Abstract
In order to overcome the instability of natural glutathione peroxidase (GPx), scientists endeavor to produce GPx mimics. The popular method first uses biological imprinting (BI) to produce the substrate binding sites and then employs chemical mutation (CM) to obtain the catalytic site. However, BICM has a drawback in that the catalytic site is not clear. Some researchers therefore tried to change the order of the method. These new GPx mimics were prepared by first producing the catalytic site through chemical mutation, and then employing biological imprinting to produce the substrate binding sites (CMBI). It has a clear catalytic site, but its determination of enzyme activity and kinetic analysis are still not elucidated. In this study, we used CMBI to synthesize a GPx mimic using trypsin as the imprinted molecule and GSSG as the template molecule and compared the enzyme activity of the four intermediates (Trypsin-SeO2H (TSeO2H), Trypsin-Se-SG (TSeSG), Imprinted Trypsin-Se-SG (ITSeSG), Cross-linked Imprinted Trypsin-Se-SG (CITSeSG), we analyzed the properties of intermediate products. All values are the means of at least four determinations, ITSeSG was produced from TSeSG through bio-imprinting, the activity of GPx mimics synthesized by CMBI was 5.7 times greater than native GPx, because of bio-imprinting make KmGSH value of the mimics decreased from 4.82 ± 0.27 mM (TSeSG) to 0.52 ± 0.05 mM (ITSeSG). This proves that bio-imprinting is the reason for increased substrate binding capability.
1. Introduction
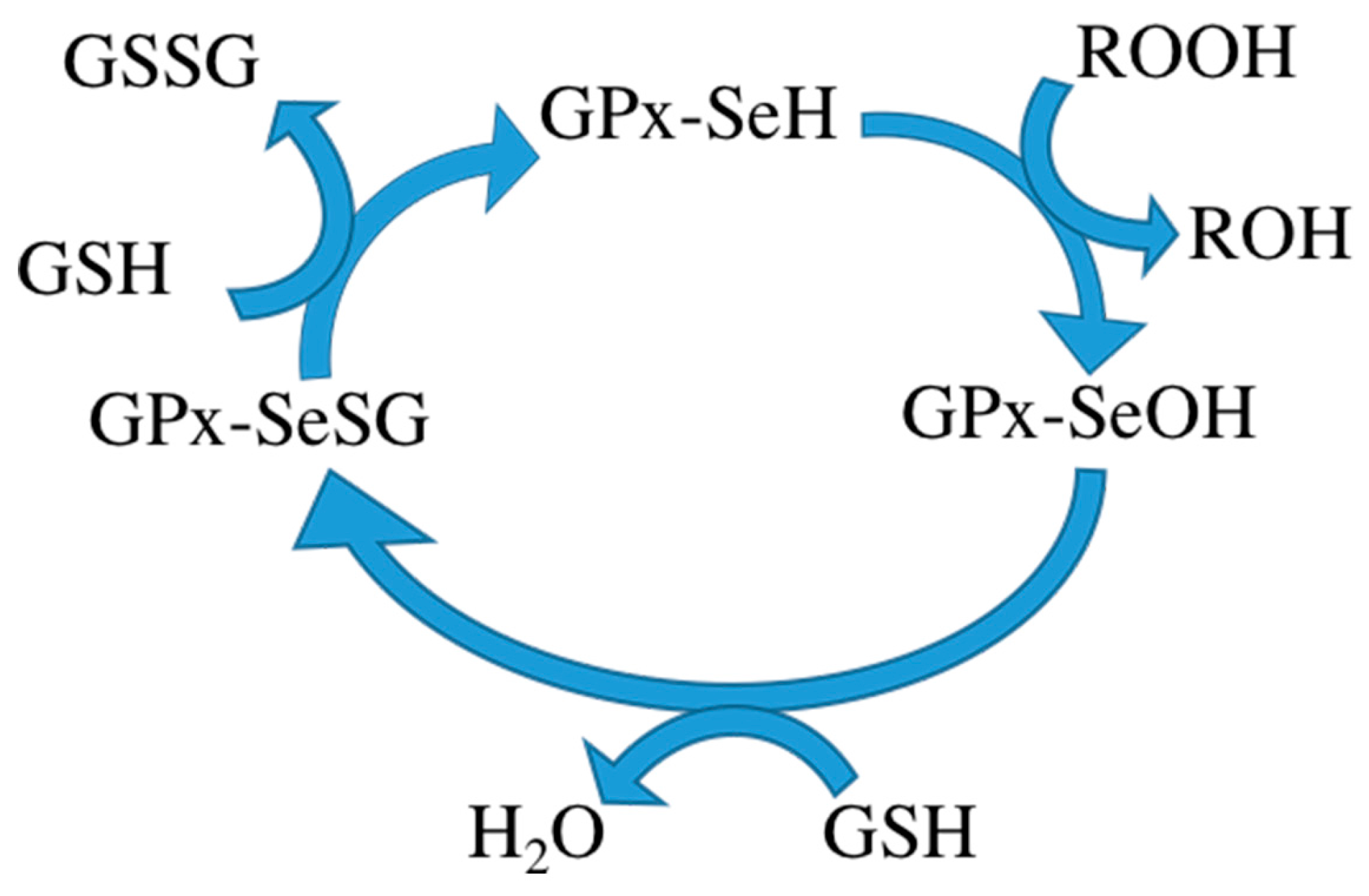
During metabolism, toxic reactive oxygen species (ROS) are produced in the cell. Some ROS can be hydrolyzed by enzymes in the body [1,2]. However, excessive ROS can cause oxidative stress, resulting in damage to many important biological macromolecules, potentially inflicting disease. Mammalian cells have a defense mechanism that prevents excessive accumulation of ROS by maintaining the balance between ROS production and degradation. Glutathione peroxidase (GPx) is one of the most important antioxidants in this defense mechanism [3]. It consists of four subunits, each subunit containing a glutathione binding site and a selenocysteine. Glutathione binding sites are responsible for binding oxidized glutathione and selenocysteine is the catalytic activity center, converting oxidized glutathione to its reduced form. Reduced glutathione in turn interacts with intracellular free radicals, thereby becoming oxidized again but detoxifying the radicals and thus protecting the body from harm of free radicals. Some researchers therefore want to use GPx to treat certain diseases. The mechanism of the GPx contains three steps: (1) oxidation of the selenol form of Sec (GPx-SeH) into a selenenic intermediate (GPx-SeOH), with simultaneous reduction of one equivalent of hydroperoxide to produce water/alcohol; (2) reduction of GPx-SeOH upon reaction with one equivalent of glutathione (GSH), thus producing water and a selenenyl sulfide intermediate (GPx-SeSG); and (3) reduction of GPx-SeSG by reaction with a second equivalent of GSH to produce oxidized glutathione (GSSG), thus resulting in the regeneration of the initial configuration of the enzyme (Scheme 1) [4,5,6].
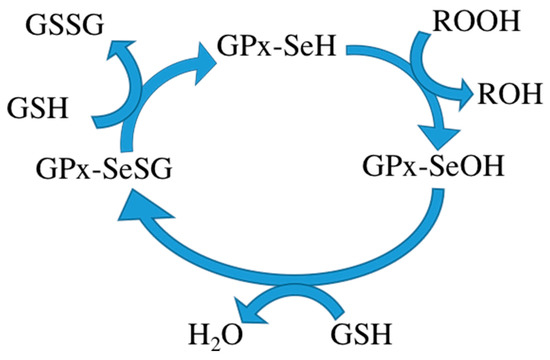
Scheme 1.
The mechanism of the GPx.
However, GPx is unstable, rare, and has a high molecular weight, making it unfavorable for therapeutic use. In order to retain the GPx antioxidant capabilities while overcoming these shortcomings, scientists have produced active selenocysteine GPx mimics by using other macromolecules [7,8,9,10].
