Abstract
The loss and/or dysregulation of several cellular and mitochondrial antioxidants’ expression or enzymatic activity, which leads to the aberrant physiological function of these proteins, has been shown to result in oxidative damage to cellular macromolecules. In this regard, it has been surmised that the disruption of mitochondrial networks responsible for maintaining normal metabolism is an established hallmark of cancer and a novel mechanism of therapy resistance. This altered metabolism leads to aberrant accumulation of reactive oxygen species (ROS), which, under specific physiological conditions, leads to a potential tumor-permissive cellular environment. In this regard, it is becoming increasingly clear that the loss or disruption of mitochondrial oxidant scavenging enzymes may be, in specific tumors, either an early event in transformation or exhibit tumor-promoting properties. One example of such an antioxidant enzyme is manganese superoxide dismutase (MnSOD, also referred to as SOD2), which detoxifies superoxide, a ROS that has been shown, when its normal physiological levels are disrupted, to lead to oncogenicity and therapy resistance. Here, we will also discuss how the acetylation of MnSOD leads to a change in detoxification function that leads to a cellular environment permissive for the development of lineage plasticity-like properties that may be one mechanism leading to tumorigenic and therapy-resistant phenotypes.
Keywords:
mitochondrial; acetylation; MnSOD; ROS; metabolism; carcinogenesis; redox; sirtuins; electron transport; metal iron metabolism 1. Introduction
A fundamental theme in precision, or personalized, cancer medicine is identifying tumors that are vulnerable to specific therapies, based on well-defined molecular biomarkers or tumor signatures [1,2,3,4]. In this regard, estrogen-receptor-positive (ER+) invasive ductal cancers (IDCs), the most common type of breast cancer, are commonly treated with selective estrogen receptor modulators (SERMs), which have been shown in multiple studies to improve clinical outcomes [5,6]. ER+ IDCs are classified as either luminal A or luminal B cancer (LuBCa). LuBCas, which account for most breast cancer deaths in America, exhibit aggressive tumor characteristics, including an elevated proliferative index (high Ki-67); are poorly differentiated (high grade); and display an increased risk of recurrence and metastasis [7,8]. The lethality in women with LuBCa is due, at least in part, to the development of resistance against SERMs and a lack of alternative systemic therapies [9].
The estrogen receptor pathway has a critical role in breast cancer and, therefore, endocrine therapy, which blocks ER signaling, is highly effective; however, over time, a small subset of ER+ tumors recur due to the development of endocrine resistance [7,8]. Multiple mechanisms of endocrine resistance have been identified, including deregulation of various components of the ER signaling pathway, altered cell cycle and cell survival processes, and the activation of escape pathways that provide tumors with alternative proliferative and survival stimuli [5,10]. While most resistance to SERMs involves one of these processes, an increasingly accepted mechanism involves the development of lineage-plasticity-like properties. In this regard, a recent NCI white paper (Beltran, 2019) [11] and a seminal review (Yuan, 2019, Cancer Discovery) [12] stated lineage plasticity, due to disruptive microenvironmental cues, stochastic genetics/epigenetics, metabolic alterations, or other therapy-imposed selective pressures, contributes to tumor heterogeneity and, importantly, to the development of resistant phenotypes.
We define lineage plasticity as a reversible or irreversible reprogramming of cellular systems, where a mature somatic cell can display plasticity, via a change in cell “identity” by dedifferentiation to a progenitor-like state or by transdifferentiation to an alternative differentiated cell type, leading to the emergence of new phenotypes [13,14,15,16,17]. Of late, it has been proposed that the disruption of mitochondrial physiology is a novel mechanism, which, under specific cellular conditions, leads to lineage plasticity and may be one way that tumor cells establish resistant phenotypes to therapeutic interventions [18,19]. Therefore, we ask, does lineage plasticity, due to altered mitochondrial detoxification enzymes such as MnSOD, also lead to a therapy resistance phenotype in a subgroup of ER+ LuBCa IDCs? Furthermore, under specific circumstances when MnSOD is dysregulated or deleted, does it contribute to aberrant cellular metabolism and/or mitochondrial bioenergetics, oncogenesis, and/or therapy resistance?
2. Post-Translational Deacetylation of MnSOD by SIRT3
Sirtuins are NAD+ dependent class III histone deacetylases that are found in various genomes from bacteria to humans [20,21,22], which deacetylate a wide range of downstream non-histone targets, including transcription factors, metabolic enzymes, and detoxification enzymes. In addition, several sirtuins have also been shown to deacetylate histones under specific conditions, including SIRT1, SIRT2, and SIRT6 [23,24,25,26]. Sirtuin proteins play important roles in metabolic and bioenergetic processes, stress resistance, cell survival, and aging [27,28]. Sirtuin genes are human and murine homologs of the Saccharomyces cerevisiae Sir2 that have been shown to regulate life span and aging in model organisms, including Saccharomyces cerevisiae, C. elegans, and Drosophila melanogaster [23,29,30,31]. The sirtuin family, which constitutes the main mammalian lysine acetyl transferases, consists of seven NAD+-dependent enzymes. These seven proteins are localized to the nucleus (Sirt1, 6, and 7), mitochondria (Sirt3, 4, and 5), and cytoplasm (Sirt2) and share a 275-amino-acid catalytic deacetylase domain [27].
Sirtuins, including the primary mitochondrial sirtuin, Sirt3 [21,22], appear to respond to changes in cellular and nutrient conditions, including mitochondrial stress, resulting in the activation of their deacetylase or ribosyltransferase activity, and consequently, in a post-translational modification (PTM) of downstream target proteins acetylation (Ac) levels [20,32,33,34,35]. Lysine acetylation neutralizes the positive charge on the lysine residue and can cause structural and functional changes in the protein due to changes in electrostatic potential. We [36], and others [34,37,38], have shown that MnSOD is an important downstream target of SIRT3 in its role and as a mitochondrial fidelity protein [21,33,39,40]. A key aspect of sirtuins is that they require NAD+ as a co-factor, which makes them metabolic sensors and connects their enzymatic activity to the energy and redox state of cells [41,42,43]. Most mitochondrial proteins are autoacetylated due to a high acetyl-CoA concentration in mitochondria, and protein functions modified by lysine acetylation are mainly regulated by the activity of deacetylation enzymes such as SIRT3 [36,44,45,46].
Thus, an important theme that has emerged in the last several years is that SIRT3, and its most important downstream targets, direct multiple mitochondrial processes. In this regard, it appears that the non-physiological levels of MnSOD acetylation (Ac), at least in part, connect metabolic and bioenergetic balance and tumor cell growth and survival. This is based on a recently identified novel mitochondrial signaling axis centered on MnSOD-Ac, which, when dysregulated, disrupts cell metabolism, leading to aberrant ROS levels [47,48]. MnSOD is a mitochondrial matrix-localized homotetrameric antioxidant enzyme with four identical subunits, each harboring a Mn2+ atom [49,50]. The primary function of MnSOD is to scavenge superoxide generated from metabolic processes, including the electron transport chain. Mammalian MnSOD appears to have four lysines that have been identified as potential SIRT3 downstream targets, including K53, K89 [34,51], K68 [38,52,53], and K122 [34,35,37,38], using different methods, including site directed acetylation mutants, physical lysine acetylation followed by mass spectrometry analyses, and acetyl-lysine specific monoclonal antibodies. However, the biochemical and physiological significance of each of these lysines, and the molecular mechanism directing MnSOD enzymatic activity and mitochondrial metabolism, remains to be fully determined. The SIRT3-MnSOD-Ac axis is an active area of research in the regulation of mammalian and human cells/organs and its dysregulation appears in several human illnesses, including cancer. It now appears quite clear that the SIRT3-MnSOD-Ac axis is a mitochondrial signaling hub that regulates how cells adapt to ROS-induced metabolic stress in addition to reprograming mitochondrial metabolism, which may play an important role in late-onset diseases [44,54,55,56,57]. However, there are limited data to demonstrate the mechanism behind this idea [36,45].
Recently, we have been focusing on MnSOD-K68-Ac as one of the key SIRT3 targets that may play an important role in the oncogenicity observed in cells lacking SIRT3. In this regard, mice lacking Sirt3, and thus containing MnSOD-Ac, develop estrogen-receptor-positive (ER+), poorly differentiated, high Ki-67 mammary gland tumors that exhibit very similar characteristics to human luminal B breast malignancies [33,36,44,56]. As compared to luminal A ER+ breast cancers, luminal B subtypes tend to have increased proliferation markers and, most importantly, can exhibit a tumor cell endocrine-resistant phenotype [45]. In this regard, there are nearly thirty publications that suggest a role for the SIRT3-MnSOD-Ac signaling axis in the dysregulation of mitochondrial function and dysregulation. Interestingly, mice that have a monoallelic knockout for MnSOD (MnSOD+/−) exhibit decreased MnSOD activity, increased oxidative stress, decreased life span, as well as aging-related phenotypes, especially carcinogenesis, which includes breast malignancies [58].
3. Lysine 68 Acetylation-Dependent Regulation of MnSOD Enzymatic Activity
The mitochondria electron transport chain (ETC) directs the flow of electrons in normal cellular metabolism, resulting in the production of superoxide radicals at less than 1% of the total rate of electron transport from NADH to oxygen [59,60,61]. A high level of superoxide can damage many components of cells, such as proteins, DNA, RNA, and lipids [62,63,64] potentially leading to cell death or dysfunction which can result in human illness permissive phenotypes. In addition, maintaining the low steady-state level of superoxide is critical for proper regulation of mitochondria function [65]. Therefore, the peroxide-removing system in mitochondria is required for respiration and to maintain metabolic homeostasis [62]. MnSOD is one of the critical mitochondrial detoxification enzymes neutralizing the toxic superoxide to less harmful and membrane-permeable hydrogen peroxide, which is converted to oxygen and water by peroxidases, such as catalase, in the cytoplasm. Two molecules of O2•− are oxidized to oxygen O2 and H2O2 through the catalytic reduction of Mn3+ to Mn2+ and restoration of Mn2+, respectively [61,66,67]. Due to the inducible oxidative stress of O2•− produced during energy production in mitochondria, MnSOD detoxification/ROS scavenging enzymic activity is critical to ATP production, maintaining mitochondrial homeostasis, and cell survival.
