Abstract
In vitro studies were conducted in a series to investigate if the ruminal microbes are capable of degrading chlorpyrifos. This in vitro study presents the results from three experiments: Exp. I was conducted without feed, while Exp II and III were conducted with feed, either with or without methanol for dissolving chlorpyrifos, respectively. A basal diet comprising finger millet straw and concentrate was prepared. Incubation medium with feed but without chlorpyrifos served as the control. A total of six replicates each of control and chlorpyrifos spiked were used for the incubation. The pesticide concentration in the incubation medium before and after 24 h of incubation was analyzed using GC-MS/MS. The genomic DNA was isolated from the incubation fluid of the individual samples, and the shotgun metagenomic sequencing was performed. The clean reads were taxonomically classified using the Kraken2 database, and microbial classification at different taxonomic ranks was separated using Pavian v1.0. The microbial genes in the metagenome data were predicted and assigned functional roles using the MetaErg v1.2.3 pipeline. The assigned KEGG Orthology (KO), EC numbers (Enzyme Commission number), Gene Ontology (GO), and corresponding NCBI taxonomy information relevant to chlorpyrifos metabolism/degradation were retrieved. Results from the study revealed that the chlorpyrifos concentration was decreased from 5.78 to 1.64 ppm over 24 h of in vitro incubation with feed. Similar alpha and beta diversity indices between control and chlorpyrifos treatments revealed that the richness and the evenness of the microbial population were not affected by the presence of chlorpyrifos in the rumen fluid. There was no difference in the microbiota affiliated to the major phyla such as Bacteroidota, Fibrobacterota, Bacillota, and Pseudomonadota. The EC 3.1.8.1, EC 3.1.3.1, EC 1.14.13.-, and EC 1.1.1.- reported for chlorpyrifos degradation were not detected in the metagenome, and only EC 3.1.1.1 was identified, which demonstrated that degradation of chlorpyrifos was carried out by the affiliated enzyme carboxylesterase. The presence of GO:0004035, GO:0004364, GO:0019637, GO:0016791, and GO:0042178 in the metagenome strengthens that the chlorpyrifos degradation in the present study was primarily assigned to the rumen microbiota. This in vitro study provided insights into the rumen microbiota involved in the chlorpyrifos degradation and the initial clue that the rumen microbes are capable of degrading chlorpyrifos. Further, the animal studies in different species with the variable levels of chlorpyrifos are also warranted to confirm the efficacy of rumen microbes in mixed syntrophy and determine the threshold capabilities of the ruminal microbes.
1. Introduction
Worldwide, 3.70 million tons of pesticides in terms of active ingredients are used in agriculture [1]. Chlorpyrifos is an organophosphate pesticide extensively used in a wide variety of food and feed crops to control insects and pests [2]. Recently, during the Stockholm Convention in Geneva to eliminate hazardous chlorpyrifos, many countries agreed to a complete ban on the use of chlorpyrifos pesticide; however, some of the countries sought exemptions for its use in selected crops for another five years [3]. In India, the Ministry of Agriculture and Farmers Welfare recently banned the usage of 46 pesticides, while nine pesticides have been kept under a watchlist with restricted use [4]. The pesticide residue could transfer from the agricultural crop to the animal feeds that are mostly obtained as by-products from the main agricultural crops [5]. The pesticide is absorbed and accumulated in the vegetative parts of the plants, such as leaves and stalks, which are used as feed for the livestock. Chlorpyrifos, despite the low aqueous solubility, highly volatile nature, and moderate persistence in soil, could transfer to the stalks, leaves, and grains at a lower concentration than the soil [6].
There are many reports on the biodegradation of chlorpyrifos in the soil [7,8,9,10]. Microbial degradation in the soil is one of the prominent mechanisms for chlorpyrifos degradation carried out by the microbial enzymes [11]. Pseudomonas, Bacillus, Enterobacter, Stenotrophomonas, Sphingomonas, and Verticillium are the prominent genera of microbes involved in the biodegradation of chlorpyrifos in the soil. Chlorpyrifos degradation by the microorganisms has been reported during the silage making over 90 days with a lesser efficiency than the degradation of beta-cypermethrin and tebuconazole [12]. Similarly, the degradation of chlorpyrifos by lactic acid bacteria during the ensiling process is also reported by Wang et al. [13]. These reports suggest that chlorpyrifos can be degraded by the soil microorganisms; however, the soil ecosystem does not resemble the rumen ecosystem, as the former is an external, aerobic, open environment driven by sunlight and rainfall, while the latter is an internal, strictly anaerobic, closed environment and host animal-dependent system. These two distinct ecosystems possess a diverse type of microbiota adapted to the specialized environment and carry different functions [14]. Similarly, the ensiling process and rumen ecosystem are also not the same, as the ensiling fermentation takes place in a silo, whereas the fermentation of ingested feed occurs in the rumen inside the animal’s body. Further, there is a distinct difference in the microbial community between the two systems. The rumen microbiome is known to degrade the xenobiotics faster than other environmental degradation pathways, as revealed through the RDX-degrading ability of the sheep rumen [15].
We did not come across any previous studies in the public domain confirming that the rumen microbes can degrade chlorpyrifos ingested through contaminated feeds, and if it is degraded, then to what extent. To the best of our knowledge, the present study is the first attempt to investigate chlorpyrifos degradation by the rumen microbial consortia and the extent of degradation.
2. Materials and Methods
The necessary approval for conducting the studies and collecting rumen fluid was obtained from the Institute Animal Ethics Committee (IAEC) of ICAR-National Institute of Animal Nutrition and Physiology, Bangalore, INDIA, and the Committee for Control and Supervision of Experiments on Animals (CPCSEA), approval no. V-11011(13)/10/2024-CPCSEA-DADF dated 14 October 2024. The ruminal fluid donors were handled in strict adherence to the protocol laid out by the IAEC.
2.1. Collection of Rumen Fluid
The donors of ruminal fluids, a source of microbial inoculum for the in vitro studies, were fed on a basal diet consisting of finger millet (Eleusine coracana) straw and concentrate mixture in 70:30 (DM basis). Two in vitro studies were conducted in a series to investigate the degradation of chlorpyrifos by the ruminal microbes over a defined incubation period. About 2 L of ruminal fluid containing both solid and liquid fractions was collected 3 h post-feeding from two cannulated Holstein Friesian male cattle (mean BW 613 kg) maintained at the experimental livestock unit of the institute. The rumen fluid was collected in a pre-warmed (39 °C) thermos flask flushed with CO2 to maintain anaerobic conditions. The rumen fluid was mixed and filtered through muslin cloth. The filtrate served as the source of ruminal microbial inoculum for the in vitro studies.
2.2. Preparation of Stock Solution
A stock solution of 1000 ppm was prepared by dissolving 10 mg of chlorpyrifos (CAS No 285138-81-0, Sigma-Aldrich Production GmbH, Buchs, Switzerland) in 10 mL of LCMS-grade methanol (CAS No 67-56-1, Sigma-Aldrich, St. Louis, MO, USA) in a volumetric flask and sonicated for 1 min. To achieve a concentration of 1 ppm, 0.1 mL stock solution (1000 ppm) was dissolved in 100 mL LCMS-grade methanol, which served as the working standard of chlorpyrifos. The standard linearity curve of 2.5, 5.0, 10.0, 25, 50.0, and 100 ppb was generated by injecting in GC-MS/MS (7000C, Agilent Technologies, Santa Clara, CA, USA). For the experiment, about 14.1 mg of chlorpyrifos was dissolved in 10 mL of LCMS-grade methanol, and the final concentration of 1383.21 ppm was achieved. From the stock solution, the working solutions were prepared for different experiments.
2.3. Experiment I
For the experiment, I prepared the working solution by taking 23.61 mL of stock solution and dissolving it in 990 mL of buffered ruminal fluid so that the final concentration of the incubation medium reached 10 ppm. This experiment was conducted without a feed sample, and the ruminal fluid, either with chlorpyrifos or without chlorpyrifos, was incubated for different time periods. A total of 30 glass syringes (100 mL, Heberle, Schwerte, Germany) were incubated in vitro to ascertain the degradation of chlorpyrifos at 0, 24, 48, 72, and 96 h. In addition, 3 syringes containing ruminal fluid without chlorpyrifos, which serve as negative controls, were also prepared for the in vitro incubation. About 30 mL of rumen fluid with or without chlorpyrifos was gently poured into the individual syringe using an automatic bottle top dispenser (Varispenser 50 mL, Eppendorf, Hamburg, Germany). The gas bubbles from the syringes were pushed out, and the position of the piston at the commencement of incubation was recorded. The incubation was set up for 96 h in a Hohenheim-type water bath shaker (Eaga Tools & Instruments, Chennai, India) at 39 °C, and the intermittent shaking was performed every 6 h. Six syringes were removed from the water bath shaker at 0, 24, 48, 72, and 96 h of incubation. To cease the fermentation, these syringes, after removal from the water bath shaker, were immediately placed on the ice. The incubation spent from the individual syringe was filtered through 0.45 μm (Milex-HV, Merck Millipore, Burlington, MA, USA) in a serum vial of 100 mL capacity and placed in the refrigerator at 4 °C until chlorpyrifos analysis.
2.4. Experiments II and III
Experiment II was similar to Experiment I, but here, along with the chlorpyrifos-containing buffered ruminal fluid at a similar concentration of 10 ppm, the feed was also incubated to see its impact on chlorpyrifos degradation. The diet for the in vitro studies was prepared by using finger millet straw and concentrate in a 70:30 ratio. About 200 mg of dry feed was weighed and placed in the glass syringes individually; thereafter, 30 mL of chlorpyrifos-containing rumen fluid was added to the syringes. A total of 12 syringes were prepared and incubated in the Hohenheim-type water bath shaker at 39 °C for 24 h. On the termination of incubation, the incubation spent from the individual syringe was filtered through 0.45 μm in a serum vial and placed in the refrigerator at 4 °C until chlorpyrifos analysis.
As methanol was used as a solvent for the spiking of chlorpyrifos in both experiments I and II, and therefore to rule out the influence of methanol on the chlorpyrifos degradation, another experiment (experiment III) was conducted where the chlorpyrifos was directly dissolved in the rumen fluid without using methanol. About 14.1 mg chlorpyrifos was dissolved in 10 mL rumen fluid by sonication for 20 min at a speed of 40 khz (Grant Ultrasonic bath, Cambridge, UK), and a concentration of 1383 ppm was achieved. To achieve the concentration of 5 ppm, about 0.13 mL from the stock solution was dissolved in 360 mL rumen fluid to be used for in vitro incubation. Experiment III was similar to experiment II in all respects except that the methanol was not used for dissolving the chlorpyrifos while preparing the stock solution. A total of 12 syringes containing 200 mg feed along with the 30 mL buffered rumen fluid, which either contained chlorpyrifos or not, were incubated in the Hohenheim-type water bath shaker at 39 °C for 24 h. On the termination of incubation, the spent was filtered and stored at 4 °C until further analysis.
