Protein Prenylation Makeovers in Plants: Insights into Substrate Diversification
Abstract
1. Modifications of Proteins by Prenyl Groups
1.1. Generalities
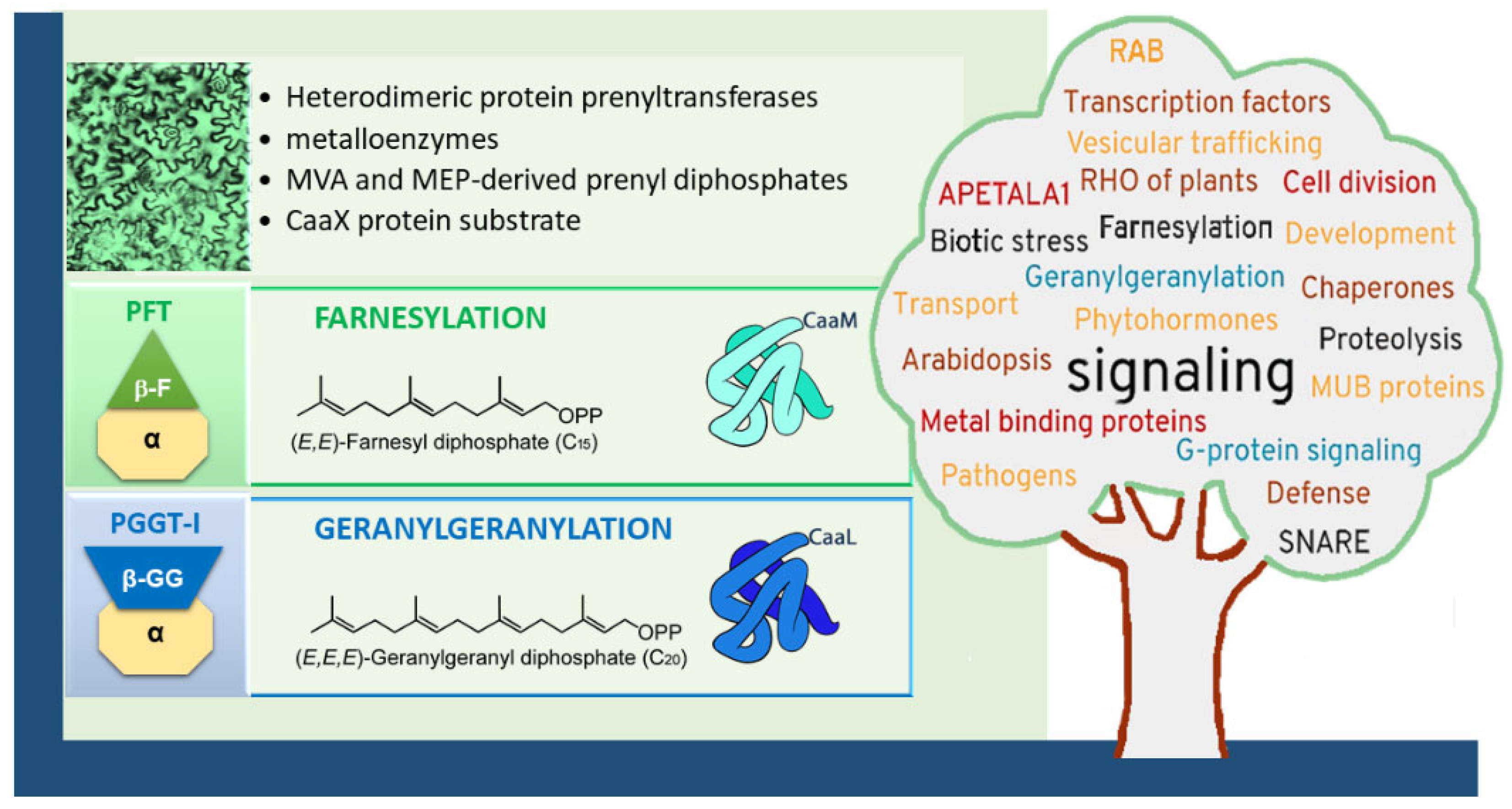
1.2. Protein Prenyltransferases
2. Prenyl Diphosphate Substrates
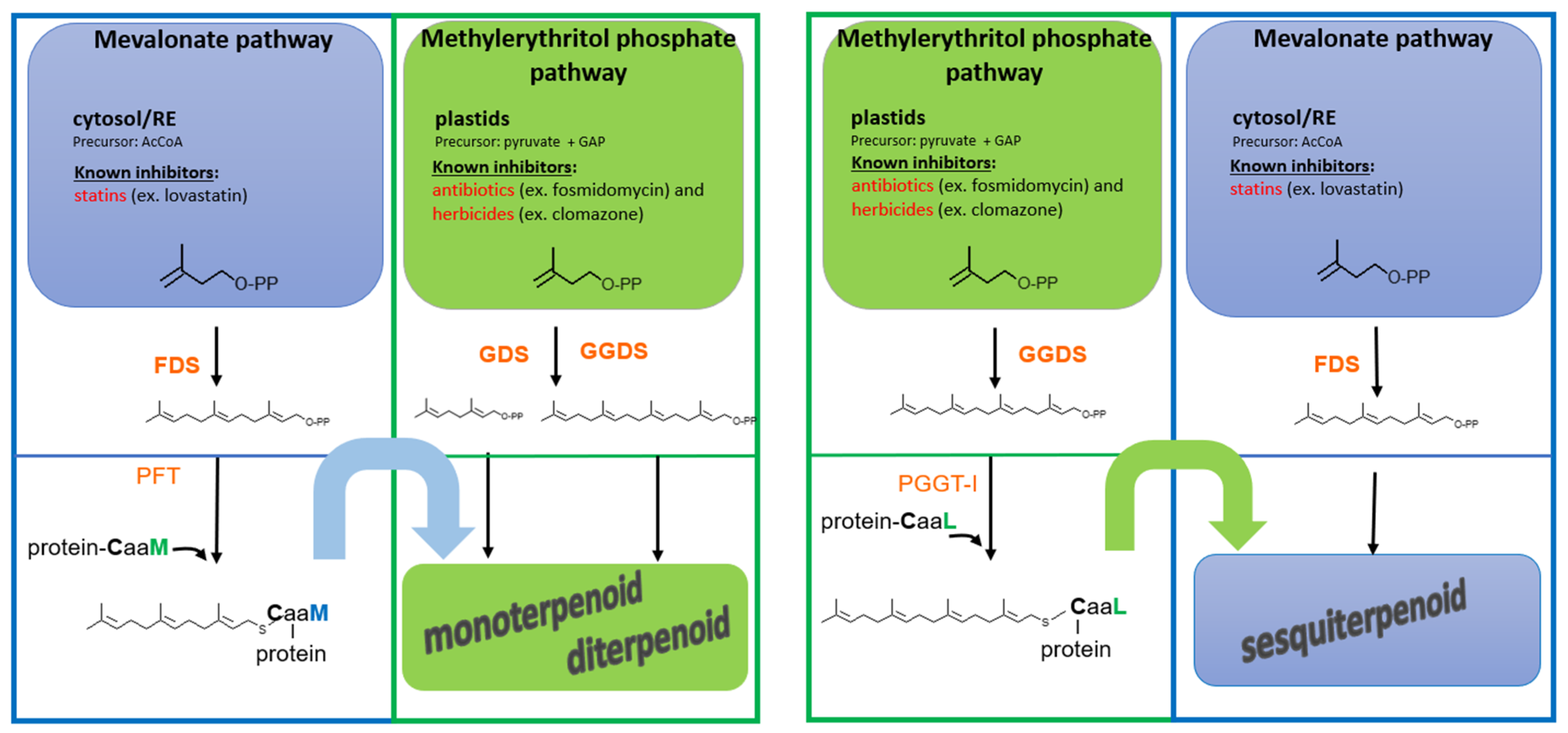
2.1. A Double Metabolic Origin in Plants
2.2. Prioritize Substrate Availability
2.3. Cross-Talk Between the MVA and MEP Pathways Mediated by Protein Prenylation
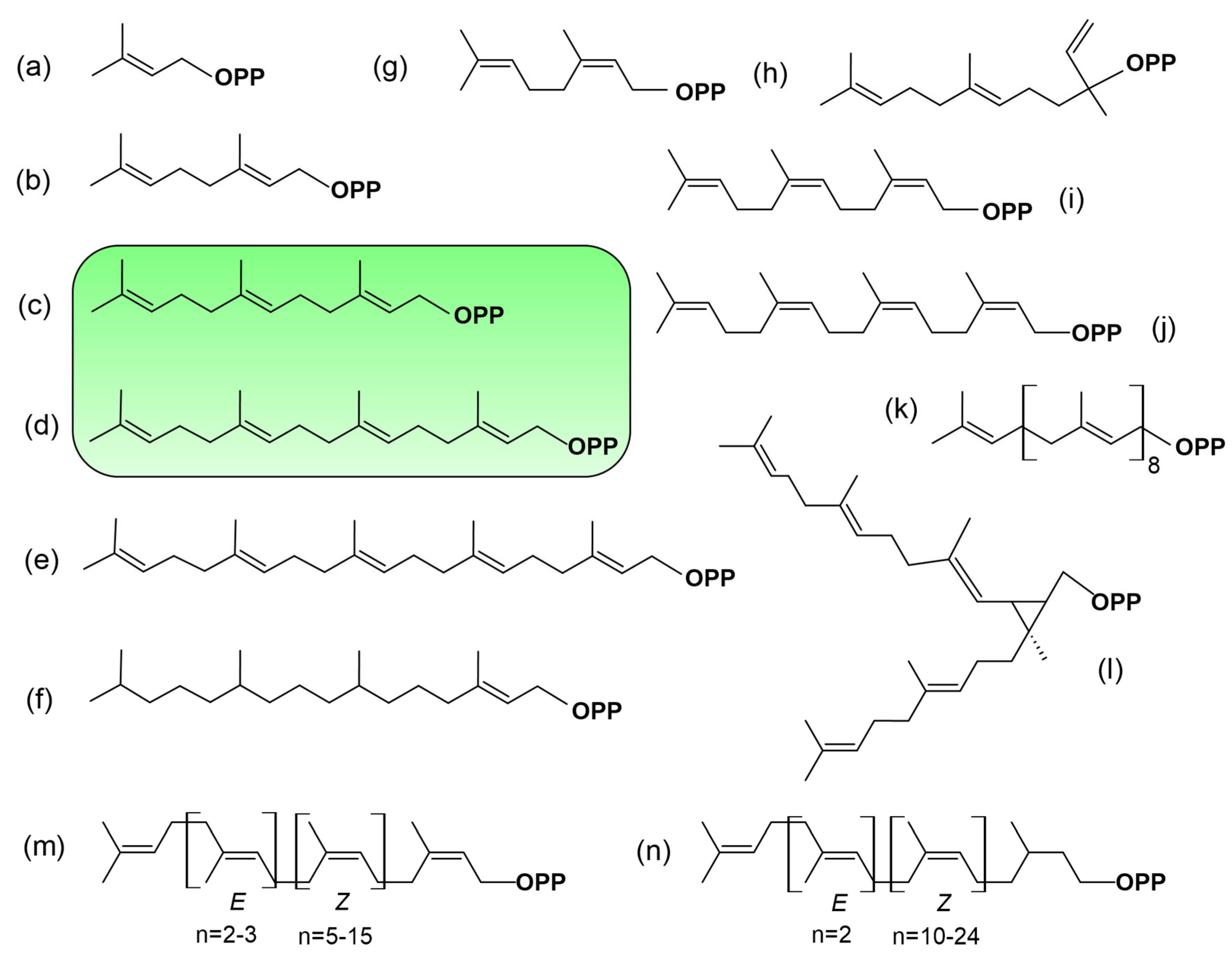
2.4. Substrates of Prenyl Diphosphate Beyond FPP and GGPP
3. CaaX Protein Substrates
3.1. CaaX Motif Proteins in Arabidopsis thaliana
3.2. Functional Conservation and Diversification of CxxX-Containing Proteins
3.3. Validation of CxxX-Containing Proteins as Substrates of PPT
3.4. Post-Transcriptional Modifications as a Mechanism to Obtain New Prenylatable Proteins
3.4.1. Gene Duplication
3.4.2. Alternative Splicing
3.4.3. Gene Fusion
4. Conclusions, Open Questions, and Challenges
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
References
- Chang, H.Y.; Cheng, T.H.; Wang, A.H.J. Structure, catalysis, and inhibition mechanism of prenyltransferase. IUBMB Life 2021, 73, 40–63. [Google Scholar] [CrossRef] [PubMed]
- Jung, D.; Bachmann, H.S. Regulation of protein prenylation. Biomed. Pharmacother. 2023, 164, 114915. [Google Scholar] [CrossRef] [PubMed]
- Sakamaki, J.I.; Mizushima, N. Cell biology of protein–lipid conjugation. Cell Struct. Funct. 2023, 48, 99–112. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Wang, Q. Protein prenylation in plants: Mechanisms and functional implications. Plants 2025, 14, 1759. [Google Scholar] [CrossRef]
- Crowell, D.N.; Huizinga, D.H. Protein isoprenylation: The fat of the matter. Trends Plant Sci. 2009, 14, 163–170, Erratum in Trends Plant Sci. 2009, 14, 181. [Google Scholar] [CrossRef]
- Galichet, A.; Gruissem, W. Protein farnesylation in plants—Conserved mechanisms but different targets. Curr. Opin. Plant Biol. 2003, 6, 530–535. [Google Scholar] [CrossRef]
- Hála, M.; Žárský, V. Protein prenylation in plant stress responses. Molecules 2019, 24, 3906. [Google Scholar] [CrossRef]
- Running, M.P. The role of lipid post–translational modification in plant developmental processes. Front. Plant Sci. 2014, 5, 50. [Google Scholar] [CrossRef]
- Rilling, H.C.; Bruenger, E.; Epstein, W.W.; Kandutsch, A.A. Prenylated proteins: Demonstration of a thioether linkage to cysteine of proteins. Biochem. Biophys. Res. Commun. 1989, 163, 143–148. [Google Scholar] [CrossRef]
- Infant, T.; Deb, R.; Ghose, S.; Nagotu, S. Post-translational modifications of proteins associated with yeast peroxisome membrane: An essential mode of regulatory mechanism. Genes Cells 2021, 26, 843–860. [Google Scholar] [CrossRef]
- Wang, T.-Y.; Wu, J.-R.; Duong, N.K.T.; Lu, C.-A.; Yeh, C.H.; Wu, S.-J. HSP70-4 and farnesylated AtJ3 constitute a specific HSP70/HSP40-based chaperone machinery essential for prolonged heat stress tolerance in Arabidopsis. J. Plant Physiol. 2021, 261, 153430. [Google Scholar] [CrossRef] [PubMed]
- Mondal, K.; Raj, S.; Thakur, K.; Verma, A.; Kharwal, N.; Chowdhury, A.; Sadhu, S.; Ram, M.; Bishnoi, P.; Dutta, S.; et al. Molecular basis of plant adaptation against aridity. In Abiotic Stress in Plants-Adaptations to Climate Change; IntechOpen: London, UK, 2023. [Google Scholar] [CrossRef]
