Abstract
The endangered ethnomedicinal plant Euchresta tubulosa harbors a valuable community of endophytic fungi, demonstrating significant potential for biotechnological applications. Endophytic fungi were isolated from E. tubulosa and identified to characterize their secondary metabolites and extracellular enzyme activities. Endophytic fungi were isolated from E. tubulosa using tissue explant culture and identified by morphological and molecular (ITS) analysis. The chemical profiles of strain fermentation products were analyzed by LC–MS/MS, while extracellular enzyme production (cellulase, protease, amylase) was assessed through chromogenic plate assays and liquid fermentation. The results indicated that a total of 55 endophytic fungi were isolated from E. tubulosa, assigned to 17 genera. Among these, three genera (Colletotrichum, Fusarium, and Penicillium) constituted the dominant groups, while four strains (including three novel species) represented potential new taxa. LC–MS/MS analysis revealed that fermentation products of the three novel endophytic fungal species contained bioactive compounds such as flavonoids and alkaloids; furthermore, bioactivity assays indicate that they exhibited significant degrees of antibacterial and antioxidant activity. Extracellular enzyme assays demonstrated that three E. tubulosa-derived endophytic strains exhibited multi-hydrolytic enzyme production capabilities. Notably, strain ETG-1-2-1 showed the highest amylase and cellulase activities (10.95 U/mL and 9.68 U/mL, respectively), while strain ETXG-1-1-1 displayed the highest protease activity (2.34 U/mL). This study provides the first systematic report on the diversity of endophytic fungi in E. tubulosa, their secondary metabolite profiles, and extracellular enzyme activities, establishing a theoretical foundation for discovering novel bioactive compounds and developing microbial resources, while also highlighting their ecological roles and biotechnological potential.
1. Introduction
The discovery of natural product-based drugs has long relied on wild plant resources. However, with growing challenges such as the depletion of these wild resources, identifying new biological sources has become an urgent need in contemporary drug development and related societal issues [1]. Endophytic fungi are a group of fungi that reside either permanently or temporarily within plant tissues (such as roots, stems, leaves, seeds, etc.), forming a symbiotic relationship with the host plant without causing overt disease symptoms [2,3,4]. In 1993, Stierle et al. isolated an endophytic fungus (Taxomyces andreanae) that produces paclitaxel from the phloem of the Pacific yew (Taxus brevifolia Nutt) [5]. This landmark discovery markedly expanded scientific understanding of the functional roles of plant endophytes.
Endophytic fungi interact with plants to shape community diversity and host specificity [6,7]. They are intricately linked to host growth and development [8], reproduction [9], repair mechanisms [10], and the secretion of metabolites [11], and are thus viewed as extensions of the host’s genetic repertoire [12]. Research by Taheri et al. revealed that endophytic fungal isolates from hairy vetch (Vicia villosa Roth) exhibit the potential to produce indole-3-acetic acid (IAA) and siderophores, along with phosphate- and potassium-solubilizing activities [13]. Among these, the newly described endophytic fungus Penicillium griseofulvum AKL25 demonstrated particularly high plant growth-promoting activity and shows promise as a microbial inoculant for enhancing the growth of hairy vetch and soybean in sustainable agriculture systems. In addition, endophytic fungi exhibit significant advantages in terms of metabolite diversity due to their rapid, reproducible, and unrestricted production, which is not affected by weather or seasonal factors [14]. Furthermore, their fermentation products exhibit diverse biological activities, including anticancer, antioxidant, and pathogen-inhibiting effects [15]. For example, a study by Sunil K. Deshmukh et al. pointed out that among the secondary metabolites produced by mangrove fungi, 91, 42, and 33 compounds have been reported to exhibit antibacterial, antifungal, and antiviral activities, respectively [16]. Separately, Zhang Yi et al. isolated endophytic fungi from Carpesium abrotanoides that demonstrated inhibitory activity against Hep G2 human liver cancer cells [17]. Among these, strain TMJ-54 exhibited the strongest suppression and significantly induced apoptosis. This finding highlights the potential of these fungi as valuable microbial resources for novel anti-hepatoma drug discovery. The above studies not only confirm the essentiality of endophytic fungi in host survival but also provide a basis for developing novel natural drug lead molecules from endophytic fungi to alleviate the shortage of plant-derived metabolites.
Endophytic fungi can also resist pathogenic bacteria by synthesizing extracellular enzymes, enhancing the host plant’s tolerance to biotic and abiotic stresses. Currently, Trichoderma species are widely deployed as biocontrol agents against soil-borne pathogens. Their secreted extracellular enzymes lyse pathogen cell walls, thereby achieving antimicrobial effects [18]. Moreover, compared to enzymes derived from animals and plants, microbial extracellular enzymes offer advantages in industrial production, including simpler manufacturing processes, lower costs, high stability, and specificity [19,20,21]. Therefore, their application in fields such as medicine, agriculture, food, and biofuels holds promising potential [22]. Hawar’s research revealed that endophytic fungi from Ziziphus spina-christi (Nabq) leaves, notably Aspergillus niger and Cladosporium sp., produce lipase and laccase [13]. Furthermore, the Cladosporium sp. exhibited high production of protease and pectinase. These enzymatic capabilities highlight the significant potential of endophytic fungi in biocontrol and clinical microbiology therapeutics.
Euchresta tubulosa Dunn, commonly known as Yapianqi, E-dougen, or Hudoulian, is a plant of the genus Euchresta. Recognized as a “prestigious ethnic medicine” in Tujia tradition, this plant is noted in Hunan Medicinal Flora for the use of its roots in treating gastric pain, abdominal discomfort, and sore throat [23]. E. tubulosa contains bioactive compounds including alkaloids, flavonoids, and terpenoids, which exhibit antitumor, antiviral, and anti-inflammatory properties, indicating significant pharmaceutical potential [24,25]. It is sporadically distributed in mid-subtropical evergreen broad-leaved forests along valleys and streams at elevations between 600 and 900 m in certain regions of Hunan, Guangdong, Guangxi, and Sichuan. This species thrives in shaded, moist, and humus-rich soils [18]. Due to its specific habitat requirements, large-scale cultivation is unfeasible. Additionally, the plant exhibits strong self-incompatibility, resulting in low natural fruiting and seed germination rates. As a consequence, its natural distribution has become extremely scarce, pushing it to the brink of extinction. It has been listed as a National Grade II Protected Plant [26]. The scarcity of medicinal material severely constrains research on its active constituents and pharmacological mechanisms, representing a major limiting factor for further exploitation.
Current research on Euchresta tubulosa Dunn primarily focuses on its chemical constituents, pharmacological activities, and clinical applications [27,28,29]. Internationally, reports on endophytic fungi isolated from E. tubulosa remain scarce. To better explore its endophytic microbial resources and discover structurally novel bioactive secondary metabolites and highly active enzymes, this study focused on the isolation and identification of endophytic fungi from E. tubulosa. Three distinctive strains were further investigated for their extracellular enzymes and secondary metabolites. This research aims to obtain functionally robust microbial isolates, thereby providing a theoretical foundation for future applications in biological control, novel drug development, and fermentation industries.
2. Materials and Methods
2.1. Biological Materials
Fresh Euchresta tubulosa Dunn plants were collected from Tianping Mountain (29°40′ N, 109°40′ E), Sangzhi County, Hunan Province, China on 13 July 2023 and 13 October 2023. Taxonomic identification was performed by Prof. Aiwen Dong. The plant materials were refrigerated at 4 °C and used for the isolation of endophytic fungi within 72 h. The strains used in the antimicrobial assay were Escherichia coli CMCC(B) 44102, Staphylococcus aureus CMCC(B) 26003, and Candida albicans CMCC(F) 98001, obtained from the Shanghai Bioresource Collection Center.
2.2. Main Reagents and Drugs
All chemicals and reagents were purchased from commercial suppliers: ethyl acetate, ethanol, and methanol from Concord Technology Co., Ltd. (Tianjin, China); sodium hypochlorite and anhydrous sodium carbonate from Tianjin Guangfu Technology Development Co., Ltd. (Tianjin, China); trichloroacetic acid, soluble starch, Congo red, and carboxymethylcellulose sodium from Tianjin Comio Chemical Reagent Co., Ltd. (Tianjin, China); casein, L-tyrosine, and Folin–Ciocalteu reagent from Ron Reagent Co., Ltd. (Shanghai, China); DNS reagent from Shanghai Yuanye Bio-Technology Co., Ltd. (Shanghai, China); Penicillin G sodium salt and kanamycin from Fuzhou Feijing Biotechnology Co., Ltd. (Fuzhou, China); lactophenol cotton blue staining solution from Qingdao Haibo Biotechnology Co., Ltd. (Qingdao, China).
Potato Dextrose Agar (PDA) and Potato Dextrose Broth (PDB) for fungal isolation and cultivation were obtained from Hope Bio-Technology Co., Ltd. (Qingdao, China).
2.3. Main Instruments
The following instruments were utilized: constant-temperature water bath (DZKW-D-6, Beijing Yongguangming Medical Instrument, Beijing, China); full-wavelength microplate reader (K6600A, Beijing Kaiwo Technology, Beijing, China); vertical autoclave (YXQ-LS-50SII, Shanghai Boxun Medical Equipment, Shanghai, China); biological microscope (PX43 FS6, Motic, Xiamen, China); constant-temperature shaker incubator (ZQPL-200, Tianjin Laiboterui Instrument, Tianjin, China); circulating water vacuum pump (SHB-III, Zhengzhou Great Wall Sci. & Tech., Zhengzhou, China); UV–Vis spectrophotometer (UV2400, Shanghai Shunyu Hengping Scientific Instrument, Shanghai, China); ultrapure water system (SMART-N30VF, Shanghai Kanglei Analytical Instrument, Shanghai, China); analytical balance (FA2004, Shanghai Shunyu Hengping Scientific Instrument, Shanghai, China); centrifuge (H1850R, Hunan Xiangyi Laboratory Instrument, Changsha, China); and rotary evaporator (RE-2000A, Shanghai Yarong Biochemical Instrument, Shanghai, China); biochemical incubator (BSP-400, Shanghai Boxun Medical Biological Instrument, Shanghai, China).
2.4. Preparation of Medium
Preparation of Solid Media
The media used in this study included both solid and liquid formulations. Solid media comprised PDA medium, inorganic salt starch medium, sodium carboxymethylcellulose (CMC-Na) medium, skim milk medium, and nutrient agar. Liquid media included PDB liquid medium, CMC-Na liquid medium, and inorganic salt starch and skim milk mixed liquid medium. Detailed compositions, preparation procedures, and sterilization conditions for all media are provided in Table S1.
2.5. Isolation of Endophytic Fungi from Euchresta tubulosa
Endophytic fungi isolated from Euchresta tubulosa fresh roots, stems, and leaves were placed firmly onto PDA (Potato Dextrose Agar) using the conventional tissue isolation method [30] and incubated for 3–7 days at 28 °C: hyphal growth around the tissue pieces was observed and recorded.
2.6. Identification and of Endophytic Fungi from Euchresta tubulosa
Colony Morphology Identification: A small amount of endophytic fungal strains preserved on PDA slant medium was streaked onto plate medium for activation (1–2 passages). Then, a 0.5 mm diameter fungal plug was inoculated onto PDA medium and incubated at 28 °C for 7–14 days. Growth characteristics including colony size, color, texture, surface patterns, margin morphology, presence of exudates, and odor were observed and recorded.
Morphological Characterization of Hyphae and Spores: Microscopic slides were prepared using the adhesive tape method [31], and the morphological examination of hyphae, conidiophores, spores, and other structures under a phase-contrast inverted microscope (model PX43 FS6).
The purified strains cultured of Identification were carried out at Chengdu Luoning Biotechnology Co., Ltd. Polymerase chain reaction (PCR) was used to amplify the internal transcribed spacer (ITS) regions 1 and 2 with the forward ITS1 5′-TCCGTAGGTGAACCTGCGG-3′and reverse ITS4 5′-TCCTCCGCTTATTGATATGC-3′PCR reaction system (50 μL): DNA template (20 ng/μL) 2 μL, forward and reverse primers (10 μmol/L) 2.5 μL each, 2 × PCR Bestaq Master Mix with dye 25 μL, ddH2O 18 μL. PCR reaction conditions: 95 °C for 5 min; 30 cycles of 94 °C for 45 s, 50 °C for 30 s, 72 °C for 45 s; 72 °C for 1 min. The PCR products were detected by 1% agarose gel electrophoresis. Concurrently, strain identification was performed by combining morphological characterization with molecular biological analysis the National Center for Biotechnology Information (NCBI) website (https://blast.ncbi.nlm.nih.gov/Blast.cgi, accessed 8 May 2025). Phylogenetic reconstruction was conducted using MEGA software (version 12.0) employing the Maximum Likelihood (ML) method with 1000 bootstrap replicates to assess branch support. This integrated approach enabled molecular-level identification of endophytic fungi from Euchresta tubulosa and determination of their taxonomic positions [32]. Endophytic fungi from E. tubulosa were taxonomically classified through molecular analysis when two or more strain sequences clustered within a monophyletic clade sharing ≥97.0–98.5% sequence similarity with reference strains at the species level. Classification thresholds were defined as: ≥90% similarity for genus-level identification, ≥85% for family-level, and ≥80% for order-level assignments [33].
2.7. Analysis of Fermentation Product Composition from Endophytic Fungi
Following established protocols [28], prepared the ethyl acetate extracted sample of the strain fermentation product and submitted to Novogene Co., Ltd. (Beijing, China) for compound identification via Liquid Chromatography–Tandem Mass Spectrometry (LC–MS/MS). Metabolites were characterized by comparing mass spectral data against public databases (GNPS, Pub-Chem, etc.).
2.8. Bioactivity Assay of Fermentation Products from Endophytic Fungi
2.8.1. Antimicrobial Activity
Prepare samples with different concentration gradients: Antimicrobial concentration gradient: 20, 10, 8, 6, 4, 2, 1 mg/mL.Antioxidant concentration gradient: 4, 2, 1, 0.8, 0.6, 0.4, 0.2 mg/mL.
Preparation of bacterial suspensions: Cultivate Escherichia coli, Staphylococcus aureus, and Candida albicans separately in beef extract peptone broth and PDB broth. Select colonies with a concentration of approximately 1.0 × 106 cfu/mL as the test bacterial suspensions. Antibacterial Activity Assay: Following the method of Wu Chunlin et al. [34], the critical value at which absorbance does not increase is used as the minimum inhibitory concentration (MIC) of the fermentation product.
2.8.2. Antioxidant Activity
Following the method described by Yan Jun et al. [35], prepare solutions of Vc and fermentation products at different concentrations. Measure the absorbance values of A1, A2, and A3 at 510 nm, respectively. Calculate the clearance rate using the following formula:
where: A1 is the absorbance value without the fermentation product solution; A2 is the absorbance value with the fermentation product solution added; A3 is the absorbance value without salicylic acid.
D = [(A1 − A2)/A3] × 100
Prepare the ABTS working solution according to the method described by Han Luoxia et al. [36] Take 100 μL each of the test solution, anhydrous ethanol, and Vc, and measure the absorbance values at 734 nm for OD1, OD2, and OD3, respectively. Calculate the scavenging rate using the following formula:
D = [1 − (OD1 − OD2)/OD0] × 100
2.9. Evaluation of Extracellular Enzyme Activity in Endophytic Fungi Using the Clear Zone Method
Extracellular enzyme activity of endophytic fungi was evaluated using the clear zone method according to Shubha Jagannath et al. [37]. The diameters of the halo zones and fungal colonies were measured daily under specific staining conditions for each substrate:
- •
- For skim milk agar, both the halo zones and colony diameters were measured directly.
- •
- For starch-containing minimal salt agar, the plates were stained with dilute iodine solution to visualize halo zones, after which the diameters of the halo zones and fungal colonies were recorded.
- •
- For CMC-Na agar, the plates were stained with Congo red reagent for 15 min, followed by destaining with NaCl solution for another 15 min. The presence of light-yellow halo zones was observed, and the diameters of the halo zones and fungal colonies were measured.
The enzyme activity zone (ZA) was calculated to assess the enzyme-producing capability of the endophytic fungi.
Enzyme activity was quantified using the enzymatic zone of activity (ZA) index [11], calculated as
Enzymatic zone of activity (ZA) = Colony Diameter/Clear Zone Diameter
2.10. Extracellular Enzyme Activity Assays
2.10.1. Preparation of Crude Enzyme Extracts
Crude enzyme extracts were prepared by aseptically transferring five 7-mm mycelial plugs from the actively growing margin of fungal colonies into 125 mL of liquid medium. Cultures were incubated in a rotary shaker (120 rpm) at 28 °C for 2–6 days. Following daily harvests from day 2 to day 6, the cultures underwent centrifugation at 10,000 r/min for 25 min at 4 °C, and the resultant supernatants served as crude enzyme extracts.