2.5. Chlorpyrifos Analysis
Rumen fluid from the donor animals, water used for the preparation of the incubation medium, and feed samples used in the in vitro incubation were analyzed for the presence of chlorpyrifos before conducting the experiments. About 10 mL of each sample was sent for chlorpyrifos analysis at the NABL-accredited laboratory, Eureka Analytical Services, Bengaluru. The chlorpyrifos assay Limit of Detection (LOD) was 0.003 mg/kg, and the Limit of Quantification was 0.01 mg/kg. From each tube, a 5.0 g sample was weighed and transferred into a 50 mL tube. Thereafter, 10.0 mL of Milli-Q water was added and vortexed for five minutes. Subsequently, 10.0 mL acidified acetonitrile was mixed with one packet of QuECHERS (EN15662, Agilent Technologies, Santa Clara, CA, USA) and vortexed for five minutes. The acidified acetonitrile was prepared by taking 0.1 mL acetic acid (Merck, Boston, MA, USA) in 10 mL acetonitrile. The centrifugation was performed at 4200× g, 4 °C for five minutes, and the supernatant (4.0 mL) was transferred to the RIA vials (Tarsons, Kolkata, India). The supernatant was preserved at −18 °C for two hours, followed by centrifugation at 4200× g, 4 °C for five minutes, and finally 3.0 mL of the supernatant was taken into the RIA vials. About 450 mg MgSO4, 75.0 mg PSA, 50.0 mg C18, and 50.0 mg GCB were sequentially added to the RIA vials containing supernatant and vortexed for two minutes. Thereafter, the content was centrifuged for five minutes at 4200× g, 4 °C, and one mL of supernatant was transferred to the RIA vials and fully evaporated under N2 gas evaporator (Athena Instruments Pvt., Ltd., Mumbai, India). The residue was reconstituted in one mL of HPLC-grade ethyl acetate (Merck, St. Louis, MO, USA), and 2 μL was injected into the GC-MS/MS (7000C, Agilent Technologies, Santa Clara, CA, USA). The chlorpyrifos was quantified by MRM (multi-reaction monitoring) mode by the MassHunter software version 2.0. The final concentration was auto-calculated considering linearity, dilution factor, and sample weight.
2.6. Statistical Analysis
Data on the chlorpyrifos degradation was analyzed using One-way ANOVA in the GraphPad prism, and the mean chlorpyrifos concentration was compared among the different in vitro incubation periods (i.e., 0, 24, 48, 72, and 96 h). The Tukey post-hoc analysis was performed to ascertain the significance among the different time points at a confidence level of 95%. The degradation of chlorpyrifos over 24 h of in vitro incubation in experiment II and III was compared by performing a t-test.
2.7. Genomic DNA Isolation
About 1.5 mL of the incubation fluid was used for the isolation of genomic DNA using the procedure of Yu and Morrison [16]. In brief, the fluid samples were initially centrifuged at 12,400× g for 15 min, and the supernatant was removed. To dissolve the pellet, 1 mL of lysis buffer was added to the tube containing the pellet. This content was transferred to a pre-sterilized screw-cap tube (BioSpec, Bartlesville, OK, USA) containing 0.5 g zirconia beads of 0.1 mm size (BioSpec, Bartlesville, OK, USA). The samples were homogenized using a bead beater (BioSpec, Bartlesville, OK, USA) at the maximum speed for three minutes, followed by incubation at 70 °C for 15 min. Thereafter, the centrifugation was carried out at 12,400× g, and the supernatant was transferred in a 2 mL tube. The protein and polysaccharide contaminations were removed by treating with 260 μL of 10 M ammonium acetate and centrifugation at 12,400× g for 10 min. The supernatant was removed in another tube, and an equal volume of isopropanol was added, followed by gentle mixing. The precipitated DNA was collected by centrifugation at 4 °C for 10 min at 12,400× g, and the pellet was washed with 70% ethanol. Finally, the DNA pellet was dissolved in 100 μL Tris-EDTA buffer, and 2 μL DNase-free RNase (10 mg/mL, Qiagen GmbH, Hilden, Germany) was added for removing the RNA contamination. Subsequently, the QIAamp DNA mini kit (Qiagen GmbH, Hilden, Germany) was used for the isolation of genomic DNA according to the manufacturer’s instructions. The DNA quality was checked with 0.8% agarose gel electrophoresis and quantified with the help of a Qubit 4.0 fluorometer (Invitrogen Corporation, Carlsbad, CA, USA).
2.8. Shotgun Sequencing
Metagenome sequencing was performed using NovaSeq 6000 (Illumina Inc., San Diego, CA, USA) at an external facility, i.e., Eurofins Genomics, Bangalore, India. The metagenomic libraries were prepared using the NEBNext® UltraTM II FS DNA Library Prep Kit (New England Biolabs, Ipswich, MA, USA). About 100–500 ng of DNA was fragmented to achieve a size of 350 base pairs. The fragmentation process was carried out by NEBNext Ultra II FS Reaction Buffer (New England Biolabs, Ipswich, MA, USA) and Ultra II FS Enzyme Mix (New England Biolabs, Ipswich, MA, USA) in a PCR. The fragmented DNA was ligated with the NEBNext adapters (New England Biolabs, Ipswich, MA, USA) by combining 35 μL of fragmented DNA with NEBNext Ultra II Ligation Master Mix (New England Biolabs, Ipswich, MA, USA). The mixture was then incubated at 20 °C for 15 min, followed by ligation with adaptors. The libraries were PCR amplified with the index primers (i5 and i7) by using the following conditions: initial denaturation at 98 °C, 30 s; denaturation at 98 °C, 10 s; annealing at 65 °C, 75 s; and final extension at 65 °C, 5 min. The libraries were evaluated on the Agilent 4150 Tape Station, pooled, and sequenced on NovaSeq 6000 (Illumina, San Diego, CA, USA) using 2 × 150 bp chemistry.
2.9. Bioinformatic Analysis
The Illumina paired-end metagenome sequencing data containing leftover adapters, low-quality bases, Ns, and polyG stretches in the reads with short inserts were cleaned using fastp v0.23.4 [17] using command line options ‘-e 30, -l 70 -g -3 -W 4 -M 20 -h’ for average read quality score, minimum length filter, removing polyG stretches, sliding window for average quality, cut mean quality, and html report generation, respectively. For removing the host contamination from the reads, the quality-filtered data were mapped against the reference created using cattle genome assembly ARS-UCD1.2 (RefSeq assembly accession: GCF_002263795.1) in Hisat2 v2.2.1 [18]. The unmapped paired-end and single-end reads were saved for further analysis using the output option ‘--un-conc-gz’. The clean reads obtained from the Hisat2 output were taxonomically classified using the Kraken2 database at BV-BRC v3.55.7 [19] following the K-mer-based approach, and the taxonomic affiliation was assigned by the lowest common ancestor method [20]. The Kraken output files were resolved to different taxonomic ranks of bacterial and archaeal domains in Pavian v1.0 [21], and the contingency tables at the phylum, order, and genus levels were prepared for downstream analysis and visualization. The contingency tables were filtered for the counts less than 4 with a minimum prevalence of the taxa in at least 20% of the samples and a low variance filter set at 10%. The filtered data were centered log-ratio transformed in MicrobiomeAnalyst v2.0 [22]. For each taxonomic level, the samples were merged into groups, and the composition bar plots were constructed by scaling the bars. Further, the abundance at different taxonomic ranks between the groups was compared for statistically different changes after applying the intrinsic normalization method of DESeq2 in MicrobiomeAnalyst v2.0. The significance cutoff was set at a false discovery rate of < 0.05. The alpha diversity was estimated at the genus level by the Shannon index, and the sample dissimilarity between the groups was estimated by beta diversity based on the Bray–Curtis dissimilarity matrix.
The alpha diversity was represented as pooled box plots, and the beta diversity was graphically represented in PCoA ordination. The statistical comparison between the alpha diversity and beta diversity was performed by the Kruskal–Wallis test and the PERMANOVA, respectively. To find the difference in the key microbial biomarkers between the groups, the linear discriminant analysis (LEfSe) was performed in MicrobiomeAnalyst v2.0 and then linear discriminant analysis for effect size normalization with an LDA score > 2.0 [23].
2.10. Metagenome Gene Annotation
The microbial genes in the metagenome data were predicted and assigned functional roles using MetaErg v1.2.3 [24]. Briefly, the clean read files, free of host contamination, were assembled into contigs using MEGAHIT (v1.2.9, [25]. The contigs below >300 bp in length were filtered out. The clean reads were mapped against the assembled contig sequences using HISAT2 (v2.2.2, [18]. The BAM files were used for generating depth files using BBmap v35.85 [26] ‘pileup.sh’ command. The functional annotation of the samples was performed in MetaErg—the metagenome annotation pipeline. The assigned KEGG Orthology (KO), Enzyme Commission (EC), and Gene Ontology (GO) and their NCBI taxonomy information relevant to chlorpyrifos metabolism/degradation were extracted, and the contribution of different phyla and orders to the detected KO, EC, and GO categories was presented in stacked bar charts, and the category-wise gene abundance was depicted in the corresponding bar plots.
3. Results
The chlorpyrifos content in the ruminal fluid, water, and feed samples used for the in vitro experiments was screened by GC MS/MS. The chlorpyrifos concentration in the above samples remained below the level of quantification (BLQ, i.e., 0.01 mg/kg), indicating that none of the inputs were contaminated with chlorpyrifos.
