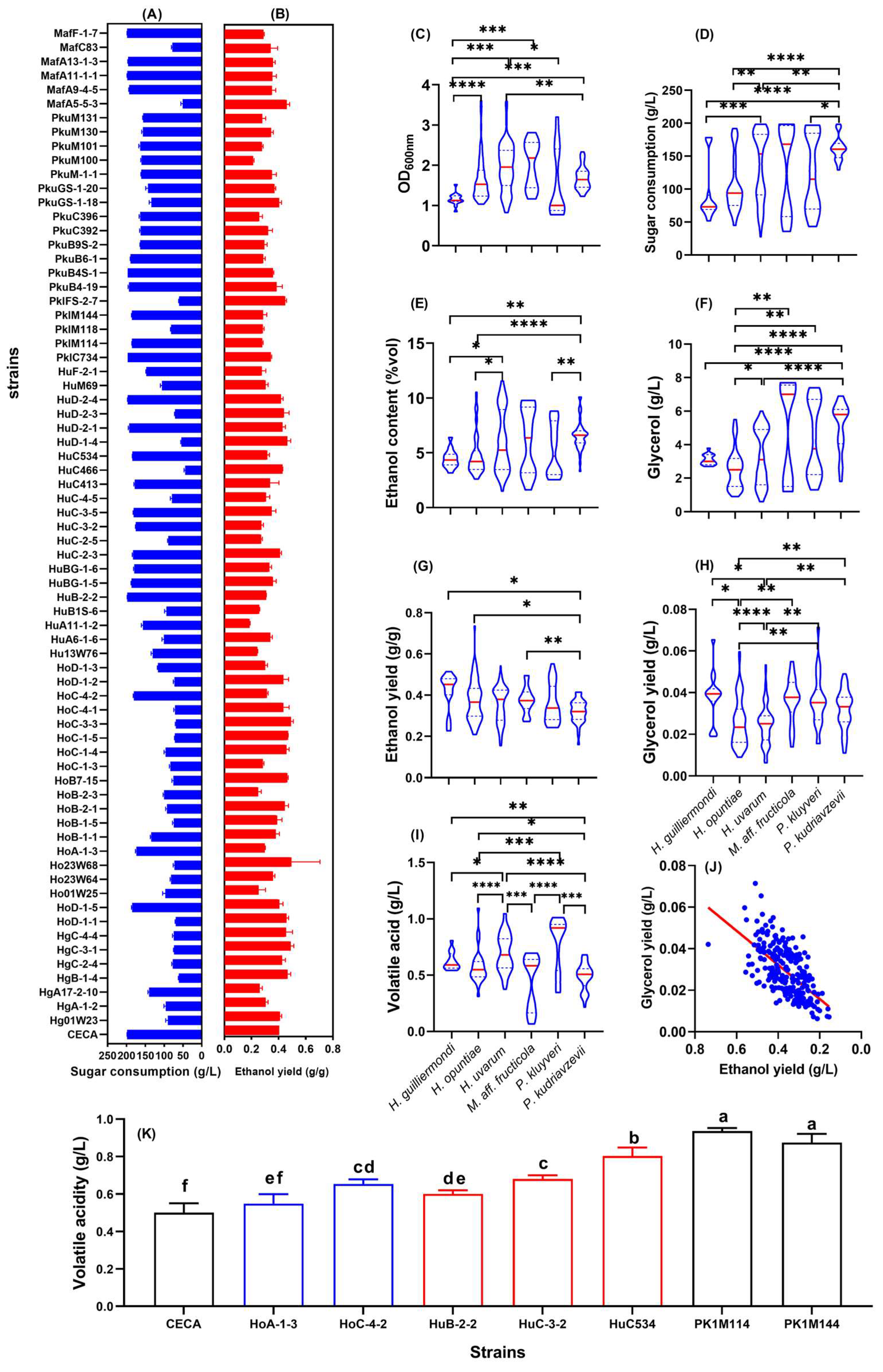
The ethanol yields of 71 indigenous non-Saccharomyces strains and S. cerevisiae CECA were assessed by fermentation in Triple M with an initial sugar concentration of 200 g/L. The physicochemical parameters of all ferments were determined, mainly to evaluate the ability of the tested strains to induce and conduct alcoholic fermentation (Figure 1). The reference strain, CECA, consumed 198.18 g/L of sugar to produce 10.16% v/v ethanol, with an ethanol yield of 0.404 g/g. By contrast, the amount of sugar consumed by the tested non-Saccharomyces isolates ranged from 43.93 to 198.15g/L (Figure 1A), and their ethanol yield ranged from 0.290 to 0.494 g/g (Figure 1B). In order to select lower-ethanol-forming non-Saccharomyces yeasts with good fermentation performance, the ethanol yield of all tested strains was compared to that of CECA at the same sugar consumption level. Here, we proposed a basic selection criterion for such low-ethanol-yield non-Saccharomyces yeasts, which includes the capability to utilize a minimum amount of 170 g/L sugar whilst having an ethanol yield of less than 0.32 g/g. Accordingly, strains HoA-1-3, HoC-4-2, HuB-2-2, HuC-3-2, HuC534, PklM114, and PklM144 were identified as candidates with the potential for ethanol reduction. Compared to CECA, the mentioned strains displayed decreased ethanol yields by 26.03%, 22.33%, 23.77%, 32.05%, 21.49%, 30.64%, and 29.24%, respectively (Figure 1A). The reduction in ethanol yields indicated a higher conversion of sugar from ethanol formation through alternate metabolic pathways. In the present study, glycerol and volatile acidity were determined after the termination of alcoholic fermentation. These parameters were further clustered at the species level, along with ethanol yield, culture density, and sugar consumption, to reveal a general picture of interspecific differences in sugar metabolism (Figure 1C–J).
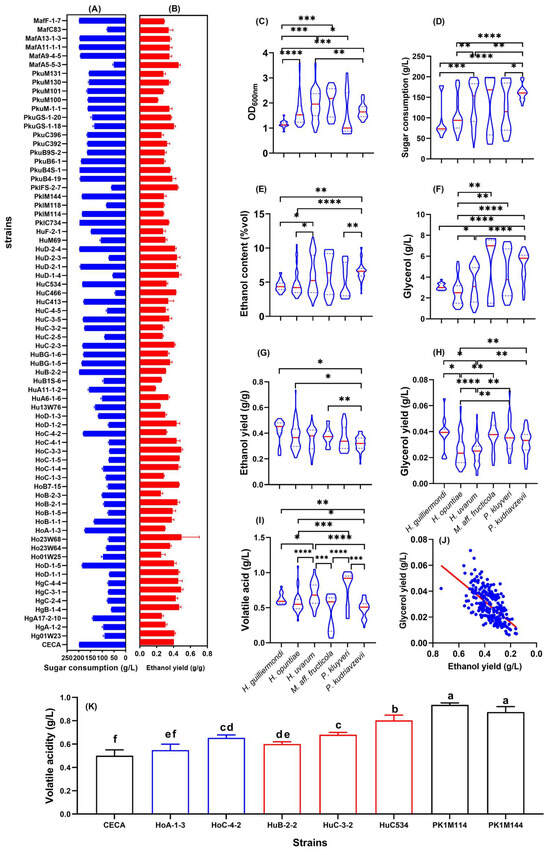
Figure 1.
Physiological parameters of fermented Triple M medium by non-Saccharomyces yeasts. (A) Sugar consumption of each tested yeast strain; (B) ethanol yield of each tested yeast strain. Graphs (C–G) are violin plots demonstrating interspecific differences of the fermentation-related parameters. (C) Culture density at 600 nm; (D) sugar consumption; (E) ethanol content; (F) glycerol content (G) ethanol yield; (H) glycerol yield; (I) volatile acidity; (J) total acidity; (K) volatile acidity of 7 candidate non-Saccharomyces strains with low ethanol yields. * p < 0.05; ** p < 0.01, *** p < 0.001; **** p < 0.0001; Different letters indicate differences among wine samples determined by the Duncan test at a 95% confidence level.
Strains belonging to H. uvarum, P. kluyveri, and M. aff. fructicola displayed better growth and more sugar consumption in Triple M compared to H. guilliermondi, H. opuntiae, and P. kudriavzevii strains (Figure 1C,D). With regard to major metabolites during alcoholic fermentation, significant differences were observed in ethanol and glycerol production among the six yeast species (Figure 1E–H). Particularly for M. aff. fructicola strains, the production of ethanol and glycerol fluctuated the most, which were within the range of 2.97–8.88% v/v and 1.37–7.60 g/L, respectively. Interestingly, the ethanol yields of the 71 non-Saccharomyces strains were negatively associated with glycerol production (Figure 1J). Volatile acidity (expressed as acetic acid) of the ferments by the end of fermentation was within the range of 0.37–0.85 g/L, among which H. uvarum and P. kluyveri resulted in a higher volatile acidity level compared to other species (Figure 1I). Notedly, the seven candidate strains with lower ethanol yields produced significantly higher amounts of volatile acidity compared to CECA, but all were nonetheless within the legal limit (1.2 g/L) (Figure 1K). Due to their lower volatile acidity yields, strains HoA-1-3, HoC-4-2, HuB-2-2, and HuC-3-2 were further shortlisted for subsequent characterization.
3.2. Oenological Properties of Low-Ethanol-Yielding Hanseniaspora Yeasts
All strains were able to tolerate an initial glucose concentration of 400 g/L (Table 2). However, a decrease in growth for Hanseniaspora strains was shown when the initial sugar concentration exceeded 300 g/L. The impacts of ethanol and temperature on the survival of the four selected Hanseniaspora strains were evaluated (Table 2). In general, increased ethanol concentration resulted in reduced yeast viability. Strains HoA-1-3 and HoC-4-2 displayed significantly reduced viability in medium with 6% v/v or more ethanol, whilst the growth density of HuB-2-2 and HuC-3-2 decreased remarkably only with 4% v/v or more ethanol (Table 2). In terms of temperature tolerance assays, growth was observed for all strains between 10 °C and 40 °C, among which the growth density was the highest for strains HoC-4-2 and HuB-2-2 at each tested temperature. Strains HoA-1-3 and HuC-3-2 showed considerable growth at typical wine fermentation temperatures (20 °C and 30 °C), whilst less population was seen at the two extreme temperatures (10 °C and 40 °C, Table 2).

Table 2.
Oenological properties of the four Hanseniaspora yeasts with low ethanol yields.
Hydrogen sulfide production by the four strains was evaluated using the plate-based method by comparing the colors of the colonies grown on the BiGGY agar. HuB-2-2 was the only strain that is capable of producing H2S, which was indicated by the darker color of all triplicate colonies (Table 2). Further, no killer activity was observed for any of the four Hanseniaspora strains against the sensitive-type strain S. cerevisiae 1296 (Table 2).
