Water Consumption Pattern Analysis Using Biclustering: When, Why and How
Abstract
1. Introduction
- overview of notorious contributions in the literature contemplating the opportunities and limitations of clustering water time series data;
- taxonomy for a structured view, principled application, and critical assessment of biclustering water consumption data;
- novel methodology for the correct application of coclustering and biclustering methods to water consumption data analysis;
- empirical validation and comprehensive discussion using a real-world case study from a WDS corresponding to a large tourist and residential resort.
Related Work
2. Background
2.1. Time Clustering
2.2. Subspace Clustering
2.2.1. Biclustering
2.2.2. Coclustering and Subspace Clustering Variants
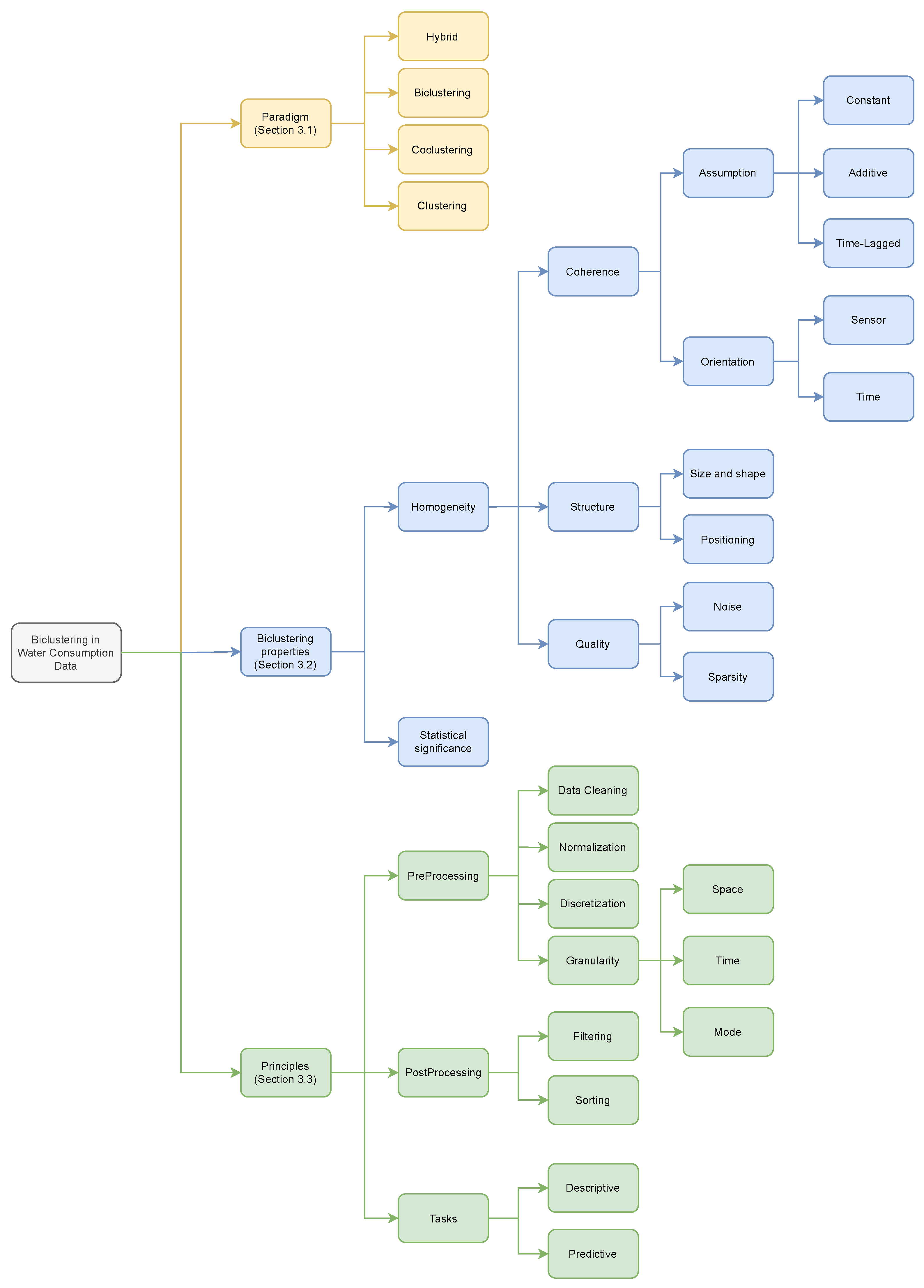
3. Solution: Biclustering for Water Consumption Pattern Mining
- biclustering-based paradigms on water consumption data (Section 3.1);
- biclustering settings (coherence, structure, quality, statistical significance) and their impact (Section 3.2);
- principles for guiding the development of biclustering-based pattern mining on time series water consumption data (Section 3.3).
3.1. Major Subspace-Clustering Paradigms
3.2. Biclustering Properties and Their Impact on the Pattern Mining Water Consumption Data
3.2.1. Biclustering Coherence
3.2.2. Biclustering Structure
3.2.3. Biclustering Quality
3.2.4. Biclustering Statistical Significance
3.3. Principles for Biclustering-Based Time Series Analysis on Water Consumption Data
4. Case Study: Water Distribution Network of Quinta Do Lago
- RQ1. Are clustering approaches adequate for water consumption profiling from time series data? What are their major limitations?
- RQ2. Does coclustering, as a more flexible clustering approach, aid the clustering analysis of water consumption data?
- RQ3. Is biclustering able to retrieve novel actionable water consumption patterns? Can biclustering address the established shortcoming of clustering and co-clustering tasks?
- RQ4. Which principles should be placed on the design and application of biclustering approaches for an effective descriptive and predictive analysis of water consumption profiles?
