Abstract
Drought and salt stresses severely impair plant growth and development worldwide. DEHYDRATION-RESPONSIVE ELEMENT BINDING proteins (DREBs), as a subfamily of the AP2/ERF transcription factor superfamily, play critical regulatory roles in plant biological processes including growth and development, as well as the adaptive response to various abiotic stresses. Based on the transcriptome data analysis of Medicago truncatula under saline-alkali stress previously conducted in our laboratory, a gene responsive to saline-alkali stress, Medtr3g110205, was identified, and its homologous gene in Arabidopsis thaliana, AtERF41 (AT5G11590), was obtained via BLAST (version BLAST+ 2.17.0.). The mutant erf41 was used to explore its biological functions in response to drought and salt stresses. The results showed that under salt and drought stress conditions, the seed germination rate, and growth status of the erf41 mutant were all better than those of the wild type. Further determination of physiological and biochemical indicators revealed that the leaf contents of superoxide dismutase (SOD) and proline (Pro) in the leaves of the mutant plants were significantly higher than those in the wild type, while the malondialdehyde (MDA) content was significantly decreased. In conclusion, the AtERF41 gene negatively regulates salt and drought tolerance in Arabidopsis thaliana, providing a potential target for the genetic improvement of crop stress tolerance. This study not only deepens our understanding of the role of DREB transcription factors in plant stress response but also provides a theoretical basis for improving crop stress tolerance using genetic engineering technology in the future.
1. Introduction
Dynamic environmental conditions such as drought and high salinity exert adverse effects on plant growth, survival, and reproduction [1]. Drought and high salt concentrations induce osmotic stress in plant cells and cause the accumulation of harmful ions, thereby leading to plant stunting or developmental retardation [2]. Additionally, drought and salt stress can induce the accumulation of reactive oxygen species (ROS), such as superoxide anions, hydrogen peroxide, and hydroxyl radicals. ROS possess strong oxidative activity that attacks cellular membranes and intracellular membrane systems, causing plasma membrane peroxidation, and leading to cell membrane damage and cell death, thereby impairing plant growth and development [3]. During long-term evolutionary adaptation, plants have developed corresponding mechanisms to cope with drought and salt stress, mitigating cellular damage to maintain survival under stressful conditions [4]. Notably, the AP2/ERF transcription factors are indispensable in mediating plant responses to osmotic stress [4].
AP2/ERF transcription factors are key regulators in plants, participating in the modulation of plant growth and development, biosynthesis, and stress responses [5]. AP2/ERF contains 1–2 AP2/ERF domains, which consist of approximately 60–70 conserved amino acids involved in DNA binding. Based on the number of AP2/ERF domains and their binding sequences, AP2/ERF transcription factors are classified into five subfamilies: AP2, ERF, DREB, RAV, and Soloist [6,7]. Specifically, the AP2 subfamily comprises two AP2/ERF domains, whereas the ERF and DREB subfamilies each harbor a single AP2/ERF domain. The RAV subfamily contains one AP2/ERF domain together with one B3 domain; while the Soloist subfamily also contains one AP2/ERF domain but exhibits significant structural differences from other subfamilies [8]. Studies have demonstrated that overexpression of OsERF19 enhances salt tolerance in rice [9]. GmERF75 is induced by multiple abiotic stresses and exogenous phytohormones, and overexpression of this gene improves osmotic stress tolerance in both Arabidopsis and soybean [10]. Similarly, in rice, OsERF115/AP2EREBP110 has been found to be induced by high-temperature and drought treatments; its overexpression enhances the heat tolerance of plants at both the seed and vegetative growth stages, and improves plant drought tolerance by increasing the accumulation of osmotic regulators [11]. Additionally, research indicates that overexpression of GmDREB1 markedly enhances drought tolerance in soybean [12].
Previous studies have identified 147 AP2/ERF genes in Arabidopsis thaliana, of which 122 belong to the ERF (including DREB) subfamily; among these 122 ERF genes, 57 are members of the DREB subfamily [13]. TINY is a member of the DEHYDRATION-RESPONSIVE ELEMENT BINDING protein A4 subfamily of the AP2/ERF transcription factor family, which comprises 17 members in Arabidopsis, and both TINY and ERF41 (also known as TINY2) are classified into this clade [13]. Studies have demonstrated that TINY transcript levels are strongly induced by multiple stresses, including dehydration, cold, and salt. Additionally, TINY overexpression is associated with elevated expression of drought-responsive genes, as well as hypersensitivity to ABA-mediated inhibition of seed germination and root growth [14,15,16]. Wei et al. demonstrated that TINY2 is induced by ABA, cold, wounding, NaCl, and drought treatments, suggesting that TINY2 may be involved in activating downstream genes in response to environmental stress [17].
In conclusion, we speculate that ERF41 plays a role in plant responses to drought and salt stresses, but its specific function remains unclear. To explore the function of AtERF41 in plant responses to drought and salt stress, we utilized the Arabidopsis mutant erf41 as experimental material to study the function of the ERF41 gene, aiming to provide theoretical insights for understanding the mechanisms of plant responses to drought and salt stress.
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
The wild-type Arabidopsis thaliana was purchased from ABRC (Arabidopsis Biological Resource Center) as the Columbia-0 (Col-0) line, and the mutant erf41 (N690869) was obtained from the Arashare website. Arabidopsis seeds were first rinsed twice with water, then treated with 75% alcohol for 1 min, followed by exposure to 1% NaClO for 10 min, and finally washed five times with distilled water. Seeds were sown on 1/2MS medium plates containing 30 g/L sucrose and 6 g/L agar. The seeded plates were vernalized at 4 °C in the dark for 2 days and then moved to 22 °C under a 16/8 light/dark cycle. When Arabidopsis sprouted, it was transplanted at a density of four plants per pot with a mixture of soil at 22 °C under a 16/8 light/dark cycle and 70% relative humidity.
2.2. Analysis of the Physicochemical Properties and Conserved Domains of AtERF41
The physicochemical properties of the AtERF41 protein were analyzed using the ExPASY ProtParam tool (https://web.expasy.org/protparam/, accessed on 3 February 2026). The transmembrane regions, hydrophilicity, and signal peptides of the protein were predicted using TMHMM 2.0 Server (https://services.healthtech.dtu.dk/services/TMHMM-2.0/, accessed on 3 February 2026), ExPASY Protscale (https://web.expasy.org/protscale/, accessed on 3 February 2026), and SignalP 4.1 Server (https://services.healthtech.dtu.dk/services/SignalP-4.1/, accessed on 3 February 2026), respectively. The secondary and tertiary structures of the protein were predicted using the online SOPMA (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_sopma.html, accessed on 3 February 2026) and SWISS-MODEL (https://swissmodel.expasy.org/, accessed on 3 February 2026) tools, respectively. The conserved domains of the amino acid sequences encoded by the gene were analyzed using the Conserved Domain Database (https://www.ncbi.nlm.nih.gov/cdd/?term=, accessed on 3 February 2026).
2.3. Identification of Arabidopsis erf41 Mutant Homozygotes
The Arabidopsis thaliana mutant seeds were cultured on 1/2 MS medium as previously described. After seedling emergence, homozygosity was confirmed by PCR.
Homozygous mutation identification was performed using the three-primer method. The LP, RP, and BP primers for the mutants were obtained from the website (http://signal.salk.edu/tdnaprimers.2.html?tdsourcetag=s_pcqq_aiomsg, accessed on 3 February 2026) for PCR detection (Table 1). PCR was conducted using the ExTaq enzyme (Aidlab Biotech, Beijing, China) with the primers LP + RP and BP + RP. The principle is as follows: the primers LP + RP and BP + RP can simultaneously amplify products of a larger molecular weight product (large fragments of the wild-type gene) and a smaller molecular weight product (small fragments of T-DNA insertion). In homozygous mutants, only the BP + RP primer can amplify the smaller molecular weight product, while the LP + RP primer fails to produce any product.

Table 1.
Primer information.
2.4. Quantitative Real-Time PCR Analysis
RNA was extracted from the leaves of Col-0 and erf41 plants using the Plant Total RNA purification kit with DNase I (Aidlab), followed by the synthesis of complementary DNA (cDNA) using the Tiangen FastQuant RT Kit (Tiangen, Beijing, China) following the manufacturer’s instructions. Subsequently, quantitative real-time PCR (RT-qPCR) was performed using SYBR Green fluorescence detection. For normalization, Arabidopsis Actin was used as an internal standard. Five biological replicates × four technical replicates were used for qRT-PCR to ensure the reproducibility of the results. The primers are listed in Table 2.

Table 2.
Primer information for RT-qPCR.
