Dynamics of Soil Bacterial Communities over Rice Growth Stages Under Different Fertilization Regimes in a Paddy Ecosystem
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Site and Design
2.2. Soil Sampling
2.3. Acquisition of Soil Properties
2.4. Analysis of Soil Bacterial Community
2.5. Statistical Analyses
3. Results
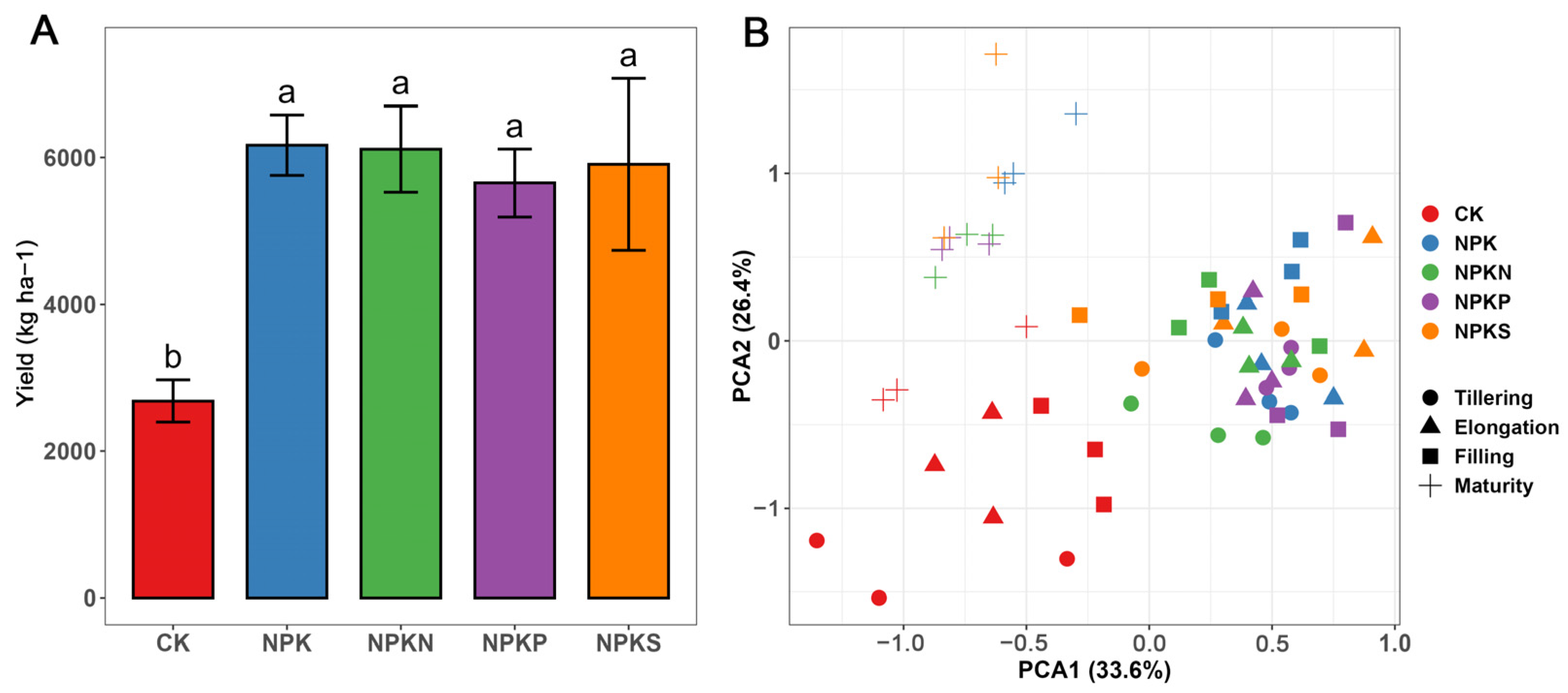
3.1. Rice Yield and Soil Physicochemical Properties
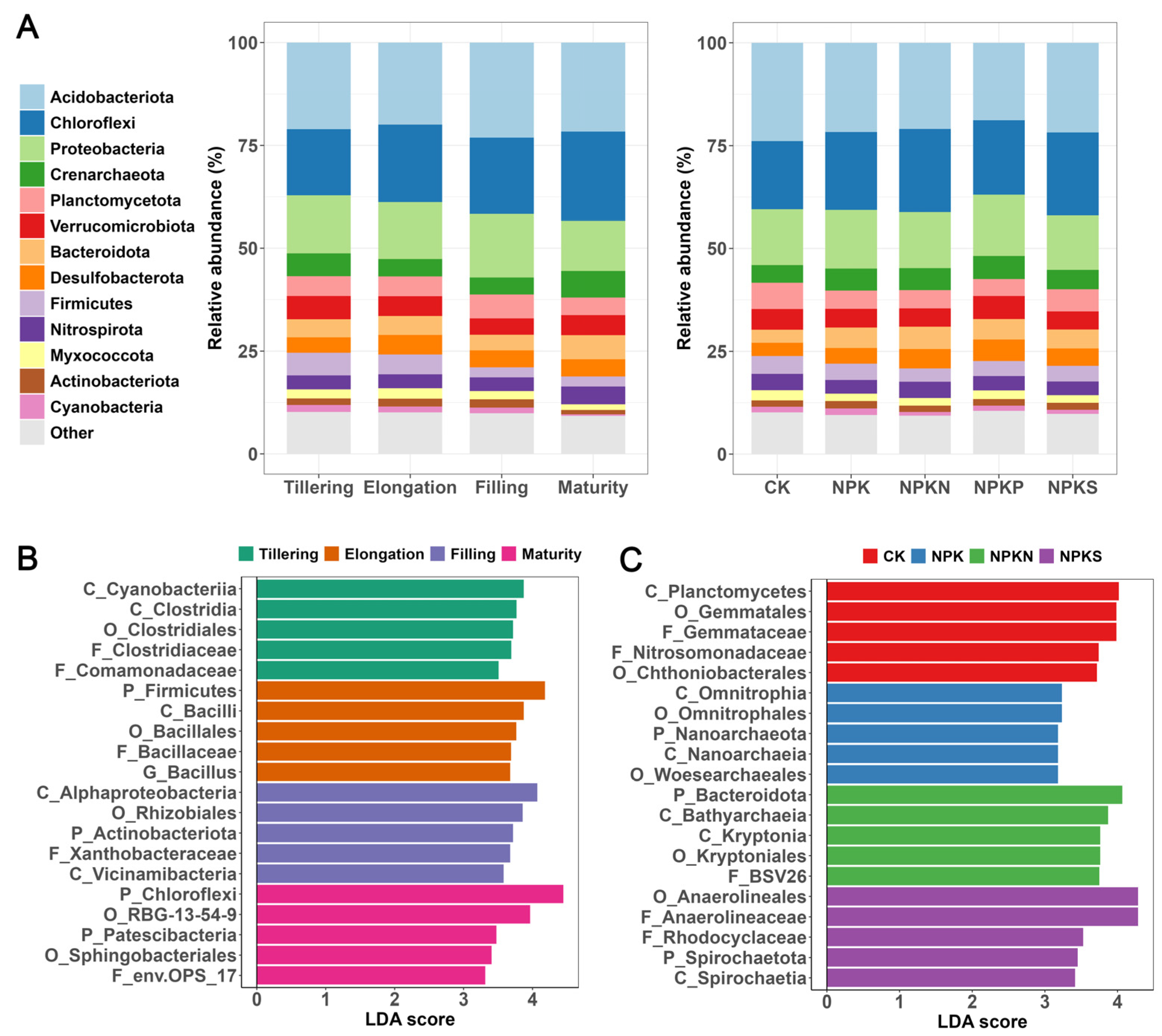
3.2. Soil Bacterial Community Composition
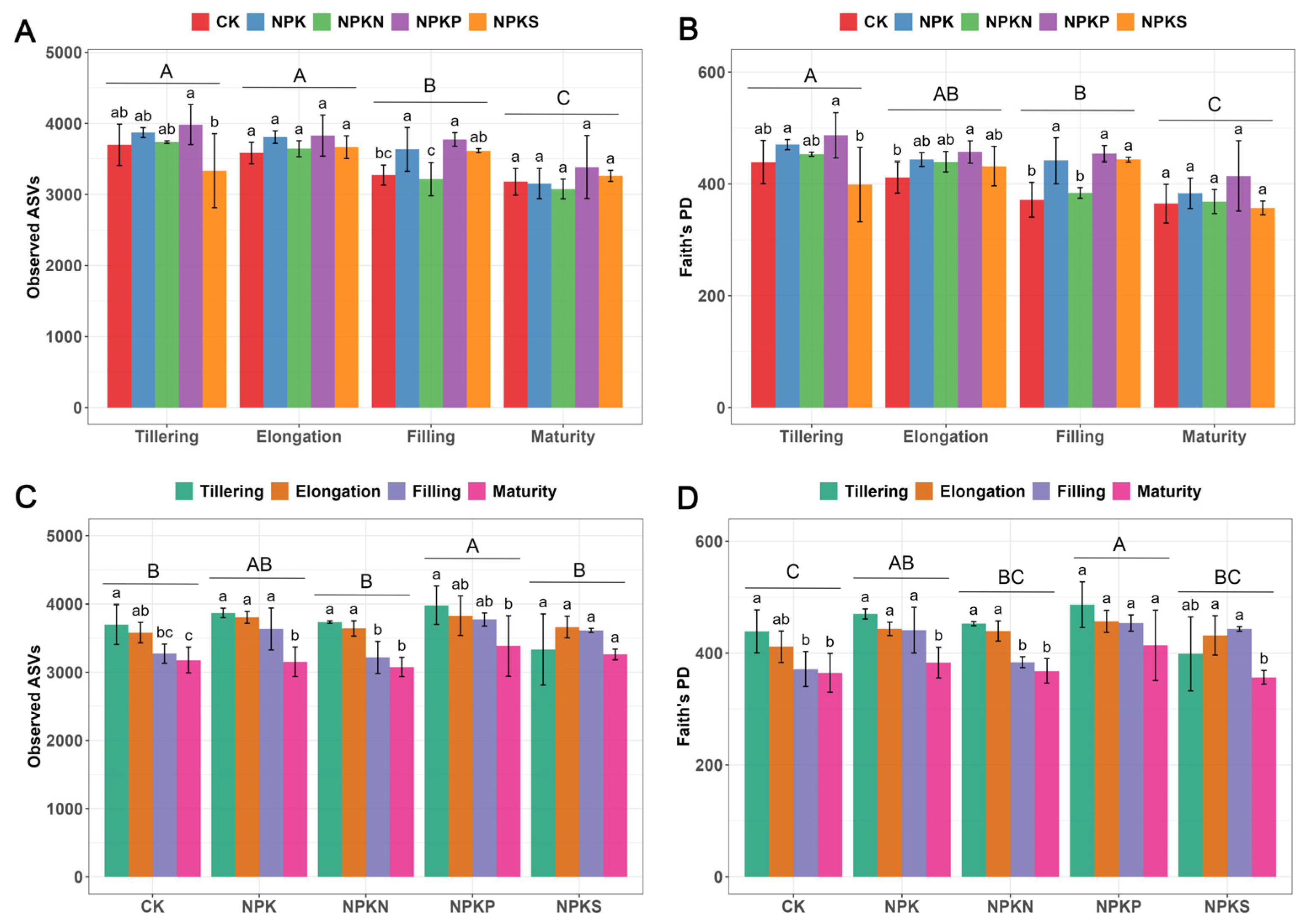
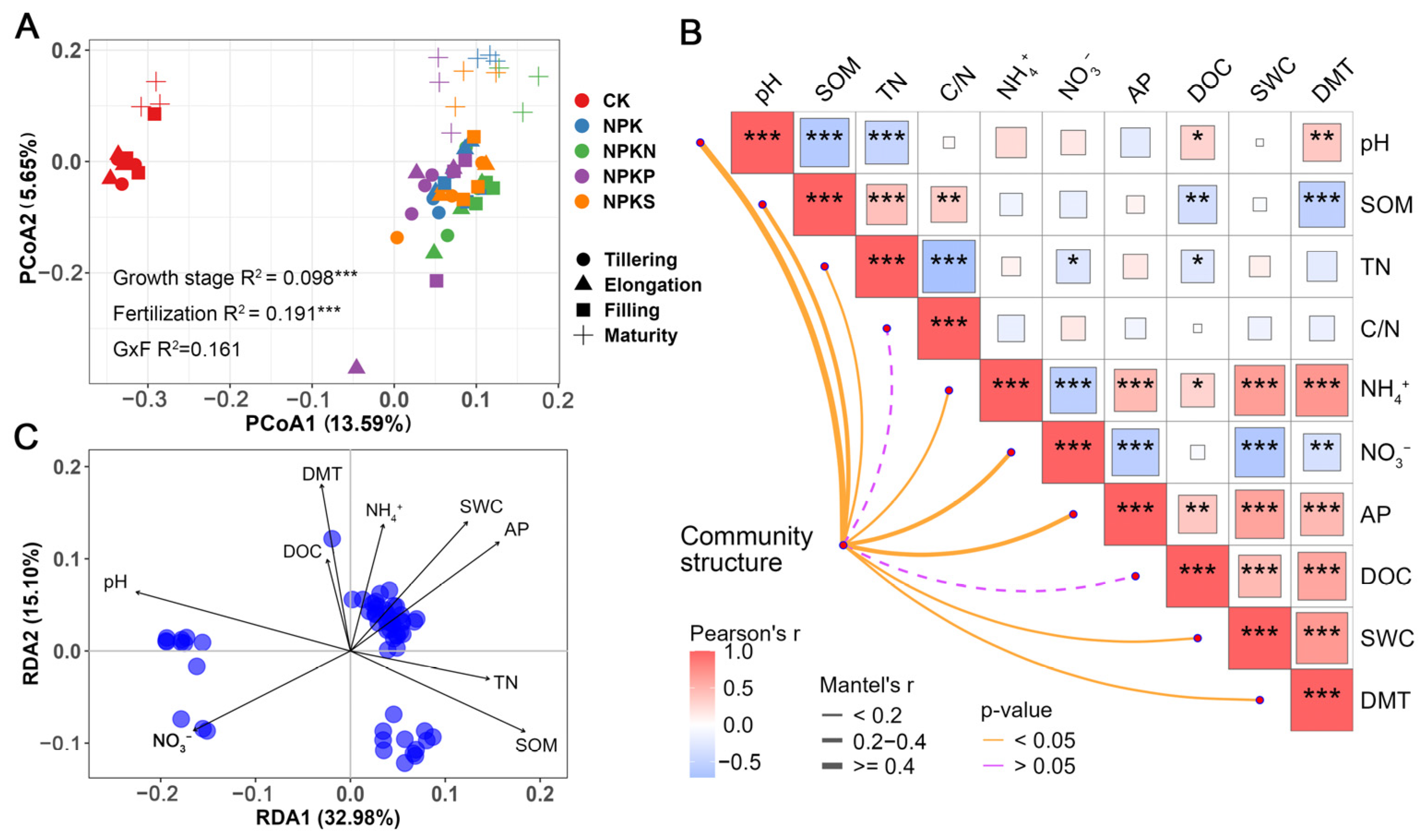
3.3. Soil Bacterial Diversity and Community Structure
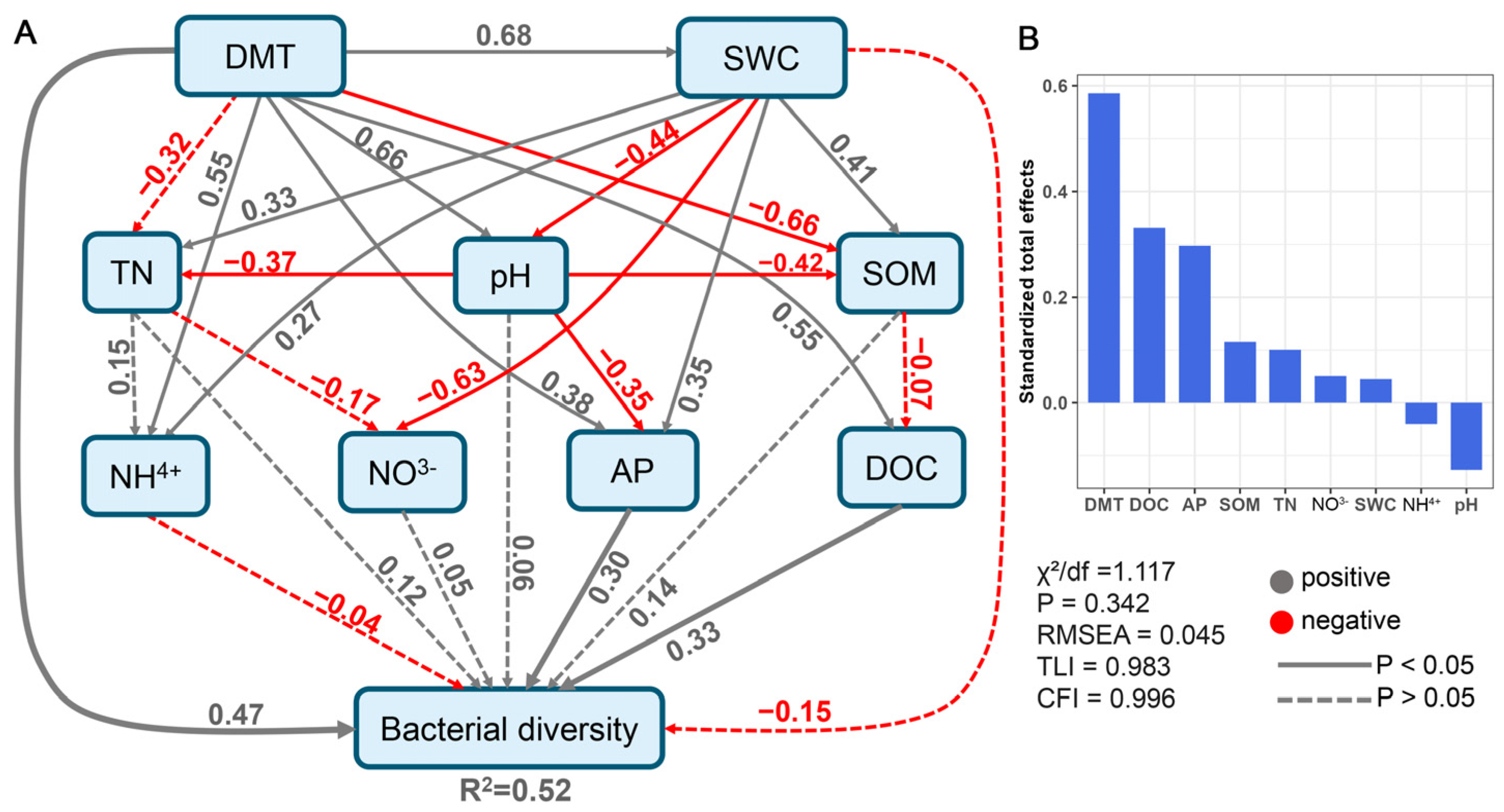
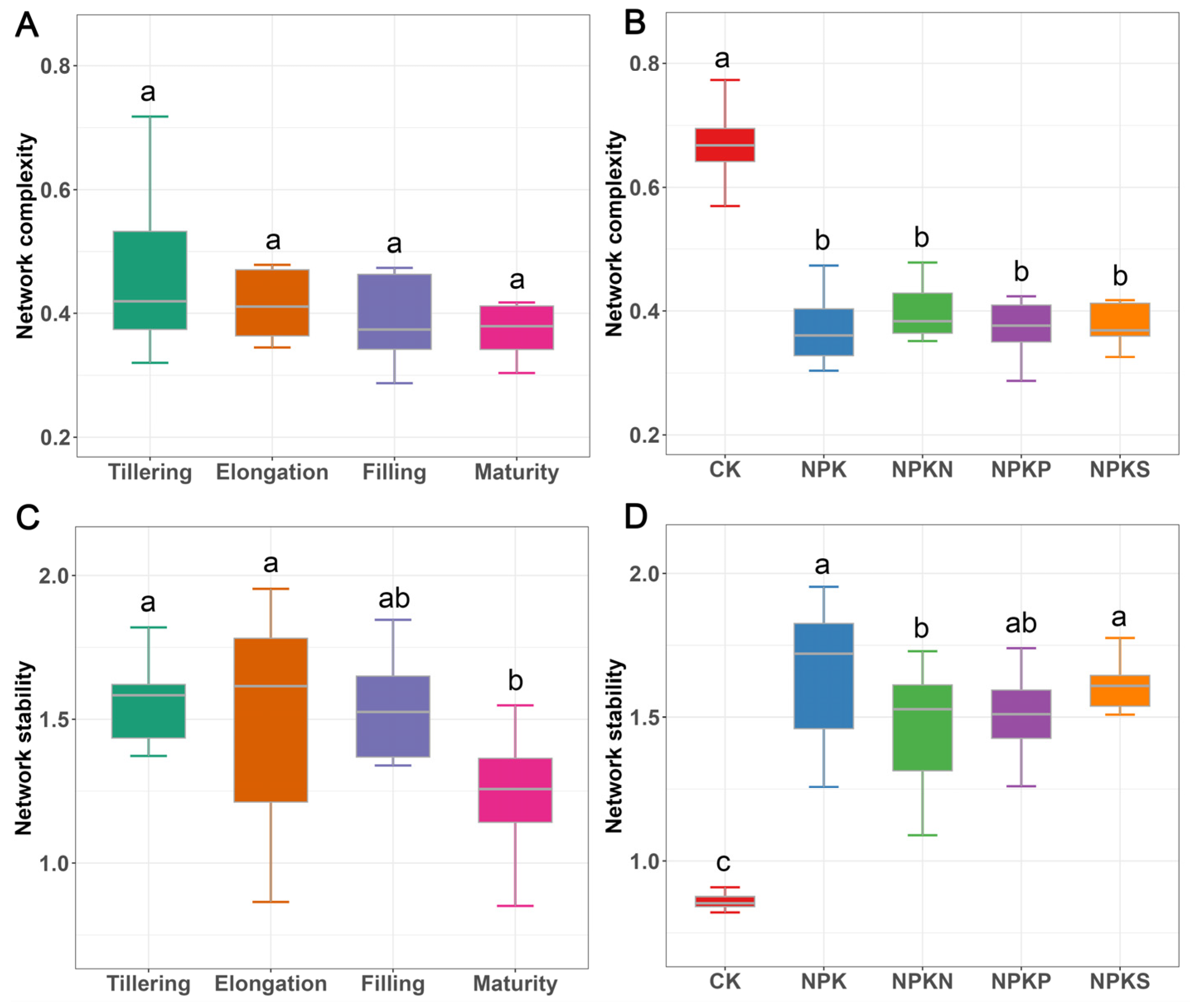
3.4. Soil Bacterial Network Complexity and Stability
4. Discussion
4.1. Rice Growth Stages Outweigh Fertilization for Soil Bacterial Diversity
4.2. Fertilization Has a Greater Effect on Soil Bacterial Community Structure
4.3. Fertilizer Addition Reduces the Complexity but Increases the Stability of Network
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| CK | No fertilization |
| NPK | Chemical fertilizer |
| NPKN | NPK supplemented with extra N |
| NPKP | NPK supplemented with extra P |
| NPKS | NPK supplemented with rice straw |
| DMT | Daily mean temperature |
| SWC | Soil water content |
| SOM | Soil organic matter |
| TN | Total nitrogen |
| AP | Available phosphorus |
| DOC | Dissolved organic carbon |
References
- Yuan, S.; Linquist, B.A.; Wilson, L.T.; Cassman, K.G.; Stuart, A.M.; Pede, V.; Miro, B.; Saito, K.; Agustiani, N.; Aristya, V.E.; et al. Sustainable intensification for a larger global rice bowl. Nat. Commun. 2021, 12, 7163. [Google Scholar] [CrossRef]
- Long, X.; Li, J.; Liao, X.; Wang, J.; Zhang, W.; Wang, K.; Zhao, J. Stable Soil Biota Network Enhances Soil Multifunctionality in Agroecosystems. Glob. Change Biol. 2025, 31, e70041. [Google Scholar] [CrossRef]
- Yan, X.; Xia, L.; Ti, C. Win-win nitrogen management practices for improving crop yield and environmental sustainability. Bull. Chin. Acad. Sci. 2018, 33, 177–183. [Google Scholar] [CrossRef]
- Yang, Q.; Zhang, M. Effect of bio-organic fertilizers partially substituting chemical fertilizers on labile organic carbon and bacterial community of citrus orchard soils. Plant Soil 2023, 483, 255–272. [Google Scholar] [CrossRef]
