From Cells to Organoids: Approaches, Regulatory Mechanisms, Applications, and Challenges of Organoids
Highlights
- This article systematically discusses the construction approaches, regulatory mechanisms, significant applications, perspectives, and challenges of organoids.
- It facilitates the development of organoids in unveiling organogenesis mechanisms and disease etiology.
- It promotes the application and development of organoids in drug screening and toxicity testing.
- It lays a foundation for the development of organoids in personalized medicine, regenerative medicine, and alternatives to animal experiments.
Abstract
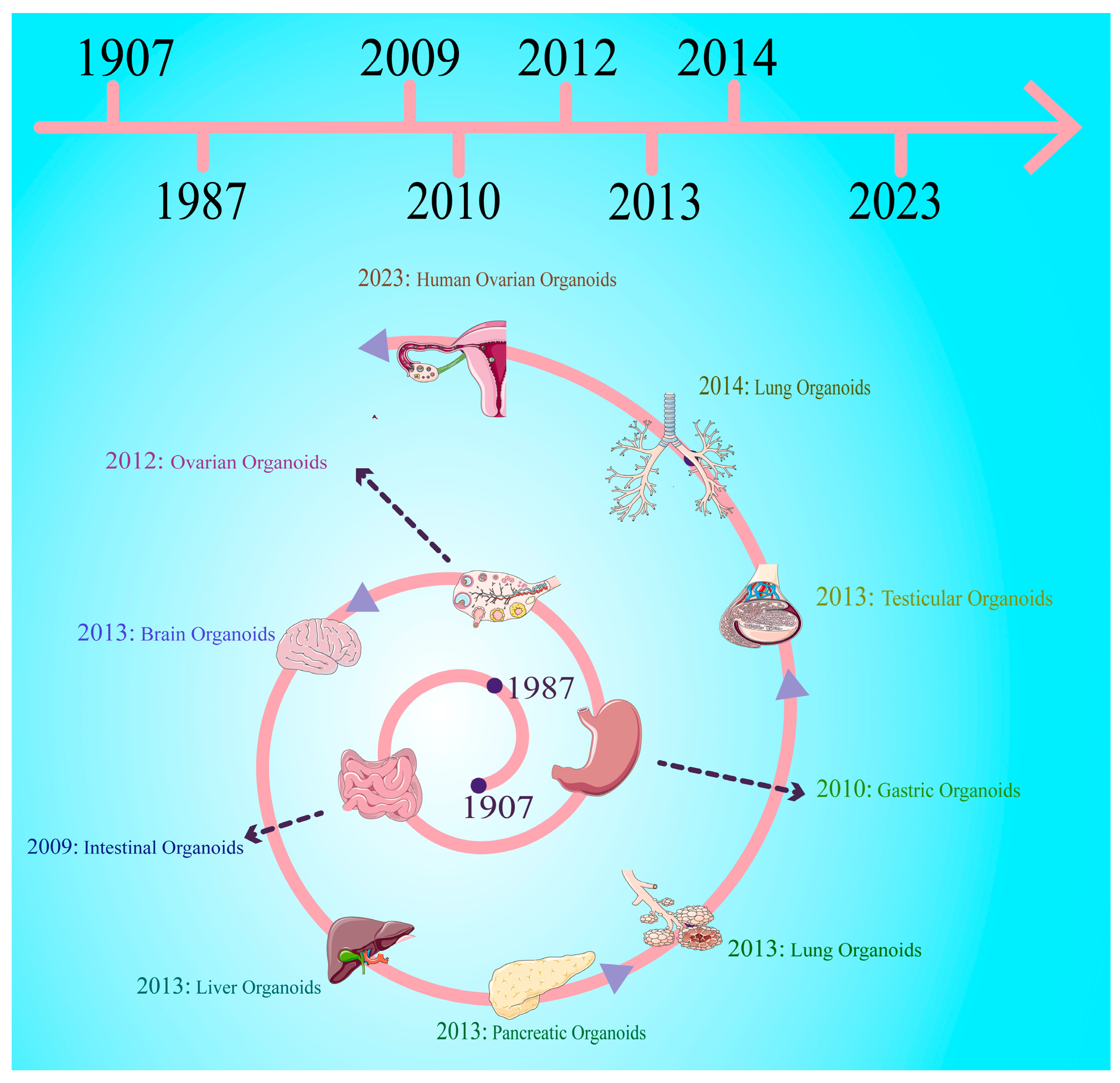
1. Introduction
2. Methods of Organoid Construction
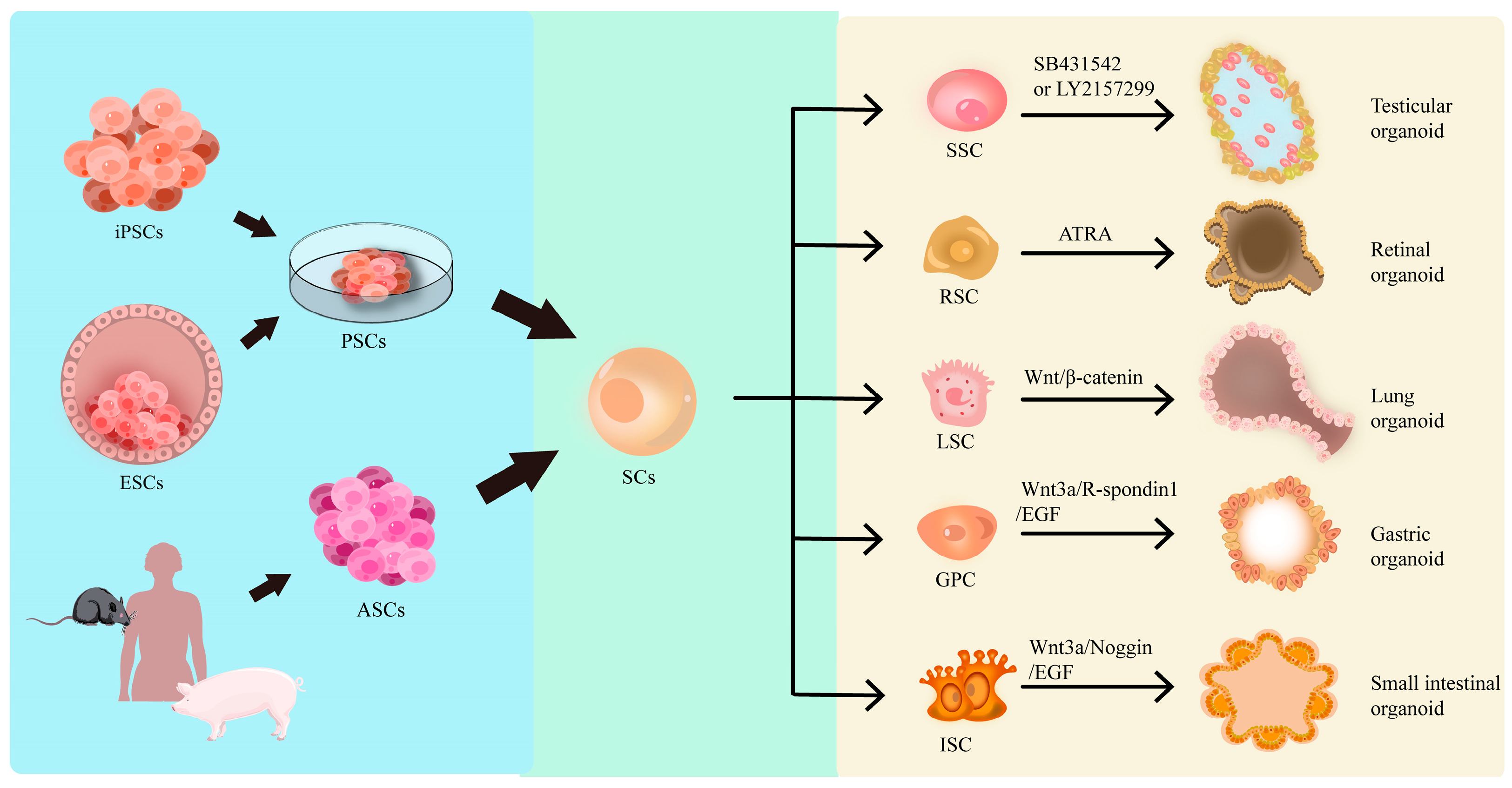
2.1. Selection and Culture of Cells
2.1.1. Pluripotent Stem Cells (PSCs): ESCs and iPSCs
2.1.2. Adult Stem Cells (ASCs)
2.1.3. Patient-Derived Cells
2.2. The Effect of the Microenvironment or Niche on Organoid Development
2.2.1. Regulation of Physical Cues Stabilizes the Organoid Microenvironment
2.2.2. Signaling Pathways in Regulating Fate Specification of Organoids
2.3. Self-Assembly and 3D Structure Formation of Organoids
2.4. Maturation Identification and Functional Characterization of Organoids
2.5. Organoid Types
3. Regulatory Mechanisms Underlying Organoid Development
3.1. Cell-Autonomous Regulation
3.2. Regulation of the Extracellular Microenvironment (ECM) in Organoids
3.3. Tissue-Specific Signaling Pathways and Intercellular Interactions
3.4. Spatiotemporal Regulation of Organoids
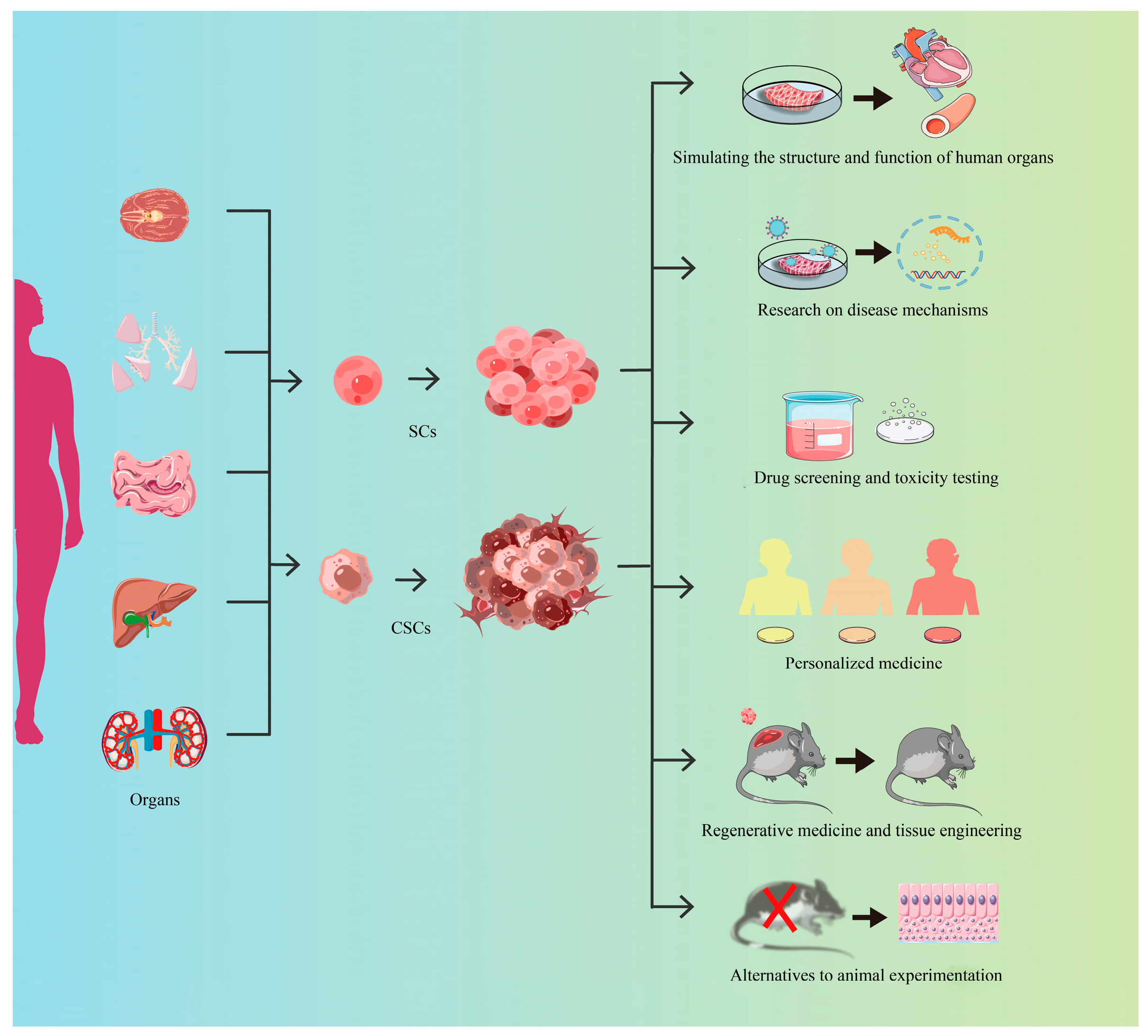
4. Applications of the Organoids Across the Spectrum of Biological Research
4.1. Simulating the Structures and Functions of Human Organs by Organoids
4.2. Disease Mechanism Research Using Organoids
4.3. Drug Screening and Toxicity Testing Using Organoids
4.4. Personalized Medicine Facilitated by the Use of Organoids
4.5. Regenerative Medicine and Tissue Engineering of Organoids
4.6. Novel Alternative Methods (NAMs) by the Use of Organoids
4.7. The Challenges and Prospects of Translational Medicine with Organoids
5. Perspectives and Challenges of Organoids
5.1. Technological Advancement and the Use of Innovative Materials
5.2. Core Challenges and Breakthroughs in the Use of Organoids
5.2.1. Core Technical Challenges
5.2.2. Technical Breakthroughs and Optimization Directions
5.3. Ethical Issues and the Establishment of Regulatory Frameworks for Organoid Use
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Sato, T.; Vries, R.G.; Snippert, H.J.; Van De Wetering, M.; Barker, N.; Stange, D.E.; Van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Lovell-Badge, R.; Anthony, E.; Barker, R.A.; Bubela, T.; Brivanlou, A.H.; Carpenter, M.; Charo, R.A.; Clark, A.; Clayton, E.; Cong, Y.; et al. ISSCR guidelines for stem cell research and clinical translation: The 2021 update. Stem Cell Rep. 2021, 16, 1398–1408. [Google Scholar] [CrossRef]
- Huang, L.; Bernink, J.H.; Giladi, A.; Krueger, D.; van Son, G.J.F.; Geurts, M.H.; Busslinger, G.; Lin, L.; Begthel, H.; Zandvliet, M.; et al. Tuft cells act as regenerative stem cells in the human intestine. Nature 2024, 634, 929–935. [Google Scholar] [CrossRef]
- Tian, C.; Yang, M.; Xu, H.; Zhu, M.-Z.; Yue, N.-N.; Zhang, Y.; Shi, R.-Y.; Yao, J.; Wang, L.-S.; Liang, Y.-J.; et al. Stem cell-derived intestinal organoids: A novel modality for IBD. Cell Death Discov. 2023, 9, 255. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Du, Y.; Kwan, W.; Yan, R.; Shi, Q.; Zang, L.; Zhu, Z.; Zhang, J.; Li, C.; Yu, Y. A quick and reliable image-based AI algorithm for evaluating cellular senescence of gastric organoids. Cancer Biol. Med. 2023, 20, 519–536. [Google Scholar] [CrossRef] [PubMed]
- Foo, M.A.; You, M.; Chan, S.L.; Sethi, G.; Bonney, G.K.; Yong, W.-P.; Chow, E.K.-H.; Fong, E.L.S.; Wang, L.; Goh, B.-C. Clinical translation of patient-derived tumour organoids-bottlenecks and strategies. Biomark. Res. 2022, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Gayibov, E.; Sychra, T.; Spalenkova, A.; Souček, P.; Oliverius, M. The use of patient-derived xenografts and patient-derived organoids in the search for new therapeutic regimens for pancreatic carcinoma. A review. Biomed. Pharmacother. 2025, 182, 117750. [Google Scholar] [CrossRef]
- Lee, S.; Kim, S.; Ahn, J.; Park, J.; Ryu, B.-Y.; Park, J.Y. Membrane-bottomed microwell array added to transwell insert to facilitate non-contact co-culture of spermatogonial stem cell and STO feeder cell. Biofabrication 2020, 12, 45031. [Google Scholar] [CrossRef]
- Eiraku, M.; Watanabe, K.; Matsuo-Takasaki, M.; Kawada, M.; Yonemura, S.; Matsumura, M.; Wataya, T.; Nishiyama, A.; Muguruma, K.; Sasai, Y. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 2008, 3, 519–532. [Google Scholar] [CrossRef]
- Wilson, H.V. A new method by which sponges may be artificially reared. Science 1907, 25, 912–915. [Google Scholar] [CrossRef]
- Rheinwatd, J.G.; Green, H. Serial cultivation of strains of human epidermal keratinocytes: The formation of keratinizing colonies from single cells. Cell 1975, 6, 331–343. [Google Scholar] [CrossRef] [PubMed]