2.5. Physiological Experiments
To determine germination rates of Col-0 and mutant erf41 seeds, 90 (30 biological replicates × 3 technical replicates) seeds from each genotype (Col-0 and mutant erf41) were sown individually on the same 1/2MS medium supplemented with or without NaCl (100 mM, 125 mM, 150 mM) and mannitol (200 mM, 300 mM), with daily recordings of germination rates. Following germination, 15 (5 biological replicates × 3 technical replicates) seedlings per genotype were selected and vertically cultured on medium with or without NaCl stress (100 mM, 125 mM, 150 mM) for 10 days, after which root lengths were measured.
To measure the activity of ROS-scavenging enzymes, Arabidopsis thaliana leaves were collected before and after treatment. Then, soluble proteins were extracted using physiological salt buffer, and the activities of MDA, SOD, Pro, and catalase (CAT) were measured using a kit from the Nanjing Jiancheng Bioengineering Institute (Nanjing, China). Here, 45 replicates (15 biological replicates × 3 technical replicates) were performed in each experiment.
2.6. Salt and Drought Experiments
For salt stress treatment, Col-0 and erf41 mutant Arabidopsis seeds were sown on 1/2 MS medium. After developing two true leaves, the seedlings were transplanted into pots (four plants per pot) containing a soil mixture, and cultured in a greenhouse at 22 °C with a 16/8 h light/dark cycle and 70% relative humidity. The seedlings were watered normally for three weeks, after which they were irrigated with a 300 mM NaCl solution every 3 days for 15 days to observe phenotypic changes.
For the drought stress tolerance test, plants were grown in well-watered soil for three weeks before water was withheld until phenotypic differences appeared among the genotypes.
2.7. Statistical Analysis of Data
Statistical analysis of experimental data was carried out using SPSS17.0 software. One-way ANOVA with Tukey–Kramer multiple comparisons tests was performed on the relative gene expression. Two-way ANOVA with Tukey–Kramer multiple comparisons was used for the analysis of seed germination rate, root length, SOD activity, MDA content, and Pro content. Image editing and processing were performed using Photoshop.
3. Results
3.1. Physicochemical Properties and Conserved Domains Analysis of AtERF41
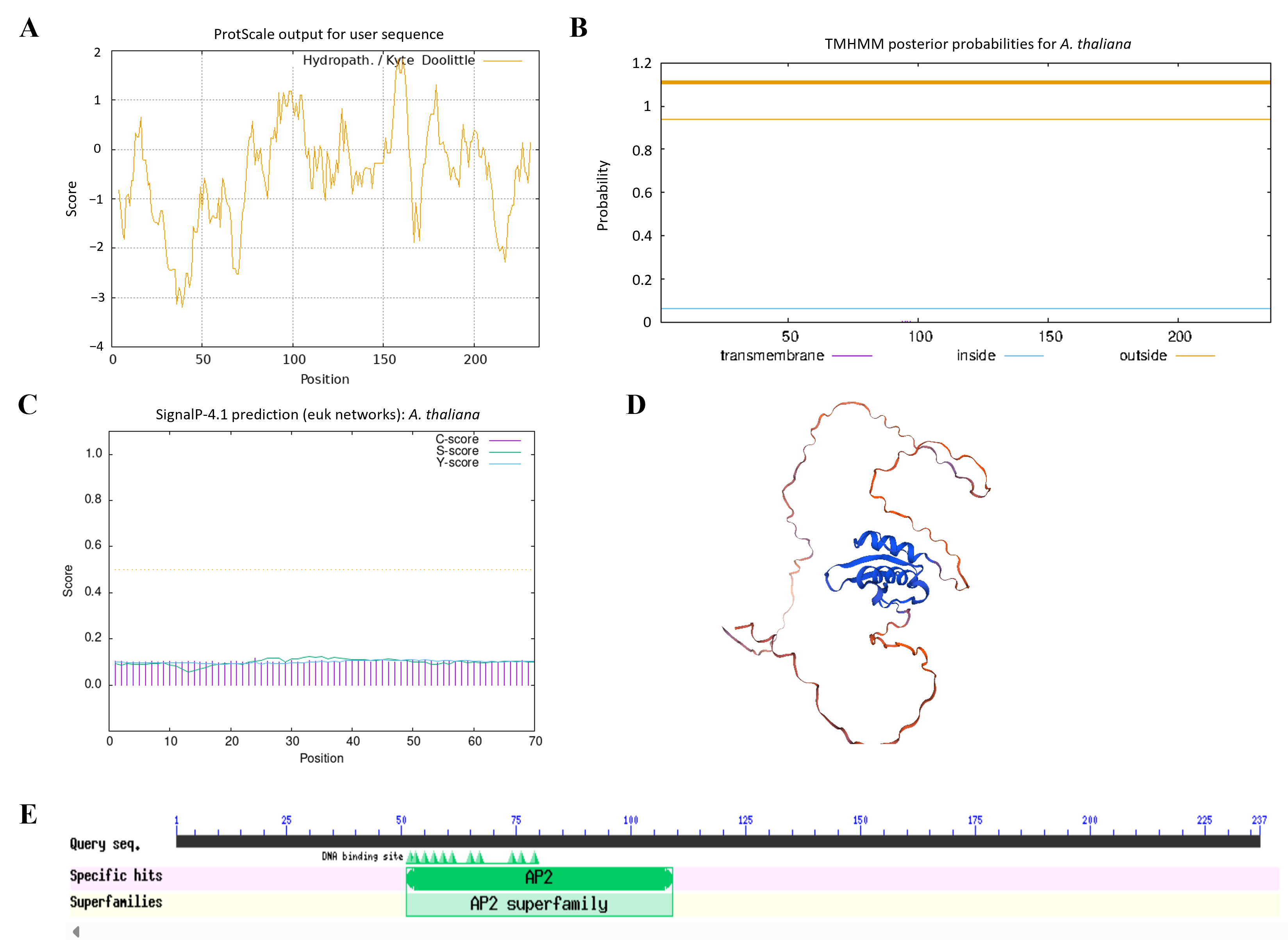
To investigate the structural characteristics of AtERF41, the physicochemical properties of the protein encoded by the AtERF41 gene were analyzed using ProtParam. The results indicated that this gene encodes a protein consisting of 236 amino acids, with the molecular formula C1143H1777N315O370S6, a molecular weight of 26.043 kDa, and an isoelectric point (pI) of 5.32. Its instability index was calculated as 54.34, classifying it as an unstable protein (proteins with an instability index below 40 are considered stable). Hydrophilicity-hydrophobicity analysis revealed a grand average of hydropathicity (GRAVY value of −0.531), indicating that this protein is hydrophilic (Figure 1A). Transmembrane structure prediction using TMHMM showed that the protein encoded by ERF41 lacks transmembrane helical regions (Figure 1B). Analysis with the SignalP 4.1 Server demonstrated that this protein does not contain a signal peptide, suggesting that it is a non-secretory protein (Figure 1C). Secondary structure prediction using the SOPMA online tool revealed that the protein is primarily composed of α-helices (25.85%), random coils (68.22%), and extended chains (5.93%), among which random coils accounted for the highest proportion. The tertiary structure of the protein encoded by MsERF41 predicted by SWISS-MODEL was consistent with the secondary structure prediction results (Figure 1D). Conserved domain analysis of the ERF41 protein using the Conserved Domain Database (CDD) identified one conserved AP2 domain, confirming that this gene belongs to the ERF transcription factor family (Figure 1E).
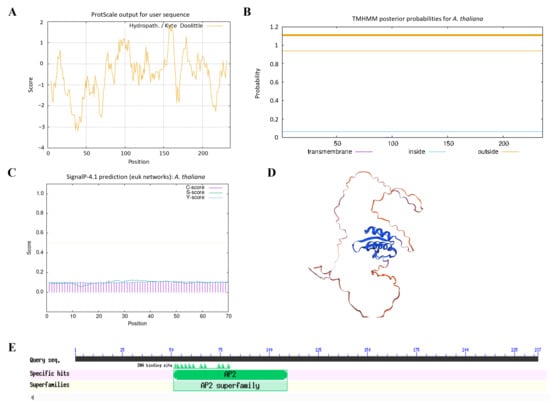
Figure 1.
Physicochemical properties and conserved domains analysis of AtERF41. Hydrophilic and hydrophobic (A), transmembrane structure (B), signal peptide prediction (C), tertiary structure prediction (D), conserved domain analysis (E) of the AtERF41 protein.