- Xiao, J.; Zhang, J.; Yuan, H.; Xie, X.; Gao, Y.; Lu, Y.; Liao, Y.; Nie, J. Long-term application of legume green manure improves rhizosphere soil bacterial stability and reduces bulk soil bacterial stability in rice. Eur. J. Soil Biol. 2024, 122, 103652. [Google Scholar] [CrossRef]
- Gu, J.; Chen, Y.; Zhang, H.; Li, Z.; Zhou, Q.; Yu, C.; Kong, X.; Liu, L.; Wang, Z.; Yang, J. Canopy light and nitrogen distributions are related to grain yield and nitrogen use efficiency in rice. Field Crops Res. 2017, 206, 74–85. [Google Scholar] [CrossRef]
- Bardgett, R.D.; van der Putten, W.H. Belowground biodiversity and ecosystem functioning. Nature 2014, 515, 505–511. [Google Scholar] [CrossRef]
- Li, Z.; Shen, Y.; Zhang, W.; Zhang, H.; Liu, L.; Wang, Z.; Gu, J.; Yang, J. Effects of long-term straw returning on rice yield and soil properties and bacterial community in a rice-wheat rotation system. Field Crops Res. 2023, 291, 108800. [Google Scholar] [CrossRef]
- Dai, Z.; Su, W.; Chen, H.; Barberán, A.; Zhao, H.; Yu, M.; Yu, L.; Brookes, P.C.; Schadt, C.W.; Chang, S.X.; et al. Long-term nitrogen fertilization decreases bacterial diversity and favors the growth of Actinobacteria and Proteobacteria in agro-ecosystems across the globe. Glob. Change Biol. 2018, 24, 3452–3461. [Google Scholar] [CrossRef]
- Shu, X.; He, J.; Zhou, Z.; Xia, L.; Hu, Y.; Zhang, Y.; Zhang, Y.; Luo, Y.; Chu, H.; Liu, W.; et al. Organic amendments enhance soil microbial diversity, microbial functionality and crop yields: A meta-analysis. Sci. Total Environ. 2022, 829, 154627. [Google Scholar] [CrossRef]
- Chen, L.; Zhou, S.; Zhang, Q.; Zou, M.; Yin, Q.; Qiu, Y.; Qin, W. Effect of organic material addition on active soil organic carbon and microbial diversity: A meta-analysis. Soil Tillage Res. 2024, 241, 106128. [Google Scholar] [CrossRef]
- Wang, W.; Luo, X.; Chen, Y.; Ye, X.; Wang, H.; Cao, Z.; Ran, W.; Cui, Z. Succession of Composition and Function of Soil Bacterial Communities During Key Rice Growth Stages. Front. Microbiol. 2019, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Qin, H.; Chen, Z.; Wu, J.; Wei, W. Effect of long-term fertilization on bacterial composition in rice paddy soil. Biol. Fertil. Soils 2011, 47, 397–405. [Google Scholar] [CrossRef]