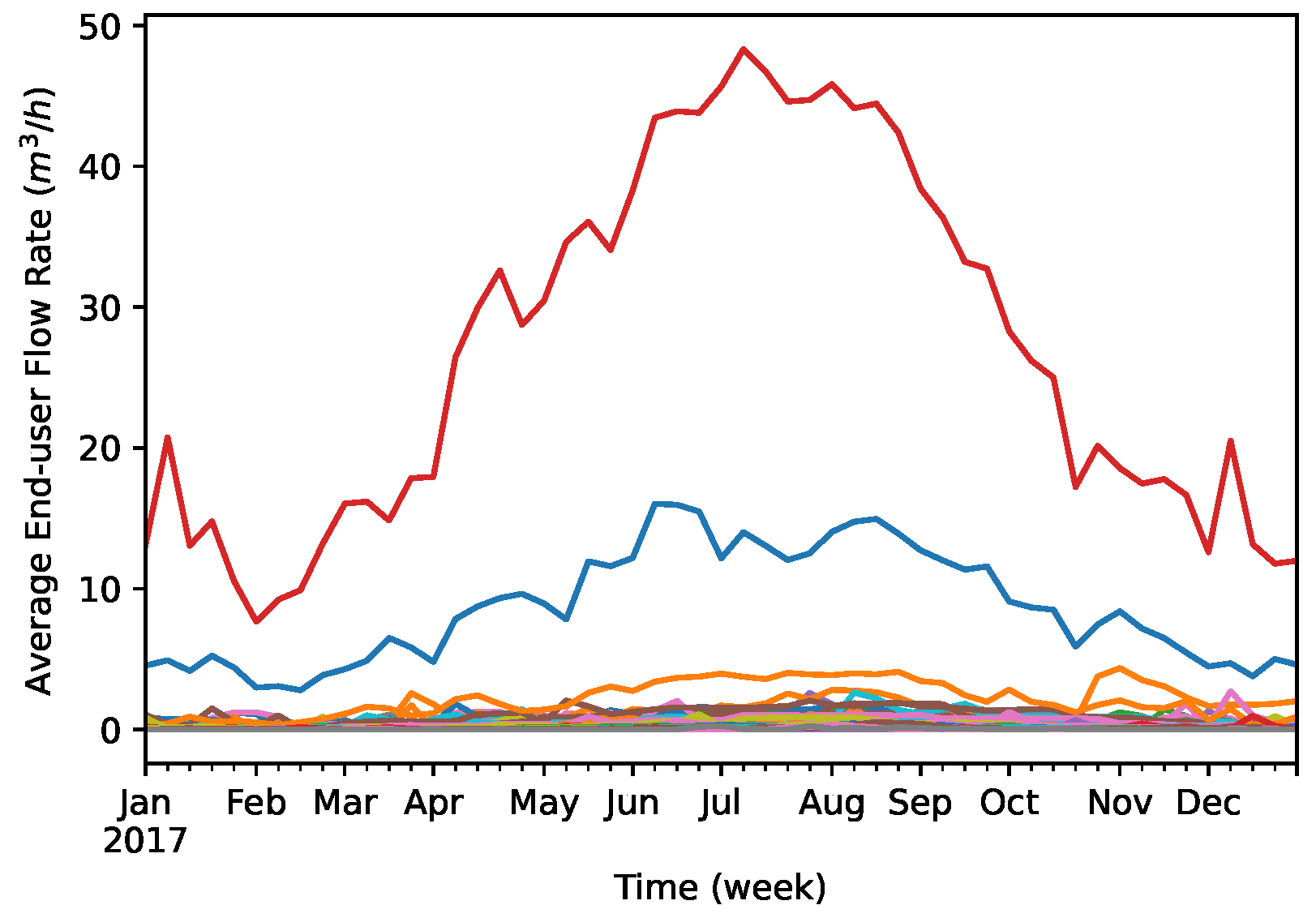
4.1. Dataset
4.2. Experimental Setting
4.3. Data Preprocessing
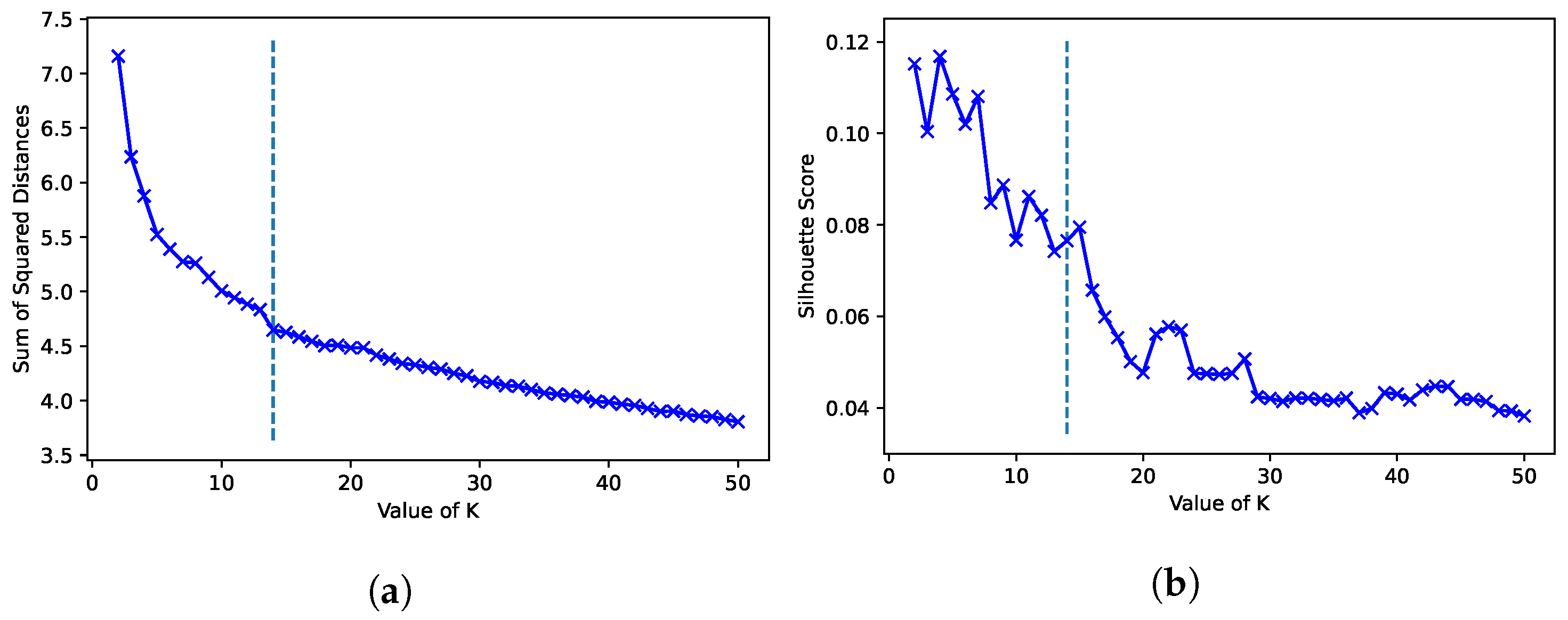
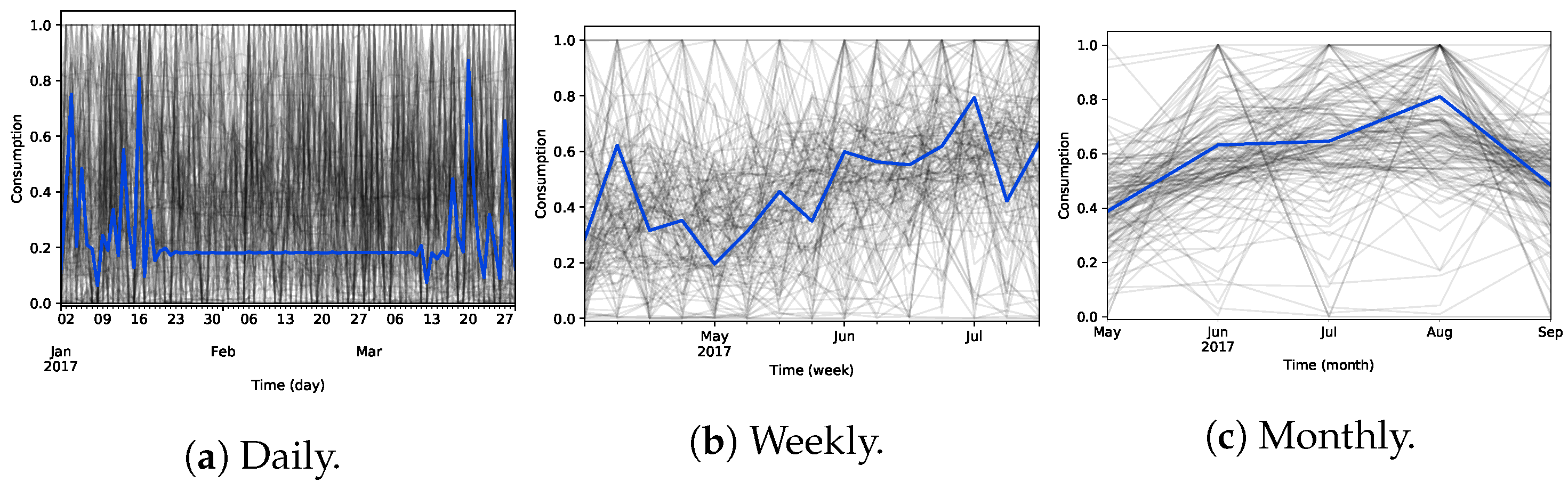
4.4. Clustering Analysis (RQ1)
- Consumption behaviour is grouped across the entire time axis, neglecting local patterns;
- Sensitive to noise and outliers requiring data transformations and cleaning procedures which are frequently not sufficient;
- Method-specific parameterization needs that considerably impact the clustering analysis, e.g., manually specifying the number of clusters in the case of K-means;
- Limited to constant relationships between time series, not considering other meaningful coherent consumption profiles explained by shifting, scaling and lagged factors.
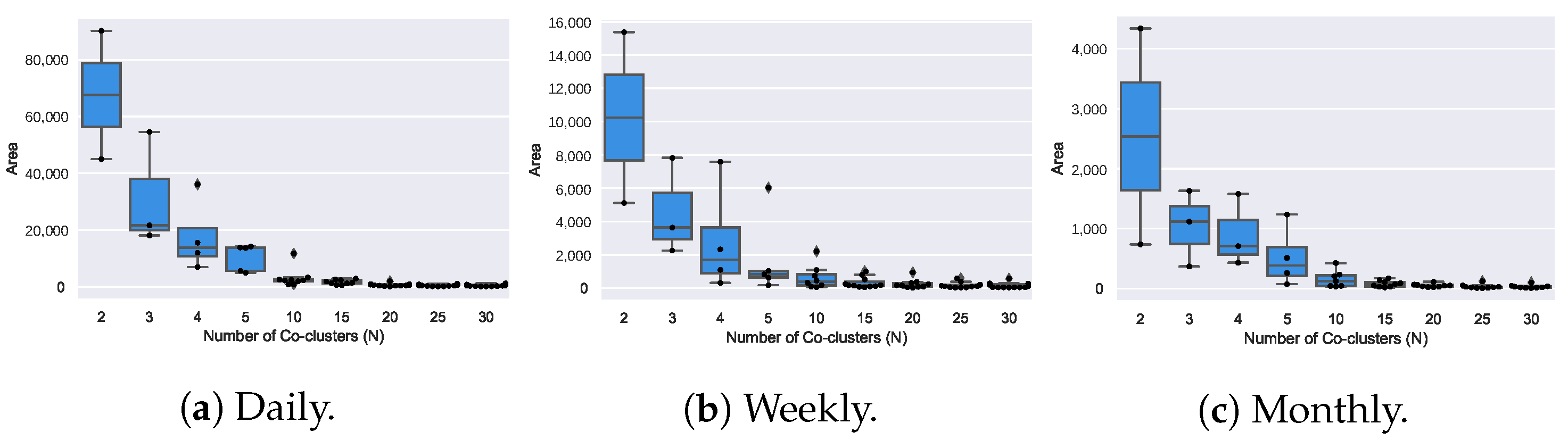
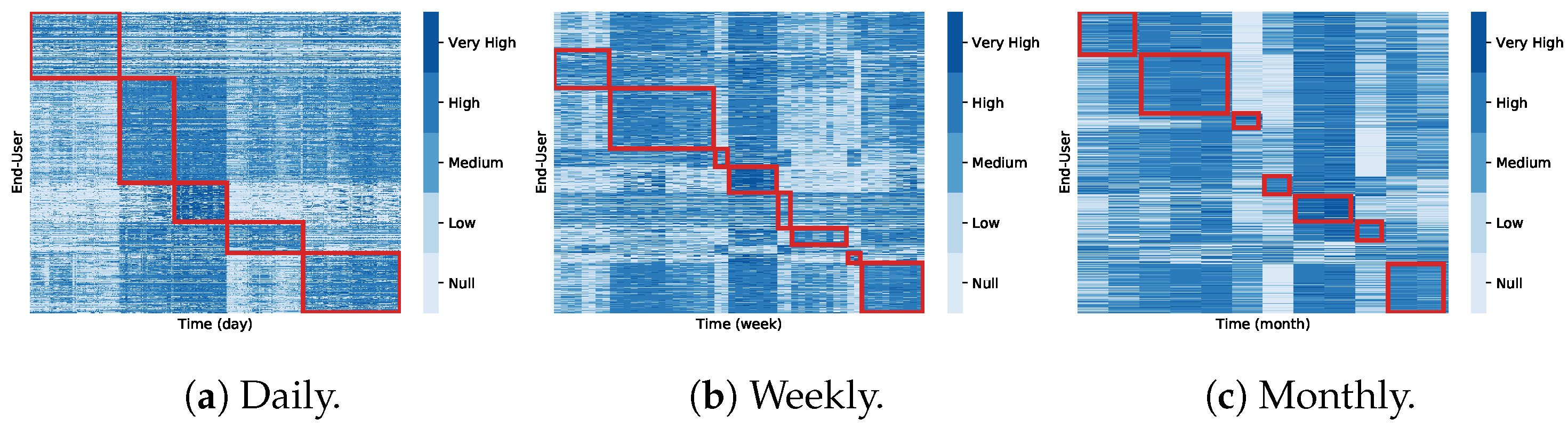
4.5. Coclustering Analysis (RQ2)
- Coclustering approaches generally disregard temporal dependencies within and across consumption signals, thus penalizing misalignments between coherent profiles as well as the inherent consumption variability along time. It further discards temporal contiguity, and as a result, water consumption patterns are generally grouped under non-sequential periods, limiting the interpretability and actionability of the gathered patterns;
- Coclustering guarantees the discovery of subspaces that can be evaluated according to a homogeneity measure, meaning that coclusters with low homogeneity can be filtered before analysis. Nevertheless, there is the need to manually specify the number of coclusters;
- Coclustering can discover groups of users with coherent consumption behavior under some periods, not limiting the search for global consumption patterns. However, coclustering assumes that each user is only associated with one consumption pattern, disregarding the possibility of associating multiple patterns with an user’s consumption profile. In addition, the partitioning of the time axis is restricting, preventing the discovery of flexibly positioned subspaces with arbitrarily-high overlaps along the time dimensions.
4.6. Biclustering Analysis (RQ3)
4.7. Guiding Biclustering Principles for Water Consumption Tasks (RQ4)
- Detection of local consumption profiles, surpassing the limitation of traditional time clustering methods that only unveil global patterns;
- Efficient search for patterns with multiple coherence assumptions and quality, instead of only assuming constant relationships between time series;
- Retrieval of well-defined consumption patterns with solid guarantees of coherence and quality, in contrast with high variability of clustering consumption profiles;
- Flexible pattern-based search that can be customized to guide and restrict the search, preventing redundant consumption patterns and ensuring efficient searches.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| WDS | Water distribution system |
| WDN | Water distribution network |
| SVM | Support vector machine |
| SOM | Self-organizing map |
| HAC | Hierarchical agglomerative clustering |
| DTW | Dynamic time warping |
| DBA | Dynamic time warping barycenter averaging |
| DWT | Discrete wavelet transform |
| PAA | Piecewise aggregate approximation |
| PLA | Piecewise linear approximation |
| SAX | Symbolic aggregate approximation |
| LCSS | Longest common sub-sequence |
| MODH | Modified hausdorff |
| HMM | Hidden markov model |
| SSE | Sum of squared error |