- Chen, L.; Kashina, A. Post-translational modifications of the protein termini. Front. Cell Dev. Biol. 2021, 9, 719590. [Google Scholar] [CrossRef] [PubMed]
- Parish, C.A.; Rando, R.R. Isoprenylation/methylation of proteins enhances membrane association by a hydrophobic mechanism. Biochemistry 1996, 35, 8473–8477. [Google Scholar] [CrossRef]
- Thompson, G.A., Jr.; Okuyama, H. Lipid-linked proteins of plants. Prog. Lipid Res. 2000, 39, 19–39. [Google Scholar] [CrossRef]
- Huizinga, D.H.; Omosegbon, O.; Omery, B.; Crowell, D.N. Isoprenylcysteine methylation and demethylation regulate abscisic acid signaling in Arabidopsis. Plant Cell 2008, 20, 2714–2728. [Google Scholar] [CrossRef]
- Smokvarska, M.; Jaillais, Y.; Martinière, A. Function of membrane domains in Rho-Of-Plant signaling. Plant Physiol. 2021, 185, 663–681. [Google Scholar] [CrossRef]
- Sanderson, J.M. Far from inert: Membrane lipids possess intrinsic reactivity that has consequences for cell biology. BioEssays 2020, 42, 1900147. [Google Scholar] [CrossRef]
- Yalovsky, S.; Rodríguez-Concepción, M.; Bracha, K.; Toledo-Ortiz, G.; Gruissem, W. Prenylation of the floral transcription factor APETALA1 modulates its function. Plant Cell 2000, 12, 1257–1266. [Google Scholar] [CrossRef]
- Tao, W.; Lijuan, L.; Zeyu, L.; Lianguang, S.; Quan, W. Cloning and characterization of protein prenyltransferase alpha subunit in rice. Rice Sci. 2021, 28, 557–566. [Google Scholar] [CrossRef]
- Cutler, S.; Ghassemian, M.; Bonetta, D.; Cooney, S.; McCourt, P. A protein farnesyl transferase involved in abscisic acid signal transduction in Arabidopsis. Science 1996, 273, 1239–1241. [Google Scholar] [CrossRef]
- Pei, Z.M.; Ghassemian, M.; Kwak, C.M.; McCourt, P.; Schroeder, J.I. Role of farnesyltransferase in ABA regulation of guard cell anion channels and plant water loss. Science 1998, 282, 287–290. [Google Scholar] [CrossRef]
- Running, M.P.; Lavy, M.; Sternberg, H.; Galichet, A.; Gruissem, W.; Hake, S.; Ori, N.; Yalovsky, S. Enlarged meristems and delayed growth in plp mutants result from lack of CaaX prenyltransferases. Proc. Natl. Acad. Sci. USA 2004, 101, 7815–7820. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.D.; Chary, N.; Chernoff, E.A.; Zeng, Q.; Running, M.P.; Crowell, D.N. Protein geranylgeranyltransferase I is involved in specific aspects of abscisic acid and auxin signaling in Arabidopsis. Plant Physiol. 2005, 139, 722–733. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, E. The small GTPase superfamily in plants: A conserved regulatory module with novel functions. Annu. Rev. Plant Biol. 2020, 71, 247–272. [Google Scholar] [CrossRef] [PubMed]
- Wilson, A.L.; Erdman, R.A.; Castellano, F.; Maltese, W.A. Prenylation of Rab8 GTPase by type I and type II geranylgeranyl transferases. Biochem. J. 1998, 333, 497–504. [Google Scholar] [CrossRef]
- Shi, W.; Zeng, Q.; Kundel, B.N.; Running, M.P. Arabidopsis Rab geranylgeranyltransferases demonstrate redundancy and broad substrate specificity in vitro. J. Biol. Chem. 2016, 291, 1398–1410. [Google Scholar] [CrossRef]
- Kuchay, S.; Wang, H.; Marzio, A.; Jain, K.; Homer, H.; Fehrenbacher, N.; Philips, M.R.; Zheng, N.; Pagano, M. GGTase3 is a newly identified geranylgeranyltransferase targeting a ubiquitin ligase. Nat. Struct. Mol. Biol. 2019, 26, 628–636. [Google Scholar] [CrossRef]
