Identification and Expression of SAUR Genes in the CAM Plant Agave
Abstract
1. Introduction
2. Materials and Methods
2.1. Sequence Retrieval and Subcellular Localization
2.2. Phylogenetic Analysis
2.3. Plant Materials and RNA Extraction
2.4. Expression Analysis
3. Results
3.1. Identification and Subcellular Localization of Agave SAUR Genes
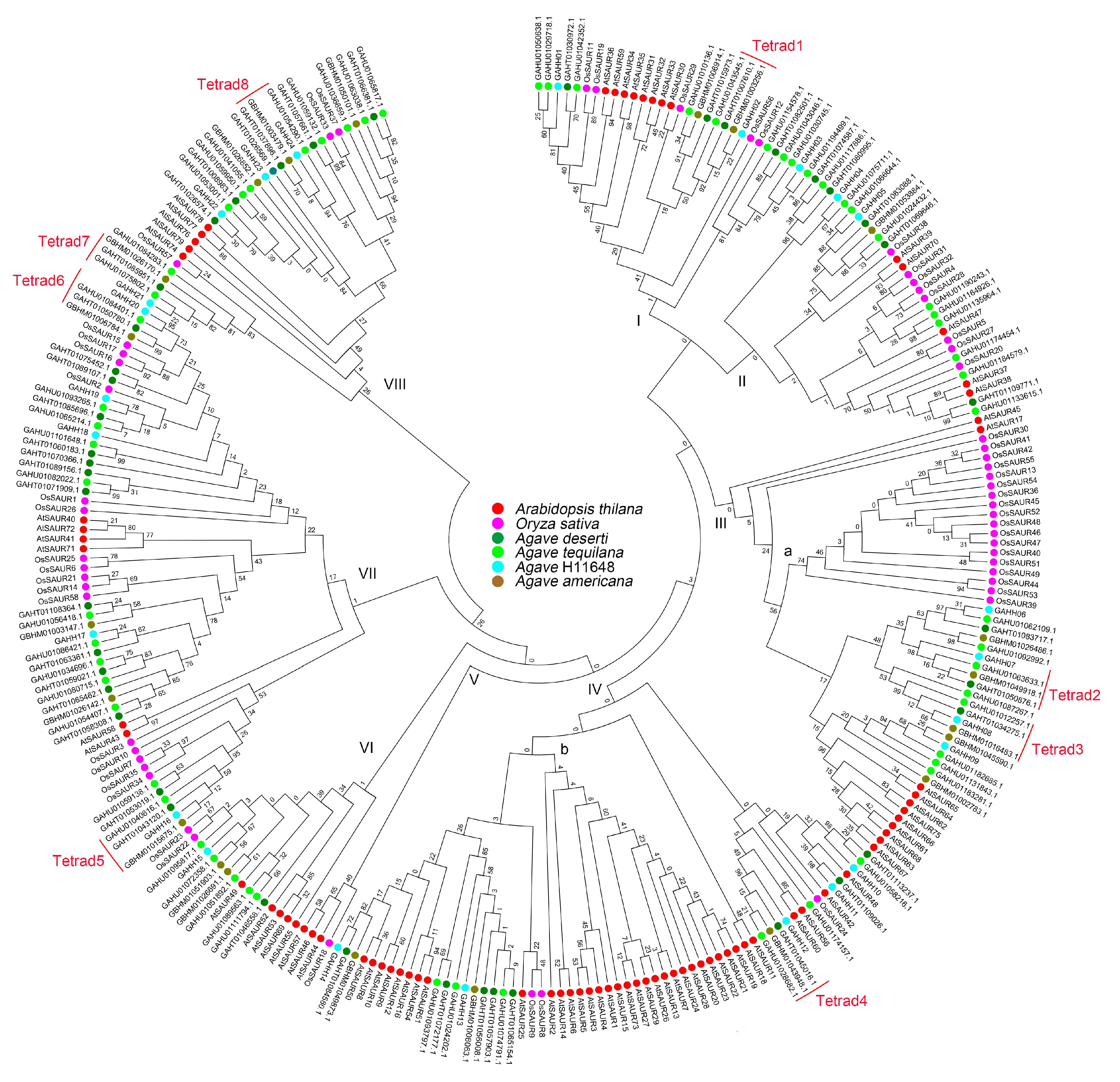
3.2. Phylogenetic Analysis of Agave SAUR Genes
3.3. In Silico Expression of SAUR Genes in Agave
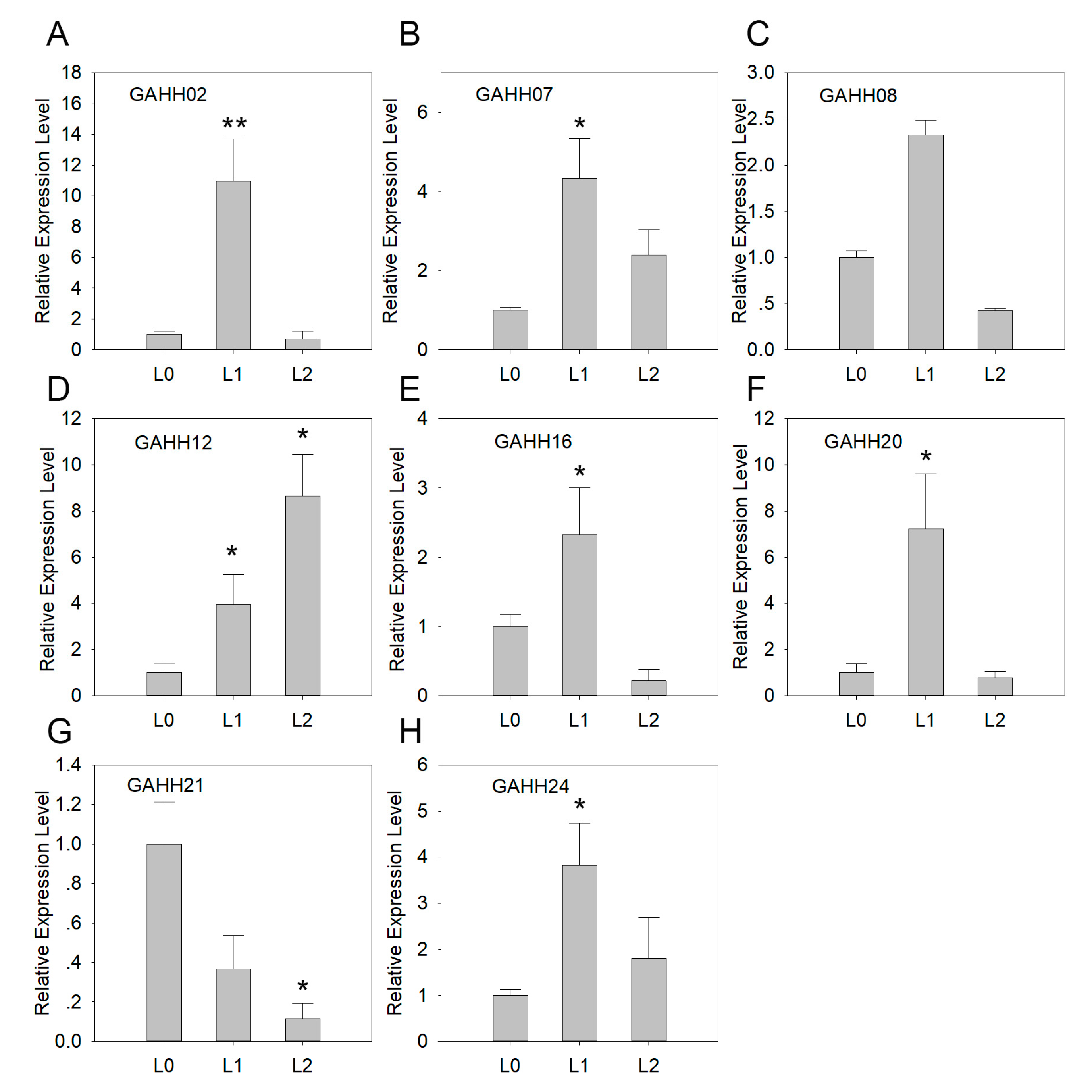
3.4. Expression of SAUR Genes in Agave during Leaf Development
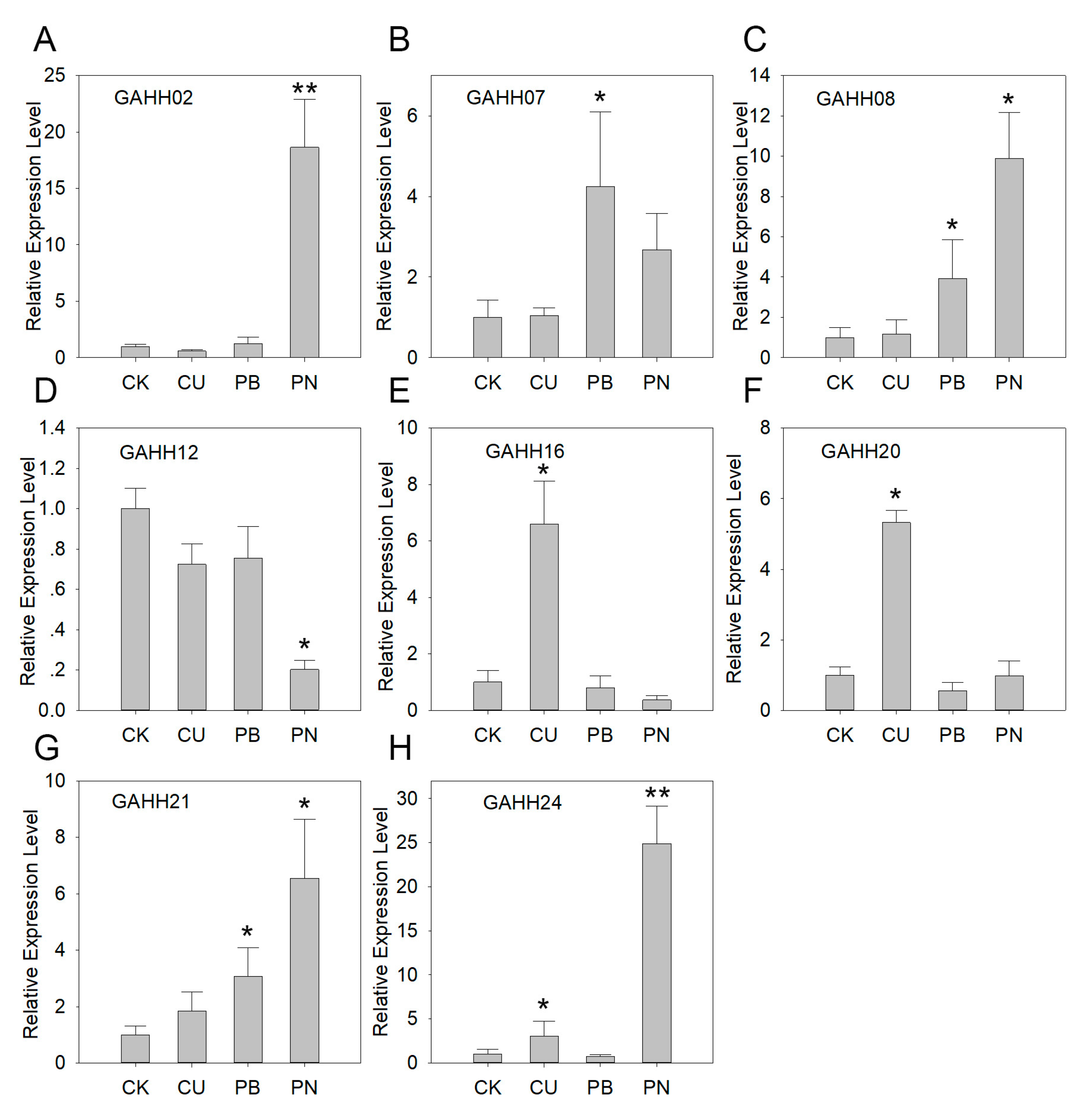
3.5. Expression of Agave SAUR Genes under Abiotic and Biotic Stresses
4. Discussion
4.1. Identification and Evolution of Agave SAUR Genes
4.2. Candidate SAUR Genes Involved in Agave Leaf Development and Stress Response
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Weijers, D.; Friml, J. SnapShot: Auxin signaling and transport. Cell 2009, 136, 1172. [Google Scholar] [CrossRef] [PubMed]
- McClure, B.; Guilfoyle, T. Characterization of a class of small auxin-inducible soybean polyadenylated RNAs. Plant Mol. Biol. 1987, 9, 611–623. [Google Scholar] [CrossRef] [PubMed]
- Gil, P.; Liu, Y.; Orbović, V.; Verkamp, E.; Poff, K.L.; Green, P.J. Characterization of the auxin-inducible SAUR-AC1 gene for use as a molecular genetic tool in Arabidopsis. Plant Physiol. 1994, 104, 777–784. [Google Scholar] [CrossRef] [PubMed]