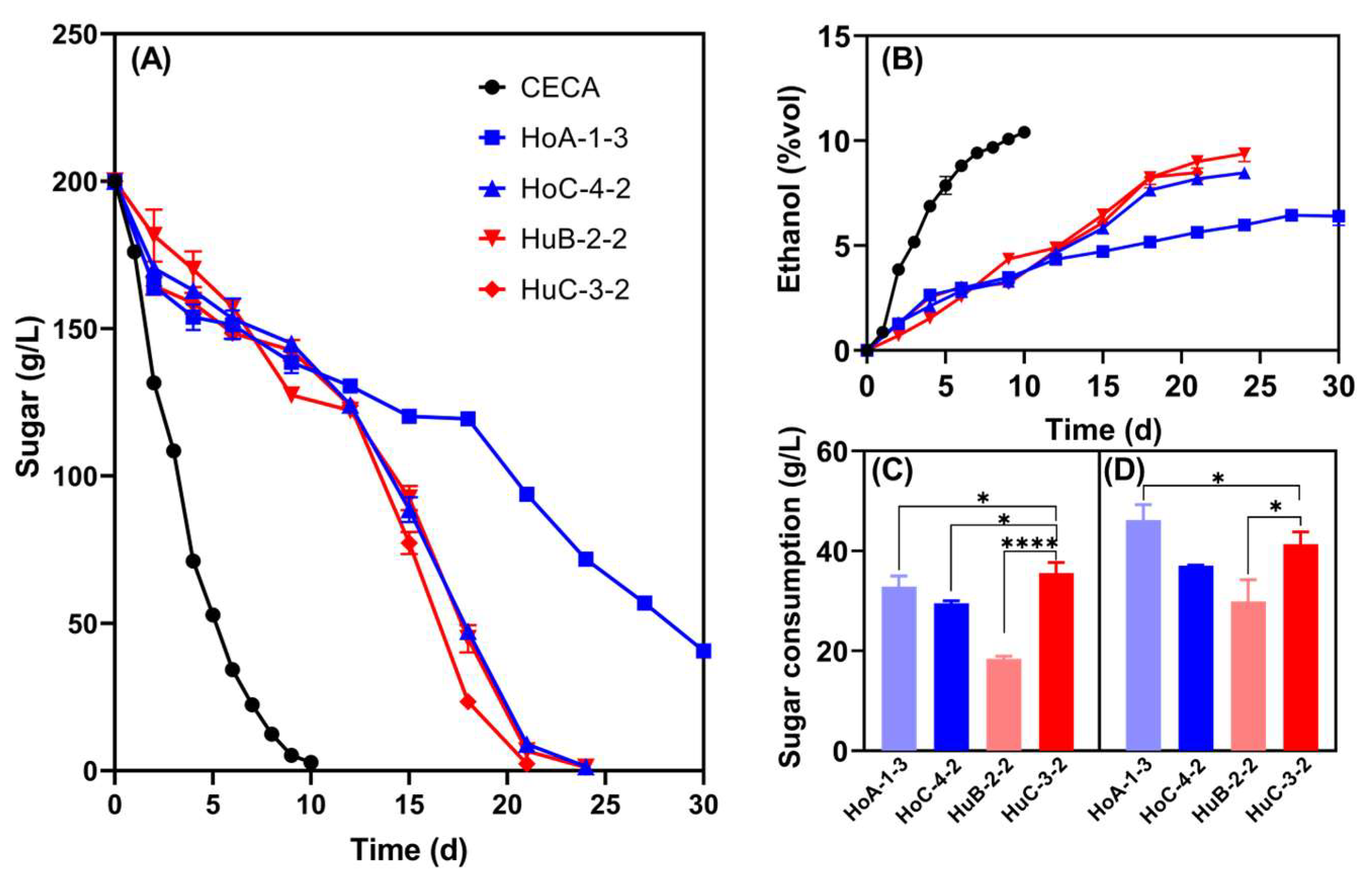
The sugar consumption and low-ethanol-production capacity of the four strains were double-checked in Triple M fermentations to ensure the stability and reliability of such traits (Figure 2). Undoubtedly, CECA showed the strongest ethanol production capacity, with 10.41% v/v ethanol being produced, which was significantly higher than that of the four Hanseniaspora strains. At the end of fermentation, the ethanol contents of HoA-1-3, HoC-4-2, HuB-2-2, and HuC-3-2 were 6.41% v/v, 8.47% v/v, 9.38% v/v, and 8.48% v/v, respectively (Figure 2B). Notedly, HoA-1-3 was unable to complete fermentation, whilst sugar was thoroughly depleted for the other three strains, but the fermentation duration doubled compared to the CECA monoculture fermentation (Figure 2A). Normally, S. cerevisiae is added into the ferments several days after the inoculation of non-Saccharomyces to ensure timely wine fermentations. Thus, the sugar consumption of the selected strains was further compared on the 2nd and the 4th day of fermentation (Figure 2C). HuC-3-2 consumed higher amounts of sugar compared to HoC-4-2 on the 2nd day, but no significant differences were observed on the 4th day (Figure 2C). Based on the above results, strain HuC-3-2 was selected as the parent strain for further reducing ethanol yield via ARTP mutagenesis.
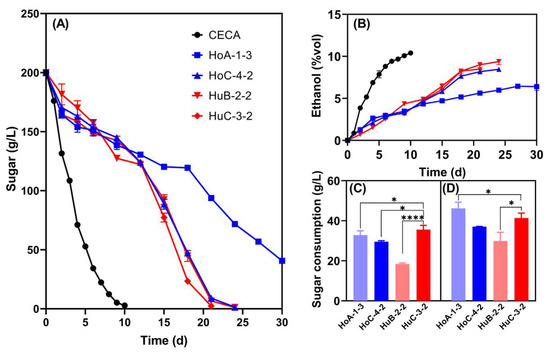
Figure 2.
Fermentation performance of the four Hanseniaspora strains in Triple M. (A) Sugar consumption kinetics; (B) ethanol yield kinetics; (C) the amount of sugar consumed on the second day of the fermentation; (D) the amount of sugar consumed on the fourth day of the fermentation. * p < 0.05; **** p < 0.0001.
3.3. ARTP Mutagenesis and Rapid Screening of Yeasts with Lower Ethanol Yield
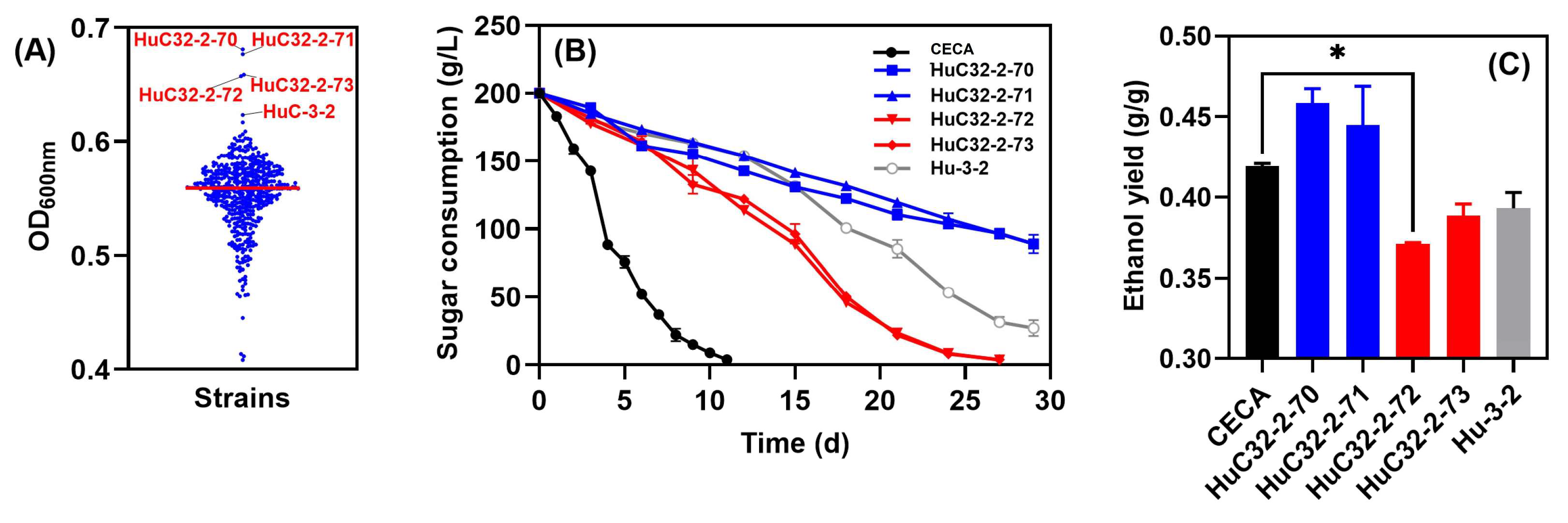
With a view to improving growth whilst further reducing the ethanol yield of the indigenous non-Saccharomyces yeast, the strain HuC-3-2 was subjected to ARTP mutagenesis. Cell counts were conducted to determine yeast viability at intervals of 45 s, 55 s, and 65 s, in which the lethality rates of the treated cells reached 80~90%. A total number of 488 ARTP-mutagenized strains were obtained, and individual isolates were evaluated for their relative performance in micro-scale screening experiments. The basis for the preliminary selection of candidate strains was an improved biomass in comparison to the parent strain, HuC-3-2. Consequently, four strains, namely, HuC32-2-70 (OD600 = 0.681), HuC32-2-71 (OD600 = 0.677), HuC32-2-72 (OD600 = 0.659), and HuC32-2-73 (OD600 = 0.657), outcompeted HuC-3-2 (OD600 = 0.623) (Figure 3A), and were further evaluated in lab-scale fermentations in Triple M (Figure 3B). For CECA fermentation, sugar utilization was rapid, finishing in 12 days, whilst strains HuC32-2-72 and HuC32-2-73 consumed all sugar in 27 days. By contrast, fermentation was stalled for strains HuC32-2-70 and HuC32-2-71 (Figure 3B). Ethanol was analyzed by the end of fermentation, and the two mutants that competed for fermentation reduced ethanol yields markedly compared to those of CECA (Figure 3C).
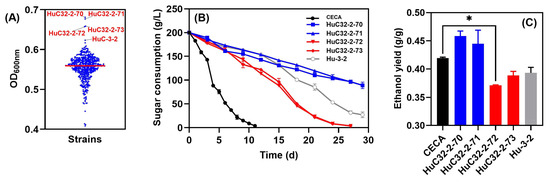
Figure 3.
Evaluation of the 488 single isolates derived from ARTP mutagenesis. (A) Culture density at 600 nm after 72 h of growth in YPD; (B) sugar consumption kinetics of the four isolates with the greatest growth; (C) ethanol yield of the four candidate isolates. * p < 0.05.
3.4. Sequential Merlot Juice Fermentations Using Low-Ethanol-Yielding Hanseniaspora Yeasts and S. cerevisiae
3.4.1. Dynamic Changes of Sugar Consumption and Yeast Population During Fermentation
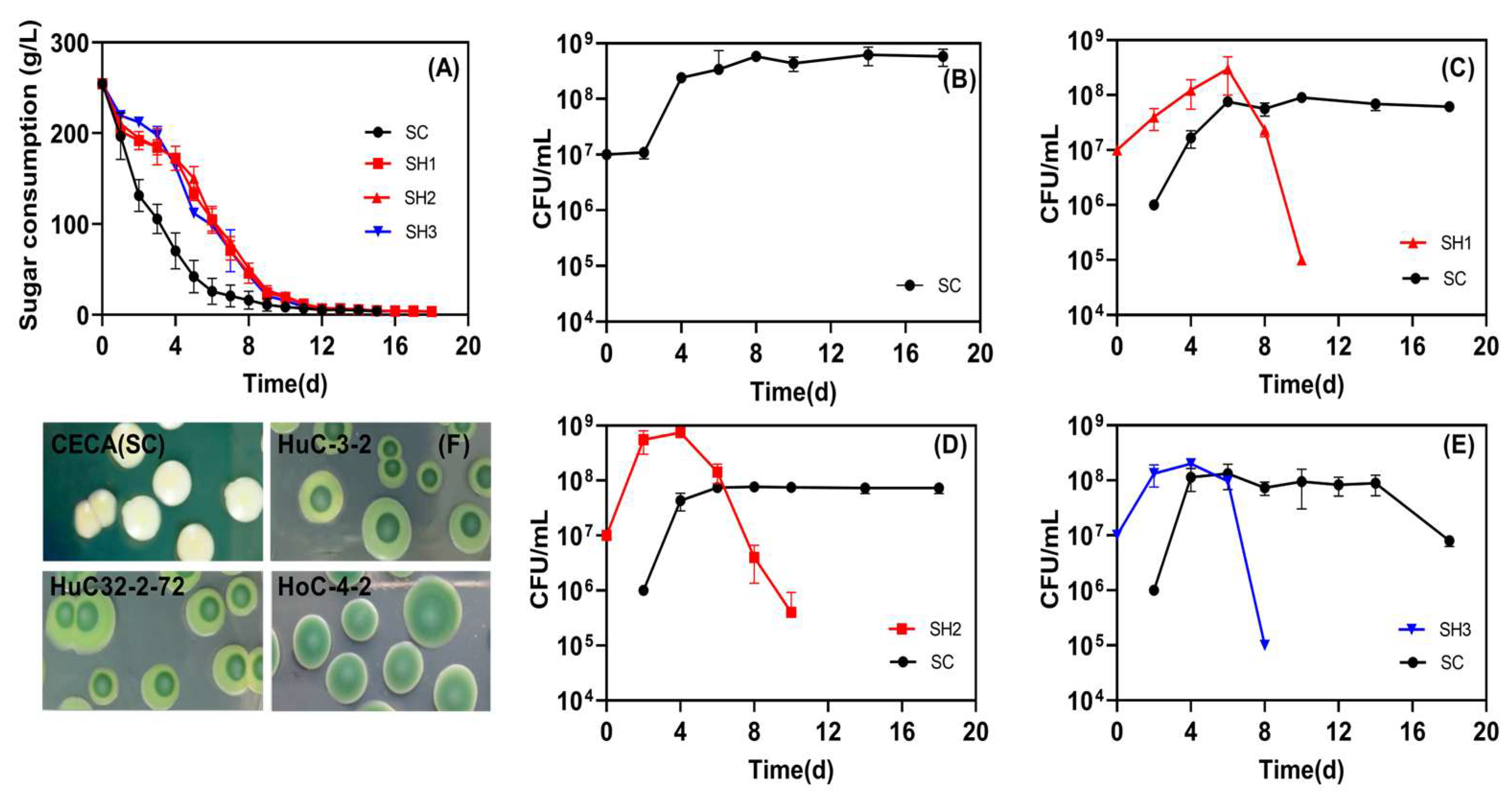
Three low-ethanol-producing isolates selected from this study, including one ARTP mutant, were evaluated for their potential application in winemaking in sequential fermentation with CECA (SH1, SH2, and SH3, respectively). The fermentation process was successfully completed for all respective treatments. Among them, the CECA monoculture fermentation exhibited the most rapid progression, whereas the sugar depletion in the mixed fermentations was observed to take place over a span of 18 days (Figure 4A), and the yeast population reached the stationary phase on Day 4, and retained over 108 CFU/mL till fermentation terminated (Figure 4B). All tested non-Saccharomyces dominated at the early stage of mixed fermentations (4~6 d), followed by a sharp drop to less than 106 CFU/mL on Day 10. Among them, the ARTP-mutagenized strain HuC32-2-72 had an OD value close to 109 CFU/mL on Day 4, which was higher than HuC-3-2 and HoC-4-2 (Figure 4C–E). In all sequential fermentations, the population of CECA increased to approximately 108 CFU/mL 2–4 days after inoculation, and the population was retained at this level afterward (Figure 4C–E).