- Shirakawa, R.; Goto-Ito, S.; Goto, K.; Wakayama, S.; Kubo, H.; Sakata, N.; Trinh, D.A.; Yamagata, A.; Sato, Y.; Masumoto, H.; et al. A SNARE geranylgeranyltransferase essential for the organization of the Golgi apparatus. EMBO J. 2020, 39, e104120. [Google Scholar] [CrossRef]
- Sakata, N.; Shirakawa, R.; Goto, K.; Trinh, D.A.; Horiuchi, H. Double prenylation of SNARE protein Ykt6 is required for lysosomal hydrolase trafficking. J. Biochem. 2021, 169, 363–370. [Google Scholar] [CrossRef]
- Tateishi, M.; Goto, K.; Hishinuma, E.; Matsukawa, N.; Kishimoto, T.; Tanaka, K.; Shirakawa, R. Double prenylation of budding yeast Ykt6 regulates cell wall integrity and autophagy. J. Biol. Chem. 2025, 301, 108384. [Google Scholar] [CrossRef]
- Ma, T.; Tan, J.R.; Lu, J.Y.; Li, S.; Zhang, Y. S-acylation of YKT61 modulates its unconventional participation in the formation of SNARE complexes in Arabidopsis. J. Genet. Genom. 2024, 51, 1079–1088. [Google Scholar] [CrossRef]
- Counihan, N.A.; Chernih, H.C.; de Koning-Ward, T.F. Post-translational lipid modifications in Plasmodium parasites. Curr. Opin. Microbiol. 2022, 69, 102196. [Google Scholar] [CrossRef] [PubMed]
- Singh, P. Plant Protein Prenylation in Environmental Responses and Protein Production. Ph.D. Thesis, University Of Louisville, Louisville, KY, USA, 2019. Paper 3264. [Google Scholar] [CrossRef]
- Suazo, K.F.; Bělíček, J.; Schey, G.L.; Auger, S.A.; Petre, A.M.; Li, L.; Błażewska, K.M.; Kopečný, D.; Distefano, M.D. Thinking outside the CaaX-box: An unusual reversible prenylation on ALDH9A1. RSC Chem. Biol. 2023, 4, 913–925. [Google Scholar] [CrossRef] [PubMed]
- Hemmerlin, A. Phosphorylation of metabolites involved in salvage pathways for isoprenoid biosynthesis in plants. Kinases Phosphatases 2023, 1, 151–166. [Google Scholar] [CrossRef]
- Marchwicka, A.; Kamińska, D.; Monirialamdari, M.; Błażewska, K.M.; Gendaszewska-Darmach, E. Protein prenyltransferases and their inhibitors: Structural and functional characterization. Int. J. Mol. Sci. 2022, 23, 5424. [Google Scholar] [CrossRef]
- Guerra, B.; Recio, C.; Aranda-Tavío, H.; Guerra-Rodríguez, M.; García-Castellano, J.M.; Fernández-Pérez, L. The mevalonate pathway, a metabolic target in cancer therapy. Front. Oncol. 2021, 11, 626971. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Hoeffler, J.-F.; Meyer, O.; Tritsch, D.; Kagan, I.A.; Grosdemange-Billiard, C.; Rohmer, M.; Bach, T.J. Crosstalk between the cytosolic mevalonate and the plastidial methylerythritol phosphate pathways in tobacco Bright Yellow-2 cells. J. Biol. Chem. 2003, 278, 26666–26676. [Google Scholar] [CrossRef]
- Gerber, E.; Hemmerlin, A.; Hartmann, M.; Heintz, D.; Hartmann, M.-A.; Mutterer, J.; Rodríguez-Concepción, M.; Boronat, A.; Van Dorsselaer, A.; Rohmer, M.; et al. The plastidial 2C-methyl-D-erythritol 4-phosphate pathway provides the isoprenyl moiety for protein geranylgeranylation in tobacco BY-2 cells. Plant Cell 2009, 21, 285–300. [Google Scholar] [CrossRef]
- Hartmann, M.; Hemmerlin, A.; Gas-Pascual, E.; Gerber, E.; Tritsch, D.; Rohmer, M.; Bach, T.J. The effect of MEP pathway and other inhibitors on the intracellular localization of a plasma membrane-targeted, isoprenylable GFP reporter protein in tobacco BY-2 cells. F1000Research 2013, 2, 170. [Google Scholar] [CrossRef]