- Roux, C.; Bilang, J.; Theunissen, B.H.; Perrot-Rechenmann, C. Identification of new early auxin markers in tobacco by mRNA differential display. Plant Mol. Biol. 1998, 37, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Bao, Y.; Wang, B.; Liu, L.; Chen, J.; Dai, L.; Baloch, S.U.; Peng, D. Identification of small auxin-up RNA (SAUR) genes in Urticales plants: Mulberry (Morus notabilis), hemp (Cannabis sativa) and ramie (Boehmeria nivea). J. Genet. 2016, 95, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Chae, K.; Isaacs, C.G.; Reeves, P.H.; Maloney, G.S.; Muday, G.K.; Nagpal, P.; Reed, J.W. Arabidopsis SMALL AUXIN UP RNA63 promotes hypocotyl and stamen filament elongation. Plant J. 2012, 71, 684–697. [Google Scholar] [CrossRef] [PubMed]
- Spartz, A.K.; Lee, S.H.; Wenger, J.P.; Gonzalez, N.; Itoh, H.; Inzé, D.; Peer, W.A.; Murphy, A.S.; Overvoorde, P.J.; Gray, W.M. The SAUR19 subfamily of SMALL AUXIN UP RNA genes promote cell expansion. Plant J. 2012, 70, 978–990. [Google Scholar] [CrossRef] [PubMed]
- Spartz, A.K.; Ren, H.; Park, M.Y.; Grandt, K.N.; Lee, S.H.; Murphy, A.S.; Sussman, M.R.; Overvoorde, P.J.; Gray, W.M. SAUR inhibition of PP2C-D phosphatases activates plasma membrane H+-ATPases to promote cell expansion in Arabidopsis. Plant Cell 2014, 26, 2129–2142. [Google Scholar] [CrossRef]
- Ren, H.; Park, M.Y.; Spartz, A.K.; Wong, J.H.; Gray, W.M. A subset of plasma membrane-localized PP2C.D phosphatases negatively regulate SAUR-mediated cell expansion in Arabidopsis. PLoS Genet. 2018, 14, e1007455. [Google Scholar] [CrossRef]
- Roig-Villanova, I.; Bou-Torrent, J.; Galstyan, A.; Carretero-Paulet, L.; Portolés, S.; Rodríguez-Concepción, M.; Martínez-García, J.F. Interaction of shade avoidance and auxin responses: A role for two novel atypical bHLH proteins. EMBO J. 2007, 26, 4756–4767. [Google Scholar] [CrossRef]