In order to regulate the activity of proteins, cellular signaling networks employ different mechanisms, including post-translational modifications (PTMs), such as acetylation [68,69,70,71]. Lysine acetylation is an extensively used PTM to regulate transcription and signaling factors, as well as histones regulated by specific sirtuins [38,72]. Non-acetylated lysine moieties are positively charged [66], while acetylation of these lysine residues, through a PTM, neutralizes this charge and alters the enzymatic function [67,73]. In general terms, the acetylome is roughly 60% that of the kinome and studies have identified the presence of more than 2000 acetylation substrates localized in both the cytosol and mitochondria, indicating the critical role of acetylation in cellular regulation [38,74,75,76,77]. In this regard, acetylation is one of the most frequent PTMs in the mitochondria.
We recently published data, using site-directed MnSOD mutants, where K68 was substituted with either a glutamine (Q) [78,79], an acetyl-lysine mimic mutant, or an arginine (R), a deacetylated lysine mimic mutant [80]. MnSOD−/− MEFs infected with lenti-MnSODK68Q decreased activity, while infection with lenti-MnSODK68R showed a significant increase in MnSOD activity, as compared to MEFs infected with wild-type (WT) lenti-MnSODWT [34,35,38]. In addition to these results, further experiments with our anti-MnSOD-K68-Ac antibody showed that (1) caloric restriction, (2) time-restricted fasting (36 h) [35], and (3) forced exercise [80] deacetylated K68, identifying K68-Ac as a physiologically relevant post-translational modification. These results led us to explore both the mechanism underlying how SIRT3 regulates acetylation of lysine-68 and directs MnSOD activity, as well as its role in oncogenicity and tumor cell resistance. Using the known crystal structure of MnSOD (PDB: 2adp), we conducted computational structural biology studies to examine both MnSODK68Q acetylation mimic mutant, as well as MnSOD-K68-Ac, to determine any differences in charge distribution, compared to the MnSOD wild-type.
We have previously published that MnSODK68Q and MnSOD-K68-Ac have nearly identical physiological and biochemical properties using tissue culture, molecular biology, and codon substitution experiments in bacteria [52]. In addition, we have also used the molecular dynamics core to calculate the surface charge density for MnSODWT and MnSOD-K68-Ac, which clearly showed that K68-Ac increases the negative surface charge distribution along the α1/α2 helices adjacent to the Mn2+ binding site/tetramerization interface of MnSOD-K68-Ac [52]. Importantly, an identical change in surface charge was also observed for MnSOD-K68-Ac and MnSOD-K68A by others, including Lu et al., 2017 (see Figure 2) [81] and Lu et al., 2015 (see Figure 7 [49]. The tetrameric interface is a four-helix bundle, symmetrically composed of two sets of helices (α1 and α2) separated by a tight turn. These calculations showed that these alpha helices, α1 and α2, which already have comparatively low surface charge, see a further decrease in surface charge upon acetylation of K68. Given that these helices make up the majority of the tetramer interface, the decreased surface charge may lead to destabilization of the quaternary structure. Thus, these results indicate that the MnSOD-K68Q mutant mimics the molecular dynamics of K68-Ac and suggest how acetylation of K68 might disrupt MnSOD detoxification activity by disrupting the organization of the homotetrameric complex. Further computational studies focusing specifically on the stability of the tetrameric interface of the MnSOD complex with respect to the acetylation status of K68 are required to conclusively demonstrate this proposed mechanism. The immediate result of the breakdown of the tetrameric structure is likely the formation of two dimeric species. However, there is evidence to suggest that the dimer is less stable and the equilibrium between the dimeric and monomeric species for eukaryotic MnSOD is rapid [82]. For consistency, we will refer to the degraded tetramer as “monomer”, even though it could exist in either monomeric or dimeric form.
Experiments with the MnSOD-K68-Ac mimic mutant, as well as the deacetylation mimic mutant, have indicated that MnSOD-K68Q loosens the homotetramer and dimeric/monomeric forms are favored. Based on these results it was surmised that MnSOD-K68-Ac may have a different function than the detoxification SOD activity observed for the homotetrameric complex. In this regard, the Flag-tagged MnSOD constructs (MnSODWT, MnSODK68R, and MnSODK68Q) were immunoprecipitated (IPed) and tested for enzymatic activity. These experiments showed that MnSOD from MEFs lacking MnSOD (MnSOD−/−) expressing MnSODK68Q exhibited a 40-fold increase in peroxidase activity, compared to IPed MnSOD-K68R or MnSOD-WT [52]. Identical results were achieved in codon substitution experiments using MnSOD-WT and MnSOD-K68-Ac recombinant proteins. Size exclusion chromatography was used to separate the tetramer (96 kDa) and monomer forms (24 kDa) and subsequent immunoblotting showed that the tetramer was enriched in MnSOD-WT, while MnSOD-K68-Ac is mostly in a monomeric form. The separate fractions were then analyzed for dismutase and peroxidase activity and overwhelmingly the tetramer showed dismutase activity, while the monomer showed peroxidase activity. Based on these results, it appears that MnSOD exhibits a dichotomous function, based on its lysine 68 (K68) acetylation status, where the deacetylated homotetrameric form acts as a protective detoxification enzyme against persistent/aberrant ROS. In contrast, K68-Ac inhibits homotetramer formation and, therefore, shifts the MnSOD equilibrium towards a predominantly monomeric form that functions as a peroxidase. As such, the working hypothesis is that K68-Ac not only inhibits its detoxification activity, but also results in a switch in enzymatic function [52,53].
4. MnSODK68Q Is a Monomer That Exhibits Oncogenic Properties
We previously showed that mice lacking Sirt3 develop ER+ luminal B-like tumor properties, suggesting that MnSOD, perhaps when acetylated, may function as a tumor promoter, instead of its more traditional function as a detoxification enzyme and tumor suppressor (TS) [83,84]. For initial experiments, an in vitro model was used, in which at least two oncogenes, i.e., c-Myc or Ras [85], are required to immortalize and/or transform primary rodent cells. MnSOD−/− primary mouse embryo fibroblasts (pMEFs) infected with lenti-MnSODK68Q, and c-Myc or Ras, became immortalized (i.e., divided beyond 15 cell passages) but not pMEFs infected with c-Myc or Ras alone. In contrast, co-infection of lenti-MnSODK68R with c-Myc or Ras failed to immortalize pMEFs [35,52]. As a positive control, pMEFs infected with both c-Myc and Ras were immortalized, as has been previously shown [85,86]. Interestingly, infection with lenti-MnSODK68R prevented immortalization of pMEFs with c-Myc and Ras, implying that MnSODK68R is a tumor suppressor. Lastly, pMEFs infected with lenti-MnSODK68Q, as compared to cells infected with MnSODK68R, exhibited increased: (1) soft agar growth, a measure of anchorage independence; (2) growth when plated at low density (250 cells/60 mm plate), a measure of proliferative capacity; (3) doubling time, a measure of proliferative rate; and (4) xenograft growth, a measure of tumorigenicity [52,53].
These established in vitro tissue culture transformation experiments support the premise that MnSOD-K68-Ac can function, under specific conditions, as a tumor promoter [52,87]. Thus, these data strongly imply that MnSODK68Q, which is proposed to model MnSOD-K68-Ac, can function as an in vitro oncogene as opposed to its more established role as a detoxification enzyme that can, under specific conditions, function as a tumor suppressor. These results may explain why initial clinical data suggest that MnSOD acts as a tumor suppressor [88]; however, surprisingly, there are also relatively new convincing data that also show that high MnSOD levels positively correlate with more aggressive tumors [89,90]. These studies, and our data, suggest a dichotomous role for MnSOD, where the tetramer acts as a TS during the early, proliferative stage of tumor initiation. However, once oncogenesis progresses, monomeric K68-acetylated MnSOD establishes a tumorigenic phenotype.
Thus, MnSOD may switch from a tetrameric dismutase to a monomeric peroxidase in response to specific cellular stress conditions, such as nutrient status and/or transformation (Figure 1). In this model, we posit that genetic or metabolic abnormalities dysregulate MnSOD-Ac and shift the balance towards higher monomer levels with decreased tetramer. The MnSOD-Ac, while accommodating the metabolic needs of the cell, may also function as a tumor promoter under specific conditions. Lastly, it is surmised that the SIRT3-MnSOD signaling axis may also be a potential therapeutic molecular target, including agents such as GC4419 (Galera Therapeutics, Inc. Malvern, PA, USA), which chemically dismutase/detoxify superoxide instead of enzymatic removal by MnSOD [87].