- Huchelmann, A.; Gastaldo, C.; Veinante, M.; Zeng, Y.; Heintz, D.; Tritsch, D.; Schaller, H.; Rohmer, M.; Bach, T.J.; Hemmerlin, A. S-Carvone suppresses cellulase-induced capsidiol production in Nicotiana tabacum by interfering with protein isoprenylation. Plant Physiol. 2014, 164, 935–950. [Google Scholar] [CrossRef]
- Huchelmann, A.; Brahim, M.S.; Gerber, E.; Tritsch, D.; Bach, T.J.; Hemmerlin, A. Farnesol-mediated shift in the metabolic origin of prenyl groups used for protein prenylation in plants. Biochimie 2016, 127, 95–102. [Google Scholar] [CrossRef]
- Kumar, S.R.; Rai, A.; Bomzan, D.P.; Kumar, K.; Hemmerlin, A.; Dwivedi, V.; Godbole, R.C.; Barvkar, V.; Shanker, K.; Shilpashree, H.B.; et al. A plastid-localized bona fide geranylgeranyl diphosphate synthase plays a necessary role in monoterpene indole alkaloid biosynthesis in Catharanthus roseus. Plant J. 2020, 103, 248–265. [Google Scholar] [CrossRef] [PubMed]
- Chevalier, Q.; Huchelmann, A.; Debié, P.; Mercier, P.; Hartmann, M.; Vonthron-Sénécheau, C.; Bach, T.J.; Schaller, H.; Hemmerlin, A. Methyl-jasmonate functions as a molecular switch promoting cross-talk between pathways for the biosynthesis of isoprenoid backbones used to modify proteins in plants. Plants 2024, 13, 1110. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Zhang, X.; Chen, X.; Aramsangtienchai, P.; Tong, Z.; Lin, H. Protein lipidation: Occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 2018, 118, 919–988. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, K.; Zimmerman, K.; Scholten, J.; Gelb, M.H. Differential prenyl pyrophosphate binding to mammalian protein geranylgeranyltransferase-I and protein farnesyltransferase and its consequence on the specificity of protein prenylation. J. Biol. Chem. 1997, 272, 3944–3952. [Google Scholar] [CrossRef]
- Long, S.B.; Casey, P.J.; Beese, L.S. Reaction path of protein farnesyltransferase at atomic resolution. Nature 2002, 419, 645–650. [Google Scholar] [CrossRef]
- Lane, K.T.; Beese, L.S. Structural biology of protein farnesyltransferase and geranylgeranyltransferase type I. J. Lipid Res. 2006, 47, 681–699. [Google Scholar] [CrossRef]
- Palsuledesai, C.C.; Distefano, M.D. Protein prenylation: Enzymes, therapeutics, and biotechnology applications. ACS Chem. Biol. 2015, 10, 51–62. [Google Scholar] [CrossRef]
- Zverina, E.A.; Lamphear, C.L.; Wright, E.N.; Fierke, C.A. Recent Advances in protein prenyltransferases: Substrate identification, regulation, and disease interventions. Curr. Opin. Chem. Biol. 2012, 16, 544–552. [Google Scholar] [CrossRef]
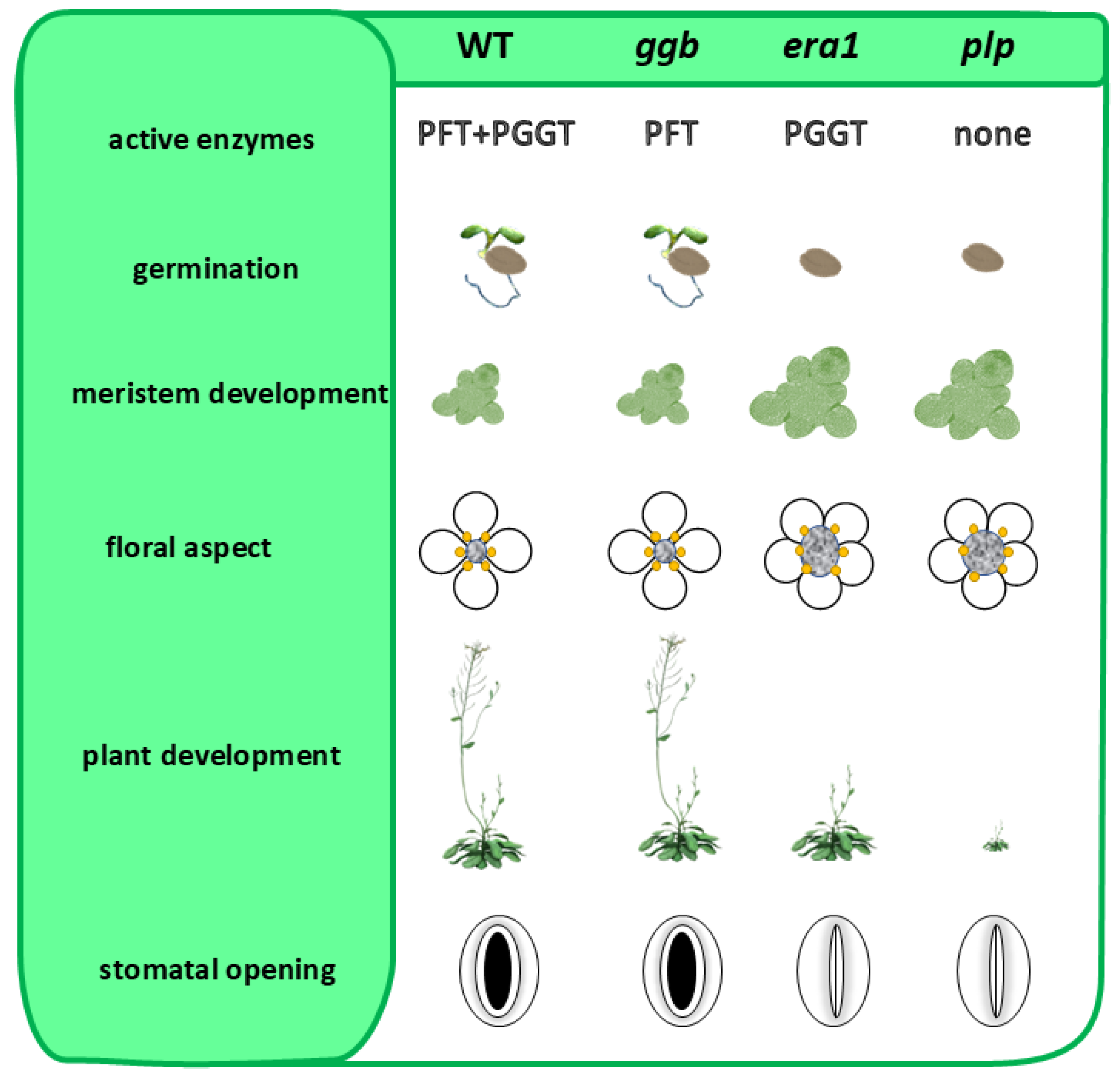
- Andrews, M.; Huizinga, D.H.; Crowell, D.N. The CaaX specificities of Arabidopsis protein prenyltransferases explain era1 and ggb phenotypes. BMC Plant Biol. 2010, 10, 118. [Google Scholar] [CrossRef]
- Hartman, H.L.; Hick, K.A.; Fierke, C.A. Peptide specificity of protein prenyltransferases is determined mainly by reactivity rather than binding affinity. Biochemistry 2005, 44, 15314–15324. [Google Scholar] [CrossRef] [PubMed]
- Whyte, D.B.; Kirschmeier, P.; Hockenberry, T.N.; Nunez-Oliva, I.; James, L.; Catino, J.J.; Bishop, W.R.; Pai, J.K. K- and N-Ras are geranylgeranylated in cells treated with farnesyl protein transferase inhibitors. J. Biol. Chem. 1997, 272, 14459–14464. [Google Scholar] [CrossRef] [PubMed]