- Sato, A.; Sasaki, S.; Matsuzaki, J.; Yamamoto, K.T. Light-dependent gravitropism and negative phototropism of inflorescence stems in a dominant Aux/IAA mutant of Arabidopsis thaliana, axr2. J. Plant Res. 2014, 127, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Bemer, M.; van Mourik, H.; Muiño, J.M.; Ferrándiz, C.; Kaufmann, K.; Angenent, G.C. FRUITFULL controls SAUR10 expression and regulates Arabidopsis growth and architecture. J. Exp. Bot. 2017, 68, 3391–3403. [Google Scholar] [CrossRef] [PubMed]
- He, S.L.; Hsieh, H.L.; Jauh, G.Y. SMALL AUXIN UP RNA62/75 are required for the translation of transcripts essential for pollen tube growth. Plant Physiol. 2018, 178, 626–640. [Google Scholar] [CrossRef] [PubMed]
- Favero, D.S.; Le, K.N.; Neff, M.M. Brassinosteroid signaling converges with SUPPRESSOR OF PHYTOCHROME B4-#3 to influence the expression of SMALL AUXIN UP RNA genes and hypocotyl growth. Plant J. 2017, 89, 1133–1145. [Google Scholar] [PubMed]
- Stamm, P.; Kumar, P.P. Auxin and gibberellin responsive Arabidopsis SMALL AUXIN UP RNA36 regulates hypocotyl elongation in the light. Plant Cell Rep. 2013, 32, 759–769. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.G.; Chen, H.W.; Li, Q.T.; Tao, J.J.; Bian, X.H.; Ma, B.; Zhang, W.K.; Chen, S.Y.; Zhang, J.S. Three SAUR proteins SAUR76, SAUR77 and SAUR78 promote plant growth in Arabidopsis. Sci. Rep. 2015, 5, 12477. [Google Scholar] [CrossRef] [PubMed]
- Xie, R.; Dong, C.; Ma, Y.; Deng, L.; He, S.; Yi, S.; Lv, Q.; Zheng, Y. Comprehensive analysis of SAUR gene family in citrus and its transcriptional correlation with fruitlet drop from abscission zone A. Funct. Integr. Genomics 2015, 15, 729–740. [Google Scholar] [CrossRef] [PubMed]