2.10.2. Protease Activity Quantification via Folin–Ciocalteu Assay
A fresh casein substrate solution (200 μg/mL) was prepared following the method of Hu [30]. A series of tyrosine standard solutions with concentrations of 0, 10, 20, 30, 40, 50, 60, and 70 μg/mL was prepared by diluting the stock solution (200 μg/mL) with de-ionized water. The optical density (OD) values of these solutions were then measured according to the method described below. A standard curve was generated by plotting the OD values (ordinate) against the mass concentrations (abscissa), and a linear regression equation was established.
For enzyme activity determination, aliquots of casein solution and crude enzyme extract were pre-incubated separately at 40 °C for 10 min, followed by transfer of 1 mL reaction mixtures to 10-mL tubes. After 15 min incubation at 40 °C, reactions were terminated with 2 mL of 0.4 M trichloroacetic acid (TCA). Following 5 min sedimentation, samples were centrifuged at 3000× g for 10 min. Subsequently, 1 mL supernatant was combined with 5 mL 0.4 M Na2CO3 and 1 mL Folin–Ciocalteu (FC) reagent, incubated at 40 °C for 20 min, and absorbance measured at 680 nm. Control reactions received TCA prior to casein addition. Protease activity (U/mL) was calculated as:
where K denotes tyrosine equivalence (μg) per unit absorbance (derived from standard curve), 4 represents total reaction volume (mL), N indicates dilution factor, 10 signifies reaction time (min), and V is crude enzyme volume (mL). One unit of protease activity was defined as the enzyme quantity liberating 1 μg tyrosine per minute under specified conditions.
Activity = (OD680 × K × 4 × N)/(10 × V)
2.10.3. Quantification of Amylase and Carboxymethyl Cellulase (CMCase) Activities via DNS Assay
Fresh substrate solutions were prepared daily: 1% (w/v) starch by suspending 1.00 g in 50 mL boiling water with continuous stirring for 10 min before dilution to 100 mL; 1% (w/v) carboxymethyl cellulose sodium (CMC-Na) by dispersing 1.00 g onto boiling water followed by hydration and dilution to 100 mL.
A glucose stock solution (50 mg/mL) was prepared in deionized water. This stock was then diluted with deionized water to obtain a series of standard solutions with concentrations of 0, 100, 200, 300, 400, 500, 600, and 700 μg/mL. Each solution was then treated according to the procedure described below, and the optical density (OD) was measured.
A linear regression equation was derived by plotting the mass concentration (as abscissa) against the OD value (as ordinate). Detection method: Refer to the method of Hu [38], For enzymatic analysis, crude enzyme extract, phosphate buffer (0.05 mol/L, pH 7.0), and 1% substrate were pre-incubated at 37 °C for 10 min. Reaction mixtures containing 1 mL enzyme extract, 2 mL buffer, and 0.5 mL substrate were incubated at 37 °C for 10 min, terminated with 1 mL of 0.4 mol/L NaOH, supplemented with 100 μL DNS reagent, and heated in a boiling water bath for 6 min. After cooling and 10–20 min equilibration, absorbance was measured at 520 nm. Control reactions received NaOH prior to substrate addition. Carboxymethyl cellulase (CMCase) activity was determined identically using CMC-Na substrate. One enzyme unit (U) was defined as the amount catalyzing liberation of 1 μmol reducing sugar per minute under assay conditions. Extracellular enzyme activity (U/mL) was calculated as:
where c represents reducing sugar concentration (μmol/mL, glucose-equivalent from standard curve), V2 denotes total reaction volume, t signifies reaction time (10 min), and V1 is crude enzyme volume.
X = (c × V1)/(t × V2)
2.10.4. Standard Curves
A calibration curve was established by quantifying glucose standard solutions using the DNS assay, with absorbance measurements taken at 520 nm. The resulting calibration equation was y = 1.3143x − 0.00051 (R2 = 0.9986), as shown in Figure S17A. Similarly, a standard curve for tyrosine was generated using the Folin method, with absorbance measured at 680 nm, yielding the regression equation y = 9.84247x − 0.00608 (R2 = 0.99956) (Figure S17B).
3. Results
3.1. Isolation and Identification of Endophytic Fungi in the Euchresta tubulosa
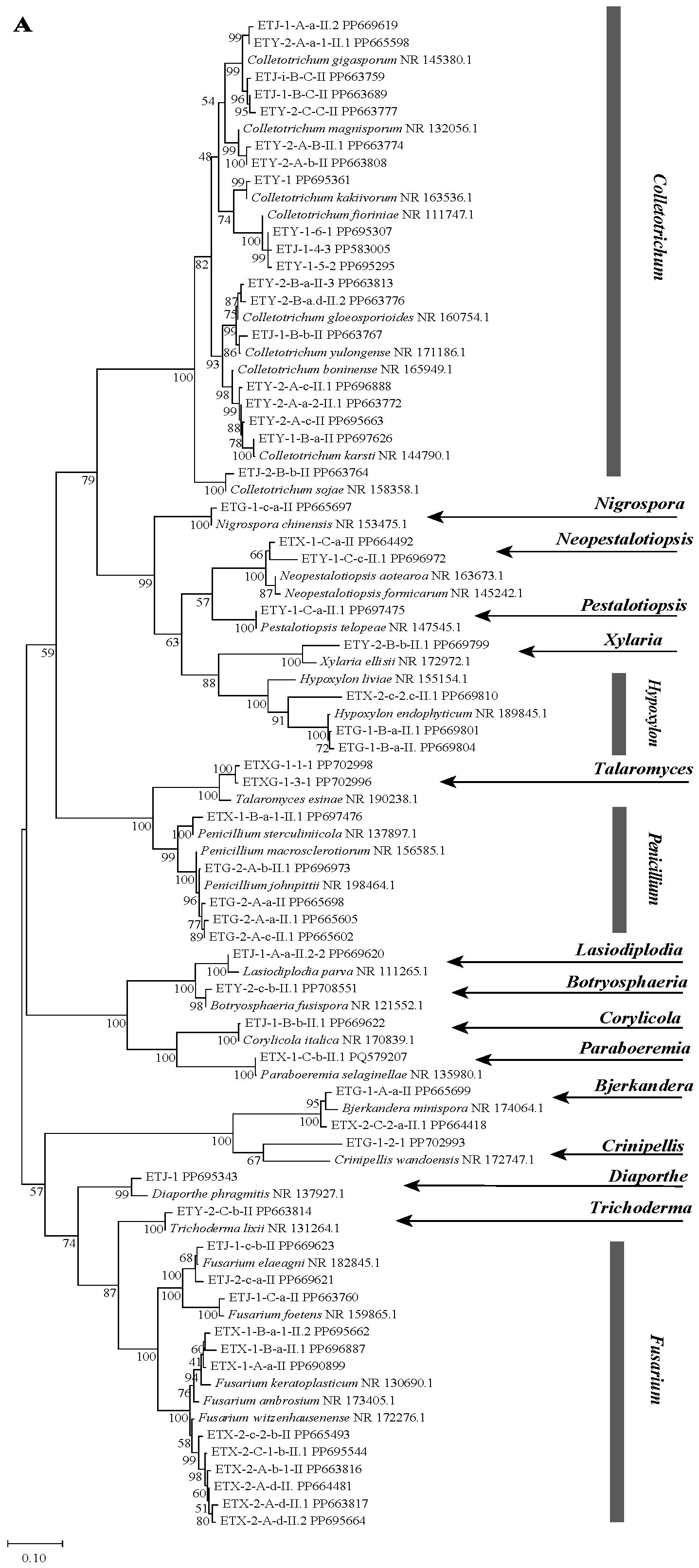
A total of 55 endophytic fungal strains were isolated from fresh Euchresta tubulosa plants. Based on preliminary rRNA-ITS sequence analysis, these strains were classified into 33 species and 17 genera, which were distributed across the following genera: Fusarium, Penicillium, Pestalotiopsis, Xylaria, Trichoderma, Bjerkandera, Neopestalotiopsis, Botryosphaeria, Nigrospora, Lasiodiplodia, Corylicola, Hypoxylon, Talaromyces, Crinipellis, Diaporthe, Paraboeremia and Colletotrichum. The dominant genera were Colletotrichum (34.5%), Fusarium (21.8%), and Penicillium (9.1%). The obtained ITS sequences were deposited in the GenBank database to acquire accession numbers. The most similar ITS region gene sequences were identified using NCBI-BLAST homology comparison, and a phylogenetic tree was constructed (Figure 1A).
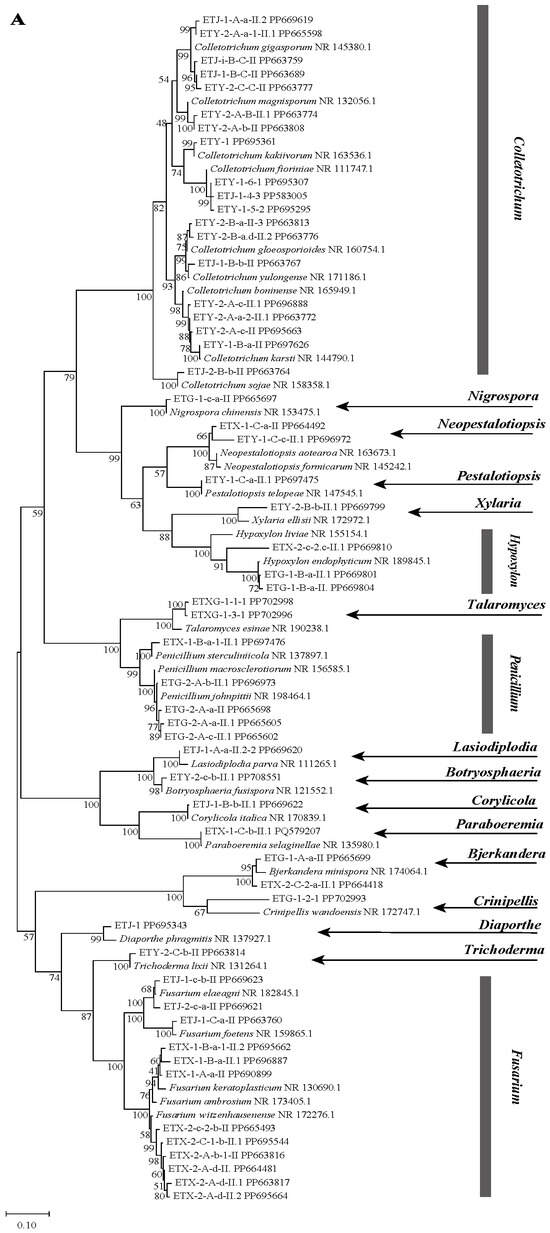
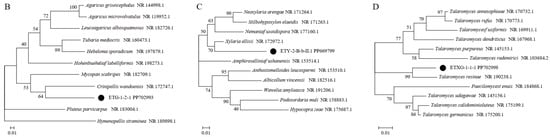
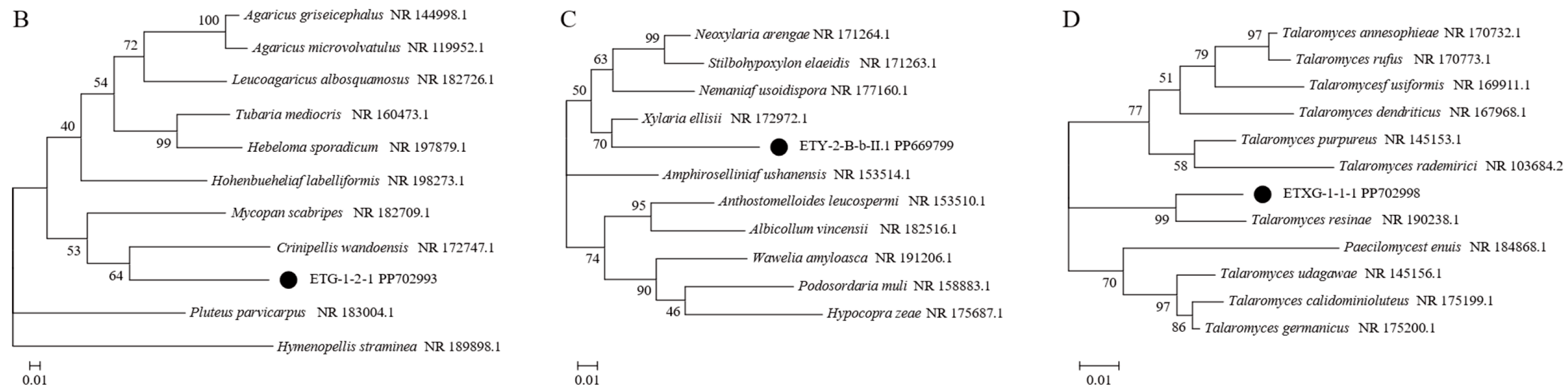
Figure 1.
Phylogenetic trees based on ITS sequences of endophytic fungi from Euchresta tubulosa (A). Trees for novel taxa: ETG-1-2-1 ((B), novel genus), ETY-2-B-b-II.1 ((C), novel species in Xylaria), and ETXG-1-1-1 ((D), novel species in Talaromyces). Numbers at branches indicate bootstrap values (>50% shown). Scale bars represent sequence divergence. Branch lengths correspond to genetic distance. Letters/numbers following strain names denote GenBank accession numbers.
NCBI-BLAST homology analysis revealed that strains ETG-1-2-1, ETY-2-B-b-II.1, ETXG-1-1-1 and ETXG-1-3-1 showed ≤97% sequence similarity to their closest relatives, indicating that these strains may represent novel taxa at the genus or species level [33]. Among them, the NCBI-BLAST homology alignment results showed that the sequence homology between strain ETG-1-2-1 and its most similar strain was not higher than 90%, leading to its identification as a novel genus. Strains ETY-2-B-b-II.1, ETXG-1-1-1, and ETXG-1-3-1 exhibited homology not exceeding 97% compared to their closest relatives, and were consequently identified as novel species. Notably, strains ETXG-1-1-1 and ETXG-1-3-1 matched the same reference sequence in the NCBI-BLAST alignment (Figures S14 and S15), and shared 99.46% sequence identity (Figure S16). Furthermore, Figure 1A demonstrated that the genetic distance between these two strains is minimal, identifying them as sister strains of a novel species within the genus Talaromyces.
Phylogenetic analysis based on ITS sequences placed all three endophytes (ETG-1-2-1, ETY-2-B-b-II.1, and ETXG-1-1-1) within distinct, well-supported clades (bootstrap values > 50%), as shown in Figure 1B–D. Specifically, ETG-1-2-1 was most closely related to Crinipellis wandoensis (NR 172747.1) with a bootstrap value of 64% and a BLAST alignment similarity of 85.28%, and was identified as a new genus of the family Marasmiaceae., It was named “Neocrinipellis albifloccum ETG-1-2-1” and deposited at the China General Microbiological Culture Collection Center with the accession number CGMCC No. 41962 [39]. Strain ETY-2-B-b-II.1 was most closely related to Xylaria ellisii (NR 172972.1) with a bootstrap value of 70% and a BLAST alignment similarity of 92.16%, and was identified as a new species of the genus Xylaria., It was named “Xylaria cinereonitens ETY-2-B-b-II.1” (CGMCC No. 42144) [40]. Strain ETXG-1-1-1 was most closely related to Talaromyces resinae (NR 190238.1) with a bootstrap value of 99% and exhibited 95.86% similarity in BLAST alignment, leading to its identification as a novel species within the genus Talaromyces. The strain was named “Talaromyces polymorphus ETXG-1-1-1” (CGMCC No. 41927) [41]. Furthermore, the three endophytic fungal strains isolated from Euchresta tubulosa were located in distinct clades in Figure 1A, indicating a distant phylogenetic relationship among them. The fungal sequencing results were submitted to the GenBank database, and the corresponding GenBank accession numbers for the tested strains were obtained, as summarized in Table 1.

Table 1.
Results of sequence comparison of ITS segments of three endophytic fungi from the Euchresta tubulosa.