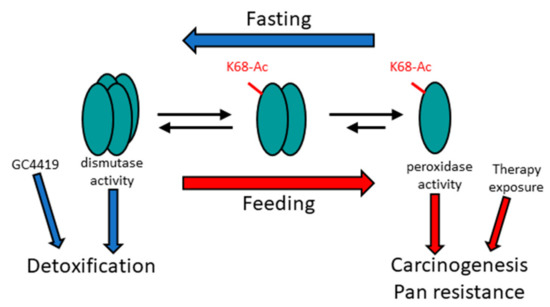
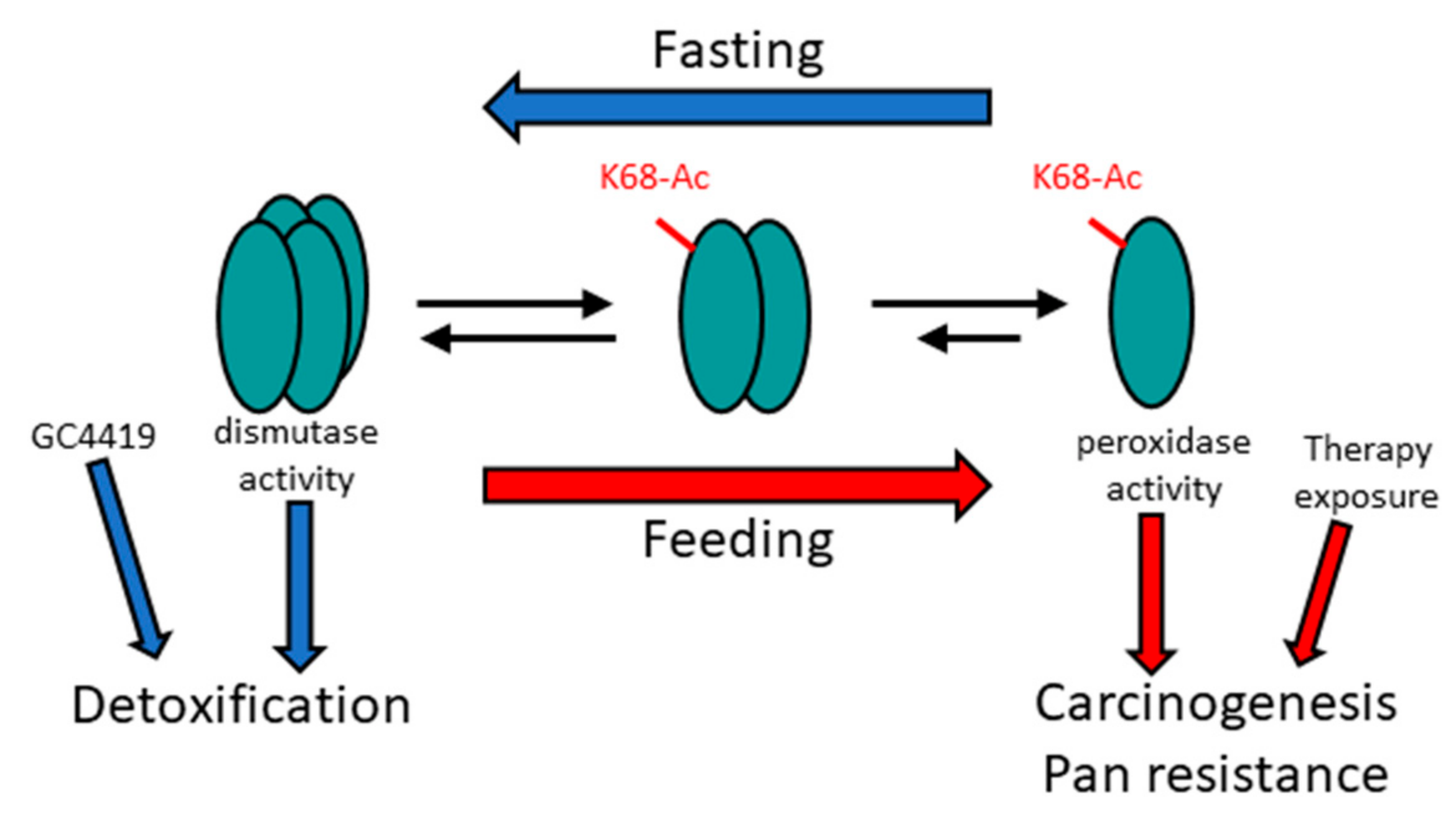
Figure 1.
Proposed mechanism of MnSOD detoxification and peroxidase roles. The tetrameric detoxification complex plays a role in a fasting physiology (blue arrows), whereas increased monomeric MnSOD, and its peroxidase activity, may be required for mitochondrial reprogramming to generate energy in a feeding state (red arrows). Other genetic and/or other types of cell stress may also create an oncogenic phenotype, which reprograms the mitochondria by increasing MnSOD-K68-Ac.
Considering the important role of MnSOD in detoxification, it is worthwhile to mention that the role of MnSOD goes beyond this activity. Van Remmen et al. revealed that MnSOD+/− heterozygous mice had a 50% reduction in SOD activity and increased 8-oxo-2-deoxyguanosine (8oxodG) in nuclear DNA [54,55,58], suggesting the enhancement in oxidative stress in monoallelic MnSOD+/− tissues. Tumor formation was observed in all MnSOD+/− mice with increased oxidative DNA damage, which indicates the correlation of MnSOD activity and carcinogenesis [59,91]. Interestingly, Sirt3−/− MEFs also showed an elevation of oxidative stress induction, which is associated with abnormal physiology changes in mitochondria. Additionally, ER/PR-positive mammary tumor development was observed in mice lacking Sirt3 (34). Other studies also showed an increase in Ki-67 in knockout Sirt3 female mice ER+ tumors, a marker for human luminal B breast malignancies [33,35,45]. As previously mentioned, we proposed the equilibrium shift model from MnSOD tetrameric form to monomeric form when K68 is acetylated [52,87]. Further results revealed that MnSOD-K68-Ac functions as a tumor promoter in cells expressing MnSODK68Q, the MnSOD acetylation mimic mutant. Together, these results show a correlation of Sirt3, MnSOD acetylation status, and ROS detoxification activity, as well as mitochondrial metabolic stress with carcinogenesis, implicating Sirt3 as a tumor suppressor.
5. Redox Signaling Involving MnSOD
As discussed above, it is proposed that there may be a mitochondrial equilibrium shift in the MnSOD tetrameric/monomeric ratio when K68 is acetylated. MnSOD catalyzes the dismutation of superoxide (O2•−) into hydrogen peroxide (H2O2); however, it has been shown to demonstrate peroxidase activity under certain conditions [52,92,93]. MnSOD is highly regulated at the transcription level by NF-kB, SP1, AP1, AP2, cytokines, and protein kinase C and is strongly correlated with cell cycle [50,94]. MnSOD is also post-translationally regulated by acetylation and phosphorylation [62,94]. Given the reversibility of the active site, tight regulation of expression and activity, and the added fact that the electron transport chain is riddled with antioxidants and only a small production of O2•−, it has been suggested that MnSOD’s second important function is as a signaling hub, as well as a superoxide dismutase [95]. This is supported by peroxidase activity [52] and the specific and selective nature in which O2•− and H2O2 oxidize targets [96,97]. H2O2 has a long half-life in aqueous solutions and can easily permeate mitochondria membranes; however, due to varied reduction potentials of amino acids, it has been shown that H2O2 selectively binds and oxidizes the active site of several phosphatases, rendering them inactive [96].
Increased MnSOD activity also leads to increased H2O2 signaling, inactivation of phosphatases under specific cellular and mitochondrial conditions, and an increase in phosphorylation activity of JNK/c-Jun and AMPK [96]. Overexpression of MnSOD in Jurkat cell lines led to increased phosphorylation of JNK and downstream c-Jun [96]. This pathway upregulates the expression of inflammatory cytokines and acts as a positive feedback loop for MnSOD as c-Jun is an AP-1 subunit that upregulates expression of MnSOD. JNK has been shown to prevent tumor initiation [98] and, through the activation of this pathway, MnSOD acts as a tumor suppressor. In addition, overexpression of MnSOD, confirmed with exogenous addition of H2O2, led to oxidation of CaMKII-M281/282, consequently activating CaMKII [90]. This leads to phosphorylation and activation of AMPK, subsequent phosphorylation, and inactivation of acetyl-CoA carboxylase (ACC), and an increase of glycolysis [90,99]. The switch to glycolysis, i.e. the Warburg effect, is a hallmark of cancer metabolism [100,101] and, in this instance, MnSOD is acting as a tumor promoter, specifically in breast cancer, as increased MnSOD expression and glycolysis both increased as severity of breast malignancies [90]. Short-term calorie restriction can also increase H2O2 levels, activating AMPK [98]. The combination of results from Han, D. et al. 2020 and Han, L. et al. 2020 [102,103] show a positive feedback loop with increased SIRT3 leading to activation of the AMPK pathway and phosphorylated AMPK upregulating SIRT3 expression. [103] Phloretin was shown to mitigate the oxidative stress induced by palmitic acid and increase phosphorylation of AMPK [103]. This also correlated to increased expression of SIRT3, which was reduced when an AMPK inhibitor (dorsomorphin) was used [103].
Alternatively, inactivation of MnSOD leads to a buildup of O2•−, which can also act as a signaling molecule that, under specific conditions, can oxidize and inactivate PTEN [104,105]. The buildup of O2•− due to the inactivation of SODs combines with nitric oxide to form peroxynitrite and nitrosylate PTEN, rendering it inactive [104,105,106]. This leads to increased phosphorylated (active) AKT to support cell growth. As a negative feedback loop, peroxynitrite can directly oxidize MnSOD-Tyr34, inactivating the enzyme [107]. In ER+ breast cancer, it has been reported that estrogen signaling relocates ER to the mitochondria, where it binds to MnSOD [105]. This blocks SIRT3 from binding and/or deacetylating MnSOD. The O2•− build-up leads to phosphorylation and activation of mTORC2, which phosphorylates and activates downstream targets AKT and PKC-α [105]. Hyper-acetylation of MnSOD can lead to over activation of AKT. Because AKT is often overactive in cancers, increasing cell survival and proliferation [108], active deacetylated MnSOD would be acting as a tumor suppressor. The totality of these results demonstrates the complexity of MnSOD’s roles in cell growth and strongly implies that PTMs can significantly determine the reparative or damaging effects of this detoxification enzyme.
6. MnSOD Signaling Linked to Lineage Plasticity
MnSOD has also been linked to signaling in cancer cells through the epithelial to mesenchymal transition (EMT), also referred to as a lineage-plasticity-like phenotypic switch, leading to more aggressive phenotypes, pan anticancer resistance properties, and a metastasis permissive tumor cell phenotype [109,110,111]. There is a positive correlation in MnSOD expression and EMT score of all subtypes of breast cancer and the subsequent knockdown of MnSOD in mesenchymal cells decreases their EMT markers and morphology [111]. Conversely, it was shown that FeTPPS (a peroxynitrite scavenger) mitigates the MET (mesenchymal–epithelial transition), the reversal of EMT, and the effect of siMnSOD. Similarly, in small-cell lung cancer (SCLC)-derived cancer stem-like cells, overexpression of MnSOD and FOXM1 mitigated the effects of genistein, decreasing stemness/lineage-plasticity-like markers and the profound migratory/invasive activity [109], whereas knockdown of MnSOD and FOXM1 enhanced the effect of genistein [109]. These studies all support the hypothesis that an increase in MnSOD can potentially lead to an increase in mitochondrial H2O2 concentrations, acting as a signaling molecule that increases expression of stemness/lineage-plasticity-like markers; however, the actual MnSOD activity in cells is not addressed in these experiments.
