Characterization and Inkjet Printing of an RNA Aptamer for Paper-Based Biosensing of Ciprofloxacin
Abstract
1. Introduction
2. Materials and Methods
2.1. In-Line Probing Experiments
2.2. Fluorescence Titration Experiments
2.3. Inkjet Printing Experiments
3. Results
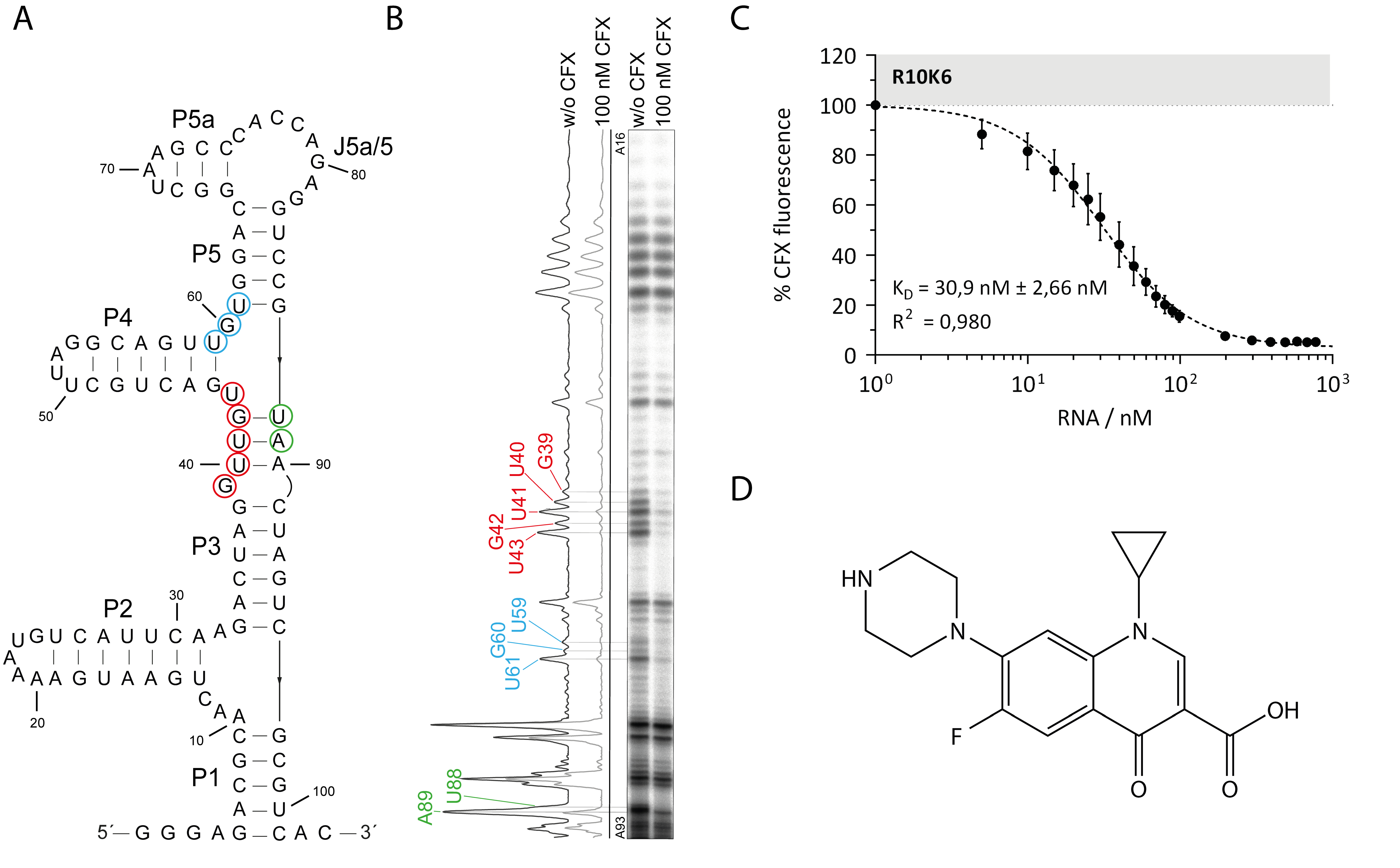
3.1. Probing the Secondary Structure of the CFX Aptamer R10K6
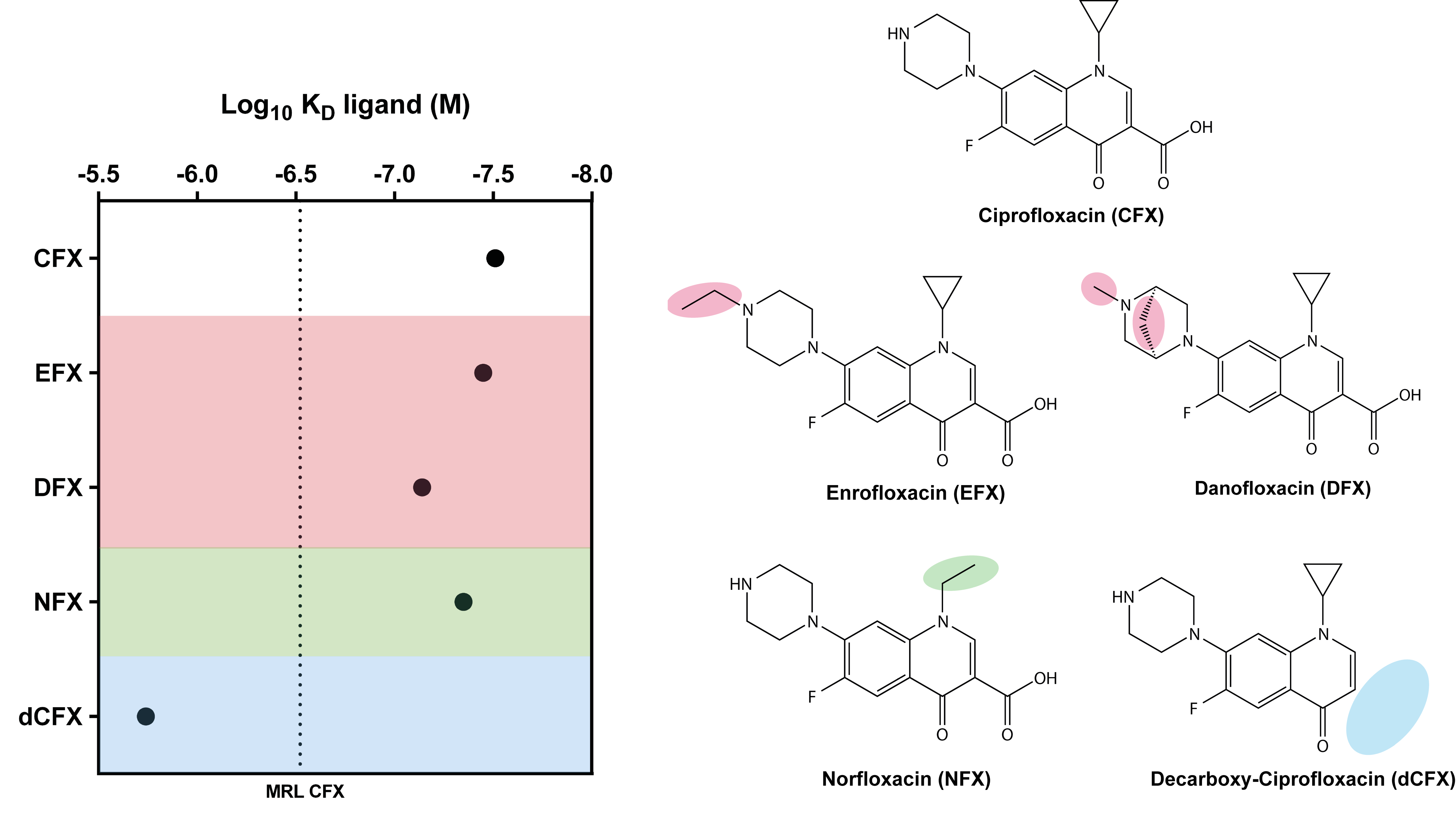
3.2. Ligand Binding Specificity of R10K6
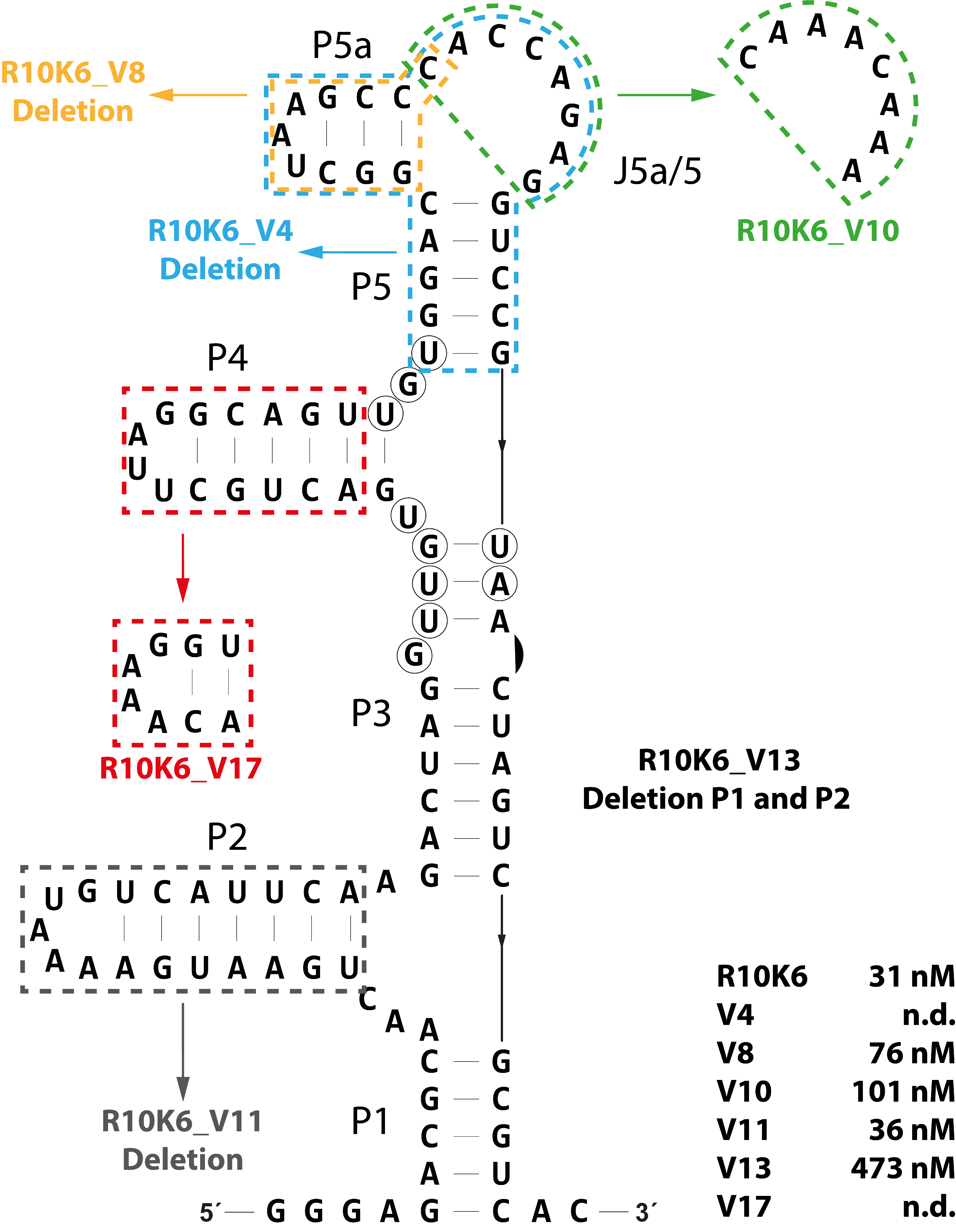
3.3. Defining a Minimal CFX-Binding Aptamer
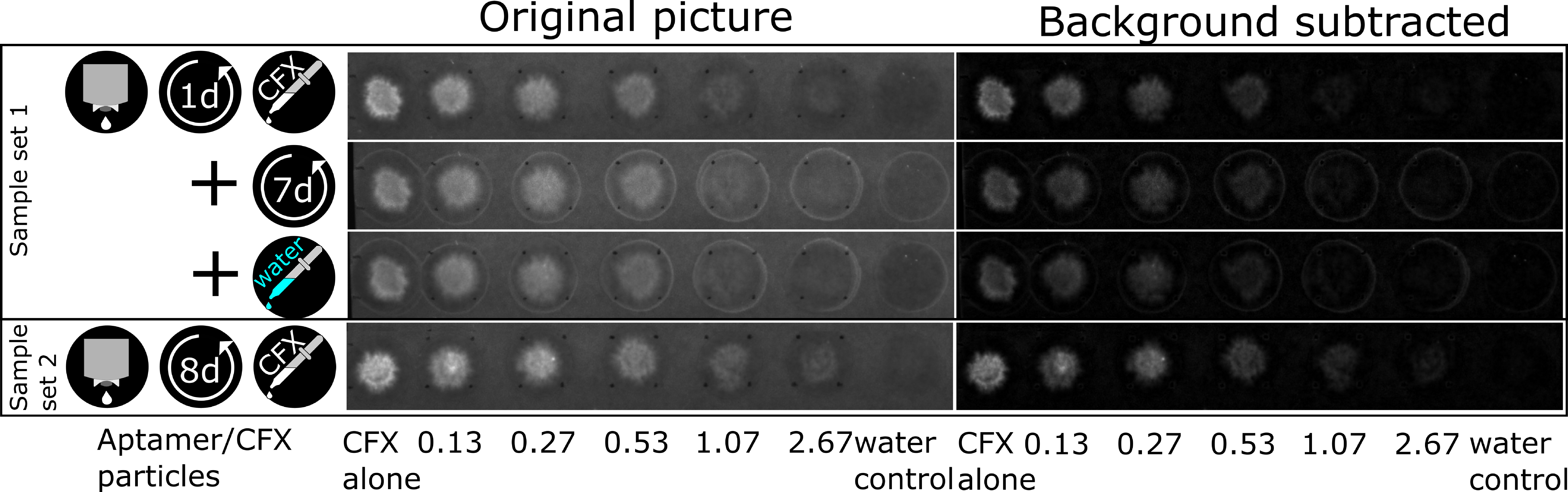
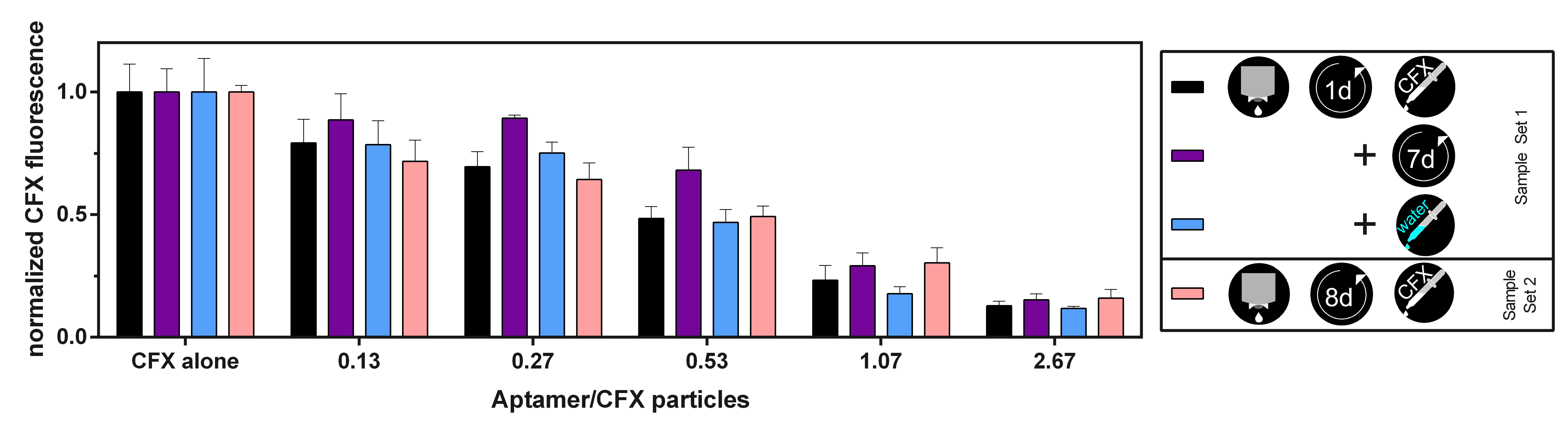
3.4. The CFX Aptamer Can Be Printed and Retains Functionality
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FDA. 2016 Summary Report on Antimicrobials Sold or Distributed for Use in Food-Producing Animals; FDA: Silver Spring, MD, USA, 2017; pp. 1–67. [Google Scholar]
- Kivirand, K.; Kagan, M.; Rinken, T. Calibrating Biosensors in Flow-Through Set-Ups: Studies with Glucose Optrodes. In State of the Art in Biosensors—General Aspects; IntechOpen: London, UK, 2013. [Google