3.2. Morphological Characterization of Selected Endophytic Fungi from Euchresta tubulosa
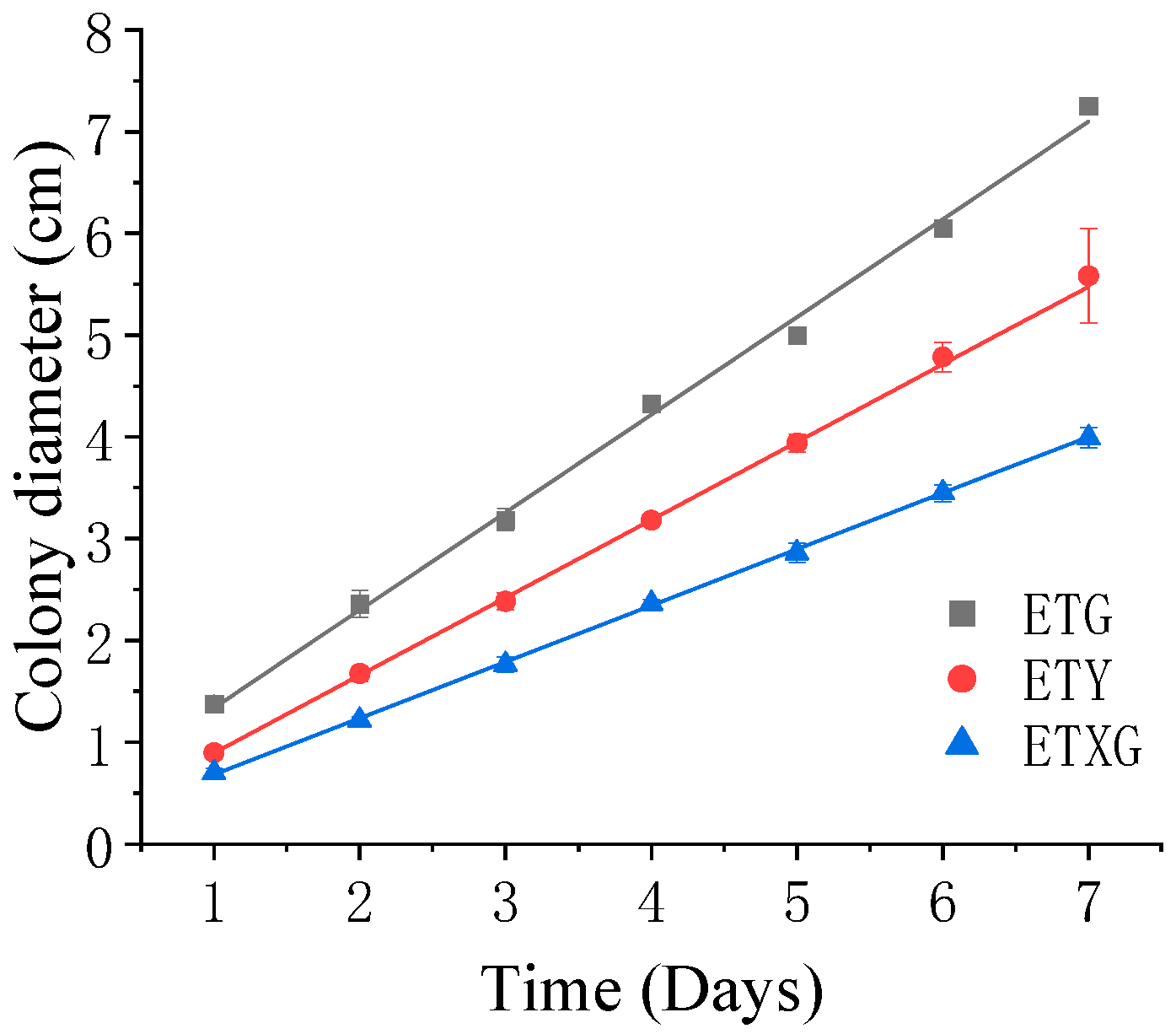
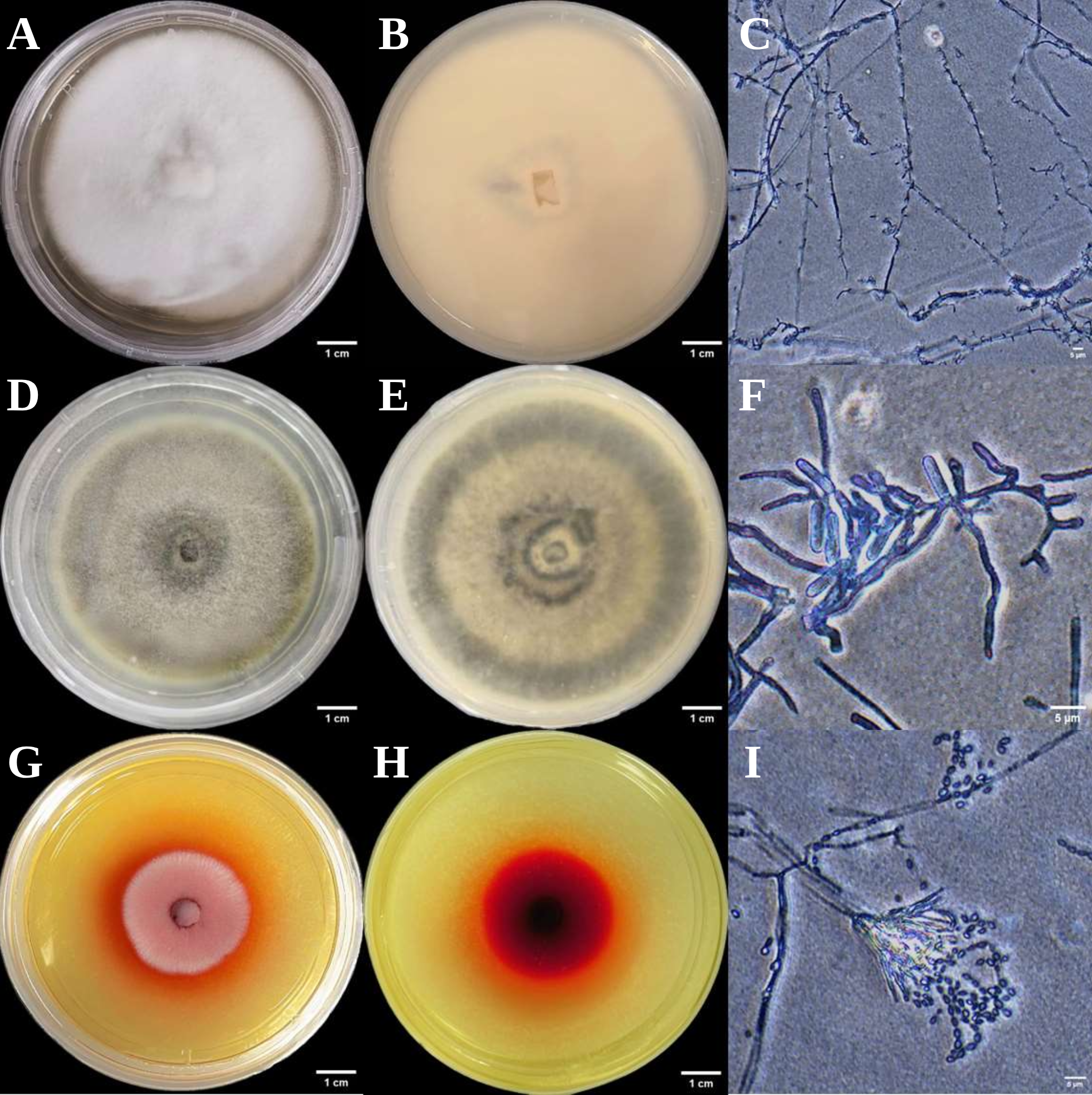
The three endophytic fungi isolated from Euchresta tubulosa were morphologically identified based on their morphological characteristics (Table 2), and their growth on PDA medium was documented (Figure 2 and Figure 3). The strain ETG-1-2-1 grew radially from the inoculation point to form a circular colony with an even margin on PDA medium. It exhibited white, dense, and villous surface mycelia, secreted no soluble pigments, and reached a diameter of 7.25 cm at 7 d. The average radial growth rate was calculated to be (9.61 ± 0.26) mm/day. On PDA medium, colony ETY-2-B-b-II.1 was initially white and turned black upon maturation, exhibiting a thin velvety texture and a relatively slow growth rate. After 7 days of incubation, the colony reached a diameter of approximately 5.58 cm, with a growth rate of (7.64 ± 0.06) mm/day. The edge of the colony was entire, with a grayish-white sporulating structure observed in the center. Concentric rings were visible on the reverse side. Microscopic examination revealed slender and uniform hyphae with distinct branching. Conidia were regularly oval-shaped, partly attached to the tips of hyphal branches and partly dispersed as individual units. On PDA medium, ETXG-1-1-1 formed colonies with dense, velvety mycelia, exhibiting a flat center and radial wrinkles. The colony displayed a white peripheral zone and a red central region. Mature spores were abundant, and easily detached. The reverse side showed a radial dark red pigmentation that was most intense in the center and faded gradually toward the margin. The strain demonstrated a slow growth rate, reaching a diameter of approximately 3.99 cm after 7 days of incubation at 28 °C, with an average radial growth rate of 5.53 ± 0.06 mm/d. Hyphal examination under inverted phase-contrast microscope revealed septate and elongated filamentous structures. Conidia were solitary or aggregated in sporodochial clusters, varying in length. The conidia have a smooth surface, round or nearly circular, arranged in chains or scattered around the spore stalk.

Table 2.
Colonial morphological characteristics of three endophytic fungi isolated from Euchresta tubulosa.
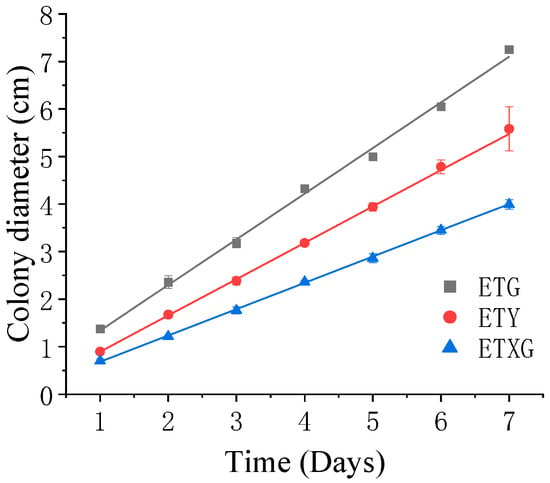
Figure 2.
Growth dynamics of endophytic fungi from Euchresta tubulosa on PDA medium.
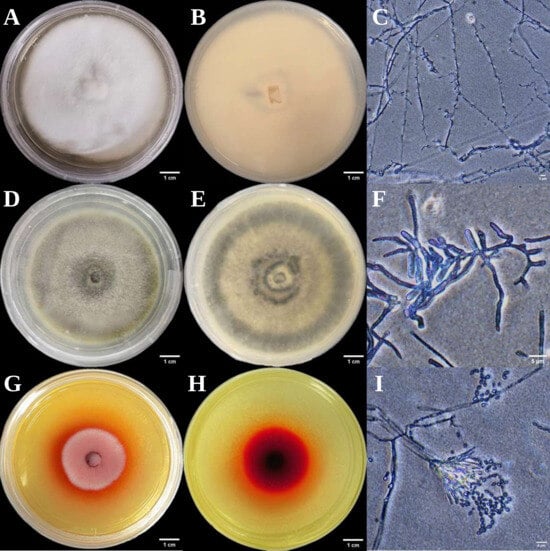
Figure 3.
Colonial and micro-morphological features of three endophytic fungal strains isolated from Euchresta tubulosa. The obverse colony morphology (A), reverse colony morphology (B), and microscopic observation of mycelia (C) of strain ETG-1-2-1; the obverse colony morphology (D), reverse colony morphology (E), and microscopic observation of mycelia (F) of strain ETY-2-B-b-II.1; the obverse colony morphology (G), reverse colony morphology (H), and microscopic observation of mycelia (I) of strain ETXG-1-1-1.
3.3. LC–MS Analysis of Secondary Metabolites from Endophytic Fungi
To investigate their chemical profiles, we analyzed the secondary metabolites of three endophytic fungal strains derived from Euchresta tubulosa using liquid chromatography–mass spectrometry (LC–MS). Total ion chromatograms (TIC) in positive (POS) and negative (NEG) ionization modes (Figure S1) and mass spectra of selected metabolites (Figure S2) were acquired. Metabolite annotation was performed by matching against the mzCloud, mzVault, and MassList databases.
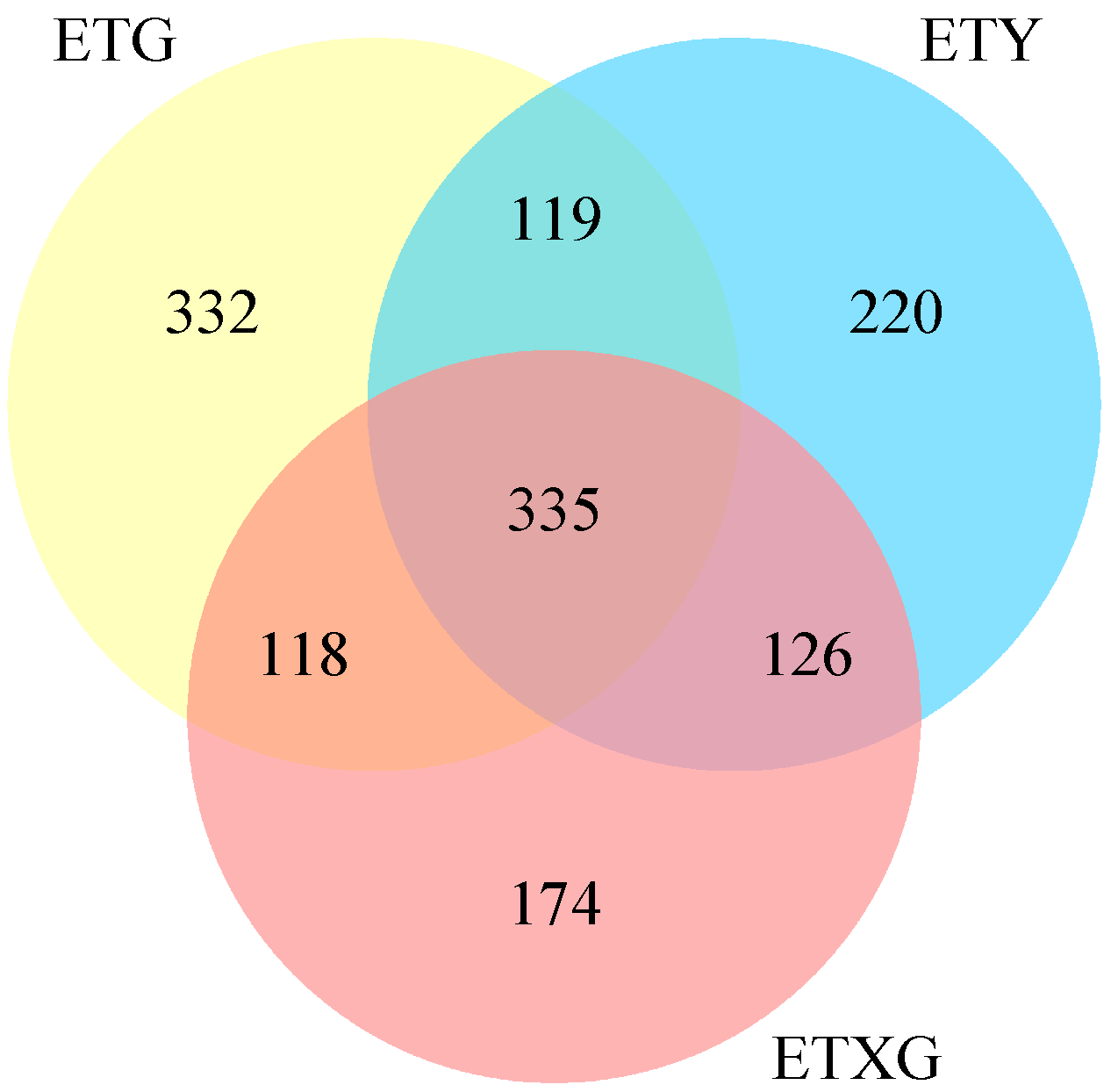
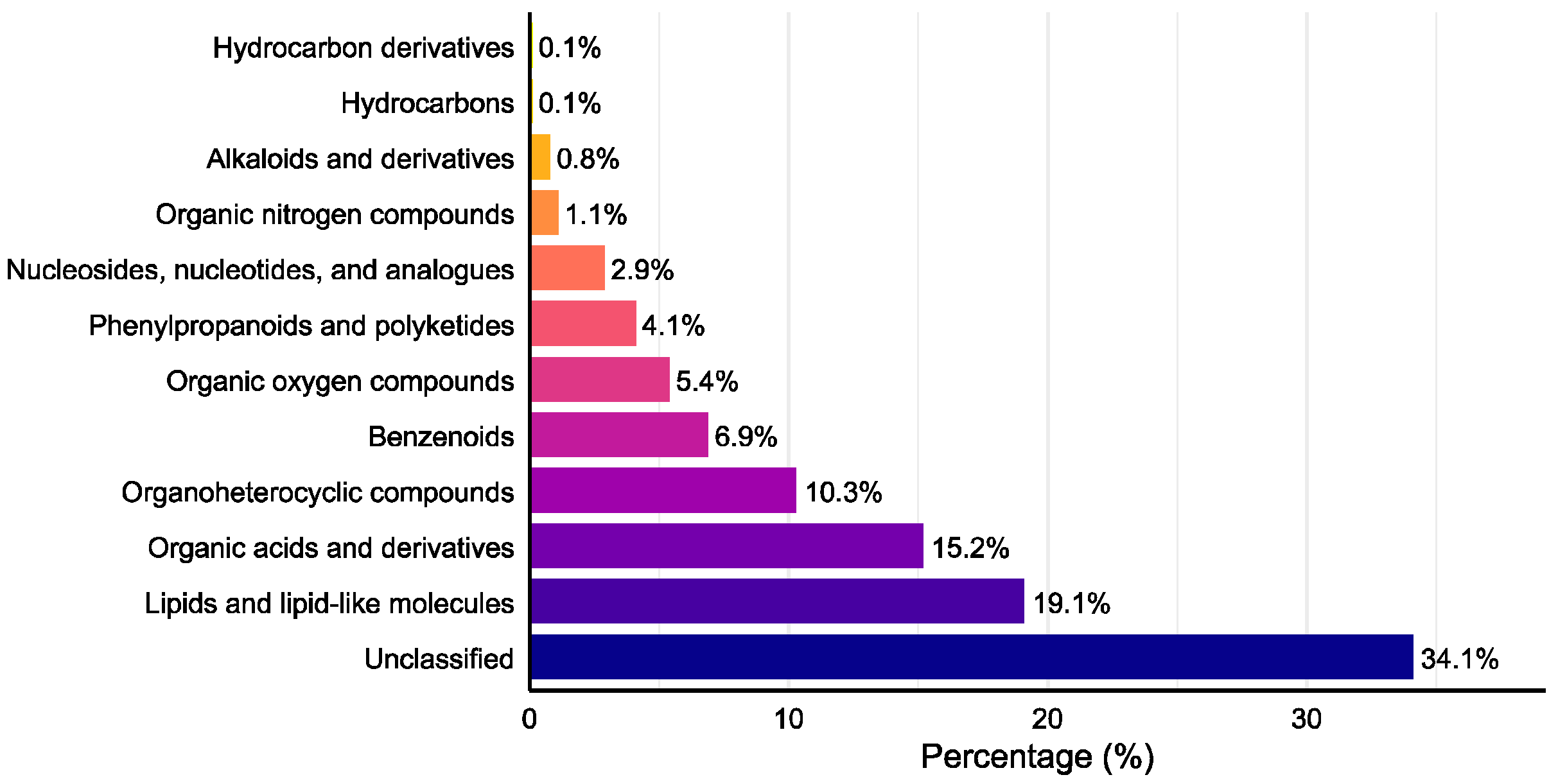
Venn diagram analysis (Figure 4) revealed 1424 total metabolites, with 698 shared compounds (49.0%) across all three strains. SuperClass classification (Figure 5) demonstrated that lipids and lipid-like molecules, organic acids and derivatives, organoheterocyclic compounds, and benzenoids represented the most abundant classes after unclassified metabolites. Additionally, we identified 11 alkaloids, 18 flavonoids, 26 terpenoids, and 59 steroids.