To date, many approaches have been tested to produce GPx mimics. These include the use of chemical macromolecules as host such as cyclodextrin, dendrimers, and hyperbranched polymers. Natural enzymes have also been used as host such as semi-synthetic selenoenzymes. Here, genetic engineering technology is used to produce the selenoenzymes. Some researchers also produce GPx mimics using monoclonal antibody, nano-micelles, macrocyclic compound scaffolds, and molecular imprinted polymers [11]. Molecular imprinting is one of the most popular methods to synthesize GPx mimics. Prior to molecular imprinting, chemical mutation [3] (chemical mutation is the method that enables the substrate to obtain specific function through engineering the specificity of existing enzymes) was commonly used. Hilvert et al. (1989) used chemical mutation for the first time, converting serine to selenocysteine [12]. Most researchers used chemical mutation to transform macromolecules (cyclodextrins, dendrimers, hyperbranched polymers, etc.) and subsequently introduced catalytic groups to obtain GPx activity mimics. Although widely used, there are still some shortcomings. Ren et al.’s study found that chemical methods provided a means for introducing into enzymes diverse functions that did not occur naturally and could not be easily incorporated by genetic engineering [13]. Some researchers therefore tried to synthesize GPx mimics by screening for antibodies or single-chain variable fragments with high substrate specificity, and then used chemical mutation. Similarly, others used the substrate binding sites of natural enzymes followed by chemical mutation to obtain GPx mimics. Although these synthetic approaches give rise to GPx mimics with high substrate binding capacity, the locations of their active sites are not clear and therefore impede detailed studies of enzymatic reaction kinetics. Ren et al. introduced selenium by chemical mutagenesis in the experiment, and the selenium content of Se-OVA (egg albumin) (OVA containing 38 serine residues) and Se-BSA (bovine serum albumin) (BSA containing 28 serine residues) was 5.6 mol, 5.2 mol, respectively. So it is not clear which selenium sites were introduced selenium. Zheng et al. [14] utilized chemical mutation to make a GPx mimic from human glutathione transferase Zeta (hGSTZ1-1). Subsequent analysis of its crystal structure showed that there were two serine (Ser 14 and Ser 15) in its active site and they played an important role in the binding and orientation of glutathione (GSH). Hence, they had to compare the activity between Ser 14 and Ser 15 in order to understand the enzymatic kinetics. It is clear that chemical mutation, used to create GPx mimics, can make research more cumbersome due to the ambiguities of the catalytic sites. Advances in the research meant that GPx mimics could now be produced using either one of two methods that rely on bio-imprinting and chemical mutation.
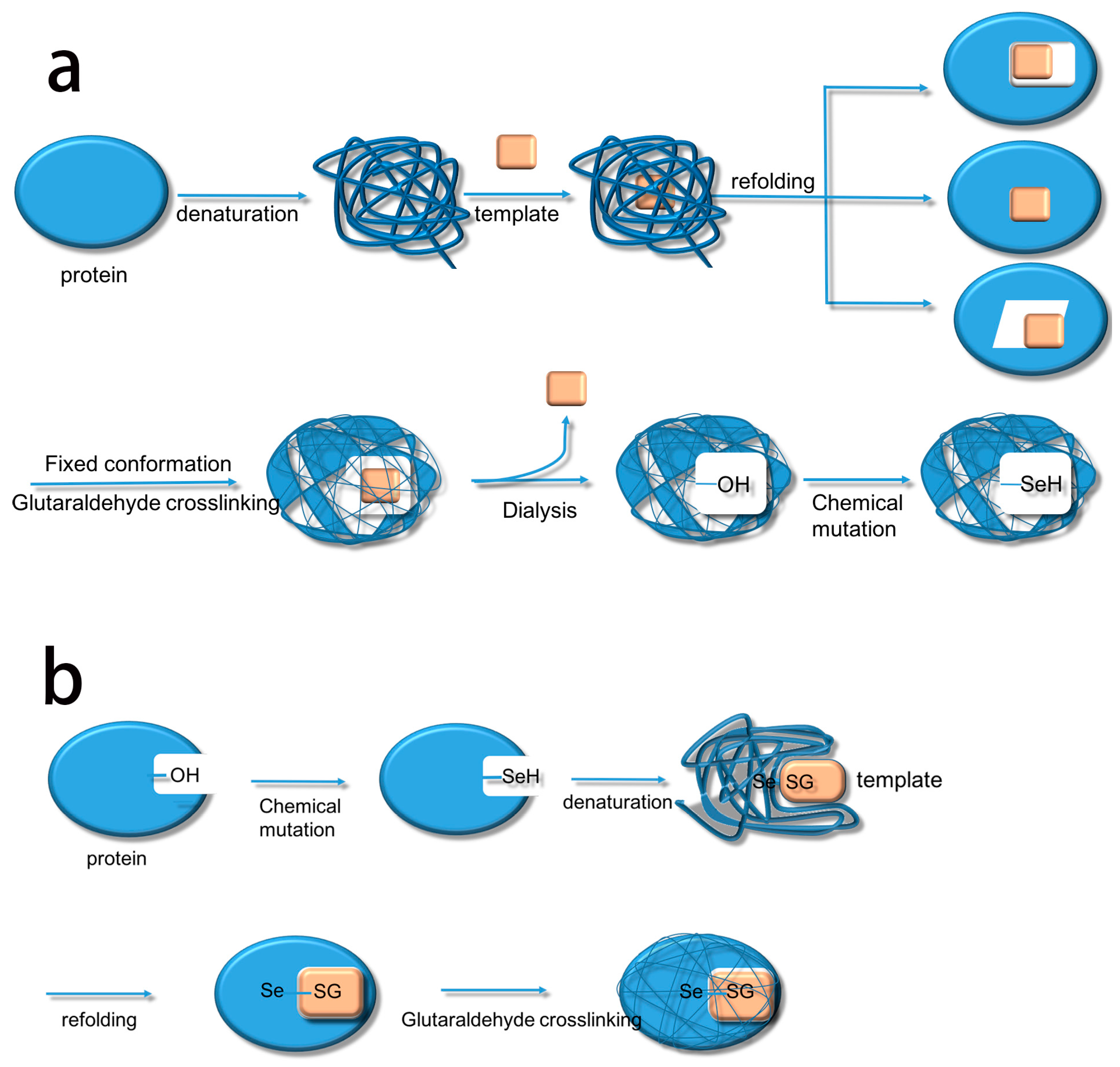
Based on this, we summarize the two methods. The first method uses biological imprinting to produce substrate binding sites and then produces a catalytic site by chemical mutation (BICM) (Scheme 2a) [15,16]. The second method first produces a catalytic site through chemical mutation and then introduces the substrate binding sites by biological imprinting (CMBI) (Scheme 2b). After several experiments, it became clear that BICM has some disadvantages: the location of the binding sites was not always clear at the time of chemical mutation, and the locations did not necessarily have a suitable hydroxyl group, meaning that the catalytic activity did not always increase. For instance, Liu et al. [15] used BICM to synthesize a GPx mimic using albumin as the imprinted molecule. The selenium content of the active site was quantitatively determined by 2-nitrobenzoic acid. The results showed that the ratio of the mimic containing selenium to the selenium content was not the same, which implied that BICM did not allow selenium to specifically connect to the active site, resulting in a waste of selenium. At the same time, it also made the active center of the mimic structure unclear and affected the reaction rate [15]. The CMBI method was also used by Liu et al. [1], this time used subtilisin as the imprinted molecule. Quantitative determination of selenium in the active site of the enzyme showed that the ratio of the imprinted selenosubtilisin in the active site to the selenium content of the system was 1:1. CMBI thus greatly improved the utilization of selenium compared with BICM, and made the active center structure clearer to improve the catalytic efficiency. CMBI therefore overcame the drawbacks of BICM but, since the method was reported [17,18], there have been no further reports using CMBI to produce GPx mimics. This was unexpected given the improvements and suggested that there may have been further complications. In this study, we therefore used the CMBI method to introduce selenium sites with trypsin as the imprinted molecule for biological imprinting and produced GPx mimics. Previously, we already determined that introducing the catalytic group of selenated trypsin through chemical mutation made a special kind of GPx mimic with a clear catalytic site. Herein, we elucidate the activity and kinetic parameters of TSeO2H (Trypsin-SeO2H), TSeSG (Trypsin-Se-SG), ITSeSG, and CITSeSG to further support our previous research.
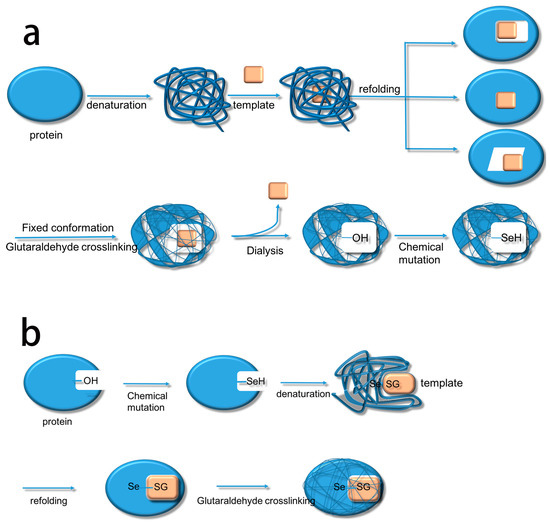
Scheme 2.
The flow charts of the two methods used to make GPx activity mimics: (a) BICM; and (b) CMBI.