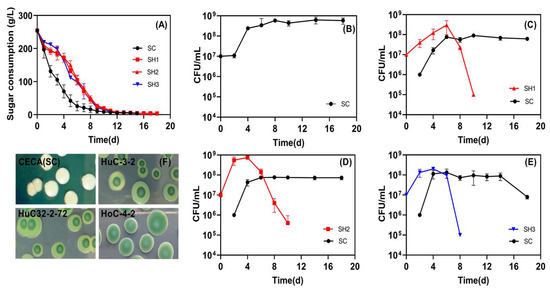
Figure 4.
Sequential inoculation fermentation in Merlot grape juice. (A) Sugar consumption kinetics; (B) viability of CECA (SC) in pure culture fermentation; (C) yeast viability in SH1 fermentations (SH1: sequential inoculation fermentation of HuC-3-2 and CECA); (D) yeast viability in SH2 fermentations (SH2: sequential inoculation fermentation of HuC32-2-72 and CECA); (E) yeast viability in SH3 fermentations (SH3: sequential inoculation fermentation of HoC-4-2 and CECA); (F) yeast counts as determined on WLN agar plates during fermentation.
3.4.2. Basic Wine Parameters
The basic chemical components of pure and mixed fermented wines are listed in Table 3, many of which were impacted by the fermentation modalities. The ethanol concentrations in the sequential wines were approximately 1.4% v/v lower than the CECA wines, resulting in a reduced ethanol yield change rate of 10.42%. The sequential wines had higher glycerol concentrations than the control, most noticeably in SH2, which exhibited a significant increase of 46.1% in comparison to CECA. Furthermore, total acid concentrations, as well as malic and succinic acids, were significantly lower in the sequential wines compared to the control. A notably low pyruvic acid concentration was identified in SH2, recording decrements of 37.4%, 25.3%, and 32.7% relative to CECA, SH1, and SH3, respectively. However, the quantities of lactic and fumaric acids displayed negligible fluctuations across all wines. Compared to CECA monoculture wines, a higher amount of acetic acid was found in all sequential wines, which were nonetheless within the limit of the international standard (1.2 g/L, OIV, 2018).

Table 3.
Concentrations of non-volatiles in Merlot wines produced with mixed Hanseniaspora/CECA and pure CECA fermentations.
3.4.3. Volatile Compounds of the Wines
A total number of 40 volatile compounds were identified and quantified in Merlot wines, of which 37 showed significant differences between the sequential and CECA wines (ANVOA; p < 0.05, Table 4).

Table 4.
Volatile compounds of the Merlot wines.
The impact of alcohol, which is a secondary metabolite generated by yeast during the process of alcoholic fermentation, on the aroma of wine can be either beneficial or detrimental [27]. Excessive contents of higher alcohols can negatively affect wine flavor [28]. Sequential inoculation caused a decrease in total higher alcohols, mainly driven by isobutanol, which was 1.4~2-fold less present than in CECA wines. The production of 2-methyl-1-propanol, 3-methyl-1-butanol, and phenylethyl alcohol during fermentation occurs via the conversion of valine, leucine, and phenylalanine via the Ehrlich pathway during fermentation [29]. Compared with CECA wines, 3-methyl-1-butanol and 2,3-butanediol were also significantly lower in SH1 and SH2 wines, but the amounts were comparable in SH3 wines. Phenethyl alcohol is a significant compound in wine, possessing a rose-honey flavor. It is produced by yeasts through the anaerobic conversion of phenylalanine [30]. In this study, phenylethyl alcohol was the only identified higher alcohol found to be higher in sequential wines than in the CECA wines, with the highest amount being seen in SH3 wines (Table 4). Farnesol and acetoin were remarkably increased in sequential wines, whilst nerol was found to be higher in the control wines than any of the sequential wines (Table 4).
Esters are formed mainly during alcoholic fermentation and are an important component of a wine’s aroma [31]. Generally, esters positively contribute to wine aroma and are considered major contributors to sweet and fruity odors [32]. In contrast to higher alcohols, in sequential wines, the acetate esters were generally higher than in the CECA wines, of which ethyl acetate was the most abundant, and it was 2~4-fold higher than in the control wines (Table 4). Isoamyl phenethyl acetates also surpassed their olfactory thresholds, and were 1.5~2-fold and 2.3~3.4-fold higher in the sequential wines, respectively (Table 4). Conversely, sequential inoculation resulted in a decrease in concentrations of most ethyl esters, particularly ethyl lactate and ethyl decanoate (Table 4). Non-Saccharomyces yeasts influence the production of ethyl esters in fruit wine, potentially due to the presence of secretases typically absent in S. cerevisiae, such as α-L-arabinofuranosidase, β-glucosidase, polygalacturonase, cellulase, and protease [33,34]. In sequential wines, the identified fatty acids were generally lower than in the CECA wines, except for octanoic and decanoic acids, which were either comparable or higher than the control (Table 4). A previous study reported that fatty acids have little to do with wine quality but play an essential role in the complexity of wine aromas [27,35]. The formation of terpenes was also affected by the fermentation matrix.
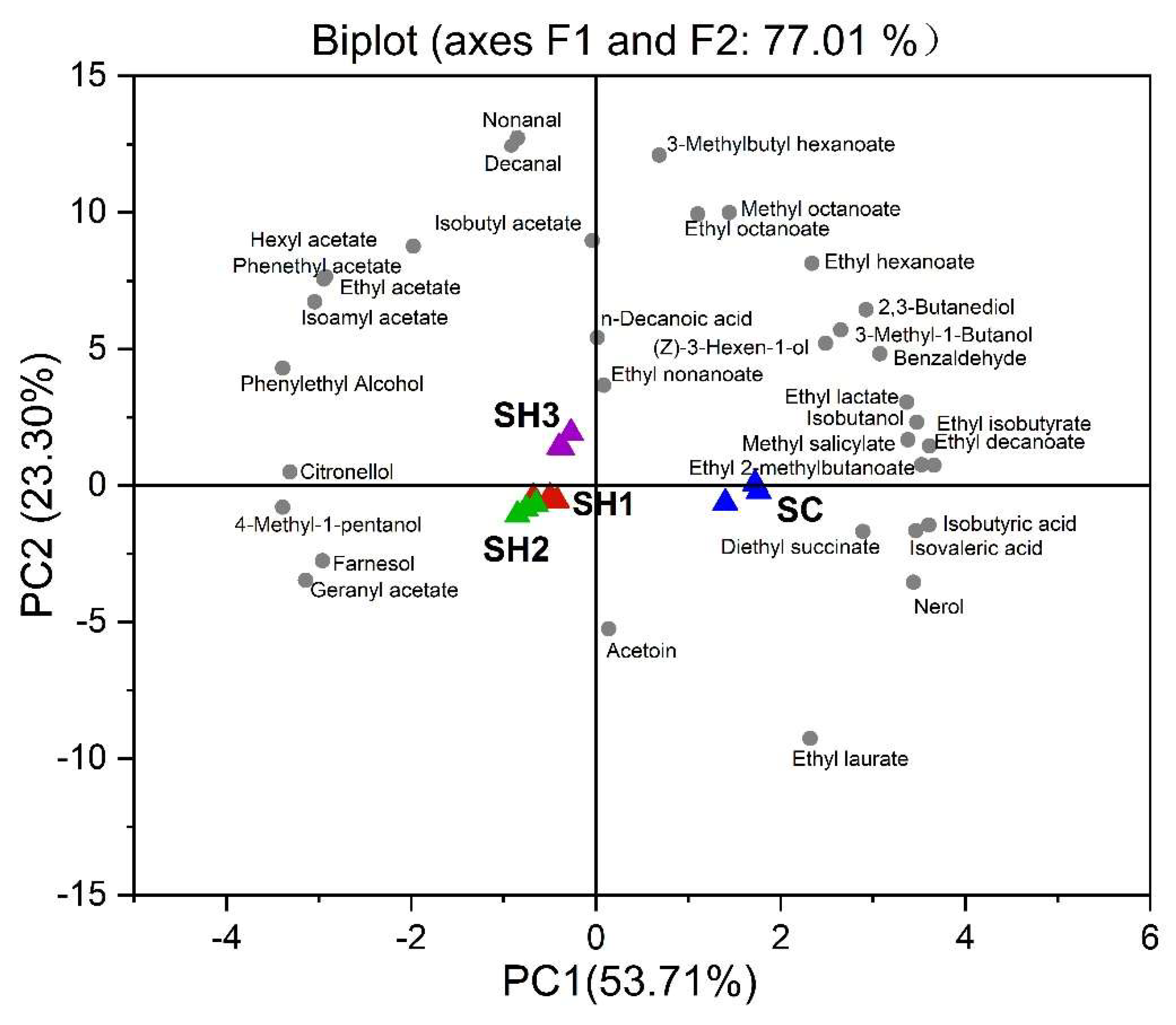
Principal component analysis (PCA) was further performed to visualize the discrimination of the entire set of volatile data of the resultant wines (Figure 5). The first two principal components (PCs) clearly separated the Merlot wines made with different fermentation modalities, and accounted for 77.01% of the variance in data (Figure 5). The first PC (F1:53.71%) separated the wine samples between sequential wines and the CECA wines, with SH1, SH2, and SH3 on the left-hand side of the plot, and SC on the right-hand side of the plot. The compounds that appear to be driving the separation along PC1 were isobutyric acid, isovaleric acid, n-decanoic acid, some ethyl esters, and many higher alcohols (Figure 5). Mixed inoculation treatments were associated with the production of acetate esters, phenylethyl alcohol, and some terpenes. The SH3 wines were separated along PC2 (higher left quadrant, Figure 5) from the remaining two mixed-inoculation wines. The SH3 wines were characterized by higher amounts of major acetate esters and phenylethyl alcohol, whereas the SH1 and SH2 wines were located at the negative axis of PC2, and were associated with 4-methyl-1-pentanol, farnesol, geranyl acetate, and acetoin (Figure 5).
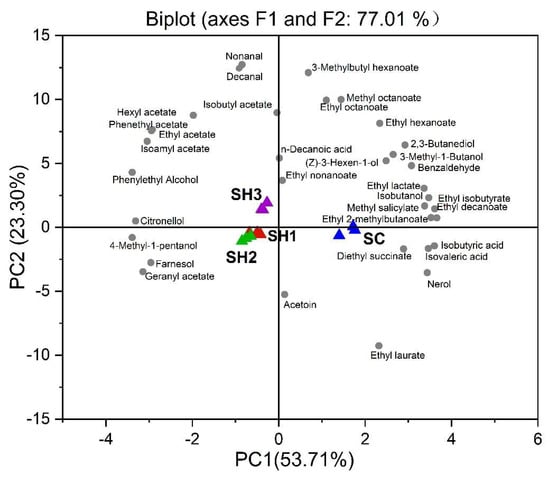
Figure 5.
Principal component analysis of the volatile profiles in Merlot wines. Yeast isolates (treatments) are presented using colored triangles, and the volatile compounds in grey dots. SC (CECA): S. cerevisiae, SH1: sequential inoculation fermentation of HuC-3-2 and CECA, SH2: sequential inoculation fermentation of HuC32-2-72 and CECA, SH3: sequential inoculation fermentation of HoC-4-2 and CECA.