Scholar]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Garst, A.D.; Edwards, A.L.; Batey, R.T. Riboswitches: Structures and mechanisms. Cold Spring Harb. Perspect. Biol. 2011, 3, a003533. [Google Scholar] [CrossRef] [PubMed]
- Feng, C.; Dai, S.; Wang, L. Optical aptasensors for quantitative detection of small biomolecules: A review. Biosens. Bioelectron. 2014, 59, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, B.; Wu, Y.; Jia, S.; Fan, T.; Zhang, Z.; Zhang, C. Fast determination of the tetracyclines in milk samples by the aptamer biosensor. Analyst 2010, 135, 2706–2710. [Google Scholar] [CrossRef] [PubMed]
- Song, K.-M.; Cho, M.; Jo, H.; Min, K.; Jeon, S.H.; Kim, T.; Han, M.S.; Ku, J.K.; Ban, C. Gold nanoparticle-based colorimetric detection of kanamycin using a DNA aptamer. Anal. Biochem. 2011, 415, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Mehlhorn, A.; Rahimi, P.; Joseph, Y. Aptamer-Based Biosensors for Antibiotic Detection: A Review. Biosensors 2018, 8, 54. [Google Scholar] [CrossRef] [PubMed]
- Carrasquilla, C.; Little, J.R.L.; Li, Y.; Brennan, J.D. Patterned Paper Sensors Printed with Long-Chain DNA Aptamers. Chemistry 2015, 21, 7369–7373. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, N.; Horii, K.; Akitomi, J.; Kato, S.; Shiratori, I.; Waga, I. An Aptamer-Based Biosensor for Direct, Label-Free Detection of Melamine in Raw Milk. Sensors 2018, 18, 3227. [Google Scholar] [CrossRef] [PubMed]
- Strehlitz, B.; Nikolaus, N.; Stoltenburg, R. Protein Detection with Aptamer Biosensors. Sensors 2008, 8, 4296–4307. [Google Scholar] [CrossRef] [PubMed]
- Cháfer-Pericás, C.; Maquieira, Á.; Puchades, R. Fast screening methods to detect antibiotic residues in food samples. TrAC Trends Anal. Chem. 2010, 29, 1038–1049. [Google Scholar] [CrossRef]
- Wu, W.; Zhao, S.; Mao, Y.; Fang, Z.; Lu, X.; Zeng, L. A sensitive lateral flow biosensor for Escherichia coli O157:H7 detection based on aptamer mediated strand displacement amplification. Anal. Chim. Acta 2015, 861, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Groher, F.; Bofill-Bosch, C.; Schneider, C.; Braun, J.; Jager, S.; Geißler, K.; Hamacher, K.; Suess, B. Riboswitching with ciprofloxacin—Development and characterization of a novel RNA regulator. Nucleic Acids Res. 2018, 46, 2121–2132. [Google Scholar] [CrossRef] [PubMed]
