Abstract
Blue mold of pome fruit, caused by Penicillium expansum, is controlled through postharvest applications of thiabendazole (TBZ), pyrimethanil (PYR), and fludioxonil (FDL). However, multi-fungicide-resistant isolates have emerged in the U.S. Pacific Northwest and their impact on decay control in long-term storage is unknown. This study evaluated the fitness of P. expansum isolates sensitive to all three postharvest fungicides (wild-types) and those resistant to TBZ (single-resistant), TBZ and PYR, or PYR and FDL (dual-resistant), and triple-resistant to the three fungicides. On nutrient-poor media, resistant isolates showed reduced conidial germination, whereas no significant differences were observed in germination, mycelial growth, or sporulation between phenotypes on nutrient-rich media at 1.5 and 20 °C. Regardless of their sensitivity phenotype, FDL-resistant isolates showed increased sensitivity to osmotic and oxidative stresses. Pathogenicity and virulence were not affected by the sensitivity phenotype on apples after six months of storage at 1.5 °C. Analysis of cumulative fitness changes indicated fitness loss under low-temperature in vitro and increased fitness under fungicide selection pressure on fruit in most resistant phenotypes. Gene expression analysis showed differential regulation of fitness-related genes, with most being up-regulated by TBZ. Overall, the results suggest that resistance in P. expansum may carry context-dependent fitness penalties, especially under high-stress conditions.
Keywords:
pome fruit; blue mold; fitness; gene expression; fludioxonil; pyrimethanil; thiabendazole; cumulative fitness 1. Introduction
Blue mold, caused by Penicillium expansum (Link) and other Penicillium spp., represents the most significant postharvest disease affecting pome fruit globally [1,2,3,4] and is estimated to account for approximately 50% and 30% of decayed apples and pears, respectively, in the U.S. Pacific Northwest (PNW) [5,6]. To effectively manage blue mold, rigorous sanitation practices in packinghouse facilities are essential to reduce the initial level of Penicillium spp. inoculum, which may significantly influence the incidence of blue mold during storage [7]. Alongside sanitation measures, chemical control remains the main approach for managing blue mold and other postharvest diseases. Thiabendazole (TBZ) has been used for over five decades, whereas pyrimethanil (PYR) and fludioxonil (FDL) have been used postharvest in packinghouses for over two decades in the PNW and other growing regions. Penicillium expansum is classified as a medium-risk fungus for fungicide resistance development [8].
The repeated application of fungicides with the same mode of action has led to the emergence of resistant fungal populations. Consequently, the resistance of P. expansum to TBZ (Fungicide Resistance Action Committee, FRAC 1), PYR (FRAC 9), and FDL (FRAC 12) has emerged in the PNW [9,10,11], as well as in other pome fruit growing regions [12,13,14]. The resistance of P. expansum to FRAC 1 fungicides was first documented in the late 1970s [15]. Specific mutations at codons 167, 198, and 200 in the target β-tubulin gene of P. expansum have been reported to confer resistance to TBZ [12,13,16,17]. Currently, the FRAC1 fungicides thiophanate-methyl and TBZ are still sprayed pre- and postharvest, respectively, in the PNW. In the early 2000s, the anilinopyrimidine PYR and the phenylpyrrole FDL were registered to control postharvest diseases in packinghouses. Although the precise mechanism of action of PYR against P. expansum is not adequately elucidated, it is posited to inhibit enzyme secretion in other fungal species [18,19], disrupt methionine biosynthesis [20], and interfere with mitochondrial functions [21]. Instances of PYR resistance in P. expansum from the PNW and Mid-Atlantic regions were first reported a decade after its registration [11,12] and have increased in subsequent years [9,22]. Resistance of P. expansum to cyprodinil, another FRAC 9 fungicide, was also reported in Greece [23,24]. By comparison, FDL is believed to induce the synthesis of polyols, specifically glycerol and mannitol, resulting in abnormal hyphal morphology and cell lysis [25,26]. Resistance to FDL has been reported in P. expansum isolates from the U.S. PNW and Mid-Atlantic regions [9,27] as well as in Greece [23,24]. Resistance to FDL in Penicillium spp. has been associated with deletions or substitutions in the OS-2 and NikA genes, a part of the group III histidine kinase of the high-osmolarity glycerol pathway [9,18,26,28,29]. The increasing prevalence of P. expansum populations resistant to TBZ, PYR, and FDL in packinghouses highlights the importance of comprehending how the evolution of fungicide resistance influences the relative fitness of these populations and its impact on decay management in long-term storage.
The fitness of filamentous fungi is attributed to their capacity to survive and reproduce in niches with a relatively low density of unoccupied space, while concurrently maximizing spore production [30]. Fluctuating impacts have been documented on the phenotypic fitness of P. expansum in relation to the development of fungicide resistance. Thus, isolates with single resistance to TBZ increased sporulation in vitro [9] and increased virulence on detached pears [12]. Conversely, PYR-resistant isolates of P. expansum demonstrated no significant differences in virulence on detached apples compared to PYR-sensitive isolates [13]. Laboratory mutants of P. expansum with single resistance to FDL showed decreased growth and increased sensitivity to osmotic stress in vitro, as well as decreased virulence and sporulation on apples [31]. Moreover, P. expansum field isolates with dual- or multiple-resistance to several preharvest fungicides used in Greek apple orchards displayed significantly decreased mycelial growth and pathogenicity on detached apples in short-term storage at room temperature, but their sporulation was greater than that of the wild-type isolates [23,24].
Fitness in filamentous fungi is often assessed at the phenotypic level by measuring traits such as spore germination, mycelial growth, sporulation, pathogenicity, virulence, and mycotoxin production. However, fitness metrics are regulated by global transcription factors that coordinate key physiological processes during infection. For example, LaeA regulates secondary metabolism such as patulin production and forms a complex with VeA to control virulence and development [32,33]. The velvet protein family, comprising VeA, VelB, VosA, and VelC, governs conidiation, secondary metabolism, and stress responses. VeA mutants exhibited defects in spore formation and failed to produce patulin and citrinin [32]. Additionally, Blistering1 plays a pivotal role in virulence, mediating vesicle-based secretion of cell wall-degrading enzymes. Mutants of P. expansum deficient of Blistering1 exhibited reduced virulence on apples [34]. Meanwhile, the transcription factor brlA is essential for conidiophore development and required for asexual reproduction. P. expansum mutants deficient in brlA lacked spore production but showed increased virulence [35]. In contrast, Pdac1, an adenylyl cyclase, regulates hyphal growth, virulence, and spore germination. Pdac1-deficient mutants produced heat-sensitive conidia with delayed germination [36]. Stress response pathways are also critical for Penicillium spp. pathogenicity. OS-2, homologous to Hog1, is a mitogen-activated protein kinase involved in osmotic stress tolerance. In P. digitatum, OS-2 mutants exhibited reduced mycelial growth and glycerol production under salt and sorbitol stress and showed altered sensitivity to fludioxonil [25,26]. However, in P. expansum, the role of OS-2 in osmotic stress response is less clear [37]. Additionally, the NADPH oxidase gene PeRacA regulates reactive oxygen species (ROS) production, growth, and pathogenicity. PeRacA-deficient mutants showed reduced growth under oxidative stress, highlighting the gene’s role in stress tolerance and infection success [38]. Collectively, these regulatory networks demonstrate the complex interplay between fungicide resistance, stress adaptation, and virulence in Penicillium spp. Assessing how fungicide resistance evolution affects the expression of key genes in P. expansum populations with different fungicide sensitivity phenotypes would provide valuable insights into phenotypic fitness and support the development of sustainable blue mold management strategies in pome fruit. Therefore, this study aimed to evaluate (i) fitness capabilities in vitro, (ii) virulence, sporulation and resistance stability on apples, and (iii) relative expression of several fitness-related genes and transcription factors in P. expansum isolates with single, dual, and triple resistance to TBZ, PYR, and FDL associated with blue mold outbreaks in packinghouses in the U.S. PNW.
2. Materials and Methods
2.1. Selection of Penicillium expansum Isolates with Different Fungicide Sensitivity Phenotypes
Fifteen P. expansum isolates with different fungicide sensitivity phenotypes were used in this study (Table 1). The isolates were obtained from decayed apples and pears collected from multiple warehouses in Washington State and Oregon between 2016 and 2018. The isolates were single-spored, stored as spore suspensions in 20% glycerol at −80 °C, and reactivated on potato dextrose agar (PDA) plates for 7 days at 20 °C prior to each experiment. The single-spored Penicillium isolates were characterized to the species level using a multi-locus sequence analysis as described previously [9,10,39].

Table 1.
Characteristics of Penicillium expansum isolates tested in this study.
The sensitivities of the single-spored isolates to TBZ (Mertect 340F, 42.3% a.i., Syngenta Crop Protection, Research Triangle, Greensboro, NC, USA) and FDL (Scholar, 20.46% a.i., Syngenta Crop Protection, Greensboro, NC, USA) were evaluated on PDA amended with 10 and 0.5 µg mL−1, respectively, and on sucrose agar (SA) amended with 0.5 µg mL−1 of PYR (Shield-Brite Penbotec 400 SC, 37.14% a.i., Pace International, Wapato, WA, USA). These doses were previously developed to discriminate resistant from sensitive isolates [40,41]. Non-amended and fungicide-amended plates were inoculated with three 10 µL droplets of spore suspensions of each isolate at 105 conidia mL−1 in 0.05% Tween-20 and incubated for 24 h at 20 °C. Concentrations of spore suspensions were determined using a hemacytometer (Hausser Scientific, Horsham, PA, USA). Two replicate plates (six values) were used for each isolate and fungicide combination. The germination and length of the germ tubes were determined microscopically, and data were used to calculate percentage germination. Conidia exhibiting germ tubes that were twice as long as the conidium diameter indicated resistance to the specific fungicide. The trial was conducted once at the onset of each parameter investigation.