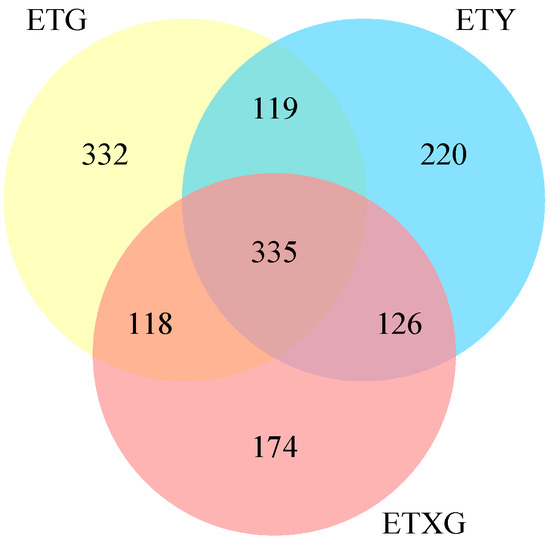
Figure 4.
Venn diagram of secondary metabolites from three endophytic fungal strains of Euchresta tubulosa.
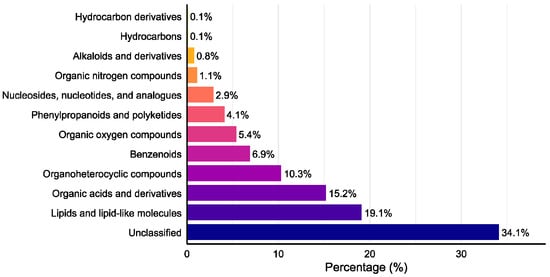
Figure 5.
SuperClass distribution of detected secondary metabolites.
The partially active components from strains were organized into a table, where the strains were denoted as ETG, ETY, and ETXG, representing ETG-1-2, ETY-2-B-b-II.1, and ETXG-1-1-1, respectively (Table 3). Meanwhile, the mass spectra of these compounds are provided in the Supplementary Material, Figures S3–S11. These active components included reserpine, known for its significant antihypertensive, tranquilizing, and antipsychotic effects; trigonelline, reported to exhibit activities such as lowering blood glucose and improving blood lipids; and oxymatrine, which possesses anti-inflammatory, anti-fibrotic, and anti-tumor properties [22]. Notably, these classes of compounds were also detected in preliminary analyses of Euchresta tubulosa plant samples conducted by our research group using LC–MS/MS technology. Li Weixin [25] identified oxymatrine in the stems of E. tubulosa through spectroscopic methods such as NMR and MS. Additionally, Lei Jiaxin performed pseudo-targeted metabolomics analysis on E. tubulosa plants and detected 145 flavonoid compounds, among which glycitein, daidzein, naringenin, formononetin, genistein, kaempferol, sakuranetin, and hesperetin were also present in the secondary metabolites of the endophytic fungi studied here. These findings indicate similarities between the secondary metabolites of these three endophytic fungal strains and the active components of E. tubulosa, suggesting a potential association with the growth process of E. tubulosa.

Table 3.
Selected metabolites detected by LC–MS/MS in endophytic fungi.
3.4. Bioactivity Assay of Fermentation Products from Endophytic Fungi
3.4.1. Antimicrobial Activity Assay
The minimum inhibitory concentrations (MICs) of ethyl acetate extracts derived from the fermentation broths of three endophytic fungi ETG-1-2-1, ETXG-1-1-1, and ETY-2-B-b-II.1 isolated from Euchresta tubulosa are summarized in Table 4 against Escherichia coli, Staphylococcus aureus, and Candida albicans. Among the tested extracts, that from strain ETG-1-2-1 exhibited the most potent activity against E. coli, with an MIC of 2.5 mg/mL. All three extracts displayed equivalent inhibitory efficacy against S. aureus, yielding identical MIC values of 2.5 mg/mL. Notably, the ETG-1-2-1-derived extract also demonstrated the lowest MIC (1.25 mg/mL) against C. albicans, indicating superior antifungal activity relative to the other two strains.

Table 4.
Minimum inhibitory concentration (MIC) of fermentation products from endophytic fungi of Euchresta tubulosa on three indicator strains.
3.4.2. Hydroxyl Radical Scavenging Capacity Assay
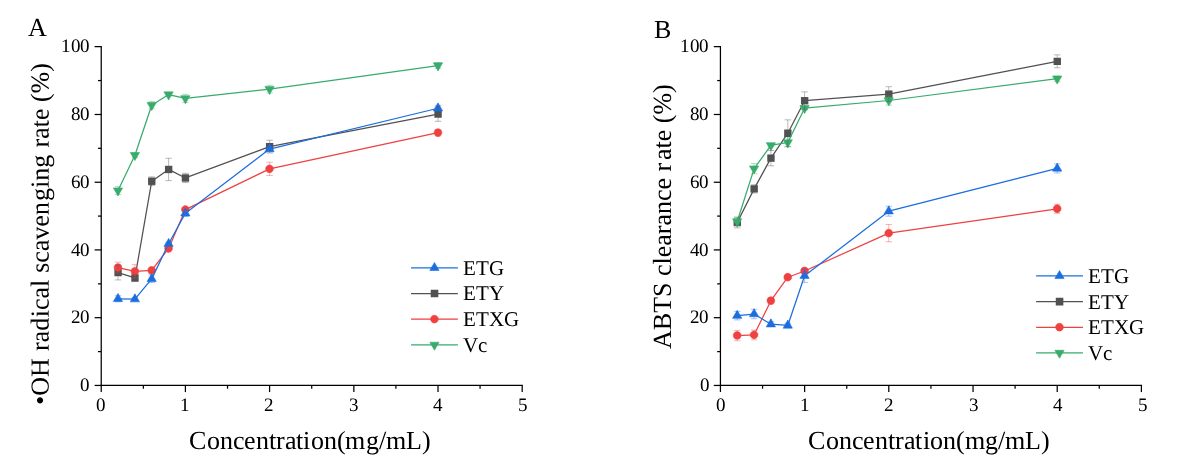
The hydroxyl radical scavenging capacities of ethyl acetate extracts derived from fermentation broths of three endophytic fungi ETG-1-2-1, ETXG-1-1-1, ETY-2-B-b-II.1, and vitamin C are presented in Figure 6A. As illustrated, the scavenging activities of the extracts from ETG-1-2-1, ETXG-1-1-1, and ETY-2-B-b-II.1 increased markedly within the concentration range of 0.4 to 1 mg/mL, followed by a marginal enhancement from 1 to 4 mg/mL. Notably, the ethyl acetate extracts of ETG-1-2-1 and ETY-2-B-b-II.1 exhibited superior scavenging rates of 81.84% and 80.10%, respectively, at 4 mg/mL. Conversely, ETXG-1-1-1 demonstrated a comparatively lower scavenging activity of 74.60%.
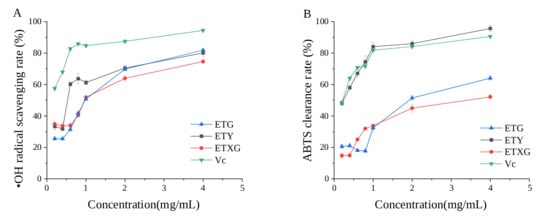
Figure 6.
Scavenging rates of hydroxyl radicals (A) and ABTS radicals (B) by fermentation products from endophytic fungi of Euchresta tubulosa at different concentrations.
3.4.3. ABTS Radical Scavenging Activity Assay
The ABTS radical scavenging capacity of ethyl acetate extracts derived from the fermentation broths of three endophytic fungi ETG-1-2-1, ETXG-1-1-1, and ETY-2-B-b-II.1 as well as that of ascorbic acid (Vc) is presented in Figure 6B. The results demonstrate that the ethyl acetate extract from the fermentation broth of strain ETY-2-B-b-II.1 exhibits exceptionally potent ABTS radical scavenging activity, achieving 95.69% scavenging at a concentration of 4 mg/mL—significantly surpassing the positive control (Vc) under identical experimental conditions. In contrast, the extracts from strains ETG-1-2-1 and ETXG-1-1-1 displayed markedly lower scavenging activities, with values of 52.14% and 64.07%, respectively.
3.5. The Extracellular Enzyme Activity of the Endophytic Fungi from Euchresta tubulosa Was Evaluated by Transparent Circle Method
3.5.1. Protease Activity
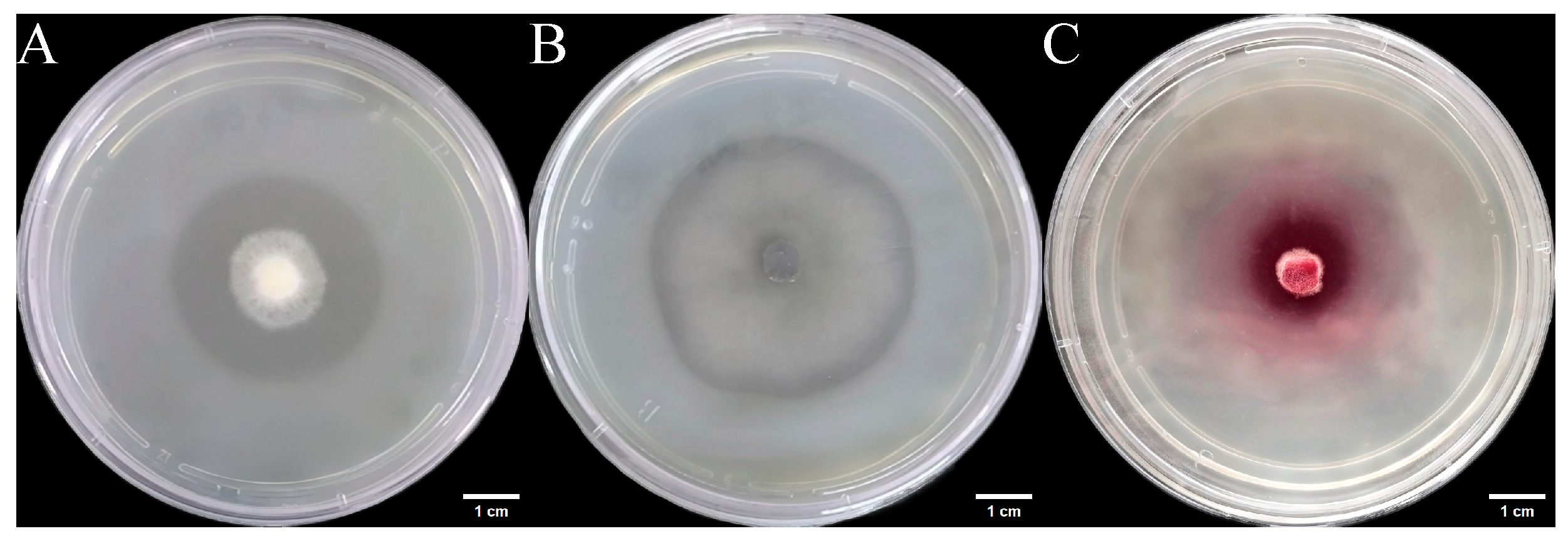
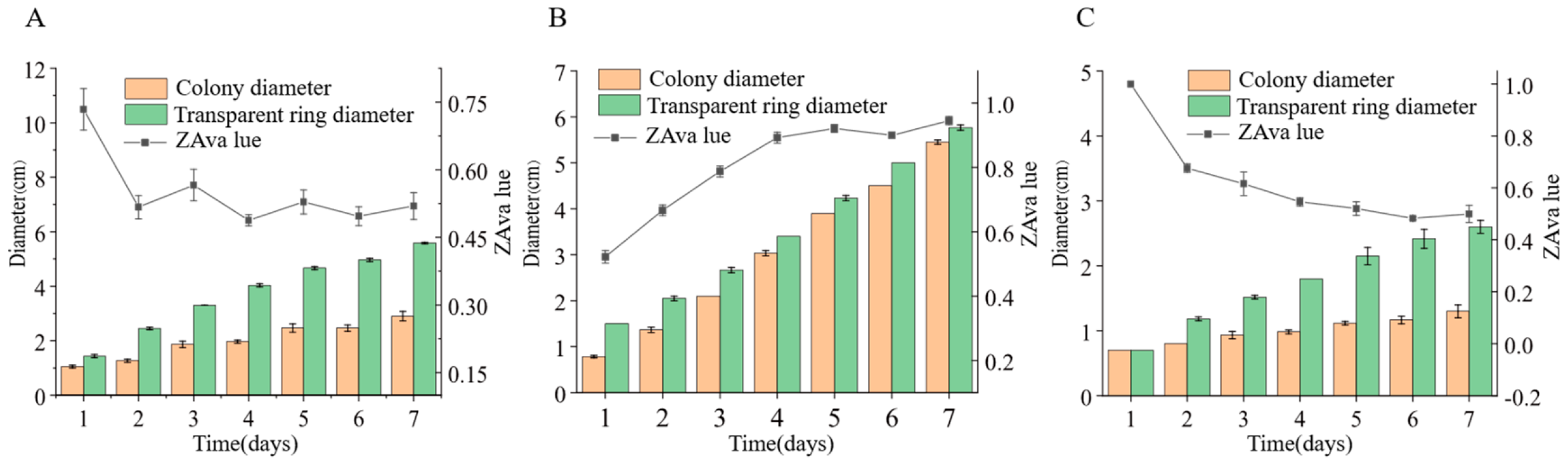
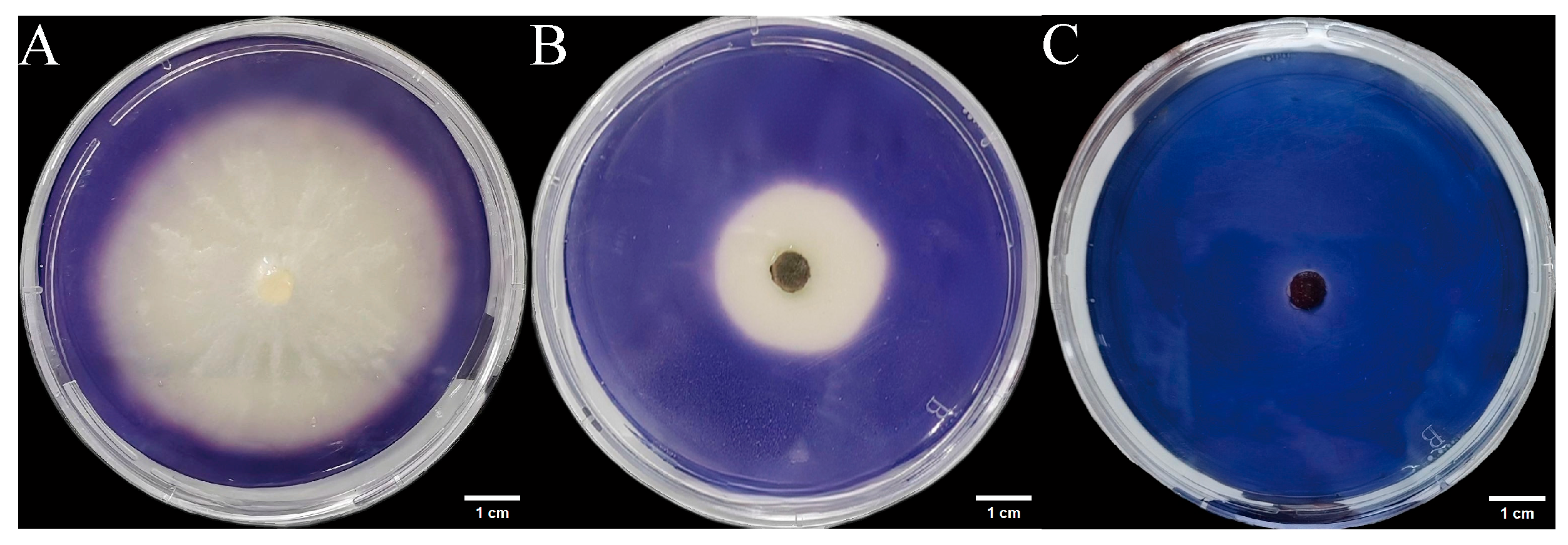
We inoculated three endophytic fungal strains isolated from Euchresta tubulosa onto skim milk agar plates. After 7-day incubation, proteolytic halos were observed (Figure 7). Halo and colony diameters were measured to calculate the enzymatic zone of activity (ZA), with results presented in dual-axis plots (Figure 8). All strains exhibited progressive increases in both colony and halo diameters, though growth kinetics differed temporally: ETG-1-2-1: Demonstrated significant ZA fluctuations during initial growth (days 1–2), stabilizing thereafter with consistently strong protease activity (ZA < 0.69 from day 2 onward). This indicates stable enzyme secretion proportional to biomass accumulation (Figure 8A). ETY-2-B-b-II.1: Displayed increasing ZA (declining enzyme activity) over time. Early-stage halo expansion exceeded colony growth, yielding strong activity (ZA < 0.69) during days 1–2 and moderate activity (ZA = 0.788) at day 3. Late-phase parallel growth of colony and halo suggested nutrient-dependent enzyme production linked to microbial proliferation (Figure 8B). ETXG-1-1-1: Showed decreasing ZA (enhancing enzyme activity) despite limited overall growth. Sustained high protease activity (ZA < 0.69 from day 2) indicated efficient catalytic stability despite low enzyme yield (Figure 8C).