3.1. Chlorpyrifos Degradation
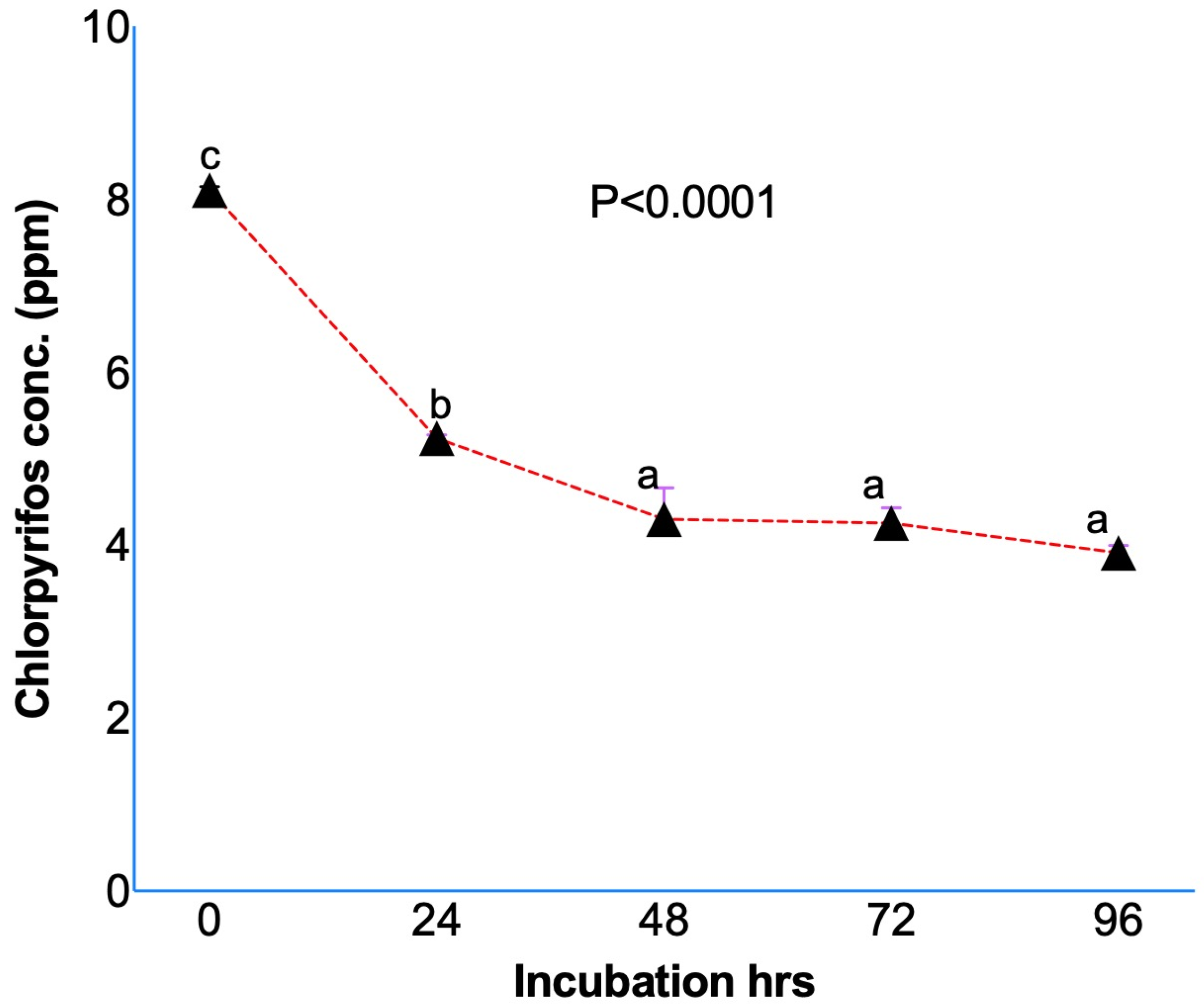
Results from the in vitro studies confirmed the degradation of chlorpyrifos while initially dissolved in methanol (Figure 1) and ruminal fluid without (Figure 2A) or with (Figure 2B) a feed sample. The initial concentration of the chlorpyrifos in the buffered rumen fluid at 0 h was 8.10 ± 0.039 ppm, which after 24 h of in vitro incubation decreased to 5.23 ± 0.042 ppm (Figure 1). The chlorpyrifos concentration was recorded as 4.29 ± 0.364, 4.25 ± 0.177, and 3.91 ± 0.082 ppm at 48, 72, and 96 h of incubation, respectively. The results from the experiment I demonstrated a significant (p < 0.0001) degradation of the chlorpyrifos over 24 h of in vitro incubation with the rumen fluid. About a 35.4% decrease in the chlorpyrifos concentration as compared to the initial concentration was recorded at 24 h of incubation. Similarly, the chlorpyrifos concentration at 48 h was further decreased by 47.0% as compared to the initial concentration at 0 h. The chlorpyrifos concentration (ppm) at 48 h of incubation was significantly (p < 0.0001) lower than the concentration at 0 and 24 h of incubation. However, the chlorpyrifos concentration at 72 and 96 h was not significantly decreased as compared to the concentration at 48 h.
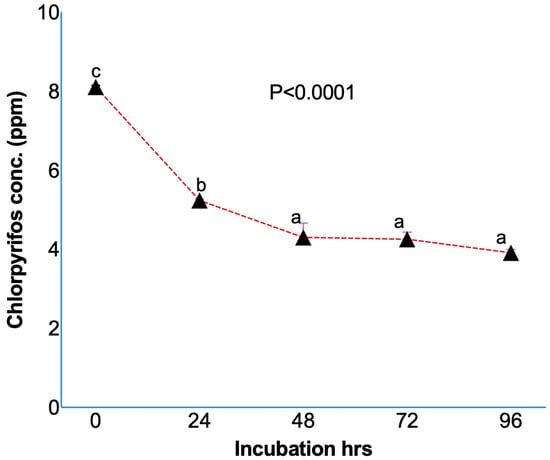
Figure 1.
Chlorpyrifos degradation during in vitro incubation over 24 h by dissolving chlorpyrifos in methanol and incubating along with buffered rumen fluid without any feed sample. Chlorpyrifos concentrations bearing different superscripts a, b, c at different incubation hours represent the significant difference among the incubation hours.
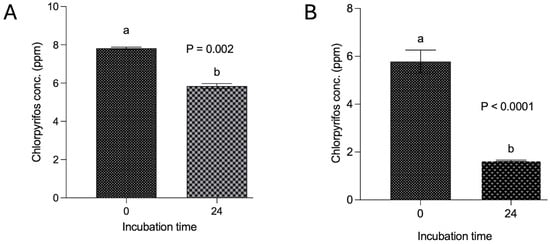
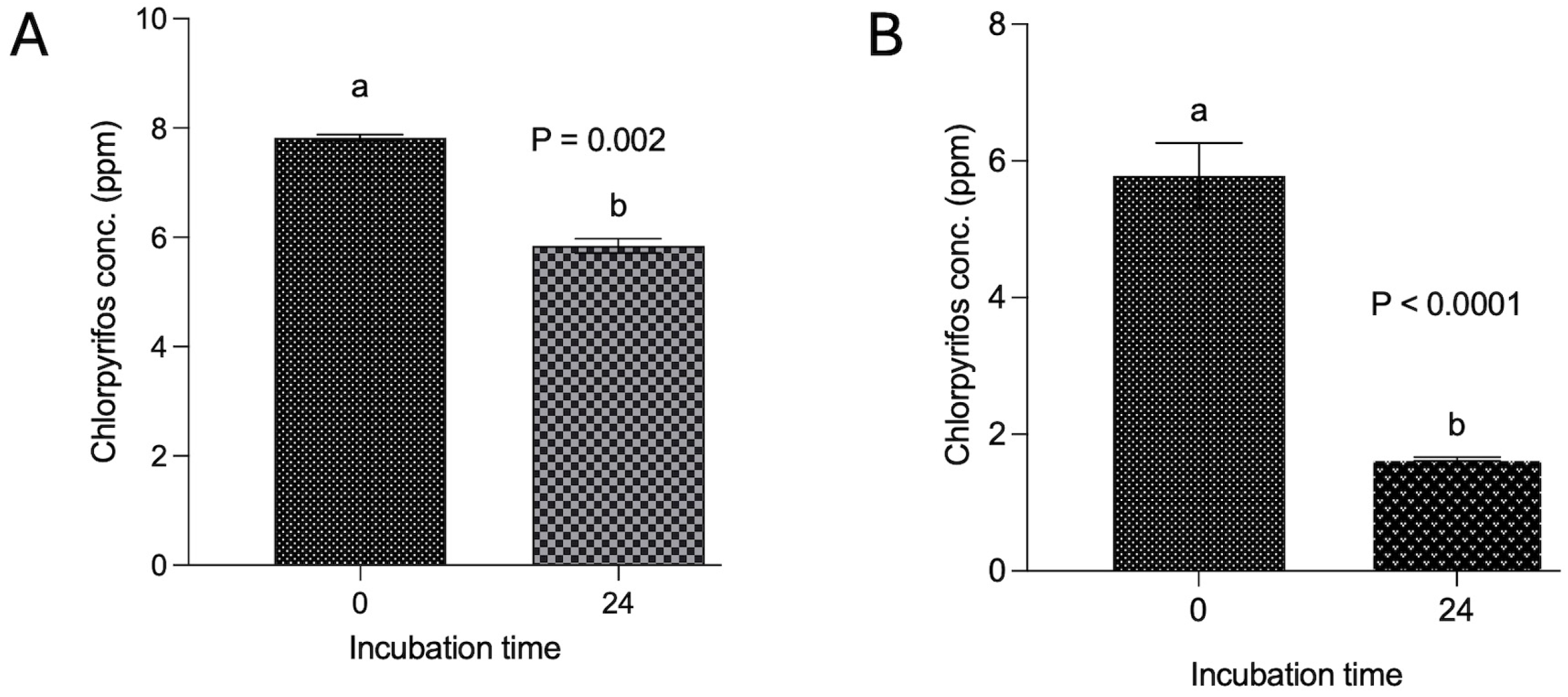
Figure 2.
Degradation of chlorpyrifos with and without methanol spiking over 24 h. Incubation in panel (A) was conducted by dissolving chlorpyrifos in methanol and then in vitro incubation along with buffered rumen fluid and 200 mg feed over 24 h, whereas in panel (B), the incubation was achieved by dissolving chlorpyrifos in ruminal fluid without using methanol and then incubation along with buffered rumen fluid and a 200 mg feed sample. Chlorpyrifos concentrations bearing different superscripts a, b at different incubation hours represent the significant difference between the incubation hours.
Since the experiment I was conducted without feed to get an idea if the chlorpyrifos can be degraded in the ruminal fluid, which possesses anaerobic microbiota, the investigation was further extended to additional experiments with feed and chlorpyrifos either dissolved in methanol or ruminal fluid.
The findings from experiment II revealed a degradation of 25.4% of chlorpyrifos over the 24 h in vitro incubation with feed and buffered ruminal fluid (Figure 2A). The decrease in the chlorpyrifos concentration (ppm) at 24 h incubation was significantly (p < 0.002) lower than the initial concentration. However, the results from another experiment where chlorpyrifos was dissolved in the ruminal fluid itself without using methanol demonstrated a substantial decrease (p < 0.0001) in the chlorpyrifos concentration from the initial 5.78 ppm to 1.64 ppm over 24 h incubation (Figure 2B). A significant (p < 0.0001) degradation of 71.6% was achieved by dissolving the chlorpyrifos in ruminal fluid and incubating with feed and ruminal fluid over 24 h.
3.2. Rumen Metagenome
In the present study, a total of 528 million sequencing reads of data were generated with an average of 44.1 million raw reads per sample (Supplementary Table S1). After quality filtration, on average 5.55% of reads per sample were dropped due to poor quality, short length, and adapter contamination. Further, 0.01% of the metagenomic reads per sample were dropped due to the host contamination removal. Subsequently, about 94.4% of the high-quality paired and single-end metagenomic reads were subjected to retrieve the information on compositional abundances.