- Eastman, R.T.; White, J.; Hucke, O.; Bauer, K.; Yokoyama, K.; Nallan, L.; Chakrabarti, D.; Verlinde, C.L.M.J.; Gelb, M.H.; Rathod, P.K.; et al. Resistance to a protein farnesyltransferase inhibitor in Plasmodium falciparum. J. Biol. Chem. 2005, 280, 13554–13559. [Google Scholar] [CrossRef] [PubMed]
- Goalstone, M.; Carel, K.; Leitner, J.W.; Draznin, B. Insulin stimulates the phosphorylation and activity of farnesyltransferase via the Ras-Mitogen-activated protein kinase pathway. Endocrinology 1997, 138, 5119–5124. [Google Scholar] [CrossRef]
- Kumar, A.; Beresini, M.H.; Dhawan, P.; Mehta, K.D. α-Subunit of farnesyltransferase is phosphorylated in vivo: Effect of protein phosphatase-1 on enzymatic activity. Biochem. Biophys. Res. Commun. 1996, 222, 445–452. [Google Scholar] [CrossRef]
- Pekel, H.; Guzel, M.; Sensoy, O. Mechanistic insight into impact of phosphorylation on the enzymatic steps of farnesyltransferase. Protein Sci. 2022, 31, e4414. [Google Scholar] [CrossRef]
- Schmitt, D.; Callan, K.; Gruissem, W. Molecular and biochemical characterization of tomato farnesyl-protein transferase. Plant Physiol. 1996, 112, 767–777. [Google Scholar] [CrossRef][Green Version]
- Thai, L.; Rush, J.S.; Maul, J.E.; Devarenne, T.; Rodgers, D.L.; Chappell, J.; Waechter, C.J. Farnesol is utilized for isoprenoid biosynthesis in plant cells via farnesyl pyrophosphate formed by successive monophosphorylation reactions. Proc. Natl. Acad. Sci. USA 1999, 96, 13080–13085. [Google Scholar] [CrossRef]
- Pu, X.; Dong, X.; Li, Q.; Chen, Z.; Liu, L. An update on the function and regulation of methylerythritol phosphate and mevalonate pathways and their evolutionary dynamics. J. Integr. Plant Biol. 2021, 63, 1211–1226. [Google Scholar] [CrossRef]
- Rolland, N.; Curien, G.; Finazzi, G.; Kuntz, M.; Marechal, E.; Matringe, M.; Ravanel, S.; Seigneurin-Berny, D. The biosynthetic capacities of the plastids and integration between cytoplasmic and chloroplast processes. Annu. Rev. Genet. 2012, 46, 233–264. [Google Scholar] [CrossRef]
- Bergman, M.E.; Kortbeek, R.W.; Gutensohn, M.; Dudareva, N. Plant terpenoid biosynthetic network and its multiple layers of regulation. Prog. Lipid Res. 2024, 95, 101287. [Google Scholar] [CrossRef] [PubMed]
- Okada, M.; Rajaram, K.; Swift, R.P.; Mixon, A.; Maschek, J.A.; Prigge, S.T.; Sigala, P.A. Critical role for isoprenoids in apicoplast biogenesis by malaria parasites. eLife 2022, 11, e73208. [Google Scholar] [CrossRef] [PubMed]
- Beck, G.; Coman, D.; Herren, E.; Ruiz-Sola, M.A.; Rodríguez-Concepción, M.; Gruissem, W.; Vranová, E. Characterization of the GGPP synthase gene family in Arabidopsis thaliana. Plant Mol. Biol. 2013, 82, 393–416. [Google Scholar] [CrossRef] [PubMed]
- Nagel, R.; Bernholz, C.; Vranová, E.; Košuth, J.; Bergau, N.; Ludwig, S.; Wessjohann, L.; Gershenzon, J.; Tissier, A.; Schmidt, A. Arabidopsis thaliana isoprenyl diphosphate synthases produce the C25 intermediate geranylfarnesyl diphosphate. Plant J. 2015, 84, 847–859. [Google Scholar] [CrossRef]
- Ruiz-Sola, M.A.; Barja, M.V.; Manzano, D.; Llorente, B.; Schipper, B.; Beekwilder, J.; Rodríguez-Concepción, M. A single Arabidopsis gene encodes two differentially targeted geranylgeranyl diphosphate synthase isoforms. Plant Physiol. 2016, 172, 1393–1402. [Google Scholar] [CrossRef]