3.2. Identification and Expression Analysis of Arabidopsis erf41 Mutant Homozygotes
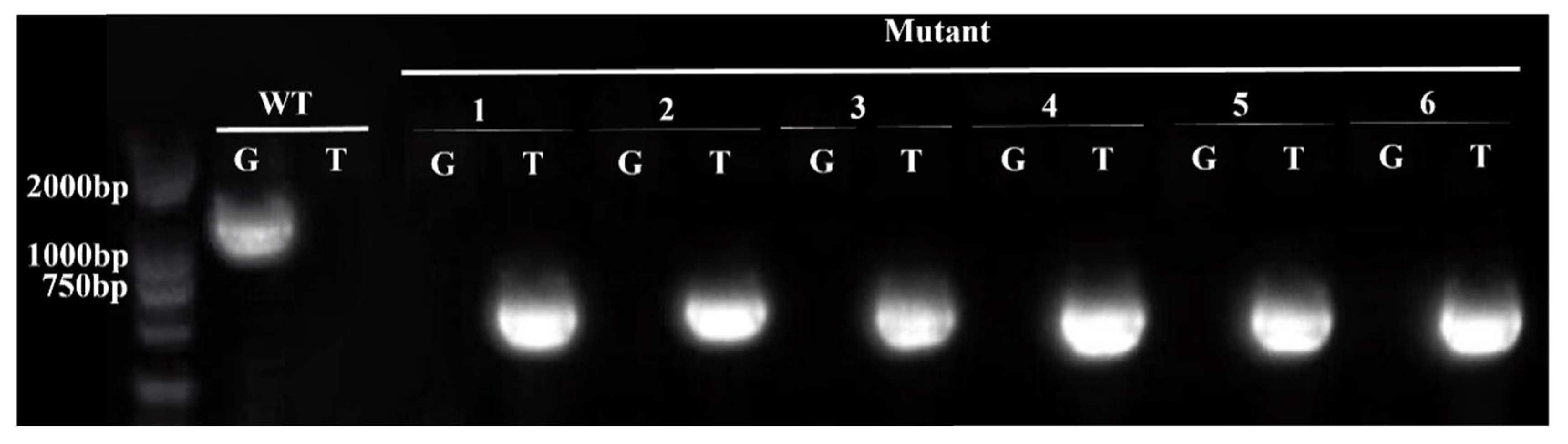
DNA was extracted from the Arabidopsis thaliana mutant erf41 and Col-0 plants, followed by PCR amplification using LP + RP and BP + RP as primer pairs (Table 1), with the expected band sizes of 1194 bp and 519 bp, respectively. As shown in Figure 2, agarose gel electrophoresis results confirmed that lines 1, 2, 3, 4, 5, and 6 were homozygous mutant plants based on the principle of the three-primer method. Seeds of the verified homozygous mutants were harvested, dried, and then stored in a dry place.
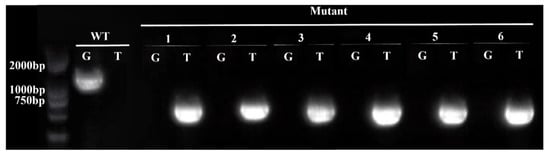
Figure 2.
Identification of homozygous erf41 mutant plants in Arabidopsis thaliana.
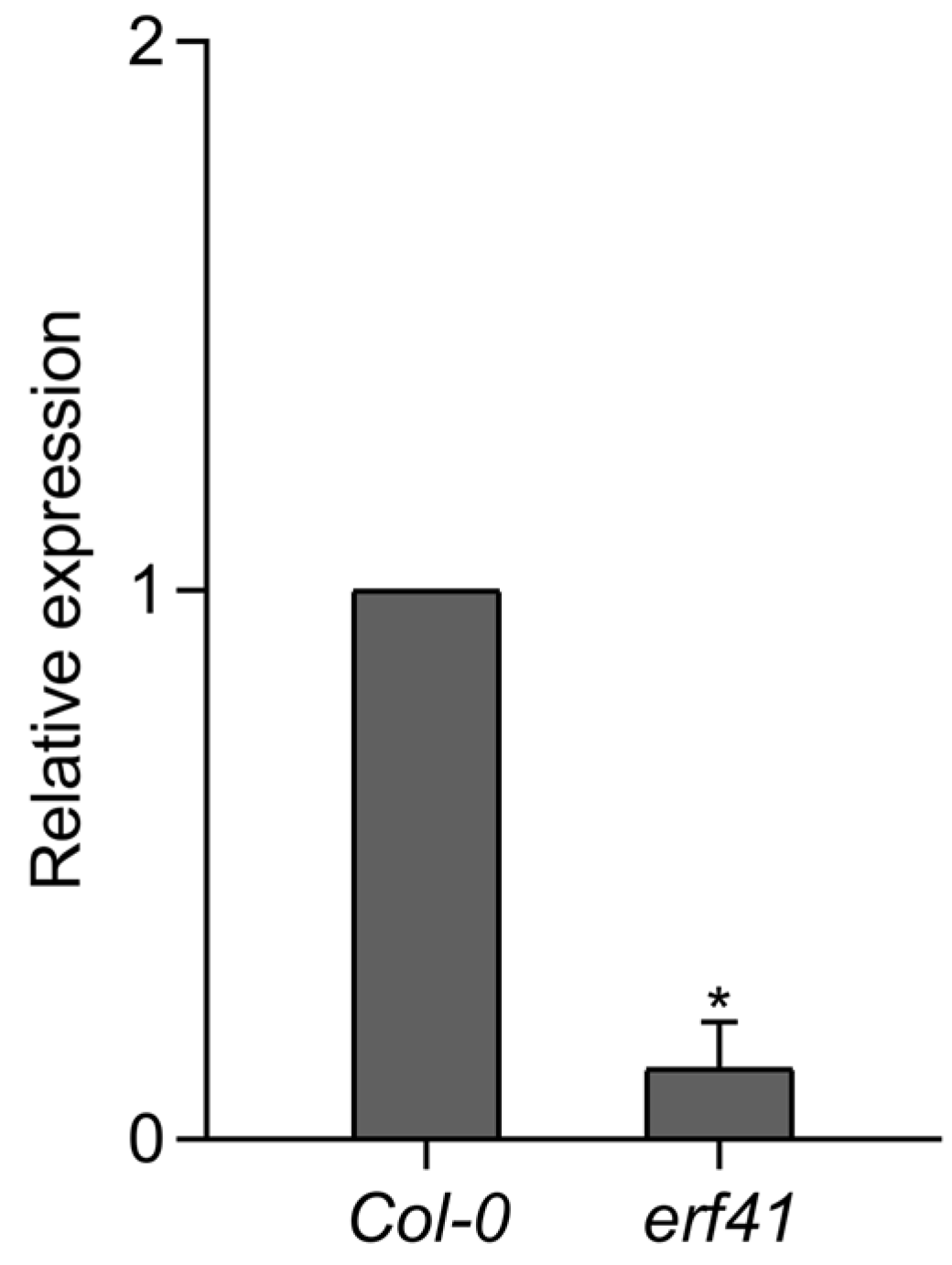
Next, to analyze the expression of the ERF41 gene in the erf41 mutant plants, RT-qPCR was performed on leaves of Col-0 and mutant plants. The results showed that ERF41 was barely expressed in the erf41 mutant plants, indicating that the ERF41 gene was disrupted and thus failed to function normally (Figure 3).
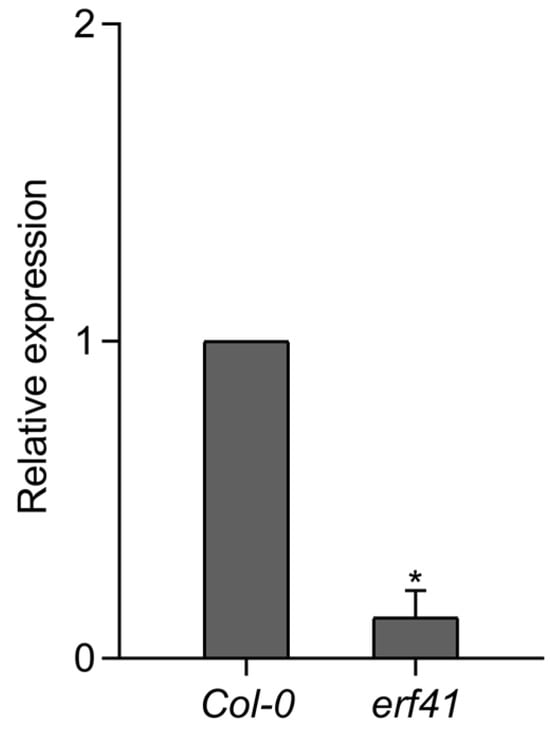
Figure 3.
Expression analysis of the AtERF41 in Col-0 and erf41 mutant plants. Data are presented as means ± SE (n = 5 biological replicates per genotype). The asterisks (*) represent significant differences (* p < 0.05).