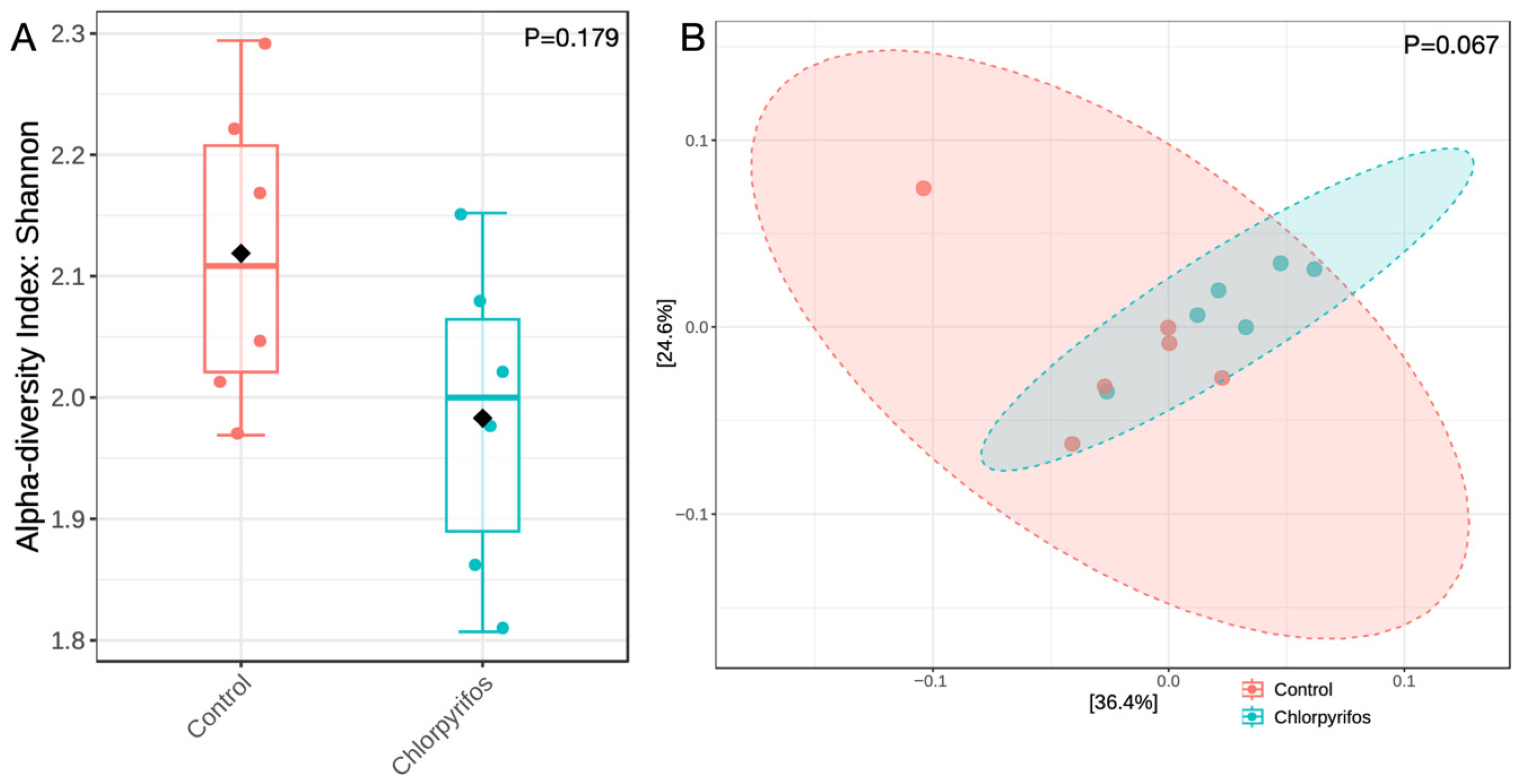
The similar alpha diversity index between the control and chlorpyrifos treatments revealed that the richness and the evenness of the microbial population were not affected (p = 0.179) by the contamination of rumen fluid with chlorpyrifos pesticide at a concentration of 5.78 ppm (Figure 3A). Similarly, beta diversity (Figure 3B) analyzed the differences in microbial composition between two treatments, and the non-significant p-value of 0.067 indicated that the microbiota composition was also not different in this study.
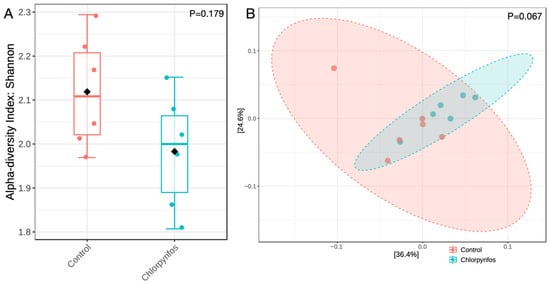
Figure 3.
Panel (A) represents the alpha diversity index of the rumen microbiota assessed through Shannon index method, whereas Panel (B) represents the beta diversity using the Bray-Curtis dissimilarity index. In control treatment, the in vitro incubation was set up using feed and buffered rumen fluid without chlorpyrifos, whereas another treatment consisted of feed, buffered rumen fluid, and chlorpyrifos for the in vitro incubation over 24 h. p-value in the panels indicates whether the alpha or beta diversity was different between the two groups.
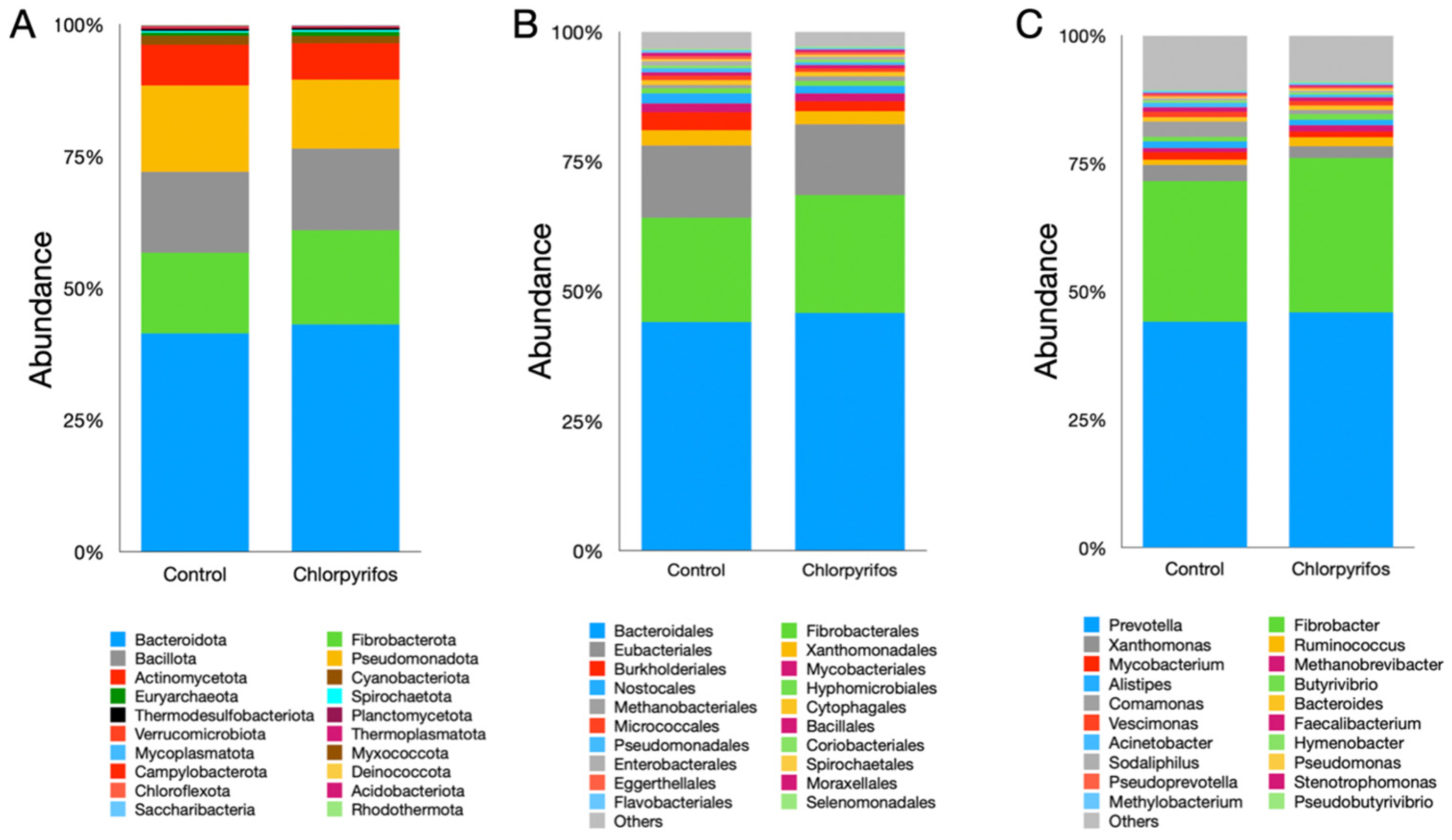
In this study, a total of 20 microbial phyla were identified (Supplementary Table S1), where Bacteroidota, Fibrobacterota, Bacillota, and Pseudomonadota constituted the largest fraction of the microbiota (Figure 4A). The Bacteroidota phylum alone represented more than 40% of the total microbiota; however, their abundance was similar (pFDR = 0.0711) between the two treatments. Similarly, the abundances of Fibrobacterota (pFDR = 0.0711), Bacillota (pFDR = 0.101), and Pseudomonadota (pFDR = 0.563) phyla also did not differ significantly between the treatments. The chlorpyrifos contamination in this study also did not exhibit any significant (pFDR = 0.0711) changes in the abundance of archaeal phylum Euryarchaeota. Likewise, the abundances of other microbial phyla were also found to be similar between the treatments (Figure 4A, Supplementary Table S1).
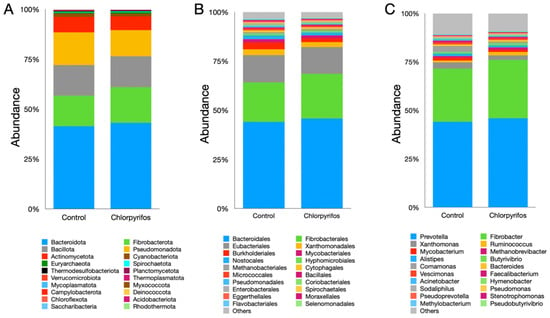
Figure 4.
Effect of contamination of ruminal fluid with chlorpyrifos on the compositional abundances of rumen microbiota. Panel (A) represents the microbial abundances at phylum level, panel (B) at the order, and panel (C) at the genus level. In control treatment, the in vitro incubation was set up using feed and buffered rumen fluid without chlorpyrifos, whereas another treatment consisted of feed, buffered rumen fluid, and chlorpyrifos for the in vitro incubation over 24 h.