- Spartz, A.K.; Lor, V.S.; Ren, H.; Olszewski, N.E.; Miller, N.D.; Wu, G.; Spalding, E.P.; Gray, W.M. Constitutive expression of Arabidopsis SMALL AUXIN UP RNA19 (SAUR19) in tomato confers auxin-independent hypocotyl elongation. Plant Physiol. 2017, 173, 1453–1462. [Google Scholar] [CrossRef]
- Xu, Y.X.; Xiao, M.Z.; Liu, Y.; Fu, J.L.; He, Y.; Jiang, D.A. The small auxin-up RNA OsSAUR45 affects auxin synthesis and transport in rice. Plant Mol. Biol. 2017, 94, 97–107. [Google Scholar] [CrossRef]
- Ma, P.; Chen, X.; Liu, C.; Meng, Y.; Xia, Z.; Zeng, C.; Lu, C.; Wang, W. MeSAUR1, encoded by a Small Auxin-Up RNA gene, acts as a transcription regulator to positively regulate ADP-Glucose Pyrophosphorylase Small Subunit1a gene in cassava. Front. Plant Sci. 2017, 8, 1315. [Google Scholar] [CrossRef]
- Jiao, W.B.; Schneeberger, K. The impact of third generation genomic technologies on plant genome assembly. Curr. Opin. Plant Biol. 2017, 36, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Tyagi, A.K.; Khurana, J.P. Genome-wide analysis, evolutionary expansion, and expression of early auxin-responsive SAUR gene family in rice (Oryza sativa). Genomics 2006, 88, 360–371. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Hao, X.; Cao, J. Small auxin upregulated RNA (SAUR) gene family in maize: Identification, evolution, and its phylogenetic comparison with Arabidopsis, rice, and sorghum. J. Integr. Plant Biol. 2014, 56, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Liu, S.; He, Y.; Guan, X.; Zhu, X.; Cheng, L.; Wang, J.; Lu, G. Genome-wide analysis of SAUR gene family in Solanaceae species. Gene 2012, 509, 38–50. [Google Scholar] [CrossRef] [PubMed]