| CD | Distance between clusters index |
| IQR | Interquartile range |
| CCC | Contiguous column coherent biclustering |
References
- Cominola, A.; Giuliani, M.; Piga, D.; Castelletti, A.; Rizzoli, A.E. Benefits and challenges of using smart meters for advancing residential water demand modeling and management: A review. Environ. Model. Softw. 2015, 72, 198–214. [Google Scholar] [CrossRef]
- Flath, C.; Nicolay, D.; Conte, T.; van Dinther, C.; Filipova-Neumann, L. Cluster Analysis of Smart Metering Data—An Implementation in Practice. Bus. Inf. Syst. Eng. 2012, 4, 31–39. [Google Scholar] [CrossRef]
- Sønderlund, A.L.; Smith, J.R.; Hutton, C.J.; Kapelan, Z.; Savic, D. Effectiveness of smart meter-based consumption feedback in curbing household water use: Knowns and unknowns. J. Water Resour. Plan. Manag. 2016, 142, 04016060. [Google Scholar] [CrossRef]
- Gurung, T.R.; Stewart, R.A.; Beal, C.D.; Sharma, A.K. Smart meter enabled water end-use demand data: Platform for the enhanced infrastructure planning of contemporary urban water supply networks. J. Clean. Prod. 2015, 87, 642–654. [Google Scholar] [CrossRef]
- Loureiro, D.; Alegre, H.; Coelho, S.; Martins, A.; Mamade, A. A new approach to improve water loss control using smart metering data. Water Sci. Technol. Water Supply 2014, 14, 618–625. [Google Scholar] [CrossRef]
- Laspidou, C.; Papageorgiou, E.; Kokkinos, K.; Sahu, S.; Gupta, A.; Tassiulas, L. Exploring patterns in water consumption by clustering. Procedia Eng. 2015, 119, 1439–1446. [Google Scholar] [CrossRef]
- Cheifetz, N.; Noumir, Z.; Samé, A.; Sandraz, A.C.; Féliers, C.; Heim, V. Modeling and clustering water demand patterns from real-world smart meter data. Drink. Water Eng. Sci. 2017, 10, 75–82. [Google Scholar] [CrossRef][Green Version]
- Ioannou, A.E.; Creaco, E.F.; Laspidou, C.S. Exploring the Effectiveness of Clustering Algorithms for Capturing Water Consumption Behavior at Household Level. Sustainability 2021, 13, 2603. [Google Scholar] [CrossRef]
- Candelieri, A. Clustering and support vector regression for water demand forecasting and anomaly detection. Water 2017, 9, 224. [Google Scholar] [CrossRef]
- Yang, A.; Zhang, H.; Stewart, R.A.; Nguyen, K. Enhancing residential water end use pattern recognition accuracy using self-organizing maps and K-means clustering techniques: Autoflow v3.1. Water 2018, 10, 1221. [Google Scholar] [CrossRef]
- Sim, K.; Gopalkrishnan, V.; Zimek, A.; Cong, G. A survey on enhanced subspace clustering. Data Min. Knowl. Discov. 2013, 26, 332–397. [Google Scholar] [CrossRef]
- Madeira, S.C.; Oliveira, A.L. Biclustering Algorithms for Biological Data Analysis: A Survey. IEEE ACM Trans. Comput. Biol. Bioinform. 2004, 1, 24–45. [Google Scholar] [CrossRef] [PubMed]
- Bougadis, J.; Adamowski, K.; Diduch, R. Short-term municipal water demand forecasting. Hydrol. Process. Int. J. 2005, 19, 137–148. [Google Scholar] [CrossRef]
- Alvisi, S.; Franchini, M.; Marinelli, A. A short-term, pattern-based model for water-demand forecasting. J. Hydroinformat. 2007, 9, 39–50. [Google Scholar] [CrossRef]
- Donkor, E.A.; Mazzuchi, T.A.; Soyer, R.; Alan Roberson, J. Urban water demand forecasting: Review of methods and models. J. Water Resour. Plan. Manag. 2014, 140, 146–159. [Google Scholar] [CrossRef]
- Brentan, B.M.; Luvizotto, E., Jr.; Herrera, M.; Izquierdo, J.; Pérez-García, R. Hybrid regression model for near real-time urban water demand forecasting. J. Comput. Appl. Math. 2017, 309, 532–541. [Google Scholar] [CrossRef]