- Yang, Y.; Shen, L.; Yanan, B.; Zhao, X.; Wang, S.; Liu, J.; Liu, X.; Tian, M.; Wangting, Y.; Jin, J.; et al. Response of potential activity, abundance and community composition of nitrite-dependent anaerobic methanotrophs to long-term fertilization in paddy soils. Environ. Microbiol. 2022, 24, 5005–5018. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, S.-L.; Chen, Q.-L.; Delgado-Baquerizo, M.; Guo, Z.-F.; Wang, F.; Xu, Y.-Y.; Zhu, Y.-G. Fertilization regulates global thresholds in soil bacteria. Glob. Change Biol. 2024, 30, e17466. [Google Scholar] [CrossRef]
- Adithya, S.; Nunna, S.A.D.; Chinnadurai, C.; Balachandar, D. Rhizosphere bacterial diversity and soil biological attributes of rice in different phenological stages and wetland cultivation methods. Pedosphere 2024, 35, 983–994. [Google Scholar] [CrossRef]
- Li, Q.; Liu, Y.; Su, N.; Tian, C.; Zhang, Y.; Tan, L.; Peng, J.; Rong, X.; Luo, G. Knowledge-based phosphorus input levels control the link between soil microbial diversity and ecosystem functions in paddy fields. Agric. Ecosyst. Environ. 2025, 379, 109352. [Google Scholar] [CrossRef]
- Degrune, F.; Theodorakopoulos, N.; Colinet, G.; Hiel, M.-P.; Bodson, B.; Taminiau, B.; Daube, G.; Vandenbol, M.; Hartmann, M. Temporal Dynamics of Soil Microbial Communities below the Seedbed under Two Contrasting Tillage Regimes. Front. Microbiol. 2017, 8, 1127. [Google Scholar] [CrossRef]
- Philippot, L.; Raaijmakers, J.M.; Lemanceau, P.; Van Der Putten, W.H. Going back to the roots: The microbial ecology of the rhizosphere. Nat. Rev. Microbiol. 2013, 11, 789–799. [Google Scholar] [CrossRef]
- Wang, J.; Xue, C.; Song, Y.; Wang, L.; Huang, Q.; Shen, Q. Wheat and Rice Growth Stages and Fertilization Regimes Alter Soil Bacterial Community Structure, But Not Diversity. Front. Microbiol. 2016, 7, 1207. [Google Scholar] [CrossRef]
- Jiao, S.; Yang, Y.; Xu, Y.; Zhang, J.; Lu, Y. Balance between community assembly processes mediates species coexistence in agricultural soil microbiomes across eastern China. ISME J. 2020, 14, 202–216. [Google Scholar] [CrossRef]
- Kuiper, J.J.; Van Altena, C.; De Ruiter, P.C.; Van Gerven, L.P.; Janse, J.H.; Mooij, W.M. Food-web stability signals critical transitions in temperate shallow lakes. Nat. Commun. 2015, 6, 7727. [Google Scholar] [CrossRef]