3.3. ERF41 Negatively Regulates Salt Tolerance in Arabidopsis thaliana
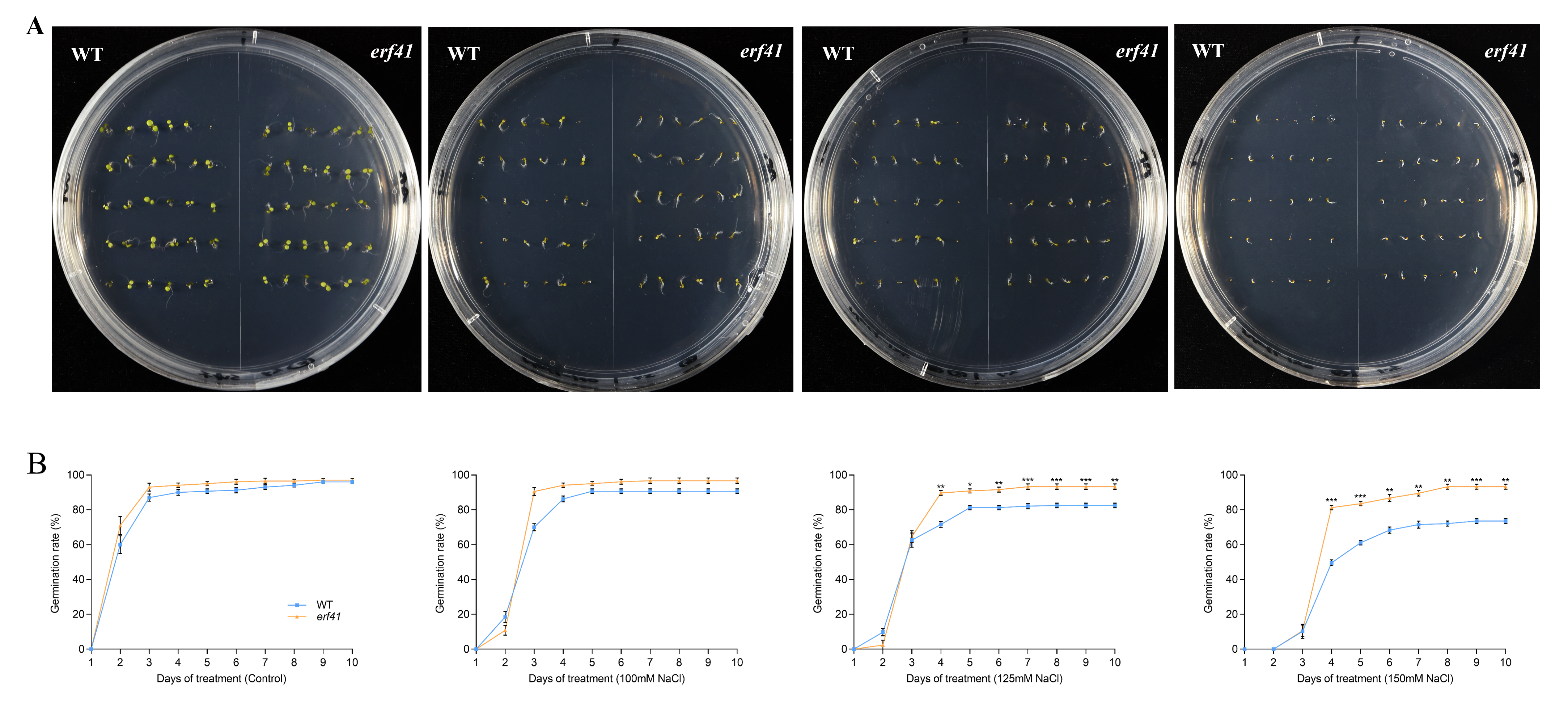
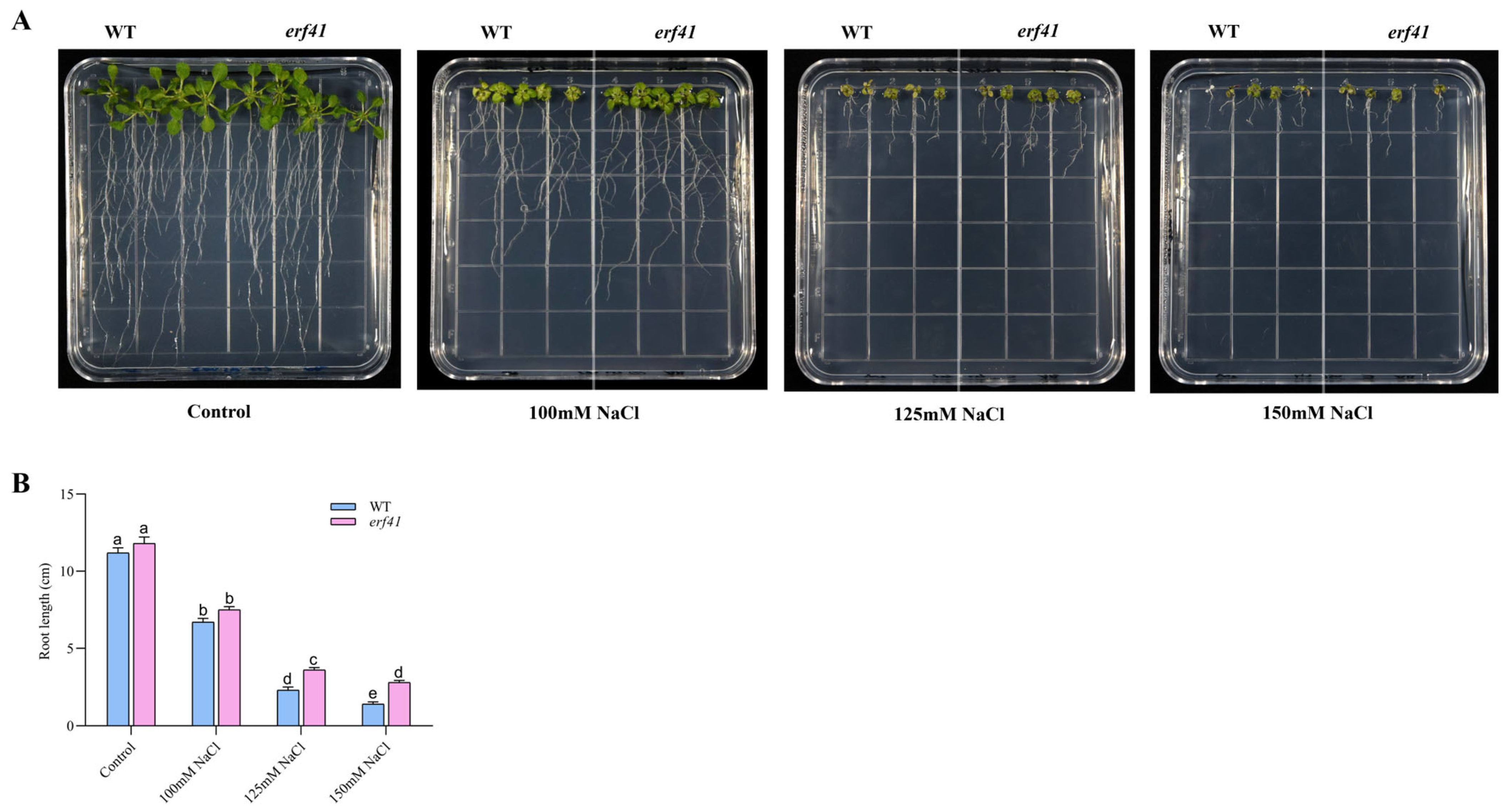
A series of experiments were performed to examine the differences in salt tolerance of Col-0 and erf41 plants. The seeds of Col-0 and erf41 were sown on 1/2 MS culture containing 100 mM, 125 mM, and 150 mM NaCl to observe germination rates. The results showed that on normal 1/2 MS medium, there was no significant difference in the seed germination rate among different Arabidopsis thaliana genotypes, all of which were close to 100%. Under the treatments of 100 mM, 125 mM and 150 mM NaCl, the germination rates of Col-0 seeds were 90.7%, 82.5% and 73.6%, respectively, while those of erf41 seeds were 96.7%, 93.3% and 93.3%, which were significantly higher than those of Col-0 seeds (Figure 4A,B). In addition, the primary root length of the seedlings differed after 10 days of vertical growth under salt stress. Results showed no significant difference in root length among genotypes on normal 1/2 MS medium. However, in salt-treated conditions, erf41 plants exhibited significantly longer roots than Col-0 plants (Figure 5A,B). It was preliminarily inferred that ERF41 plays a role under salt stress and negatively regulates the salt tolerance of Arabidopsis thaliana.