4. Discussion
Yeast strains that are capable of reducing ethanol levels in wine whilst enhancing the overall wine quality are highly demanded by winemakers. In this study, four low-ethanol-producing strains, H. uvarum HuC-3-2 and HuB-2-2, and H. opuntiae HoC-4-2 and HoA-1-3, were primarily selected from 71 indigenous non-Saccharomyces strains obtained from four wine-producing regions in China (Figure 1B and Table A1). These four Hanseniaspora strains required more than 24 g/L of consumed sugar to produce 1% v/v of ethanol in Triple M medium (Table A2). Contrarily, it is documented that S. cerevisiae yeast uses 16.83 g/L to 17 g/L on average [36]. Hanseniaspora has been reported as one of the predominant yeast species present during the early stages of wine fermentation, of which many strains were able to lower wine ethanol [37,38]. The ethanol yield of Hanseniaspora spp. exhibited both interspecies and intraspecies variability, as shown in Figure 1. In this study, the ethanol yields of H. uvarum and H. opuntiae were found to be 0.185–0.464 g/g and 0.250–0.492 g/g, respectively (Table A2). There is also a finding that the ethanol yield from the isolates of H. uvarum and H. opuntiae, collected from South African grape must and vineyards, ranged between 0.28 and 0.53 g/g and 0.44 and 0.54 g/g, respectively [39].
In addition to the superiority in reducing ethanol yields of the four Hanseniaspora strains, it is also important to highlight that both strains completed fermentation in a medium with 200 g/L sugar and produced 7–9% v/v ethanol (Table A2). This indicated their better tolerance to higher concentrations of ethanol during fermentation, whilst cell growth was observed in the presence of 8% v/v ethanol (Table 2). A higher tolerance to ethanol would be beneficial for glucose uptake and ensure normal fermentation rates under oenological conditions since ethanol alters cell membrane structure [40]. Apart from ethanol tolerance, the four Hanseniaspora strains can also resist low fermentation temperatures (Table 2), indicating their potential applicability in producing low-ethanol white wines with desirable aroma profiles [41]. Additionally, three strains (except HuB-2-2) did not produce H2S, which has a negative organoleptic impact in wines [42,43], nor did they present any killer activities against S. cerevisiae (Table 2).
Strain HuC-3-2 outcompeted the other three strains due to its shorter fermentation duration and greater sugar consumption during the first two days in the Triple M medium (Figure 2). This strain was further subjected to ARTP mutagenesis, and a mutant strain, HuC32-2-72, with both low-ethanol-producing capability and rapid growth, was finally selected (Figure 3). ARTP, as a newly developed mutagenesis system, has been widely utilized in microorganism breeding. In addition to being environmentally friendly and easy to operate, ARTP is more convenient and effective in obtaining a higher mutation rate of microbes due to its stronger DNA damage capacity than traditional methods [42]. The mutant strains derived via ARTP mutagenesis are more able to obtain the target phenotypes, e.g., having an improved specific growth rate than the wild-type strains [21]. Nonetheless, there is a lack of studies on utilizing ARTP to reduce the ethanol yields of yeast strains. To the best of our knowledge, we reported the first successful case of using ARTP mutagenesis in wine yeast for more rapid growth whilst further reducing its ethanol yield (Figure 3).
The low-ethanol-producing phenotype of HuC-3-2, HuC32-2-72, and HoC-4-2 was then validated in a series of mixed inoculation treatments in Merlot juice with a higher initial sugar concentration, where ethanol modulation becomes further relevant. Fermentations inoculated with S. cerevisiae CECA alone finished first (Figure 4A), whereas the addition of S. cerevisiae induced a rapid population decline in Hanseniaspora (Figure 4C–E). Such a decrease might be attributed to the interaction between S. cerevisiae and Hanseniaspora, and the inhibitory impact of the multi-stressor wine environment, in particular, ethanol stress and the competition for nutrients [43]. The three selected Hanseniaspora strains allowed for an ethanol reduction of 1.4% v/v compared to S. cerevisiae CECA (Table 3). For Saccharomyces spp., alcoholic fermentation is usually efficient, with comparable ethanol yield between strains.
Strains belonging to Hanseniaspora have been reported to be able to reduce wine ethanol by 0.6–2.5% v/v when co-inoculated with S. cerevisiae [38]. In this study, sequential fermentations of H. uvarum HuC-3-2, H. uvarum HuC32-2-72, and H. opuntiae HoC-4-2 with S. cerevisiae in Merlot juice consistently resulted in a 1.4% v/v decrease in ethanol yield. The fate of carbon transferred during the fermentation of low-ethanol-producing non-Saccharomyces strains remains largely elusive. A decreased ethanol yield can be due to the respiratory metabolism in aerobic cultures [44]. In anaerobiosis, non-Saccharomyces yeast can transfer the carbon source in sugar metabolism to the non-ethanol metabolism terminal and lead to the formation of secondary metabolites, such as glycerol, organic acids, higher alcohols, and esters [15,16,45,46], or sinks that remain undetected. In the present study, we observed that H. uvarum HuC-3-2, H. uvarum HuC32-2-72, and H. opuntiae HoC-4-2 resulted in a decrease in ethanol content along with an increase in glycerol in both Triple M and Merlot juice fermentations (Figure 1J, Table 3). Similar results were reported before in wine fermentations [15], but our finding disagrees with others [47]. Our results indicated that glycerol yield, like other fermentation features, was an interspecies and strain-related trait.
Higher contents of acetic and lactic acids were found in sequential wines (Table 3), which was in agreement with previous studies [46,47]. In this study, a significant decrease in ethanol production was observed in mixed fermentations, even without any intervention on the oxygen level during fermentation, and without a detrimental increase in acetic acid (Table 3). Acetic acid is produced from the oxidation of acetaldehyde by aldehyde dehydrogenase or the hydrolysis of acetyl-CoA derived from pyruvate [48]. However, the sequential wines contained less pyruvic acid than the CECA wines (Table 3). Previous studies have also observed that the reduction in the ethanol yield by some non-Saccharomyces strains could not be fully explained by the overproduction of glycerol or organic acids, suggesting that respiration would be responsible, at least in part, for the decreased ethanol yield observed for these strains [49,50]. Sugars were probably partially consumed through the oxidative pathway to produce biomass and other unknown products.
Significant increases in acetate esters and ethyl esters in the sequential wines (Table 4) agree with a number of previous studies [47,51,52]. In general, esters produced during fermentation were dependent on the rate of ester synthesis and hydrolysis [53] In fact, esters remain the important carbon sink in a range of lower-ethanol yeasts, and have either negligible or positive sensory contributions. C2-alcohol O-acyl-transferases (AATs), which are encoded by ATF1 and ATF2, are critical and versatile enzymes that can utilize alcohols and acyl-CoA to form acetate esters [54]. A previous study suggested that overexpression of ATF1 enhanced the total ethyl acetate whilst decreasing the yield of ethanol slightly [55]. All of these findings highlighted the need for further research on ATF1 expression and AATs activity by Hanseniaspora, so as to offer a partial explanation for ethanol decrease. Of particular interest was the 2-3-fold increase in ethyl acetate in the sequential wines compared to the CECA wines (Table 4), which was close to or surpassed the level at which ethyl acetate could impart spoilage to the resultant wines. Thus, the influence of elevated ethyl acetate on the olfactory quality of the sequential wines requires further investigation. In addition, our study also demonstrated that Hanseniaspora can vigorously produce several key aromatic compounds (phenylethyl acetate and isoamyl acetate) in mixed inoculation fermentation with S. cerevisiae, which is in accord with some previous studies [51,56].
5. Conclusions
This study described the selection, optimization (via ARTP mutagenesis), and comprehensive characterization of three Hanseniaspora strains for their potential use in low-ethanol-producing starter cultures. The characterization of oenological properties indicated that the Hanseniaspora strains might be suitable for initiating fermentation without the risk of producing off-flavors (e.g., H2S). The three strains finally obtained resulted in approximately 1.4% v/v less ethanol in mixed fermentation wines than the S. cerevisiae monoculture wines. The evaluation of the yeast population and yeast metabolites highlighted glycerol and some esters as potential carbon sinks, partially accounting for the ethanol reduction. Further analysis on volatile compounds in the wine suggested that these strains have promise for use in combination with S. cerevisiae to manage ethanol levels and modulate aromas in wine, laying the groundwork for their broader application. Considering that these positive benefits were observed in sterile laboratory-scale fermentations, future studies on industrial-scale fermentation and sensory tasting should be undertaken to assess the effects of Hanseniaspora strains on the flavor profiles of the wines. Furthermore, optimizing the fermentation parameters, such as yeast cell concentration, fermentation time, or temperature, among others, of Hanseniaspora strains in fermentation will be essential, because these yeasts might be used in the production of wines that will be appreciated by the average consumer.
Author Contributions
Conceptualization, H.Y., W.F., H.Z., J.J. and Y.Q.; Data curation, H.Y., W.F. and J.J.; Formal analysis, H.Y., Y.W. and H.Z.; Investigation, Y.W.; Methodology, H.Y., Y.W., W.F., H.Z., J.J. and Y.Q.; Project administration, Y.Q.; Resources, W.F. and Y.Q.; Software, H.Y., Y.W. and W.F.; Supervision, J.J. and Y.Q.; Validation, Y.W. and H.Z.; Visualization, H.Y.; Writing—original draft, H.Y.; Writing—review and editing, Y.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research was Supported by the Ningxia Hui Autonomous Region Key R&D Project (2023BCF01027), the Changji State Science and Technology Support High-Quality Industrial Development Project (2022Z03), the Earmarked Fund for CARS (CARS-29-jg-3), and the Fundamental Research Funds for the Central Universities (2452020177).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding authors.
Acknowledgments
The Northwest A&F University provided support and facilities.
Conflicts of Interest
The authors declare no conflicts of interest.
Appendix A

Table A1.
26S D1/D2 fragment sizes of non-Saccharomyces yeasts and their identities compared to the reference yeasts.
Table A1.
26S D1/D2 fragment sizes of non-Saccharomyces yeasts and their identities compared to the reference yeasts.