- Barker, N.; Huch, M.; Kujala, P.; Van De Wetering, M.; Snippert, H.J.; Van Es, J.H.; Sato, T.; Stange, D.E.; Begthel, H.; van den Born, M.; et al. Lgr5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 2010, 6, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Takebe, T.; Sekine, K.; Enomura, M.; Koike, H.; Kimura, M.; Ogaeri, T.; Zhang, R.; Ueno, Y.; Zheng, Y.-W.; Koike, N.; et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 2013, 499, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Xia, H.; Meng, T.; Fan, Y.; Tang, X.; Li, Y.; Zhang, T.; Deng, J.; Yao, B.; Huang, Y.; et al. Construction of human pluripotent stem cell-derived testicular organoids and their use as humanized testis models for evaluating the effects of semaglutide. Theranostics 2025, 15, 2597–2623. [Google Scholar] [CrossRef]
- Iwasawa, K.; Takebe, T. Organogenesis in vitro. Curr. Opin. Cell Biol. 2021, 73, 84–91. [Google Scholar] [CrossRef]
- Fan, X.; Hou, K.; Liu, G.; Shi, R.; Wang, W.; Liang, G. Strategies to overcome the limitations of current organoid technology—Engineered organoids. J. Tissue Eng. 2025, 16, 20417314251319475. [Google Scholar] [CrossRef]
- Yang, H.; Wang, Y.; Wang, P.; Zhang, N.; Wang, P. Tumor organoids for cancer research and personalized medicine. Cancer Biol. Med. 2021, 19, 319–332. [Google Scholar] [CrossRef]
- Fang, Z.; Li, P.; Du, F.; Shang, L.; Li, L. The role of organoids in cancer research. Exp. Hematol. Oncol. 2023, 12, 69. [Google Scholar] [CrossRef]
- Zhang, Z.; Wu, Y.; Xuan, Z.; Xu, H.; Yin, S.; Meng, Z. Self-assembly of three-dimensional liver organoids: Virtual reconstruction via endocytosed polymer dots for refactoring the fine structure. Biomater. Sci. 2023, 11, 7867–7883. [Google Scholar] [CrossRef]
- Keshara, R.; Kim, Y.H.; Grapin-Botton, A. Organoid imaging: Seeing development and function. Annu. Rev. Cell Dev. Biol. 2022, 38, 447–466. [Google Scholar] [CrossRef]
- Geng, Y.; Chen, Z.; Luo, T.; Liu, Y.; Kong, S.; Yan, X.; Bai, H.; Wang, Y. Innovative construction and application of bile duct organoids: Unraveling the complexity of bile duct diseases and potential therapeutic strategies. Cancer Lett. 2025, 618, 217619. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.J.; Kaufman, M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature 1981, 292, 154–156. [Google Scholar] [CrossRef] [PubMed]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic stem cell lines derived from human blastocysts. Science 1998, 282, 1145–1147. [Google Scholar] [CrossRef] [PubMed]
- Keller, G. Embryonic stem cell differentiation: Emergence of a new era in biology and medicine. Genes. Dev. 2005, 19, 1129–1155. [Google Scholar] [CrossRef]
- de Jongh, D.; Massey, E.K.; Bunnik, E.M. Organoids: A systematic review of ethical issues. Stem Cell Res. Ther. 2022, 13, 337. [Google Scholar] [CrossRef]
- Gaykema, L.H.; van Nieuwland, R.Y.; Lievers, E.; Moerkerk, W.B.J.; de Klerk, J.A.; Dumas, S.J.; Kers, J.; Zaldumbide, A.; Berg, C.W.v.D.; Rabelink, T.J. T-cell mediated immune rejection of beta-2-microglobulin knockout induced pluripotent stem cell-derived kidney organoids. Stem Cells Transl. Med. 2024, 13, 69–82. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- Hirami, Y.; Mandai, M.; Sugita, S.; Maeda, A.; Maeda, T.; Yamamoto, M.; Uyama, H.; Yokota, S.; Fujihara, M.; Igeta, M.; et al. Safety and stable survival of stem-cell-derived retinal organoid for 2 years in patients with retinitis pigmentosa. Cell Stem Cell 2023, 30, 1585–1596. [Google Scholar] [CrossRef]
- Kustermann, C.; Narbute, K.; Movcana, V.; Parfejevs, V.; Rūmnieks, F.; Kauķis, P.; Priedols, M.; Mikilps-Mikgelbs, R.; Mihailova, M.; Andersone, S.; et al. IPSC-derived lung and lung cancer organoid model to evaluate cisplatin encapsulated autologous iPSC-derived mesenchymal stromal cell-isolated extracellular vesicles. Stem Cell Res. Ther. 2024, 15, 246. [Google Scholar] [CrossRef]
- Zhu, X.; Huang, L.; Li, G.; Deng, B.; Wang, X.; Yang, H.; Zhang, Y.; Wen, Q.; Wang, C.; Zhang, J.; et al. Genome-wide silencer screening reveals key silencer modulating reprogramming efficiency in mouse induced pluripotent stem cells. Adv. Sci. 2025, 12, e2408839. [Google Scholar] [CrossRef]
- Bai, X. Stem cell-based disease modeling and cell therapy. Cells 2020, 9, 2193. [Google Scholar] [CrossRef]
- Maggiore, J.C.; LeGraw, R.; Przepiorski, A.; Velazquez, J.; Chaney, C.; Vanichapol, T.; Streeter, E.; Almuallim, Z.; Oda, A.; Chiba, T.; et al. A genetically inducible endothelial niche enables vascularization of human kidney organoids with multilineage maturation and emergence of renin expressing cells. Kidney Int. 2024, 106, 1086–1100. [Google Scholar] [CrossRef]
- Yamanaka, S. Pluripotent stem cell-based cell therapy-promise and challenges. Cell Stem Cell 2020, 27, 523–531. [Google Scholar] [CrossRef]
- Zeevaert, K.; Elsafi Mabrouk, M.H.; Wagner, W.; Goetzke, R. Cell mechanics in embryoid bodies. Cells 2020, 9, 2270. [Google Scholar] [CrossRef]
- Augustyniak, J.; Kozlowska, H.; Buzanska, L. Genes involved in DNA repair and mitophagy protect embryoid bodies from the toxic effect of methylmercury chloride under physioxia conditions. Cells 2023, 12, 390. [Google Scholar] [CrossRef]