A total of 57 microbial orders were identified in the present study (Figure 4B, Supplementary Table S1). Among the orders, the Bacteroidales represented the largest fraction; however, the chlorpyrifos contamination did not affect (pFDR = 0.120) their abundance between treatments. Similarly, the abundances of other prominent microbial orders such as Fibrobacterales (pFDR = 0.381), Eubacteriales (pFDR = 0.464), and Xanthomonadales (pFDR = 0.941) were also similar between the treatments and not affected by the chlorpyrifos contamination of the ruminal fluid. Likewise, the abundance of the prominent archaeal order Methanobacteriales was also similar (pFDR = 0.120) between the control and chlorpyrifos treatments. Overall, no significant difference in the microbiota at the order level was reported in the present study.
A metagenomic study identified a total of 120 microbial genera in the ruminal fluid. Among the genera, Prevotella was the single largest genus, which constituted 44–45% of the total ruminal microbiota (Figure 4C, Supplementary Table S1). The contamination of ruminal fluid with chlorpyrifos did not affect (pFDR = 0.120) the abundance of the Prevotella genus in the present study. The Fibrobacter (pFDR = 0.126) and Xanthomonas (pFDR = 0.162) were the next abundant microbial genera reported with a similar abundance between the two treatments. At the genus level, the abundance of Ruminococcus was significantly different (pFDR = 0.007) between the control and chlorpyrifos treatments (Figure 4C, Supplementary Table S1). Among the archaeal genera, Methanobrevibacter was the most prominent genus, which constituted 0.77–1.14% of the total microbiota; however, there was no significant difference (pFDR = 0.468) between the two treatments in their abundance. Similarly, the abundances of other genera also did not differ between the two treatments (Supplementary Table S1).
3.3. Gene Annotation
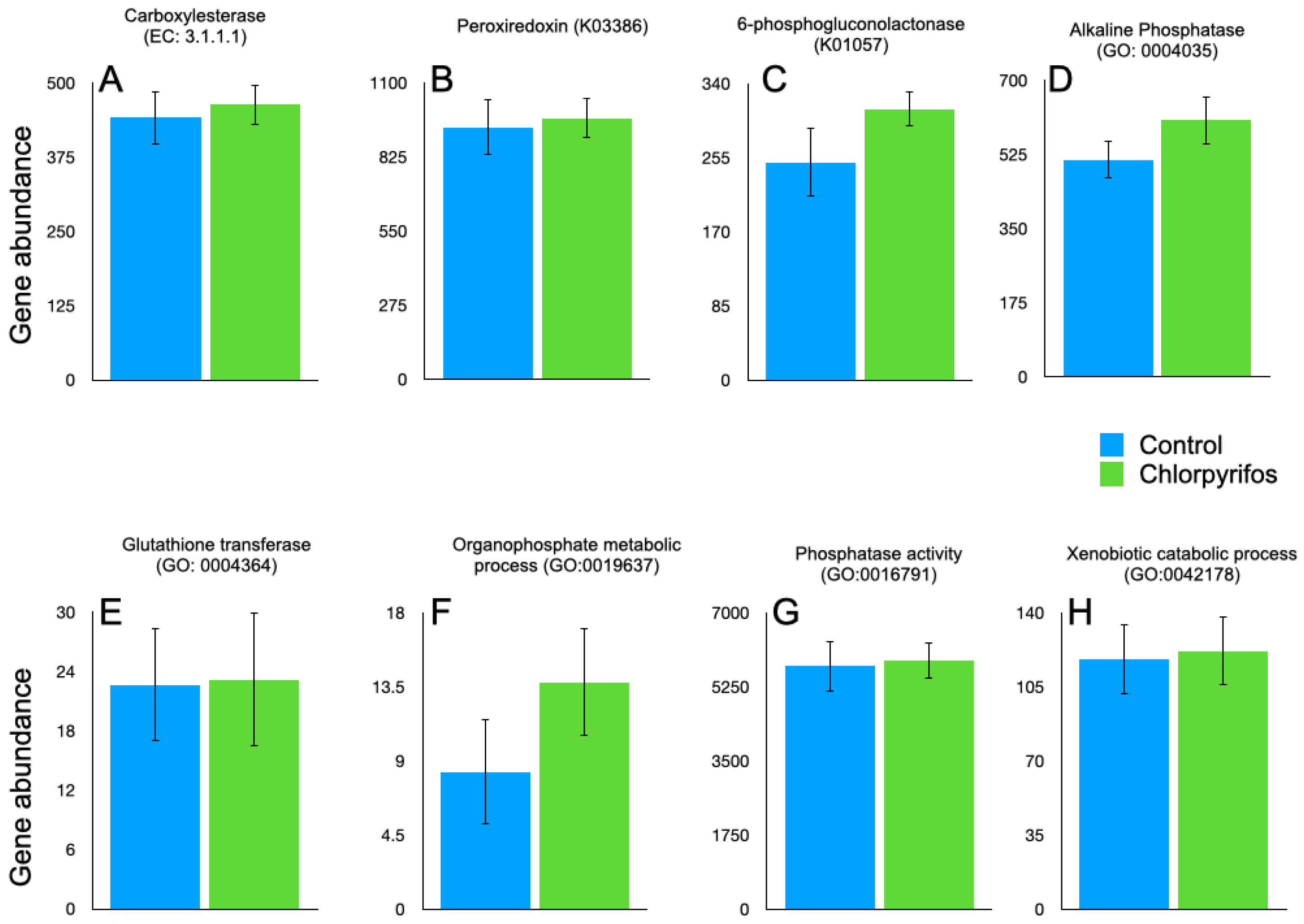
In annotated metagenome data, we have tried to identify the genes involved in the enzymatic degradation of chlorpyrifos with their EC numbers. The search was made for the previously reported EC 3.1.8.1 (organophosphorus hydrolase), EC 3.1.3.1 (phosphatase), EC 1.14.13.- (monooxygenases), EC 1.1.1.- (dehydrogenases), and EC 3.1.1.- (esterases). However, the genes affiliated with EC 3.1.8.1, EC 3.1.3.1, EC 1.14.13.-, and EC 1.1.1.- were not present in our metagenome data. In the present study, only carboxylesterase (EC 3.1.1.1) was present in the metagenome data (Figure 5). The gene abundances for EC 3.1.1.1 were 442 ± 44.1 and 464 ± 34.4, respectively, in the control and chlorpyrifos treatments. The gene abundances for EC 3.1.1.1 were numerically higher in the chlorpyrifos treatment but not significantly (p = 0.693) different from the control (Supplementary Table S1). The genes for carboxylesterase in the metagenome were contributed by the microbes affiliated with the two major phyla Bacteroidota and Firmicutes (Figure 6, Supplementary Table S1). At the order level, Bacteroidales, Oscillopirales, Lachnospirales, and Christensenellales contributed to the genes encoding carboxylesterase (Figure 6, Supplementary Table S1).
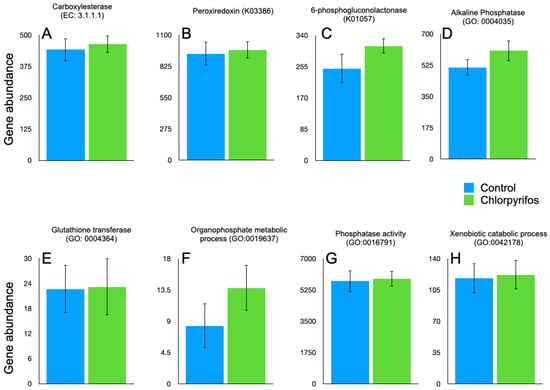
Figure 5.
Abundance of the EC, GO, and KOs associated with chlorpyrifos degradation in the rumen metagenome data. The abundances are calculated from the metagenomic contig coverage values estimated by mapping the sequencing data against the contigs, and the predicted genes from the contigs were mapped against the EC, KO, and GO databases. (A): abundance of carboxylesterase (EC 3.1.1.1); (B): peroxiredoxin (K03386); (C): 6-phosphogluconolactonase (K01057); (D): alkaline phosphatase (GO:0004035); (E): glutathione transferase (GO:0004364); (F): organophosphate metabolic process (GO:0019637); (G): phosphatase activity (GO:0016791); (H): xenobiotic catabolic process (GO:0042178).
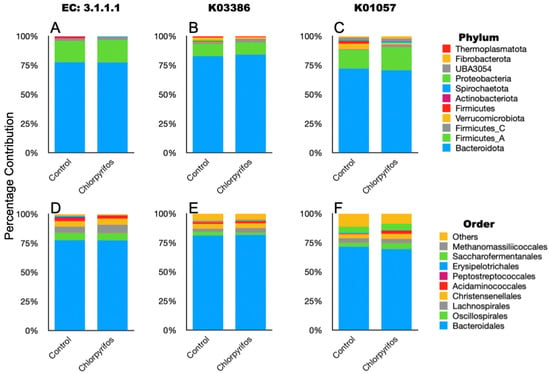
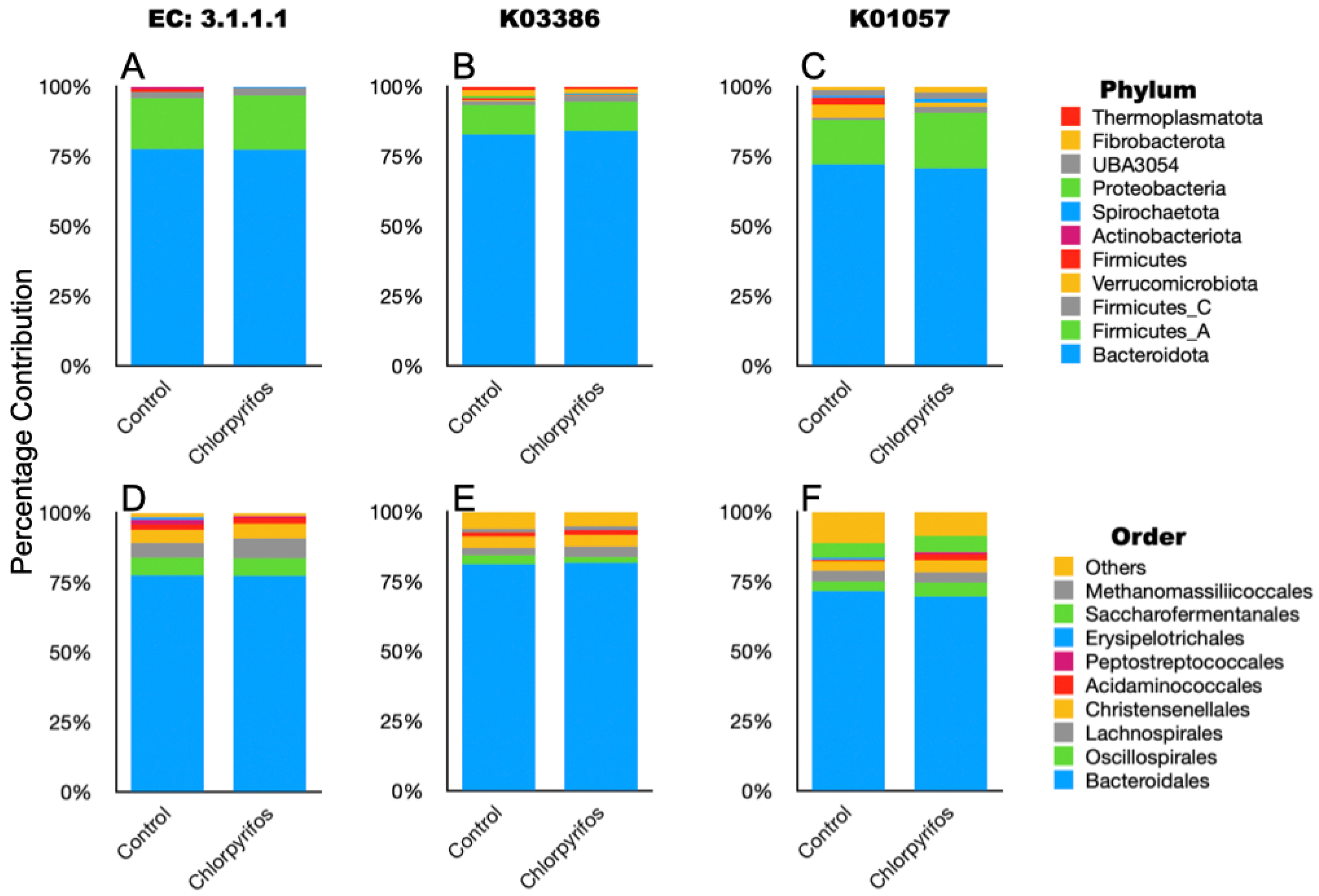
Figure 6.