- Van Den Hoogen, J.; Geisen, S.; Routh, D.; Ferris, H.; Traunspurger, W.; Wardle, D.A.; De Goede, R.G.; Adams, B.J.; Ahmad, W.; Andriuzzi, W.S. Soil nematode abundance and functional group composition at a global scale. Nature 2019, 572, 194–198. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, Q.; Zhang, Z.; Li, W.; Liu, W.; Xiao, N.; Liu, H.; Wang, L.; Li, Z.; Ma, J.; et al. Decreased soil multifunctionality is associated with altered microbial network properties under precipitation reduction in a semiarid grassland. iMeta 2023, 2, e106. [Google Scholar] [CrossRef]
- Wu, M.-H.; Chen, S.-Y.; Chen, J.-W.; Xue, K.; Chen, S.-L.; Wang, X.-M.; Chen, T.; Kang, S.-C.; Rui, J.-P.; Thies, J.E.; et al. Reduced microbial stability in the active layer is associated with carbon loss under alpine permafrost degradation. Proc. Natl. Acad. Sci. USA 2021, 118, e2025321118. [Google Scholar] [CrossRef]
- Nelson, D.W.; Sommers, L.E. Total Carbon, Organic Carbon, and Organic Matter. In Methods of Soil Analysis; SSSA Book Series; American Society of Agronomy and Soil Science Society of America: Madison, WI, USA, 1996; pp. 961–1010. [Google Scholar]
- Zhang, J.; Zhu, T.; Meng, T.; Zhang, Y.; Yang, J.; Yang, W.; Müller, C.; Cai, Z. Agricultural land use affects nitrate production and conservation in humid subtropical soils in China. Soil Biol. Biochem. 2013, 62, 107–114. [Google Scholar] [CrossRef]
- Olsen, S.R.; Sommers, L.E. Phosphorus. In Methods of Soil Analysis; Agronomy Monographs; American Society of Agronomy and Soil Science Society of America: Madison, WI, USA, 1982; pp. 403–430. [Google Scholar]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Meth. 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Conway, J.; Lex, A.; Gehlenborg, N. UpSetR: An R Package For The Visualization Of Intersecting Sets And Their Properties. Bioinformatics 2017, 33, 2938–2940. [Google Scholar] [CrossRef] [PubMed]
- Xu, A.; Guo, Z.; Pan, K.; Wang, C.; Zhang, F.; Liu, J.; Pan, X.J. Increasing land-use durations enhance soil microbial deterministic processes and network complexity and stability in an ecotone. Appl. Soil Ecol. 2023, 181, 104630. [Google Scholar] [CrossRef]
- Luo, F.; Zhong, J.; Yang, Y.; Scheuermann, R.H.; Zhou, J. Application of random matrix theory to biological networks. Phys. Lett. A 2006, 357, 420–423. [Google Scholar] [CrossRef]