In a quite interesting study, it was shown that the forced overexpression of MnSOD also led to an increase in acetylation of lysine 68 in breast cancer cells [53]. He et al. showed that the acetylation of MnSOD correlates with the upregulation of stemness/lineage plasticity-like biomarkers Oct4, Sox2, and Nanog through HIF2α signaling. However, He et al. also showed that the inhibition of H2O2 signaling with overexpression of catalase reduces the expression of HIF2α, as well as stemness biomarkers, including Oct4 and Nanog [53]. Based on these results, they postulated that MnSOD-K68-Ac leads to higher H2O2 levels, which adds intriguing possibilities when compared to previous reports [83,112,113]. Interestingly, the MnSODK68Q mutant, which is a biochemically validated acetylation mimic, also showed an increase in stemness markers [53] and provided more supporting data regarding the increased MnSOD acetylation hypothesis. A study in squamous carcinoma cells showed that the invasive cell line (A431-III) had inherently more expression of MnSOD and higher H2O2 levels than the parental cell line (A431-P) [110]. Using siRNA, the knockdown of MnSOD led to an increase in H2O2 and invasiveness of the cells in both the parental and metastatic cell line [110]. Treatment with antioxidants, diphenyleneiodonium, and N-acetyl-l-cysteine blocked H2O2 signaling and reversed the effects of the siRNA [110]. These two papers showed that either the acetylation or the downregulation of MnSOD leads to higher mitochondrial H2O2 concentrations, which in turn leads to a potential lineage-plasticity-like switch to cells that exhibit stemness or EMT biomarkers.
Shedding light on the seemingly opposing nature of MnSOD, as it was previously discussed when cells are manipulated to express MnSOD-K68-Ac experimentally by using the acetylation mimic MnSODK68Q or made in bacteria using a codon expansion method, the concentration of monomeric MnSOD also increased as a proportion and/or change in the tetrameric to monomeric ratio [52]. In addition, the monomer form of MnSOD shows peroxidase activity instead of the more established superoxide dismutase activity [52,92]. Instead of acetylation being an on/off switch, the working hypothesis is that acetylation can cause a biological change of function. This hypothesis supports the theory of MnSOD being a signaling hub. Expressing the MnSODK68Q acetylation mimic mutant or silencing SIRT3 also shows a higher proportion of monomer and peroxidase activity [52]. A potential extension of these results could be that when MnSOD is overexpressed beyond its physiologically normal levels, a disproportionate increase in monomeric MnSOD occurs, which functions as a peroxidase [52]. Therefore, when it is silenced, as in Fan et al. [110], H2O2 concentration increases. The conditional nature of MnSOD all falls back to whether it is acetylated or not, and on the functionality of SIRT3.
7. The Cofactor of MnSOD and Peroxidase Activity, Another Layer of Regulation
MnSOD is an old, evolutionally conserved protein and is required for organisms that make ATP through respiration, primarily coming in two varieties that are categorized based on the cofactor identity. Eukaryotic MnSOD utilizes a Mn2+ cofactor, whereas bacteria can utilize both MnSOD and a distinct but similar FeSOD using Fe3+ as a cofactor. The essential structure of these enzymes is remarkably similar, especially in the active site region. In both, the active site region contains the metal cofactor ligated by three histidine residues, one aspartate residue, and one hydroxide ion that is stabilized by a highly conserved glutamine residue. In MnSOD, the glutamine arises from the C-terminal domain (Q146 E. coli), but in FeSOD, the glutamine arises from the N-terminal domain (Q69 E. coli) [114]. Despite the similarity of the enzyme structures, the proteins behave very differently with respect to incorporation of the metal cofactor.
It has been well established that substitution of iron into MnSOD, (Fe(Mn)SOD), leads to a virtually complete loss of dismutase activity for the enzyme. It is also well understood that the structure of the enzyme is virtually unchanged upon incorporation of the “wrong” metal and the loss of activity most likely arises from a deficiency in the reduction potential of the Fe-substituted enzyme. Vance et al. suggested that Fe2+(Mn)SOD is able to catalyze the reduction of O2•−, but the Fe3+(Mn)SOD generated from Fe2+(Mn)SOD during the previous reaction is not able to oxidize O2•−, and the catalytic cycle cannot be completed [115]. A similar argument can be made for Mn(Fe)SOD, which is also inactive as a dismutase. Thus, metalation is very important to these enzymes and various studies have been conducted to determine the consequences of mis-metalation. The specificity of MnSOD for the Mn cofactor is quite low. In fact, in bacteria, Fe(Mn)SOD is in 10–100 times greater abundance than MnSOD and the ratio of MnSOD/Fe(Mn)SOD is only increased under conditions of oxidative stress, such as the time when the catalytically active enzyme is needed [116]. However, within mitochondria, the enzyme is almost exclusively bound to the Mn2+ cofactor under normal conditions. Studies with S. cerevisiae have suggested that the exclusivity of MnSOD within mitochondria arises because the metalation process is coupled with the import of the unfolded polypeptide MnSOD into the mitochondria [116], and once the cofactor is incorporated into the enzyme, its removal has a high energy barrier.
Mis-metalation can be induced in S. cerevisiae by disruption of manganese trafficking factor (mtm1) [117]. Mtm1 plays a role in the insertion of Mn into the MnSOD enzyme and deletion of mtm1 has been shown in vivo to increase the concentration of Fe(Mn)SOD without reducing the concentration of mitochondrial manganese. Disruption of mitochondrial iron homeostasis, specifically via knockdown of ssq1 or grx5 [118], has also been shown to affect the MnSOD/Fe(Mn)SOD ratio in cells. Mis-metalation of MnSOD is also observed under conditions of Mn scarcity as the Fe-incorporated inactive enzyme is preferred over the apo-enzyme [119]. This is especially the case in cells where MnSOD is overexpressed [93,115,118].
Recent developments suggest that Fe(Mn)SOD has deleterious effects, not only because of its loss of dismutase function, but also because of a gained pro-oxidant/peroxidase function [120]. In early studies, this peroxidase function was attributed simply to overexpressed MnSOD, based on the oxidation of Amplex Red when MnSOD is incubated with H2O2 [92]. However, that interpretation has drawn some criticism [121] and several reports have shown that MnSOD is unable to oxidize ABTS∙+ [120] or Amplex Red [122], whereas Fe(Mn)SOD is able to oxidize those reagents. Ganini et al. showed that, in bacteria, the peroxidase activity is highly correlated to the concentration of Fe(Mn)SOD and the dismutase activity is highly correlated to the concentration of MnSOD [93]. Further, Ganini explored the metalation of MnSOD in mammalian cells cultivated in media with different iron to manganese ratios [122]. Cells grown in Mn-deficient media were observed to have significant iron incorporation, whereas cells grown in Mn-supplemented media were mainly bound to Mn. The cultivated Fe(Mn)SOD was then shown to catalyze the oxidation of Amplex Red in the presence of H2O2 and supported protein radical formation. The cultivated Mn-bound MnSOD was unable to oxidize Amplex Red and the formation of protein radicals was not observed [122]. Thus, the current evidence suggests that Fe incorporation into the MnSOD enzyme is a critical factor in the observed peroxidase activity and the role of overexpression of MnSOD is simply that it leads to a Mn deficiency in cells and makes mis-metalation more likely.
8. Conclusions
In conclusion, the data in this review article suggest that MnSOD-Ac, including K68-Ac, may function as an oncoprotein in an advanced stage of cancer, in contrast to the more traditional premise that MnSOD is a detoxification TS (Figure 2), and this dual function may arise from the presence of two structural forms of MnSOD. As such, it is surmised that acetylation status of MnSOD-K68 is a molecular switch that directs its enzymatic detoxification from its antioxidant activity (i.e., metabolic stress/protective function) as a tumor suppressor (tetrameric) or as a oncoprotein (monomeric) and, most importantly, that may promote a tumor-permissive phenotype through lineage-plasticity-like properties. This idea has, over the last decade, led to multiple publications suggesting a potential link between mitochondrial function, and dysregulation, in the lineage cell fate and plasticity in both stem cells, as well as tumor cells. It also expands the theme that the mitochondria, perhaps through small reactive oxygen species (ROS), may relay signals to the other parts of the cell to maintain both metabolic and bioenergetic equilibrium. The disruption of these pathways, through one of many potential processes, may lead to permissive phenotypes for human illness, including malignancies.