Phenotyping yielded three isolates sensitive to all three postharvest fungicides, three with single resistance to TBZ (TBZR), three isolates with dual resistance to TBZ and PYR (TBZRPYRR), three with dual resistance to PYR and FDL (PYRRFDLR), and three isolates with triple resistance (TBZRPYRRFDLR) to all three fungicides (Table 1). Additional screenings to identify P. expansum isolates with single resistance to PYR (PYRR) and FDL (FDLR) or dual TBZ-FDL resistance (TBZRFDLR) were unsuccessful.
2.2. Evaluation of Fitness In Vitro
Six fitness parameters, i.e., conidial germination, mycelial growth, sporulation, sensitivity to osmotic stress (OSS), and reactive oxygen species (ROS), as well as the stability of the sensitivity phenotypes, were evaluated for the 15 P. expansum isolates at two distinct temperatures to assess fitness capabilities of the isolates under various conditions. Conidial germination was assessed on PDA, 2% water agar (WA), and intermediate medium (IM; 2 g MgSO4·7H2O, 4 g glucose, 4 g peptone, 0.75 g KH2PO4, 2 g citric acid, 12 g agar, and 0.2 mL Tween 20 in 1 L of distilled water) [38]. Three 10 µL droplets of each isolate, at 105 spores mL−1, were deposited equidistantly in a triangular arrangement on each plate and the plates were sealed with parafilm and incubated in the dark at 1.5 or 20 °C. Spore germination was assessed after 24 h on PDA and IM and after 48 h on WA at 20 °C. Germination at 1.5 °C was assessed after 24, 48, 72, and 96 h on PDA and IM and up to 10 days on WA. A total of 100 spores were examined by placing the plates upside down under a microscope and data were used to calculate the percentage of germination.
The sealed plates used for germination assessment were used to assess mycelial growth except on WA. Colony diameter was measured after 5 and 10 days of incubation at 20 °C and at 20-day intervals for up to 120 days at 1.5 °C in the dark. The sporulation on PDA and IM at 20 and 1.5 °C was assessed after 15 and 120 days, respectively, on the same plates used for mycelial growth assay. Ten milliliters of sterile distilled water with 0.05% Tween 20 were added to each plate and scraped using a sterile inoculation loop. Spore concentration was determined using a hemacytometer (Hausser Scientific, Horsham, PA). The sensitivities to OSS and ROS were assessed at 20 and 1.5 °C on PDA amended with 12% NaCl and 30 mM of methyl viologen (Mv, Acros Organics, Fair Lawn, NJ, USA), respectively, following the methodology used for the mycelial growth assay. For the above fitness parameters, three replicate plates were used for each isolate, medium, and temperature treatment combination across two trials.
The stability of fungicide sensitivity phenotypes of the 15 P. expansum isolates in the absence of fungicide selection pressure was assessed on PDA at 1.5 and 20 °C. In the 20 °C trial, conidia were prepared as described previously [42] and were transferred weekly to fresh PDA for 10 weeks. Due to the absence of sporulation at 1.5 °C, stability was evaluated through monthly transfers of 5 mm mycelial plugs taken from the edge of 30-day-old colonies grown on PDA. The plugs were placed upside down on fresh PDA plates and transferred for 10 consecutive months. Isolates subjected to 10 weekly transfers at 20 °C or 10 monthly transfers at 1.5 °C were designated as Gw10 and Gm10, respectively, while G0 referred to the original isolates before any transfer. To assess potential shifts in fungicide sensitivity between G0 and G10, the effective concentrations required to inhibit 50% germination and growth (EC50) were determined. Sensitivity to TBZ and FDL was assessed on PDA amended with TBZ at 0.0, 1, 5, 10, 50, 100, and 500 µg mL−1 and FDL at 0.0, 0.05, 0.5, 1, 5, 10, and 50 µg mL−1. Pyrimethanil sensitivity was assessed through germination inhibition on SA and mycelial growth inhibition on L-Asparagine agar (1 g K2HPO4; 1 g MgSO4; 0.5 g FeSO4·7H2O; 2 g L-Asparagine; 2 g glucose; 22 g agar) amended with PYR at 0.0, 0.05, 0.5, 1, 5, 10.0, and 50 µg mL−1. Three replicate plates were used for each isolate and the test was conducted once. Plate inoculation was conducted following previously described methods [9,43], spore germination and mycelial growth were evaluated after 24 h and 5 days, respectively, and data were used to calculate EC50 values as described previously [9,10].
2.3. Evaluation of Fitness on Apple Fruit
Three fitness parameters, i.e., virulence, sporulation, and stability of sensitivity phenotypes, were evaluated on detached apples in the absence and presence of fungicide selection pressure during prolonged cold storage to mimic conditions typically found in commercial packinghouses. Fitness parameters were evaluated in ten P. expansum isolates, i.e., two isolates representing each phenotype (Table 1).
Fuji apples harvested at commercial maturity in October 2022 and 2023 were sterilized and prepared for inoculation as described previously [43]. For trials without selection pressure, punctured apples (two punctures 4 mm wide and 4 mm deep per fruit) were inoculated with 20 µL of spore suspensions at 105 spores mL−1. To assess the virulence and sporulation under selection pressure, punctured fruits were dipped in suspensions of Mertect 340F (TBZ), Scholar SC (FDL), or Shield-Brite Penbotec 400 SC (PYR) at the label rate of 1.25 mL L−1 for 1 min, allowed to dry for 4 h, and then inoculated with 20 µL of spore suspensions at 105 spores mL−1. For each isolate, inoculation method, and year, four replicates, consisting of three apples each, were used. Inoculated apples were placed in clean plastic clamshells, each holding 12 fruits and stored at 1.5 °C. Blue mold incidence was monitored, and lesion diameters were measured in two perpendicular directions monthly up to 6 months. Sporulation was assessed after 6 months by resuspending the spores collected from the lesions of each fruit in 20 mL of sterile distilled water containing 0.01% Tween 20. The suspensions were filtered through cheesecloth, diluted appropriately, and used to enumerate spores in triplicate using a hemacytometer.
The stability of sensitivity phenotypes in vivo was assessed with and without selection pressure conditions on Fuji apples prepared and treated as described above. For each selection pressure condition, three apples with two inoculated punctures each were used for each isolate in 12-fruit clamshells, and the fruits were stored at 1.5 °C in a regular atmosphere. Every 60 days, a plug was dissected from the inner side of the decayed lesion and used to inoculate surface sterilized and punctured apples. This process was repeated four times, using apples that had been punctured and treated with TBZ, PYR, and FDL at 1.25 mL L−1, and a control treatment with water. At each transfer, the diameter of the lesions was measured. At the fourth and final transfer (eight months; Gm8), spores were collected from the lesions using a sterile cotton swab. The sensitivity of the Gm8 isolates to TBZ, PYR, and FDL was evaluated using the discriminatory doses used above.
2.4. Expression Analysis of Fitness-Related Genes in P. expansum Isolates
Seven fitness-related target genes were selected, i.e., Ac1 (PEX2_003910), Blistering1 (PEX2_008940), BrlA (PEX2_076900), LaeA (PEX2_005650), OS-2 (PEX2_061660), RacA (PEX2_19970), and VeA (PEX2_043190). These genes have been documented to influence various fitness parameters examined in this study. To measure gene expression relative to two reference genes, 28s (NG_069649) and CaM (DQ911134), the 15 P. expansum isolates were grown either on PDA without elicitation or with elicitation on PDA amended with 0.13, 0.01, and 0.01 µg mL−1 of TBZ, PYR, and FDL, equivalent of 1/75, 1/50, and 1/50 of their respective discriminatory doses, established as the highest doses allowing growth of wild-type isolates in preliminary experiments. The plates were inoculated with 10 µL of spore suspensions at 105 conidia mL−1 of each isolate and incubated for 7 days at 20 °C.
Total RNA was extracted from mycelia and conidia harvested from the 7-day-old PDA cultures using Quick-RNA Fungal/Bacterial Miniprep TM kit (Zymo Research, Irvine, CA, USA) following the manufacturer’s protocol. The RNA was suspended in nuclease-free water (VWR, Radnor, PA, USA) and residual genomic DNA was eliminated through two rounds of RQ1 RNase-Free DNase treatment (Promega, Madison, WI, USA). The 260/280 ratio of the RNA samples was determined by nanodrop to be between 1.94 and 2.3. First-strand cDNA synthesis was performed using a qScript cDNA synthesis kit (Quantabio, Beverly, MA, USA) and stored in nuclease-free water at −20 °C. Quantitative PCR (qPCR) reactions were performed in a Bio-Rad CFX96 TM Real-Time PCR detection system (Bio-Rad Inc., Hercules, CA, USA). The reaction mixtures consisted of 10 µL of PerfeCTa SYBR Green FastMix (Quantabio, Beverly, MA, USA), 5 µL of cDNA template at 0.6 ng µL−1, primers (Table 2), and nuclease-free water (VWR, Radnor, PA, USA) to a final reaction volume of 20 µL.

Table 2.
Fitness-related genes and primers used in RT-qPCR relative expression analysis.
The recommended thermal cycling protocol for PerfeCTa SYBR Green FastMix was used with an annealing temperature of 60 °C, except for Ac1, LaeA, and BrlA (Supplementary Table S1). Reaction conditions were optimized independently for each gene to achieve efficiencies of 90 to 110%. CFX Maestro TM software version 2.1 was used to analyze qPCR data. Relative gene expression was calculated using the 2−ΔΔCt method, as described previously [44,45]. The expression data presented herein are averages of three biological replicates with three technical replicates per gene and treatment combination. The “sample maximization” experimental set-up for multi-plate qPCR studies [45] was used to minimize technical variation between samples.