- Pedenla Bomzan, D.; Sharma, A.; Lemos Cruz, P.; Carqueijeiro, I.; Bellenger, L.; Rai, A.; Thippesh, A.K.; Chinnegowda, V.S.; Parihar, D.; Ducos, E.; et al. ROP GTPases with a geranylgeranylation motif modulate alkaloid biosynthesis in Catharanthus roseus. Plant Physiol. 2024, 195, 2213–2233. [Google Scholar] [CrossRef]
- Sharma, A.; Mohan, S.; Gupta, P.; Parihar, D.; Nagegowda, D.A. Three Rho of Plant (ROP)-GTPase regulatory proteins control ROP-mediated alkaloid biosynthesis in Catharanthus roseus. Plant Physiol. 2025, 197, kiaf115. [Google Scholar] [CrossRef]
- Demurtas, O.C.; Nicolia, A.; Diretto, G. Terpenoid transport in plants: How far from the final picture? Plants 2023, 12, 634. [Google Scholar] [CrossRef]
- Manne, V.; Roberts, D.; Tobin, A.; O’Rourke, E.; De Virgilio, M.; Meyers, C.; Ahmed, N.; Kurz, B.; Resh, M.; Kung, H.F.; et al. Identification and preliminary characterization of protein-cysteine farnesyltransferase. Proc. Natl. Acad. Sci. USA 1990, 87, 7541–7545. [Google Scholar] [CrossRef]
- Tong, H.; Wiemer, A.J.; Neighbors, J.D.; Hohl, R.J. Quantitative determination of farnesyl and geranylgeranyl diphosphate levels in mammalian tissue. Anal. Biochem. 2008, 378, 138–143. [Google Scholar] [CrossRef]
- Zhao, Y.; Wu, T.Y.; Zhao, M.F.; Li, C.J. The balance of protein farnesylation and geranylgeranylation during the progression of nonalcoholic fatty liver disease. J. Biol. Chem. 2020, 295, 5152–5162. [Google Scholar] [CrossRef]
- Schumacher, M.M.; DeBose-Boyd, R.A. Posttranslational regulation of HMG-CoA reductase, the rate-limiting enzyme in synthesis of cholesterol. Annu. Rev. Biochem. 2021, 90, 659–679. [Google Scholar] [CrossRef] [PubMed]
- Gutensohn, M.; Hartzell, E.; Dudareva, N. Another level of complexity: The role of metabolic channeling and metabolons in plant terpenoid metabolism. Front. Plant Sci. 2022, 13, 954083. [Google Scholar] [CrossRef] [PubMed]
- Dahmani, I.; Qin, K.; Zhang, Y.; Fernie, A.R. The formation and function of plant metabolons. Plant J. 2023, 114, 1080–1092. [Google Scholar] [CrossRef]
- Hajj Chehade, M.; Pelosi, L.; Fyfe, C.D.; Loiseau, L.; Rascalou, B.; Brugière, S.; Kazemzadeh, K.; Vo, C.D.T.; Ciccone, L.; Aussel, L.; et al. A soluble metabolon synthesizes the isoprenoid lipid ubiquinone. Cell. Chem. Biol. 2019, 26, 482–492. [Google Scholar] [CrossRef]
- Bassard, J.E.; Halkier, B.A. How to prove the existence of metabolons? Phytochem. Rev. 2018, 17, 211–227. [Google Scholar] [CrossRef]
- Bassard, J.E.; Laursen, T. Molecular snapshots of dynamic membrane-bound metabolons. Methods Enzymol. 2019, 617, 1–27. [Google Scholar] [CrossRef]
- Maeda, H.A. Harnessing evolutionary diversification of primary metabolism for plant synthetic biology. J. Biol. Chem. 2019, 294, 16549–16566. [Google Scholar] [CrossRef]
- Chappell, J. The biochemistry and molecular biology of isoprenoid metabolism. Plant Physiol. 1995, 107, 1–6. [Google Scholar] [CrossRef]
- Zhang, Y.; Fernie, A.R. Metabolons, enzyme–enzyme assemblies that mediate substrate channeling, and their roles in plant metabolism. Plant Commun. 2021, 2, 100081. [Google Scholar] [CrossRef]
- Camagna, M.; Grundmann, A.; Bär, C.; Koschmieder, J.; Beyer, P.; Welsch, R. Enzyme fusion removes competition for geranylgeranyl diphosphate in carotenogenesis. Plant Physiol. 2019, 179, 1013–1027. [Google Scholar] [CrossRef]