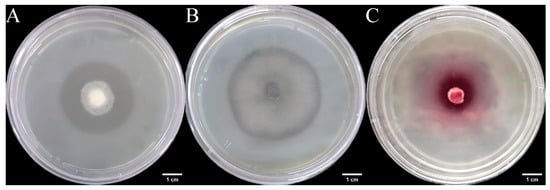
Figure 7.
Detection of protease activity of endophytic fungi from Euchresta tubulosa. The incubation time of strains ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C) was 4 d, 6 d and 5 d, respectively. The transparent area in the middle of the culture medium is the enzyme active zone, and the white area on the outer ring of the culture medium is the inactive zone. In addition, the deep red area in the middle of the culture medium in Figure C represents the enzyme activity zone.
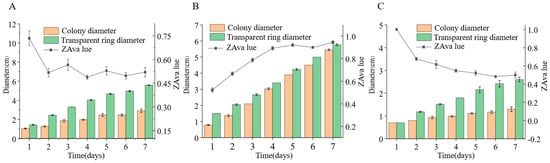
Figure 8.
Comparison of protease activities in endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C) from Euchresta tubulosa.
3.5.2. Amylase Activity
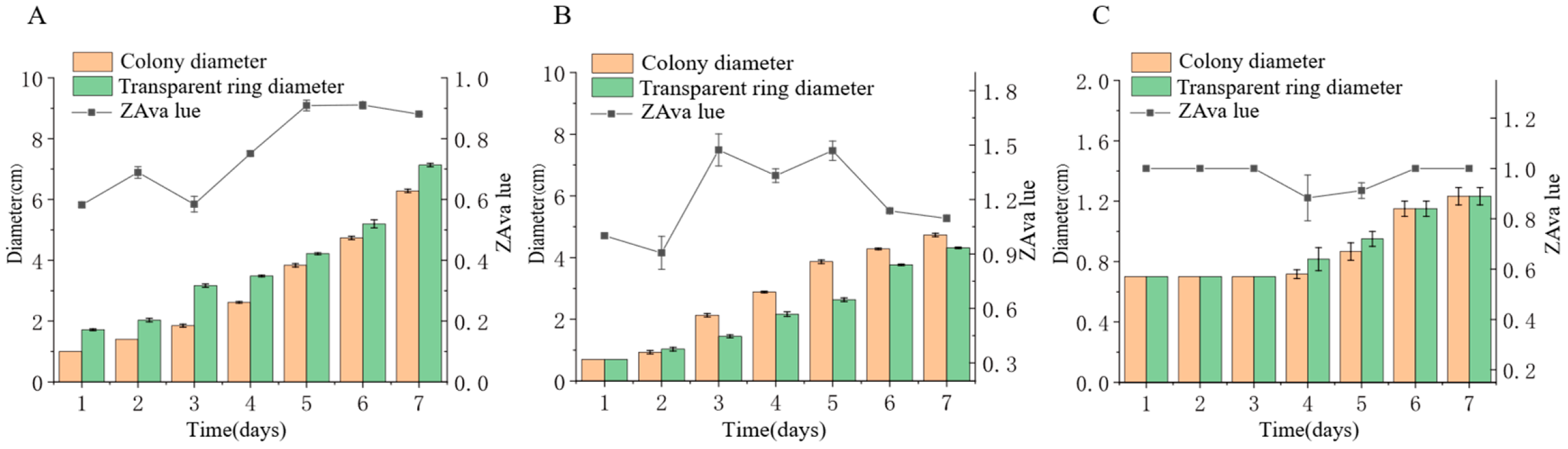
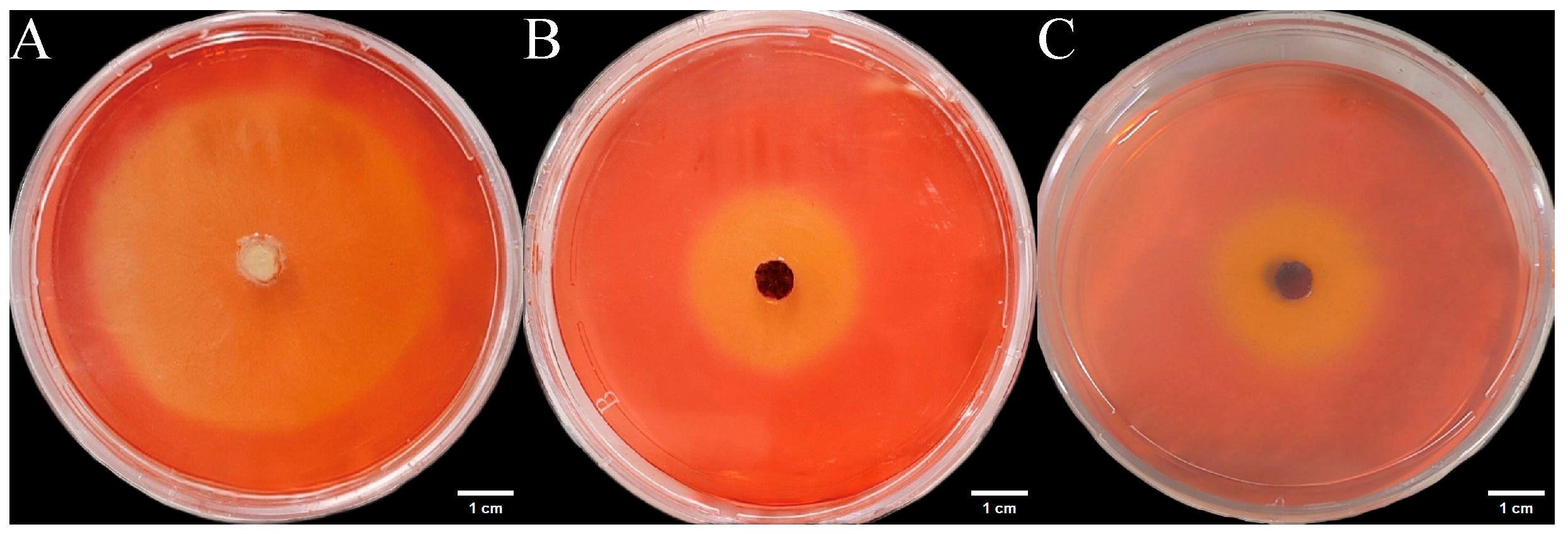
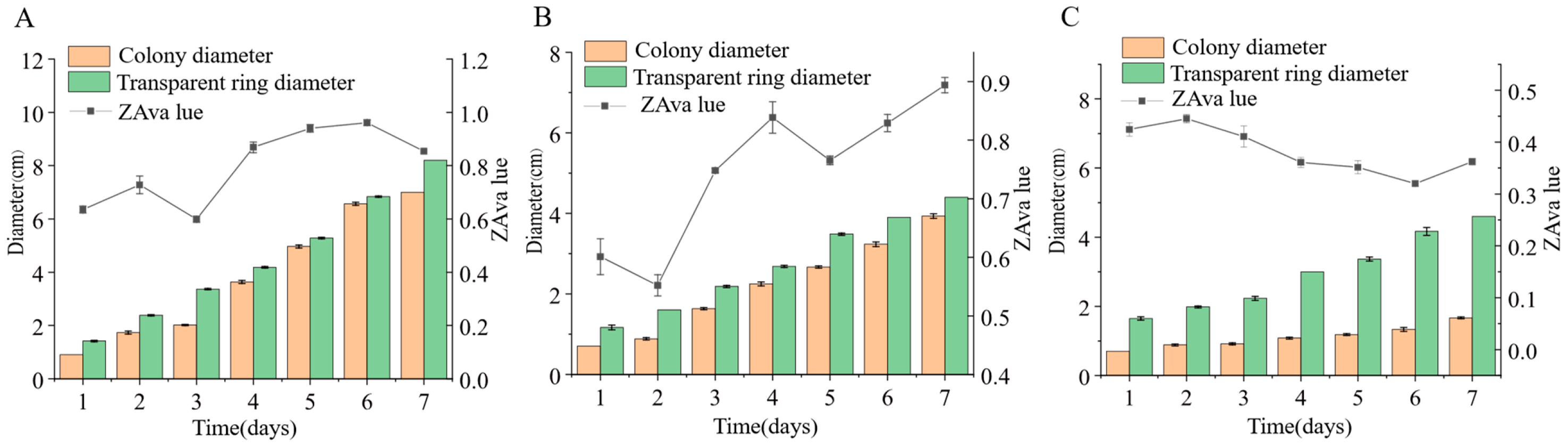
Three endophytic fungal strains isolated from Euchresta tubulosa were inoculated onto mineral-starch agar plates. After 7-day incubation, iodine staining revealed clear zones (Figure 9) Colony and clear zone diameters were measured to calculate hydrolytic coefficients (ZA), with results plotted on dual-axis graphs (Figure 10). Strain ETG-1-2-1 exhibited slow initial colony growth but rapid clear zone expansion during days 1–3 (ZA < 0.69), indicating strong amylase activity and a lag phase for environmental adaptation; subsequent colony acceleration coincided with fluctuating ZA, reflecting stage-dependent variations in exoenzyme efficiency (Figure 10A). Strain ETY-2-B-b-II.1 showed sustained increases in both parameters, but colony diameters exceeded clear zones from day 3 onward (ZA > 1), precluding enzymatic assessment by this metric—potentially due to efficient utilization of hydrolytic products and rapid growth (Figure 10B). Strain ETXG-1-1-1 demonstrated negligible growth until day 4 with minimal expansion (ZA > 0.85 throughout), suggesting limited amylase production, weak activity, and inadequate substrate degradation under these nutritional conditions (Figure 10C).
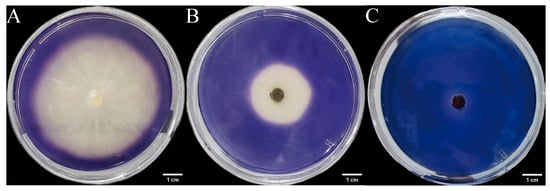
Figure 9.
Amylase activity of endophytic fungi isolated from Euchresta tubulosa. The incubation time of strains ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C) was 7 d, 5 d and 5 d, respectively. The transparent central area of the culture medium represents the enzymatically active zone, while the blue outer ring indicates the inactive zone.
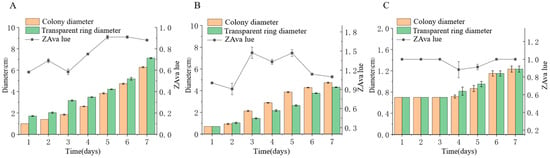
Figure 10.
Comparison of amylase activities in endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C) from Euchresta tubulosa.
3.5.3. Carboxymethyl Cellulase (CMCase) Activity
We inoculated three endophytic fungi derived from Euchresta tubulosa onto carboxymethyl cellulose sodium (CMC-Na) agar plates. After 7 days of culture, Congo red staining and NaCl destaining revealed hydrolysis zones (Figure 11). Colony diameters (CD) and hydrolysis zone diameters (HZD) were measured to calculate enzymatic activity zones (ZA = CD/HZD), with photographic documentation in Figure 12. Strain ETG-1-2-1 exhibited characteristic growth phases: lag, exponential, and stationary. Significant ZA fluctuations occurred during cultivation, particularly in the initial 48 h, indicating physiological instability during adaptation to the novel environment through enzymatic and transport system restructuring. Optimal ZA (0.599) occurred at day 3 (Figure 12A). Strain ETY-2-B-b-II.1 showed progressive increases in both CD and HZD throughout cultivation. However, its ZA profile displayed biphasic fluctuations: an initial decrease followed by recovery. This pattern suggests: early phase (days 1–2): rapid carboxymethyl cellulase (CMCase) synthesis facilitated carbon acquisition, triggering exponential growth. Mid-late phase (>day 4): Nutrient limitation induced secondary enzyme induction, though metabolic waste accumulation ultimately suppressed both enzymatic activity and growth rate. Minimum ZA (0.552, optimal value) was recorded at day 2 (Figure 12B). Strain ETXG-1-1-1 demonstrated severely restricted growth on CMC-Na agar plates: 73.68% reduction in growth rate compared with PDA controls CD expansion significantly lagged behind other strains. Conversely, HZD progressively increased (notably during days 2–4), driving accelerated ZA decline (0.44530→0.41073→0.36111). Critically, ZA consistently remained within the high-activity range (0.30–0.45), confirming this strain’s exceptional capacity for sustained CMCase hyperproduction and stability (Figure 12C).
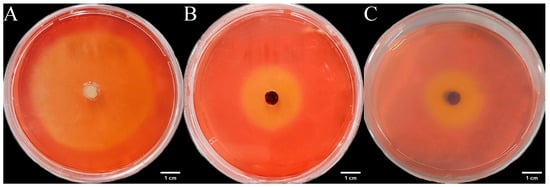
Figure 11.
Detection of CMCase activity in endophytic fungi from Euchresta tubulosa. The incubation time of strains ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C) was 7 d, 5 d and 4 d, respectively. The light yellow central area of the culture medium is the enzymatically active zone, while the red outer ring is the inactive zone.
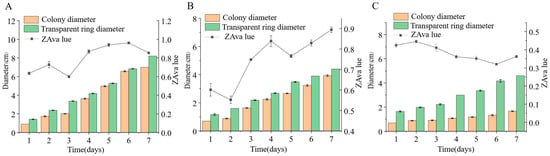
Figure 12.
Comparison of CMCase activities in endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C). from Euchresta tubulosa.