2.5. Data Analysis
Data from repeat trials were combined for statistical analysis for all fitness parameters studied except conidial germination at 1.5 °C on intermediate media as well as virulence and sporulation on detached Fuji, which were analyzed by year. Normal distribution of the data was assessed using the Shapiro–Wilk test. All normal datasets were analyzed in R using type I ANOVA followed by Tukey’s post hoc test (p = 0.05). Relative gene expression values were box–cox-transformed and, if normalized, were analyzed via type II ANOVA followed by Tukey’s post hoc test to assess the interaction between the treatment (elicitation vs. non-elicitation) and isolate effects. For non-normal datasets, Kruskal–Wallis followed by Dunn’s test with Benjamini–Hochberg p-value adjustment was used (p = 0.05). A fitness change was calculated for the resistant isolates relative to the average of the three wild-type isolates for each fitness parameter. A fitness change (Fc) was calculated using the formula FC = (XR − XWT)/((XR − XWT)/2) × 100 [46], where FC = fitness change (%), XR = the mean value of the given parameter in the resistant isolate, and XWT = the mean value of the three wild-type isolates for the same parameter. A cumulative fitness change (CFC) was then expressed as the sum of the percentage of gain or loss across all parameters for each isolate.
3. Results
3.1. Comparison of Fitness Capabilities of P. expansum Isolates In Vitro
After 24 h incubation at 20 °C, no significant differences (p > 0.05) in germination rates were observed among isolates on IM, with all isolates germinating at rates ≥99.4% (Table 3) regardless of their phenotype, which were comparable to the germination rates on PDA (Supplementary Table S2). Germination was low in all isolates after 24 h on 2% WA at 20 °C, whereas after 48 h, all resistant isolates, except the dual-resistant TBZRPYRR isolate Pe-08 and PYRRFDLR isolates Pe-153 and Pe-2501, displayed significantly (p < 0.05) lower germination rates (<80%) compared to the wild-type isolates (Table 3). At 1.5 °C, germination on IM and PDA was only observed after 48 h with the dual-resistant TBZRPYRR isolates exhibiting the lowest germination rates (1.2 to 25.9%), in contrast to the dual-resistant PYRRFDLR isolates, which had the greatest germination rates ranging from 30.8 to 51.8% (Table 4). Germination on WA at 1.5 °C was absent up to 96 h. After 10 days of incubation, the resistant isolates, except for the single TBZR isolate Pe-23 and PYRRFDLR isolate Pe-2501, exhibited a significant reduction in germination compared to the wild-type isolates (Table 4).

Table 3.
Germination, mycelial growth, sporulation, and sensitivity to exogenic stresses of 15 Penicillium expansum isolates with different sensitivity phenotypes in vitro at 20 °C.

Table 4.
Germination, mycelial growth, sporulation, and sensitivity to exogenic stresses of 15 Penicillium expansum isolates with different sensitivity phenotypes in vitro at 1.5 °C.
Most isolates grew faster on IM compared to PDA at both 20 and 1.5 °C (Table 3 and Table 4). After 15 days at 20 °C, all isolates grew similarly on IM regardless of their sensitivity phenotype (Table 3) with no differences seen between isolates at 5 and 10 days (Supplementary Table S2). On PDA, the growth of the TBZR isolates Pe-184 and Pe-219 was significantly greater than that of the wild-type isolates (p < 0.05), but no difference in growth was observed among the 10 other resistant isolates (Table 3). There were no significant growth differences between resistant isolates and the three wild-type isolates on IM or PDA after 120 days of incubation at 1.5 °C (Table 4) as well as 30, 60, and 90 days (Supplementary Table S2). Sporulation was not affected by the isolates regardless of their sensitivity phenotype after 15 days at 20 °C or after 120 days at 1.5 °C, regardless of the growth medium (Table 3 and Table 4).
Sensitivities to OSS and ROS were assessed on PDA amended with 12% NaCl or 30 mM of methyl viologen, respectively. After 15 days at 20 °C, the three single TBZR, two dual TBZRPYRR isolates, and one PYRRFDLR isolate grew significantly (p < 0.05) faster than the wild-type isolates on PDA amended with 12% NaCl (Table 3). At 1.5 °C, the triple-resistant isolates showed significantly reduced growth than the wild-type isolates after 120 days on PDA amended with 12% NaCl (Table 4). Moreover, increased sensitivity to ROS was observed in some dual- and triple-resistant isolates compared to the single TBZR and wild-type isolates after 15 days at 20 °C. Notably, two of the TBZRPYRRFDLR-resistant isolates, Pe-2754 and Pe-3045, grew 6 and 16 mm, respectively, compared to >40 mm in the remaining isolates (Table 3). No growth was observed for any of the 15 isolates on PDA amended with 30 mM of methyl viologen after 120 days at 1.5 °C (Table 4).
The stability of the sensitivity phenotypes assessed after 10 monthly transfers on PDA at 1.5 °C showed no significant shifts in EC50 values for TBZ and FDL between G0 and Gm10 in either the spore germination or mycelial growth inhibition assays, except for isolate Pe-2517, which showed an increase in EC50 for FDL based on the spore germination assay (Supplementary Table S2). In contrast, EC50 values for PYR increased by 4- to 25-fold, 1- to 5-fold, and 1- to 9-fold in TBZRPYRR, PYRRFDLR, and TBZRPYRRFDLR isolates, respectively, according to the spore germination assay. However, increases in EC50 values were lower or minor based on the mycelial growth assay (Supplementary Table S3).
3.2. Virulence, Sporulartion, and Resistance Stability in P. expansum Isolates on Detached Apples
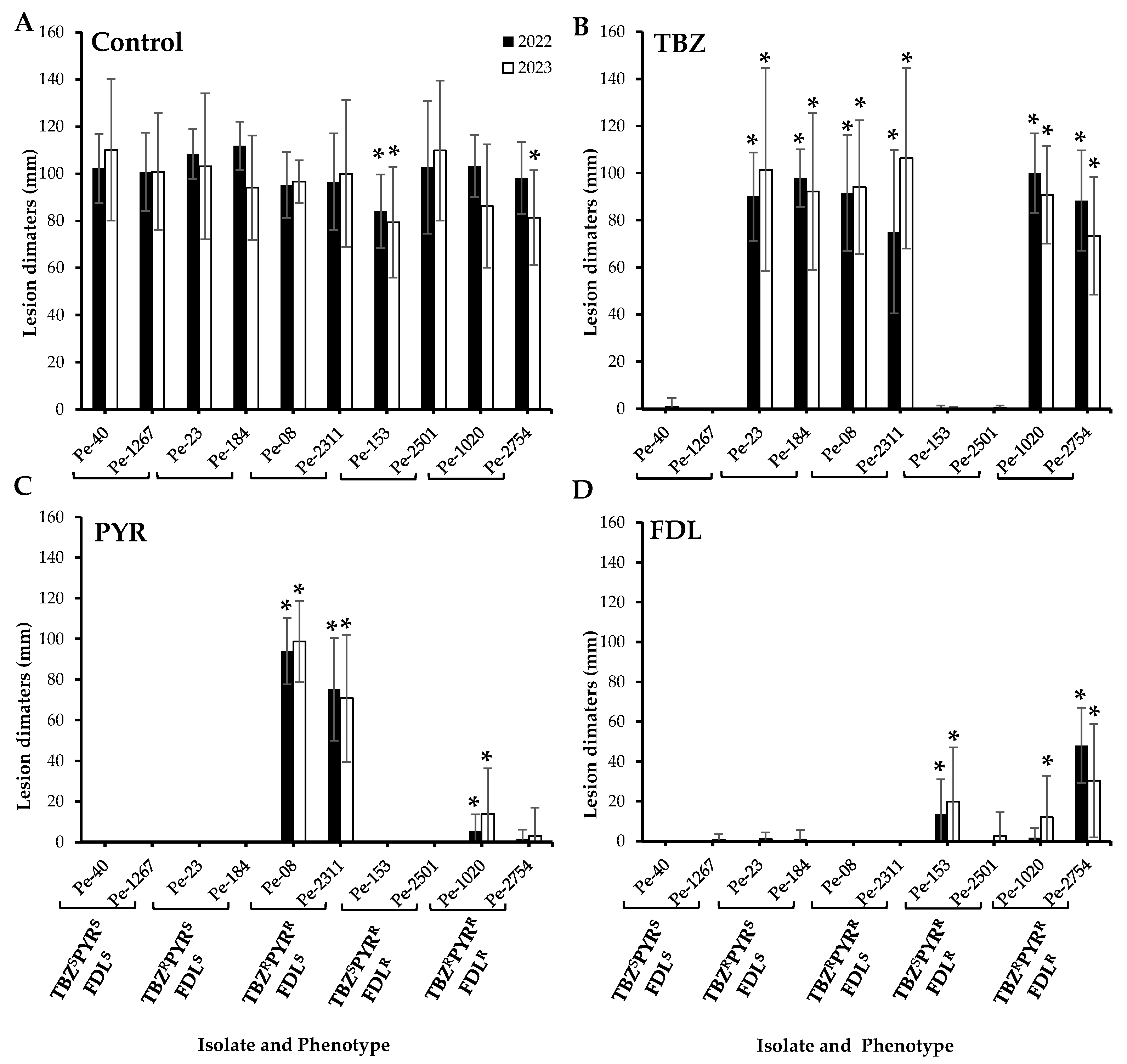
Regardless of their sensitivity phenotype, all ten tested P. expansum isolates caused 100% blue mold incidence on untreated Fuji apples at all inspection times (monthly between two to six months) at 1.5 °C, but some differences were observed in their virulence after six months of storage. For instance, the dual-resistant PYRRFDLR isolate Pe-153 was significantly less virulent compared to the wild-type isolates in both the 2022 and 2023 trials, while the triple-resistant TBZRPYRRFDLR Pe-2754 isolate was less virulent only in the 2023 trial (p < 0.05) (Figure 1A). Under fungicide selection pressure, only isolates with corresponding resistance phenotypes were able to infect the Fuji apples after six months of cold storage (Figure 1A). There were no significant differences in the size of lesions caused by the isolates that were resistant to TBZ on TBZ-treated apples (Figure 1B). On PYR-treated apples, the dual-resistant TBZRPYRR isolates exhibited a significantly greater lesion diameter compared to the other isolates. The triple-resistant isolates Pe-1020 and Pe-2754 caused small lesions on PYR-treated apples (Figure 1C), but no lesions were observed on apples infected by the dual-resistant PYRRFDLR isolates on PYR-treated apples in both years (Figure 1C). Overall, the FDLR isolates caused smaller lesions on FDL-treated apples. Although, the dual-resistant PYRRFDLR isolate Pe-2501 remained avirulent in 2022, both the PYRRFDLR and the triple-resistant isolates produced lesions ranging from 1.7 to 48.0 mm in 2022 and 2.5 to 30.4 mm in 2023 on FDL-treated apples (Figure 1D).