- Xun, W.; Liu, Y.; Li, W.; Ren, Y.; Xiong, W.; Xu, Z.; Zhang, N.; Miao, Y.; Shen, Q.; Zhang, R.J.M. Specialized metabolic functions of keystone taxa sustain soil microbiome stability. Microbiome 2021, 9, 35. [Google Scholar] [CrossRef]
- Li, H.; Yang, X.; Weng, B.; Su, J.; Nie, S.a.; Gilbert, J.A.; Zhu, Y.-G. The phenological stage of rice growth determines anaerobic ammonium oxidation activity in rhizosphere soil. Soil Biol. Biochem. 2016, 100, 59–65. [Google Scholar] [CrossRef]
- Huang, X.; Wang, J.; Dumack, K.; Anantharaman, K.; Ma, B.; He, Y.; Liu, W.; Di, H.; Li, Y.; Xu, J. Temperature-dependent trophic associations modulate soil bacterial communities along latitudinal gradients. ISME J. 2024, 18, wrae145. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Deng, Y.; Shen, L.; Wen, C.; Yan, Q.; Ning, D.; Qin, Y.; Xue, K.; Wu, L.; He, Z.; et al. Temperature mediates continental-scale diversity of microbes in forest soils. Nat. Commun. 2016, 7, 12083. [Google Scholar] [CrossRef] [PubMed]
- Wood, T.E.; Tucker, C.; Alonso-Rodríguez, A.M.; Loza, M.I.; Grullón-Penkova, I.F.; Cavaleri, M.A.; O’Connell, C.S.; Reed, S.C. Warming induces unexpectedly high soil respiration in a wet tropical forest. Nat. Commun. 2025, 16, 8222. [Google Scholar] [CrossRef]
- Zhang, X.; Johnston, E.R.; Barberán, A.; Ren, Y.; Lü, X.; Han, X. Decreased plant productivity resulting from plant group removal experiment constrains soil microbial functional diversity. Glob. Change Biol. 2017, 23, 4318–4332. [Google Scholar] [CrossRef]
- Li, H.; Wang, Z.; Feng, B.; Shi, J.; Liao, M.; He, K.; Tian, H.; Megharaj, M.; He, W. Arsenic stress on soil microbial nutrient metabolism interpreted by microbial utilization of dissolved organic carbon. J. Hazard. Mater. 2024, 470, 134232. [Google Scholar] [CrossRef]
- Guo, L.; Shi, J.; Lin, W.; Liang, J.; Lu, Z.; Tang, X.; Liu, Y.; Wu, P.; Li, C. Soil Bacteria Mediate Soil Organic Carbon Sequestration under Different Tillage and Straw Management in Rice-Wheat Cropping Systems. Agriculture 2022, 12, 1552. [Google Scholar] [CrossRef]
- Yang, Y.; Chen, X.; Liu, L.; Li, T.; Dou, Y.; Qiao, J.; Wang, Y.; An, S.; Chang, S. Nitrogen fertilization weakens the linkage between soil carbon and microbial diversity: A global meta-analysis. Glob. Change Biol. 2022, 28, 6446–6461. [Google Scholar] [CrossRef]