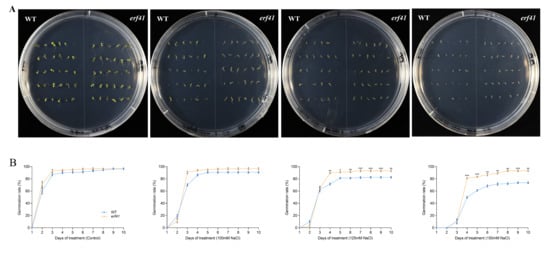
Figure 4.
Germination rate of wild-type and erf41 mutant plants under salt stress. Germination performance (A) and germination rate (B) of wild-type and mutant seeds under salt stress. The data are represented as means ± SE (n = 30 biological replicates per genotype). The asterisks (*, ** and ***) represent significant differences (* p < 0.05, ** p < 0.01, and *** p < 0.001) compared with WT plants at the same time point.
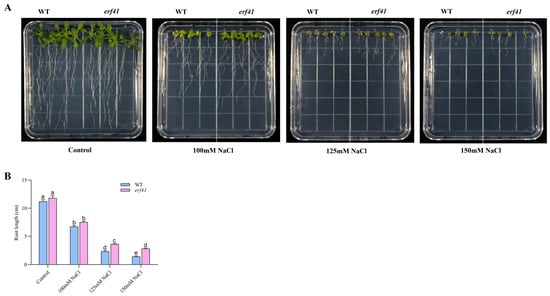
Figure 5.
Morphological analysis of roots from wild-type and erf41mutant plants grown on 1/2 MS medium with or without NaCl. (A) Morphology analysis. (B) The primary root length. The data are represented as means ± SE (n = 5 biological replicates per genotype). Different letters represent significant differences (p < 0.05) compared to the control.
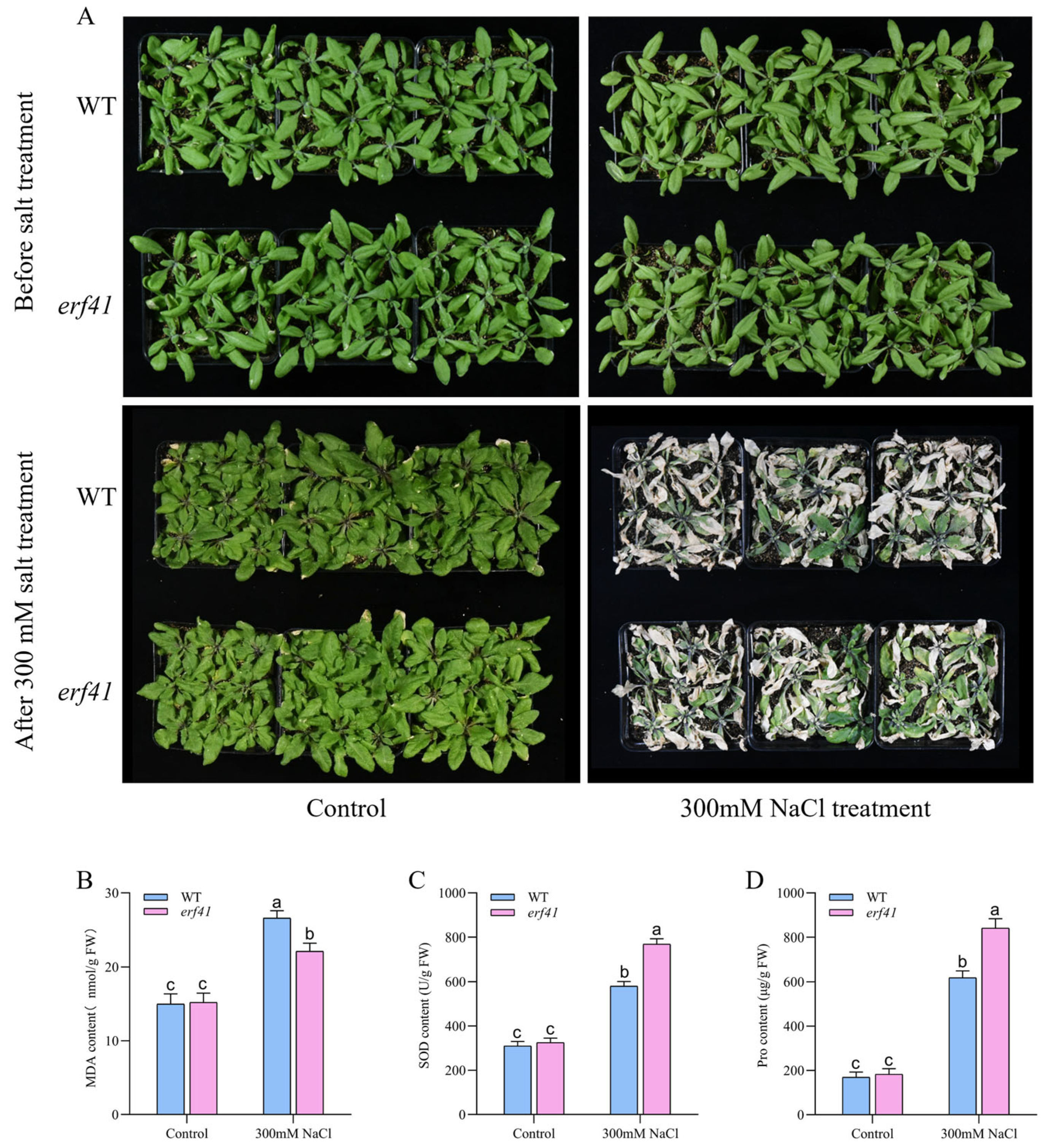
To further investigate the function of ERF41 in Arabidopsis under salt stress, potted plants were experimentally treated. Arabidopsis wild-type (Col-0) and mutant (erf41) plants were treated with a 300 mM salt solution. Phenotypic analysis revealed that under normal conditions, the growth status of Col-0 and erf41 plants showed no statistically significant difference, both of which could grow normally. After salt treatment, leaves of the Col-0 plants exhibited wilting and drying, and the plants were nearly dead, whereas leaves of the erf41 plants only exhibited mild wilting, with only the basal leaves withering, but they could still grow normally (Figure 6A). Then, the physiological indicators of MDA, SOD, and Pro in leaves of wild-type (Col-0) and mutant (erf41) plants under normal and salt treatment conditions were measured (Figure 6B–D). The results showed that under normal watering conditions, there were no significant differences in these physiological indicators between Col-0 and erf41 plants. Under salt solution treatment, the contents of MDA, SOD, and Pro in leaves of all genotypes increased, but the contents of SOD and Pro in mutant plant leaves were significantly higher than those in Col-0 plants, while the content of MDA was significantly lower than that in Col-0 plants. This further indicates that the ERF41 gene exerts a negative regulatory effect on salt tolerance in Arabidopsis thaliana.
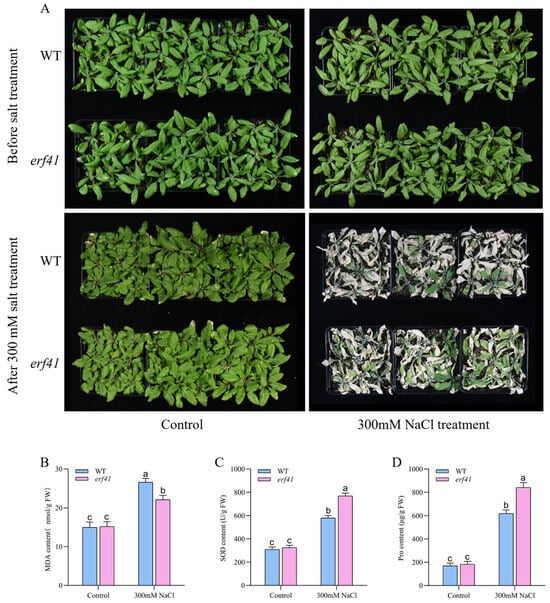
Figure 6.
AtERF41 negatively regulates salt tolerance in Arabidopsis thaliana. (A) Morphological differences in plants under salt experiments. The content of MDA (B), SOD (C), Pro (D). Data are means ± SE (n = 15 biological replicates per genotype). Different letters denote significant differences: p < 0.05.