- Zeevaert, K.; Goetzke, R.; Elsafi Mabrouk, M.H.; Schmidt, M.; Maaßen, C.; Henneke, A.-C.; He, C.; Gillner, A.; Zenke, M.; Wagner, W. YAP1 is essential for self-organized differentiation of pluripotent stem cells. Biomater. Adv. 2023, 146, 213308. [Google Scholar] [CrossRef] [PubMed]
- Eicher, A.K.; Kechele, D.O.; Sundaram, N.; Berns, H.M.; Poling, H.M.; Haines, L.E.; Sanchez, J.G.; Kishimoto, K.; Krishnamurthy, M.; Han, L.; et al. Functional human gastrointestinal organoids can be engineered from three primary germ layers derived separately from pluripotent stem cells. Cell Stem Cell 2022, 29, 36–51. [Google Scholar] [CrossRef] [PubMed]
- Koppes, A.N.; Koppes, R.A.; Nichols, K.N. These organoids have some nerve: Uniting three germ layers in a human gastric model system. Cell Stem Cell 2022, 29, 5–6. [Google Scholar] [CrossRef]
- Huang, M.; Chen, M.; Yuan, G.; Cui, Y.; Shen, B.; Liu, Z.; Zhang, B.; Chen, J.; Chen, D.; Qiu, S.; et al. Establishment of human gastrulating stem cells with the capacity of stable differentiation into multiple gastrulating cell types. Cell Res. 2025, 35, 719–734. [Google Scholar] [CrossRef] [PubMed]
- Artegiani, B.; Hendriks, D. Organoids from pluripotent stem cells and human tissues: When two cultures meet each other. Dev. Cell 2025, 60, 493–511. [Google Scholar] [CrossRef]
- Huch, M.; Gehart, H.; van Boxtel, R.; Hamer, K.; Blokzijl, F.; Verstegen, M.M.; Ellis, E.; van Wenum, M.; Fuchs, S.A.; de Ligt, J.; et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 2015, 160, 299–312. [Google Scholar] [CrossRef]
- Prakash, N.; Kim, J.; Jeon, J.; Kim, S.; Arai, Y.; Bello, A.B.; Park, H.; Lee, S.H. Progress and emerging techniques for biomaterial-based derivation of mesenchymal stem cells (MSCs) from pluripotent stem cells (PSCs). Biomater. Res. 2023, 27, 31. [Google Scholar] [CrossRef]
- Merlos-Suarez, A.; Barriga, F.M.; Jung, P.; Iglesias, M.; Céspedes, M.V.; Rossell, D.; Sevillano, M.; Hernando-Momblona, X.; da Silva-Diz, V.; Muñoz, P.; et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 2011, 8, 511–524. [Google Scholar] [CrossRef] [PubMed]
- Atanasova, V.S.; de Jesus Cardona, C.; Hejret, V.; Tiefenbacher, A.; Mair, T.; Tran, L.; Pfneissl, J.; Draganić, K.; Binder, C.; Kabiljo, J.; et al. Mimicking tumor cell heterogeneity of colorectal cancer in a patient-derived organoid-fibroblast model. Cell. Mol. Gastroenterol. Hepatol. 2023, 15, 1391–1419. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, R.; Liu, Y.; Lin, X.; Chen, J. Cancer patient-derived organoids: Novel models for the study of natural products. Int. J. Biol. Sci. 2025, 21, 4485–4503. [Google Scholar] [CrossRef]
- Cho, J.; Park, J.J.; Seo, E.; Lee, O.-H.; Cho, T.-J.; Kim, J.Y.; Bae, H.C.; Lee, E.; Park, Y.; Jang, H.; et al. Self-assembled organoid-tissue modules for scalable organoid engineering: Application to chondrogenic regeneration. Acta Biomater. 2025, 197, 152–166. [Google Scholar] [CrossRef]
- Huo, Y.; Ci, Z.; Wu, S.; Feng, S.; Sun, Y.; Li, G.; Liu, Y.; Hua, Y.; Zhang, Y.; Zhou, G. Scaffold-free three-dimensional cartilage regeneration based on cartilaginous organoids bioassembly technology. Aggregate 2024, 5, e619. [Google Scholar] [CrossRef]
- Kim, J.W.; Nam, S.A.; Yi, J.; Kim, J.Y.; Lee, J.Y.; Park, S.; Sen, T.; Choi, Y.; Lee, J.Y.; Kim, H.L.; et al. Kidney decellularized extracellular matrix enhanced the vascularization and maturation of human kidney organoids. Adv. Sci. 2022, 9, e2103526. [Google Scholar] [CrossRef]
- Zhang, D.; Jin, W.; Cui, Y.; He, Z. Establishment and characterization of testis organoids with proliferation and differentiation of spermatogonial stem cells into spermatocytes and spermatids. Cells 2024, 13, 1642. [Google Scholar] [CrossRef]
- Lewis-Israeli, Y.R.; Wasserman, A.H.; Gabalski, M.A.; Volmert, B.D.; Ming, Y.; Ball, K.A.; Yang, W.; Zou, J.; Ni, G.; Pajares, N.; et al. Self-assembling human heart organoids for the modeling of cardiac development and congenital heart disease. Nat. Commun. 2021, 12, 5142. [Google Scholar] [CrossRef]
- Lai, W.; Geliang, H.; Bin, X.; Wang, W. Effects of hydrogel stiffness and viscoelasticity on organoid culture: A comprehensive review. Mol. Med. 2025, 31, 83. [Google Scholar] [CrossRef] [PubMed]
- Ding, H.; Chen, D.; Tan, X.; Xu, X.; Li, G.; Xu, L.; Zhang, Y.; Song, J. Enhanced bone repair using callus organoids derived from urine-derived stem cells with silk fibroin. Adv. Healthc. Mater. 2025, 14, e2501852. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.N.; Ishahak, M.; Chaimov, D.; Velraj, A.; LaShoto, D.; Hagan, D.W.; Buchwald, P.; Phelps, E.A.; Agarwal, A.; Stabler, C.L. Organoid microphysiological system preserves pancreatic islet function within 3d matrix. Sci. Adv. 2021, 7, eaba5515. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Kimura, M.; Shrestha, S.; Lewis, P.; Lee, S.; Cai, Y.; Joshi, P.; Acharya, P.; Liu, J.; Yang, Y.; et al. A pillar and perfusion plate platform for robust human organoid culture and analysis. Adv. Healthc. Mater. 2024, 13, e2302502. [Google Scholar] [CrossRef]