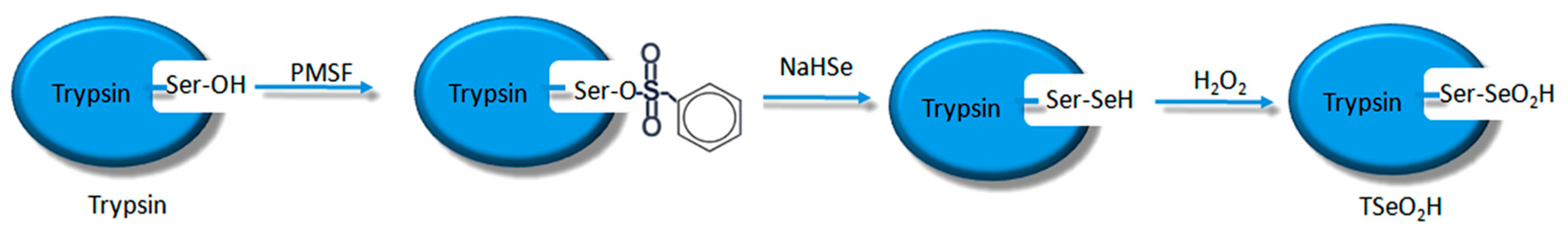
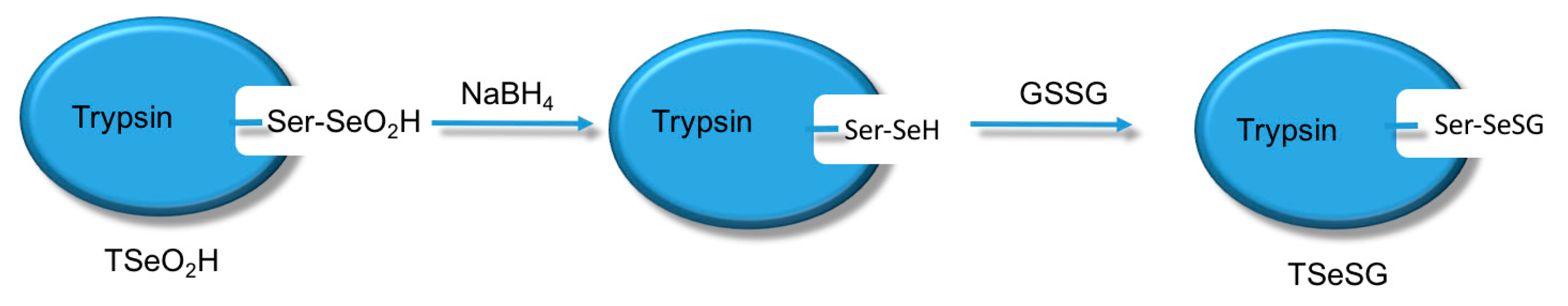
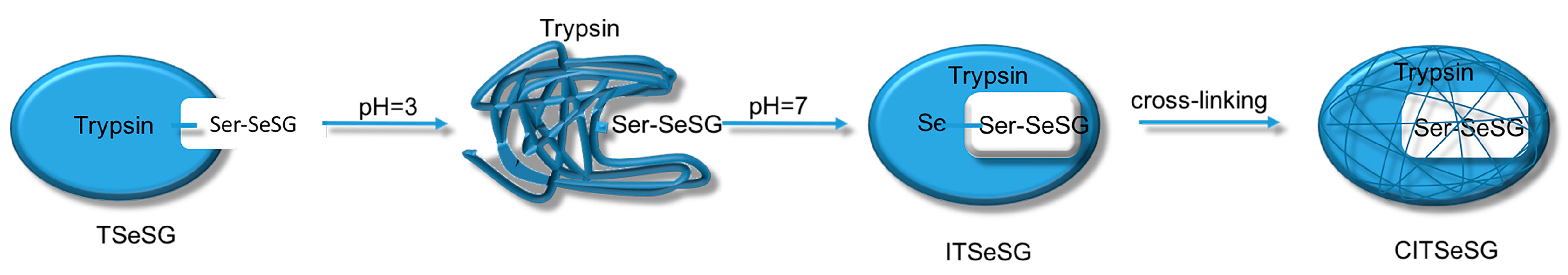
The process of synthesizing molecular imprinted polymers can be divided into three steps: (1) template molecules and functional monomers are combined; (2) imprinting molecules are polymerized with complexes formed by template molecules and functional monomers; (3) template molecules are removed from the polymer [19]. Scientists have already demonstrated that bio-imprinting, an integral part of molecular imprinting, is an effective means to introduce binding sites into proteins or other biomolecules. In this study, imprinting molecules were synthesized by bio-imprinting, and the GPx mimic with high catalytic activity and a clear catalytic site was prepared using the above three steps. In our paper, we synthesized GPx mimic through the following route: the hydroxyl group of Ser was site-specifically activated by the addition of phenylmethanesulfonyl fluoride (PMSF) to form a sulfonylated trypsin. Then selenium was introduced into the active site by reaction of the sulfonated trypsin with NaHSe. Adding H2O2 subsequently to the resulting selenol form of the trypsin yields the seleninic acid form of TSeO2H. Then through adding NaBH4 and GSSG (template molecule) TSeO2H changes into TSeSG, the process is that the S–S of GSSG combines with the SeH group of trypsin and synthesizes a Se–S bond. This process leads GSH to combine with trypsin in the form of Se–S bond. Finally through the change of pH, imprinted molecules stretch and refold to let GSH wrap in the trypsin. The product of this process called ‘ITSeSG’. After crosslinking of glutaraldehyde, ITSeSG changes into CITSeSG.
2. Results and Discussion
2.1. Preparation and Characterization of Mimics
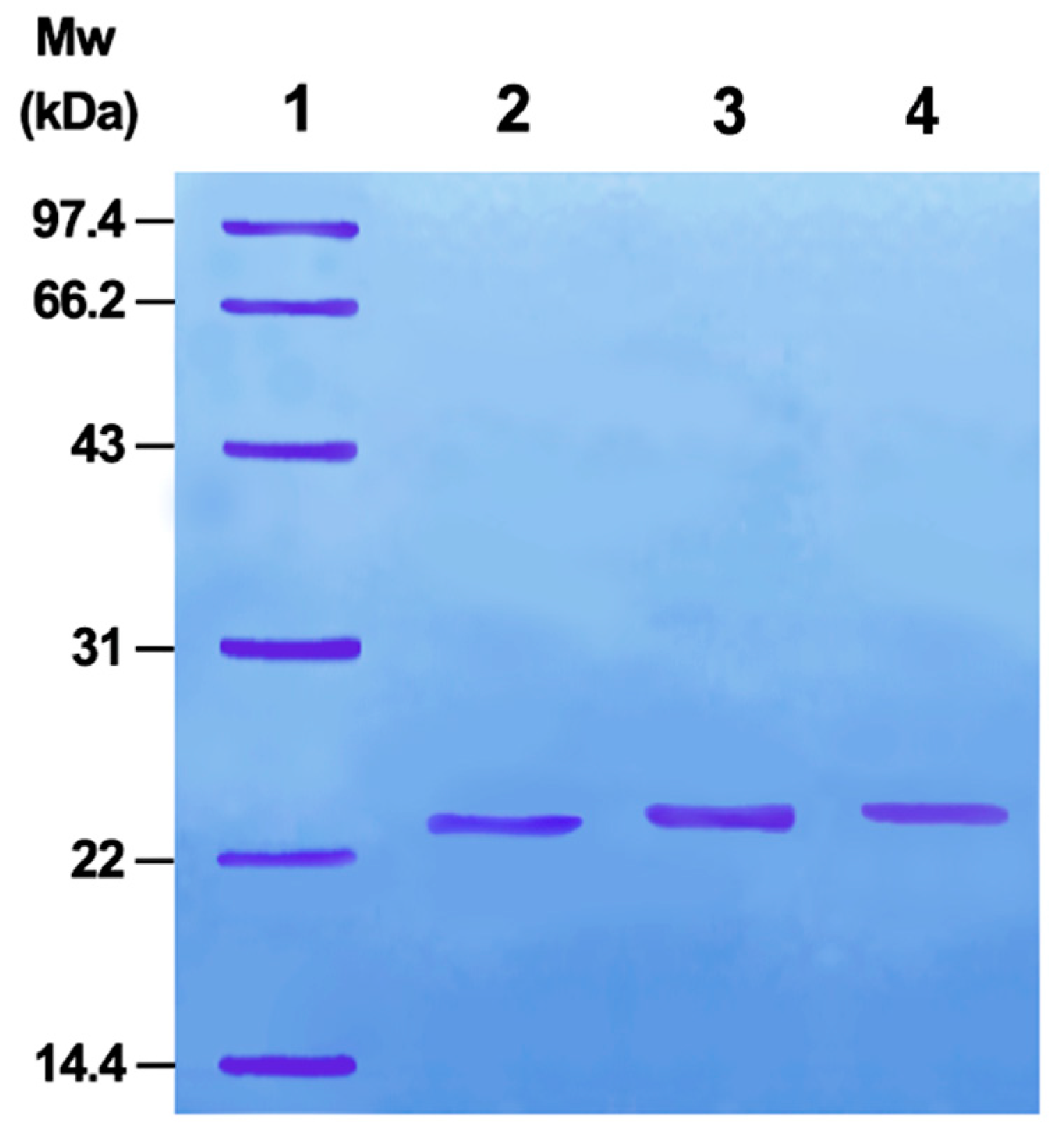
According to Luo [20], PMSF addition can activate the serine side chain -OH in the trypsin active site. Further, activated byproducts do not affect the following selenide substitution reaction. Therefore, once activated, the selenide substitution reaction can directly occur without purification. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) indicated that the product is a single protein (Figure 1). Selenoprotein (TSeO2H) interacted with DTNB and the absorbance of the new product was measured, indicating 0.96 SeH per TSeO2H. Trypsin activity of TSeO2H was determined using N-benzoyl-l-tyrosine ethyl ester as substrate (PBS, pH 7.4). TSeO2H completely abolished trypsin activity, illustrating that the serine side chain -OH in the active site of trypsin had essentially translated into SeO2H with GPx catalytic activity.