- Linnenbrügger, L.; Doering, L.; Lansing, H.; Fischer, K.; Eirich, J.; Finkemeier, I.; von Schaewen, A. Alternative splicing of Arabidopsis G6PD5 recruits NADPH-producing OPPP reactions to the endoplasmic reticulum. Front. Plant Sci. 2022, 13, 909624. [Google Scholar] [CrossRef]
- Baranyi, M.; Buday, L.; Hegedűs, B. K-Ras prenylation as a potential anticancer target. Cancer Metastasis Rev. 2020, 39, 1127–1141. [Google Scholar] [CrossRef] [PubMed]
- Randall, S.K.; Marshall, M.S.; Crowell, D.N. Protein isoprenylation in suspension-cultured tobacco cells. Plant Cell 1993, 5, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Verdaguer, I.B.; Crispim, M.; Hernández, A.; Katzin, A.M. The biomedical importance of the missing pathway for farnesol and geranylgeraniol salvage. Molecules 2022, 27, 8691. [Google Scholar] [CrossRef] [PubMed]
- Selwal, N.; Goutam, U.; Akhtar, N.; Sood, M.; Kukreja, S. Elicitation: “A Trump Card” for enhancing secondary metabolites in plants. J. Plant Growth Regul. 2024, 43, 3027–3047. [Google Scholar] [CrossRef]
- Eisenreich, W.; Menhard, B.; Hylands, P.J.; Zenk, M.H.; Bacher, A. Studies on the biosynthesis of taxol: The taxane carbon skeleton is not of mevalonoid origin. Proc. Natl. Acad. Sci. USA 1996, 93, 6431–6436. [Google Scholar] [CrossRef]
- Cusidó, R.M.; Palazón, J.; Bonfill, M.; Expósito, O.; Moyano, E.; Piñol, M.T. Source of isopentenyl diphosphate for taxol and baccatin III biosynthesis in cell cultures of Taxus baccata. Biochem. Eng. J. 2007, 33, 159–167. [Google Scholar] [CrossRef]
- Contin, A.; van der Heijden, R.; Lefeber, A.W.; Verpoorte, R. The iridoid glucoside secologanin is derived from the novel triose phosphate/pyruvate pathway in a Catharanthus roseus cell culture. FEBS Lett. 1998, 434, 413–416. [Google Scholar] [CrossRef]
- Imbault, N.; Thiersault, M.; Dupéron, P.; Benabdelmouna, A.; Doireau, P. Pravastatin: A tool for investigating the availability of mevalonate metabolites for primary and secondary metabolism in Catharanthus roseus cell suspensions. Physiol. Plant. 1996, 98, 803–809. [Google Scholar] [CrossRef]
- Kumar, M.; Carr, P.; Turner, S. An atlas of Arabidopsis protein S-acylation reveals its widespread role in plant cell organization and function. Nat. Plants 2022, 8, 670–681. [Google Scholar] [CrossRef]
- Resh, M.D. Lipid modification of proteins. In Biochemistry of Lipids, Lipoproteins and Membranes; Elsevier: Amsterdam, The Netherlands, 2021; pp. 429–456. [Google Scholar] [CrossRef]
- Sorek, N.; Gutman, O.; Bar, E.; Abu-Abied, M.; Feng, X.; Running, M.P.; Lewinsohn, E.; Ori, N.; Sadot, E.; Henis, Y.I.; et al. Differential effects of prenylation and S-acylation on type I and II ROPS membrane interaction and function. Plant Physiol. 2011, 155, 706–720. [Google Scholar] [CrossRef] [PubMed]
- Kopcsayová, D.; Vranová, E. Functional gene network of prenyltransferases in Arabidopsis thaliana. Molecules 2019, 24, 4556. [Google Scholar] [CrossRef] [PubMed]
- Barja, M.V.; Rodriguez-Concepcion, M. Plant geranylgeranyl diphosphate synthases: Every (gene) family has a story. aBIOTECH 2021, 2, 289–298. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Li, J.; Ma, Y.; Yuan, W.; Zhang, P.; Wang, G. Occurrence and biosynthesis of plant sesterterpenes (C25), a new addition to terpene diversity. Plant Commun. 2021, 2, 100184. [Google Scholar] [CrossRef]