Taxonomic composition of the ruminal microbes that contributed to the selected EC and KOs associated with chlorpyrifos degradation in the rumen metagenome data. (A): phylum-level composition of metagenome members contributed to EC 3.1.1.1; (B): phylum-level composition of metagenome members contributed to K03386; (C): phylum-level composition of metagenome members contributed to K01057; (D): order-level composition of metagenome members contributed to EC 3.1.1.1; (E): order-level composition of metagenome members contributed to K03386; (F): order-level composition of metagenome members contributed to K01057.
Based on the reported KEGG orthologs (KO) involved in the chlorpyrifos degradation, we searched the metagenome gene catalogue for a total of seven KO (K07099, K01077, K00540, K01093, K00128, K03386, and K01057) in the annotated metagenome data. The K07099, K01077, K00540, K01093, and K00128 that encode for organophosphorus hydrolase, phosphatase, monooxygenases, esterases, and dehydrogenases were not identified in this study. However, K03386 and K01057, encoding peroxiredoxin and 6-phosphogluconolactonase, respectively, were detected in the metagenome (Figure 5). The Bacteroidota, Firmicutes, Fibrobacterota, and Thermoplasmatota phyla contributed to the K03386, whereas the K01057 was mostly associated with the Bacteroidota, Firmicutes, Verrucomicrobiota, Fibrobacterota, and UBA3054 phyla (Figure 6, Supplementary Table S1).
In the metagenome, the gene ontology (GO) for the chlorpyrifos degradation, such as GO:0046434, GO:0006805, GO:0009636, GO:0016787, GO:0003824, GO:0004035, GO:0004364, GO:0019637, GO:0016791, and GO:0042178, was identified. The GO:0046434, GO:0006805, GO:0009636, GO:0016787, and GO:0003824 were absent in the metagenome. On the contrary, GO:0004035, GO:0004364, GO:0019637, GO:0016791, and GO:0042178 were present in the metagenome (Figure 5). The GO present in the metagenome encodes for the enzymes alkaline phosphatase, glutathione transferase, organophosphate metabolic process, phosphatase activity, and xenobiotic catabolic process, respectively. However, the abundances of these GO were similar between the treatments (Supplementary Table S1).
4. Discussion
Chlorpyrifos, a broad-spectrum organophosphorus pesticide, due to its wide use, is considered as one of the major feed and environmental contaminants [27]. The persistence of chlorpyrifos in soil is reported to be moderate to high [28] depending upon the soil type, soil pH, microbiota, temperature, and moisture [29], and due to low water solubility and a strong tendency for binding with organic matter, it can persist for quite a long time [30]. The pesticide is transferred from soil to the plants through root uptake, direct adsorption on plant surfaces, and contamination during application. Accumulation of chlorpyrifos in feed and fodder poses serious health implications, particularly to livestock. The contaminating pesticide residues in animal feed could be a source of pesticide residues in milk [31,32]. Although chlorpyrifos degradation is extensively studied in the soil [7,8,33], studies investigating the chlorpyrifos degradation in the rumen and its mechanism are still scanty.
The rumen is strictly an anaerobic compartment harboring diverse microbiota, including bacteria, protozoa, fungi, archaea, and bacteriophages. These microbes act syntrophically in the rumen ecosystem and perform important functions such as feed fermentation, volatile fatty acid production, microbial protein synthesis, and removal of fermentative gases [34,35]. In addition, the rumen microbial consortia are also reported to have metabolic capabilities for degrading the xenobiotic compounds, including pesticides, and the ability can be acquired by the ruminal microflora through continuous exposure [36] and can be transferred by transfaunation [37,38]. The ruminal environment can be ideal for anaerobic degradation of chlorpyrifos, as the degradation is effective over a narrow range (5.5–8.5) of pH [39] and temperature between 30 and 37 °C [40,41]. The degradation of chlorpyrifos up to a significant extent (71%) over 24 h incubation was demonstrated in the present study. This rate of degradation is faster than the anaerobic degradation of chlorpyrifos in soils [42]. Further, the degradation of chlorpyrifos in the rumen fluid inoculum in experiment I was 64% as compared to 71% in experiment II, where additional feed material was incubated with the chlorpyrifos-spiked buffered rumen fluid. This result is substantiated by in-soil degradation of chlorpyrifos; the degradation is faster in the presence of other nutrients that support faster growth of microbes [39,43]. While comparison of chlorpyrifos degradation in experiment II, wherein chlorpyrifos was solubilized in methanol versus directly in rumen fluid, revealed that the degradation over a 24 h period was less in incubation where methanol was used as a solvent (25.4) compared to the other incubation without using methanol as a solvent (71.6%). The presence of hydrocarbons/alcohols reduced the chlorpyrifos degradation [44]. Results of this study clearly established that the rumen microbes can degrade chlorpyrifos through the microbial enzymes. The degradation of chlorpyrifos due to the poor water solubility is not possible, as it is almost insoluble in water [45]. The microbial degradation of chlorpyrifos is in consonance with Megharaj et al. [46], who reported that the heat sterilization of rumen fluid lost the hydrolytic and degradation activity. Hydrolysis is the predominant mechanism of chlorpyrifos degradation [47]. The hydrolytic degradation of chlorpyrifos is majorly performed by the organophosphorus hydrolases through a nucleophilic strike on the phosphorus atom [48,49]. However, there is no report in the public domain confirming the occurrence of organophosphorus hydrolases and degradation of chlorpyrifos in the rumen. In addition, certain bacteria and fungi also have capabilities to perform the oxidative desulfuration of chlorpyrifos [50]. However, the anaerobic conditions of the rumen restrict the oxidative transformations, and therefore, the reductive dichlorination appears to be the complementary pathway for chlorpyrifos degradation in the anaerobic environment [51]. Further, the dichlorination is faster in anaerobic conditions [52]. The sulfate-reducing bacteria in the rumen [53] may also contribute to the reductive degradation of chlorpyrifos [54].
Pseudomonas have been reported as the most efficient degraders of chlorpyrifos through their organophosphorus hydrolases enzyme [55]. Further, Enterobacter [10] and Bacillus [56] are also potential degraders of the chlorpyrifos. The detection of the Pseudomonadota phylum in our metagenome confirmed that the chlorpyrifos-degrading Pseudomonas and Enterobacter were present in the rumen. Similarly, Bacillus affiliated with the phylum Bacillota (Firmicutes) was also detected in the rumen metagenome. The presence of chlorpyrifos-degrading microbial phylum indicated that the rumen possesses microbes capable of degrading chlorpyrifos pesticide. The absence of EC 3.1.8.1, EC 3.1.3.1, EC 1.14.13.-, and EC 1.1.1.- in the metagenome revealed that the organophosphorus hydrolase, phosphatase, monooxygenase, and dehydrogenase enzymes were not involved in the degradation of chlorpyrifos, and only EC 3.1.1.1 carried out the degradation of chlorpyrifos in the present study, as only the carboxylesterase was detected in the present study. Singh [57] reported that the carboxylesterases are important in degrading the xenobiotic, including organophosphorus pesticides. Carboxylesterases are reported to hydrolyze the phosphotriester bond in chlorpyrifos molecules and produce less toxic alcohols and acids [58,59]. The substantiative evidence of chlorpyrifos degradation by carboxylesterase was established by Wang et al. [60]; the authors have shown that the immobilized carboxylesterases degraded 90% of chlorpyrifos content in the farmland water. Further, the presence of GO:0004035, GO:0004364, GO:0019637, GO:0016791, and GO:0042178 in the metagenome strengthens that the chlorpyrifos degradation in the present study was primarily assigned to the rumen microbiota. These GO categories are related to the enzymes alkaline phosphatase, glutathione transferase, organophosphate metabolic process, phosphatase activity, and xenobiotic catabolic process, which is associated with the chlorpyrifos degradation.
Out of the seven KOs, two (K03386 and K01057) were detected in the annotated rumen metagenome. Therefore, the KO for organophosphorus hydrolase, phosphatase, monooxygenase, esterases, and dehydrogenases remains undetected in the metagenome. The two Kos, i.e., K03386 and K01057, detected in the metagenome encode for peroxiredoxin and 6-phosphogluconolactonase, respectively. During the bacterial degradation of chlorpyrifos, the enzyme peroxiredoxin helps in neutralizing the reactive oxygen species [61] so that the detoxification process is continued. In the metagenome, the major contributors to the peroxiredoxin were Bacteroidota, Firmicutes, Fibrobacterota, and Thermoplasmatota phyla, whereas Bacteroidota, Firmicutes, Verrucomicrobiota, Fibrobacterota, and UBA3054 phyla contributed to the K01057.