- Arunrat, N.; Sansupa, C.; Sereenonchai, S.; Hatano, R. Stability of soil bacteria in undisturbed soil and continuous maize cultivation in Northern Thailand. Front. Microbiol. 2023, 14, 1285445. [Google Scholar] [CrossRef] [PubMed]
- Philippot, L.; Chenu, C.; Kappler, A.; Rillig, M.C.; Fierer, N. The interplay between microbial communities and soil properties. Nat. Rev. Microbiol. 2024, 22, 226–239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Chen, W.; Dong, L.; Wang, W. Grassland degradation amplifies the negative effect of nitrogen enrichment on soil microbial community stability. Glob. Change Biol. 2024, 30, e17217. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Guan, Y. Consistent responses of soil bacterial communities to bioavailable silicon deficiency in croplands. Geoderma 2022, 408, 115587. [Google Scholar] [CrossRef]
- Liu, X.; Chen, M.; Li, Y.; Bai, L.; Guo, J. Modeling phosphorus dynamics in rice irrigation systems: Integrating O-Fe-P coupling and regional water cycling. J. Hydrol. 2025, 649, 132405. [Google Scholar] [CrossRef]
- Dash, B.; Nayak, S.; Pahari, A.; Nayak, S. Verrucomicrobia in soil: An agricultural perspective. In Frontiers in Soil and Environmental Microbiology; CRC Press: Boca Raton, FL, USA, 2020; pp. 37–46. [Google Scholar]
- Huang, R.; McGrath, S.P.; Hirsch, P.R.; Clark, I.M.; Storkey, J.; Wu, L.; Zhou, J.; Liang, Y. Plant–microbe networks in soil are weakened by century-long use of inorganic fertilizers. Microb. Biotechnol. 2019, 12, 1464–1475. [Google Scholar] [CrossRef]
- Yao, M.; Rui, J.; Li, J.; Dai, Y.; Bai, Y.; Heděnec, P.; Wang, J.; Zhang, S.; Pei, K.; Liu, C.; et al. Rate-specific responses of prokaryotic diversity and structure to nitrogen deposition in the Leymus chinensis steppe. Soil Biol. Biochem. 2014, 79, 81–90. [Google Scholar] [CrossRef]
- Potapov, A.M.; Drescher, J.; Darras, K.; Wenzel, A.; Janotta, N.; Nazarreta, R.; Kasmiatun; Laurent, V.; Mawan, A.; Utari, E.H. Rainforest transformation reallocates energy from green to brown food webs. Nature 2024, 627, 116–122. [Google Scholar] [CrossRef]
- Xie, G.; Sun, C.; Luo, W.; Gong, Y.; Tang, X. Distinct ecological niches and community dynamics: Understanding free-living and particle-attached bacterial communities in an oligotrophic deep lake. Appl. Environ. Microbiol. 2024, 90, e00714–e00724. [Google Scholar] [CrossRef]