3.6. Extracellular Enzyme Activity
3.6.1. Extracellular Protease Activity
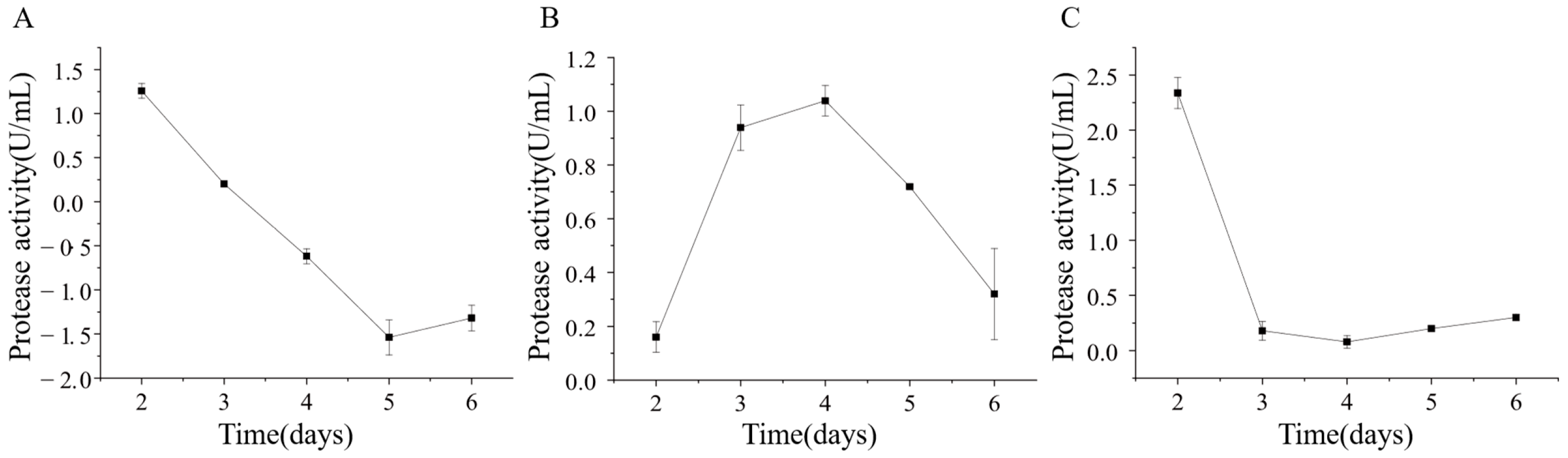
The extracellular protease production of strains ETG-1-2-1, ETY-2-B-b-II.1, and ETXG-1-1-1 was evaluated by fermentation in a skim milk-based liquid medium. Crude enzyme extracts were collected at various time points and subsequently assayed for protease activity, as summarized in Figure 13 The protease activity of strain ETG-1-2-1 reached its highest value of 1.26 ± 0.08 U/mL on day 2 of culture, after which it began to decline until the viability was undetectable. The protease activity of strain ETY-2-B-b-II.1 increased first and then decreased, reaching a peak of 1.04 ± 0.06 U/mL on the 4th day. The protease activity of strain ETXG-1-1-1 had a maximum value of 2.34 ± 0.14 U/mL on the second day, followed by a sharp decrease on the third day, and then a slight upward trend.
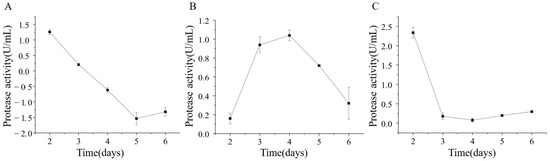
Figure 13.
Changes of extracellular protease activity of three endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C). from Euchresta tubulosa.
Analysis revealed that the extracellular protease activity in strains ETG-1-2-1 and ETXG-1-1-1 primarily occurred during the early cultivation period. Based on the mycelial growth throughout the liquid culture process, it is proposed that the nutrient profile of the skim milk-based medium strongly supported initial fungal proliferation, promoting rapid and vigorous growth that facilitated high levels of enzyme synthesis. In contrast, the peak enzyme production of strain ETY-2-B-b-II.1 exhibited delayed peak enzyme production, possibly due to differences in temporal nutrient utilization or distinct physiological regulation mechanisms.
3.6.2. Extracellular Amylase Activity
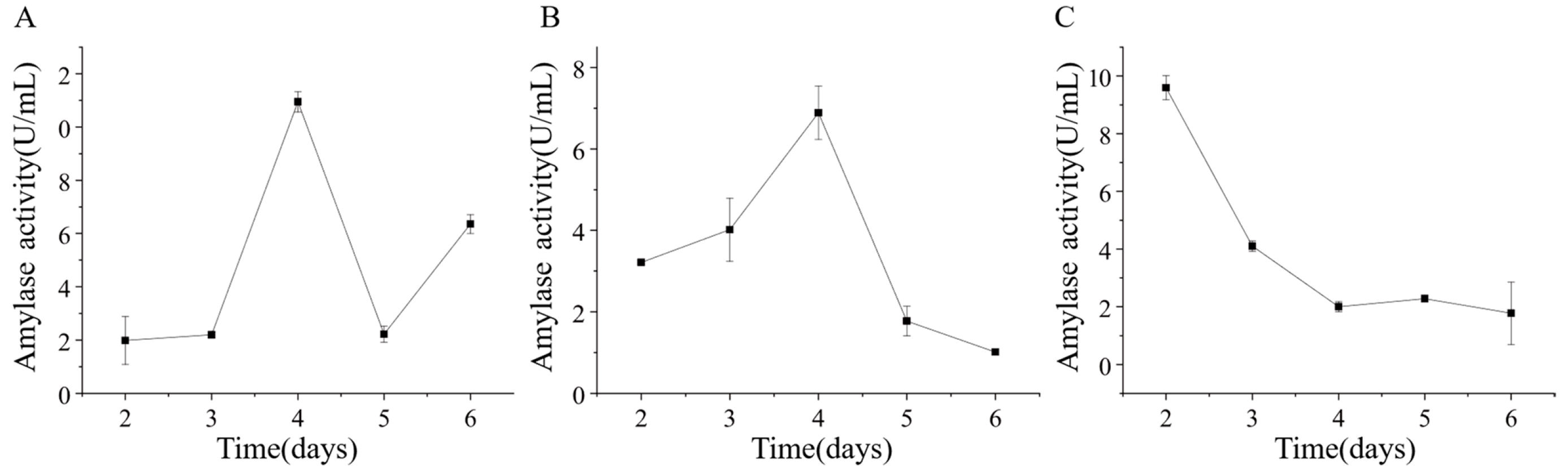
Strains ETG-1-2-1, ETY-2-B-b-II.1, and ETXG-1-1-1 were cultured in mineral-starch broth to assess extracellular amylase production. Crude enzyme extracts collected at various time points exhibited distinct amylolytic activity profiles, as shown in Figure 14 ETG-1-2-1 exhibited gradual activity increase from day 2, surged sharply during days 3–4 (peak: 10.95 ± 0.38 U/mL on day 4), followed by rapid decline and a secondary increase on day 6 (6.35 ± 0.35 U/mL); ETY-2-B-b-II.1 showed unimodal activity peaking at 6.89 ± 0.66 U/mL on day 4; whereas ETXG-1-1-1 reached maximal activity (9.60 ± 0.42 U/mL) on day 2, followed by rapid decline.
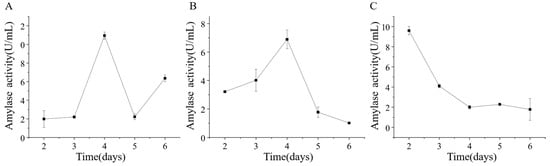
Figure 14.
Changes of extracellular amylase activity of three endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C). from Euchresta tubulosa.
3.6.3. Extracellular Carboxymethyl Cellulase (CMCase) Activity
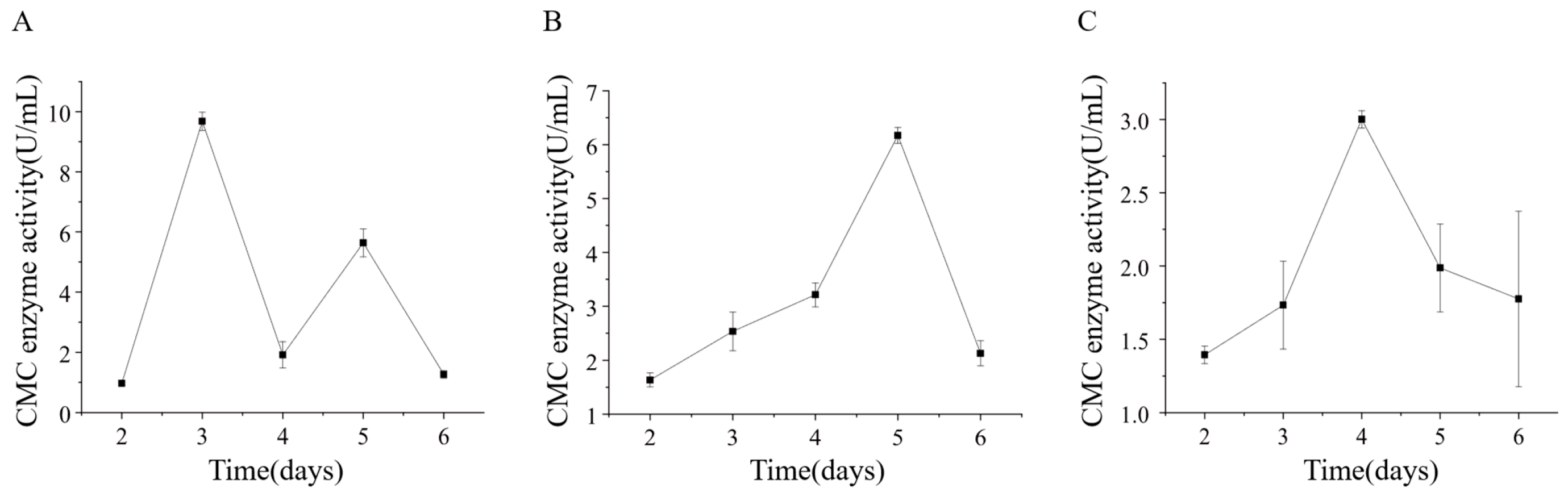
Strains ETG-1-2-1, ETY-2-B-b-II.1, and ETXG-1-1-1 were cultivated in sodium carboxymethyl cellulose (CMC-Na) liquid medium under submerged fermentation (SmF) conditions to evaluate extracellular cellulase production. Crude enzyme extracts collected at various time points were assayed for carboxymethyl cellulase (CMCase) activity, as presented in Figure 15 ETG-1-2-1 reached its peak CMCase activity (9.68 ± 0.30 U/mL) on day 3, followed by a sharp decline and a subsequent rebound to 5.64 ± 0.47 U/mL by day 5. Both ETXG-1-1-1 and ETY-2-B-b-II.1 exhibited unimodal activity profiles: ETY-2-B-b-II.1 achieved maximum activity (6.17 ± 0.15 U/mL) on day 5, while ETXG-1-1-1 peaked earlier (3.00 ± 0.06 U/mL) on day 4.
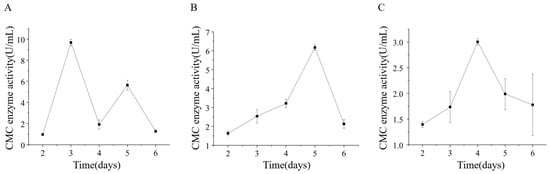

Figure 15.
Changes of extracellular carboxymethyl cellulase (CMCase) activity of three endophytic fungi: ETG−1−2−1 (A), ETY−2−B−b−II.1 (B) and ETXG−1−1−1 (C). from Euchresta tubulosa.
ETG-1-2-1 demonstrated a bimodal carboxymethyl cellulase (CMCase) production pattern. According to Cao Chunlei et al. [15], this phenomenon may stem from the strain’s limited cellulose utilization capacity. During early cultivation, mycelia prioritize nucleic acid and protein synthesis for subsequent growth, triggering initial CMCase induction. Subsequent rapid mycelial expansion depletes intracellular amino acid pools, impairing CMCase synthesis. As readily metabolizable glucose diminishes in the medium, secondary CMCase secretion is induced until nutrient exhaustion ultimately arrests growth.
4. Discussion
Owing to the long-term co-evolution of endophytic fungi with host plants, these fungi engage in synergistic interactions that enhance the host’s adaptive responses to biotic and abiotic stresses. This mutual adaptation has led to the establishment of unique, host-specific endophytic assemblages [3]. A total of 55 endophytic fungal strains were isolated from different tissues of Euchresta tubulosa, comprising 18 from leaves, 12 from stems, and 25 from roots. The morphological typing method based on macro-morphological characteristics is insufficient to fully reflect the phylogenetic relationships of endophytic fungi, especially for non-sporulating fungal species, which presents significant limitations. The ITS region is currently the standard DNA marker for fungal identification [42,43]. Based on combined morphological and molecular analyses, these isolates were assigned to 17 genera, with Colletotrichum, Fusarium, and Penicillium identified as the predominant genera. The detected ITS sequences were subjected to NCBI-BLAST alignment, with the following criteria applied for taxonomic identification: a similarity of ≥97.0–98.5% corresponds to species-level identification, ≥90% to genus-level identification, ≥85% to family-level identification, and ≥80% to order-level identification [33]. Based on these criteria, strain ETG-1-2-1 belonged to a novel genus within the family Marasmiaceae, while strains ETY-2-B-b-II.1 and ETXG-1-1-1 were identified as novel species belonging to the genera Xylaria and Talaromyces, respectively.
Endophytic fungi constitute a major reservoir of unexplored fungal diversity and are capable of producing a vast array of diverse secondary metabolites [44]. They have garnered significant scientific attention due to their potential as novel biological sources of natural active compounds, their role in mediating intraspecific and interspecific signaling, and their ability to alleviate abiotic and biotic stresses in plants [25,45]. Secondary metabolites from three endophytic fungal strains isolated from different tissues of Euchresta tubulosa, were analyzed using LC–MS/MS. Chemical profiling revealed a diverse spectrum of metabolites, including alkaloids, flavonoids, terpenoids, stilbenes, tetracyclines, and steroids. Consistent with previous reports, endophytic fungi from plants not only biosynthesized compounds identical or analogous to those of their host medicinal plant but also produced structurally novel metabolites with putative bioactivities [25]. Notably, strains ETG-1-2-1 and ETXG-1-1-1 were found to produce oxymatrine, a compound also present in E. tubulosa, which exhibits broad pharmacological properties such as anti-inflammatory, antiviral, antifibrotic, and antitumor effects [18]. Li Danyang et al. reported that alkaloid compounds primarily exert antibacterial effects by disrupting key structural components of microbial cells, including the cell wall and cytoplasmic membrane [46]. Building upon this finding, Liu Mengxiao et al. demonstrated that oxymatrine significantly inhibits fungal growth through multiple mechanisms: suppression of biofilm formation, impairment of fungal cell membrane integrity, and reduction of conidial adhesion capacity [47]. Fermentation products of these strains demonstrated significant inhibitory activity against both Staphylococcus aureus and Bacillus subtilis. Against this background, the present study evaluated the antibacterial activity of acetone-soluble extracts derived from the fermentation broths of three endophytic fungal strains isolated from E. tubulosa. The results revealed varying degrees of inhibitory activity against Escherichia coli, Staphylococcus aureus, and Candida albicans, with strain ETG-1-2-1 exhibiting the strongest antimicrobial effect. Supporting this observation, previous studies have shown that oxymatrine, purified from the rhizomes of E. tubulosa, displays potent antibacterial activity against both S. aureus and E. coli, as evidenced by low minimum inhibitory concentrations (MICs) and minimum bactericidal concentrations (MBCs) [48]. Collectively, these findings suggest that the observed antibacterial activity of the fungal fermentation products may be attributable, at least in part, to the production or modulation of oxymatrine or structurally/functionally related secondary metabolites.
Furthermore, the flavonoids detected in the fungal fermentation product, including kaempferol, sakuranetin, and hesperetin, have also been reported in the plant Euchresta tubulosa. These compounds hold significant potential for medicinal research, as accumulating evidence suggests they possess various pharmacological activities, such as anti-inflammatory, antioxidant, neuroprotective, and cardiovascular disease-fighting effects. [49,50,51]. This study further evaluated the hydroxyl (•OH) and ABTS radical scavenging capacities of extracts derived from three endophytic fungal strains. Results revealed that all three strains exhibited concentration-dependent scavenging activity against both radicals; however, their •OH radical scavenging capacity was consistently lower than that of the positive control (ascorbic acid, Vc). Notably, the ethyl acetate extract obtained from the fermentation broth of ETY-2-B-b-II.1, an endophytic fungus isolated from the leaves of E. tubulosa, demonstrated significantly stronger ABTS radical scavenging activity than the Vc control at concentrations exceeding 1 mg/mL. Taken together with previous findings showing that the total flavonoid extract from E. tubulosa leaves exhibits markedly superior ABTS radical scavenging activity relative to Vc, these results suggest that the antioxidant properties of fermentation products derived from E. tubulosa endophytes may be attributable, at least in part, to flavonoid compounds present in their secondary metabolites [18].