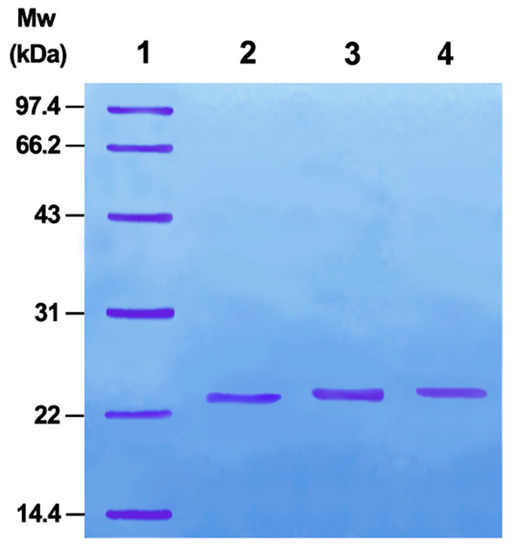

Figure 1.
Discontinuous SDS-PAGE analysis of mimics. (1) Marker proteins; (2) TSeO2H; (3) TSeSG; (4) ITSeSG. SDS-PAGE was performed on a Pharmacia gel system following standard procedures.
Biological imprinting technologies have improved the imprinting capabilities of molecules. By adjusting the solution to pH 3.0, trypsin was first partially stretched and its amino acid structure was unfolded. The solution was then slowly adjusted to pH 7.0 to allow for molecular recognition of the amino acids. This induced the conformation with -Se-SG [1] so that the protein folded again with a substrate binding site, thereby obtaining ITSeSG.
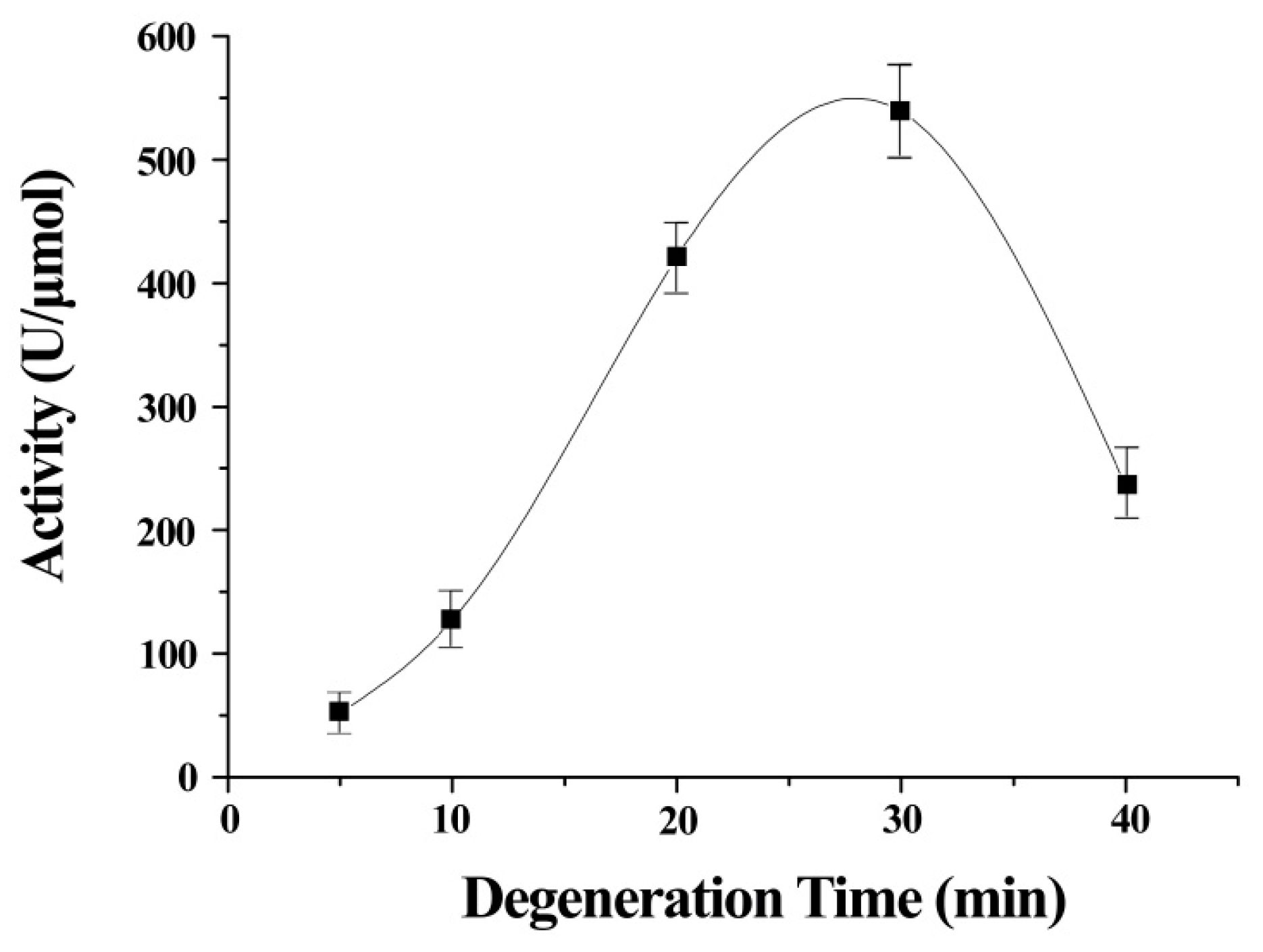
The stretch time was an important contributor to the activity of ITSeSG (Figure 2). This is possibly because the stretch time is positively correlated with the extent of protein stretch. A shorter stretch time may therefore lead to inadequate protein unfolding and thus less interaction with -Se-SG and omitting the substrate binding site. Decreased GPx activity was indicative of excessive ITSeSG degeneration (Figure 2). Therefore, we inferred that a partially stretched protein creates a suitable binding site and that this did not affect the formation of catalytic intermediates. The experimental results showed that a stretch time of 20 to 30 min was needed to reach the highest GPx activity.
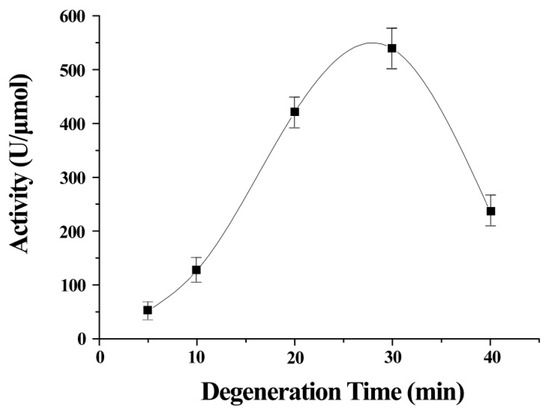
Figure 2.
Relationship between GPx activity and the degeneration time of ITSeSG. (All values are the means of five determinations).
Error bars represent the mean ± standard error (n = 4).