- Ya, S.; Ding, W.; Li, S.; Du, K.; Zhang, Y.; Li, C.; Liu, J.; Li, F.; Li, P.; Luo, T.; et al. On-chip construction of liver lobules with self-assembled perfusable hepatic sinusoid networks. ACS Appl. Mater. Interfaces 2021, 13, 32640–32652. [Google Scholar] [CrossRef]
- Gerasimova, E.; Beenen, A.C.; Kachkin, D.; Regensburger, M.; Zundler, S.; Blumenthal, D.B.; Lutzny-Geier, G.; Winner, B.; Prots, I. Novel co-culture model of t cells and midbrain organoids for investigating neurodegeneration in parkinson’s disease. npj Park. Dis. 2025, 11, 36. [Google Scholar] [CrossRef]
- de Lau, W.B.M.; Wijnakker, J.J.A.P.; van Son, G.J.F.; Krueger, D.; Wang, D.; Abendroth, M.S.; Schreurs, R.; Janda, C.Y.; van Rijt, F.L.H.; Chalopin, B.J.P.; et al. A single-chain derivative of an integrin-activating antibody potentiates organoid growth in matrigel and collagen hydrogels. Nat. Biotechnol. 2025; online ahead of print. [Google Scholar] [CrossRef]
- Matkovic Leko, I.; Schneider, R.T.; Thimraj, T.A.; Schrode, N.; Beitler, D.; Liu, H.-Y.; Beaumont, K.; Chen, Y.-W.; Snoeck, H.-W. A distal lung organoid model to study interstitial lung disease, viral infection and human lung development. Nat. Protoc. 2023, 18, 2283–2312. [Google Scholar] [CrossRef]
- Wang, M.; Yu, H.; Zhang, T.; Cao, L.; Du, Y.; Xie, Y.; Ji, J.; Wu, J. In-depth comparison of matrigel dissolving methods on proteomic profiling of organoids. Mol. Cell. Proteom. 2022, 21, 100181. [Google Scholar] [CrossRef]
- Tan, R.; Zhang, Z.; Ding, P.; Liu, Y.; Liu, H.; Lu, M.; Chen, Y.-G. A growth factor-reduced culture system for colorectal cancer organoids. Cancer Lett. 2024, 588, 216737. [Google Scholar] [CrossRef]
- Nuciforo, S.; Heim, M.H. Organoids to model liver disease. JHEP Rep. 2021, 3, 100198. [Google Scholar] [CrossRef]
- Altmaier, S.; Meiser, I.; Lemesre, E.; Chanrion, B.; Steeg, R.; Leonte, L.E.; Holst, B.; Nielsen, B.S.; Clausen, C.; Schmidt, K.; et al. Human iPSC-derived hepatocytes in 2d and 3d suspension culture for cryopreservation and in vitro toxicity studies. Reprod. Toxicol. 2022, 111, 68–80. [Google Scholar] [CrossRef]
- Wang, X.; Liu, N.; Zhang, H.; Yin, Z.-S.; Zha, Z.-G. From cells to organs: Progress and potential in cartilaginous organoids research. J. Transl. Med. 2023, 21, 926. [Google Scholar] [CrossRef]
- Yang, R.; Qi, Y.; Zhang, X.; Gao, H.; Yu, Y. Living biobank: Standardization of organoid construction and challenges. Chin. Med. J. 2024, 137, 3050–3060. [Google Scholar] [CrossRef]
- Birey, F.; Andersen, J.; Makinson, C.D.; Islam, S.; Wei, W.; Huber, N.; Fan, H.C.; Metzler, K.R.C.; Panagiotakos, G.; Thom, N.; et al. Assembly of functionally integrated human forebrain spheroids. Nature 2017, 545, 54–59. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Imaizumi, K.; Jurjut, O.; Kelley, K.W.; Wang, D.; Thete, M.V.; Hudacova, Z.; Amin, N.D.; Levy, R.J.; Scherrer, G.; et al. Human assembloid model of the ascending neural sensory pathway. Nature 2025, 642, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Zushin, P.-J.H.; Mukherjee, S.; Wu, J.C. FDA modernization act 2.0: Transitioning beyond animal models with human cells, organoids, and AI/ML-based approaches. J. Clin. Investig. 2023, 133, e175824. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.; Liu, M.; Li, J.; Zhang, W.; Hu, R.; Yang, G.; Deng, J.; Li, Y.; Li, X.; Liu, Y. Musculoskeletal organoids-on-chip uncover muscle-bone communication under intermittent hypoxia. Natl. Sci. Rev. 2025, 12, nwaf214. [Google Scholar] [CrossRef]
- Sun, C.; Wu, G.; Wu, D.; Wu, L.; Lu, Q.; Hu, W.; Du, Q.; Wang, J.; Xie, A.; Xia, M.; et al. Determining the optimal transplantation window in hepatic organoids via real-time biosensing of vascularization and metabolic maturation utilizing the integrated organoid-on-a-chip platform. Biosens. Bioelectron. 2026, 292, 118057. [Google Scholar] [CrossRef]
- Ramani, A.; Pasquini, G.; Gerkau, N.J.; Jadhav, V.; Vinchure, O.S.; Altinisik, N.; Windoffer, H.; Muller, S.; Rothenaigner, I.; Lin, S.; et al. Reliability of high-quantity human brain organoids for modeling microcephaly, glioma invasion and drug screening. Nat. Commun. 2024, 15, 10703. [Google Scholar] [CrossRef]
- Abilez, O.J.; Yang, H.; Guan, Y.; Shen, M.; Yildirim, Z.; Zhuge, Y.; Venkateshappa, R.; Zhao, S.R.; Gomez, A.H.; El-Mokahal, M.; et al. Gastruloids enable modeling of the earliest stages of human cardiac and hepatic vascularization. Science 2025, 388, eadu9375. [Google Scholar] [CrossRef]
- Lee, J.; Kim, Y.; Jung, H.; Lim, J.; Kwak, B.S. Channel-assembling tumor microenvironment on-chip for evaluating anticancer drug efficacy. J. Control. Release 2025, 377, 376–384. [Google Scholar] [CrossRef]
- Tang, X.; Wu, S.; Wang, D.; Chu, C.; Hong, Y.; Tao, M.; Hu, H.; Xu, M.; Guo, X.; Liu, Y. Human organoids in basic research and clinical applications. Signal Transduct. Target. Ther. 2022, 7, 168. [Google Scholar] [CrossRef] [PubMed]
- Birtele, M.; Lancaster, M.; Quadrato, G. Modelling human brain development and disease with organoids. Nat. Rev. Mol. Cell Biol. 2025, 26, 389–412. [Google Scholar] [CrossRef] [PubMed]