These findings suggest that the endophytic fungi of Euchresta tubulosa could serve as a sustainable alternative resource to alleviate the shortage of plant resources. The bioactive metabolites produced by these fungi exhibit significant potential for development and promising application prospects in the research and development of novel pharmaceuticals, as well as in the investigation of their biological activities.
Extracellular enzymes, since their discovery, have been extensively studied for their roles in pathogenicity and substrate degradation. Notably, microbial production of proteases, amylases, carboxymethyl cellulases has been widely applied in fields such as edible mushroom cultivation, baking, food processing, biofuels, and healthcare [17,22,52,53]. As a result, they have gradually become the primary source of industrial enzyme preparations. Referring to the method of Shubha Jagannath et al. [37], specific criteria for the enzymatic zone of activity (ZA) of endophytic fungi were established. Notably, ZA less than 0.69 was defined as indicative of strong enzyme-producing capability. Our extracellular enzyme activity assays revealed that three Euchresta tubulosa endophytic strains possess a rich repertoire of extracellular enzymes.
The strains ETG-1-2-1 and ETXG-1-1-1 both exhibit high protease production capabilities. After 2 days of cultivation, their ZA consistently remained below 0.69, with the optimal ZA as low as 0.48, indicating that their protease production capacity at peak performance surpasses that of many other strains, including Diaporthe sp. KaL-5 [54], 26 strains of Dendrobium officinale seed endophytic fungi [55], and 54 strains of Baliospermum montanum endophytic fungi [37]. Furthermore, referencing the enzyme index (EI) evaluation criteria used by Chien Hao Chai et al. [56] or Tang [55], the assertion that these two fungal strains are strong protease producers is further validated.
Strain ETG-1-2-1 exhibited strong amylase production capacity during the early cultivation stage, by ZA consistently remaining below 0.69 within the first three days. Previous studies have reported that Crinipellis perniciosa, another member of the Marasmiaceae family, also demonstrated varying degrees of amylase production in solid media [57]. However, Marasmius sp. CE25, a strain known for its exceptionally high laccase activity, tested negative for amylase activity [58]. Furthermore, few strains within the Marasmiaceae family have been documented to possess high amylase production capabilities, and no studies have yet been conducted to optimize their enzyme production conditions. Given the strong amylase production capacity demonstrated by strain ETG-1-2-1 in this study, it is hypothesized that further optimization of enzyme production conditions could establish it as a model strain for high-yield amylase production within the Marasmiaceae family.
Three endophytic fungal from Euchresta tubulosa demonstrated a strong capacity for producing carboxymethyl cellulase (CMCase). Notably, strain ETXG-1-1-1 maintained exceptionally high CMCase production throughout its entire growth cycle, with the ZA consistently below 0.45. This performance surpassed that of three other Talaromyces fungi strains isolated from maize in terms of CMCase production capability [59]. Research by Shivam Aggarwal et al. [60] indicates that Talaromyces sp. is a novel fungus characterized by highly efficient cellulose degradation, with cellulases present on its spore surface. Their study showed that adding an inducer with a cellobiose-to-gentiobiose stoichiometric ratio of 2.5:1 to the culture medium could enhance the CMCase synthesis capability of Talaromyces sp. by more than 1.6-fold. This finding further supports the view that cellulases are adaptive enzymes whose expression is influenced by soluble inducers. Furthermore, studies have suggested that enzyme preparations from different fungi are superior to enzyme mixtures from a single strain [61]. Therefore, to obtain an optimal cellulose-degrading enzyme system, it is necessary to further explore and screen for CMCase-producing strains from E. tubulosa.
Endophytic fungi possess significant potential for producing industrially valuable enzymes. Based on our findings, the endophytic fungi isolated from Euchresta tubulosa exhibit considerable diversity in hydrolytic enzymes. Further quantitative analysis via submerged fermentation (SmF) demonstrated significant variations in the activities of protease, amylase, and carboxymethyl cellulase (CMCase) produced by three strains of E. tubulosa endophytic fungi.
The strains exhibited comparable protease-producing activities, with peak protease activities of ETG-1-2-1, ETY-2-B-b-II.1, and ETXG-1-1-1 being 1.26 ± 0.08 U/mL, 1.04 ± 0.06 U/mL, and 2.34 ± 0.14 U/mL, respectively. The differences in protease activity among the three strains are similar to those observed during solid-state fermentation (SSF). However, in comparison with studies by Camila Agnes Lumi Abe [59], Zhang [62], Cai [63], et al., the protease activities produced by these three endophytic fungi are considered to be at a moderate activity level. Further optimization of culture conditions for enzyme production is still required. Additionally, this further validates that even among same microbial origin different strains may exhibit functional divergence in nitrogen utilization and protein degradation [64].
Strain ETG-1-2-1 demonstrates exceptional polysaccharide-degrading capacity, with both its amylase and carboxymethyl cellulase (CMCase) activities being significantly higher than those of the other two strains. Amylase activity peaked at 10.95 ± 0.38 U/mL on day 4, while CMCase activity reached its maximum of 9.68 ± 0.305 U/mL on day 3. Compared with studies by Shivam Aggarwal [60], Honghai Zhang [65], and Shan Zhang [62], the amylase and CMCase activities of this strain have reached a strong activity level. In contrast, strain ETY-2-B-b-II.1 exhibited amylase and CMCase activities of 6.89 ± 0.66 U/mL and 6.17 ± 0.15 U/mL, respectively, with its amylase activity classified as strong, meanwhile its CMCase activity remained at a moderate level. The difference in polysaccharide degradation capabilities between the two strains likely reflects their occupancy of distinct ecological niches within the host microenvironment.
Notably, the amylase produced by strain ETXG-1-1-1 exhibited high activity during submerged fermentation (SmF), whereas the strain not only grew slowly but also showed negligible amylase activity in solid-state fermentation (SSF). For carboxymethyl cellulase (CMCase), the variation in ZA in solid culture indicated that this strain had an extremely high enzyme-producing potential; however, its activity only peaked at 3.00 ± 0.06 U/mL on the 4 d of liquid culture, representing the lowest activity among the three fungal strains tested. Given that the medium components were identical across all experiments, it is inferred that such differences in enzyme activity were caused by the variations in culture methods. This is consistent with the findings of numerous previous studies. For instance, Duan et al. [66] SSF and liquid—state fermentation (LSF) for cellulase production by Trichoderma viride, and the results showed that the yield of SSF was 25% higher than that of SmF, with a correspondingly higher enzyme output. In another study conducted by Cai et al. [63], the protease activity produced by Rhizopus chinensis under SSF was 9.2 times higher than that under SmF. Additionally, Hiroyuki Imanaka et al. [67] demonstrated that agar-plate cultivation or near-solid-state membrane-surface liquid cultivation could significantly stimulate Aspergillus oryzae to produce higher levels of protease and amylase. Furthermore, Cai et al. [63] employed two-dimensional gel electrophoresis combined with mass spectrometry to separate and identify extracellular proteins produced by Rhizopus chinensis under solid-state and submerged cultivation. The results demonstrated that the extracellular proteins generated by Rhizopus chinensis under the two cultivation systems exhibited significant differences in both types and expression levels, with approximately 70% being specific proteins produced under either solid-state or submerged cultivation, among which hydrolases accounted for a relatively large proportion. In addition, building on the findings of Hiroyuki Imanaka et al. [67], which demonstrated that most genes specifically transcribed under membrane-surface liquid cultivation were upregulated by 10-fold or more compared with those specifically transcribed under shake-flask cultivation—it can be concluded that the differences in enzyme activity detected in strain ETXG-1-1-1 under solid-state and submerged cultivation are mainly attributed to the distinct enzyme-producing mechanisms of the strain under different culture methods.
The results of this study further confirm the potential of endophytic fungi in the production of industrial enzymes. Further in-depth analysis of the regulatory mechanisms governing extracellular enzyme activity in endophytic fungi and scientific optimization of the optimal enzyme-producing conditions for strains are required, so as to lay a solid scientific foundation for the application of endophytic fungi in the food processing industry, pharmaceutical manufacturing, and various biomedical therapies.
5. Conclusions
This study presents a systematic investigation of endophytic fungi isolated from Euchresta tubulosa Dunn, an endangered ethnomedicinal plant, and encompassing their isolation, taxonomic identification, secondary metabolites profiling, extracellular enzyme activities assays. Using the tissue explant culture approach, 55 endophytic fungal strains were obtained and classified into 17 genera, based on morphological and molecular characteristics, with Colletotrichum, Fusarium, and Penicillium representing the dominant genera. Among these, four strains were proposed as putative new species, including three novel taxa. Metabolite analysis revealed of fermentation broths from three potentially novel species (or genera) revealed abundant medically relevant bioactive compounds, such as flavonoids and alkaloids. Combined with bioactivity assays, the fermentation products of the three bacterial strains exhibited varying degrees of antibacterial and antioxidant activity. Analysis revealed that the secondary metabolites of these strains are closely associated with their biological activities. Extracellular enzyme activity assays indicated that some strains exhibited broad-spectrum substrate hydrolysis capabilities. Notably, strain ETG-1-2-1 exhibited outstanding polysaccharide-degrading capacity, demonstrating the highest amylase and cellulase activities (10.95 U/mL and 9.68 U/mL, respectively), while strain ETXG-1-1-1 demonstrated prominent protease activity (2.34 U/mL) and maintained a strong ability to produce carboxymethyl cellulase (CMCase) over prolonged periods under solid-state fermentation (SSF) conditions. This study provides the first comprehensive report on the diversity, metabolic characteristics, and ecological functions of endophytic fungi in E. tubulosa. These findings establish a foundation for the future exploration of bioactive natural products, development of enzymatic preparations, and utilization of microbial resources derived from endophytic fungi.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microorganisms14030664/s1, Figure S1: TIC profiles of sample ETG-1-2-1. (Top) Positive ESI mode (targeting cations); (bottom) negative ESI mode (targeting anions). Figure S2: TIC profiles of sample ETY-2-B-b-II.1. (Top) Positive ESI mode (targeting cations); (bottom) nnegative ESI mode (targeting anions). Figure S3: TIC profiles of sample ETXG-1-1-1. (Top) Positive ESI mode (targeting cations). Figure S4: Mass spectrum of pilocarpine. (TOP) Full scan mass spectrum of pilocarpine; (Bottom) MS/MS spectrum of pilocarpine. Figure S5: Mass spectrum of trigonelline. (Top) Full scan mass spectrum of trigonelline; (bottom) MS/MS spectrum of trigonelline. Figure S6: Mass spectrum of oxymatrine. (Top) Full scan mass spectrum of oxymatrine; (bottom) MS/MS spectrum of oxymatrine. Figure S7: Mass spectrum of glycitein. (Top) Full scan mass spectrum of glycitein; (bottom) MS/MS spectrum of glycitein. Figure S8: Mass spectrum of 4′,7-Dihydroxyflavanone. (Top) Full scan mass spectrum of 4′,7-Dihydroxyflavanone; (bottom) MS/MS spectrum of 4′,7-Dihydroxyflavanone. Figure S9: Mass spectrum of naringenin. (Top) Full scan mass spectrum of naringenin; (bottom) MS/MS spectrum of naringenin. Figure S10: Mass spectrum of formononetin. (Top) Full scan mass spectrum of formononetin; (bottom) MS/MS spectrum of formononetin. Figure S11: Mass spectrum of minocycline. (Top) Full scan mass spectrum of minocycline; (bottom) MS/MS spectrum of minocycline. Figure S12. Results of sequence comparison of ITS segments of endophytic fungi from ETG-1-2-1. Figure S13. Results of sequence comparison of ITS segments of endophytic fungi from ETY-2-B-b-II.1. Figure S14. Results of sequence comparison of ITS segments of endophytic fungi from ETXG-1-1-1. Figure S15. Results of sequence comparison of ITS segments of endophytic fungi from ETXG-1-3-1. Figure S16. Results of ITS sequence alignment from ETXG-1-1-1 and ETXG-1-3-1. Figure S17. Standard curve of determination of content glucose (A) and tyrosine (B). Table S1. Preparation of Medium.