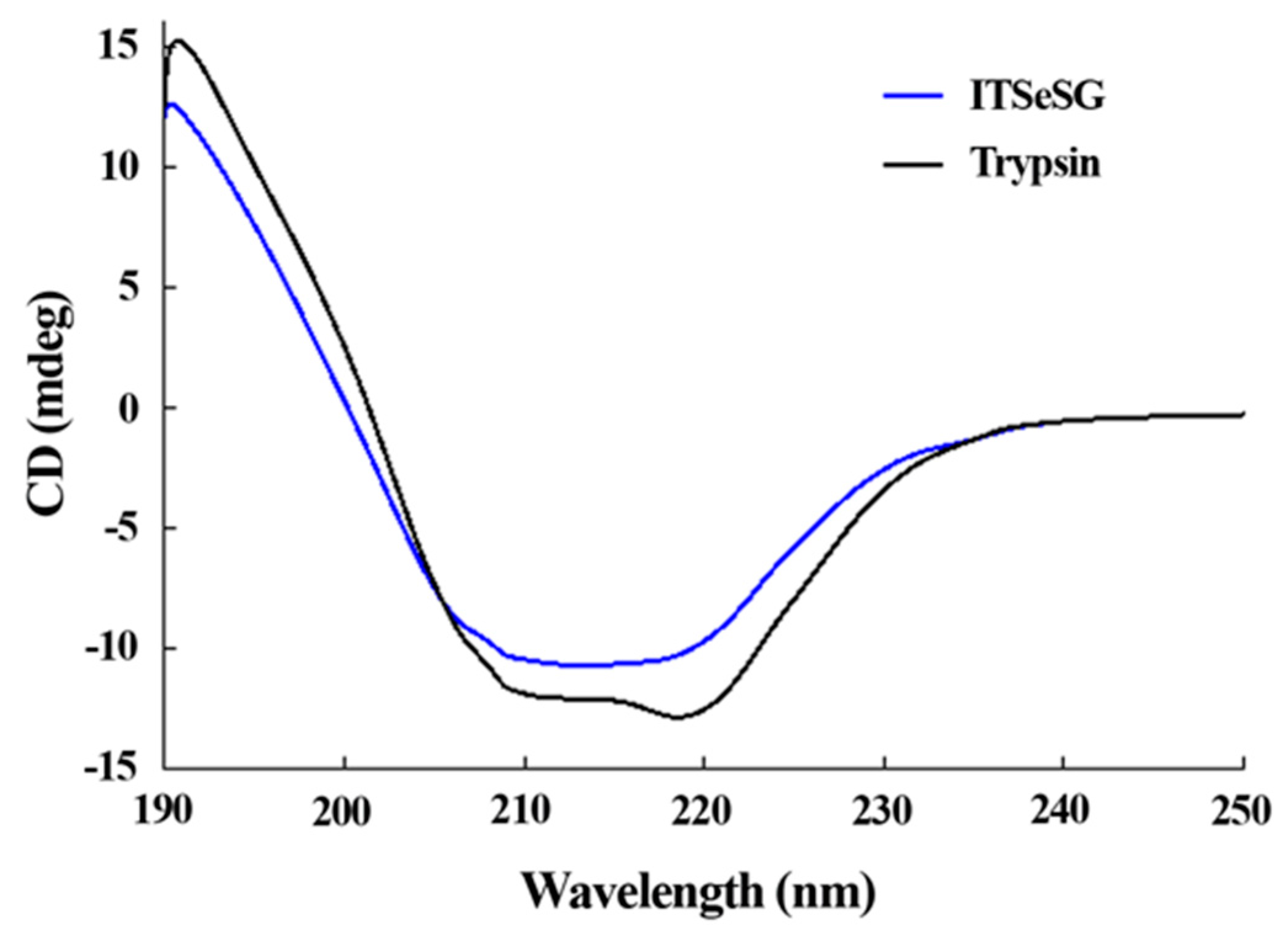
The changing of the secondary structure of trypsin and ITSeSG was detected by circular dichroism spectra (Figure 3).
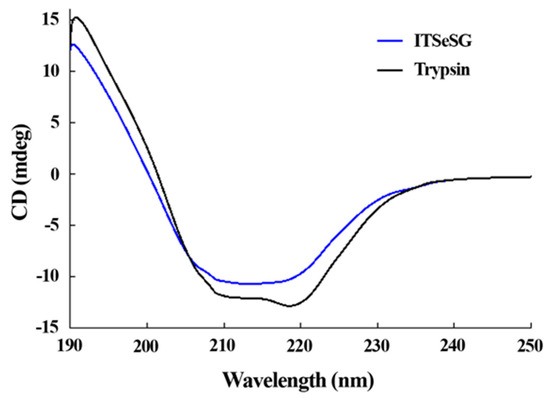
Figure 3.
Circular dichroism spectra of ITSeSG (blue) and Trypsin (black) monitored between 190 and 260 nm.
The proportion of α-helices and β-folds was 19.2% and 48.8% for trypsin, and 16.3% and 39.39% for ITSeSG, respectively. The proportion of loose structure changed from 27.3% (trypsin) to 36.3% (ITSeSG) (Table 1).

Table 1.
ITSeSG and trypsin secondary structure. Data were obtained from the plots in Figure 3.
2.2. The GPx Activity of Mimic
GPx activities of the GPx mimics catalyzed the reduction of H2O2 by GSH and was determined as a change in NADPH absorption at 340 nm (Equations (1) and (2)).
The trypsin active site serine was replaced with selenium cysteine and a catalytic site was introduced to get TSeO2H by chemical mutation. As expected, because TSeO2H had no substrate GSH binding sites, it showed low GPx activity. When transformed to ITSeSG, GPx activity increased 10-fold from 51 U μmol−1 (TSeO2H) to 536 U μmol−1 (ITSeSG). Although the activity of ITSeSG has greatly improved using this technique, it is still very different from natural GPx. GPx activity of ITSeSG is 10.8-fold less than that of native GPx (5780 U μmol−1 from rabbit liver), and is 540 times more active than that of the selenoorganic compound 2-phenyl-1,2-benzoisoselenazol-3(H)-one (Ebselen). The result showed in the Table 2.

Table 2.
Comparison of GPx activities of the GPx mimic-catalysed reduction of H2O2 by GSH and native GPx.
2.3. Kinetic Properties of Mimic
The steady state kinetics was observed for substrates H2O2 and GSH. The initial velocities for reduction of H2O2 by GSH were determined as a function of substrate concentration at 37 °C and pH 7.0, varying one substrate concentration while another was fixed. The relevant steady-state equation for the mimic reaction is shown as Equation (3) [22].
where v0 is the initial reaction rate, [E]0 is the initial enzyme mimic concentration, kmax is a pseudo-first-order rate constant and KH2O2 and KGSH are the Michaelis–Menten constants (Km) for the H2O2 and GSH, respectively.
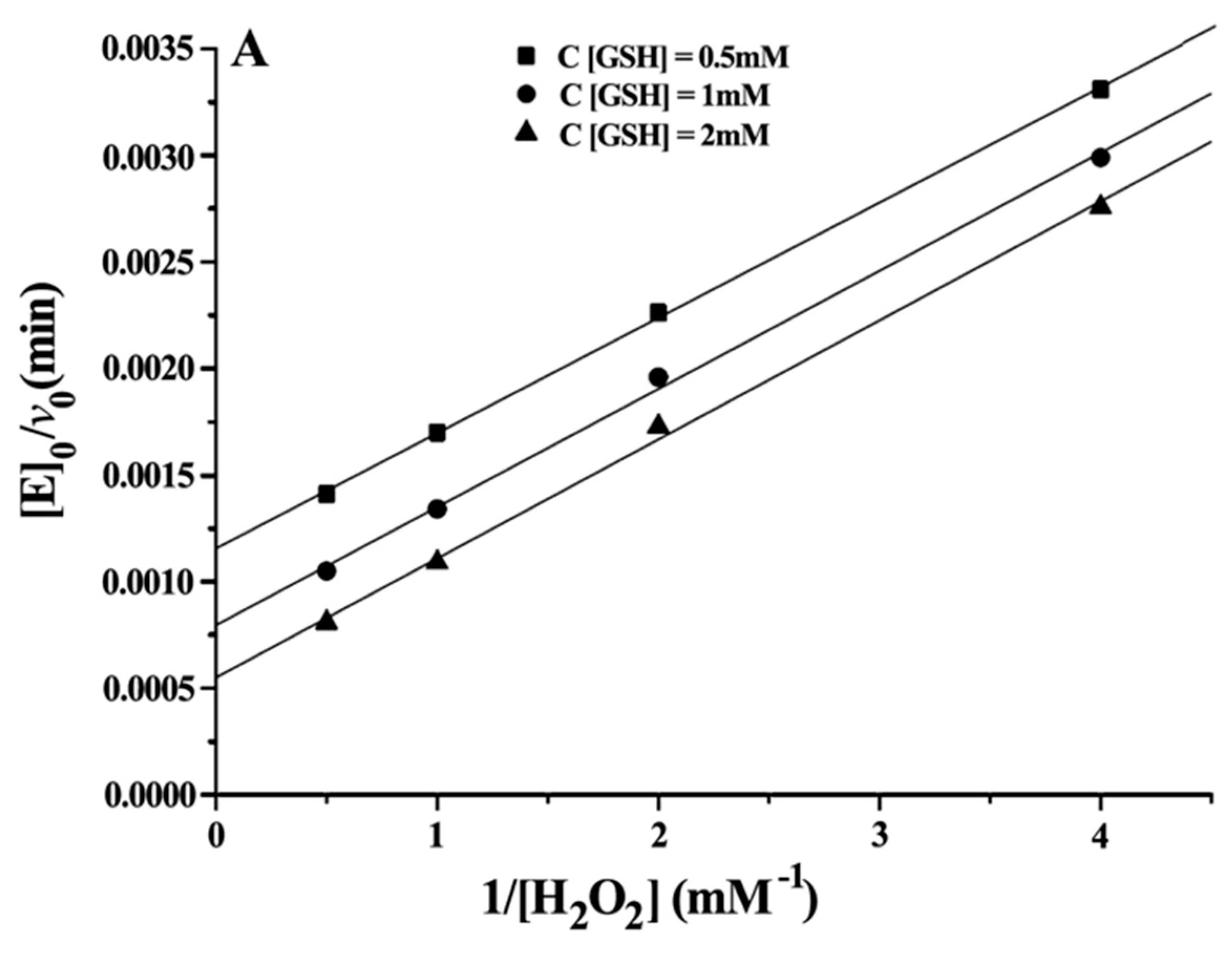
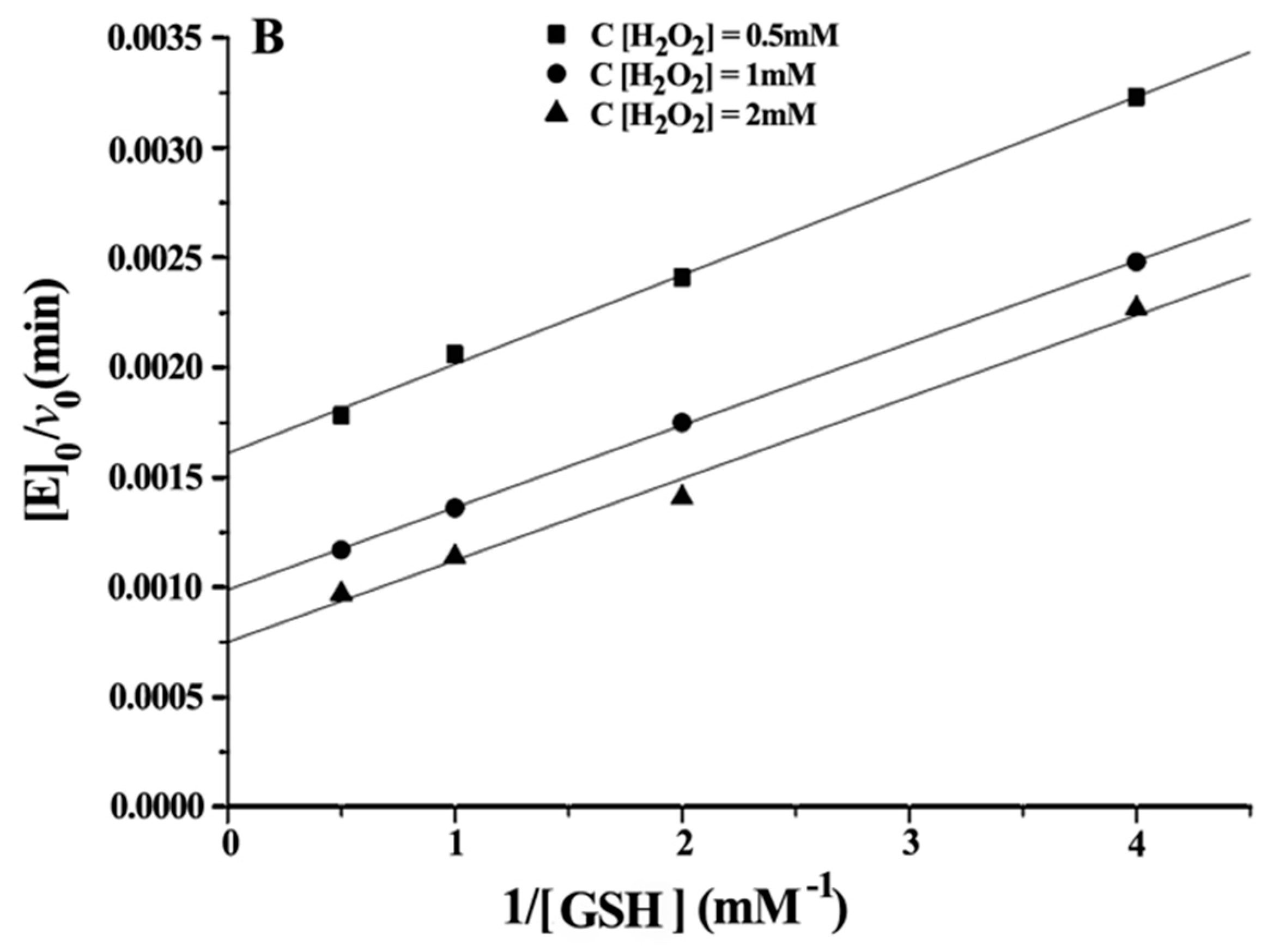