This in vitro study provides insights into the rumen microbiota involved in the chlorpyrifos degradation and the initial clue that the rumen microbes are capable of degrading chlorpyrifos. However, studies with the pure microbial isolates of rumen origin affiliated with the phyla Pseudomonadota and Bacillota are warranted to determine the efficiency of different microbes for chlorpyrifos degradation. Apart from the bacteria and archaea, the ruminal microbiome also contains protozoa and anaerobic fungi. We did not come across any report on protozoal degradation of chlorpyrifos. Although studies in soil and other environments did report aerobic fungi in the degradation of chlorpyrifos and TCP. The rumen contains only anaerobic fungi belonging to the phylum Neocallimastigomycota, and the fungi are limited to only 6 genera, namely, Neocallimastix, Piromyces, Orpinomyces, Caecomyces, Cyllamyces, and Anaeromyces. No report is available on the chlorpyrifos degradation by the ruminal fungi. Further, the animal studies in different species with the variable levels of chlorpyrifos are also warranted to confirm the efficacy of rumen microbes in mixed syntrophy and also determine the threshold capabilities of the ruminal microbes.
5. Conclusions
From the in vitro study, it can be inferred that the chlorpyrifos pesticide can be degraded in the rumen to a significant extent. Rumen microbiota affiliated with Pseudomonadota and Bacillota are actively involved in the ruminal degradation of chlorpyrifos via carboxylesterase (EC 3.1.1.1). Since the concentration of the chlorpyrifos used in the study did not influence the microbial diversity or composition between the control and chlorpyrifos-spiked samples, it indicates the innate ability of the ruminal microbiome to degrade the chlorpyrifos. This study was restricted to an in vitro setup; hence, the animal studies in different species with the variable levels of chlorpyrifos are also warranted to confirm the efficacy of rumen microbes in mixed syntrophy and also determine the threshold capabilities of the ruminal microbes. The metagenomic approach can only reveal the presence of degrading genes; however, their actual expression needs to be evaluated through the metatranscriptomic approach. The metabolites of the chlorpyrifos degradation were not estimated in the present study, which limits the identification of the actual chlorpyrifos degradation mechanism in rumen. Tracing the chlorpyrifos degradation metabolites in different organs and milk would be helpful in establishing the ability of ruminants to offset the chlorpyrifos or its residue contamination in the animal-origin products.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microorganisms14030581/s1, Table S1: Read stats, taxonomic affiliation, and functional analysis data of the microbiota.
Author Contributions
Conceptualization, P.K.M.; methodology, P.K.M.; software, A.P.K.; validation, P.K.M. and A.P.K.; formal analysis, P.K.M.; investigation, S.T. and A.M.; resources, A.S.; data curation, S.T. and A.P.K.; writing—original draft preparation, P.K.M.; writing—review and editing, A.S. and R.B.; visualization, S.T. and A.M.; supervision, P.K.M.; project administration, P.K.M.; funding acquisition, P.K.M. and A.S. All authors have read and agreed to the published version of the manuscript.
Funding
The work was carried out under the institute research project ‘Effect of graded chlorpyrifos on rumen fermentation, microbial degradation, and bioaccumulation in visceral organs in lactating goats’. The research received no external funding.
Institutional Review Board Statement
The necessary approval for conducting the studies and collecting rumen fluid was obtained from the Institute Animal Ethics Committee (IAEC) and the Committee for Control and Supervision of Experiments on Animals (CPCSEA), approval no. V-11011(13)/10/2024-CPCSEA-DADF dated 14 October 2024. The donor animals for the ruminal fluids were handled in strict adherence to the procedure laid out by the IAEC.
Informed Consent Statement
Not applicable.
Data Availability Statement
The datasets presented in this study can be found in online repositories with the accession number PRJNA1393601. The metagenome data with accession number(s) is available in the repository/repositories and can be found at: https://www.ncbi.nlm.nih.gov/sra/PRJNA1393601, accessed on 25 December 2025.
Conflicts of Interest
The authors declare no conflicts of interest, and the research was conducted in the absence of any commercial or financial relationships that could be construed as potential conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| KO | KEGG Orthologs |
| GO | Gene Ontology |
| EC | Enzyme Commission (numerical classification of enzymes based on chemical reactions they catalyze) |
| QuECHERS | Quick, Easy, Cheap, Effective, Rugged, and Safe, sample preparation for GC-MS |
| ppm | Parts per million (mg/kg) |
| pFDR | False Discovery Rate corrected p value |
References
- FAO. Pesticides Use and Trade—1990–2022; FAOSTAT Analytical Briefs, No. 89; FAO: Rome, Italy, 2024; Volume 89. [Google Scholar]
- Christensen, K.; Harper, B.; Luukinen, B.; Buhl, K.; Stone, D. Chlorpyrifos Technical Fact Sheet; National Pesticide Information Center, Oregon State University Extension Services. Available online: https://npic.orst.edu/factsheets/archive/chlorptech.html (accessed on 24 January 2026).
- Mohan, V. Global Agreement Reached to Eliminate Hazardous Pesticide Chlorpyrifos, India to Use It for Another Five Years. Times of India, 6 May 2025. [Google Scholar]
- Ministry of Agriculture & Farmers Welfare. Banning of Pesticides; Press Information Bureau: New Delhi, India, 2023.
- Li, R.; Wei, W.; He, L.; Hao, L.; Ji, X.; Zhou, Y.; Wang, Q. Chlorpyrifos Residual Behaviors in Field Crops and Transfers during Duck Pellet Feed Processing. J. Agric. Food Chem. 2014, 62, 10215–10221. [Google Scholar] [CrossRef] [PubMed]
- Zu’Amah, H.; Harsanti, E.S.; Wihardjaka, A.; Sukarjo; Ardiwinata, A.N. Distribution of Chlorpyrifos Residue in Maize (Zea mays). In Proceedings of the IOP Conference Series: Earth and Environmental Science; Institute of Physics: Bristol, UK, 2023; Volume 1230. [Google Scholar]
- Farhan, M.; Ahmad, M.; Kanwal, A.; Butt, Z.A.; Khan, Q.F.; Raza, S.A.; Qayyum, H.; Wahid, A. Biodegradation of Chlorpyrifos Using Isolates from Contaminated Agricultural Soil, Its Kinetic Studies. Sci. Rep. 2021, 11, 10320. [Google Scholar] [CrossRef] [PubMed]
- Akbar, S.; Sultan, S. Soil Bacteria Showing a Potential of Chlorpyrifos Degradation and Plant Growth Enhancement. Braz. J. Microbiol. 2016, 47, 563–570. [Google Scholar] [CrossRef]
- Saengsanga, T.; Phakratok, N. Biodegradation of Chlorpyrifos by Soil Bacteria and Their Effects on Growth of Rice Seedlings under Pesticide-Contaminated Soil. Plant Soil Environ. 2023, 69, 210–220. [Google Scholar] [CrossRef]
- Singh, B.K.; Walker, A.; Morgan, J.A.W.; Wright, D.J. Biodegradation of Chlorpyrifos by Enterobacter Strain B-14 and Its Use in Bioremediation of Contaminated Soils. Appl. Environ. Microbiol. 2004, 70, 4855–4863. [Google Scholar] [CrossRef]
- Minj, K.; Sohal, J.K. Isolation and Characterization of Chlorpyrifos Degrading Microorganisms from Agriculture Soil: A Review. Curr. Agric. Res. J. 2025, 13, 443–452. [Google Scholar] [CrossRef]
- Ge, J.; Fu, W.; Bai, M.; Zhang, L.; Guo, B.; Qiao, Q.; Tao, R.; Kou, J. The Degradation of Residual Pesticides and the Quality of White Clover Silage Are Related to the Types and Initial Concentrations of Pesticides. J. Pestic. Sci. 2021, 46, 342–351. [Google Scholar] [CrossRef]
- Wang, Y.S.; Wu, T.H.; Yang, Y.; Zhu, C.L.; Ding, C.L.; Dai, C.C. Binding and Detoxification of Chlorpyrifos by Lactic Acid Bacteria on Rice Straw Silage Fermentation. J. Environ. Sci. Health Part B 2016, 51, 316–325. [Google Scholar]
- Attwood, G.T.; Wakelin, S.A.; Leahy, S.C.; Rowe, S.; Clarke, S.; Chapman, D.F.; Muirhead, R.; Jacobs, J.M.E. Applications of the Soil, Plant and Rumen Microbiomes in Pastoral Agriculture. Front. Nutr. 2019, 6, 107. [Google Scholar] [CrossRef]
- Perumbakkam, S.; Craig, A.M. Anaerobic Transformation of Octahydro-1,3,5,7-Tetranitro-1,3,5,7-Tetrazocine (HMX) by Ovine Rumen Microorganisms. Res. Microbiol. 2012, 163, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Morrison, M. Improved Extraction of PCR-Quality Community DNA from Digesta and Fecal Samples. Biotechniques 2004, 36, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Chen, S. Ultrafast One-Pass FASTQ Data Preprocessing, Quality Control, and Deduplication Using Fastp. iMeta 2023, 2, e107. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Olson, R.D.; Assaf, R.; Brettin, T.; Conrad, N.; Cucinell, C.; Davis, J.J.; Dempsey, D.M.; Dickerman, A.; Dietrich, E.M.; Kenyon, R.W.; et al. Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): A Resource Combining PATRIC, IRD and ViPR. Nucleic Acids Res. 2023, 51, D678–D689. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved Metagenomic Analysis with Kraken 2. Genome Biol. 2019, 20, 257. [Google Scholar] [CrossRef]
- Breitwieser, F.P.; Salzberg, S.L. Pavian: Interactive Analysis of Metagenomics Data for Microbiome Studies and Pathogen Identification. Bioinformatics 2020, 36, 1303–1304. [Google Scholar] [CrossRef]