- Yan, Y.; Zhou, X.; Liu, L.; Cai, Z.; Penuelas, J.; Huang, X. Soil Nutrient Enrichment Induces Trade-Offs in Bacterial Life-History Strategies Promoting Plant Productivity. Adv. Sci. 2025, e10066. [Google Scholar] [CrossRef]
- Chen, Y.; Li, Y.; Qiu, T.; He, H.; Liu, J.; Duan, C.; Cui, Y.; Huang, M.; Wu, C.; Fang, L. High nitrogen fertilizer input enhanced the microbial network complexity in the paddy soil. Soil Ecol. Lett. 2023, 6, 230205. [Google Scholar] [CrossRef]
- Yang, L.; Sun, R.; Li, J.; Zhai, L.; Cui, H.; Fan, B.; Wang, H.; Liu, H. Combined organic-inorganic fertilization builds higher stability of soil and root microbial networks than exclusive mineral or organic fertilization. Soil Ecol. Lett. 2022, 5, 220142. [Google Scholar] [CrossRef]
- Sun, S.; Qiao, Z.; Tikhonenkov, D.V.; Gong, Y.; Li, H.; Li, R.; Sun, K.; Huo, D. Temporal Dynamics and Adaptive Mechanisms of Microbial Communities: Divergent Responses and Network Interactions. Microb. Ecol. 2025, 88, 94. [Google Scholar] [CrossRef]
- Barka Essaid, A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Klenk, H.-P.; Clément, C.; Ouhdouch, Y.; van Wezel Gilles, P. Taxonomy, Physiology, and Natural Products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2015, 80, 1–43. [Google Scholar] [CrossRef]
- Gao, X.; Xiao, Y.; Wang, Z.; Zhao, H.; Yue, Y.; Nair, S.; Zhang, Z.; Zhang, Y. Adaptive traits of Planctomycetota bacteria to thrive in macroalgal habitats and establish mutually beneficial relationship with macroalgae. Limnol. Oceanogr. Lett. 2024, 9, 745–753. [Google Scholar] [CrossRef]

| Nitrogen (kg·ha−1) | Phosphorus (P2O5 kg·ha−1) | Potassium (K2O kg·ha−1) | Straw (kg·ha−1) | |
|---|---|---|---|---|
| CK | 0 | 0 | 0 | 0 |
| NPK | 166.05 | 29.85 | 72 | 0 |
| NPKN | 249.15 | 29.85 | 72 | 0 |
| NPKP | 166.05 | 44.85 | 72 | 0 |
| NPKS | 166.05 | 29.85 | 72 | 3600 |
| Growth Stage (G) | Fertilization Treatment (F) | G × F | ||||
|---|---|---|---|---|---|---|
| F | p | F | p | F | p | |
| pH | 14.581 | <0.001 | 33.834 | <0.001 | 2.911 | 0.006 |
| SOM | 11.673 | <0.001 | 10.616 | <0.001 | 0.194 | 0.998 |
| TN | 5.622 | 0.003 | 5.790 | 0.001 | 0.940 | 0.519 |
| C/N | 3.040 | 0.040 | 0.358 | 0.837 | 0.964 | 0.498 |
| NH4+ | 20.876 | <0.001 | 1.108 | 0.366 | 1.403 | 0.205 |
| NO3− | 19.254 | <0.001 | 17.998 | <0.001 | 3.142 | 0.003 |