3.4. ERF41 Negatively Regulates Drought Tolerance in Arabidopsis thaliana
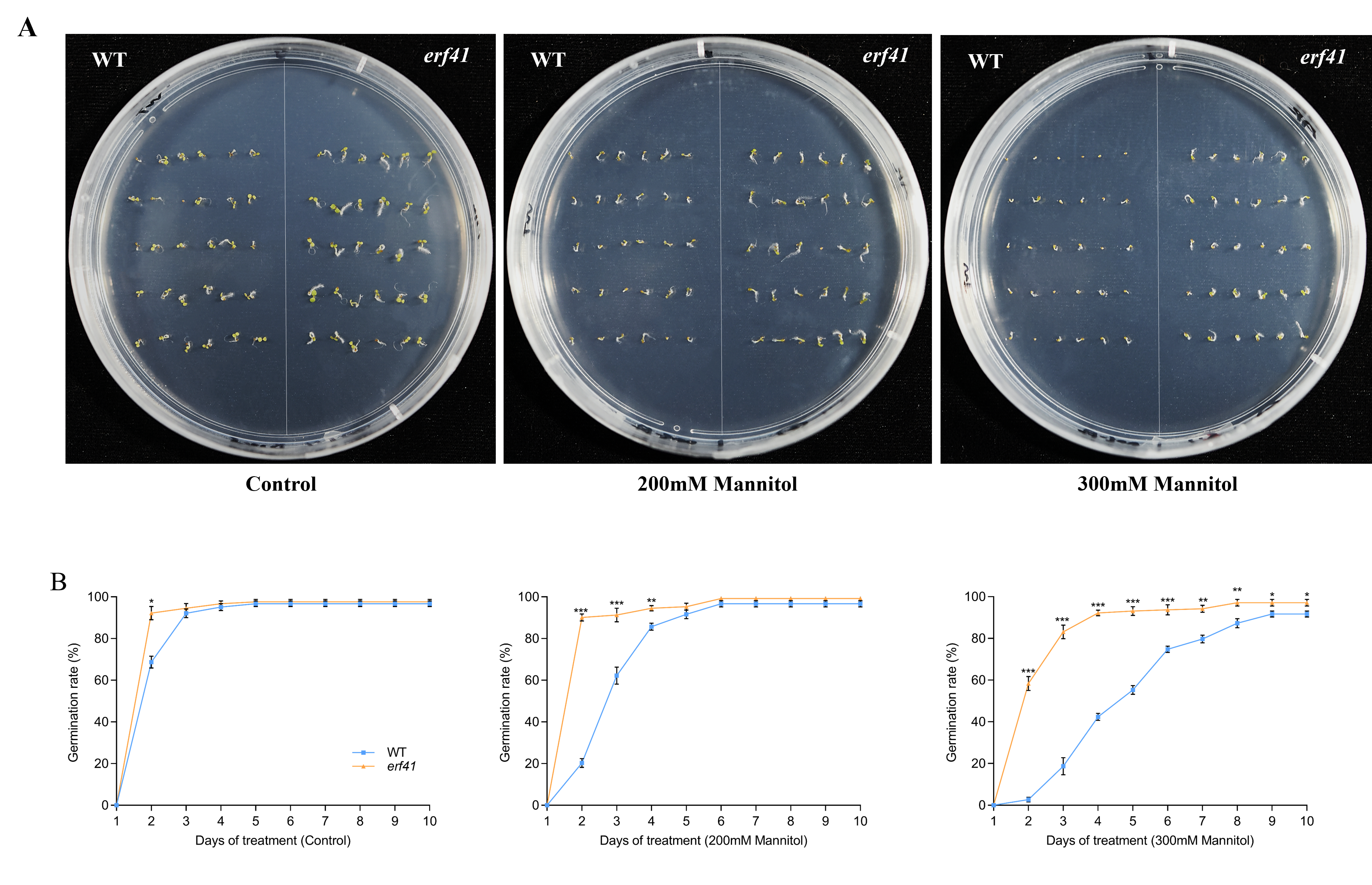
To explore the function of ERF41 in Arabidopsis thaliana responses to drought stress, we statistically analyzed the seed germination of wild-type (Col-0) and mutant (erf41) plants under treatments with different concentrations of mannitol (Figure 7A). The results showed that on normal 1/2 MS medium, there was no significant difference in the seed germination rate among different Arabidopsis genotypes, with all rates close to 100%. Under the treatments of 200 mM and 300 mM mannitol, the germination rates of Col-0 seeds were 96.7% and 91.7%, respectively, whereas those of erf41 seeds were 99.2% and 97.1%, which were significantly higher than those of Col-0 seeds (Figure 7B). It was preliminarily inferred that ERF41 exerts a function under drought stress and negatively regulates the drought tolerance of Arabidopsis thaliana.
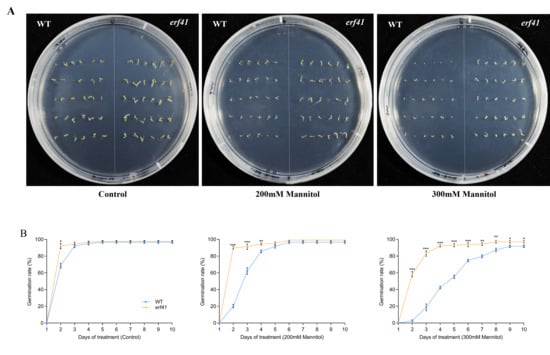
Figure 7.
Germination conditions of WT and erf41 mutant under drought stress. Germination performance (A) and germination rate (B) of wild-type and mutant seeds under drought stress. The data are represented as means ± SE (n = 30 biological replicates per genotype). The asterisks (*, ** and ***) represent significant differences (* p < 0.05, ** p < 0.01, and *** p < 0.001) compared with WT plants at the same time point.
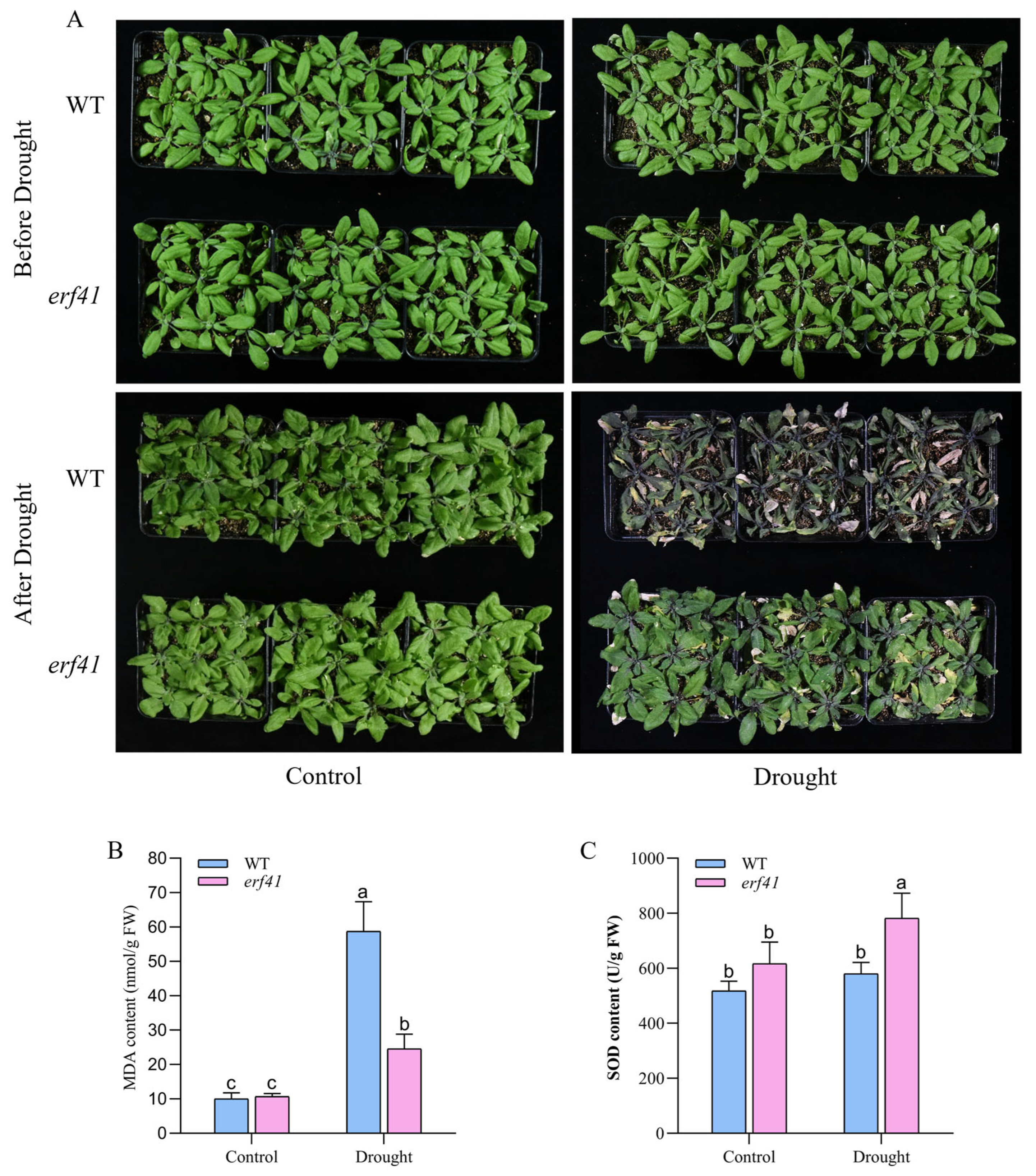
To further explore the role of the ERF41 gene in Arabidopsis thaliana responses to drought stress, wild-type (Col-0) and mutant (erf41) Arabidopsis plants were subjected to drought treatment. Phenotypic analysis revealed that under normal growth conditions, Col-0 and erf41 plants exhibited similar growth phenotypes with no significant difference, with both genotypes growing normally. After drought treatment, the leaves of wild-type Arabidopsis exhibited severe curling and chlorosis, followed by wilting and desiccation, and the plants were nearly dead. In contrast, the leaves of erf41 plants showed less chlorosis than those of the wild type and maintained a much better growth status. These results indicated that the erf41 mutant had enhanced drought tolerance under drought stress conditions. (Figure 8A). Subsequently, the physiological indices of MDA and SOD in the leaves of wild-type (Col-0) and mutant (erf41) plants were determined under both normal and drought conditions (Figure 8B,C). The results showed that there were no significant differences in these physiological indices between the leaves of Col-0 and mutant erf41 plants under well-watered conditions. Under drought treatment, the MDA content in the leaves of erf41 mutants was significantly lower than that in Col-0 plants, whereas the SOD content in the mutant leaves was significantly higher than that in Col-0 plants. These findings demonstrate that ERF41 plays a role in response to drought stress and negatively regulates drought tolerance in Arabidopsis thaliana.