| Strain | Size (bp) | Related Yeast | Type Strain | Identity (%) | Isolate Source |
|---|---|---|---|---|---|
| Hg01W23 | 629 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 99.67 | Jingyang, Shaanxi |
| HgA-1-2 | 628 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 100.00 | Jingyang, Shaanxi |
| HgA17-2-10 | 640 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 100.00 | Shanshan, Xinjiang |
| HgB-1-4 | 634 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 99.50 | Jingyang, Shaanxi |
| HgC-2-4 | 634 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 100.00 | Jingyang, Shaanxi |
| HgC-3-1 | 633 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 99.83 | Jingyang, Shaanxi |
| HgC-4-4 | 630 | Hanseniaspora guilliermondii TY14 | FJ972216.1 | 99.50 | Jingyang, Shaanxi |
| HgD-1-1 | 635 | Hanseniaspora guilliermondii TY20 | FJ972220.1 | 99.18 | Jingyang, Shaanxi |
| HgD-1-5 | 631 | Hanseniaspora guilliermondii TY14 | FJ972216.1 | 99.34 | Jingyang, Shaanxi |
| Ho01W25 | 623 | Hanseniaspora opuntiae HCM-NM44 | MK101218.1 | 99.51 | Jingyang, Shaanxi |
| Ho23W64 | 627 | Hanseniaspora opuntiae 11-1139 | MH465522.1 | 99.83 | Jingyang, Shaanxi |
| Ho23W68 | 625 | Hanseniaspora opuntiae HCM-NM44 | MK101218.1 | 99.67 | Jingyang, Shaanxi |
| HoA-1-3 | 618 | Hanseniaspora opuntiae JEY269 | KC111446.1 | 99.68 | Jingyang, Shaanxi |
| HoB-1-1 | 633 | Hanseniaspora opuntiae 11-1184 | MH465493.1 | 99.35 | Jingyang, Shaanxi |
| HoB-1-5 | 632 | Hanseniaspora opuntiae JW11-4 | KU316739.1 | 99.51 | Jingyang, Shaanxi |
| HoB-2-1 | 629 | Hanseniaspora opuntiae 11-1194 | MH465395.1 | 99.83 | Jingyang, Shaanxi |
| HoB-2-3 | 632 | Hanseniaspora opuntiae HCM-NM44 | MK101218.1 | 99.67 | Jingyang, Shaanxi |
| HoB7-15 | 631 | Hanseniaspora opuntiae JPMK66 | MH118515.1 | 100.00 | Jingyang, Shaanxi |
| HoC-1-3 | 633 | Hanseniaspora opuntiae JW11-4 | KU316739.1 | 99.67 | Jingyang, Shaanxi |
| HoC-1-4 | 634 | Hanseniaspora opuntiae SM10UFAM | MN268780.1 | 99.67 | Jingyang, Shaanxi |
| HoC-1-5 | 636 | Hanseniaspora opuntiae CEC 31C-9 | KF263943.1 | 99.83 | Jingyang, Shaanxi |
| HoC-3-3 | 631 | Hanseniaspora opuntiae SM10UFAM | MN268780.1 | 100.00 | Jingyang, Shaanxi |
| HoC-4-1 | 634 | Hanseniaspora opuntiae 11-1139 | MH465522.1 | 99.51 | Jingyang, Shaanxi |
| HoC-4-2 | 633 | Hanseniaspora opuntiae JEY269 | KC111446.1 | 99.83 | Jingyang, Shaanxi |
| HoD-1-2 | 635 | Hanseniaspora opuntiae CEC 31C-9 | KF263943.1 | 99.50 | Jingyang, Shaanxi |
| HoD-1-3 | 630 | Hanseniaspora opuntiae CEC 31C-9 | KF263943.1 | 99.34 | Jingyang, Shaanxi |
| Hu13W76 | 631 | Hanseniaspora uvarum X21-10 | MN337251.1 | 99.51 | Jingyang, Shaanxi |
| HuA6-1-6 | 637 | Hanseniaspora uvarum NS-O-182 | KT922901.1 | 100.00 | Shanshan, Xinjiang |
| HuA11-1-2 | 620 | Hanseniaspora uvarum CEC 32SA-51 | KF263942.1 | 99.35 | Shanshan, Xinjiang |
| HuB1S-6 | 628 | Hanseniaspora uvarum NS-PDC-10 | KT922480.1 | 99.67 | Jingyang, Shaanxi |
| HuB-2-2 | 634 | Hanseniaspora uvarum CBS:2589 | KY107852.1 | 99.67 | Jingyang, Shaanxi |
| HuBG-1-5 | 633 | Hanseniaspora uvarum NS-EM-77 | KT922349.1 | 99.67 | Jingyang, Shaanxi |
| HuBG-1-6 | 636 | Hanseniaspora uvarum NS-O-241 | KT922960.1 | 100.00 | Jingyang, Shaanxi |
| HuC-2-3 | 632 | Hanseniaspora uvarum 11-1148 | MH465536.1 | 99.83 | Jingyang, Shaanxi |
| HuC-2-5 | 635 | Hanseniaspora uvarum HN-NM-67 | MK101217.1 | 99.35 | Jingyang, Shaanxi |
| HuC-3-2 | 629 | Hanseniaspora uvarum HuW1 | KF992155.1 | 99.51 | Jingyang, Shaanxi |
| HuC-3-5 | 634 | Hanseniaspora uvarum X21-10 | MN337251.1 | 99.18 | Jingyang, Shaanxi |
| HuC-4-5 | 631 | Hanseniaspora uvarum X22-9 | MN337255.1 | 99.67 | Jingyang, Shaanxi |
| HuC413 | 633 | Hanseniaspora uvarum NS-O-51 | KT922774.1 | 99.83 | Gansu Mogao Winery |
| HuC466 | 637 | Hanseniaspora uvarum J15-5 | MN337257.1 | 99.66 | Gansu Mogao Winery |
| HuC534 | 639 | Hanseniaspora uvarum NS-EM-122 | KT922393.1 | 100.00 | Gansu Mogao Winery |
| HuD-1-4 | 630 | Hanseniaspora uvarum NS-O-51 | KT922744.1 | 99.83 | Jingyang, Shaanxi |
| HuD-2-1 | 623 | Hanseniaspora uvarum J15-5 | MN337257.1 | 99.51 | Jingyang, Shaanxi |
| HuD-2-3 | 633 | Hanseniaspora uvarum NS-O-16 | KT922739.1 | 99.67 | Jingyang, Shaanxi |
| HuD-2 -4 | 635 | Hanseniaspora uvarum X21-10 | MN337251.1 | 100.00 | Jingyang, Shaanxi |
| HuM69 | 635 | Hanseniaspora uvarum NS-EM-82 | KT922353.1 | 100.00 | Manas, Xinjiang |
| HuF-2-1 | 637 | Hanseniaspora uvarum NS-O-241 | KT922960.1 | 99.49 | Ningxia Royal Horse Winery |
| MafA5-5-3 | 558 | Metschnikowia aff. fructicola D3895 | AM286804.1 | 99.81 | Shanshan, Xinjiang |
| MafA9-4-5 | 578 | Metschnikowia aff. fructicola C83 | EU373448.1 | 99.80 | Shanshan, Xinjiang |
| MafA11-1-1 | 629 | Metschnikowia aff. fructicola D3895 | AM286804.1 | 99.81 | Shanshan, Xinjiang |
| MafA13-1-3 | 559 | Metschnikowia aff. fructicola D3895 | AM286804.1 | 99.81 | Shanshan, Xinjiang |
| MafC83 | 555 | Metschnikowia aff. fructicola D3895 | AM286804.1 | 99.81 | Gansu Mogao Winery |
| MafF-1-7 | 558 | Metschnikowia aff. fructicola D3895 | AM286804.1 | 99.62 | Ningxia Royal Horse Winery |
| PklC734 | 616 | Pichia kluyveri X27-5 | MN337254.1 | 99.51 | Gansu Mogao Winery |
| PklM114 | 617 | Pichia kluyveri X23-10 | MN337239.1 | 99.66 | Manas, Xinjiang |
| PklM118 | 631 | Pichia kluyveri X23-10 | MN337239.1 | 99.66 | Manas, Xinjiang |
| PklM144 | 625 | Pichia kluyveri CBS:7274 | KY108823.1 | 99.66 | Manas, Xinjiang |
| PklFS-2-7 | 630 | Pichia kluyveri YG14 | MF045458.1 | 99.83 | Ningxia Royal Horse Winery |
| PkuB4-19 | 625 | Pichia kudriavzevii CK12 | MN712334.1 | 99.66 | Jingyang, Shaanxi |
| PkuB4S-1 | 616 | Pichia kudriavzevii CBS:5147 | MH545928.1 | 99.83 | Jingyang, Shaanxi |
| PkuB6-1 | 619 | Pichia kudriavzevii ZJ-13 | KY283161.1 | 99.66 | Jingyang, Shaanxi |
| PkuB9S-2 | 618 | Pichia kudriavzevii CK12 | MN712334.1 | 100.00 | Jingyang, Shaanxi |
| PkuC392 | 617 | Pichia kudriavzevii CK12 | MN712334.1 | 99.17 | Gansu Mogao Winery |
| PkuC396 | 570 | Pichia kudriavzevii CK12 | MN712334.1 | 100.00 | Gansu Mogao Winery |
| PkuGS-1-18 | 622 | Pichia kudriavzevii CK12 | MN712334.1 | 100.00 | Helan Mountain, Ningxia |
| PkuGS-1-20 | 621 | Pichia kudriavzevii CK12 | MN712334.1 | 99.83 | Helan Mountain, Ningxia |
| PkuM-1-1 | 621 | Pichia kudriavzevii CK12 | MN712334.1 | 99.83 | Helan Mountain, Ningxia |
| PkuM100 | 621 | Pichia kudriavzevii 4_5 | MF461004.1 | 100.00 | Manas, Xinjiang |
| PkuM101 | 622 | Pichia kudriavzevii 4_5 | MF461004.1 | 100.00 | Manas, Xinjiang |
| PkuM130 | 623 | Pichia kudriavzevii CBS:5147 | MH545928.1 | 100.00 | Manas, Xinjiang |
| PkuM131 | 625 | Pichia kudriavzevii CBS:5147 | MH545928.1 | 100.00 | Manas, Xinjiang |

Table A2.
Fermentation parameters and secondary metabolites of the 71 non-Saccharomyces strains and S. cerevisiae CECA in synthetic grape juice *.
Table A2.
Fermentation parameters and secondary metabolites of the 71 non-Saccharomyces strains and S. cerevisiae CECA in synthetic grape juice *.
| Strain | OD600nm | Residual Sugar (g/L) | Sugar Consumption (g/L) | Ethanol (%v/v) | Ethanol Yield (g/g) | Changes in Ethanol Yield (%) | Glycerol (g/L) | Glycerol Yield (g/g) | Total Acid a (g/L) | Volatile Acid b (g/L) |
|---|---|---|---|---|---|---|---|---|---|---|
| Saccharomyces cerevisiae | ||||||||||
| CECA c | 3.39 ± 0.04 | 1.82 ± 0.10 | 198.18 ± 0.10 | 10.16 ± 0.01 | 0.404 ± 0.000 | 0 | 5.30 ± 0.00 | 0.026 ± 0.000 | 7.31 ± 0.11 | 0.50 ± 0.01 |
| Hanseniaspora guilliermondii | ||||||||||
| Hg01W23 | 1.39 ± 0.13 | 110.04 ± 5.91 | 89.96 ± 5.91 | 3.25 ± 0.20 | 0.409 ± 0.014 | −4.30 | 1.25 ± 0.07 | 0.014 ± 0.001 | 6.57 ± 0.47 | 0.52 ± 0.02 |
| HgA-1-2 | 1.58 ± 0.25 | 104.17 ± 4.47 | 95.83 ± 4.47 | 3.70 ± 0.78 | 0.303 ± 0.020 | −32.42 | 1.45 ± 0.05 | 0.015 ± 0.003 | 6.74 ± 0.16 | 0.49 ± 0.05 |
| HgA17-2-10 | 2.21 ± 0.19 | 60.07 ± 2.94 | 139.93 ± 2.94 | 4.64 ± 0.51 | 0.261 ± 0.018 | −40.11 | 2.50 ± 0.06 | 0.018 ± 0.003 | 6.06 ± 0.20 | 0.43 ± 0.03 |
| HgB-1-4 | 1.10 ± 0.02 | 139.01 ± 1.10 | 60.99 ± 1.10 | 3.58 ± 0.37 | 0.465 ± 0.024 | −1.96 | 3.10 ± 0.26 | 0.052 ± 0.002 | 6.22 ± 0.10 | 0.57 ± 0.01 |
| HgC-2-4 | 1.24 ± 0.04 | 123.06 ± 2.46 | 76.94 ± 2.46 | 4.13 ± 0.67 | 0.425 ± 0.025 | −13.94 | 3.03 ± 0.12 | 0.040 ± 0.006 | 6.69 ± 0.21 | 0.62 ± 0.05 |