- Divina, F.; Goméz Vela, F.A.; García Torres, M. Biclustering of smart building electric energy consumption data. Appl. Sci. 2019, 9, 222. [Google Scholar] [CrossRef]
- Divina, F.; Aguilar-Ruiz, J.S. A multi-objective approach to discover biclusters in microarray data. In Proceedings of the Genetic and Evolutionary Computation Conference, GECCO 2007, London, UK, 7–11 July 2007; Lipson, H., Ed.; ACM: New York, NY, USA, 2007; pp. 385–392. [Google Scholar] [CrossRef]
- Han, J.; Kamber, M.; Pei, J. Data Mining: Concepts and Techniques, 3rd ed.; Morgan Kaufmann: Burlington, MA, USA, 2011. [Google Scholar]
- Ernst, J.; Nau, G.J.; Bar-Joseph, Z. Clustering short time series gene expression data. In Proceedings of the Thirteenth International Conference on Intelligent Systems for Molecular Biology 2005, Detroit, MI, USA, 25–29 June 2005; pp. 159–168. [Google Scholar] [CrossRef]
- Fu, T.C.; Chung, F.L.; Ng, V.; Luk, R. Pattern discovery from stock time series using self-organizing maps. In Workshop Notes of KDD2001 Workshop on Temporal Data Mining; Springer: New York, NY, USA, 2001; Volume 1. [Google Scholar]
- Ruiz, L.G.B.; del Carmen Pegalajar Jiménez, M.; Arcucci, R.; Molina-Solana, M. A time-series clustering methodology for knowledge extraction in energy consumption data. Expert Syst. Appl. 2020, 160, 113731. [Google Scholar] [CrossRef]
- Saas, A.; Guitart, A.; Perianez, A. Discovering playing patterns: Time series clustering of free-to-play game data. In Proceedings of the IEEE Conference on Computational Intelligence and Games, CIG 2016, Santorini, Greece, 20–23 September 2016; pp. 1–8. [Google Scholar] [CrossRef]
- Aghabozorgi, S.R.; Shirkhorshidi, A.S.; Teh, Y.W. Time-series clustering—A decade review. Inf. Syst. 2015, 53, 16–38. [Google Scholar] [CrossRef]
- Liao, T.W. Clustering of time series data—A survey. Pattern Recognit. 2005, 38, 1857–1874. [Google Scholar] [CrossRef]
- Sakoe, H.; Chiba, S. Dynamic programming algorithm optimization for spoken word recognition. IEEE Trans. Acoust. Speech, Signal Process. 1978, 26, 43–49. [Google Scholar] [CrossRef]
- Hautamäki, V.; Nykänen, P.; Fränti, P. Time-series clustering by approximate prototypes. In Proceedings of the 19th International Conference on Pattern Recognition (ICPR 2008), Tampa, FL, USA, 8–11 December 2008; IEEE Computer Society: Washington, DC, USA, 2008; pp. 1–4. [Google Scholar] [CrossRef]
- Keogh, E.J.; Lonardi, S.; Ratanamahatana, C.A. Towards parameter-free data mining. In Proceedings of the Tenth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Seattle, WA, USA, 22–25 August 2004; Kim, W., Kohavi, R., Gehrke, J., DuMouchel, W., Eds.; ACM: New York, NY, USA, 2004; pp. 206–215. [Google Scholar] [CrossRef]
- Petitjean, F.; Ketterlin, A.; Gançarski, P. A global averaging method for dynamic time warping, with applications to clustering. Pattern Recognit. 2011, 44, 678–693. [Google Scholar] [CrossRef]
- Henriques, R.; Antunes, C.; Madeira, S.C. A structured view on pattern mining-based biclustering. Pattern Recognit. 2015, 48, 3941–3958. [Google Scholar] [CrossRef]
- Zhang, Y.; Zha, H.; Chu, C. A Time-Series Biclustering Algorithm for Revealing Co-Regulated Genes. In Proceedings of the International Symposium on Information Technology: Coding and Computing (ITCC 2005), Las Vegas, NA, USA, 4–6 April 2005; IEEE Computer Society: Washington, DC, USA, 2005; Volume 1, pp. 32–37. [Google Scholar] [CrossRef]