- Maisumu, G.; Willerth, S.; Nestor, M.W.; Waldau, B.; Schülke, S.; Nardi, F.V.; Ahmed, O.; Zhou, Y.; Durens, M.; Liang, B.; et al. Brain organoids: Building higher-order complexity and neural circuitry models. Trends Biotechnol. 2025, 43, 1583–1598. [Google Scholar] [CrossRef]
- Zhou, Z.; Van der Jeught, K.; Li, Y.; Sharma, S.; Yu, T.; Moulana, I.; Liu, S.; Wan, J.; Territo, P.R.; Opyrchal, M.; et al. A t cell-engaging tumor organoid platform for pancreatic cancer immunotherapy. Adv. Sci. 2023, 10, e2300548. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.; Wang, P.; Zhang, D.; Fang, Z.; Feng, Y.; Chen, J.; Chen, J.; Fu, Y.; Yang, B.; Yu, S.; et al. A novel organoid model retaining the glioma microenvironment for personalized drug screening and therapeutic evaluation. Bioact. Mater. 2025, 53, 205–217. [Google Scholar] [CrossRef]
- Lin, L.; DeMartino, J.; Wang, D.; van Son, G.J.F.; van der Linden, R.; Begthel, H.; Korving, J.; Andersson-Rolf, A.; Brink, S.v.D.; Lopez-Iglesias, C.; et al. Unbiased transcription factor CRISPR screen identifies ZNF800 as master repressor of enteroendocrine differentiation. Science 2023, 382, 451–458. [Google Scholar] [CrossRef]
- Mule, P.; Fernandez-Perez, D.; Amato, S.; Manganaro, D.; Oldani, P.; Brandini, S.; Diaferia, G.; Cuomo, A.; Recordati, C.; Soriani, C.; et al. WNT oncogenic transcription requires MYC suppression of lysosomal activity and EPCAM stabilization in gastric tumors. Gastroenterology 2024, 167, 903–918. [Google Scholar] [CrossRef]
- Cammareri, P.; Raponi, M.; Hong, Y.; Billard, C.V.; Peckett, N.; Zhu, Y.; Velez-Bravo, F.D.; Younger, N.T.; Dunican, D.S.; Pohl, S.Ö.-G.; et al. Loss of colonic fidelity enables multilineage plasticity and metastasis. Nature 2025, 644, 547–556. [Google Scholar] [CrossRef]
- Chanda, A.; Sarkar, A.; Deng, L.; Bonni, A.; Bonni, S. Sumoylated SnoN interacts with HDAC1 and p300/CBP to regulate EMT-associated phenotypes in mammary organoids. Cell Death Dis. 2023, 14, 405. [Google Scholar] [CrossRef]
- Martens-de Kemp, S.R.; Komor, M.A.; Hegi, R.; Bolijn, A.S.; Tijssen, M.; de Groen, F.L.; Depla, A.; van Leerdam, M.; Meijer, G.A.; Fijneman, R.J.; et al. Overexpression of the mir-17-92 cluster in colorectal adenoma organoids causes a carcinoma-like gene expression signature. Neoplasia 2022, 32, 100820. [Google Scholar] [CrossRef]
- Kim, S.; Min, S.; Choi, Y.S.; Jo, S.-H.; Jung, J.H.; Han, K.; Kim, J.; An, S.; Ji, Y.W.; Kim, Y.-G.; et al. Tissue extracellular matrix hydrogels as alternatives to matrigel for culturing gastrointestinal organoids. Nat. Commun. 2022, 13, 1692. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Jiao, L.; Feng, D.; Yuan, Y.; Yang, X.; Li, J.; Jiang, D.; Chen, H.; Meng, Q.; Chen, R.; et al. Human apical-out nasal organoids reveal an essential role of matrix metalloproteinases in airway epithelial differentiation. Nat. Commun. 2024, 15, 143. [Google Scholar] [CrossRef] [PubMed]
- Chrisnandy, A.; Lutolf, M.P. An extracellular matrix niche secreted by epithelial cells drives intestinal organoid formation. Dev. Cell 2025, 60, 3116–3130.e8. [Google Scholar] [CrossRef] [PubMed]
- Monckton, C.P.; Brougham-Cook, A.; Underhill, G.H.; Khetani, S.R. Modulation of human iPSC-derived hepatocyte phenotype via extracellular matrix microarrays. Acta Biomater. 2022, 153, 216–230. [Google Scholar] [CrossRef]
- Moraveji, S.; Erfanian, S.; Ghanian, M.H.; Baharvand, H. Harnessing TGF-beta signaling to improve testicular organoid development from dissociated testicular cells. Stem Cell Res. Ther. 2025, 16, 443. [Google Scholar] [CrossRef]
- Miao, Y.; Pek, N.M.; Tan, C.; Jiang, C.; Yu, Z.; Iwasawa, K.; Shi, M.; Kechele, D.O.; Sundaram, N.; Pastrana-Gomez, V.; et al. Co-development of mesoderm and endoderm enables organotypic vascularization in lung and gut organoids. Cell 2025, 188, 4295–4313. [Google Scholar] [CrossRef]
- Isla-Magrane, H.; Zufiaurre-Seijo, M.; Garcia-Arumi, J.; Duarri, A. All-trans retinoic acid modulates pigmentation, neuroretinal maturation, and corneal transparency in human multiocular organoids. Stem Cell Res. Ther. 2022, 13, 376. [Google Scholar] [CrossRef]
- Wang, X.; Li, S.; Liu, Y.; Kuang, X.; Chen, J.; Yu, P.; Zhao, L.; Zhang, Z.; Huang, M.; Liu, L.; et al. Positional BMP signaling orchestrates villus length in the small intestine. Nat. Commun. 2025, 16, 5461. [Google Scholar] [CrossRef]
- Brownfield, D.G.; de Arce, A.D.; Ghelfi, E.; Gillich, A.; Desai, T.J.; Krasnow, M.A. Alveolar cell fate selection and lifelong maintenance of at2 cells by FGF signaling. Nat. Commun. 2022, 13, 7137. [Google Scholar] [CrossRef]
- Lim, K.; Donovan, A.P.A.; Tang, W.; Sun, D.; He, P.; Pett, J.P.; Teichmann, S.A.; Marioni, J.C.; Meyer, K.B.; Brand, A.H.; et al. Organoid modeling of human fetal lung alveolar development reveals mechanisms of cell fate patterning and neonatal respiratory disease. Cell Stem Cell 2023, 30, 20–37. [Google Scholar] [CrossRef]
- Sankoda, N.; Tanabe, W.; Tanaka, A.; Shibata, H.; Woltjen, K.; Chiba, T.; Haga, H.; Sakai, Y.; Mandai, M.; Yamamoto, T.; et al. Epithelial expression of gata4 and sox2 regulates specification of the squamous-columnar junction via MAPK/ERK signaling in mice. Nat. Commun. 2021, 12, 560. [Google Scholar] [CrossRef]