Double reciprocal plots of initial velocity versus the concentration of substrates gave a family of parallel lines (Figure 4), indicating that the reaction mechanism belongs to the ping-pong mechanism. This result demonstrated that the GPx mimic, has the same catalytic mechanism as that of native GPx. From the steady-state equation, the kinetic parameters were obtained (Table 3).
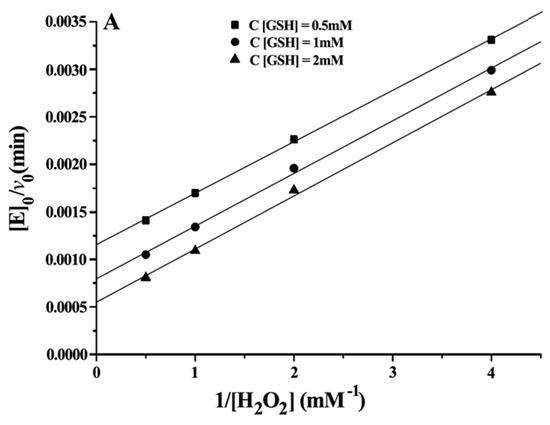
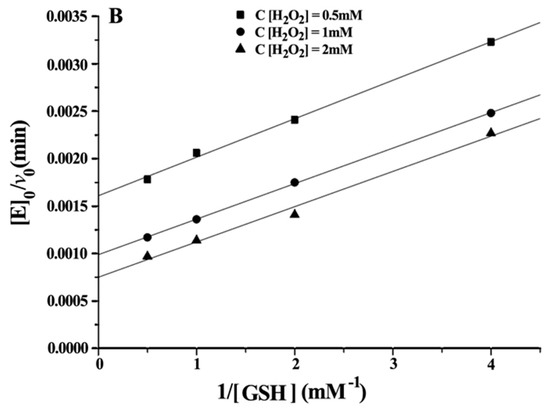
Figure 4.
Double reciprocal plots for the reduction of H2O2 by GSH catalysed by 5 μM CITSeSG. (A) [E]0/v0 versus 1/[H2O2] (mM−1) at [GSH] 0.5 mM (■), 1 mM (●) and 2 mM (▼); (B) [E]0/v0 versus 1/[GSH] (mM−1) at [H2O2] 0.5 mM (●), 1 mM (▲) and 2 mM (▼).

Table 3.
Kinetic parameters of GPx mimics.
With H2O2 as the substrate: kmax·Km−1 H2O2 (ITSeSG) > kmax·Km−1 H2O2 (CITSeSG) > kmax·Km−1 H2O2 (TSeSG) > kmax·Km−1 H2O2 (TSeO2H). ITSeSG had the fastest reaction rate to H2O2, and the reaction rate of TSeO2H to H2O2 was the slowest. The KmH2O2 exhibited by the four mimics is close to that of the substrate H2O2 because the H2O2 molecule is very small; it freely diffuses into the binding site and is less susceptible to the spatial structure of the mimic.
With GSH as the substrate: Km GSH (TSeSG) > Km GSH (TSeO2H) > Km GSH (CITSeSG) > Km GSH (ITSeSG) can be found by comparing the kinetic parameters of the four intermediates. Similarly, kmax·Km−1 GSH (ITSeSG) > kmax·Km−1 GSH (CITSeSG) > kmax·Km−1 GSH (TSeSG) > kmax·Km−1 GSH (TSeO2H). Since the Km value is a coefficient related to substrate affinity, a large Km value indicates low affinity, whereas a small Km value is indicative of high substrate affinity. Therefore, it can be concluded that TSeSG has the lowest affinity for GSH, simply because TSeSG does not have a GSH binding site and can therefore not easily interact with GSH. GSH substrate affinity of TSeSG is similar to TSeO2H. However, due to its -SeSG structure, TSeSG directly enters into the catalytic cycle and the kmax showed a significant increase. The -SeOOH in the TSeO2H molecule must be reduced from GSH to -SeOH or -SeSG to enter the catalytic cycle and thus kmax is the lowest. ITSeSG has the highest GSH substrate affinity. ITSeSG has a GSH binding site and a flexible molecular structure, allowing for the strong binding of GSH. CITSeSG also has a substrate binding site, but due to the glutaraldehyde cross-linking, has a less flexible molecular structure than ITSeSG; moreover, the cross-linked structure will to some extent also affect the release of GSH substrate, and the transformation number was also higher.