- Madeira, S.C.; Oliveira, A.L. A Linear Time Biclustering Algorithm for Time Series Gene Expression Data. In Proceedings of the Lecture Notes in Computer Science, Algorithms in Bioinformatics, 5th International Workshop, WABI 2005, Mallorca, Spain, 3–6 October 2005; Casadio, R., Myers, G., Eds.; Springer: Berlin/Heidelberg, Germany, 2005; Volume 3692. [Google Scholar] [CrossRef]
- Madeira, S.C.; Oliveira, A.L. A polynomial time biclustering algorithm for finding approximate expression patterns in gene expression time series. Algorithms Mol. Biol. 2009, 4, 8. [Google Scholar] [CrossRef] [PubMed]
- Madeira, S.C.; Teixeira, M.C.; Sá-Correia, I.; Oliveira, A.L. Identification of Regulatory Modules in Time Series Gene Expression Data Using a Linear Time Biclustering Algorithm. IEEE ACM Trans. Comput. Biol. Bioinform. 2010, 7, 153–165. [Google Scholar] [CrossRef]
- Gonçalves, J.P.; Madeira, S.C.; Oliveira, A.L. BiGGEsTS: Integrated environment for biclustering analysis of time series gene expression data. BMC Res. Notes 2009, 2, 1–11. [Google Scholar] [CrossRef]
- Xue, Y.; Liao, Z.; Li, M.; Luo, J.; Hu, X.; Luo, G.; Chen, W. A New Biclustering Algorithm for Time-Series Gene Expression Data Analysis. In Proceedings of the Tenth International Conference on Computational Intelligence and Security, CIS 2014, Kunming, China, 15–16 November 2014; IEEE Computer Society: Washington, DC, USA, 2014; pp. 268–272. [Google Scholar] [CrossRef]
- Denitto, M.; Farinelli, A.; Bicego, M. Biclustering of time series data using factor graphs. In Proceedings of the Symposium on Applied Computing, SAC 2017, Marrakech, Morocco, 3–7 April 2017; Seffah, A., Penzenstadler, B., Alves, C., Peng, X., Eds.; ACM: New York, NY, USA, 2017; pp. 28–30. [Google Scholar] [CrossRef]
- Lee, J.H.; Lee, Y.R.; Jun, C.H. A biclustering method for time series analysis. Ind. Eng. Manag. Syst. 2010, 9, 131–140. [Google Scholar] [CrossRef]
- Ji, L.; Tan, K.L. Identifying time-lagged gene clusters using gene expression data. Bioinformatics 2005, 21, 509–516. [Google Scholar] [CrossRef]
- Gonçalves, J.P.; Madeira, S.C. LateBiclustering: Efficient Heuristic Algorithm for Time-Lagged Bicluster Identification. IEEE ACM Trans. Comput. Biol. Bioinform. 2014, 11, 801–813. [Google Scholar] [CrossRef]
- Henriques, R.; Madeira, S.C. BSig: Evaluating the statistical significance of biclustering solutions. Data Min. Knowl. Discov. 2018, 32, 124–161. [Google Scholar] [CrossRef]
- Henriques, R.; Madeira, S.C. BicPAM: Pattern-based biclustering for biomedical data analysis. Algorithms Mol. Biol. 2014, 9, 27. [Google Scholar] [CrossRef] [PubMed]
- Peeters, R. The maximum edge biclique problem is NP-complete. Discret. Appl. Math. 2003, 131, 651–654. [Google Scholar] [CrossRef]
- Horta, D.; Campello, R.J.G.B. Similarity Measures for Comparing Biclusterings. IEEE ACM Trans. Comput. Biol. Bioinform. 2014, 11, 942–954. [Google Scholar] [CrossRef] [PubMed]
- Tanay, A.; Sharan, R.; Shamir, R. Discovering statistically significant biclusters in gene expression data. In Proceedings of the Tenth International Conference on Intelligent Systems for Molecular Biology, Edmonton, AB, Canada, 3–7 August 2002; pp. 136–144. [Google Scholar]
- Gupta, N.; Aggarwal, S. MIB: Using mutual information for biclustering gene expression data. Pattern Recognit. 2010, 43, 2692–2697. [Google Scholar] [CrossRef]