| HgC-3-1 | 1.22 ± 0.04 | 126.49 ± 1.39 | 73.51 ± 1.39 | 4.55 ± 0.30 | 0.489 ± 0.023 | −1.04 | 2.97 ± 0.25 | 0.040 ± 0.001 | 6.62 ± 0.21 | 0.56 ± 0.03 |
| HgC-4-4 | 1.12 ± 0.01 | 126.07 ± 3.33 | 73.93 ± 3.33 | 4.25 ± 0.27 | 0.455 ± 0.048 | −8.01 | 2.90 ± 0.17 | 0.039 ± 0.001 | 6.55 ± 0.21 | 0.62 ± 0.02 |
| Hanseniaspora opuntiae | ||||||||||
| HoD-1-1 | 1.22 ± 0.02 | 131.10 ± 1.68 | 68.9 ± 1.68 | 4.03 ± 0.14 | 0.459 ± 0.015 | −7.15 | 2.40 ± 0.17 | 0.035 ± 0.007 | 6.65 ± 0.18 | 0.53 ± 0.04 |
| HoD-1-5 | 2.24 ± 0.13 | 15.08 ± 1.51 | 184.92 ± 1.51 | 9.49 ± 0.54 | 0.405 ± 0.027 | −4.00 | 4.97 ± 0.09 | 0.027 ± 0.002 | 7.59 ± 0.18 | 0.82 ± 0.02 |
| Ho01W25 | 1.83 ± 0.10 | 103.41 ± 6.99 | 96.59 ± 6.99 | 3.05 ± 0.40 | 0.252 ± 0.052 | −43.82 | 1.07 ± 0.15 | 0.011 ± 0.002 | 6.01 ± 0.30 | 0.47 ± 0.02 |
| Ho23W64 | 1.43 ± 0.11 | 118.63 ± 2.58 | 81.37 ± 2.58 | 3.53 ± 0.14 | 0.358 ± 0.015 | −16.24 | 1.27 ± 0.15 | 0.016 ± 0.001 | 6.24 ± 0.25 | 0.46 ± 0.06 |
| Ho23W68 | 1.63 ± 0.17 | 127.7 ± 3.58 | 72.30 ± 3.58 | 4.11 ± 0.21 | 0.494 ± 0.211 | −0.03 | 1.77 ± 0.15 | 0.027 ± 0.003 | 6.58 ± 0.12 | 0.51 ± 0.04 |
| HoA-1-3 | 2.46 ± 0.10 | 26.11 ± 2.13 | 173.89 ± 2.13 | 6.58 ± 0.39 | 0.299 ± 0.003 | −30.98 | 3.25 ± 0.07 | 0.019 ± 0.002 | 6.27 ± 0.20 | 0.56 ± 0.07 |
| HoB-1-1 | 1.54 ± 0.21 | 65.65 ± 2.06 | 134.35 ± 2.06 | 6.38 ± 0.41 | 0.378 ± 0.028 | −13.35 | 3.13 ± 0.15 | 0.023 ± 0.004 | 7.12 ± 0.30 | 0.50 ± 0.08 |
| HoB-1-5 | 1.15 ± 0.11 | 126.12 ± 3.75 | 73.88 ± 3.75 | 3.63 ± 0.20 | 0.389 ± 0.036 | −21.38 | 3.23 ± 0.23 | 0.044 ± 0.001 | 6.32 ± 0.44 | 0.58 ± 0.06 |
| HoB-2-1 | 1.17 ± 0.05 | 107.81 ± 3.72 | 92.19 ± 3.72 | 5.15 ± 0.26 | 0.445 ± 0.028 | −0.84 | 2.43 ± 0.06 | 0.027 ± 0.003 | 6.85 ± 0.05 | 0.48 ± 0.07 |
| HoB-2-3 | 1.70 ± 0.19 | 99.91 ± 2.40 | 100.09 ± 2.40 | 3.18 ± 0.32 | 0.250 ± 0.022 | −44.17 | 1.33 ± 0.15 | 0.013 ± 0.001 | 6.54 ± 0.17 | 0.49 ± 0.03 |
| HoB7-15 | 1.48 ± 0.08 | 124.65 ± 3.74 | 75.35 ± 3.74 | 4.26 ± 0.43 | 0.464 ± 0.006 | −6.16 | 1.63 ± 0.15 | 0.023 ± 0.005 | 10.11 ± 0.39 | 1.02 ± 0.03 |
| HoC-1-3 | 1.65 ± 0.11 | 116.74 ± 2.65 | 83.26 ± 2.65 | 2.96 ± 0.40 | 0.283 ± 0.008 | −33.77 | 1.27 ± 0.21 | 0.015 ± 0.003 | 6.76 ± 0.28 | 0.47 ± 0.04 |
| HoC-1-4 | 1.36 ± 0.17 | 103.72 ± 3.54 | 96.28 ± 3.54 | 5.61 ± 0.42 | 0.459 ± 0.018 | 2.28 | 2.83 ± 0.12 | 0.029 ± 0.003 | 6.48 ± 0.03 | 0.54 ± 0.02 |
| HoC-1-5 | 1.23 ± 0.04 | 128.54 ± 1.01 | 71.46 ± 1.01 | 4.08 ± 0.61 | 0.469 ± 0.001 | −5.11 | 2.80 ± 0.10 | 0.042 ± 0.005 | 6.38 ± 0.18 | 0.56 ± 0.04 |
| HoC-3-3 | 1.23 ± 0.04 | 131.31 ± 1.65 | 68.69 ± 1.65 | 4.30 ± 0.47 | 0.492 ± 0.016 | −0.43 | 3.27 ± 0.12 | 0.048 ± 0.008 | 6.85 ± 0.34 | 0.63 ± 0.04 |
| HoC-4-1 | 1.10 ± 0.01 | 128.71 ± 2.77 | 71.29 ± 2.77 | 3.92 ± 0.39 | 0.434 ± 0.043 | −12.21 | 2.73 ± 0.15 | 0.038 ± 0.003 | 6.46 ± 0.28 | 0.59 ± 0.03 |
| HoC-4-2 | 2.40 ± 0.03 | 19.74 ± 1.65 | 180.26 ± 1.65 | 7.13 ± 0.70 | 0.314 ± 0.009 | −25.66 | 4.97 ± 0.25 | 0.028 ± 0.001 | 7.44 ± 0.07 | 0.75 ± 0.03 |
| HoD-1-2 | 1.24 ± 0.01 | 127.08 ± 2.47 | 72.92 ± 2.47 | 4.01 ± 0.23 | 0.435 ± 0.039 | −12.00 | 2.87 ± 0.06 | 0.039 ± 0.001 | 6.52 ± 0.11 | 0.54 ± 0.03 |
| HoD-1-3 | 1.89 ± 0.14 | 84.00 ± 1.89 | 116.00 ± 1.89 | 4.23 ± 0.47 | 0.301 ± 0.017 | −21.53 | 1.75 ± 0.09 | 0.015 ± 0.002 | 5.98 ± 0.52 | 0.60 ± 0.03 |
| Hanseniaspora uvarum | ||||||||||
| Hu13W76 | 2.47 ± 0.19 | 69.44 ± 3.67 | 130.56 ± 3.67 | 4.12 ± 0.69 | 0.244 ± 0.002 | −42.19 | 1.93 ± 0.17 | 0.014 ± 0.001 | 6.17 ± 0.50 | 0.67 ± 0.03 |
| HuA6-1-6 | 1.85 ± 0.20 | 99.8 ± 5.47 | 100.2 ± 5.47 | 4.30 ± 0.18 | 0.339 ± 0.014 | −24.39 | 1.93 ± 0.12 | 0.019 ± 0.000 | 6.05 ± 0.26 | 0.61 ± 0.02 |
| HuA11-1-2 | 2.02 ± 0.14 | 43.85 ± 4.13 | 156.15 ± 4.13 | 3.66 ± 0.24 | 0.185 ± 0.005 | −58.01 | 1.43 ± 0.09 | 0.009 ± 0.003 | 6.11 ± 0.58 | 0.55 ± 0.04 |
| HuB1S-6 | 1.84 ± 0.26 | 106.31 ± 4.79 | 93.69 ± 4.79 | 3.03 ± 0.20 | 0.257 ± 0.004 | −42.73 | 1.35 ± 0.07 | 0.014 ± 0.000 | 7.15 ± 0.05 | 0.44 ± 0.01 |
| HuB-2-2 | 2.56 ± 0.09 | 2.46 ± 0.26 | 197.54 ± 0.26 | 7.72 ± 0.59 | 0.308 ± 0.001 | −24.13 | 5.20 ± 0.17 | 0.028 ± 0.001 | 7.16 ± 0.09 | 0.85 ± 0.03 |
| HuBG-1-5 | 2.46 ± 0.20 | 13.58 ± 1.28 | 186.42 ± 1.28 | 8.52 ± 0.04 | 0.360 ± 0.022 | −12.49 | 4.90 ± 0.28 | 0.026 ± 0.002 | 7.73 ± 0.03d | 0.81 ± 0.01 |
| HuBG-1-6 | 2.15 ± 0.07 | 20.64 ± 1.55 | 179.36 ± 1.55 | 7.56 ± 0.35 | 0.332 ± 0.016 | −21.52 | 4.60 ± 0.14 | 0.026 ± 0.002 | 7.62 ± 0.21 | 0.76 ± 0.04 |
| HuC-2-3 | 2.50 ± 0.10 | 17.11 ± 1.64 | 182.89 ± 1.64 | 9.52 ± 0.26 | 0.411 ± 0.010 | −2.73 | 4.90 ± 0.14 | 0.025 ± 0.001 | 7.52 ± 0.14 | 0.81 ± 0.03 |
| HuC-2-5 | 1.89 ± 0.16 | 110.91 ± 1.72 | 89.09 ± 1.72 | 3.01 ± 0.38 | 0.271 ± 0.008 | −36.59 | 1.20 ± 0.06 | 0.013 ± 0.002 | 6.76 ± 0.32 | 0.51 ± 0.05 |
| HuC-3-2 | 2.32 ± 0.20 | 24.63 ± 0.68 | 175.37 ± 0.68 | 6.13 ± 0.29 | 0.274 ± 0.014 | −33.67 | 5.43 ± 0.25 | 0.030 ± 0.001 | 7.19 ± 0.38 | 0.75 ± 0.10 |
| HuC-3-5 | 2.31 ± 0.17 | 18.88 ± 1.07 | 181.12 ± 1.07 | 8.02 ± 0.07 | 0.349 ± 0.031 | −17.39 | 3.80 ± 0.02 | 0.022 ± 0.003 | 7.69 ± 0.34 | 0.86 ± 0.03 |
| HuC-4-5 | 1.47 ± 0.12 | 120.88 ± 4.31 | 79.12 ± 4.31 | 2.98 ± 0.38 | 0.307 ± 0.027 | −28.03 | 1.00 ± 0.00 | 0.013 ± 0.000 | 6.88 ± 0.09 | 0.44 ± 0.06 |
| HuC413 | 2.11 ± 0.15 | 21.87 ± 2.05 | 178.13 ± 2.05 | 7.63 ± 0.59 | 0.337 ± 0.065 | −19.13 | 4.83 ± 0.31 | 0.027 ± 0.001 | 7.42 ± 0.12 | 0.87 ± 0.05 |
| HuC466 | 0.97 ± 0.05 | 156.07 ± 2.93 | 43.93 ± 2.93 | 2.39 ± 0.26 | 0.429 ± 0.002 | 7.13 | 1.35 ± 0.07 | 0.036 ± 0.002 | 11.03 ± 0.73 | 0.90 ± 0.03 |
| HuC534 | 2.16 ± 0.03 | 16.00 ± 0.43 | 184.00 ± 0.43 | 7.42 ± 0.15 | 0.317 ± 0.015 | −24.86 | 4.90 ± 0.26 | 0.027 ± 0.002 | 7.44 ± 0.27 | 0.70 ± 0.04 |
| HuD-1-4 | 1.01 ± 0.13 | 145.83 ± 1.83 | 54.17 ± 1.83 | 3.19 ± 0.25 | 0.464 ± 0.026 | 11.84 | 2.53 ± 0.23 | 0.047 ± 0.006 | 6.53 ± 0.21 | 0.55 ± 0.06 |
| HuD-2-1 | 2.42 ± 0.17 | 7.34 ± 2.30 | 192.66 ± 2.30 | 10.47 ± 0.63 | 0.429 ± 0.021 | 6.18 | 5.07 ± 0.21 | 0.026 ± 0.001 | 7.41 ± 0.09 | 0.89 ± 0.03 |
| HuD-2-3 | 1.29 ± 0.06 | 129.46 ± 1.65 | 70.54 ± 1.65 | 3.91 ± 0.37 | 0.441 ± 0.037 | −10.77 | 2.63 ± 0.09 | 0.038 ± 0.002 | 6.72 ± 0.23 | 0.63 ± 0.00 |
| HuD-2-4 | 2.34 ± 0.16 | 3.32 ± 0.62 | 196.68 ± 0.62 | 10.42 ± 0.41 | 0.418 ± 0.015 | 2.82 | 5.57 ± 0.38 | 0.028 ± 0.002 | 7.51 ± 0.34 | 1.00 ± 0.01 |
| HuM69 | 1.66 ± 0.01 | 94.56 ± 4.31 | 105.44 ± 4.31 | 4.06 ± 0.24 | 0.305 ± 0.018 | −26.42 | 1.55 ± 0.07 | 0.015 ± 0.001 | 7.28 ± 0.08 | 0.67 ± 0.06 |
| HuF-2-1 | 2.61 ± 0.73 | 53.1 ± 1.58 | 146.90 ± 1.58 | 5.11 ± 0.45 | 0.276 ± 0.030 | −34.81 | 2.53 ± 0.38 | 0.017 ± 0.002 | 6.16 ± 0.37 | 0.58 ± 0.01 |
| Pichia kluyveri | ||||||||||
| PklC734 | 2.09 ± 0.14 | 3.68 ± 0.12 | 196.32 ± 0.12 | 8.55 ± 0.10 | 0.343 ± 0.006 | −15.39 | 6.90 ± 0.20 | 0.035 ± 0.001 | 8.00 ± 0.24 | 0.98 ± 0.04 |
| PklM114 | 2.99 ± 0.15 | 14.7 ± 0.31 | 185.30 ± 0.31 | 6.61 ± 1.47 | 0.280 ± 0.005 | −31.81 | 5.50 ± 0.49 | 0.029 ± 0.007 | 7.81 ± 0.15 | 0.94 ± 0.02 |
| PklM118 | 0.96 ± 0.06 | 93.81 ± 1.76 | 81.37 ± 1.76 | 2.93 ± 0.05 | 0.284 ± 0.009 | −31.47 | 1.65 ± 0.09 | 0.020 ± 0.007 | 7.49 ± 0.14 | 0.50 ± 0.07 |
| PklM144 | 2.30 ± 0.14 | 14.94 ± 1.33 | 185.06 ± 1.33 | 6.70 ± 0.60 | 0.286 ± 0.027 | −30.42 | 6.67 ± 0.15 | 0.036 ± 0.001 | 7.66 ± 0.13 | 0.87 ± 0.05 |
| PklFS-2-7 | 0.88 ± 0.09 | 139.86 ± 1.04 | 60.14 ± 1.04 | 3.35 ± 0.14 | 0.449 ± 0.010 | −5.47 | 2.40 ± 0.06 | 0.041 ± 0.001 | 10.01 ± 0.16 | 0.98 ± 0.04 |
| Pichia kudriavzevii | ||||||||||
| PkuB4-19 | 2.12 ± 0.22 | 6.50 ± 2.02 | 193.50 ± 2.02 | 9.43 ± 1.16 | 0.384 ± 0.044 | −3.39 | 6.60 ± 0.06 | 0.034 ± 0.002 | 8.10 ± 0.21 | 0.65 ± 0.04 |
| PkuB4S-1 | 2.14 ± 0.05 | 3.28 ± 0.07 | 196.72 ± 0.07 | 8.94 ± 0.22 | 0.359 ± 0.005 | −11.75 | 5.93 ± 0.15 | 0.030 ± 0.000 | 7.34 ± 0.26 | 0.66 ± 0.02 |
| PkuB6-1 | 1.99 ± 0.20 | 10.56 ± 0.70 | 189.44 ± 0.70 | 6.88 ± 0.37 | 0.287 ± 0.014 | −29.27 | 5.97 ± 0.45 | 0.031 ± 0.002 | 8.53 ± 0.25 | 0.57 ± 0.02 |
| PkuB9S-2 | 1.48 ± 0.09 | 36.42 ± 0.52 | 163.58 ± 0.52 | 6.11 ± 0.25 | 0.295 ± 0.020 | −34.03 | 5.40 ± 0.44 | 0.033 ± 0.004 | 8.51 ± 0.45 | 0.51 ± 0.03 |
| PkuC392 | 1.41 ± 0.21 | 37.7 ± 2.42 | 162.3 ± 2.42 | 6.62 ± 0.27 | 0.322 ± 0.033 | −28.12 | 4.21 ± 0.20 | 0.026 ± 0.006 | 7.49 ± 0.18 | 0.48 ± 0.02 |
| PkuC396 | 1.81 ± 0.18 | 36.38 ± 2.76 | 163.62 ± 2.76 | 5.32 ± 0.46 | 0.258 ± 0.023 | −42.42 | 4.07 ± 0.25 | 0.025 ± 0.002 | 7.21 ± 0.16 | 0.31 ± 0.02 |