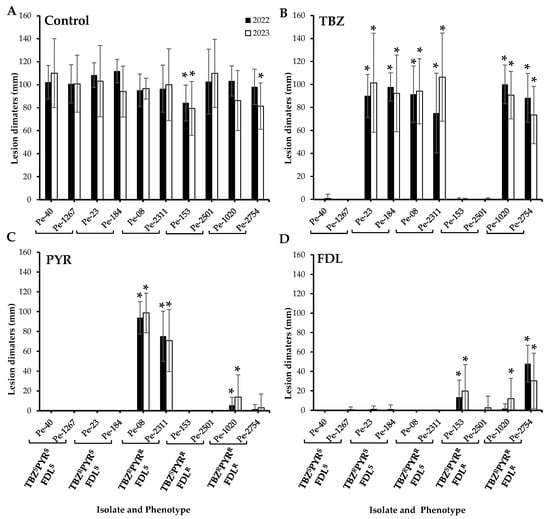
Figure 1.
Virulence (lesion diameters in mm) of ten Penicillium expansum isolates on wounded untreated Fuji apples (control, (A)) and apples treated with thiabendazole (TBZ, (B)), pyrimethanil (PYR, (C)), and fludioxonil (FDL, (D)) at the label rates then inoculated with spore suspensions from each isolate. Lesion diameters were measured after six months at 1.5 °C. The 2022 and 2023 trials were analyzed separately. Error bars indicate the standard deviations of the means. Asterisks indicate statistical difference from the TBZSPYRSFDLS (wild-type) isolates based on Kruskal–Wallis and Dunn’s tests at p < 0.05. S and R denote sensitive and resistant isolates, respectively.
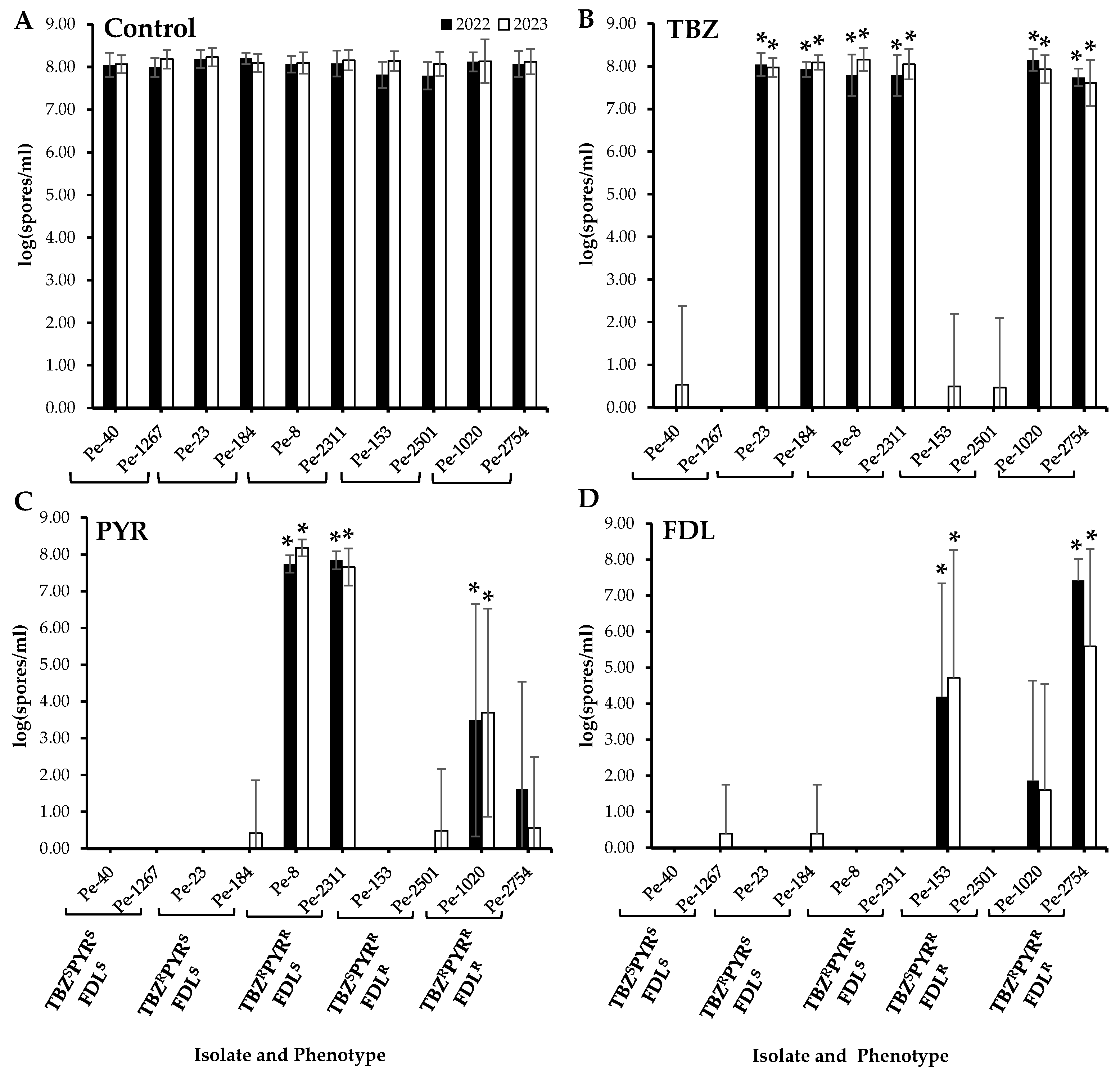
No significant differences in sporulation were observed between the wild-type and the resistant isolates on untreated apples after six months of cold storage (Figure 2A). On TBZ-treated apples, the sporulation of all TBZR isolates, i.e., those with single, dual, and triple resistance, was significantly higher than that of the TBZS isolates (Figure 2B). On PYR-treated fruit, significantly increased sporulation (p < 0.05) was observed in the two TBZRPYRR isolates as well as the triple-resistant isolate Pe-1020 (Figure 2C), while only the PYRRFDLR isolate Pe-153 and the triple-resistant isolate Pe-2754 exhibited significantly increased spore production (p < 0.05) on FDL-treated Fuji apples (Figure 2D).
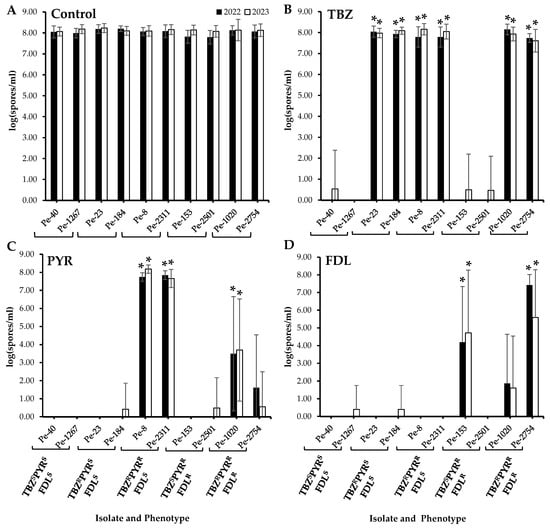
Figure 2.
Sporulation (spores mL−1) of Penicillium expansum isolates with different sensitivity phenotypes on wounded untreated Fuji apples (control, (A)) and on apples treated with thiabendazole (TBZ, (B)), pyrimethanil (PYR, (C)), and fludioxonil (FDL, (D)) at the label rates then inoculated with spore suspensions from each isolate. Lesion diameters were measured after six months at 1.5 °C. Error bars indicate the standard deviations of the means. Asterisks indicate statistical difference from the TBZSPYRSFDLS (wild-type) isolates based on Kruskal–Wallis and Dunn’s tests at p < 0.05. S and R denote sensitive and resistant isolates, respectively.
Consistent with the in vitro findings, no significant changes in phenotypic sensitivity for TBZ and FDL were observed after eight months (four transfers at two-month intervals) on detached Fuji apples at 1.5 °C, based on the spore germination assay and discriminatory doses used for isolate characterization. For PYR, isolate Pe-153 exhibited reduced sensitivity after four transfers on apples.
3.3. Cumulative Phenotypic Fitness Changes in P. expansum-Resistant Isolates
A cumulative fitness change (CFC) expressed as the sum of the percentage of gain or loss across all parameters in the resistant isolates relative to the wild-type isolates was calculated for each isolate and phenotype. All resistant isolates, regardless of their sensitivity phenotypes, showed a cumulative fitness loss on WA with mean cumulative fitness changes (CFC) ranging from −72 to −86% and −15 to −26% at 1.5 and 20 °C, respectively, with no major difference among the phenotypes (Table 5). On the IM at 1.5 °C, the dual-resistant PYRRFDLR isolates exhibited a mean fitness gain of 36%, to the contrary of the TBZR, TBZRPYRR, and TBZRPYRRFDLR phenotypes with respective mean fitness losses of −70, −103, and −8%. At 20 °C, only minor gains or losses were observed on IM. On the nutrient-rich medium PDA, all fungicide sensitivity phenotypes registered fitness losses at 1.5 °C, whereas fitness loss was only observed in PYRRFDLR and the TBZRPYRRFDLR phenotypes at 20 °C. On detached untreated apples, the single TBZR isolates exhibited a fitness gain of 7%, while the three other phenotypes exhibited moderate fitness losses between −5 and −14% (Table 5). On fungicide-treated apples, large fitness gains were observed across most phenotypes, except for the TBZRPYRR phenotypes on FDL-treated apples (Table 5).