- Murali, T.M.; Kasif, S. Extracting Conserved Gene Expression Motifs from Gene Expression Data. In Proceedings of the 8th Pacific Symposium on Biocomputing, PSB 2003, Lihue, HI, USA, 3–7 January 2003; Altman, R.B., Dunker, A.K., Hunter, L., Klein, T.E., Eds.; World Scientific: Toh Tuck Link, Singapore, 2003; pp. 77–88. [Google Scholar]
- Yang, J.; Wang, H.; Wang, W.; Yu, P.S. Enhanced Biclustering on Expression Data. In Proceedings of the 3rd IEEE International Symposium on BioInformatics and BioEngineering (BIBE 2003), Bethesda, MD, USA, 10–12 March 2003; IEEE Computer Society: Washington, DC, USA, 2003; pp. 321–327. [Google Scholar] [CrossRef]
- Dhillon, I.S. Co-clustering documents and words using bipartite spectral graph partitioning. In Proceedings of the seventh ACM SIGKDD international conference on Knowledge discovery and data mining, San Francisco, CA, USA, 26–29 August 2001; Lee, D., Schkolnick, M., Provost, F.J., Srikant, R., Eds.; ACM: New York, NY, USA, 2001; pp. 269–274. [Google Scholar] [CrossRef]
- Alqadah, F.; Reddy, C.K.; Hu, J.; Alqadah, H.F. Biclustering neighborhood-based collaborative filtering method for top-n recommender systems. Knowl. Inf. Syst. 2015, 44, 475–491. [Google Scholar] [CrossRef]
- Dolnicar, S.; Kaiser, S.; Lazarevski, K.; Leisch, F. Biclustering: Overcoming data dimensionality problems in market segmentation. J. Travel Res. 2012, 51, 41–49. [Google Scholar] [CrossRef]
- Izenman, A.J.; Harris, P.W.; Mennis, J.; Jupin, J.; Obradovic, Z. Local spatial biclustering and prediction of urban juvenile delinquency and recidivism. Stat. Anal. Data Mining Asa Data Sci. J. 2011, 4, 259–275. [Google Scholar] [CrossRef]
- Dhamodharavadhani, S.; Rathipriya, R. Biclustering Analysis of Countries Using COVID-19 Epidemiological Data. In Internet of Things; Springer: Berlin/Heidelberg, Germany, 2021; pp. 93–114. [Google Scholar]
- Kluger, Y.; Basri, R.; Chang, J.T.; Gerstein, M. Spectral biclustering of microarray data: Coclustering genes and conditions. Genome Res. 2003, 13, 703–716. [Google Scholar] [CrossRef]
- Dhillon, I.S.; Mallela, S.; Modha, D.S. Information-theoretic co-clustering. In Proceedings of the Ninth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Washington, DC, USA, 24–27 August 2003; Getoor, L., Senator, T.E., Domingos, P.M., Faloutsos, C., Eds.; ACM: New York, NY, USA, 2003; pp. 89–98. [Google Scholar] [CrossRef]
- Henriques, R.; Madeira, S.C. Triclustering Algorithms for Three-Dimensional Data Analysis: A Comprehensive Survey. ACM Comput. Surv. 2019, 51, 1–43. [Google Scholar] [CrossRef]
- Moritz, S.; Bartz-Beielstein, T. imputeTS: Time Series Missing Value Imputation in R. R J. 2017, 9, 207. [Google Scholar] [CrossRef]
- Henriques, R.; Madeira, S.C. FleBiC: Learning classifiers from high-dimensional biomedical data using discriminative biclusters with non-constant patterns. Pattern Recognit. 2021, 115, 107900. [Google Scholar] [CrossRef]
- Soares, D.; Henriques, R.; Gromicho, M.; Pinto, S.; Carvalho, M.d.; Madeira, S.C. Towards triclustering-based classification of three-way clinical data: A case study on predicting non-invasive ventilation in als. In Proceedings of the International Conference on Practical Applications of Computational Biology & Bioinformatics, L´Aquila, Italy, 17–19 June 2020; Springer: Berlin/Heidelberg, Germany, 2020; pp. 112–122. [Google Scholar]
- Soares, D.F.; Henriques, R.; Gromicho, M.; de Carvalho, M.; C Madeira, S. Prognostic Prediction in ALS: Triclustering-Based Classification of Longitudinal Data Targeting Relevant Clinical Endpoints. Available online: https://ssrn.com/abstract=4102493 (accessed on 11 May 2022).