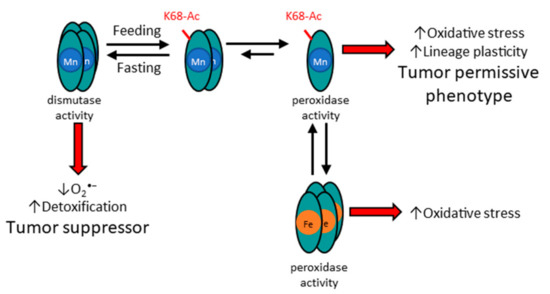
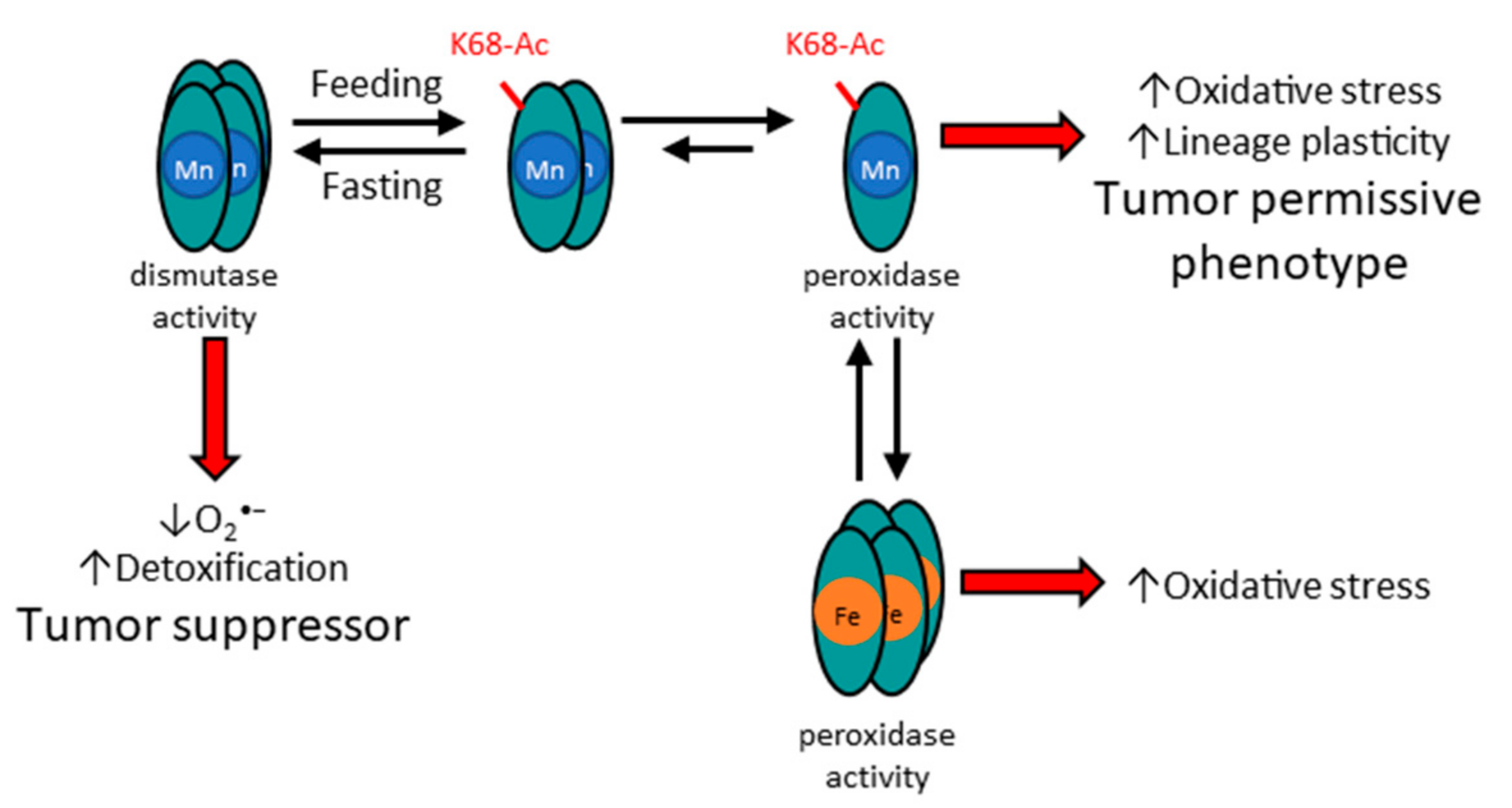
Figure 2.
Acetylation status of SOD2 affects structure and function, as well as metalation status. The potential mechanism for how acetylation of MnSOD lysine 68 alters the structure and function of MnSOD enzymatic activity. In this regard, it is proposed that acetylation, for example through feeding, leads to the formation of a dimer. In addition, it is surmised that mitochondrial damage via oxidative stress, or other forms of stress, may lead to iron incorporation of MnSOD.
Funding
This research was funded by the National Cancer Institute grant numbers R01CA152601, R01CA214025, and R01CA257148 and by the Cancer Prevention and Research Institute of Texas grant number RR20012. Meredith Ogle and Joe Schell are funded by the National Institute of Aging grant number 5T32AG021890-18.
Acknowledgments
D.G. is supported by R01CA152601, R01CA214025, and R01CA257148 from the National Cancer Institute, as well as the Cancer Prevention and Research Institute of Texas (CPRIT) grant no. RR20012. M.O. and J.S. are supported by a T32 T32AG021890 training grant from the NIA.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Sachdev, J.C.; Sandoval, A.C.; Jahanzeb, M. Update on precision medicine in breast cancer. Cancer Treat Res. 2019, 178, 45–80. [Google Scholar] [PubMed]
- Tsuchida, J.; Rothman, J.; McDonald, K.A.; Nagahashi, M.; Takabe, K.; Wakai, T. Clinical target sequencing for precision medicine of breast cancer. Int. J. Clin. Oncol. 2019, 24, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Meisel, J.L.; Venur, V.A.; Gnant, M.; Carey, L. Evolution of targeted therapy in breast cancer: Where precision medicine began. Am. Soc. Clin. Oncol. Educ. Book 2018, 38, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Naito, Y.; Urasaki, T. Precision medicine in breast cancer. Chin. Clin. Oncol. 2018, 7, 29. [Google Scholar] [CrossRef]
- Osborne, C.K.; Schiff, R. Mechanisms of endocrine resistance in breast cancer. Annu. Rev. Med. 2011, 62, 233–247. [Google Scholar] [CrossRef]
- Szostakowska, M.; Trebinska-Stryjewska, A.; Grzybowska, E.A.; Fabisiewicz, A. Resistance to endocrine therapy in breast cancer: Molecular mechanisms and future goals. Breast Cancer Res. Treat. 2019, 173, 489–497. [Google Scholar] [CrossRef]
- Ades, F.; Zardavas, D.; Bozovic-Spasojevic, I.; Pugliano, L.; Fumagalli, D.; de Azambuja, E.; Viale, G.; Sotiriou, C.; Piccart, M. Luminal b breast cancer: Molecular characterization, clinical management, and future perspectives. J. Clin. Oncol. 2014, 32, 2794–2803. [Google Scholar] [CrossRef]
- Creighton, C.J. The molecular profile of luminal b breast cancer. Biologics 2012, 6, 289–297. [Google Scholar] [CrossRef]
- Sotgia, F.; Fiorillo, M.; Lisanti, M.P. Mitochondrial markers predict recurrence, metastasis and tamoxifen-resistance in breast cancer patients: Early detection of treatment failure with companion diagnostics. Oncotarget 2017, 8, 68730–68745. [Google Scholar] [CrossRef]
- Rodriguez, D.; Ramkairsingh, M.; Lin, X.; Kapoor, A.; Major, P.; Tang, D. The central contributions of breast cancer stem cells in developing resistance to endocrine therapy in estrogen receptor (er)-positive breast cancer. Cancers (Basel) 2019, 11, 1028. [Google Scholar] [CrossRef]
- Beltran, H.; Hruszkewycz, A.; Scher, H.I.; Hildesheim, J.; Isaacs, J.; Yu, E.Y.; Kelly, K.; Lin, D.; Dicker, A.P.; Arnold, J.T.; et al. The role of lineage plasticity in prostate cancer therapy resistance. Clin. Cancer Res. 2019, 25, 6916–6924. [Google Scholar]
- Yuan, S.; Norgard, R.J.; Stanger, B.Z. Cellular plasticity in cancer. Cancer Discov. 2019, 9, 837–851. [Google Scholar] [CrossRef] [PubMed]
- Witt, A.E.; Lee, C.W.; Lee, T.I.; Azzam, D.J.; Wang, B.; Caslini, C.; Petrocca, F.; Grosso, J.; Jones, M.; Cohick, E.B.; et al. Identification of a cancer stem cell-specific function for the histone deacetylases, hdac1 and hdac7, in breast and ovarian cancer. Oncogene 2017, 36, 1707–1720. [Google Scholar] [CrossRef] [PubMed]
- Wahl, G.M.; Spike, B.T. Cell state plasticity, stem cells, emt, and the generation of intra-tumoral heterogeneity. NPJ Breast Cancer 2017, 3, 14. [Google Scholar] [CrossRef] [PubMed]
- Spike, B.T.; Engle, D.D.; Lin, J.C.; Cheung, S.K.; La, J.; Wahl, G.M. A mammary stem cell population identified and characterized in late embryogenesis reveals similarities to human breast cancer. Cell Stem Cell 2012, 10, 183–197. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.M.; Giltnane, J.M.; Balko, J.M.; Schwarz, L.J.; Guerrero-Zotano, A.L.; Hutchinson, K.E.; Nixon, M.J.; Estrada, M.V.; Sanchez, V.; Sanders, M.E.; et al. Myc and mcl1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab. 2017, 26, 633–647. [Google Scholar] [CrossRef]
- Yan, Y.; Liu, F.; Han, L.; Zhao, L.; Chen, J.; Olopade, O.I.; He, M.; Wei, M. Hif-2alpha promotes conversion to a stem cell phenotype and induces chemoresistance in breast cancer cells by activating wnt and notch pathways. J. Exp. Clin. Cancer Res. 2018, 37, 256. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Sundaresan, N.R.; Samant, S.A.; Pillai, V.B.; Rajamohan, S.B.; Gupta, M.P. Sirt3 is a stress responsive deacetylase in cardiomyocytes that protects cells from stress-mediated cell death by deacetylation of ku-70. Mol. Cell. Biol. 2008, 20, 6384–6401. [Google Scholar] [CrossRef]
- Finkel, T.; Deng, C.X.; Mostoslavsky, R. Recent progress in the biology and physiology of sirtuins. Nature 2009, 460, 587–591. [Google Scholar] [CrossRef] [PubMed]
- Ahn, B.H.; Kim, H.S.; Song, S.; Lee, I.H.; Liu, J.; Vassilopoulos, A.; Deng, C.X.; Finkel, T. A role for the mitochondrial deacetylase sirt3 in regulating energy homeostasis. Proc. Natl. Acad. Sci. USA 2008, 105, 14447–14452. [Google Scholar] [CrossRef] [PubMed]
- Imai, S.; Johnson, F.B.; Marciniak, R.A.; McVey, M.; Park, P.U.; Guarente, L. Sir2: An nad-dependent histone deacetylase that connects chromatin silencing, metabolism, and aging. Cold Spring Harb. Symp. Quant. Biol. 2000, 65, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Landry, J.; Sutton, A.; Tafrov, S.T.; Heller, R.C.; Stebbins, J.; Pillus, L.; Sternglanz, R. The silencing protein sir2 and its homologs are nad-dependent protein deacetylases. Proc. Natl. Acad. Sci. USA 2000, 97, 5807–5811. [Google Scholar] [CrossRef] [PubMed]
- Sebastian, C.; Zwaans, B.M.; Silberman, D.M.; Gymrek, M.; Goren, A.; Zhong, L.; Ram, O.; Truelove, J.; Guimaraes, A.R.; Toiber, D.; et al. The histone deacetylase sirt6 is a tumor suppressor that controls cancer metabolism. Cell 2012, 151, 1185–1199. [Google Scholar] [CrossRef]
- Kim, H.S.; Vassilopoulos, A.; Wang, R.H.; Lahusen, T.; Xiao, Z.; Xu, X.; Li, C.; Veenstra, T.D.; Li, B.; Yu, H.; et al. Sirt2 maintains genome integrity and suppresses tumorigenesis through regulating apc/c activity. Cancer Cell 2011, 20, 487–499. [Google Scholar] [CrossRef]
- Huang, J.Y.; Hirschey, M.D.; Shimazu, T.; Ho, L.; Verdin, E. Mitochondrial sirtuins. Biochim. Biophys. Acta. 2010, 1804, 1645–1651. [Google Scholar] [CrossRef]
- Hirschey, M.D.; Shimazu, T.; Huang, J.Y.; Verdin, E. Acetylation of mitochondrial proteins. Methods Enzym. 2009, 457, 137–147. [Google Scholar]
- Knyphausen, P.; de Boor, S.; Kuhlmann, N.; Scislowski, L.; Extra, A.; Baldus, L.; Schacherl, M.; Baumann, U.; Neundorf, I.; Lammers, M. Insights into lysine deacetylation of natively folded substrate proteins by sirtuins. J. Biol. Chem. 2016, 291, 14677–14694. [Google Scholar] [CrossRef]