Table 5.
Cumulative fitness change (CFC in %) in resistant isolates of Penicillium expansum with different sensitivity phenotypes relative to sensitive wild-type isolates in vitro and on apples.
3.4. Gene Expression of Fitness-Related Genes in P. expansum Isolates with Different Fungicide Sensitivity Phenotypes
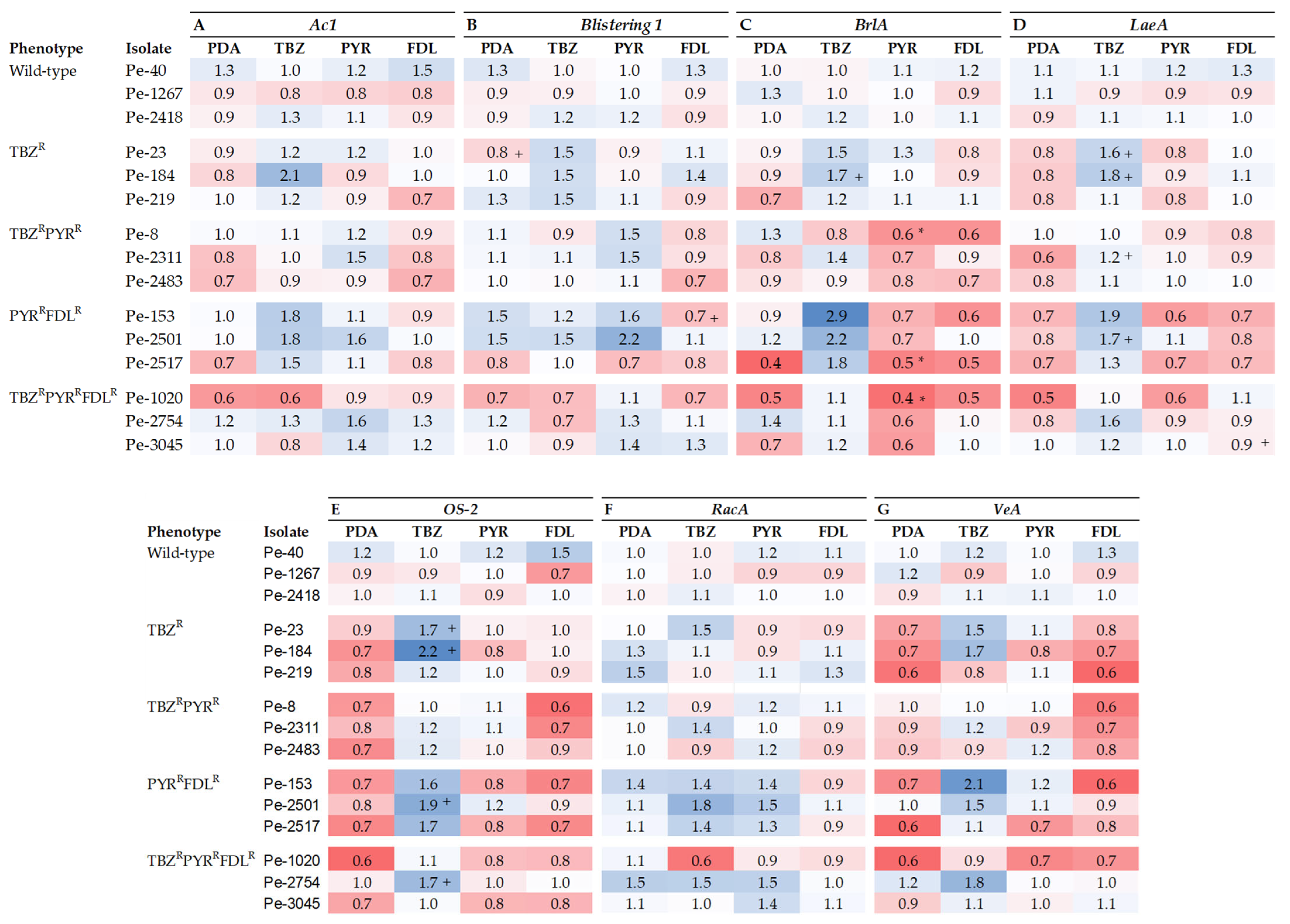
The relative expression (RE) of seven fitness-related genes was measured in vitro on PDA with and without elicitation via sub-lethal doses of TBZ, PYR, or FDL. The Ac1 gene, which codes for adenylyl cyclase, is linked to germination, growth, and virulence in fungi. In a cross-isolate comparison, Ac1 was significantly up-regulated (2-fold) in the TBZR isolate Pe-184 by TBZ elicitation compared to the control (Figure 3A). Ac1 expression was significantly (p < 0.05) down- and up-regulated in the TBZRPYRRFDLR and TBZR phenotypes, respectively, whereas its expression was up-regulated by PYR in the TBZRPYR phenotype (Supplementary Figure S1A). Likewise, the expression of the Blistering1 gene, linked to growth, virulence, and patulin production, was largely unaffected in most isolates except for the TBZR isolate Pe-23 and the PYRRFDLR isolate Pe-153, which were, respectively, up- and down-regulated (p < 0.05) by TBZ and FDL compared to the control (Figure 3B). A sensitivity phenotype effect (p < 0.05) was observed in the TBZR phenotype where expression was up-regulated by TBZ (Supplementary Figure S1B).
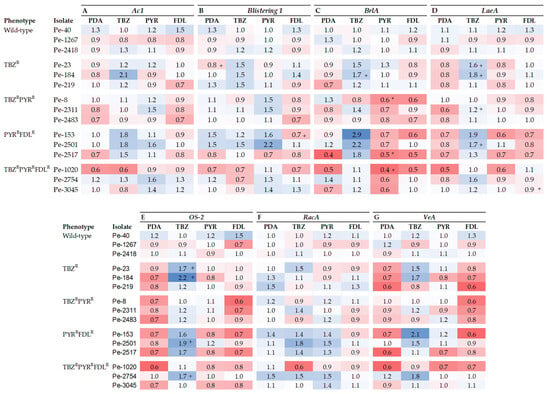
Figure 3.
Heatmap representing the expression of Ac1 (A), Blistering1 (B), OS-2 (C), RacA (D), BrlA (E), LaeA (F), and VeA (G) genes relative to the reference genes 28s and CMD in Penicillium expansum isolates with different sensitivity phenotypes after 7 days at 20 °C on PDA (control) and on PDA amended with thiabendazole (TBZ), pyrimethanil (PYR), and fludioxonil (FDL) at 0.13, 0.01, and 0.01 µg mL−1, respectively. Data bars are the means of nine values from three biological replicates. Error bars indicate the standard deviations of the means. S and R denote sensitive and resistant isolates, respectively. Pluses indicate statistical difference between isolates as a factor of fungicide elicitation compared to the control. Asterisks indicate statistical differences in resistant isolates from the wild-type isolates; + indicates statistical differences in elicitated samples compared to the PDA treatment alone. (A–G) and (E,F) were analyzed with type II ANOVA/Tukey-HSD and type I ANOVA/Tukey-HSD or Kruskal–Wallis and Dunn’s tests, respectively, at p ≤ 0.05.
The expression of the OS-2 gene, implicated in osmotic stress tolerance, was significantly up-regulated by TBZ in four isolates, Pe-23 and Pe-184 (TBZR), Pe-2483 (TBZRPYRR), and Pe-2501 (PYRRFDLR), across three phenotypes (Figure 3C). The expression of OS-2 was down-regulated by FDL in the TBZRPYRR phenotype and was up-regulated by PYR in the TBZRPYRR phenotype and by TBZ in all resistant phenotypes (Figure S1C). Moreover, the RE of the RacA gene, linked to oxidative stress response and patulin production, was significantly down-regulated (p < 0.05) by TBZ in the TBZR isolate Pe-219 and the TBZRPYRRFDLR isolate Pe-1020 (Figure 3D), whereas no differences were observed between phenotypes (Figure S1D).
The RE of BrlA, a transcription factor important for sporulation in P. expansum, was significantly down-regulated by PYR in isolates Pe-8 (TBZRPYRR), Pe-2517 (PYRRFDLR), and Pe-1020 (TBZRPYRRFDLR) compared to the sensitive isolates (Figure 3E). There was also a significant phenotype effect as BrlA RE was down-regulated (p < 0.05) in the TBZRPYRR, PYRRFDLR, and TBZRPYRRFDLR phenotypes compared to the sensitive phenotype (Figure S1E). BrlA expression was significantly up-regulated (p < 0.05) in the TBZR and PYRRFDLR phenotypes by TBZ elicitation compared to the control (Figure S1E). LaeA is another transcription factor linked to virulence, sporulation, and patulin production. A cross-isolate comparison showed LaeA to be up-regulated by TBZ in isolates Pe-23 and Pe-184 (TBZR), Pe-2311 (TBZRPYRR), and Pe-2501 (PYRRFDLR), whereas the gene was over-expressed (p < 0.05) by FDL in the triple-resistant isolate Pe-1020 compared to the non-elicited control (Figure 3F). A comparison between phenotypes showed that LaeA was down-regulated in all non-elicited resistant phenotypes. PYR down-regulated LaeA expression (p < 0.05) in the PYRRFDLR and TBZRPYRRFDLR phenotypes, while expression was down-regulated by FDL in the PYRRFDLR phenotype compared to the sensitive phenotype (Supplementary Figure S1F).