- Bai, Q.; Hou, D.; Li, L.; Cheng, Z.; Ge, W.; Liu, J.; Li, X.; Mu, S.; Gao, J. Genome-wide analysis and expression characteristics of small auxin-up RNA (SAUR) genes in moso bamboo (Phyllostachys edulis). Genome 2017, 60, 325–336. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Huang, X.; Bao, Y.; Wang, B.; Zeng, H.; Cheng, W.; Tang, M.; Li, Y.; Ren, J.; Sun, Y. Genome-wide identification of SAUR genes in watermelon (Citrullus lanatus). Physiol. Mol. Biol. Plants 2017, 23, 619–628. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, G.; Geng, Y.; Wu, M.; Pei, W.; Zhai, H.; Zang, X.; Li, X.; Zhang, J.; Yu, S.; et al. A genome-wide analysis of the small auxin-up RNA (SAUR) gene family in cotton. BMC Genom. 2017, 18, 815. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Yan, H.; Luo, S.; Pan, F.; Wang, Y.; Xiang, Y. Genome-wide analysis of poplar SAUR gene family and expression profiles under cold, polyethylene glycol and indole-3-acetic acid treatments. Plant Physiol. Biochem. 2018, 128, 50–65. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Gray, W.M. SAUR proteins as effectors of hormonal and environmental signals in plant growth. Mol. Plant 2015, 8, 1153–1164. [Google Scholar] [CrossRef]
- Stortenbeker, N.; Bemer, M. The SAUR gene family: The plant’s toolbox for adaptation of growth and development. J. Exp. Bot. 2019, 70, 17–27. [Google Scholar] [CrossRef]
- Huang, X.; Xiao, M.; Xi, J.; He, C.; Zheng, J.; Chen, H.; Gao, J.; Zhang, S.; Wu, W.; Liang, Y.; et al. De novo transcriptome assembly of Agave H11648 by Illumina sequencing and identification of cellulose synthase genes in Agave species. Genes 2019, 10, 103. [Google Scholar] [CrossRef] [PubMed]