- Willet, S.G.; Thanintorn, N.; McNeill, H.; Huh, S.-H.; Ornitz, D.M.; Huh, W.J.; Hoft, S.G.; DiPaolo, R.J.; Mills, J.C. SOX9 governs gastric mucous neck cell identity and is required for injury-induced metaplasia. Cell. Mol. Gastroenterol. Hepatol. 2023, 16, 325–339. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Ding, H.; Wu, S.; Xiong, N.; Hong, Y.; Zhu, W.; Chen, X.; Han, X.; Tao, M.; Wang, Y.; et al. Artificial meshed vessel-induced dimensional breaking growth of human brain organoids and multiregional assembloids. ACS Nano 2024, 18, 26201–26214. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Liu, S.; Yang, Z.; Yang, L.; Yang, Y.; Han, P.; Miao, Y.; Lin, L.; Xu, L.; Li, Y.; et al. Construction of vascularized liver microtissues recapitulates angiocrine-mediated hepatocytes maturation and enhances therapeutic efficacy for acute liver failure. Bioact. Mater. 2025, 50, 525–539. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, J.F.; Wiegerinck, C.L.; de Jonge, H.R.; Bronsveld, I.; Janssens, H.M.; De Winter-de Groot, K.M.; Brandsma, A.M.; de Jong, N.W.; Bijvelds, M.J.; Scholte, B.J.; et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 2013, 19, 939–945. [Google Scholar] [CrossRef]
- Tran, H.; Shin, M.; Yeo, X.Y.; Jung, S.; Junaid, M.; Bin Lim, S.; Kim, J.; Woo, H.G.; Denman, C.R.; Han, R.-R.; et al. A human striatal-midbrain assembloid model of alpha-synuclein propagation. Brain 2025, awaf326, online ahead of print. [Google Scholar] [CrossRef]
- Hu, L.; Han, M.; Yang, J.; Yu, L.; Qin, M.; Xu, J.; Zhang, H.; Chen, X.; Li, Y.; Meng, H.; et al. Engineering cardiobundles with dynamic microenvironmental control as advanced cardiac modeling for heart-on-a-chip. Small 2025, 21, e2505617. [Google Scholar] [CrossRef]
- Das Adhikari, U.; Froehle, L.M.; Pipkin, A.N.; Baharlou, H.; Linder, A.H.; Shah, P.; Hussey, A.; Zhang, Q.; Nyquist, S.; Khawaled, S.; et al. Immunometabolic defects of CD8+ t cells disrupt gut barrier integrity in people with HIV. Cell 2025, 188, 5666–5679. [Google Scholar] [CrossRef]
- Shin, H.; An, Y.; Lee, J.; Kim, J.H.; Prasad, R.; Kim, K.; Kang, H.; Sun, W.; Cho, S.; Cho, I. One-step drug screening system utilizing electrophysiological activity in multiple brain organoids. Adv. Sci. 2025, 12, e04913. [Google Scholar] [CrossRef]
- Luo, X.; Chen, M.; Shan, H.; Yu, X.; Lin, Q.; Tao, Q.; Wei, X.; Lv, C.; Chen, Z.; Zhuo, F.; et al. Label-free 3d photoacoustic imaging of tumor organoids for volumetric drug screening. Adv. Sci. 2025, 12, e17226. [Google Scholar] [CrossRef]
- Kang, H.; Liu, X.; Ge, D.; Zeng, Y. Revolutionizing bladder cancer research: Harnessing 3d organoid technology to decode tumor heterogeneity and propel personalized therapeutics. Biochim. Biophys. Acta Rev. Cancer 2025, 1880, 189454. [Google Scholar] [CrossRef]
- Hui, W.; Lei, K.; Liu, Y.; Huang, X.; Zhong, Y.; Chen, X.; Wei, M.; Yan, J.; Shen, R.; Mak, P.; et al. Identification and drug screening of single cells from human tumors on semiconductor chip for cancer precision medicine. Adv. Sci. 2025, 12, e2503131. [Google Scholar] [CrossRef] [PubMed]
- Arino, S.; Ferrer-Lorente, R.; Serrano, G.; Zanatto, L.; de la Torre, R.A.M.-G.; Gratacós-Ginès, J.; Rubio, A.B.; Pérez, M.; Mateos-Sánchez, C.; Aguilar-Bravo, B.; et al. Patient-derived liver organoids recapitulate liver epithelial heterogeneity and enable precision modeling of alcohol-related liver disease. J. Hepatol. 2025; online ahead of print. [Google Scholar] [CrossRef]
- Feng, Z.; Zhou, B.; Shuai, Q.; Mu, J.; Xie, J. Applications and limitations of pluripotent stem cell-derived liver organoids. Chin. Med. J. 2025; online ahead of print. [Google Scholar] [CrossRef]
- Zhao, Y.C.; Wang, Z.; Nasser, A.; Sun, A.; Wang, Z.; Zhang, Y.; Ren, J.; Zhao, H.; Yap, N.A.; Wang, Y.; et al. Rapid glass-substrate digital light 3d printing enables anatomically accurate stroke patient-specific carotid artery-on-chips for personalized thrombosis investigation. Adv. Mater. 2025, e08890. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Zhang, Y.; Zhu, Y.; Lee, U.; Luo, A.C.; Li, X.; Wang, X.; Chen, D.; Pu, W.T.; Lin, R.-Z.; et al. Rapid generation of functional vascular organoids via simultaneous transcription factor activation of endothelial and mural lineages. Cell Stem Cell 2025, 32, 1200–1217. [Google Scholar] [CrossRef] [PubMed]
- Clerkin, S.; Singh, K.; Davis, J.L.; Treacy, N.J.; Krupa, I.; Reynaud, E.G.; Lees, R.M.; Needham, S.R.; MacWhite-Begg, D.; Wychowaniec, J.K.; et al. Tuneable gelatin methacryloyl (GelMA) hydrogels for the directed specification of renal cell types for hiPSC-derived kidney organoid maturation. Biomaterials 2025, 322, 123349. [Google Scholar] [CrossRef]
- Kojima, H.; Ishii, T.; Fukumitsu, K.; Ogiso, S.; Tomofuji, K.; Oshima, Y.; Horie, H.; Ito, T.; Wakama, S.; Makino, K.; et al. In vivo regeneration of tubular small intestine with motility: A novel approach by orthotopic transplantation of decellularized scaffold. Transplantation 2023, 107, 1955–1964. [Google Scholar] [CrossRef]
- Bealer, E.; Crumley, K.; Clough, D.; King, J.; Behrend, M.; Annulis, C.; Li, F.; Soleimanpour, S.; Shea, L.D. Extrahepatic transplantation of 3d cultured stem cell-derived islet organoids on microporous scaffolds. Biomater. Sci. 2023, 11, 3645–3655. [Google Scholar] [CrossRef]