The last gene investigated was VeA, reported to influence growth, virulence, sporulation, and patulin production. Its expression was very similar among the 15 P. expansum isolates except in the TBZR isolates Pe-23 and Pe-184, which showed significantly increased expression (p < 0.05) compared to the wild-type isolates (Figure 3G). VeA expression among phenotypes was down-regulated in TBZR, PYRRFDLR, and TBZRPYRRFDLR in non-elicited treatment as well as by FDL elicitation in the TBZR, TBZRPYRR, and PYRRFDLR phenotypes compared to the sensitive phenotype (Figure S1G). In contrast, TBZ elicitation up-regulated VeA RE significantly in the TBZR and PYRRFDLR phenotypes.
4. Discussion
P. expansum isolates with varying fungicide sensitivity phenotypes, i.e., isolates sensitive to TBZ, PYR, and FDL or resistant to one, two, or all three fungicides, were collected from commercial packinghouses in the U.S. PNW states of WA and OR. Their germination, growth, sporulation, response to osmotic and oxidative stress, and phenotype stability were assessed in vitro, while virulence, sporulation, and stability were evaluated on detached apples under different temperatures and fungicide selection pressures. Resistant isolates exhibited significantly reduced conidial germination on a nutrient-deficient medium (WA) at both 20 and 1.5 °C compared to the wild-type isolates. However, this reduction was pronounced on nutrient-rich media like PDA and IM, which simulates plant surface nutrient availability [38]. While a correlation between fungicide resistance and reduced germination has been documented in Aspergillus parasiticus [47], only one previous study using P. expansum lab mutants found no effect of resistance to PYR and FDL on germination on PDA after 24 h [31]. Our study is the first to provide more insights into such a correlation in wild-type and field-resistant isolates. Given that germination is crucial for fungal infection, impaired germination may hinder the successful establishment of resistant isolates. These findings suggest that P. expansum isolates resistant to TBZ, PYR, and/or FDL may pose lower disease risk compared to wild-type isolates in cold and nutrient-deficient environments such as bins, storage rooms, and flume water, provided that organic matter sources (e.g., remaining decayed fruit) are absent. This highlights the importance of thoroughly cleaning and sanitizing postharvest facilities and equipment to minimize the risk of decay by fungicide-resistant isolates.
In contrast with spore germination, the growth of P. expansum isolates on three agar media was not affected by their fungicide sensitivity phenotype, either after 120 days at 1.5 °C or after 15 days at 20 °C. Previous studies have reported mixed findings regarding growth differences among isolates with varying resistance profiles. For instance, lab mutants and field P. expansum isolates resistant to TBZ exhibited no growth differences on PDA after 7 days at 20 °C but displayed higher severity than the sensitive isolates on detached pears and apples [12,13]. Conversely, P. expansum lab mutants resistant to TBZ, PYR, or/and FDL exhibited variable growth on PDA after one week and 10 weeks at 20 and 0 °C, respectively [31]. Specifically, lab mutants resistant to FDL, PYR, and dual-resistant to TBZ and FDL showed reduced growth compared to the parental isolates, whereas the triple-resistant isolates grew at rates comparable to the wild-type. Similarly, Greek field isolates with single resistance to cyprodinil (FRAC 9) exhibited similar growth on PDA and virulence on Red Delicious apples compared to the wild-type isolates, whereas isolates with triple resistance to FDL, iprodione (FRAC 2), and tebuconazole (FRAC3) exhibited reduced growth on PDA and reduced virulence after 7 days at 20 °C [22]. Discrepancies between studies may stem from the use of lab mutants, which often display reduced fitness, as well as differences in resistance phenotypes investigated, experimental methods, or environmental conditions. Nevertheless, when considering both the current findings and previous research on various fitness parameters, P. expansum isolates resistant to one or more fungicide appear capable of maintaining adequate growth across a range of nutrient environments, suggesting that resistance does not necessarily compromise growth capacity under the tested conditions.
The ability of plant pathogens to withstand external stressors, such as osmotic (OSS) and oxidative (ROS) stress, plays a critical role in their survival under adverse conditions and their potential to cause disease. Fludioxonil is believed to disrupt the high-osmolarity glycerol (HOG) pathway, which regulates glycerol synthesis, an essential osmoregulatory mechanism [18,26]. As a result, the sensitivity to OSS is commonly evaluated in FDL-resistant fungal pathogens. In this study, the fungicide resistance phenotype of P. expansum field isolates did not significantly affect their OSS response after 15 days at 20 °C. However, dual PYRRFDLR- and triple TBZRPYRRFDLR-resistant isolates showed significantly reduced growth on PDA amended with 12% NaCl at 1.5 °C after 120 days. These findings align with previous reports of increased OSS sensitivity in FDLR isolates of P. expansum and B. cinerea [31,48] but not in P. digitatum [18]. The difference observed in OSS sensitivity between P. expansum and P. digitatum could be explained by the relatively higher dose of NaCL2 used in our study compared to that used by Kanetis et al. [18]. Furthermore, ROS plays a crucial role in fungal infection, signaling, and pathogenicity [49]. While ROS sensitivity did not differ significantly among wild-type and resistant isolates, the dual PYRRFDLR isolates Pe-2501 and Pe-2517, as well as the triple-resistant isolates Pe-2754 and Pe-3045, exhibited the lowest growth among the isolates after 15 days on PDA amended with 30 mM of methyl viologen. Although ROS sensitivity in P. expansum has not been extensively studied, its link to the HOG pathway has been demonstrated in FDL-resistant strains of B. cinerea and Saccharomyces cerevisiae [50,51]. In contrast, the deletion of Hog1, a key kinase in the HOG pathway, did not affect ROS sensitivity in P. digitatum. The heightened OSS and ROS sensitivity in FDLR isolates may explain their low prevalence in PNW packinghouses after nearly 20 years of FDL use. Notably, no substantial shift in EC50 to FDL, TBZ, or PYR among the resistant isolates occurred after ten weekly transfers, suggesting that unstable resistance mechanisms are unlikely to explain their limited occurrence. Nevertheless, continued monitoring and further investigation into stress response mechanisms are essential to mitigate the risk of sudden increase in resistance.
Understanding the pathogenic fitness of fungicide-resistant populations is important for developing management strategies. In 80% of P. expansum isolates inoculated on fungicide-untreated Fuji apples, no significant differences in lesion size were observed after 140 days at 1.5 °C, even if some dual PYRRFDLR- and triple TBZRPYRRFDLR-resistant isolates produced smaller lesions compared to the wild-type isolates after six months of cold storage. Previous studies reported that TBZR P. expansum isolates from Italy and Greece were more virulent than the wild-type isolates [12,13]. In this study, TBZR isolates behaved differently across years with larger lesions than the wild-type isolates in 2022, but not in 2023. Previously, laboratory PYRR mutants showed virulence comparable to their parental isolates, but FDLR and TBZRFDLR mutants exhibited temperature-dependent virulence, which was reduced at 0 °C but not at 20 °C [31]. Field isolates resistant to FDL and other preharvest fungicides, not labeled for blue mold management, have also demonstrated reduced virulence in short-term storage at 20 °C [23,24]. Despite some variation in virulence, all resistant phenotypes, i.e., single, dual, and triple, could infect apples, produce substantial lesions, and sporule abundantly under selection pressure. Moreover, these isolates maintained stable resistance over 10 months in vitro and eight months on fruits. These findings align with previous reports [27,49] and reinforce concerns that continued fungicide use may accelerate the selection for additional resistant P. expansum populations in commercial packinghouses.
Comparing individual fitness parameters within and across studies to assess the overall impact of fungicide resistance on fitness is inherently challenging due to methodological and biological variability. In natural environments, however, fitness costs likely result from the combined effect of multiple components, such as germination, growth, sporulation, stress tolerance, and virulence, rather than from a single trait. To better capture these cumulative effects, we calculated a cumulative fitness cost change (CFc), representing the overall gain or loss in fitness of resistant relative to the wild-type isolates. This analysis revealed clear overall fitness reductions in the resistant phenotypes under most in vitro conditions, especially under cold storage, compared to the wild-type isolates. These reductions were not always apparent when examining individual fitness components separately. Furthermore, the CFc analysis highlighted a negative impact of prolonged selection pressure on P. expansum virulence on apples. The CFc approach corresponds with the observed low prevalence of dual- and triple-resistant phenotypes, especially those resistant to FDL, in PNW packinghouses, suggesting that cumulative fitness costs, rather than isolated trait penalties, may limit their persistence. While further mathematical and statistical refinement is needed, CFc offers a promising framework for estimating fitness costs in plant pathogens under natural conditions.
To investigate whether resistance to a single or multiple fungicides affects the expression and functionality of genes associated with key fitness traits, the expression of seven fitness-related genes was evaluated in the absence and presence of fungicide selection pressure. Exposure to TBZ significantly elevated the expression of LaeA and OS2, which play roles in virulence, sporulation, and sensitivity to osmotic stress (OSS), across all resistant phenotypes, as well as increased expression of Ac1, Blistering1, BrlA, and VeA in select phenotypes. While differences in OSS sensitivity were observed between isolates and phenotypes in vitro, the differential expression of certain genes may help explain the absence of fitness penalties, such as in growth, virulence, or sporulation, both in vitro and on detached fruit. Previous research reported differential gene expressions due to fungicide application, particularly in resistant isolates [52]. Notably, this gene expression induction was observed only with TBZ. In contrast, PYR treatment elicited only minor changes in expression at the phenotype level and no discernible impact at the isolate level. FDL treatment had no observable effect on gene expression at the phenotype level and only a minor impact at the isolate level. This lack of response is particularly surprising for OS-2, given the well-established role of the HOG pathway in mediating fludioxonil responses across several fungal species [9,18,26,29]. Fludioxonil is thought to stimulate polyol (e.g., glycerol) production via the HOG pathway and OS-2 in response to osmotic stress; however, this mechanism may differ in P. expansum, as prior studies have not linked OS-2 expression to osmotic stress in this species [38] nor were mutations detected in OS genes of P. expansum and other Penicillium spp. [9].