- Chong, J.; Liu, P.; Zhou, G.; Xia, J. Using MicrobiomeAnalyst for Comprehensive Statistical, Functional, and Meta-Analysis of Microbiome Data. Nat. Protoc. 2020, 15, 799–821. [Google Scholar] [CrossRef]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic Biomarker Discovery and Explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef]
- Dong, X.; Strous, M. An Integrated Pipeline for Annotation and Visualization of Metagenomic Contigs. Front. Genet. 2019, 10, 999. [Google Scholar] [CrossRef]
- Li, D.; Liu, C.M.; Luo, R.; Sadakane, K.; Lam, T.W. MEGAHIT: An Ultra-Fast Single-Node Solution for Large and Complex Metagenomics Assembly via Succinct de Bruijn Graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Bushnell, B. BBMap: A Fast, Accurate, Splice-Aware Aligner; Lawrence Berkeley National Laboratory LBL Publications: Berkeley, CA, USA, 2014. [Google Scholar]
- Tudi, M.; Yang, L.; Wang, L.; Lv, J.; Gu, L.; Li, H.; Peng, W.; Yu, Q.; Ruan, H.; Li, Q.; et al. Environmental and Human Health Hazards from Chlorpyrifos, Pymetrozine and Avermectin Application in China under a Climate Change Scenario: A Comprehensive Review. Agriculture 2023, 13, 1683. [Google Scholar] [CrossRef]
- Chai, L.K.; Wong, M.H.; Hansen, H.C.B. Degradation of Chlorpyrifos in Humid Tropical Soils. J. Environ. Manag. 2013, 125, 28–32. [Google Scholar] [CrossRef] [PubMed]
- Kamrin, M.A. Pesticide Profiles: Toxicity, Environmental Impact, and Fate; Kamrin, M.A., Ed.; CRC Press: Boca Raton, FL, USA; Lewis Publishers: New York, NY, USA, 1997. [Google Scholar]
- Lu, M.X.; Jiang, W.W.; Wang, J.L.; Jian, Q.; Shen, Y.; Liu, X.J.; Yu, X.Y. Persistence and Dissipation of Chlorpyrifos in Brassica Chinensis, Lettuce, Celery, Asparagus Lettuce, Eggplant, and Pepper in a Greenhouse. PLoS ONE 2014, 9, e100556. [Google Scholar] [CrossRef] [PubMed]
- Bedi, J.S.; Gill, J.P.S.; Kaur, P.; Aulakh, R.S. Pesticide Residues in Milk and Their Relationship with Pesticide Contamination of Feedstuffs Supplied to Dairy Cattle in Punjab (India). J. Anim. Feed Sci. 2018, 27, 18–25. [Google Scholar] [CrossRef]
- Yuan, S.; Yang, F.; Yu, H.; Xie, Y.; Guo, Y.; Yao, W. Degradation Mechanism and Toxicity Assessment of Chlorpyrifos in Milk by Combined Ultrasound and Ultraviolet Treatment. Food Chem. 2022, 383, 132550. [Google Scholar] [CrossRef]
- Singh, B.; Kaur, J.; Singh, K. Microbial Degradation of an Organophosphate Pesticide, Malathion. Crit. Rev. Microbiol. 2014, 40, 146–154. [Google Scholar] [CrossRef]
- Gagen, E.J.; Denman, S.E.; McSweeney, C.S. Acetogenesis as an Alternative to Methanogenesis in Rumen. In Livestock Production and Climate Change; Malik, P.K., Bhatta, R., Takahashi, J., Kohn, R.A., Prasad, C.S., Eds.; CABI: Wallingford, Oxfordshire, UK, 2015; pp. 292–303. [Google Scholar]
- Malik, P.K.; Bhatta, R.; Soren, N.M.; Sejian, V.; Mech, A.; Prasad, K.S.; Prasad, C.S. Feed-Based Approaches in Enteric Methane Amelioration. In Livestock Production and Climate Change; Malik, P.K., Bhatta, R., Takahashi, J., Kohn, R.A., Prasad, C.S., Eds.; CABI: Wallingford, Oxfordshire, UK, 2015; pp. 336–359. [Google Scholar]
- Loh, Z.H.; Ouwerkerk, D.; Klieve, A.V.; Hungerford, N.L.; Fletcher, M.T. Toxin Degradation by Rumen Microorganisms: A Review. Toxins 2020, 12, 664. [Google Scholar] [CrossRef]
- Jones, R.J.; Megarrity, R.G. Successful Transfer of DHP-Degrading Bacteria from Hawaiian Goats to Australian Ruminants to Overcome the Toxicity of Leucaena. Aust. Vet. J. 1986, 63, 259–262. [Google Scholar] [CrossRef]
- Duarte, A.L.; Maria, R.; Medeiros, T.; Riet-Correa, F. Poisoning by Amorimia spp. in Ruminants/Intoxicacao Por Amorimia spp. Em Ruminantes. Cienc. Rural 2013, 43, 1294. [Google Scholar]
- Yadav, M.; Shukla, A.K.; Srivastva, N.; Upadhyay, S.N.; Dubey, S.K. Utilization of Microbial Community Potential for Removal of Chlorpyrifos: A Review. Crit. Rev. Biotechnol. 2016, 36, 727–742. [Google Scholar] [CrossRef] [PubMed]
- Bosu, S.; Rajamohan, N.; Al Salti, S.; Rajasimman, M.; Das, P. Biodegradation of Chlorpyrifos Pollution from Contaminated Environment—A Review on Operating Variables and Mechanism. Environ. Res. 2024, 248, 118212. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Chen, S.; Hu, M.; Hu, Q.; Luo, J.; Li, Y. Purification and Characterization of a Novel Chlorpyrifos Hydrolase from Cladosporium cladosporioides Hu-01. PLoS ONE 2012, 7, e38137. [Google Scholar] [CrossRef] [PubMed]
- Vidya Lakshmi, C.; Kumar, M.; Khanna, S. Biodegradation of Chlorpyrifos in Soil by Enriched Cultures. Curr. Microbiol. 2009, 58, 35–38. [Google Scholar] [CrossRef]
- Castillo, M.D.P.; Torstensson, L. Effect of Biobed Composition Moisture and Temperature on the Degradation of Pesticides. J. Agric. Food Chem. 2007, 55, 5725–5733. [Google Scholar] [CrossRef]
- Khalid, S.; Hashmi, I.; Khan, S.J. Bacterial Assisted Degradation of Chlorpyrifos: The Key Role of Environmental Conditions, Trace Metals and Organic Solvents. J. Environ. Manag. 2016, 168, 1–9. [Google Scholar] [CrossRef]
- Rathod, A.L.; Garg, R.K. Chlorpyrifos Poisoning and Its Implications in Human Fatal Cases: A Forensic Perspective with Reference to Indian Scenario. J. Forensic Leg. Med. 2017, 47, 29–34. [Google Scholar] [CrossRef]
- Megharaj, M.; Ramakrishnan, B.; Venkateswarlu, K.; Sethunathan, N.; Naidu, R. Bioremediation Approaches for Organic Pollutants: A Critical Perspective. Environ. Int. 2011, 37, 1362–1375. [Google Scholar] [CrossRef]
- Chen, S.; Liu, C.; Peng, C.; Liu, H.; Hu, M.; Zhong, G. Biodegradation of Chlorpyrifos and Its Hydrolysis Product 3,5,6-Trichloro-2-Pyridinol by a New Fungal Strain Cladosporium cladosporioides Hu-01. PLoS ONE 2012, 7, e47205. [Google Scholar] [CrossRef]
- Shi, T.; Fang, L.; Qin, H.; Chen, Y.; Wu, X.; Hua, R. Rapid Biodegradation of the Organophosphorus Insecticide Chlorpyrifos by Cupriavidus nantongensis X1T. Int. J. Environ. Res. Public Health 2019, 16, 4593. [Google Scholar] [CrossRef]
- Li, C.; Ma, Y.; Mi, Z.; Huo, R.; Zhou, T.; Hai, H.; Kwok, L.Y.; Sun, Z.; Chen, Y.; Zhang, H. Screening for Lactobacillus plantarum Strains That Possess Organophosphorus Pesticide-Degrading Activity and Metabolomic Analysis of Phorate Degradation. Front. Microbiol. 2018, 9, 2048. [Google Scholar] [CrossRef]
- Chambers, J.E.; Chambers, H.W. Oxidative Desulfuration of Chlorpyrifos, Chlorpyrifos-Methyl, and Leptophos by Rat Brain and Liver. J. Biochem. Mol. Toxicol. 1989, 4, 201–203. [Google Scholar] [CrossRef] [PubMed]
- Dutta, N.; Usman, M.; Ashraf, M.A.; Luo, G.; Zhang, S. Efficacy of Emerging Technologies in Addressing Reductive Dechlorination for Environmental Bioremediation: A Review. J. Hazard. Mater. Lett. 2022, 3, 100065. [Google Scholar] [CrossRef]
- D’Angelo, E.; Reddy, K. Aerobic and Anaerobic Transformations of Pentachlorophenol in Wetland Soils. Soil Sci. Soc. Am. J. 2000, 64, 933–943. [Google Scholar] [CrossRef]
- Shah, N.S.; Khan, J.A.; Sayed, M.; Khan, Z.U.H.; Iqbal, J.; Imran, M.; Murtaza, B.; Zakir, A.; Polychronopoulou, K. Nano Zerovalent Zinc Catalyzed Peroxymonosulfate Based Advanced Oxidation Technologies for Treatment of Chlorpyrifos in Aqueous Solution: A Semi-Pilot Scale Study. J. Clean. Prod. 2020, 246, 119032. [Google Scholar] [CrossRef]
- Xu, Y.; Xue, L.; Ye, Q.; Franks, A.E.; Zhu, M.; Feng, X.; Xu, J.; He, Y. Inhibitory Effects of Sulfate and Nitrate Reduction on Reductive Dechlorination of PCP in a Flooded Paddy Soil. Front. Microbiol. 2018, 9, 567. [Google Scholar] [CrossRef]
- Singh, B.K.; Walker, A.; Wright, D.J. Bioremedial Potential of Fenamiphos and Chlorpyrifos Degrading Isolates: Influence of Different Environmental Conditions. Soil Biol. Biochem. 2006, 38, 2682–2693. [Google Scholar] [CrossRef]
- Anwar, S.; Liaquat, F.; Khan, Q.M.; Khalid, Z.M.; Iqbal, S. Biodegradation of Chlorpyrifos and Its Hydrolysis Product 3,5,6-Trichloro-2-Pyridinol by Bacillus pumilus Strain C2A1. J. Hazard. Mater. 2009, 168, 400–405. [Google Scholar] [CrossRef]
- Singh, B. Review on Microbial Carboxylesterase: General Properties and Role in Organophosphate Pesticides Degradation. Biochem. Mol. Biol. 2014, 2, 1–6. [Google Scholar] [CrossRef]
- Bhatt, P.; Bhatt, K.; Huang, Y.; Lin, Z.; Chen, S. Esterase Is a Powerful Tool for the Biodegradation of Pyrethroid Insecticides. Chemosphere 2020, 244, 125507. [Google Scholar] [CrossRef]
- Hatfield, M.J.; Umans, R.A.; Hyatt, J.L.; Edwards, C.C.; Wierdl, M.; Tsurkan, L.; Taylor, M.R.; Potter, P.M. Carboxylesterases: General Detoxifying Enzymes. Chem. Biol. Interact. 2016, 259, 327–331. [Google Scholar] [CrossRef]
- Wang, B.; Chen, J.; Wu, S.; Fang, J.; Li, Q.; Wang, G. Reusable Carboxylesterase Immobilized in ZIF for Efficient Degradation of Chlorpyrifos in Enviromental Water. Pestic. Biochem. Physiol. 2023, 194, 105519. [Google Scholar] [CrossRef]
- Shi, G.Q.; Zhang, Z.; Jia, K.L.; Zhang, K.; An, D.X.; Wang, G.; Zhang, B.L.; Yin, H.N. Characterization and Expression Analysis of Peroxiredoxin Family Genes from the Silkworm Bombyx mori in Response to Phoxim and Chlorpyrifos. Pestic. Biochem. Physiol. 2014, 114, 24–31. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.