- Kaeberlein, M.; McVey, M.; Guarente, L. The sir2/3/4 complex and sir2 alone promote longevity in saccharomyces cerevisiae by two different mechanisms. Genes Dev. 1999, 13, 2570–2580. [Google Scholar] [CrossRef]
- Guarente, L. Sir2 links chromatin silencing, metabolism, and aging. Genes Dev. 2000, 14, 1021–1026. [Google Scholar] [CrossRef] [PubMed]
- Hirschey, M.D.; Shimazu, T.; Goetzman, E.; Jing, E.; Schwer, B.; Lombard, D.B.; Grueter, C.A.; Harris, C.; Biddinger, S.; Ilkayeva, O.R.; et al. Sirt3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation. Nature 2010, 464, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Patel, K.; Muldoon-Jacobs, K.; Bisht, K.S.; Aykin-Burns, N.; Pennington, J.D.; van der Meer, R.; Nguyen, P.; Savage, J.; Owens, K.M.; et al. Sirt3 is a mitochondria-localized tumor suppressor required for maintenance of mitochondrial integrity and metabolism during stress. Cancer Cell 2010, 17, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Brown, K.; Hirschey, M.D.; Verdin, E.; Chen, D. Calorie restriction reduces oxidative stress by sirt3-mediated sod2 activation. Cell Metab. 2010, 12, 662–667. [Google Scholar] [CrossRef] [PubMed]
- Tao, R.; Coleman, M.C.; Pennington, J.D.; Ozden, O.; Park, S.H.; Jiang, H.; Kim, H.S.; Flynn, C.R.; Hill, S.; Hayes McDonald, W.; et al. Sirt3-mediated deacetylation of evolutionarily conserved lysine 122 regulates mnsod activity in response to stress. Molecular Cell 2010, 40, 893–904. [Google Scholar] [CrossRef] [PubMed]
- Tao, R.; Vassilopoulos, A.; Parisiadou, L.; Yan, Y.; Gius, D. Regulation of mnsod enzymatic activity by sirt3 connects the mitochondrial acetylome signaling networks to aging and carcinogenesis. Antioxid. Redox Signal. 2014, 20, 1646–1654. [Google Scholar] [CrossRef]
- Guarente, G.P. Hook, Line & Sinker; Razorbill: New York, NY, USA, 2005; p. 198. [Google Scholar]
- Chen, Y.; Zhang, J.; Lin, Y.; Lei, Q.; Guan, K.L.; Zhao, S.; Xiong, Y. Tumour suppressor sirt3 deacetylates and activates manganese superoxide dismutase to scavenge ros. EMBO Rep. 2011, 12, 534–541. [Google Scholar] [CrossRef]
- Finley, L.W.; Carracedo, A.; Lee, J.; Souza, A.; Egia, A.; Zhang, J.; Teruya-Feldstein, J.; Moreira, P.I.; Cardoso, S.M.; Clish, C.B.; et al. Sirt3 opposes reprogramming of cancer cell metabolism through hif1alpha destabilization. Cancer Cell 2011, 19, 416–428. [Google Scholar] [CrossRef]
- Haigis, M.C.; Deng, C.X.; Finley, L.W.; Kim, H.S.; Gius, D. Sirt3 is a mitochondrial tumor suppressor: A scientific tale that connects aberrant cellular ros, the warburg effect, and carcinogenesis. Cancer Res. 2012, 72, 2468–2472. [Google Scholar] [CrossRef]
- Schwer, B.; North, B.J.; Frye, R.A.; Ott, M.; Verdin, E. The human silent information regulator (sir)2 homologue hsirt3 is a mitochondrial nicotinamide adenine dinucleotide-dependent deacetylase. J. Cell Biol. 2002, 158, 647–657. [Google Scholar] [CrossRef]
- Scher, M.B.; Vaquero, A.; Reinberg, D. Sirt3 is a nuclear nad+-dependent histone deacetylase that translocates to the mitochondria upon cellular stress. Genes Dev. 2007, 21, 920–928. [Google Scholar] [CrossRef] [PubMed]
- Pirinen, E.; Lo Sasso, G.; Auwerx, J. Mitochondrial sirtuins and metabolic homeostasis. Best Pr. Res. Clin. Endocrinol. Metab. 2012, 26, 759–770. [Google Scholar] [CrossRef] [PubMed]
- Ozden, O.; Park, S.H.; Kim, H.S.; Jiang, H.; Coleman, M.C.; Spitz, D.R.; Gius, D. Acetylation of mnsod directs enzymatic activity responding to cellular nutrient status or oxidative stress. Aging 2011, 3, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Zou, X.; Santa-Maria, C.A.; O’Brien, J.; Gius, D.; Zhu, Y. Manganese superoxide dismutase acetylation and dysregulation, due to loss of sirt3 activity, promote a luminal b-like breast carcinogenic-permissive phenotype. Antioxid. Redox Signal. 2016, 25, 326–336. [Google Scholar] [CrossRef]
- Zhu, Y.; Yan, Y.; Principe, D.R.; Zou, X.; Vassilopoulos, A.; Gius, D. Sirt3 and sirt4 are mitochondrial tumor suppressor proteins that connect mitochondrial metabolism and carcinogenesis. Cancer Metab. 2014, 2, 15. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Dean, A.E.; Horikoshi, N.; Heer, C.; Spitz, D.R.; Gius, D. Emerging evidence for targeting mitochondrial metabolic dysfunction in cancer therapy. J. Clin. Investig. 2018, 128, 3682–3691. [Google Scholar] [CrossRef]
- Zhu, Y.; Paul, P.; Lee, S.; Craig, B.T.; Rellinger, E.J.; Qiao, J.; Gius, D.R.; Chung, D.H. Antioxidant inhibition of steady-state reactive oxygen species and cell growth in neuroblastoma. Surgery 2015, 158, 827–836. [Google Scholar] [CrossRef][Green Version]
- Lu, J.; Cheng, K.; Zhang, B.; Xu, H.; Cao, Y.; Guo, F.; Feng, X.; Xia, Q. Novel mechanisms for superoxide-scavenging activity of human manganese superoxide dismutase determined by the k68 key acetylation site. Free Radic. Biol. Med. 2015, 85, 114–126. [Google Scholar] [CrossRef]
- Zelko, I.N.; Mariani, T.J.; Folz, R.J. Superoxide dismutase multigene family: A comparison of the cuzn-sod (sod1), mn-sod (sod2), and ec-sod (sod3) gene structures, evolution, and expression. Free Radic. Biol. Med. 2002, 33, 337–349. [Google Scholar] [CrossRef]
- Brown, K.; Xie, S.; Qiu, X.; Mohrin, M.; Shin, J.; Liu, Y.; Zhang, D.; Scadden, D.T.; Chen, D. Sirt3 reverses aging-associated degeneration. Cell Rep. 2013, 3, 319–327. [Google Scholar] [CrossRef]
- Zhu, Y.; Zou, X.; Dean, A.E.; Brien, J.O.; Gao, Y.; Tran, E.L.; Park, S.H.; Liu, G.; Kieffer, M.B.; Jiang, H.; et al. Lysine 68 acetylation directs mnsod as a tetrameric detoxification complex versus a monomeric tumor promoter. Nat. Commun. 2019, 10, 2399. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Danes, J.M.; Hart, P.C.; Zhu, Y.; Huang, Y.; de Abreu, A.L.; O’Brien, J.; Mathison, A.J.; Tang, B.; Frasor, J.M.; et al. Sod2 acetylation on lysine 68 promotes stem cell reprogramming in breast cancer. Proc. Natl. Acad. Sci. USA 2019, 116, 23534–23541. [Google Scholar] [CrossRef] [PubMed]
- Van Remmen, H.; Salvador, C.; Yang, H.; Huang, T.T.; Epstein, C.J.; Richardson, A. Characterization of the antioxidant status of the heterozygous manganese superoxide dismutase knockout mouse. Arch. Biochem. Biophys 1999, 363, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Van Remmen, H.; Qi, W.; Sabia, M.; Freeman, G.; Estlack, L.; Yang, H.; Mao Guo, Z.; Huang, T.T.; Strong, R.; Lee, S.; et al. Multiple deficiencies in antioxidant enzymes in mice result in a compound increase in sensitivity to oxidative stress. Free Radic. Biol. Med. 2004, 36, 1625–1634. [Google Scholar] [CrossRef]
- Holley, A.K.; Dhar, S.K.; St Clair, D.K. Curbing cancer’s sweet tooth: Is there a role for mnsod in regulation of the warburg effect? Mitochondrion 2013, 13, 170–188. [Google Scholar] [CrossRef]
- Miriyala, S.; Holley, A.K.; St Clair, D.K. Mitochondrial superoxide dismutase--signals of distinction. Anticancer. Agents Med. Chem. 2011, 11, 181–190. [Google Scholar] [CrossRef]
- Van Remmen, H.; Ikeno, Y.; Hamilton, M.; Pahlavani, M.; Wolf, N.; Thorpe, S.R.; Alderson, N.L.; Baynes, J.W.; Epstein, C.J.; Huang, T.T.; et al. Life-long reduction in mnsod activity results in increased DNA damage and higher incidence of cancer but does not accelerate aging. Physiol Genom. 2003, 16, 29–37. [Google Scholar] [CrossRef]
- Turrens, J.F. Mitochondrial formation of reactive oxygen species. J. Physiol. 2003, 552, 335–344. [Google Scholar] [CrossRef]