Comparisons between PYR-resistant and -sensitive phenotypes revealed consistently lower expression of BrlA in resistant isolates treated with PYR or FDL. BrlA, a C2H2-type zinc-finger transcription factor, is essential for conidiophore development and sporulation [35]. Additional differences in LaeA and VeA expression were observed between resistant and sensitive phenotypes; however, no consistent differences emerged at the individual isolate level. These results did not align with in vitro and in vivo sporulation studies, which showed no significant differences between isolates or phenotypes. This suggests that assessing fitness penalties at the molecular level, by measuring the expression of fitness-related genes, may be more effective in detecting potential fitness costs associated with fungicide resistance development. Although the precise resistance mechanisms to PYR and FDL in P. expansum remain unclear, if resistance involves increased metabolic demand, fungicide exposure may redirect energy toward detoxification or efflux pump activity at the expense of reproductive functions, leading to reduced expression of genes such as BrlA [53]. The relative expression analysis in this study highlights the complex nature of fungal fitness. Numerous transcription factors and regulatory pathways influence the life cycle and virulence of P. expansum, underscoring the need for broader approaches, such as RNA sequencing, to investigate both known and novel fitness-related genes.
5. Conclusions
Our study established variable associations between fungicide resistance and fitness in P. expansum. We demonstrated that under low-nutrient conditions, the germination capacity of fungicide-resistant isolates is significantly reduced, and that dual- and triple-resistant isolates may be more vulnerable to osmotic and oxidative stresses. Furthermore, we developed a cumulative fitness cost (CFc) metric that revealed reduced overall fitness in resistant phenotypes. However, no association was found between fungicide resistance and virulence, suggesting that resistant phenotypes may persist and cause a greater risk in commercial packinghouses.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microorganisms13081846/s1. Figure S1: Expression of Ac1 (A), Blistering1 (B), OS2 (C), RacA (D), BrlA (E), LaeA (F), and VeA (G) genes relative to reference genes 28s and CMD in Penicillium expansum of different sensitivity phenotypes after 7 days at 20 °C on PDA (control) and on PDA amended with thiabendazole (TBZ), pyrimethanil (PYR), and fludioxonil (FDL) at 0.13, 0.01, and 0.01 µg/mL, respectively. Table S1: Thermocycler conditions and primer efficiencies for fitness gene expression analysis via qPCR. Table S2: Germination and mycelial growth of 15 Penicillium isolates on different media and at various incubation periods. Table S3: Effective concentrations inhibiting 50% germination or growth (EC50) of thiabendazole, pyrimethanil, and fludioxonil in 15 Penicillium expansum isolates.
Author Contributions
Conceptualization, A.A.; Data Curation, J.T.P.; Formal Analysis, J.T.P.; Funding Acquisition, A.A.; Investigation, J.T.P.; Project Administration, A.A.; Supervision, A.A.; Writing—Original Draft, J.T.P.; Writing—Review and Editing, J.T.P. and A.A. All authors have read and agreed to the published version of the manuscript.
Funding
This work was funded by the Washington Tree Fruit Research Commission Grant #AP-20-101A and by the Washington State Department of Agriculture (WSDA) Specialty Crop Block Grant #K3935.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in this study are included in the article and Supplementary Materials. Further inquiries can be directed to the corresponding author.
Acknowledgments
The authors are grateful for Valentin Leannec-Rialland for his technical assistance with the gene expression analyses.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| TBZ | Thiabendazole |
| PYR | Pyrimethanil |
| FDL | Fludioxonil |
| TBZSPYRSFDLS | Wild-type isolates sensitive to all three fungicides |
| TBZR | Single-resistant isolate to thiabendazole |
| TBZRPYR | Dual-resistant isolate to thiabendazole and pyrimethanil |
| PYRRFDLR | Dual-resistant isolate to pyrimethanil and fludioxonil |
| TBZRPYRRFDLR | Triple-resistant isolates to all three fungicides |
| Fc | Fitness change relative to wild-type isolates |
| CFc | Cumulative fitness change |
| FRAC | Fungicide Resistance Action Committee |
| Gw or Gm | Generation at a given week or month following transfer of isolates |
| OSS | Osmotic stress |
| ROS | Reactive oxygen species |
References
- Amiri, A.; Bompeix, G. Diversity and population dynamics of Penicillium spp. on apples in pre- and postharvest environments: Consequences for decay development. Plant Pathol. 2005, 54, 74–81. [Google Scholar] [CrossRef]
- Rosenberger, D.A. Blue mold. In Compendium of Apple and Pear Diseases; Jones, A.L., Aldwinkle, H.S., Eds.; American Phytopathological Society: St. Paul, MN, USA, 1990; pp. 54–55. [Google Scholar]
- Errampalli, D.; Crnko, N. Control of blue mold caused by Penicillium expansum on apples “Empire” with fludioxonil and cyprodinil. Can. J. Plant Pathol. 2004, 26, 70–75. [Google Scholar] [CrossRef]
- Wang, K.; Ngea, G.L.N.; Godana, E.A.; Shi, Y.; Lanhuang, B.; Zhang, X.; Zhao, L.; Yang, Q.; Wang, S.; Zhang, H. Recent advances in Penicillium expansum infection mechanisms and current methods in controlling P. expansum in postharvest apples. Crit. Rev. Food Sci. Nutr. 2023, 63, 2598–2611. [Google Scholar] [CrossRef]
- Amiri, A.; Ali, E.M. Prevalence of Storage Decays of Apple: Lessons from the 2016 Statewide Survey. Available online: https://treefruit.wsu.edu/article/prevalence-of-storage-decays-of-apple-lessons-from-the-2016-statewide-survey/ (accessed on 11 April 2025).
- Sanderson, P.G.; Spotts, R.A. Postharvest decay of winter pear and apple fruit caused by species of Penicillium. Phytopathology 1995, 85, 103–110. [Google Scholar] [CrossRef]
- Dutoit, M.; Nelson, L.M.; Tyson, R. Predicting the spread of postharvest disease in stored fruit, with application to apples. Postharvest Biol. Technol. 2013, 85, 45–56. [Google Scholar] [CrossRef]
- Fungicide Resistance Action Committee Pathogen Risk List 2019. Available online: https://www.frac.info/publications/all-downloads/#open-tour (accessed on 11 April 2025).
- Pandey, M.; Amiri, A. High resistance levels to pyrimethanil and fludioxonil among fourteen Penicillium spp. from pome fruits in the U.S. Pacific Northwest. Pestic. Biochem. Physiol. 2024, 206, 106206. [Google Scholar] [CrossRef]
- Pandey, M.; Haskell, C.L.; Cowell, J.D.; Amiri, A. Sensitivity to the Demethylation Inhibitor Difenoconazole Among Baseline Populations of Various Penicillium spp. Causing Blue Mold of Apples and Pears. J. Fungi 2025, 11, 61. [Google Scholar] [CrossRef]
- Xiao, C.L.; Kim, K.K.; Boal, R.J. First report of occurrence of pyrimethanil resistance in Penicillium expansum from stored apples in Washington State. Plant Dis. 2011, 95, 72. [Google Scholar] [CrossRef]
- Baraldi, E.; Mari, M.; Chierici, E.; Pondrelli, M.; Bertolini, P.; Pratella, G.C. Studies on thiabendazole resistance of Penicillium expansum of pears: Pathogenic fitness and genetic characterization. Plant Pathol. 2003, 52, 362–370. [Google Scholar] [CrossRef]
- Malandrakis, A.A.; Markoglou, A.N.; Konstantinou, S.; Doukas, E.C.; Kalmpokis, J.F.; Karaoglanidis, G.S. Molecular characterization, fitness and mycotoxin production of benzimidazole-resistant isolates of Penicillium expansum. Int. J. Food Microbiol. 2013, 162, 237–244. [Google Scholar] [CrossRef]
- Yan, H.J.; Gaskins, V.L.; Vico, I.; Luo, Y.G.; Jurick, W.M., II. First report of Penicillium expansum isolates resistant to pyrimethanil from stored apple fruit in Pennsylvania. Plant Dis. 2014, 98, 1004. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, P.F.; Saulie-Carter, J.L. The occurrence of benomyl-tolerant strains of Penicillium expansum and Botrytis cinerea in the mid-Columbia region of Oregon and Washington. Plant Dis. Rep. 1978, 62, 302–305. [Google Scholar]
- Cabañas, R.; Castellá, G.; Lourdes Abaarca, M.; Rosa Bragulat, M.; Javier Cabañes, F. Thiabendazole resistance and mutations in the ß-tubulin gene of Penicillium expansum strains isolated from apples and pears with blue mold decay. FEMS Microbiol. Lett. 2009, 297, 189–195. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yin, Y.N.; Xiao, C.L. Molecular characterization and a multiplex allele-specific PCR method for detection of thiabendazole resistance in Penicillium expansum from apple. Eur. J. Plant Pathol. 2013, 136, 703–713. [Google Scholar] [CrossRef]