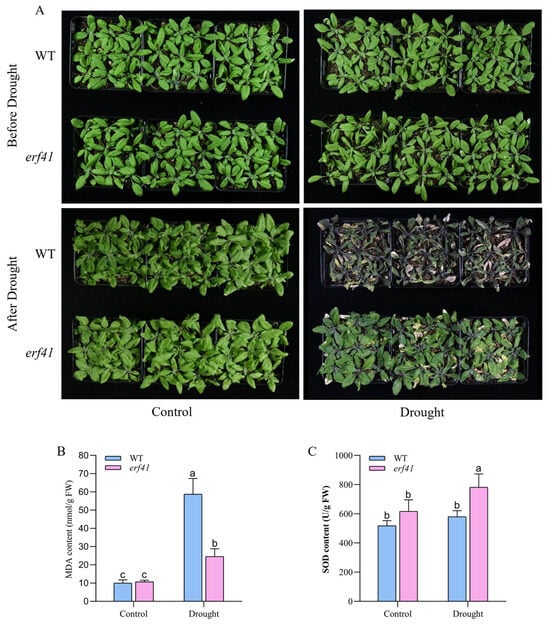
Figure 8.
AtERF41 negatively regulates drought tolerance in Arabidopsis thaliana. (A) Morphological differences in plants under drought experiments. The content of MDA (B), SOD (C). Data are means ± SE (n = 15 biological replicates per genotype). Different letters denote significant differences: p < 0.05.
4. Discussion
Abiotic stresses such as drought, high temperature, and high salinity are major limiting factors for global agricultural production, which inhibit plant growth and development and ultimately lead to significant reductions in crop biomass and yield [18]. To survive under adverse environmental conditions, plants have evolved sophisticated and precise regulatory networks to perceive stress signals, activate downstream stress-responsive genes, and initiate a series of physiological and biochemical responses to mitigate stress damage. The AP2/ERF transcription factors are key regulators of ethylene-responsive gene expression, participating in plant growth and development, primary and secondary metabolism, as well as responses to environmental stresses [5]. Previously, our laboratory identified an AP2/ERF transcription factor member, Medtr3g110205, which responds to salt-alkali stress, via transcriptome analysis of alfalfa under salt-alkali stress. In this study, we identified the homolog of Medtr3g110205 in Arabidopsis thaliana, designated ERF41, and performed functional characterization of AtERF41. Conserved domain analysis of the ERF41 protein revealed that it contains one conserved AP2 domain (Figure 1E). Moreover, previous studies have demonstrated that ERF41, also known as TINY2, belongs to the dehydration-responsive element-binding protein A4 (DREB A4) subfamily within the AP2/ERF transcription factor family [13]. And, ERF41 is induced by ABA, cold, wounding, NaCl, and drought treatments [17]. However, the function of ERF41 in plant responses to drought and salt stresses remains unclear.
Through genetic experiments and physiological and biochemical analyses, this study found that the Arabidopsis thaliana mutant erf41 exhibited enhanced tolerance to both salt and drought stresses. Under drought and salt stress conditions, plant cells rapidly produce and accumulate substantial amounts of reactive oxygen species (ROS), which induce severe oxidative damage [19,20,21]. Malondialdehyde (MDA) serves as a key physiological marker for assessing the extent of plasma membrane peroxidation [20], while proline—a widely accumulated osmolyte under such stresses—plays a critical role in maintaining protein stability [22]. Additionally, superoxide dismutase (SOD) functions to scavenge ROS generated in plants exposed to abiotic stress, thereby facilitating survival under adverse environmental conditions [23]. Therefore, these stress-related physiological indicators enable the rapid and accurate evaluation of plant tolerance to abiotic stresses such as drought and salt. In this study, compared with wild-type plants, the erf41 mutant plants showed significantly increased SOD and proline contents and significantly decreased MDA content in leaves under both salt and drought stresses. These results provide direct evidence that AtERF41 negatively regulates salt and drought tolerance in Arabidopsis thaliana (Figure 6 and Figure 8). In addition, the erf41 mutant plants exhibited significantly improved seed germination and growth performance under salt and drought stresses, which further confirmed that ERF41 negatively regulates salt and drought tolerance in Arabidopsis thaliana. Previous studies have shown that overexpression of the TaERF6-3A gene in Arabidopsis thaliana leads to the downregulation of genes related to salt stress and antioxidation, thereby negatively regulating salt and drought tolerance in Arabidopsis thaliana [24].
Numerous studies have demonstrated that overexpression of OsERF19 can improve salt and drought tolerance in rice [9]. Overexpression of the GhERF13.12 gene in Arabidopsis thaliana resulted in transgenic plants with stronger salt tolerance compared with the control group. In contrast, silencing of the GhERF13.12 gene increased the sensitivity of cotton to salt stress, indicating that GhERF13.12 positively regulates salt tolerance in both Arabidopsis thaliana and cotton [25]. Different from most reports that ERF transcription factors positively regulate stress resistance, the negative regulatory role of AtERF41 reveals the functional diversity of the ERF family. AtERF41 is closely related to TINY, which also belongs to the DREB A4 subfamily, in terms of evolutionary relationship [13]. However, TINY can positively regulate plant drought tolerance by activating drought-responsive genes and actively promoting ABA-mediated stomatal closure [14], whereas AtERF41 exerts the opposite negative regulatory function. This phenomenon fully suggests that there is significant functional differentiation among members of the DREB A4 subfamily, which is worthy of further investigation. We speculate that ERF41 may directly or indirectly suppress the expression of stress-responsive genes, including those involved in ABA signal transduction, reactive oxygen species scavenging, osmotic adjustment, and stress protection. Therefore, the loss of function of ERF41 releases this inhibition, leading to elevated expression of stress-related genes and enhanced adaptation to salt and drought stresses. Further investigation into the direct target genes of ERF41 will help clarify the precise molecular regulatory network underlying ERF41-mediated stress responses.
This study expands the understanding of the role of ERF transcription factors in plant stress responses, and proposes that ERF genes may achieve the adaptive balance of plants to environmental changes through the precise regulation of the expression of stress-responsive genes. However, this study was mainly conducted in Arabidopsis thaliana, and its conclusions need to be verified in more species. Future research should further explore the specific mechanism of action of AtERF41, including its interacting proteins and downstream target genes. Meanwhile, field trials should be carried out to evaluate the practical application effects in agricultural production.
5. Conclusions
In this study, we thoroughly investigated the function of the AtERF41 gene in salt and drought tolerance of Arabidopsis thaliana via bioinformatics analysis, genetic experiments, and the determination of physiological and biochemical indices. Furthermore, we successfully identified and obtained homozygous erf41 mutant plants of Arabidopsis thaliana. Phenotypic analysis under treatments with different concentrations of salt and mannitol revealed that the erf41 mutant exhibited distinct growth characteristics compared with the Col-0 under both salt and drought stresses. Specifically, the seed germination rate of the erf41 mutant was higher than that of the Col-0 under salt and drought stresses. Moreover, its leaves showed milder wilting and maintained normal growth after exposure to salt and drought treatments. Meanwhile, the results of physiological and biochemical index determination indicated that under salt and drought stresses, the SOD content in the leaves of the mutant plants was significantly higher than that of the wild type, whereas the MDA content was significantly lower. These findings further confirmed the negative regulatory role of the AtERF41 gene in salt and drought tolerance of Arabidopsis thaliana.