- Turrens, J.F.; Boveris, A. Generation of superoxide anion by the nadh dehydrogenase of bovine heart mitochondria. Biochem. J. 1980, 191, 421–427. [Google Scholar] [CrossRef]
- Buettner, G.R. Superoxide dismutase in redox biology: The roles of superoxide and hydrogen peroxide. Anticancer. Agents Med. Chem. 2011, 11, 341–346. [Google Scholar] [CrossRef]
- Zhu, Y.; Park, S.H.; Ozden, O.; Kim, H.S.; Jiang, H.; Vassilopoulos, A.; Spitz, D.R.; Gius, D. Exploring the electrostatic repulsion model in the role of sirt3 in directing mnsod acetylation status and enzymatic activity. Free Radic. Biol. Med. 2012, 53, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Oberley, L.W.; Spitz, D.R. Assay of superoxide dismutase activity in tumor tissue. Methods Enzym. 1984, 105, 457–464. [Google Scholar]
- Cai, M.Y.; Tong, Z.T.; Zheng, F.; Liao, Y.J.; Wang, Y.; Rao, H.L.; Chen, Y.C.; Wu, Q.L.; Liu, Y.H.; Guan, X.Y.; et al. Ezh2 protein: A promising immunomarker for the detection of hepatocellular carcinomas in liver needle biopsies. Gut 2011, 60, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Willson, R.L. Iron, zinc, free radicals and oxygen in tissue disorders and cancer control. In Ciba Foundation Symposium 51-Iron Metabolism; John Wiley & Sons, Ltd.: Chichester, UK, 1977; pp. 331–369. [Google Scholar]
- Hosp, F.; Lassowskat, I.; Santoro, V.; De Vleesschauwer, D.; Fliegner, D.; Redestig, H.; Mann, M.; Christian, S.; Hannah, M.A.; Finkemeier, I. Lysine acetylation in mitochondria: From inventory to function. Mitochondrion 2017, 33, 58–71. [Google Scholar] [CrossRef] [PubMed]
- Weinert, B.T.; Wagner, S.A.; Horn, H.; Henriksen, P.; Liu, W.R.; Olsen, J.V.; Jensen, L.J.; Choudhary, C. Proteome-wide mapping of the drosophila acetylome demonstrates a high degree of conservation of lysine acetylation. Sci. Signal. 2011, 4, ra48. [Google Scholar] [CrossRef]
- Blander, G.; Guarente, L. The sir2 family of protein deacetylases. Annu. Rev. Biochem. 2004, 73, 417–435. [Google Scholar] [CrossRef]
- Motta, M.C.; Divecha, N.; Lemieux, M.; Kamel, C.; Chen, D.; Gu, W.; Bultsma, Y.; McBurney, M.; Guarente, L. Mammalian sirt1 represses forkhead transcription factors. Cell 2004, 116, 551–563. [Google Scholar] [CrossRef]
- Tanner, K.G.; Landry, J.; Sternglanz, R.; Denu, J.M. Silent information regulator 2 family of nad- dependent histone/protein deacetylases generates a unique product, 1-o-acetyl-adp-ribose. Proc. Natl. Acad. Sci. USA 2000, 97, 14178–14182. [Google Scholar] [CrossRef]
- Imai, S.; Armstrong, C.M.; Kaeberlein, M.; Guarente, L. Transcriptional silencing and longevity protein sir2 is an nad-dependent histone deacetylase. Nature 2000, 403, 795–800. [Google Scholar] [CrossRef]
- Yang, X.J.; Seto, E. Lysine acetylation: Codified crosstalk with other posttranslational modifications. Molecular Cell 2008, 31, 449–461. [Google Scholar] [CrossRef]
- Schwer, B.; Eckersdorff, M.; Li, Y.; Silva, J.C.; Fermin, D.; Kurtev, M.V.; Giallourakis, C.; Comb, M.J.; Alt, F.W.; Lombard, D.B. Calorie restriction alters mitochondrial protein acetylation. Aging Cell 2009, 8, 604–606. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.C.; Sprung, R.; Chen, Y.; Xu, Y.; Ball, H.; Pei, J.; Cheng, T.; Kho, Y.; Xiao, H.; Xiao, L.; et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Molecular Cell 2006, 23, 607–618. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, C.; Kumar, C.; Gnad, F.; Nielsen, M.L.; Rehman, M.; Walther, T.C.; Olsen, J.V.; Mann, M. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 2009, 325, 834–840. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Xu, W.; Jiang, W.; Yu, W.; Lin, Y.; Zhang, T.; Yao, J.; Zhou, L.; Zeng, Y.; Li, H.; et al. Regulation of cellular metabolism by protein lysine acetylation. Science 2010, 327, 1000–1004. [Google Scholar] [CrossRef] [PubMed]
- Guan, K.L.; Xiong, Y. Regulation of intermediary metabolism by protein acetylation. Trends Biochem. Sci. 2011, 36, 108–116. [Google Scholar] [CrossRef]
- Schwer, B.; Bunkenborg, J.; Verdin, R.O.; Andersen, J.S.; Verdin, E. Reversible lysine acetylation controls the activity of the mitochondrial enzyme acetyl-coa synthetase 2. Proc. Natl. Acad. Sci. USA 2006, 103, 10224–10229. [Google Scholar] [CrossRef]
- Li, X.; Zhang, S.; Blander, G.; Tse, J.G.; Krieger, M.; Guarente, L. Sirt1 deacetylates and positively regulates the nuclear receptor lxr. Molecular Cell 2007, 28, 91–106. [Google Scholar] [CrossRef]
- Vassilopoulos, A.; Pennington, J.D.; Andresson, T.; Rees, D.M.; Bosley, A.D.; Fearnley, I.M.; Ham, A.; Flynn, C.R.; Hill, S.; Rose, K.L.; et al. Sirt3 deacetylates atp synthase f1 complex proteins in response to nutrient- and exercise-induced stress. Antioxid. Redox. Signal. 2014, 21, 551–564. [Google Scholar] [CrossRef]
- Lu, J.; Zhang, H.; Chen, X.; Zou, Y.; Li, J.; Wang, L.; Wu, M.; Zang, J.; Yu, Y.; Zhuang, W.; et al. A small molecule activator of sirt3 promotes deacetylation and activation of manganese superoxide dismutase. Free Radic. Biol. Med. 2017, 112, 287–297. [Google Scholar] [CrossRef]
- Sheng, Y.; Durazo, A.; Schumacher, M.; Gralla, E.B.; Cascio, D.; Cabelli, D.E.; Valentine, J.S. Tetramerization reinforces the dimer interface of mnsod. PLoS ONE 2013, 8, e62446. [Google Scholar] [CrossRef]
- Oberley, L.W. Mechanism of the tumor suppressive effect of mnsod overexpression. Biomed. Pharmacother. 2005, 59, 143–148. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, S.; Jiang, X.; Weydert, C.; Zhang, Y.; Zhang, H.J.; Goswami, P.C.; Ritchie, J.M.; Oberley, L.W.; Buettner, G.R. Manganese superoxide dismutase overexpression inhibits the growth of androgen-independent prostate cancer cells. Oncogene 2005, 24, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Land, H.; Chen, A.C.; Morgenstern, J.P.; Parada, L.F.; Weinberg, R.A. Behavior of myc and ras oncogenes in transformation of rat embryo fibroblasts. Mol. Cell. Biol. 1986, 6, 1917–1925. [Google Scholar] [PubMed]
- Land, H.; Parada, L.F.; Weinberg, R.A. Tumorigenic conversion of primary embryo fibroblasts requires at least two cooperating oncogenes. Nature 1983, 304, 596–602. [Google Scholar] [CrossRef]
- Gao, Y.; Zhu, Y.; Tran, E.L.; Tokars, V.; Dean, A.E.; Quan, S.; Gius, D. Mnsod lysine 68 acetylation leads to cisplatin and doxorubicin resistance due to aberrant mitochondrial metabolism. Int. J. Biol. Sci. 2021, 17, 1203–1216. [Google Scholar] [CrossRef]
- He, T.; Weintraub, N.L.; Goswami, P.C.; Chatterjee, P.; Flaherty, D.M.; Domann, F.E.; Oberley, L.W. Redox factor-1 contributes to the regulation of progression from g0/g1 to s by pdgf in vascular smooth muscle cells. Am. J. Physiol. Heart Circ. Physiol. 2003, 285, H804–H812. [Google Scholar] [CrossRef]
- Kattan, Z.; Minig, V.; Leroy, P.; Dauca, M.; Becuwe, P. Role of manganese superoxide dismutase on growth and invasive properties of human estrogen-independent breast cancer cells. Breast Cancer Res. Treat. 2008, 108, 203–215. [Google Scholar] [CrossRef]
- Hart, P.C.; Mao, M.; de Abreu, A.L.; Ansenberger-Fricano, K.; Ekoue, D.N.; Ganini, D.; Kajdacsy-Balla, A.; Diamond, A.M.; Minshall, R.D.; Consolaro, M.E.; et al. Mnsod upregulation sustains the warburg effect via mitochondrial ros and ampk-dependent signalling in cancer. Nat. Commun. 2015, 6, 6053. [Google Scholar] [CrossRef]
- Gius, D.; Spitz, D.R. Redox signaling in cancer biology. Antioxid. Redox Signal. 2006, 8, 1249–1252. [Google Scholar] [CrossRef]