The data confirm that the GPx mimic synthesized by molecular imprinting is very effective. Further, the affinity of CITSeSG was only slightly decreased from ITSeSG, suggesting that cross-linking of glutaraldehyde affects GPx activity.
3. Experimental Section
3.1. Materials
Trypsin (from bovine pancreas), phenylmethanesulfonyl fluoride (PMSF), oxidized glutathione (GSSG), 5,5-dithiobis(2-nitrobenzoic acid) (DTNB), and reduced nicotinamide adenine dinucleotide phosphate (NADPH) were purchased from Sigma-Aldrich Sephadex G-25 was obtained from Pharmacia Biotech Inc. (Shanghai, China). All other chemicals were of the highest purity commercially available and were used without further purification.
3.2. Preparation of Trypsin-SeO2H (TSeO2H)
NaHSe was synthesized [23] as follows: 4NaBH4 + 2Se + 7H2O→2NaHSe + 14H2↑ + Na2B4O7. Trypsin (238 mg) was dissolved in phosphate buffer (25 mL, 50 mmol L−1, pH 7.0). 240 μL acetonitrile solution (20 mg mL−1 PMSF) [1] was added slowly and the reaction was kept at 25 °C for 3 h. The reaction mixture was then bubbled through with pure nitrogen gas for 40 min, and then 1.5 mL NaHSe (1 mol L−1) solution was added with a syringe. The mixture was kept for 36 h at room temperature without air. To oxidize it, the mixture was kept at 4 °C for 5 h in air. After centrifugation to remove the solid selenium, concentrated supernatant was desalted using Sephadex G-25 equilibrated in phosphate buffered saline (PBS) (50 mmol L−1, pH 7.0). Next, the protein solution was dialyzed in PBS (10 mmol L−1 H2O2, pH 7.0) at 4 °C for 4 h. The -SeH was oxidized to form -SeO2H, and concentrated by dialysis in PBS (10 mmol L−1, pH 7.0) at 4 °C for 4 h. The dialysis was repeated three times before freeze-drying the final solution. Synthesis of TSeO2H is shown in Scheme 3.
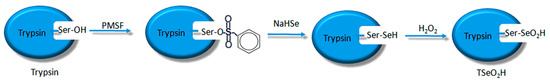
Scheme 3.
Synthesis route of TSeO2H.
3.3. Preparation of Trypsin-Se-SG (TSeSG)
TSeO2H (40 mg) was dissolved in PBS (40 mL, 10 mmol L−1, pH 7.0) and bubbled through with nitrogen gas for 30 min. NaBH4 solution (2 mL, 2 mg mL−1, used within six hours after being configured under sterile sealing conditions) was then added and the reaction was kept at room temperature for 25 min. The solution was slowly added to degassed HCl in order to quench the excessive unreacted NaBH4. At this time, the solution produced a lot of bubbles. Once bubble formation stopped, the solution was adjusted to pH 7.0 with degassed imidazole solution. The above steps reduced -SeO2H to -SeH, ready for the preparation of selenium sulfur compounds. GSSG (40 mL, 40 mmol L−1) was subsequently added and the mixture was kept for 2 h without air. When the reaction was complete, ultrafiltration of the solution ensured that any unreacted GSSG was quenched. Finally, the solution was freeze-dried. Synthesis of TSeSG is shown in Scheme 4.
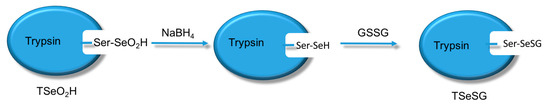
Scheme 4.
Synthesis of TSeSG.
3.4. Preparation of Imprinted Trypsin-Se-SG (ITSeSG) and Cross-Linked Imprinted Trypsin-Se-SG (CITSeSG)
TSeSG (20 mg) was dissolved in PBS (10 mmol L−1, pH 7.0). The solution was carefully adjusted to pH 3.0 with 0.1 M HCl and stirred at 4 °C for 30 min at 200 revolutions min−1 using an electromagnetic stirrer. Next PBS (50 mmol L−1, pH 7.0) was used to adjust the solution to pH 7.0. After resting the solution for 24 h, it was desalted using Sephadex G-25 and the protein peak solution was freeze-dried, thereby obtaining the imprinted enzyme dry powder (ITSeSG). PBS (10 mmol L−1, pH 7.0) was again used to adjust the pH to 7.0. By adding 200 μL glutaraldehyde solution (1%), ITSeSG was cross-linked to fix the new conformation of the protein (CITSeSG). Also, CITSeSG was freeze-dried. Synthesis of ITSeSG and CITSeSG is shown in Scheme 5.
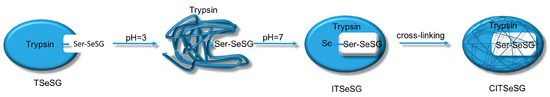
Scheme 5.
Synthesis of CITSeSG.
3.5. Determination of Selenium Content of Mimic
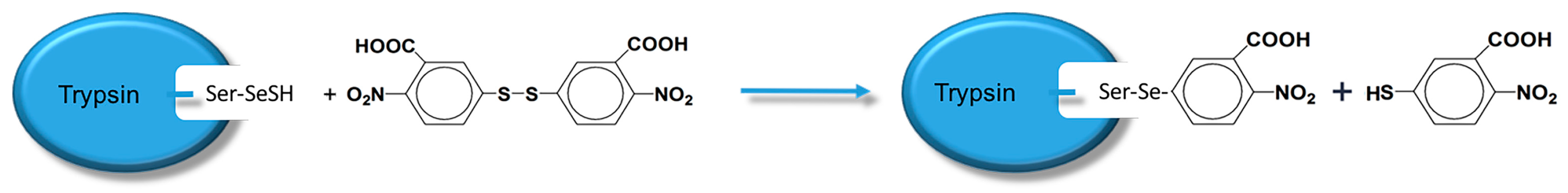
Mimic (10 mg) was dissolved in 5 mL PBS (10 mM pH 7.0), the mixture was bubbled through with pure nitrogen gas for 20 min and an excess of NaBH4 was added. The solution was kept for 20 min before adding degassed 1 M HCl in order to quench the reducing NaBH4. The solution was eventually adjusted to pH 7.0 with 1 M degassed imidazole solution [24]. The selenium content was determined by measuring the absorption of 3-carboxyl-4-nitrobenzenethiolate at 410 nm (ε = 11,400 M−1 cm−1, pH 7.0) of a reaction mixture of 50 μL reduced mimic, 950 μL PBS (50 mM, pH 7.0), and 200 μM DTNB. The absorption equation is shown in Scheme 6.

Scheme 6.
The absorption equation used to measure the selenium content of the GPx mimic.
3.6. Determination of GPx-Like Activity
The catalytic activities were determined according to Wilson et al. [21]. The reaction was carried out at 37 °C in 700 μL of solution containing 50 mM PBS (pH 7.0), 1 mM EDTA, 1 mM sodium azide, 1 mM GSH, 0.25 mM NADPH, 1 unit of glutathione reductase, and 5 μM of mimic. The reaction was initiated by addition of 0.5 mM H2O2. The activity was measured as a decrease of NADPH absorption at 340 nm (ε = 6220 M−1 cm−1). Background absorption of the non-catalytic reaction was measured without mimic and subtracted. The activity unit of the enzyme mimic was defined as the amount of enzyme mimic, which utilizes 1 μmol of NADPH per minute. The concentration of mimic was determined by measuring the UV absorbance at 280 nm and pH 7.0 (ε = 23,500 M−l cm−l) [25].
3.7. Kinetics of the Mimic
The assay to assess the kinetics was similar to that for native GPx [4]. The initial rates for reduction of H2O2 by GSH were determined by observing the change of NADPH absorption at 340 nm at 37 °C and pH 7.0, varying one substrate concentration while another is fixed. All kinetic experiments are performed at 37 °C in 700 μL of the reaction solution containing 0.5–2.0 mM GSH, 0.5–2.0 mM H2O2, 50 mM potassium phosphate buffer (pH 7.0), 1mM EDTA, 0.25 mM NAPDH, 1 U of GSH reductase, and 5 μM mimic. Background absorption of the non-catalytic reaction was measured without mimic and subtracted.