Author Contributions
X.Y. (Xinlian Yin): Writing—original draft, Investigation, Methodology, Data curation, Funding acquisition. W.G.: Writing—original draft, Methodology, Data curation, Investigation, Formal analysis. Q.W.: Data curation, Investigation, Visualization, Funding acquisition. R.N.: Investigation, Formal analysis, Funding acquisition. D.Q.: Investigation, Data curation, Visualization. Y.H. and S.H.: Investigation, Formal analysis, Visualization. L.W. and X.Y. (Xiaolin Ye): Investigation, Data curation. S.Y.: Writing–review and editing, Conceptualization, Funding acquisition, Project administration, Supervision, Resources. J.C.: Conceptualization, Funding acquisition, Writing—review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the project supported by the Natural Science Foundation of Hunan Province, China (2026JJ80741, 2024JJ7409), Scientific Research Fund of Hunan Provincial Education Department (25B0496), the College Student Innovation and Entrepreneurship Training Program of Hunan Provincial (S202410531046, 202510531017), and the Scientific Research Project of Jishou University (Jdx23031).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors on request.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| E. tubulosa | Euchresta tubulosa Dunn |
| LC–MS/MS | Liquid Chromatography–Tandem Mass Spectrometry |
| PDA | Potato Dextrose Agar |
| PDB | Potato Dextrose Broth |
| ZA | Enzymatic zone of activity |
| ETG | ETG-1-2-1 |
| ETY | ETY-2-B-b-II.1 |
| ETXG | ETXG-1-1-1 |
| CMC-Na | Carboxymethyl cellulose sodium |
| CMCase | Carboxymethyl cellulase |
| SSF | Solid-state fermentation |
| SmF | Submerged fermentation |
References
- Tiwari, P.; Bae, H. Endophytic fungi: Key insights, emerging prospects, and challenges in natural product drug discovery. Microorganisms 2022, 10, 360. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Zhu, J.; Zhao, X.; Shi, J.; Jiang, C.; Shao, D. Beneficial effects of endophytic fungi colonization on plants. Appl. Microbiol. Biotechnol. 2019, 103, 3327–3340. [Google Scholar] [CrossRef] [PubMed]
- Flores, A.C.; Kadowaki, M.K.; Silva, J.L.d.C.; Bianchini, I.d.A.; Sene, L. Enzymatic potential of endophytic fungi: Xylanase production by Colletotrichum boninense from sugarcane biomass. Braz. J. Microbiol. 2023, 54, 2705–2718. [Google Scholar] [CrossRef]
- Saini, H.P.; Meena, M.; Sahoo, A.; Mehta, T. A review on fungal endophytes of the family Fabaceae, their metabolic diversity and biological applications. Heliyon 2025, 11, e42153. [Google Scholar] [CrossRef]
- Stierle, A.; Strobel, G.; Stierle, D. Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of Pacific yew. Science 1993, 260, 214–216. [Google Scholar] [CrossRef]
- Wang, Q.; Yang, X.R.; Zheng, X.; Shi, P.J. Research Progress on Endophytic Fungal Diversity and Metabolites of Medicinal Plants. Plant Fiber Sci. China 2024, 46, 344–353, 360. [Google Scholar] [CrossRef]
- Correia, A.M.; Lira, S.P.; Assis, M.A.; Rodrigues, A. Fungal Endophyte Communities in Begonia Species from the Brazilian Atlantic Rainforest. Curr. Microbiol. 2018, 75, 441–449. [Google Scholar] [CrossRef]
- Huang, M.; Gao, D.; Lin, L.; Wang, S.; Xing, S. Spatiotemporal dynamics and functional characteristics of the composition of the main fungal taxa in the root microhabitat of Calanthe sieboldii (Orchidaceae). BMC Plant Biol. 2022, 22, 556. [Google Scholar] [CrossRef]
- Patchett, A.; Newman, J.A. Comparison of Plant Metabolites in Root Exudates of Lolium perenne Infected with Different Strains of the Fungal Endophyte Epichloë festucae var. lolii. J. Fungi 2021, 7, 148. [Google Scholar] [CrossRef]
- Nandy, S.; Das, T.; Tudu, C.K.; Pandey, D.K.; Dey, A.; Ray, P. Fungal endophytes: Futuristic tool in recent research area of phytoremediation. S. Afr. J. Bot. 2020, 134, 285–295. [Google Scholar] [CrossRef]
- Taheri, P.; Dastogeer, K.M.G.; Ratu, S.T.N.; Agyekum, D.V.A.; Yasuda, M.; Fujii, Y.; Okazaki, S. Diversity and plant growth promotion potential of endophytic fungi isolated from hairy vetch in Japan. Front. Plant Sci. 2024, 15, 1476200. [Google Scholar] [CrossRef]
- Mo, Q.G.; Fu, H.Y.; Zhou, G.; Wang, Y.W. Research Progresses of Endophytic Fungi from Dendrobium huoshanense and Its Secondary Metabolite. J. Wuhan Univ. (Nat. Sci. Ed.) 2019, 65, 333–339. [Google Scholar] [CrossRef]
- Hawar, S. Extracellular Enzyme of Endophytic Fungi Isolated from Ziziphus spina Leaves as Medicinal Plant. Int. J. Biomater. 2022, 2022, 2135927. [Google Scholar] [CrossRef]
- Agrawal, S.; Samanta, S.; Deshmukh, S.K. The antidiabetic potential of endophytic fungi: Future prospects as therapeutic agents. Biotechnol. Appl. Biochem. 2022, 69, 1159–1165. [Google Scholar] [CrossRef]
- Cao, C.L.; Cui, B.K.; Qin, W.M. Activity changes of several extracellular enzymes in liquid culture of Phellinus mori. Mycosystema 2011, 30, 275–280. [Google Scholar] [CrossRef]
- Deshmukh, S.K.; Agrawal, S.; Prakash, V.; Gupta, M.K.; Reddy, M.S. Anti-infectives from mangrove endophytic fungi. S. Afr. J. Bot. 2020, 134, 237–263. [Google Scholar] [CrossRef]
- Zhang, Y.; Yao, S.F.; Sun, W.H.; OuYang, L.; Zhao, B.Q.; Zhou, X.J. Isolation, identification, and anti-hepatoma activity of endophytic fungi from Carpesium abrotanoides. J. Hunan Univ. Chin. Med. 2024, 44, 47–53. [Google Scholar] [CrossRef]
- Lei, J.X. Extraction, Isolation and Identification of Flavonoids from Euchresta tubulosa Dunn. Master’s Thesis, Jishou University, Jishou, China, 29 May 2024. [Google Scholar]
- Ahmed, T.; Juhász, A.; Bose, U.; Terefe, N.S.; Colgrave, M.L. The role of fungal proteases in bioactive peptide production and analytical approaches for tracking their activity. TrAC Trends Anal. Chem. 2025, 192, 118359. [Google Scholar] [CrossRef]
- Pan, J.Q.; Chen, L.M.; Lao, L.Y.; Zhou, X.J. Purification and characterization of extracellular proteases from Bacillus pumilus. Food Ferment. Ind. 2025, 51, 63–70. [Google Scholar] [CrossRef]
- Sopalun, K.; Iamtham, S. Isolation and screening of extracellular enzymatic activity of endophytic fungi isolated from Thai orchids. S. Afr. J. Bot. 2020, 134, 273–279. [Google Scholar] [CrossRef]
- Bhadra, F.; Gupta, A.; Vasundhara, M.; Reddy, M.S. Endophytic fungi: A potential source of industrial enzyme producers. 3 Biotech 2022, 12, 86. [Google Scholar] [CrossRef]
- Cai, G.X.; Pu, X.C.; Chen, L.F.; Chen, H.J.; Zhou, Z.G. Hunan Materia Medica; Hunan Science and Technology Press: Changsha, China, 2004; Volume 7, p. 5078. [Google Scholar]
- Fu, Y.M.; Yu, D.X.; Wang, S.N.; Yang, L.Y.; Deng, Z.P. Research progress on pharmacological effects and mechanisms of flavonoids from Sophorae Tonkinensis Radix et Rhizoma. Tradit. Herb. Drugs 2022, 53, 6234–6244. [Google Scholar] [CrossRef]
- Li, W.; Wang, H.; Dong, A. Preparative Separation of Alkaloids from Stem of Euchresta tubulosa Dunn. by High-Speed Counter-Current Chromatography Using Stepwise Elution. Molecules 2019, 24, 4602. [Google Scholar] [CrossRef]
- Zhuo, W.; Ren, F.; Wang, L.; Chen, S.; Chen, Y.; Huang, H. Characterization of the first chloroplast genome of Euchresta tubulosa Dunn and its phylogenetic analysis. Mitochondrial DNA Part B Resour. 2021, 6, 2884–2885. [Google Scholar] [CrossRef]
- Li, F.; Liu, X.N.; Wei, Y.M.; Hua, Y.L. Analysis of Chemical Constituents of Different Extracts from Kudouzi (Sophora alopecuroides L.). Chin. Arch. Tradit. Chin. Med. 2023, 41, 45–52, 280-281. [Google Scholar] [CrossRef]
- Liu, T.T.; Zhang, Y.L.; Ma, L.N.; Mao, L.Y.; He, T.; Cao, J.L. Progress of Neurotoxicity Induced by Irrational Use of Sophorae tonkinensis Radix et Rhizoma. Eval. Anal. Drug-Use Hosp. China 2022, 22, 1149–1152. [Google Scholar] [CrossRef]
- Luo, F.X.; Anaer, M.H.M.T.; Luo, J.Y.; Gao, L.; Yan, M. Inhibitory Effect of Sophora Alopecuroides Alkaloids on PC12 Cells and Its Mechanism. Guid. J. Tradit. Chin. Med. Pharmacol. 2024, 30, 5–8, 25. [Google Scholar] [CrossRef]
- Ke, S.W.; Lv, J.H.; Chen, P.; Zhang, R.P.; Yu, Y. Isolation and Identification of Flavonoids-producing Endophytic Fungi in Papaya. Biotechnol. Bull. 2019, 35, 47–52. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Põlme, S.; Kõljalg, U.; Yorou, N.S.; Wijesundera, R.; Villarreal Ruiz, L.; Vasco-Palacios, A.M.; Thu, P.Q.; Suija, A.; et al. Fungal biogeography. Global diversity and geography of soil fungi. Science 2014, 346, 1256688. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Wang, Z.; Zhu, Q.; Tian, P. Systematic Investigations of the Huperzine A—Producing Endophytic Fungi of Huperzia serrata in China and Fermentation Optimization Using OSMAC Strategy. Molecules 2025, 30, 2704. [Google Scholar] [CrossRef]
- Nie, R.S.; Liu, Y.T.; Wang, X.Y.; Chen, Y.; Hu, J.T.; Yin, X.L.; Cheng, J.; Yao, S.F. Activities and components of fermentation products of the endophytic fungi of Euchresta tubulosa Dunn. Microbiol. China 2025, 52, 5316–5335. [Google Scholar] [CrossRef]
- Wu, C.L.; Huang, T.Q.; Lan, T.R.; Lin, L.; Zhang, L.G.; Li, J.; Luo, Z.B.; Wu, Y.J. Isolation, Identification and Minimum Inhibitory Concentration Determinationof Mycoplasma gallisepticum from White-feathered Broilers in Fujian. China Anim. Husb. Vet. Med. 2021, 48, 1489–1497. [Google Scholar] [CrossRef]
- Yan, J.; Gou, X.J.; Zhou, Q.F.; Ji, X.M.; Cui, X.Y.; Lan, H.R.; Li, X. Detem ination of H ydroxyl Radical G enerating from Fenton Reaction by Spectrophotometry. J. Chengdu Univ. (Nat. Sci. Ed.) 2009, 28, 91–93, 103. [Google Scholar] [CrossRef]
- Han, L.X.; Ji, P.; Mai, Z.H.; Zhang, Y.L.; Yang, R.; Hua, L.; Wei, Y.M. Analysis of Antioxidant Activity of Shaoyao Decoction and Its Protective Effect against H2O2-induced Oxidative Damage in Caco-2 Cells. Prog. Vet. Med. 2025, 46, 15–21. [Google Scholar] [CrossRef]
- Jagannath, S.; Konappa, N.; Lokesh, A.; Dasegowda, T.; Udayashankar, A.C.; Chowdappa, S.; Cheluviah, M.; Satapute, P.; Jogaiah, S. Bioactive compounds guided diversity of endophytic fungi from Baliospermum montanum and their potential extracellular enzymes. Anal. Biochem. 2021, 614, 114024. [Google Scholar] [CrossRef]
- Hu, Y.Y.; Tan, P.; Yang, T.; He, Y.N.; Yang, X.B.; Lin, J.Z.; Xu, R.C.; Zhang, D.K. Study on determination methods of biological activity of quality of four kinds of Chinese herbal medicine for digestion: Selection of digestive enzymes and determination of activity potency. Chin. Tradit. Herb. Drugs 2023, 54, 2106–2113. Available online: https://link.cnki.net/urlid/12.1108.R.20230329.1450.002 (accessed on 3 February 2026).
- Yao, S.F.; Yin, X.L.; Cheng, J.; Wang, Q.; Guo, W.; Nie, R.S.; Hu, Y.; Wang, L.X.; Ye, X.L.; Yang, Z.J. Euchresta tubulosa Dunn Endophytic Fungus Neocrinipelis albiflorocum ETG-1-2-1, Its Cultivation Method and Application. CN120682943A, 24 June 2025. [Google Scholar]
- Cheng, J.; Wang, Q.; Yao, S.F.; Yin, X.L.; Zhang, C.; Wang, Y.; Yang, Z.J.; Hu, S.S.; Qing, D.J.; Zhang, Z.Y. A Strain of Euchresta tubulosa Dunn Endophytic Fungus Xylaria cineeronitense ETY-2-B-b-II. 1, and Its Cultivation Method and Application. CN121046216A , 27 August 2025. [Google Scholar]
- Yao, S.F.; Li, R.; Cheng, J.; Jia, Y.L.; Xie, W.; Yin, X.L.; Wang, Q.; Guo, W.; Liu, Y.T.; Qin, D.J. A Strain of Euchresta tubulosa Dunn Endophytic Fungus Talaromyces polymorphus ETXG-1-1-1, and Its Culture Method Law and Its Applications. CN121046215A, 26 August 2025. [Google Scholar]
- Dos Reis, J.B.A.; Lorenzi, A.S.; Do Vale, H.M.M. Methods used for the study of endophytic fungi: A review on methodologies and challenges, and associated tips. Arch. Microbiol. 2022, 204, 675. [Google Scholar] [CrossRef]
- Iquebal, M.A.; Jaiswal, S.; Mishra, V.K.; Jasrotia, R.S.; Angadi, U.B.; Singh, B.P.; Passari, A.K.; Deka, P.; Prabha, R.; Singh, D.P. Fungal genomic resources for strain identification and diversity analysis of 1900 fungal species. J. Fungi 2021, 7, 288. [Google Scholar] [CrossRef]
- Bertini, L.; Perazzolli, M.; Proietti, S.; Capaldi, G.; Savatin, D.V.; Bigini, V.; Longa, C.M.O.; Basaglia, M.; Favaro, L.; Casella, S. Biodiversity and bioprospecting of fungal endophytes from the antarctic plant Colobanthus quitensis. J. Fungi 2022, 8, 979. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Fasoyin, O.E.; Molnár, I.; Xu, Y. Secondary metabolites from hypocrealean entomopathogenic fungi: Novel bioactive compounds. Nat. Prod. Rep. 2020, 37, 1181–1206. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.M.; Li, X.; Zhang, C.H.; Lv, L.J.; Gao, B.; Li, M.H. Research Progress on Antibacterial Activities and Mechanisms of Natural Alkaloids: A Review. Antibiotics 2021, 10, 318. [Google Scholar] [CrossRef]
- Liu, M.X.; Chen, L.L.; Wang, Y.; Zhang, J.P. Research progress on Sophora Flavescens of traditional Chinese medicine. J. Pharm. Pract. Serv. 2025, 43, 156–162. [Google Scholar] [CrossRef]
- Mo, J.W. Isolation, Purification and Activity of Alkaloids from the Stem of Euchresta tubulosa Dunn. Master’s Thesis, Jishou University, Jishou, China, 22 May 2025. [Google Scholar]
- Wang, Z.; He, C.H.; Yang, H.Y. Research progress on the mechanism of kaempferol against atherosclerosis. Chin. J. Arterioscler. 2026, 34, 171–177. [Google Scholar] [CrossRef]
- Xiao, L.Y.; Chen, Y.; Duan, T.; Sun, Y.; Xu, Y.B.; Zhao, Y.J.; Song, X.; Yan, X.Z.; Hu, J.G. Effect of Sakuranetin on Microglia-Mediated Neuroinflammation After Spinal Cord Injury. ACTA Acad. Med. Sin. 2024, 46, 836–848. [Google Scholar] [CrossRef]
- Yu, Q.L. Research Progress on Extraction Methods and Bioactivities of Citrus Peel Flavonoids. Agric. Prod. Process. 2026, 3, 105–110. [Google Scholar] [CrossRef]
- Chen, X.X. Building a Microfluidic-Mass Spectrometry Platform for High-Throughput Screening. Master’s Thesis, Tianjin University of Science and Technology, Tianjin, China, 1 June 2024. [Google Scholar]
- Di, C.Y.; Guo, X.J.; Liu, H.L.; Guo, W.; Zhu, B.C. Cloning and expression of β-1,4-endoglucanase gene of cellulose derading bacteria N2-10. J. Hebei Univ. (Nat. Sci. Ed.) 2018, 38, 403–409. [Google Scholar] [CrossRef]
- Kubayi, S.; Makola, R.T.; Dithebe, K. Exploring the Antimicrobial, Antioxidant and Extracellular Enzymatic Activities of Culturable Endophytic Fungi Isolated from the Leaves of Kirkia acuminata Oliv. Microorganisms 2025, 13, 692. [Google Scholar] [CrossRef]
- Tang, Y.J. Preliminary Study on the Specificity and Mechanism of Symbiotic Germination Between Medicinal Dendrobium Seeds and Mycorrhizal Fungi. Master’s Thesis, Peking Union Medical College, Beijin, China, 1 May 2021. [Google Scholar]
- Chai, C.H.; Hong, C.F.; Huang, J.W. Identification and characterization of a multifunctional biocontrol agent, Streptomyces griseorubiginosus LJS06, against cucumber anthracnose. Front. Microbiol. 2022, 13, 923276. [Google Scholar] [CrossRef]
- Bastos, C.N. Produção de enzimas extracelulares por Crinipellis perniciosa. Fitopatol. Bras. 2005, 30, 286–288. [Google Scholar] [CrossRef]
- Peraza-Jiménez, K.; De la Rosa-García, S.; Huijara-Vasconselos, J.J.; Reyes-Estebanez, M.; Gómez-Cornelio, S. Enzymatic bioprospecting of fungi isolated from a tropical rainforest in Mexico. J. Fungi 2021, 8, 22. [Google Scholar] [CrossRef] [PubMed]
- Lumi Abe, C.A.; Bertechini Faria, C.; Fernandes de Castro, F.; De Souza, S.R.; dos Santos, F.C.; Novais da Silva, C.; Tessmann, D.J.; Barbosa-Tessmann, I.P. Fungi isolated from maize (Zea mays L.) grains and production of associated enzyme activities. Int. J. Mol. Sci. 2015, 16, 15328–15346. [Google Scholar] [CrossRef]
- Aggarwal, S.; Dorairaj, S.; Adlakha, N. Stoichiometric balance ratio of cellobiose and gentiobiose induces cellulase production in Talaromyces cellulolyticus. Biotechnol. Biofuels Bioprod. 2023, 16, 48. [Google Scholar] [CrossRef]
- Borin, G.P.; Sanchez, C.C.; De Souza, A.P.; De Santana, E.S.; De Souza, A.T.; Leme, A.F.P.; Squina, F.M.; Buckeridge, M.; Goldman, G.H.; Oliveira, J.V.d.C. Comparative secretome analysis of Trichoderma reesei and Aspergillus niger during growth on sugarcane biomass. PLoS ONE 2015, 10, e0129275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Xiong, Z.P.; Zhang, J.; Nie, Z.L.; Chen, X.X.; Ling, X.; Xiong, Z. Isolation and identification of intestinal fungi in the larvae of Dendrolimu kikuchii Matsumura and their hydrolase activity. J. Environ. Entomol. 2019, 41, 413–419. [Google Scholar] [CrossRef]
- Cai, J.; Wang, D.; Xu, Y. Comparison of extracellular proteome from Rhizopus chinensis in solid-state and submerged cultures. Microbiol. China 2017, 44, 402–410. [Google Scholar] [CrossRef]
- Jacoby, R.P.; Succurro, A.; Kopriva, S. Nitrogen substrate utilization in three rhizosphere bacterial strains investigated using proteomics. Front. Microbiol. 2020, 11, 784. [Google Scholar] [CrossRef]
- Zhang, H.H.; Wang, Y.; Brunecky, R.; Yao, B.; Xie, X.M.; Zheng, F.; Luo, H.Y. A swollenin from Talaromyces leycettanus JCM12802 enhances cellulase hydrolysis toward various substrates. Front. Microbiol. 2021, 12, 658096. [Google Scholar] [CrossRef] [PubMed]
- Duan, J.Z.; Zhang, A.L.; Cao, D.J. The Compare of SSF and LSF with Trichodema Viride for Cellulase Production in Yield and Catalysis Property. Mod. J. Anim. Husb. Vet. Med. 2000, 3, 5–6. [Google Scholar] [CrossRef]
- Imanaka, H.; Tanaka, S.; Feng, B.; Imamura, K.; Nakanishi, K. Cultivation characteristics and gene expression profiles of Aspergillus oryzae by membrane-surface liquid culture, shaking-flask culture, and agar-plate culture. J. Biosci. Bioeng. 2010, 109, 267–273. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.