- Maceckova Brymova, A.; Rodriguez-Jimenez, F.J.; Konrad, A.; Nemesh, Y.; Thottappali, M.A.; Artero-Castro, A.; Nyshchuk, R.; Kolesnikova, A.; Müller, B.; Studenovska, H.; et al. Delivery of human iPSC-derived RPE cells in healthy minipig retina results in interaction between photoreceptors and transplanted cells. Adv. Sci. 2025, 12, e2412301. [Google Scholar] [CrossRef]
- Sugita, S.; Mandai, M.; Kamao, H.; Takahashi, M. Immunological aspects of RPE cell transplantation. Prog. Retin. Eye Res. 2021, 84, 100950. [Google Scholar] [CrossRef]
- Beilmann, M.; Adkins, K.; Boonen, H.C.M.; Hewitt, P.; Hu, W.; Mader, R.; Moore, S.; Rana, P.; Steger-Hartmann, T.; Villenave, R.; et al. Application of new approach methodologies for nonclinical safety assessment of drug candidates. Nat. Rev. Drug Discov. 2025, 24, 705–725. [Google Scholar] [CrossRef]
- Gu, Y.; Zhang, W.; Wu, X.; Zhang, Y.; Xu, K.; Su, J. Organoid assessment technologies. Clin. Transl. Med. 2023, 13, e1499. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Zheng, Y.; Yang, G.; Zhang, H.; Lu, M.; Ma, H.; Wu, C.; Lu, H. A bioactive calcium silicate nanowire-containing hydrogel for organoid formation and functionalization. Mater. Horiz. 2024, 11, 2957–2973. [Google Scholar] [CrossRef] [PubMed]
- Harris, A.R.; Walker, M.J.; Gilbert, F. Ethical and regulatory issues of stem cell-derived 3-dimensional organoid and tissue therapy for personalised regenerative medicine. BMC Med. 2022, 20, 499. [Google Scholar] [CrossRef] [PubMed]
- Verstegen, M.M.A.; Coppes, R.P.; Beghin, A.; De Coppi, P.; Gerli, M.F.M.; de Graeff, N.; Pan, Q.; Saito, Y.; Shi, S.; Zadpoor, A.A.; et al. Clinical applications of human organoids. Nat. Med. 2025, 31, 409–421. [Google Scholar] [CrossRef]
- Jiang, S.; Zhao, H.; Zhang, W.; Wang, J.; Liu, Y.; Cao, Y.; Zheng, H.; Hu, Z.; Wang, S.; Zhu, Y.; et al. An automated organoid platform with inter-organoid homogeneity and inter-patient heterogeneity. Cell Rep. Med. 2020, 1, 100161. [Google Scholar] [CrossRef]
- Gao, Q.; Wang, J.; Zhang, H.; Wang, J.; Jing, Y.; Su, J. Organoid vascularization: Strategies and applications. Adv. Healthc. Mater. 2025, 14, e2500301. [Google Scholar] [CrossRef]
- Mao, R.; Zhang, J.; Qin, H.; Liu, Y.; Xing, Y.; Zeng, W. Application progress of bio-manufacturing technology in kidney organoids. Biofabrication 2025, 17, 022007. [Google Scholar] [CrossRef]
- Xia, X.; Hu, M.; Zhou, W.; Jin, Y.; Yao, X. Engineering cardiology with miniature hearts. Mater. Today Bio 2025, 31, 101505. [Google Scholar] [CrossRef]
- Shin, T.H.; Noh, J.; Choi, S.; Song, M.; Kang, M.; Song, M.-H.; Heo, R.; Jeon, Y.K.; Kim, S.J.; Kim, S.-J.; et al. Three-dimensional magnetic torque stimulation enhances functional structural maturation in developing human cardiac organoids. Acta Biomater. 2025, 208, 350–361. [Google Scholar] [CrossRef]
- Sun, L.; Duan, Y.; Lan, Q.; Wang, Y.; Wong, A.S.; Ni, Y.; Liu, D. Organoids in host pathology: Unveiling microbial-induced therapeutic responses. Microbiol. Res. 2025, 301, 128313. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Zhou, D.; Wang, F.; Chen, X.; Li, M.; Su, J. Organoids for tissue repair and regeneration. Mater. Today Bio 2025, 33, 102013. [Google Scholar] [CrossRef] [PubMed]
- Septiana, W.L.; Pawitan, J.A. Potential use of organoids in regenerative medicine. Tissue Eng. Regen. Med. 2024, 21, 1125–1139. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Bo, Q.; Wang, C.; Xu, Y.; Fei, X.; Chen, R. Single BMSC-derived cartilage organoids for gradient heterogeneous osteochondral regeneration by leveraging native vascular microenvironment. J. Nanobiotechnol. 2025, 23, 325. [Google Scholar] [CrossRef]
- Dabbagh Moghaddam, F.; Anvar, A.; Ilkhani, E.; Dadgar, D.; Rafiee, M.; Ranjbaran, N.; Mortazavi, P.; Ghoreishian, S.M.; Huh, Y.S.; Makvandi, P. Advances in engineering immune-tumor microenvironments on-a-chip: Integrative microfluidic platforms for immunotherapy and drug discovery. Mol. Cancer 2025, 24, 271. [Google Scholar] [CrossRef]
- Huang, T.; Huang, W.; Bian, Q. Organoids as predictive platforms: Advancing disease modeling, therapeutic innovation, and drug delivery systems. J. Control. Release 2025, 387, 114222. [Google Scholar] [CrossRef]
- Saiki, N.; Nio, Y.; Yoneyama, Y.; Kawamura, S.; Iwasawa, K.; Kawakami, E.; Araki, K.; Fukumura, J.; Sakairi, T.; Kono, T.; et al. Self-organization of sinusoidal vessels in pluripotent stem cell-derived human liver bud organoids. Nat. Biomed. Eng. 2025, 9, 1869–1885. [Google Scholar] [CrossRef]
- Jung, Y.H.; Choi, D.; Park, K.; Lee, S.-B.; Kim, J.; Kim, H.; Jeong, H.-W.; Yang, J.H.; Kim, J.-A.; Chung, S.; et al. Drug screening by uniform patient derived colorectal cancer hydro-organoids. Biomaterials 2021, 276, 121004. [Google Scholar] [CrossRef]
- Mun, S.J.; Hong, Y.; Shin, Y.; Lee, J.; Cho, H.-S.; Kim, D.-S.; Chung, K.-S.; Son, M.J. Efficient and reproducible generation of human induced pluripotent stem cell-derived expandable liver organoids for disease modeling. Sci. Rep. 2023, 13, 22935. [Google Scholar] [CrossRef]