- Ansenberger-Fricano, K.; Ganini, D.; Mao, M.; Chatterjee, S.; Dallas, S.; Mason, R.P.; Stadler, K.; Santos, J.H.; Bonini, M.G. The peroxidase activity of mitochondrial superoxide dismutase. Free Radic. Biol. Med. 2013, 54, 116–124. [Google Scholar] [CrossRef]
- Ganini, D.; Petrovich, R.M.; Edwards, L.L.; Mason, R.P. Iron incorporation into mnsod a (bacterial mn-dependent superoxide dismutase) leads to the formation of a peroxidase/catalase implicated in oxidative damage to bacteria. Biochim. Biophys. Acta. 2015, 1850, 1795–1805. [Google Scholar] [CrossRef] [PubMed]
- Dhar, S.K.; St Clair, D.K. Manganese superoxide dismutase regulation and cancer. Free Radic. Biol. Med. 2012, 52, 2209–2222. [Google Scholar] [CrossRef] [PubMed]
- Zou, X.; Zhu, Y.; Park, S.H.; Liu, G.; O’Brien, J.; Jiang, H.; Gius, D. Sirt3-mediated dimerization of idh2 directs cancer cell metabolism and tumor growth. Cancer Res. 2017, 77, 3990–3999. [Google Scholar] [CrossRef]
- Gill, T.; Levine, A.D. Mitochondria-derived hydrogen peroxide selectively enhances t cell receptor-initiated signal transduction. J. Biol. Chem. 2013, 288, 26246–26255. [Google Scholar] [CrossRef]
- Lim, C.S. Is sirt6 a new biomarker for oxidative stress and longevity assurance gene? Med. Hypotheses 2007, 69, 231. [Google Scholar] [CrossRef]
- Garcia-Prieto, C.F.; Gil-Ortega, M.; Plaza, A.; Manzano-Lista, F.J.; Gonzalez-Blazquez, R.; Alcala, M.; Rodriguez-Rodriguez, P.; Viana, M.; Aranguez, I.; Gollasch, M.; et al. Caloric restriction induces h2o2 formation as a trigger of ampk-enos-no pathway in obese rats: Role for camkii. Free Radic. Biol. Med. 2019, 139, 35–45. [Google Scholar] [CrossRef]
- Zhou, Z.; Song, X.; Chi, J.J.; Gius, D.R.; Huang, Y.; Cristofanilli, M.; Wan, Y. Regulation of klf4 by posttranslational modification circuitry in endocrine resistance. Cell. Signal. 2020, 70, 109574. [Google Scholar] [CrossRef]
- Samudio, I.; Fiegl, M.; Andreeff, M. Mitochondrial uncoupling and the warburg effect: Molecular basis for the reprogramming of cancer cell metabolism. Cancer. Res. 2009, 69, 2163–2166. [Google Scholar] [CrossRef]
- Warburg, O. Iron, the oxygen-carrier of respiration-ferment. Science 1925, 61, 575–582. [Google Scholar] [CrossRef]
- Han, D.; Jiang, L.; Gu, X.; Huang, S.; Pang, J.; Wu, Y.; Yin, J.; Wang, J. Sirt3 deficiency is resistant to autophagy-dependent ferroptosis by inhibiting the ampk/mtor pathway and promoting gpx4 levels. J. Cell. Physiol. 2020, 235, 8839–8851. [Google Scholar] [CrossRef]
- Han, L.; Li, J.; Li, J.; Pan, C.; Xiao, Y.; Lan, X.; Wang, M. Activation of ampk/sirt3 pathway by phloretin reduces mitochondrial ros in vascular endothelium by increasing the activity of mnsod via deacetylation. Food Funct. 2020, 11, 3073–3083. [Google Scholar] [CrossRef]
- Vidimar, V.; Gius, D.; Chakravarti, D.; Bulun, S.E.; Wei, J.J.; Kim, J.J. Dysfunctional mnsod leads to redox dysregulation and activation of prosurvival akt signaling in uterine leiomyomas. Sci. Adv. 2016, 2, e1601132. [Google Scholar] [CrossRef]
- Lone, M.U.; Baghel, K.S.; Kanchan, R.K.; Shrivastava, R.; Malik, S.A.; Tewari, B.N.; Tripathi, C.; Negi, M.P.; Garg, V.K.; Sharma, M.; et al. Physical interaction of estrogen receptor with mnsod: Implication in mitochondrial o2(.-) upregulation and mtorc2 potentiation in estrogen-responsive breast cancer cells. Oncogene 2017, 36, 1829–1839. [Google Scholar] [CrossRef]
- Lim, E.H.; Zhang, S.L.; Li, J.L.; Yap, W.S.; Howe, T.C.; Tan, B.P.; Lee, Y.S.; Wong, D.; Khoo, K.L.; Seto, K.Y.; et al. Using whole genome amplification (wga) of low-volume biopsies to assess the prognostic role of egfr, kras, p53, and cmet mutations in advanced-stage non-small cell lung cancer (nsclc). J. Thorac. Oncol. 2009, 4, 12–21. [Google Scholar] [CrossRef]
- Demicheli, V.; Moreno, D.M.; Radi, R. Human mn-superoxide dismutase inactivation by peroxynitrite: A paradigm of metal-catalyzed tyrosine nitration in vitro and in vivo. Metallomics 2018, 10, 679–695. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Gupta Vallur, P.; Phaeton, R.; Mythreye, K.; Hempel, N. Insights into the dichotomous regulation of sod2 in cancer. Antioxidants (Basel) 2017, 6, 86. [Google Scholar] [CrossRef]
- Fu, Z.; Cao, X.; Liu, L.; Cao, X.; Cui, Y.; Li, X.; Quan, M.; Ren, K.; Chen, A.; Xu, C.; et al. Genistein inhibits lung cancer cell stem-like characteristics by modulating mnsod and foxm1 expression. Oncol. Lett. 2020, 20, 2506–2515. [Google Scholar] [CrossRef]
- Fan, J.J.; Hsu, W.H.; Hung, H.H.; Zhang, W.J.; Lee, Y.A.; Chen, K.C.; Chu, C.Y.; Ko, T.P.; Lee, M.T.; Lin, C.W.; et al. Reduction in mnsod promotes the migration and invasion of squamous carcinoma cells. Int. J. Oncol. 2019, 54, 1639–1650. [Google Scholar] [CrossRef]
- Loo, S.Y.; Hirpara, J.L.; Pandey, V.; Tan, T.Z.; Yap, C.T.; Lobie, P.E.; Thiery, J.P.; Goh, B.C.; Pervaiz, S.; Clement, M.V.; et al. Manganese superoxide dismutase expression regulates the switch between an epithelial and a mesenchymal-like phenotype in breast carcinoma. Antioxid Redox Signal. 2016, 25, 283–299. [Google Scholar] [CrossRef]
- Pani, G.; Colavitti, R.; Bedogni, B.; Fusco, S.; Ferraro, D.; Borrello, S.; Galeotti, T. Mitochondrial superoxide dismutase: A promising target for new anticancer therapies. Curr. Med. Chem. 2004, 11, 1299–1308. [Google Scholar] [CrossRef]
- Zhao, Y.; Oberley, T.D.; Chaiswing, L.; Lin, S.M.; Epstein, C.J.; Huang, T.T.; St Clair, D. Manganese superoxide dismutase deficiency enhances cell turnover via tumor promoter-induced alterations in ap-1 and p53-mediated pathways in a skin cancer model. Oncogene 2002, 21, 3836–3846. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, J.W. The irony of manganese superoxide dismutase. Biochem. Soc. Trans. 2003, 31, 1318–1321. [Google Scholar] [CrossRef] [PubMed]
- Vance, C.K.; Miller, A.F. Novel insights into the basis for escherichia coli superoxide dismutase’s metal ion specificity from mn-substituted fesod and its very high e(m). Biochemistry 2001, 40, 13079–13087. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, J.D.; Culotta, V.C. Battles with iron: Manganese in oxidative stress protection. J. Biol. Chem. 2012, 287, 13541–13548. [Google Scholar] [CrossRef] [PubMed]
- Luk, E.; Carroll, M.; Baker, M.; Culotta, V.C. Manganese activation of superoxide dismutase 2 in saccharomyces cerevisiae requires mtm1, a member of the mitochondrial carrier family. Proc. Natl. Acad. Sci. USA 2003, 100, 10353–10357. [Google Scholar] [CrossRef]
- Yang, M.; Cobine, P.A.; Molik, S.; Naranuntarat, A.; Lill, R.; Winge, D.R.; Culotta, V.C. The effects of mitochondrial iron homeostasis on cofactor specificity of superoxide dismutase 2. EMBO J. 2006, 25, 1775–1783. [Google Scholar] [CrossRef]
- Pierrel, F.; Cobine, P.A.; Winge, D.R. Metal ion availability in mitochondria. Biometals 2007, 20, 675–682. [Google Scholar] [CrossRef]
- Yamakura, F.; Kobayashi, K.; Furukawa, S.; Suzuki, Y. In vitro preparation of iron-substituted human manganese superoxide dismutase: Possible toxic properties for mitochondria. Free Radic. Biol. Med. 2007, 43, 423–430. [Google Scholar] [CrossRef]
- Liochev, S.I.; Fridovich, I. Peroxidase activity by mnsod? Free Radic. Biol. Med. 2013, 65, 1535. [Google Scholar] [CrossRef]
- Ganini, D.; Santos, J.H.; Bonini, M.G.; Mason, R.P. Switch of mitochondrial superoxide dismutase into a prooxidant peroxidase in manganese-deficient cells and mice. Cell Chem. Biol. 2018, 25, 413–425. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).