- Robert, M.L.; Lim, K.Y.; Hanson, L.; Sanchez-Teyer, F.; Bennett, M.D.; Leitch, A.R.; Leitch, I.J. Wild and agronomically important Agave species (Asparagaceae) show proportional increases in chromosome number, genome size, and genetic markers with increasing ploidy. Bot. J. Linn. Soc. 2010, 158, 215–222. [Google Scholar] [CrossRef]
- Gross, S.M.; Martin, J.A.; Simpson, J.; Abraham-Juarez, M.J.; Wang, Z.; Visel, A. De novo transcriptome assembly of drought tolerant CAM plants, Agave deserti and Agave tequilana. BMC Genomics 2013, 14, 563. [Google Scholar] [CrossRef] [PubMed]
- Abraham, P.E.; Yin, H.; Borland, A.M.; Weighill, D.; Lim, S.D.; De Paoli, H.C.; Engle, N.; Jones, P.C.; Agh, R.; Weston, D.J.; et al. Transcript, protein and metabolite temporal dynamics in the CAM plant Agave. Nat. Plants 2016, 2, 16178. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Huang, X.; Wang, B.; Xi, J.; Zhang, Y.; He, C.; Zheng, J.; Gao, J.; Chen, H.; Zhang, S.; Wu, W.; et al. Transcriptome comparison reveals distinct selection patterns in domesticated and wild Agave species, the important CAM plants. Int. J. Genom. 2018, 2018, 5716518. [Google Scholar] [CrossRef]
- Rombel, I.T.; Sykes, K.F.; Rayner, S.; Johnston, S.A. ORF-FINDER: A vector for high-throughput gene identification. Gene 2002, 282, 33–41. [Google Scholar] [CrossRef]
- Yu, C.S.; Chen, Y.C.; Lu, C.H.; Hwang, J.K. Prediction of protein subcellular localization. Proteins 2006, 64, 643–651. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- DNAMAN-Bioinformatics Solutions. Available online: www.lynnon.com (accessed on 30 March 2019).