Author Contributions
Conceptualization, Y.Z.; methodology, J.W. and M.L.; formal analysis and investigation, H.X.; writing—original draft preparation, J.W., M.L. and Y.Z.; writing—review and editing, J.W. and M.L.; funding acquisition, Y.Z. All authors have read and agreed to the published version of the manuscript.
Funding
The research was financially supported by the Natural Science Foundation of Shandong Province (Grant No. ZR2023QC143).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF Transcription Factor Regulatory Networks in Hormone and Abiotic Stress Responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef]
- Rai, G.K.; Mishra, S.; Chouhan, R.; Mushtaq, M.; Chowdhary, A.A.; Rai, P.K.; Kumar, R.R.; Kumar, P.; Perez-Alfocea, F.; Colla, G.; et al. Plant salinity stress, sensing, and its mitigation through WRKY. Front. Plant Sci. 2023, 14, 1238507. [Google Scholar] [CrossRef]
- Mittler, R.; Zandalinas, S.I.; Fichman, Y.; Van Breusegem, F. Reactive oxygen species signalling in plant stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 663–679. [Google Scholar] [CrossRef]
- Wang, K.; Guo, H.Q.; Yin, Y.H. AP2/ERF transcription factors and their functions in Arabidopsis responses to abiotic stresses. Environ. Exp. Bot. 2024, 222, 105763. [Google Scholar] [CrossRef]
- Feng, K.; Hou, X.L.; Xing, G.M.; Liu, J.X.; Duan, A.Q.; Xu, Z.S.; Li, M.Y.; Zhuang, J.; Xiong, A.S. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 2020, 40, 750–776. [Google Scholar] [CrossRef]
- Sakuma, Y.; Liu, Q.; Dubouzet, J.G.; Abe, H.; Shinozaki, K.; Yamaguchi-Shinozaki, K. DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem. Biophys. Res. Commun. 2002, 290, 998–1009. [Google Scholar] [CrossRef]
- Phukan, U.J.; Jeena, G.S.; Tripathi, V.; Shukla, R.K. Regulation of Apetala2/Ethylene Response Factors in Plants. Front. Plant Sci. 2017, 8, 150. [Google Scholar] [CrossRef]
- Yue, M.F.; Zhang, C.; Wu, Z.Y. Research Progress in the Structural and Functional Analysis of PlantTranscription Factor AP2/ERF Protein Family. Biotechnol. Bull. 2022, 38, 11–26. [Google Scholar]
- Huang, S.; Ma, Z.; Hu, L.; Huang, K.; Zhang, M.; Zhang, S.; Jiang, W.; Wu, T.; Du, X. Involvement of rice transcription factor OsERF19 in response to ABA and salt stress responses. Plant Physiol. Biochem. 2021, 167, 22–30. [Google Scholar] [CrossRef]
- Zhao, M.J.; Yin, L.J.; Liu, Y.; Ma, J.; Zheng, J.C.; Lan, J.H.; Fu, J.D.; Chen, M.; Xu, Z.S.; Ma, Y.Z. The ABA-induced soybean ERF transcription factor gene GmERF75 plays a role in enhancing osmotic stress tolerance in Arabidopsis and soybean. BMC Plant Biol. 2019, 19, 506. [Google Scholar] [CrossRef]
- Park, S.I.; Kwon, H.J.; Cho, M.H.; Song, J.S.; Kim, B.G.; Baek, J.; Kim, S.L.; Ji, H.; Kwon, T.R.; Kim, K.H.; et al. The OsERF115/AP2EREBP110 Transcription Factor Is Involved in the Multiple Stress Tolerance to Heat and Drought in Rice Plants. Int. J. Mol. Sci. 2021, 22, 7181. [Google Scholar] [CrossRef]
- Chen, K.; Tang, W.; Zhou, Y.; Chen, J.; Xu, Z.; Ma, R.; Dong, Y.; Ma, Y.; Chen, M. AP2/ERF transcription factor GmDREB1 confers drought tolerance in transgenic soybean by interacting with GmERFs. Plant Physiol. Biochem. 2022, 170, 287–295. [Google Scholar] [CrossRef]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef]
- Xie, Z.; Nolan, T.; Jiang, H.; Tang, B.; Zhang, M.; Li, Z.; Yin, Y. The AP2/ERF Transcription Factor TINY Modulates Brassinosteroid-Regulated Plant Growth and Drought Responses in Arabidopsis. Plant Cell 2019, 31, 1788–1806. [Google Scholar] [CrossRef]
- Sun, S.; Yu, J.P.; Chen, F.; Zhao, T.J.; Fang, X.H.; Li, Y.Q.; Sui, S.F. TINY, a dehydration-responsive element (DRE)-binding protein-like transcription factor connecting the DRE- and ethylene-responsive element-mediated signaling pathways in Arabidopsis. J. Biol. Chem. 2008, 283, 6261–6271. [Google Scholar] [CrossRef]
- Coego, A.; Brizuela, E.; Castillejo, P.; Ruíz, S.; Koncz, C.; del Pozo, J.C.; Piñeiro, M.; Jarillo, J.A.; Paz-Ares, J.; León, J.; et al. The TRANSPLANTA collection of Arabidopsis lines: A resource for functional analysis of transcription factors based on their conditional overexpression. Plant J. 2014, 77, 944–953. [Google Scholar] [CrossRef]
- Wei, G.; Pan, Y.; Lei, J.; Zhu, Y.X. Molecular cloning, phylogenetic analysis, expressional profiling and in vitro studies of TINY2 from Arabidopsis thaliana. J. Biochem. Mol. Biol. 2005, 38, 440–446. [Google Scholar] [CrossRef]
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop Production under Drought and Heat Stress: Plant Responses and Management Options. Front. Plant Sci. 2017, 8, 1147. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, Y.Y.; Zhang, D.; Tang, X.L.; Zheng, L.; Shen, C.; Han, X.; Deng, W.H.; Yin, W.L.; Xia, X.L. PtrWRKY75 overexpression reduces stomatal aperture and improves drought tolerance by salicylic acid-induced reactive oxygen species accumulation in poplar. Environ. Exp. Bot. 2020, 176, 104117. [Google Scholar] [CrossRef]
- He, W.J.; Yan, K.; Zhang, Y.; Han, G.X.; Su, Y.J.; Yang, X.Y.; Sun, L.N.; Wang, F.J.; Wang, X. Contrasting photosynthesis, photoinhibition and oxidative damage in honeysuckle (Lonicera japonica Thunb.) under iso-osmotic salt and drought stresses. Environ. Exp. Bot. 2020, 182, 104313. [Google Scholar] [CrossRef]
- Cheng, X.; He, Q.; Tang, S.; Wang, H.; Zhang, X.; Lv, M.; Liu, H.; Gao, Q.; Zhou, Y.; Wang, Q.; et al. The miR172/IDS1 signaling module confers salt tolerance through maintaining ROS homeostasis in cereal crops. New Phytol. 2021, 230, 1017–1033. [Google Scholar] [CrossRef]
- Székely, G.; Abrahám, E.; Cséplo, A.; Rigó, G.; Zsigmond, L.; Csiszár, J.; Ayaydin, F.; Strizhov, N.; Jásik, J.; Schmelzer, E.; et al. Duplicated P5CS genes of Arabidopsis play distinct roles in stress regulation and developmental control of proline biosynthesis. Plant J. 2008, 53, 11–28. [Google Scholar] [CrossRef]
- Li, X.; Tang, Y.; Li, H.; Luo, W.; Zhou, C.; Zhang, L.; Lv, J. A wheat R2R3 MYB gene TaMpc1-D4 negatively regulates drought tolerance in transgenic Arabidopsis and wheat. Plant Sci. 2020, 299, 110613. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Yu, M.; Zhang, S.; Song, T.; Zhang, M.; Zhou, H.; Wang, Y.; Xiang, J.; Zhang, X. Transcriptomic Identification of Wheat AP2/ERF Transcription Factors and Functional Characterization of TaERF-6-3A in Response to Drought and Salinity Stresses. Int. J. Mol. Sci. 2022, 23, 3272. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Qanmber, G.; Li, J.; Pu, M.; Chen, G.; Li, S.; Liu, L.; Qin, W.; Ma, S.; Wang, Y.; et al. Identification and Characterization of the ERF Subfamily B3 Group Revealed GhERF13.12 Improves Salt Tolerance in Upland Cotton. Front. Plant Sci. 2021, 12, 705883. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.