| PkuGS-1-18 | 1.33 ± 0.09 | 66.07 ± 5.59 | 133.93 ± 5.59 | 6.83 ± 0.11 | 0.403 ± 0.018 | −7.61 | 6.10 ± 0.10 | 0.046 ± 0.001 | 6.63 ± 0.15 | 0.30 ± 0.01 |
| PkuGS-1-20 | 1.46 ± 0.11 | 57.02 ± 5.65 | 142.98 ± 5.65 | 6.71 ± 0.12 | 0.370 ± 0.008 | −12.62 | 6.33 ± 0.35 | 0.044 ± 0.004 | 9.71 ± 0.18 | 0.56 ± 0.03 |
| PkuM-1-1 | 1.59 ± 0.11 | 39.55 ± 1.21 | 160.45 ± 1.21 | 7.12 ± 0.37 | 0.351 ± 0.032 | −20.39 | 6.53 ± 0.47 | 0.041 ± 0.004 | 9.03 ± 0.26 | 0.52 ± 0.04 |
| PkuM100 | 1.64 ± 0.23 | 40.37 ± 2.44 | 159.63 ± 2.44 | 4.30 ± 0.14 | 0.213 ± 0.006 | −51.83 | 3.03 ± 0.17 | 0.019 ± 0.001 | 6.33 ± 0.16 | 0.41 ± 0.03 |
| PkuM101 | 1.62 ± 0.19 | 36.72 ± 3.82 | 163.28 ± 3.82 | 5.74 ± 0.30 | 0.277 ± 0.008 | −38.02 | 4.37 ± 0.32 | 0.027 ± 0.001 | 7.29 ± 0.18 | 0.40 ± 0.07 |
| PkuM130 | 1.56 ± 0.11 | 43.76 ± 2.99 | 156.24 ± 2.99 | 6.82 ± 0.33 | 0.344 ± 0.017 | −22.02 | 5.87 ± 0.06 | 0.038 ± 0.000 | 9.77 ± 0.26 | 0.57 ± 0.06 |
| PkuM131 | 1.72 ± 0.17 | 44.40 ± 1.12 | 155.60 ± 1.12 | 5.48 ± 0.44 | 0.279 ± 0.026 | −36.80 | 3.30 ± 0.13 | 0.021 ± 0.002 | 7.23 ± 0.35 | 0.35 ± 0.04 |
| Metschnikowia aff. fructicola | ||||||||||
| MafA5-5-3 | 1.32 ± 0.15 | 149.3 ± 4.79 | 50.70 ± 4.79 | 2.97 ± 0.11 | 0.460 ± 0.021 | 10.72 | 2.60 ± 0.00 | 0.052 ± 0.005 | 6.66 ± 0.20 | 0.16 ± 0.02 |
| MafA9-4-5 | 2.25 ± 0.01 | 7.02 ± 0.91 | 192.98 ± 0.91 | 8.62 ± 0.96 | 0.351 ± 0.028 | −13.15 | 7.40 ± 0.36 | 0.038 ± 0.001 | 7.69 ± 0.22 | 0.60 ± 0.02 |
| MafA11-1-1 | 2.72 ± 0.11 | 2.47 ± 0.76 | 197.53 ± 0.76 | 8.87 ± 1.21 | 0.354 ± 0.029 | −12.75 | 7.20 ± 0.22 | 0.036 ± 0.003 | 7.31 ± 0.56 | 0.63 ± 0.03 |
| MafA13-1-3 | 2.31 ± 0.11 | 4.38 ± 0.55 | 195.62 ± 0.55 | 8.88 ± 0.54 | 0.358 ± 0.017 | −11.88 | 7.33 ± 0.25 | 0.037 ± 0.001 | 7.28 ± 0.40 | 0.64 ± 0.05 |
| MafC83 | 1.51 ± 0.15 | 121.98 ± 2.42 | 78.02 ± 2.42 | 3.35 ± 0.31 | 0.342 ± 0.051 | −30.93 | 1.37 ± 0.15 | 0.018 ± 0.003 | 7.36 ± 0.22 | 0.10 ± 0.03 |
| MafF-1-7 | 2.53 ± 0.11 | 1.85 ± 0.15 | 198.15 ± 0.15 | 7.25 ± 0.37 | 0.289 ± 0.005 | −28.64 | 7.60 ± 0.10 | 0.038 ± 0.000 | 7.63 ± 0.37 | 0.67 ± 0.02 |
* The initial sugar concentration of synthetic must was 200 g/L. a Expressed as tartaric acid. b Expressed as acetic acid. c S. cerevisiae CECA was used as the control.
References
- Keller, M. Climate Change Impacts on Vineyards in Warm and Dry Areas: Challenges and Opportunities: From the ASEV Climate Change Symposium Part 1–Viticulture. Am. J. Enol. Vitic. 2023, 74, 23024. [Google Scholar] [CrossRef]
- Ubeda, C.; Hornedo-Ortega, R.; Cerezo, A.B.; Garcia-Parrilla, M.C.; Troncoso, A.M. Chemical Hazards in Grapes and Wine, Climate Change and Challenges to Face. Food Chem. 2020, 314, 126222. [Google Scholar] [CrossRef] [PubMed]
- Mavrommati, M.; Economou, C.N.; Kallithraka, S.; Papanikolaou, S.; Aggelis, G. Simultaneous Improvement of Fructophilicity and Ethanol Tolerance of Saccharomyces cerevisiae Strains through a Single Adaptive Laboratory Evolution Strategy. Carbon Resour. Convers. 2024, 100270. [Google Scholar] [CrossRef]
- Jiang, J.; Sumby, K.M.; Sundstrom, J.F.; Grbin, P.R.; Jiranek, V. Directed Evolution of Oenococcus oeni Strains for More Efficient Malolactic Fermentation in a Multi-Stressor Wine Environment. Food Microbiol. 2018, 73, 150–159. [Google Scholar] [CrossRef]
- Moreno-Olivares, J.D.; Giménez-Bañón, M.J.; Ruiz-García, L.; Cebrián-Pérez, A.; Bleda-Sánchez, J.A.; Gil-Muñoz, R. New Wines Characterised by Their Low Alcohol and Extraordinary Polyphenolic Content. Eur. Food Res. Technol. 2025, 251, 233–246. [Google Scholar] [CrossRef]
- Shaw, C.L.; Dolan, R.; Corsi, A.M.; Goodman, S.; Pearson, W. Exploring the Barriers and Triggers towards the Adoption of Low- and No-Alcohol (NOLO) Wines. Food Qual. Prefer. 2023, 110, 104932. [Google Scholar] [CrossRef]
- Gutiérrez, A.R.; Portu, J.; López, R.; Garijo, P.; González-Arenzana, L.; Santamaría, P. Carbonic Maceration Vinification: A Tool for Wine Alcohol Reduction. Food Chem. 2023, 426, 136558. [Google Scholar] [CrossRef]
- Bedoya, K.; Buetas, L.; Rozès, N.; Mas, A.; Portillo, M.C. Influence of Different Stress Factors during the Elaboration of Grape Must’s Pied de Cuve on the Dynamics of Yeast Populations during Alcoholic Fermentation. Food Microbiol. 2024, 123, 104571. [Google Scholar] [CrossRef]
- Garrigós, V.; Vallejo, B.; Mollà-Martí, E.; Picazo, C.; Peltier, E.; Marullo, P.; Matallana, E.; Aranda, A. Up-Regulation of Retrograde Response in Yeast Increases Glycerol and Reduces Ethanol during Wine Fermentation. J. Biotechnol. 2024, 390, 28–38. [Google Scholar] [CrossRef]
- Montevecchi, G.; Ricci, A.; Masino, F.; Ferrari, V.; Versari, A.; Antonelli, A. Profile of Red Wine Partially Dealcoholized with a Membrane-Based Technique and Strategies to Mitigate the Loss of Volatile Compounds. Curr. Res. Food Sci. 2024, 8, 100776. [Google Scholar] [CrossRef]
- Jolly, N.; Mehlomakulu, N.N.; Nortje, S.; Beukes, L.; Hoff, J.; Booyse, M.; Erten, H. Non-Saccharomyces Yeast for Lowering Wine Alcohol Levels: Partial Aeration versus Standard Conditions. FEMS Yeast Res. 2022, 22, foac002. [Google Scholar] [CrossRef] [PubMed]
- Benavides, S.; Franco, W.; Lecco, C.C.D.; Durán, A.; Urtubia, A. Evaluation of Indigenous Candida oleophila and Candida boidinii in Monoculture and Sequential Fermentations: Impact on Ethanol Reduction and Chemical Profile in Chilean Sauvignon Blanc Wines. J. Fungi 2022, 8, 259. [Google Scholar] [CrossRef] [PubMed]
- Mančić, S.; Stamenković Stojanović, S.; Danilović, B.; Djordjević, N.; Malićanin, M.; Lazić, M.; Karabegović, I. Oenological Characterization of Native Hanseniaspora uvarum Strains. Fermentation 2022, 8, 92. [Google Scholar] [CrossRef]
- Urbina, Á.; Calderón, F.; Benito, S. The Combined Use of Lachancea thermotolerans and Lactiplantibacillus plantarum (Former Lactobacillus plantarum) in Wine Technology. Foods 2021, 10, 1356. [Google Scholar] [CrossRef]
- Hranilovic, A.; Gambetta, J.M.; Jeffery, D.W.; Grbin, P.R.; Jiranek, V. Lower-Alcohol Wines Produced by Metschnikowia pulcherrima and Saccharomyces cerevisiae Co-fermentations: The Effect of Sequential Inoculation Timing. Int. J. Food Microbiol. 2020, 329, 108651. [Google Scholar] [CrossRef] [PubMed]
- Capece, A.; Pietrafesa, A.; Pietrafesa, R.; Garrigós, V.; Tedesco, F.; Romano, P.; Matallana, E.; Siesto, G.; Aranda, A. Impact of Starmerella bacillaris and Zygosaccharomyces bailii on Ethanol Reduction and Saccharomyces cerevisiae Metabolism during Mixed Wine Fermentations. Food Res. Int. 2022, 159, 111649. [Google Scholar] [CrossRef]
- Wang, B.; Tan, F.; Chu, R.; Li, G.; Li, L.; Yang, T.; Zhang, M. The Effect of Non-Saccharomyces Yeasts on Biogenic Amines in Wine. Trends Food Sci. Technol. 2021, 116, 1029–1040. [Google Scholar] [CrossRef]
- Fernandes, T.; Osório, C.; Sousa, M.J.; Franco-Duarte, R. Contributions of Adaptive Laboratory Evolution towards the Enhancement of the Biotechnological Potential of Non-conventional Yeast Species. J. Fungi 2023, 9, 186. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, M.L.; Chimeno, S.V.; Mercado, L.A.; Ciklic, I.F. Hybridization and Spore Dissection of Native Wine Yeasts for Improvement of Ethanol Resistance and Osmotolerance. World J. Microbiol. Biotechnol. 2022, 38, 225. [Google Scholar] [CrossRef]
- Zhang, Q.; Miao, R.; Feng, R.; Yan, J.; Wang, T.; Gan, Y.; Zhao, J.; Lin, J.; Gan, B. Application of Atmospheric and Room-Temperature Plasma (ARTP) to Microbial Breeding. Curr. Issues Mol. Biol. 2023, 45, 6466–6484. [Google Scholar] [CrossRef]
- Nie, X.; Xing, Y.; Li, Q.; Gao, F.; Wang, S.; Liu, P.; Li, X.; Tan, Z.; Wang, P.; Shi, H. ARTP Mutagenesis Promotes Selenium Accumulation in Saccharomyces boulardii. LWT 2022, 168, 113916. [Google Scholar] [CrossRef]
- Yun, J.; Zabed, H.M.; Zhang, Y.; Zhang, G.; Zhao, M.; Qi, X. Improving Tolerance and 1,3-Propanediol Production of Clostridium butyricum Using Physical Mutagenesis, Adaptive Evolution and Genome Shuffling. Bioresour. Technol. 2022, 363, 127967. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, Y.; Ye, D.; Song, Y.; Shi, J.; Qin, Y.; Liu, Y. Impact of Serine and Serine Synthesis Genes on H2S Release in Saccharomyces cerevisiae during Wine Fermentation. Food Microbiol. 2022, 103, 103961. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, Y.; Liu, M.; Qin, Y.; Liu, Y. Saccharomyces cerevisiae Isolates with Extreme Hydrogen Sulfide Production Showed Different Oxidative Stress Resistances Responses during Wine Fermentation by RNA Sequencing Analysis. Food Microbiol. 2019, 79, 147–155. [Google Scholar] [CrossRef]