- Li, F.; Zhang, L.; Li, X.; Guo, B.; Chen, L.; Qi, Z. Sisal tolerance of cupreous and its accumulation preliminary explore. Chin. Agric. Sci. Bull. 2006, 22, 417–420. [Google Scholar]
- Chen, L.; Zhang, L.; L, F.; Guo, B.; Li, X.; Liao, X.; Qi, Z. A primary research on sisal’s uptake property and the accumulation rule to Pb ions. J. Agro Environ. Sci. 2007, 26, 1879–1883. [Google Scholar]
- Wang, P.; Gao, J.; Yang, F.; Zheng, J.; Liu, Q.; Chen, H.; Yi, K. Transcriptome of sisal leaf pretreated with Phytophthora nicotianae Breda. Chin. J. Trop. Crops 2014, 35, 576–582. [Google Scholar]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Wang, L.; You, F.M.; Rodriguez, J.C.; Deal, K.R.; Chen, L.; Li, J.; Chakraborty, S.; Balan, B.; Jiang, C.Z.; et al. Sequencing a Juglans regia × J. microcarpa hybrid yields high-quality genome assemblies of parental species. Hortic. Res. 2019, 6, 55. [Google Scholar] [CrossRef] [PubMed]
- Aubry, S.; Aresheva, O.; Reyna-Llorens, I.; Smith-Unna, R.D.; Hibberd, J.M.; Genty, B. A Specific Transcriptome Signature for Guard Cells from the C4 Plant Gynandropsis gynandra. Plant Physiol. 2016, 170, 1345–1357. [Google Scholar] [CrossRef]
- Jain, M.; Khurana, J.P. Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J. 2009, 276, 3148–3162. [Google Scholar] [CrossRef]
- Hara, M.; Kashima, D.; Horiike, T.; Kuboi, T. Metal-binding characteristics of the protein which shows the highest histidine content in the Arabidopsis genome. Plant Biotechnol. 2010, 27, 475–480. [Google Scholar] [CrossRef][Green Version]
- Bacete, L.; Mélida, H.; Miedes, E.; Molina, A. Plant cell wall-mediated immunity: Cell wall changes trigger disease resistance responses. Plant J. 2018, 93, 614–636. [Google Scholar] [CrossRef]
- Jalmi, S.K.; Bhagat, P.K.; Verma, D.; Noryang, S.; Tayyeba, S.; Singh, K.; Sharma, D.; Sinha, A.K. Traversing the Links between Heavy Metal Stress and Plant Signaling. Front. Plant Sci. 2018, 9, 12. [Google Scholar] [CrossRef]

| Subcellular Position | A. deserti | A. tequilana | A. H11648 | A. americana |
|---|---|---|---|---|
| Chloroplast | 3 | 3 | 2 | 1 |
| Cytoplasm | 7 | 3 | 4 | 2 |
| Extracellular | 0 | 2 | 0 | 0 |
| Mitochondria | 12 | 16 | 7 | 7 |
| Nucleus | 20 | 34 | 9 | 10 |
| Plasma Membrane | 1 | 2 | 2 | 1 |
| Total | 43 | 60 | 24 | 21 |
| Species | I | II | III | IV | V | VI | VII | VIII | Total |
|---|---|---|---|---|---|---|---|---|---|
| A. thaliana | 8 | 5 | 11 | 36 | 3 | 5 | 6 | 5 | 79 |
| O. sativa | 5 | 8 | 18 | 3 | 1 | 2 | 16 | 3 | 56 |
| A. deserti | 4 | 5 | 3 | 8 | 0 | 1 | 15 | 7 | 43 |
| A. tequilana | 9 | 10 | 8 | 6 | 0 | 5 | 12 | 10 | 60 |
| A. H11648 | 3 | 2 | 4 | 5 | 0 | 1 | 5 | 4 | 24 |
| A. americana | 2 | 1 | 5 | 3 | 0 | 2 | 4 | 4 | 21 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Deng, G.; Huang, X.; Xie, L.; Tan, S.; Gbokie, T., Jr.; Bao, Y.; Xie, Z.; Yi, K. Identification and Expression of SAUR Genes in the CAM Plant Agave. Genes 2019, 10, 555. https://doi.org/10.3390/genes10070555
Deng G, Huang X, Xie L, Tan S, Gbokie T Jr., Bao Y, Xie Z, Yi K. Identification and Expression of SAUR Genes in the CAM Plant Agave. Genes. 2019; 10(7):555. https://doi.org/10.3390/genes10070555
Chicago/Turabian StyleDeng, Gang, Xing Huang, Li Xie, Shibei Tan, Thomas Gbokie, Jr., Yaning Bao, Zhouli Xie, and Kexian Yi. 2019. "Identification and Expression of SAUR Genes in the CAM Plant Agave" Genes 10, no. 7: 555. https://doi.org/10.3390/genes10070555
APA StyleDeng, G., Huang, X., Xie, L., Tan, S., Gbokie, T., Jr., Bao, Y., Xie, Z., & Yi, K. (2019). Identification and Expression of SAUR Genes in the CAM Plant Agave. Genes, 10(7), 555. https://doi.org/10.3390/genes10070555