- Bollinger, J.; May, E.; Mathews, D.; Donowitz, M.; Sugarman, J. Patients’ perspectives on the derivation and use of organoids. Stem Cell Rep. 2021, 16, 1874–1883. [Google Scholar] [CrossRef]
- Ravn, T.; Sorensen, M.P.; Capulli, E.; Kavouras, P.; Pegoraro, R.; Picozzi, M.; Saugstrup, L.I.; Spyrakou, E.; Stavridi, V. Public perceptions and expectations: Disentangling the hope and hype of organoid research. Stem Cell Rep. 2023, 18, 841–852. [Google Scholar] [CrossRef]
- Harrison, T.R.; Faber, S.C.; Zare, M.; Fontaine, M.; Williams, M.T. Wolves among sheep: Sexual violations in psychedelic-assisted therapy. Am. J. Bioeth. 2025, 25, 40–55. [Google Scholar] [CrossRef]
- MacDuffie, K.E.; Stein, J.L.; Doherty, D.; Jayadev, S.; Girault, J.B.; Emmons, K.A.; Glass, M.R.; Dempsey, J.C.; Marrus, N.; Botteron, K.N.; et al. Donor perspectives on informed consent and use of biospecimens for brain organoid research. Stem Cell Rep. 2023, 18, 1389–1393. [Google Scholar] [CrossRef]
| Culturing Technologies | Cell Types | Principles | Benefits | Application Fields | References |
|---|---|---|---|---|---|
| 3D cell culture | ASCs/iPSCs | ASCs self-organize; iPSCs differentiate | High physiological relevance; personalized construction | The construction of liver, kidney, and intestinal organoid models; personalized medicine | [15,16] |
| ECM scaffold materials like matrigel simulate the in vivo microenvironment and promote 3D cell self-organization | Provides a near-in vivo extracellular matrix environment and regulates stiffness and biocompatibility | Facilitation of the construction of intestinal and liver organoids and support for cancer research | [17] | ||
| Self-formation of organoids | Replicates the natural development process | Developmental biology | [18] | ||
| Multi-cellular co-culture for simulating intercellular interactions | Enhances organoid function | Immune responses, regeneration, viral infections, and other aspects of organs, e.g., the liver and kidneys | [19] | ||
| 3D bioprinting | Printing layer by layer using bioink | Precisely controls the spatial distribution of cells to construct organ microstructures containing microvessels | Drug screening and regenerative medicine for liver organoids | [20] | |
| Organoid-on-a- chip | A microfluidic organoid chip and microchannels mimicking vascular networks to offer a dynamic microenvironment for enhanced organoid functionality | Culture conditions can be dynamically controlled | Research on drug toxicity and immune regulation; construction of liver and lung organoids | [21] |
| Organoid Types | Cell Sources | Supporting Matrix | Strengths | Limitations | Major Applications | References |
|---|---|---|---|---|---|---|
| ASCOs | ASCs | Matrigel/ECM/synthetic hydrogel | Rapidly established, preserving specific structure/function | Lineage differentiation potential is limited | Adult disease modeling, regenerative medicine | [73] |
| PSCOs | ESCs/iPSCs | Matrigel/ECM/synthetic hydrogel | Pluripotent differentiation, simulate early organ development | Long culture cycle, insufficient maturity | Developmental biology, genetic disease mechanisms, gene editing | [73,74,75] |
| PDOs/PDTOs | Patient- derived tissues | Matrigel | Preserve tumor genetics/microenvironment, high clinical relevance | Lack complete immune microenvironment (insufficient stroma cells) | Personalized tumor therapy, high-throughput drug screening | [44,45,76,77] |
| Assembloids | iPSCs | Matrigel/ECM/synthetic hydrogel | Simulate multi-tissue interactions, cross-tissue signaling | Complex technology, hard to standardize, coordinating developmental rhythm required | Research on tissue interactions, cross-organ signal transduction | [65,66] |
| Organoid-on-a- chip | Cells/tissues | Matrigel/synthetic hydrogel scaffold/chip | Dynamic physicochemical regulation, enhances vascularization/metabolism | High equipment dependence | Drug toxicity testing, permeability prediction, physiological simulation | [68,69,72] |
| Gastruloids | PSCs | Synthetic hydrogel scaffold/biomaterial | Simulate process of gastrulation in early embryos | Incomplete structure and function | Early development simulation, vascularization research | [39,71] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, L.; Chen, A.; Zhang, D.; He, Z. From Cells to Organoids: Approaches, Regulatory Mechanisms, Applications, and Challenges of Organoids. Cells 2025, 14, 1898. https://doi.org/10.3390/cells14231898
Wang L, Chen A, Zhang D, He Z. From Cells to Organoids: Approaches, Regulatory Mechanisms, Applications, and Challenges of Organoids. Cells. 2025; 14(23):1898. https://doi.org/10.3390/cells14231898
Chicago/Turabian StyleWang, Lihong, Anqi Chen, Dong Zhang, and Zuping He. 2025. "From Cells to Organoids: Approaches, Regulatory Mechanisms, Applications, and Challenges of Organoids" Cells 14, no. 23: 1898. https://doi.org/10.3390/cells14231898
APA StyleWang, L., Chen, A., Zhang, D., & He, Z. (2025). From Cells to Organoids: Approaches, Regulatory Mechanisms, Applications, and Challenges of Organoids. Cells, 14(23), 1898. https://doi.org/10.3390/cells14231898