- Seetharaman, S.; Zivarts, M.; Sudarsan, N.; Breaker, R.R. Immobilized RNA switches for the analysis of complex chemical and biological mixtures. Nat. Biotechnol. 2001, 19, 336. [Google Scholar] [CrossRef] [PubMed]
- Regulski, E.E.; Breaker, R.R. In-Line Probing Analysis of Riboswitches. In Post-Transcriptional Gene Regulation; Wilusz, J., Totowa, N.J., Eds.; Humana Press: New York, NY, USA, 2008; pp. 53–67. [Google Scholar]
- Stamm, J.; Spiehl, D.; Jaeger, J.; Groher, F.; Meckel, T.; Suess, B.; Dörsam, E. DNA and DNA Staining as a Test System for the Development of an Aptamer-based Biosensor for Sensing Antibiotics. In Proceedings of the 45th International Research Conference of IARIGAI, Warsaw, Poland, 3–7 October 2018. [Google Scholar]
- Waugh, A.; Gendron, P.; Altman, R.; Brown, J.W.; Case, D.; Gautheret, D.; Harvey, S.C.; Leontis, N.; Westbrook, J.; Westhof, E.; et al. RNAML: A standard syntax for exchanging RNA information. RNA 2002, 8, 707–717. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M.; Jacobson, A.B. Using reliability information to annotate RNA secondary structures. RNA 1998, 4, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Jaén-Oltra, J.; Salabert-Salvador, M.T.; García-March, F.J.; Pérez-Giménez, F.; Tomás-Vert, F. Artificial Neural Network Applied to Prediction of Fluorquinolone Antibacterial Activity by Topological Methods. J. Med. Chem. 2000, 43, 1143–1148. [Google Scholar] [CrossRef] [PubMed]
- Rhône, R.-B. Current EU Antibiotic Maximum Residue Limits; R-Biopharm Rhône Ltd.: Glasgow, UK, 2011. [Google Scholar]
- Barroso, J.M. COMMISSION REGULATION (EU) No 37/2010 of 22 December 2009 on pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin. Off. J. Eur. Union 2009, 1–72. [Google Scholar]
| Fluoroquinolone | Excitation (nm) * | Emission (nm) * |
|---|---|---|
| Ciprofloxacin (CFX) | 335 | 420 |
| Danofloxacin (DFX) | 325 | 425 |
| Decarboxy-Ciprofloxacin (dCFX) | 320 | 420 |
| Enrofloxacin (EFX) | 330 | 420 |
| Norfloxacin (NFX) | 330 | 425 |
| Dropspace/mm | Grid Size | mL/m2 | µL | Aptamer/CFX Particles |
|---|---|---|---|---|
| 0.253 | 28 | 5 | 0.25 | 0.13 |
| 0.177 | 40 | 10 | 0.5 | 0.27 |
| 0.124 | 57 | 20 | 1 | 0.53 |
| 0.088 | 80 | 40 | 2 | 1.07 |