- Kanetis, L.; Forster, H.; Jones, C.A.; Borkovich, K.A.; Adaskaveg, J.E. Characterization of genetic and biochemical mechanisms of fludioxonil and pyrimethanil resistance in field isolates of Penicillium digitatum. Phytopathology 2008, 98, 205–214. [Google Scholar] [CrossRef]
- Zhang, Y.; Fu, Y.; Luo, C.; Zhu, F. Pyrimethanil sensitivity and resistance mechanisms in Penicillium digitatum. Plant Dis. 2021, 105, 1758–1764. [Google Scholar] [CrossRef]
- Fritz, R.; Lanen, C.; Colas, V.; Leroux, P. Inhibition of methionine biosynthesis in Botrytis cinerea by the anilinopyrimidine fungicide pyrimethanil. Pestic. Sci. 1997, 49, 40–46. [Google Scholar] [CrossRef]
- Mosbach, A.; Edel, D.; Farmer, A.D.; Widdison, S.; Barchietto, T.; Dietrich, R.A.; Corran, A.; Scalliet, G. Anilinopyrimidine resistance in Botrytis cinerea is linked to mitochondrial function. Front. Microbiol. 2017, 8, 2361. [Google Scholar] [CrossRef]
- Amiri, A.; Ali, M.E.; De Angelis, D.R.; Mulvaney, K.A.; Pandit, L.K. Prevalence and distribution of Penicillium expansum and Botrytis cinerea in apple packinghouses across Washington State and their sensitivity to the postharvest fungicide pyrimethanil. Acta Hortic. 2021, 1323, 167–172. [Google Scholar] [CrossRef]
- Karaoglanidis, G.S.; Markoglou, A.N.; Bardas, G.A.; Doukas, E.G.; Konstantinou, S.; Kalampokis, J.F. Sensitivity of Penicillium expansum field isolates to tebuconazole, iprodione, fludioxonil and cyprodinil and characterization of fitness parameters and patulin production. Int. J. Food Microbiol. 2011, 145, 195–204. [Google Scholar] [CrossRef]
- Samaras, A.; Ntasiou, P.; Myresiotis, C.; Karaoglandis, G. Multidrug resistance of Penicillium expansum to fungicides: Whole transcriptome analysis of MDR strains reveals overexpression of efflux transporter genes. Int. J. Food Microbiol. 2020, 335, 108896. [Google Scholar] [CrossRef]
- Gandia, M.; Garrigues, S.; Hernanz-Koers, M.; Manzanares, P. Differential roles, crosstalk and response to the antifungal protein AfpB in the three mitogen-activated protein kinases (MAPK) pathways of the citrus postharvest pathogen Penicillium digitatum. Fungal Genet. Biol. 2019, 124, 17–28. [Google Scholar] [CrossRef]
- Zhang, Y.; Lamm, R.; Pillonel, C.; Lam, S.; Xu, J.-R. Osmoregulation and fungicide resistance: The Neurospore crassa os-2 gene encodes a HOG1 mitogen-activated protein kinase homologue. Appl. Environ. Microbiol. 2002, 68, 532–538. [Google Scholar] [CrossRef] [PubMed]
- Gaskins, V.L.; Yu, I.V.; Jurick, W.M., II. First Report of Penicillium expansum isolates with reduced sensitivity to fludioxonil from a commercial packinghouse in Pennsylvania. Plant Dis. 2015, 99, 1182. [Google Scholar] [CrossRef]
- Oiki, S.; Yaguchi, T.; Urayama, S.I.; Hagiwara, D. Wide distribution of resistance to the fungicides fludioxonil and iprodione in Penicillium species. PLoS ONE 2022, 17, e0262521. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Chen, C.; Zhu, C.; Sun, X.; Ruan, R. Os2 MAP kinase-mediated osmostress tolerance in Penicillium digitatum is associated with its positive regulation on glycerol synthesis and negative regulation on ergosterol synthesis. Microbiol. Res. 2014, 169, 511–521. [Google Scholar] [CrossRef]
- Gilchrist, M.A.; Sulsky, D.L.; Pringle, A. Identifying fitness and optimal life-history strategies for an asexual filamentous fungus. Evolution 2006, 60, 970–979. [Google Scholar] [CrossRef]
- Li, H.X.; Xiao, C.L. Characterization of fludioxonil-resistant and pyrimethanil-resistant phenotypes of Penicillium expansum from apple. Phytopathology 2008, 98, 427–435. [Google Scholar] [CrossRef]
- El Hajj Assaf, C.; Snini, S.P.; Tadrist, S.; Bailly, S.; Naylies, C.; Oswald, I.P.; Lorber, S.; Puel, O. Impact of veA on the development, aggressiveness, dissemination and secondary metabolism of Penicillium expansum. Mol. Plant Pathol. 2018, 19, 1971–1983. [Google Scholar] [CrossRef]
- Kumar, D.; Barad, S.; Chen, Y.; Luo, X.; Tannous, J.; Dubey, A.; Matana, N.; Tian, S.; Li, B.; Keller, N.; et al. LaeA regulation of secondary metabolism modulates virulence in Penicillium expansum and is mediated by sucrose. Mol. Plant Pathol. 2017, 18, 1150–1163. [Google Scholar] [CrossRef]
- Jurick, W.M., II; Peng, H.; Beard, H.S.; Garrett, W.M.; Lichtner, F.J.; Luciano-Rosario, D.; Macarisin, O.; Liu, Y.; Peter, K.A.; Gaskins, V.L.; et al. Blistering1 modulates Penicillium expansum virulence via vesicle-mediated protein secretion. Mol. Cell. Proteom. 2020, 19, 344–361. [Google Scholar] [CrossRef]
- Zetina-Serrano, C.; Rocher, O.; Naylies, C.; Lippi, Y.; Oswald, I.P.; Lorber, S.; Purl, O. The brlA gene deletion reveals that patulin biosynthesis is not related to conidiation in Penicillium expansum. Int. J. Mol. Sci. 2020, 21, 6660. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, M.; Wang, J.; Zhu, C.; Chung, K.-R.; Li, H. Adenylyl cyclase is required for cAMP production, growth, conidial germination, and virulence in the citrus green mold pathogen Penicillium digitatum. Microbiol. Res. 2016, 192, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, A.; Medina, Á.; Córdoba, J.J.; Magan, N. Development of a HOG-based real-time PCR method to detect stress response changes in mycotoxigenic moulds. Food Microbiol. 2016, 57, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Monteau, S.; Abouna, S.; Lambert, B.; Legendre, L. Differential regulation by ambient pH of putative virulence factor secretion by the phytopathogenic fungus Botrytis cinerea. FEMS Microbiol. Ecol. 2003, 43, 359–366. [Google Scholar] [CrossRef]
- Zhang, X.; Zong, Y.; Gong, D.; Yu, L.; Sionov, E.; Bi, Y.; Prusky, D. NADPH oxidase regulates the growth and pathogenicity of Penicillium expansum. Front. Plant Sci. 2021, 12, 696210. [Google Scholar] [CrossRef]
- Visagie, C.M.; Houbraken, J.; Frisvad, J.C.; Hong, S.-B.; Klaassen, C.H.W.; Perrone, G.; Seifert, K.A.; Varga, J.; Yaguchi, T.; Samson, R.A. Identification and nomenclature of the genus Penicillium. Stud. Mycol. 2014, 78, 343–371. [Google Scholar] [CrossRef]
- Cabañas, R.; Abarca, M.L.; Bragulat, M.R.; Cabañes, F.J. Comparison of methods to detect resistance of Penicillium expansum to thiabendazole. Lett. Appl. Microbiol. 2009, 48, 241–246. [Google Scholar] [CrossRef]
- Li, H.X.; Xiao, C.L. Baseline sensitivities to fludioxonil and pyrimethanil in Penicillium expansum populations from apple in Washington State. Postharvest Biol. Technol. 2008, 47, 239–245. [Google Scholar] [CrossRef]
- Ali, E.M.; Amiri, A. Selection pressure pathways and mechanisms of resistance to demethylation inhibitor-Difenoconazole in Penicillium expansum. Front. Microbiol. 2018, 9, 2472. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research/0034.1. [Google Scholar] [CrossRef]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef]
- Cole, T.J.; Altman, D.G. Statistics notes: What is a percentage difference? BMJ 2017, 358, j3663. [Google Scholar] [CrossRef] [PubMed]
- Markoglou, A.N.; Doukas, E.G.; Malandrakis, A.A. Effect of anilinopyrimidine resistance on aflatoxin production and fitness parameters in Aspergillus parasiticus Speare. Int. J. Food Microbiol. 2011, 146, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Hiber, U.W.; Schüepp, H.; Schwinn, F.J. Resistance risk evaluation of fludioxonil, a new phenylpyrrole fungicide. In Fungicide Resistance, Monograph No.60; Heaney, S., Slawson, D., Hollomon, D.W., Smith, M., Russel, P.E., Perry, D.W., Eds.; British Crop Protection Council: Surrey, UK, 1994; pp. 397–402. [Google Scholar]
- Tudzynski, P.; Heller, J.; Siegmund, U. Reactive oxygen species generation in fungal development and pathogenesis. Curr. Opin. Microbiol. 2012, 15, 653–659. [Google Scholar] [CrossRef]
- Bilsland, E.; Molin, C.; Swaminathan, S.; Ramne, A.; Sunnerhagen, P. Rck1 and Rck2 MAPKAP kinases and the HOG pathway are required for oxidative stress resistance. Mol. Microbiol. 2004, 53, 1743–1756. [Google Scholar] [CrossRef] [PubMed]
- Segmüller, N.; Ellendorf, U.; Tudzynski, B.; Tudzynski, P. BcSAK1, a stress-activated mitogen-activated protein kinase, is involved in vegetative differentiation and pathogenicity in Botrytis cinerea. Eukaryot. Cells 2007, 6, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Cao, Q.; Li, N.; Liu, D.; Yuan, Y. Transcriptome analysis of fungicide-responsive gene expression profiles in two Penicillium italicum strains with different responses to the sterol demethylation inhibitor (DMI) fungicide prochloraz. BMC Genom. 2020, 21, 156. [Google Scholar] [CrossRef]
- Hawkins, N.J.; Fraaije, B.A. Fitness penalties in the evolution of fungicide resistance. Annu. Rev. Phytopathol. 2018, 56, 339–360. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).