4. Conclusions
We used CMBI to prepare CITSeSG from trypsin. It has proved that CMBI can be used for other proteins, indicating that CMBI is a versatile technique. Our work explored, for the first time, the use of bio-imprinting for the preparation of efficient GPx mimics from the perspective of reaction kinetics parameters. Bio-imprinting of converted TSeSG to ITSeSG, the KmGSH value of the mimic decreased from 4.82 ± 0.27 mM (TSeSG) to 0.52 ± 0.05 mM (ITSeSG), proving that we produced a highly efficient substrate recognition cavities. Compared with the non-imprinted mimic (TSeSG), the activity of ITSeSG increased 5.7 times. Importantly, this article focused on the activity of the intermediates at various steps leading to the synthetic GPx. In order to confirmed the advantages of CMBI, we compared our GPx mimic’s activity with the activity of natural GPx, Ebselen [26] (the well GPx mimic known), and we found that ITSeSG displays a high GPx like activity and is only 10 times less than that of the native GPx (5780 U/μmol). However, when comparing with Ebselen (0.99 U/μmol), the GPx activity of CITSeSG is 382 times that for Ebselen. After crosslinking of glutaraldehyde, the activity of CITSeSG has a little lower. Comparing with GPx mimic produced by Liu [1] through CMBI which used Subtilisin as imprinted molecular, the activity of ITSeSG is similar to imprinted selenosubtilisin. We believe that this work has laid the foundation for the future design of efficient biological imprinting enzyme.
Acknowledgments
Thanks to the Natural Science Foundation of Jilin Province, China (No. 20130101159JC) and the National Key Scientific Instrument and Equipment Development Project of China (No. 2013YQ470781) for their support of this study.
Author Contributions
Yi Huang, Dan-Yang Ge, Hui Zong, Shao-Wu Lv designed and completed the main framework of this experiment. Yi Huang, Dan-Yang Ge measured and analyzed most of the data obtained by the experiment (including the enzyme activity of the intermediates and the kinetic parameters, etc.) and wrote this part of the content. Xiao-Nan Qu compared the structure of ITSeSG and trypsin. Ju-Xin Yin mainly involved in the revision of article.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Liu, L.; Mao, S.Z.; Liu, X.; Huang, X.; Xu, J.; Liu, J.; Luo, G.; Shen, J. Functional mimicry of the active site of glutathione peroxidase by glutathione imprinted selenium-containing protein. Biomacromolecules 2008, 9, 363–368. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Huang, X.; Miao, L.; Zhu, J.; Yin, L.; Luo, Q.; Xu, J.; Liu, J. A supramolecular bifunctional artificial enzyme with superoxide dismutase and glutathione peroxidase activities. Bioorg. Chem. 2010, 38, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Huang, X.; Lv, C.; Wang, L.; Yu, S.; Luo, Q.; Xu, J.; Liu, J. Construction of an artificial glutathione peroxidase active site on copolymer vesicles. Macromol. Biosci. 2010, 10, 1505–1516. [Google Scholar] [CrossRef] [PubMed]
- Flohé, L.; Loschen, G.; Günzler, W.A.; Eichele, E. Glutathione peroxidase, V. The kinetic mechanism. Hoppe Seylers Z. Physiol. Chem. 1972, 353, 987–1000. [Google Scholar]
- Mao, S.; Dong, Z.; Liu, J.; Li, X.; Liu, X.; Luo, G.; Shen, J. Semisynthetic tellurosubtilisin with glutathione peroxidase activity. J. Am. Chem. Soc. 2005, 127, 11588–11589. [Google Scholar] [CrossRef] [PubMed]
- Mugesh, G.; Singh, H.B. Synthetic organoselenium compounds as antioxidants:glutathione peroxidase activity. Chem. Soc. Rev. 2000, 29, 347–357. [Google Scholar] [CrossRef]
- Xin, X.; Tian, Q.; Yin, G.; Chen, X.; Zhang, J.; Ng, S.; Lu, X. Reduced mitochondrial and ascorbate-glutathione activity after artificial ageing in soybean seed. J. Plant Physiol. 2014, 171, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Liang, K.; Wang, C.; Huang, X.; Mao, S.; Li, X.; Xu, J.; Liu, J.; Luo, G.; Shen, J. A study of biomimetic system: Exploration of factors modulating the catalytic capacity of glutathione peroxidase mimics. J. Mol. Catal. A Chem. 2007, 277, 193–201. [Google Scholar] [CrossRef]
- Xu, H.; Cao, W.; Zhang, X. Selenium-containing polymers: Promising biomaterials for controlled release and enzyme mimics. Acc. Chem. Res. 2013, 46, 1647–1658. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Wang, L.; Jin, H.; Liu, J. Construction of a smart glutathione peroxidase mimic with temperature responsive activity based on block copolymer. Soft Matter 2011, 7, 2521–2529. [Google Scholar] [CrossRef]
- Huang, X.; Liu, X.; Luo, Q.; Liu, J.; Shen, J. ChemInform abstract: Artificial selenoenzymes: Designed and redesigned. ChemInform 2011, 42. [Google Scholar] [CrossRef]
- Wu, Z.P.; Hilvert, D. Conversion of a protease into an acyl transferase: Selenolsubtilisin. J. Am. Chem. Soc. 2002, 111, 453–460. [Google Scholar] [CrossRef]
- Ren, X.; Jemth, P.; Board, P.G.; Luo, G.; Mannervik, B.; Liu, J.; Zhang, K.; Shen, J. A semisynthetic glutathione peroxidase with high catalytic efficiency: Selenoglutathione transferase. Chem. Biol. 2002, 9, 789–794. [Google Scholar] [CrossRef]
- Zheng, K.; Board, P.G.; Fei, X.; Sun, Y.; Lv, S.; Yan, G.; Liu, J.; Shen, J.; Luo, G. A novel selenium-containing glutathione transferase zeta1-1, the activity of which surpasses the level of some native glutathione peroxidases. Int. J. Biochem. Cell Biol. 2008, 40, 2090–2097. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Luo, G.; Gao, S.; Zhang, K.; Chen, X.; Shen, J. Generation of a glutathione peroxidase-like mimic using bioimprinting and chemical mutation. Chem. Commun. 1999, 199, 199–200. [Google Scholar] [CrossRef]
- Shen, N.; Yan, F.; Guo, Y.; Lu, S.; Gong, P.; Xu, Y.; Yan, G.; Mu, Y.; Luo, G. Imprinted human serum albumin with antioxidant activity. Chem. Res. Chin. Univ. 2011, 27, 258–263. [Google Scholar]
- Liu, J.; Zhang, K.; Ren, X.; Luo, G.; Shen, J. Bioimprinted protein exhibits glutathione peroxidase activity. Anal. Chim. Acta 2004, 504, 185–189. [Google Scholar] [CrossRef]
- Huang, X.; Yin, Y.; Liu, Y.; Bai, X.; Zhang, Z.; Xu, J.; Shen, J.; Liu, J. Incorporation of glutathione peroxidase active site into polymer based on imprinting strategy. Biosens. Bioelectron. 2009, 25, 657–660. [Google Scholar] [CrossRef] [PubMed]
- Haupt, K. Microchimica Acta special issue MIP2012 on Molecularly imprinted polymers—Science and technology. Microchim. Acta 2013, 180, 1357–1358. [Google Scholar] [CrossRef]
- Luo, G.M.; Zhu, Z.Q.; Ding, L.; Gao, G.; Sun, Q.A.; Liu, Z.; Yang, T.S.; Shen, J.C. Generation of selenium-containing abzyme by using chemical mutation. Biochem. Biophys. Res. Commun. 1994, 198, 1240–1247. [Google Scholar] [CrossRef] [PubMed]
- Wilson, S.R.; Zucker, P.A.; Huang, R.R.C.; Spector, A. Development of synthetic compounds with glutathione peroxidase activity. ChemInform 1989, 20, 603–612. [Google Scholar] [CrossRef]
- Liu, J.Q.; Jiang, M.S.; Luo, G.M.; Yan, G.L.; Shen, J.C. Conversion of trypsin into a selenium-containing enzyme by using chemical mutation. Biotechnol. Lett. 1998, 20, 693–696. [Google Scholar] [CrossRef]
- Klayman, D.L.; Griffin, T.S. Reaction of selenium with sodium borohydride in protic solvents. A Facile Method for the introduction of selenium into organic molecules. J. Am. Chem. Soc. 1973, 95, 9111–9120. [Google Scholar] [CrossRef]
- Cavallini, D.; Graziani, M.T.; Dupré, S. Determination of disulphide groups in proteins. Nature 1966, 212, 294–295. [Google Scholar] [CrossRef] [PubMed]
- Bell, I.M.; Fisher, M.L.; Wu, Z.P.; Hilvert, D. Kinetic studies on the peroxidase activity of selenosubtilisin. Biochemistry 1993, 32, 3754–3762. [Google Scholar] [CrossRef] [PubMed]
- Antony, S.; Bayse, C.A. Modeling the mechanism of the glutathione peroxidase mimic ebselen. Inorg. Chem. 2011, 50, 12075–12084. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).