- Liu, N.; Qin, Y.; Song, Y.; Ye, D.; Yuan, W.; Pei, Y.; Xue, B.; Liu, Y. Selection of Indigenous Saccharomyces cerevisiae Strains in Shanshan County (Xinjiang, China) for Winemaking and Their Aroma-Producing Characteristics. World J. Microbiol. Biotechnol. 2015, 31, 1781–1792. [Google Scholar] [CrossRef]
- Chen, Y.; Jiang, J.; Song, Y.; Zang, X.; Wang, G.; Pei, Y.; Song, Y.; Qin, Y.; Liu, Y. Yeast Diversity during Spontaneous Fermentations and Oenological Characterisation of Indigenous Saccharomyces cerevisiae for Potential as Wine Starter Cultures. Microorganisms 2022, 10, 1455. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.; Liu, Y.; Wang, X.; Gao, C.; Mu, J.; Wang, W.; Wang, J. Correlations between Microbes with Volatile Compounds and Physicochemical Indicators of Cabernet Sauvignon Wines Fermented with Different Starters. LWT-Food Sci. Technol. 2024, 198, 115918. [Google Scholar] [CrossRef]
- Yang, L.; Zhu, X.; Mao, Y.; Zhang, X.; Xu, B.; Yang, X. Effect of Different Inoculation Strategies of Mixed Culture Saccharomyces cerevisiae/Oenococcus oeni on the Aroma Quality of Chardonnay Wine. Food Res. Int. 2024, 190, 114636. [Google Scholar] [CrossRef]
- Stribny, J.; Querol, A.; Pérez-Torrado, R. Differences in Enzymatic Properties of the Saccharomyces kudriavzevii and Saccharomyces uvarum Alcohol Acetyltransferases and Their Impact on Aroma-Active Compounds Production. Front. Microbiol. 2016, 7, 897. [Google Scholar] [CrossRef]
- Carpena, M.; Fraga-Corral, M.; Otero, P.; Nogueira, R.A.; Garcia-Oliveira, P.; Prieto, M.A.; Simal-Gandara, J. Secondary Aroma: Influence of Wine Microorganisms in Their Aroma Profile. Foods 2021, 10, 51. [Google Scholar] [CrossRef]
- Tekarslan-Sahin, S.H. Adaptive Laboratory Evolution of Yeasts for Aroma Compound Production. Fermentation 2022, 8, 372. [Google Scholar] [CrossRef]
- Wei, J.; Zhang, Y.; Wang, Y.; Ju, H.; Niu, C.; Song, Z.; Yuan, Y.; Yue, T. Assessment of Chemical Composition and Sensorial Properties of Ciders Fermented with Different Non-Saccharomyces Yeasts in Pure and Mixed Fermentations. Int. J. Food Microbiol. 2020, 318, 108471. [Google Scholar] [CrossRef] [PubMed]
- van Wyk, N.; Grossmann, M.; Wendland, J.; von Wallbrunn, C.; Pretorius, I.S. The Whiff of Wine Yeast Innovation: Strategies for Enhancing Aroma Production by Yeast during Wine Fermentation. J. Agric. Food Chem. 2019, 67, 13496–13505. [Google Scholar] [CrossRef]
- Martin, V.; Valera, M.J.; Medina, K.; Boido, E.; Carrau, F. Oenological Impact of the Hanseniaspora/Kloeckera Yeast Genus on Wines-A Review. Fermentation 2018, 4, 76. [Google Scholar] [CrossRef]
- Coletta, A.; Toci, A.T.; Pati, S.; Ferrara, G.; Grieco, F.; Tufariello, M.; Crupi, P. Effect of Soil Management and Training System on Negroamaro Wine Aroma. Foods 2021, 10, 454. [Google Scholar] [CrossRef] [PubMed]
- Mestre, M.V.; Maturano, Y.P.; Gallardo, C.; Combina, M.; Mercado, L.; Toro, M.E.; Carrau, F.; Vazquez, F.; Dellacassa, E. Impact on Sensory and Aromatic Profile of Low Ethanol Malbec Wines Fermented by Sequential Culture of Hanseniaspora uvarum and Saccharomyces cerevisiae Native Yeasts. Fermentation 2019, 5, 65. [Google Scholar] [CrossRef]
- van Wyk, N.; Badura, J.; von Wallbrunn, C.; Pretorius, I.S. Exploring Future Applications of the Apiculate Yeast Hanseniaspora. Crit. Rev. Biotechnol. 2024, 44, 100–119. [Google Scholar] [CrossRef]
- Gonzalez, R.; Guindal, A.M.; Tronchoni, J.; Morales, P. Biotechnological Approaches to Lowering the Ethanol Yield during Wine Fermentation. Biomolecules 2021, 11, 1569. [Google Scholar] [CrossRef]
- Rossouw, D.; Bauer, F.F. Exploring the Phenotypic Space of Non-Saccharomyces Wine Yeast Biodiversity. Food Microbiol. 2016, 55, 32–46. [Google Scholar] [CrossRef]
- Navarro-Tapia, E.; Querol, A.; Pérez-Torrado, R. Membrane Fluidification by Ethanol Stress Activates Unfolded Protein Response in Yeasts. Microb. Biotechnol. 2018, 11, 465–475. [Google Scholar] [CrossRef]
- Du, Q.; Ye, D.; Zang, X.; Nan, H.; Liu, Y. Effect of Low Temperature on the Shaping of Yeast-Derived Metabolite Compositions during Wine Fermentation. Food Res. Int. 2022, 162, 112016. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Liu, E.; Yin, Y.; Yang, L.; Huang, Q.; Chen, S.; Ho, C.-T. Enhancing Activities of Salt-Tolerant Proteases Secreted by Aspergillus oryzae Using Atmospheric and Room-Temperature Plasma Mutagenesis. J. Agric. Food Chem. 2020, 68, 2757–2764. [Google Scholar] [CrossRef]
- Wang, C.; Mas, A.; Esteve-Zarzoso, B. Interaction between Hanseniaspora uvarum and Saccharomyces cerevisiae during Alcoholic Fermentation. Int. J. Food Microbiol. 2015, 206, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Quirós, M.; Rojas, V.; Gonzalez, R.; Morales, P. Selection of Non-Saccharomyces Yeast Strains for Reducing Alcohol Levels in Wine by Sugar Respiration. Int. J. Food Microbiol. 2014, 181, 85–91. [Google Scholar] [CrossRef]
- Hranilovic, A.; Albertin, W.; Capone, D.L.; Gallo, A.; Grbin, P.R.; Danner, L.; Bastian, S.E.P.; Masneuf-Pomarede, I.; Coulon, J.; Bely, M.; et al. Impact of Lachancea thermotolerans on Chemical Composition and Sensory Profiles of Merlot Wines. Food Chem. 2021, 349, 129015. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hong, M.; Qi, B. Impact of Torulaspora delbrueckii During Fermentation on Aromatic Profile of Vidal Blanc Icewine. Front. Microbiol. 2022, 13, 860128. [Google Scholar] [CrossRef]
- Vaquero, C.; Escott, C.; Heras, J.M.; Carrau, F.; Morata, A. Co-Inoculations of Lachancea thermotolerans with Different Hanseniaspora Spp.: Acidification, Aroma, Biocompatibility, and Effects of Nutrients in Wine. Food Res. Int. 2022, 161, 111891. [Google Scholar] [CrossRef]
- Jiang, X.; Lu, Y.; Liu, S.Q. Effects of Different Yeasts on Physicochemical and Oenological Properties of Red Dragon Fruit Wine Fermented with Saccharomyces cerevisiae, Torulaspora delbrueckii and Lachancea thermotolerans. Microorganisms 2020, 8, 315. [Google Scholar] [CrossRef]
- Gonzalez, R.; Quirós, M.; Morales, P. Yeast Respiration of Sugars by Non-Saccharomyces Yeast Species: A Promising and Barely Explored Approach to Lowering Alcohol Content of Wines. Trends Food Sci. Technol. 2013, 29, 55–61. [Google Scholar] [CrossRef]
- Magyar, I.; Tóth, T. Comparative Evaluation of Some Oenological Properties in Wine Strains of Candida stellata, Candida zemplinina, Saccharomyces uvarum and Saccharomyces cerevisiae. Food Microbiol. 2011, 28, 94–100. [Google Scholar] [CrossRef]
- Hu, K.; Jin, G.-J.; Mei, W.-C.; Li, T.; Tao, Y.-S. Increase of Medium-Chain Fatty Acid Ethyl Ester Content in Mixed H. uvarum/S. cerevisiae Fermentation Leads to Wine Fruity Aroma Enhancement. Food Chem. 2018, 239, 495–501. [Google Scholar] [CrossRef]
- Zhang, B.; Tang, C.; Yang, D.; Liu, H.; Xue, J.; Duan, C.; Yan, G. Effects of Three Indigenous Non-Saccharomyces Yeasts and Their Pairwise Combinations in Co-fermentation with Saccharomyces cerevisiae on Volatile Compounds of Petit Manseng Wines. Food Chem. 2022, 368, 130807. [Google Scholar] [CrossRef] [PubMed]
- Sumby, K.M.; Grbin, P.R.; Jiranek, V. Microbial Modulation of Aromatic Esters in Wine: Current Knowledge and Future Prospects. Food Chem. 2010, 121, 1–16. [Google Scholar] [CrossRef]
- Kruis, A.J.; Bohnenkamp, A.C.; Patinios, C.; van Nuland, Y.M.; Levisson, M.; Mars, A.E.; van den Berg, C.; Kengen, S.W.M.; Weusthuis, R.A. Microbial Production of Short and Medium Chain Esters: Enzymes, Pathways, and Applications. Biotechnol. Adv. 2019, 37, 107407. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Lin, L.; Chen, Z.; Wang, K.; Sun, J.; Zhu, T. The Same Genetic Regulation Strategy Produces Inconsistent Effects in Different Saccharomyces cerevisiae Strains for 2-Phenylethanol Production. Appl. Microbiol. Biotechnol. 2022, 106, 4041–4052. [Google Scholar] [CrossRef]
- Viana, F.; Belloch, C.; Vallés, S.; Manzanares, P. Monitoring a Mixed Starter of Hanseniaspora vineae–Saccharomyces cerevisiae in Natural Must: Impact on 2-Phenylethyl Acetate Production. Int. J. Food Microbiol. 2011, 151, 235–240. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).