| 0.056 | 127 | 100 | 5 | 2.67 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jaeger, J.; Groher, F.; Stamm, J.; Spiehl, D.; Braun, J.; Dörsam, E.; Suess, B. Characterization and Inkjet Printing of an RNA Aptamer for Paper-Based Biosensing of Ciprofloxacin. Biosensors 2019, 9, 7. https://doi.org/10.3390/bios9010007
Jaeger J, Groher F, Stamm J, Spiehl D, Braun J, Dörsam E, Suess B. Characterization and Inkjet Printing of an RNA Aptamer for Paper-Based Biosensing of Ciprofloxacin. Biosensors. 2019; 9(1):7. https://doi.org/10.3390/bios9010007
Chicago/Turabian StyleJaeger, Jeannine, Florian Groher, Jacqueline Stamm, Dieter Spiehl, Johannes Braun, Edgar Dörsam, and Beatrix Suess. 2019. "Characterization and Inkjet Printing of an RNA Aptamer for Paper-Based Biosensing of Ciprofloxacin" Biosensors 9, no. 1: 7. https://doi.org/10.3390/bios9010007
APA StyleJaeger, J., Groher, F., Stamm, J., Spiehl, D., Braun, J., Dörsam, E., & Suess, B. (2019). Characterization and Inkjet Printing of an RNA Aptamer for Paper-Based Biosensing of Ciprofloxacin. Biosensors, 9(1), 7. https://doi.org/10.3390/bios9010007