- Gomes, S.C.; Vinga, S.; Henriques, R. Spatiotemporal Correlation Feature Spaces to Support Anomaly Detection in Water Distribution Networks. Water 2021, 13, 2551. [Google Scholar] [CrossRef]
- Castanho, E.N.; Aidos, H.; Madeira, S.C. Biclustering fMRI time series: A comparative study. BMC Bioinform. 2022, 23, 192. [Google Scholar] [CrossRef] [PubMed]
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; Haberland, M.; Reddy, T.; Cournapeau, D.; Burovski, E.; Peterson, P.; Weckesser, W.; Bright, J.; et al. SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python. Nat. Methods 2020, 17, 261–272. [Google Scholar] [CrossRef] [PubMed]
- Tavenard, R.; Faouzi, J.; Vandewiele, G.; Divo, F.; Androz, G.; Holtz, C.; Payne, M.; Yurchak, R.; Rußwurm, M.; Kolar, K.; et al. Tslearn, A Machine Learning Toolkit for Time Series Data. J. Mach. Learn. Res. 2020, 21, 1–6. [Google Scholar]
- Keogh, E.J.; Pazzani, M.J. Scaling up dynamic time warping for datamining applications. In Proceedings of the sixth ACM SIGKDD international Conference on Knowledge Discovery and Data Mining, Boston, MA, USA, 20–23 August 2000; Ramakrishnan, R., Stolfo, S.J., Bayardo, R.J., Parsa, I., Eds.; ACM: New York, NY, USA, 2000; pp. 285–289. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- Satopaa, V.; Albrecht, J.R.; Irwin, D.E.; Raghavan, B. Finding a “Kneedle” in a Haystack: Detecting Knee Points in System Behavior. In Proceedings of the 31st IEEE International Conference on Distributed Computing Systems Workshops (ICDCS 2011 Workshops), Minneapolis, MN, USA, 20–24 June 2011; IEEE Computer Society: Washington, DC, USA, 2011; pp. 166–171. [Google Scholar] [CrossRef]
- Divina, F.; Pontes, B.; Giráldez, R.; Aguilar-Ruiz, J.S. An effective measure for assessing the quality of biclusters. Comput. Biol. Med. 2012, 42, 245–256. [Google Scholar] [CrossRef]
- Henriques, R.; Madeira, S.C. BicNET: Flexible module discovery in large-scale biological networks using biclustering. Algorithms Mol. Biol. 2016, 11, 1–30. [Google Scholar] [CrossRef]
- de França, F.O.; Coelho, G.P.; Zuben, F.J.V. Predicting missing values with biclustering: A coherence-based approach. Pattern Recognit. 2013, 46, 1255–1266. [Google Scholar] [CrossRef]
- Agrawal, R.; Gehrke, J.; Gunopulos, D.; Raghavan, P. Automatic Subspace Clustering of High Dimensional Data for Data Mining Applications. In Proceedings of the SIGMOD 1998, ACM SIGMOD International Conference on Management of Data, Seattle, WA, USA, 2–4 June 1998; Haas, L.M., Tiwary, A., Eds.; ACM Press: New York, NY, USA, 1998; pp. 94–105. [Google Scholar] [CrossRef]
- Singh, M.; Mehrotra, M. Impact of biclustering on the performance of Biclustering based Collaborative Filtering. Expert Syst. Appl. 2018, 113, 443–456. [Google Scholar] [CrossRef]






















| Dataset | ID | #Users | #Time Points (First, Last) |
|---|---|---|---|
| Daily | 0 | 161 | 88 (0, 87) |
| Weekly | 2 | 147 | 15 (14, 28) |
| Monthly | 1 | 142 | 3 (4, 8) |
| Solution | Post-Processed | ||||||
|---|---|---|---|---|---|---|---|
| Dataset | (min #Users, min #Time Points) | #bics | #bics | p-Value < 0.05 | p-Value < | ||
| Daily | (20, 7) | 18,666 | 65.2 ± 43.5 | 26.0 ± 20.7 | 655 | 655 | 655 |
| Weekly | (20, 4) | 1310 | 69.8 ± 65.9 | 9.4 ± 5.8 | 263 | 168 | 133 |
| Monthly | (20, 3) | 221 | 50.7 ± 51.5 | 4.5 ± 1.5 | 94 | 23 | 10 |