| AP | 26.894 | <0.001 | 52.164 | <0.001 | 2.080 | 0.041 |
| DOC | 15.057 | <0.001 | 0.832 | 0.513 | 1.567 | 0.141 |
| SWC | 36.854 | <0.001 | 13.915 | <0.001 | 1.714 | 0.100 |
| Growth Stage (G) | Fertilization Treatment (F) | G × F | ||||
|---|---|---|---|---|---|---|
| F | p | F | p | F | p | |
| Observed ASVs | 15.818 | <0.001 | 4.356 | 0.005 | 1.244 | 0.289 |
| Faith’s PD | 14.771 | <0.001 | 6.169 | 0.001 | 1.303 | 0.255 |
| Growth Stage (G) | Fertilization Treatment (F) | G × F | ||||
|---|---|---|---|---|---|---|
| F | p | F | p | F | p | |
| Network complexity | 1.006 | 0.400 | 144.135 | <0.001 | 1.634 | 0.121 |
| Network stability | 10.781 | <0.001 | 54.846 | <0.001 | 1.885 | 0.067 |
| Network Complexity | Network Stability | |
|---|---|---|
| Acidobacteriota | 0.421 ** | −0.150 |
| Chloroflexi | −0.167 | 0.206 |
| Proteobacteria | −0.067 | −0.038 |
| Crenarchaeota | −0.106 | 0.125 |
| Planctomycetota | 0.376 ** | −0.288 * |
| Verrucomicrobiota | 0.034 | −0.117 |
| Bacteroidota | −0.449 ** | 0.2137 |
| Desulfobacterota | −0.331 ** | 0.046 |
| Firmicutes | −0.178 | 0.092 |
| Nitrospirota | 0.175 | −0.227 |
| Myxococcota | 0.127 | −0.138 |
| Actinobacteriota | −0.163 | 0.382 ** |
| Cyanobacteria | −0.059 | −0.143 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, A.; Zheng, X.; Zhang, Y.; Chen, Q.; Wang, H. Dynamics of Soil Bacterial Communities over Rice Growth Stages Under Different Fertilization Regimes in a Paddy Ecosystem. Agriculture 2025, 15, 2466. https://doi.org/10.3390/agriculture15232466
Xu A, Zheng X, Zhang Y, Chen Q, Wang H. Dynamics of Soil Bacterial Communities over Rice Growth Stages Under Different Fertilization Regimes in a Paddy Ecosystem. Agriculture. 2025; 15(23):2466. https://doi.org/10.3390/agriculture15232466
Chicago/Turabian StyleXu, Aiai, Xiangzhou Zheng, Yushu Zhang, Qianqian Chen, and Huangping Wang. 2025. "Dynamics of Soil Bacterial Communities over Rice Growth Stages Under Different Fertilization Regimes in a Paddy Ecosystem" Agriculture 15, no. 23: 2466. https://doi.org/10.3390/agriculture15232466
APA StyleXu, A., Zheng, X., Zhang, Y., Chen, Q., & Wang, H. (2025). Dynamics of Soil Bacterial Communities over Rice Growth Stages Under Different Fertilization Regimes in a Paddy Ecosystem. Agriculture, 15(23), 2466. https://doi.org/10.3390/agriculture15232466