| Dataset | ID | #Users | #Time Points (First, Last) | p-Value |
|---|---|---|---|---|
| Daily | 9964 | 20 | 7 (331, 337) | 0.0029 |
| Weekly | 245 | 27 | 13 (31, 43) | 1.09 × 10 |
| Monthly | 210 | 21 | 9 (1, 9) | 5.17 × 10 |
| Solution | Post-Processed | ||||||
|---|---|---|---|---|---|---|---|
| Dataset | (min #Users, min #Time Points) | #bics | #bics | p-Value < 0.05 | p-Value 1 × 10 | ||
| Daily | (20, 7) | 786,232 | 72.3 ± 44.8 | 28.9 ± 21.7 | 2347 | 839 | 744 |
| Weekly | (20, 4) | 55,073 | 70.1 ± 63.3 | 11.7 ± 6.6 | 4304 | 279 | 160 |
| Monthly | (20, 3) | 6441 | 57.5 ± 59.8 | 5.6 ± 1.8 | 942 | 18 | 6 |
| Dataset | ID | #Users | #Time Points (First, Last) | p-Value |
|---|---|---|---|---|
| Daily | 197,684 | 20 | 65 (206, 270) | 2.86 × 10 |
| Weekly | 33,405 | 47 | 25 (19, 43) | 1.83 × 10 |
| Monthly | 412 | 22 | 10 (1, 10) | 8.40 × 10 |
| Solution | Post-Processed | |||||||
|---|---|---|---|---|---|---|---|---|
| Dataset | (min #Users, min #Time Points) | L-Shift | #bics | #bics | p-Value < 0.05 | p-Value < 10 | ||
| Daily | (20, 7) | 1 | 38,933 | 87.0 ± 51.8 | 23.6 ± 16.4 | 625 | 593 | 588 |
| 2 | 46,308 | 111.4 ± 64.6 | 25.4 ± 17.1 | 383 | 340 | 330 | ||
| 3 | 32,669 | 124.6 ± 63.8 | 29.6 ± 21.9 | 367 | 332 | 323 | ||
| 4 | 16,033 | 114.8 ± 66.6 | 37.6 ± 26.5 | 345 | 310 | 301 | ||
| Weekly | (20, 4) | 1 | 2828 | 76.7 ± 74.0 | 8.2 ± 4.8 | 404 | 193 | 178 |
| 2 | 2743 | 96.2 ± 90.6 | 8.3 ± 5.0 | 369 | 153 | 131 | ||
| 3 | 1677 | 109.4 ± 97.5 | 10.0 ± 6.7 | 360 | 144 | 123 | ||
| 4 | 1391 | 84.5 ± 82.13 | 10.9 ± 7.0 | 357 | 141 | 121 | ||
| Monthly | (20, 3) | 1 | 372 | 56.3 ± 57.0 | 4.1 ± 1.4 | 113 | 33 | 21 |
| 2 | 318 | 65.2 ± 69.4 | 4.2 ± 1.4 | 108 | 27 | 18 | ||
| 3 | 251 | 60.3 ± 65.7 | 4.5 ± 1.5 | 108 | 30 | 21 | ||
| 4 | 245 | 55.4 ± 55.8 | 4.5 ± 1.6 | 108 | 30 | 21 | ||
| Dataset | ID | #Users | #Time Points (First, Last) | p-Value |
|---|---|---|---|---|
| Daily | 141 | 23 | 8 (358, 364) | 0.002 |
| Weekly | 478 | 26 | 14 (30, 43) | 2.27 × 10 |
| Monthly | 239 | 21 | 9 (1, 9) | 5.17× 10 |
| Solution | Post-Processed | ||||||
|---|---|---|---|---|---|---|---|
| Dataset | (min #Users, min #TimePoints) | #bics | #bics | p-Value < 0.05 | p-Value < 10 | ||
| Daily | (20, 7) | 15,844 | 56.1 ± 51.3 | 26.5 ± 21.8 | 1471 | 1471 | 1471 |
| Weekly | (20, 4) | 1738 | 60.6 ± 61.9 | 10.2 ± 6.3 | 393 | 393 | 393 |
| Monthly | (20, 3) | 243 | 61.3 ± 66.2 | 4.8 ± 1.6 | 99 | 98 | 98 |
| Dataset | ID | #Users | #Time Points (First, Last) | p-Value |
|---|---|---|---|---|
| Daily | 8476 | 32 | 112 | 0 * |
| Weekly | 965 | 44 | 29 | 1.26× 10 |
| Monthly | 120 | 25 | 9 | 1.65 × 10−9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, M.G.; Madeira, S.C.; Henriques, R. Water Consumption Pattern Analysis Using Biclustering: When, Why and How. Water 2022, 14, 1954. https://doi.org/10.3390/w14121954
Silva MG, Madeira SC, Henriques R. Water Consumption Pattern Analysis Using Biclustering: When, Why and How. Water. 2022; 14(12):1954. https://doi.org/10.3390/w14121954
Chicago/Turabian StyleSilva, Miguel G., Sara C. Madeira, and Rui Henriques. 2022. "Water Consumption Pattern Analysis Using Biclustering: When, Why and How" Water 14, no. 12: 1954. https://doi.org/10.3390/w14121954
APA StyleSilva, M. G., Madeira, S. C., & Henriques, R. (2022). Water Consumption Pattern Analysis Using Biclustering: When, Why and How. Water, 14(12), 1954. https://doi.org/10.3390/w14121954