- Hoshino, Y.; Villanueva, L. Four billion years of microbial terpenome evolution. FEMS Microbiol. Rev. 2023, 47, fuad008. [Google Scholar] [CrossRef]
- Zhou, F.; Pichersky, E. More is better: The diversity of terpene metabolism in plants. Curr. Opin. Plant Biol. 2020, 55, 1–10. [Google Scholar] [CrossRef]
- Manaswiyoungkul, P.; de Araujo, E.D.; Gunning, P.T. Targeting prenylation inhibition through the mevalonate pathway. RSC Med. Chem. 2020, 11, 51–71. [Google Scholar] [CrossRef]
- Mori, T. Enzymatic studies on aromatic prenyltransferases. J. Nat. Med. 2020, 74, 501–512. [Google Scholar] [CrossRef]
- Bergman, M.E.; Dudareva, N. Plant specialized metabolism: Diversity of terpene synthases and their products. Curr. Opin. Plant Biol. 2024, 81, 102607. [Google Scholar] [CrossRef]
- Yazaki, K.; Sasaki, K.; Tsurumaru, Y. Prenylation of aromatic compounds, a key diversification of plant secondary metabolites. Phytochemistry 2009, 70, 1739–1745. [Google Scholar] [CrossRef]
- Pais, A.C.S.; Saraiva, J.A.; Rocha, S.M.; Silvestre, A.J.D.; Santos, S.A.O. Current research on the bioprospection of linear diterpenes from Bifurcaria bifurcate: From extraction methodologies to possible applications. Mar. Drugs 2019, 17, 556. [Google Scholar] [CrossRef]
- Shipton, C.A.; Parmryd, I.; Swiezewska, E.; Andersson, B.; Dallner, G. Isoprenylation of plant proteins in vivo: Isoprenylated proteins are abundant in the mitochondria and nuclei of spinach. J. Biol. Chem. 1995, 270, 566–572. [Google Scholar] [CrossRef]
- Parmryd, I.; Andersson, B.; Dallner, G. Protein prenylation in spinach chloroplasts. Proc. Natl. Acad. Sci. USA 1999, 96, 10074–10079. [Google Scholar] [CrossRef] [PubMed]
- Parmryd, I.; Dallner, G. In vivo prenylation of rat proteins: Modification of proteins with penta- and hexaprenyl groups. Arch. Biochem. Biophys. 1999, 364, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Gutkowska, M.; Bieńkowski, T.; Hung, V.S.; Wanke, M.; Hertel, J.; Danikiewicz, W.; Swiezewska, E. Proteins are polyisoprenylated in Arabidopsis thaliana. Biochem. Biophys. Res. Comm. 2004, 322, 998–1004. [Google Scholar] [CrossRef] [PubMed]
- Lipko, A.; Pączkowski, C.; Perez-Fons, L.; Fraser, P.D.; Kania, M.; Hoffman-Sommer, M.; Danikiewicz, W.; Rohmer, M.; Poznanski, J.; Swiezewska, E. Divergent contribution of the MVA and MEP pathways to the formation of polyprenols and dolichols in Arabidopsis. Biochem. J. 2023, 480, 495–520. [Google Scholar] [CrossRef]
- Vom Dorp, K.; Hölzl, G.; Plohmann, C.; Eisenhut, M.; Abraham, M.; Weber, A.P.M.; Hanson, A.D.; Dörmann, P. Remobilization of phytol from chlorophyll degradation is essential for tocopherol synthesis and growth of Arabidopsis. Plant Cell 2015, 27, 2846–2859. [Google Scholar] [CrossRef]
- Huchelmann, A. Etude de L’implication des Deux Voies de Biosynthèse des Isoprénoïdes Pour la Spécificité et la Régulation de la Prénylation des Protéines Chez Les Plantes. Ph.D. Thesis, University of Strasbourg, Strasbourg, France, 2013. [Google Scholar]
- Mann, R.K.; Beachy, P.A. Cholesterol modification of proteins. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2000, 1529, 188–202. [Google Scholar] [CrossRef]
- Purohit, R.; Peng, D.S.; Vielmas, E.; Ondrus, A.E. Dual roles of the sterol recognition region in Hedgehog protein modification. Commun. Biol. 2020, 3, 250, Correction in Commun. Biol. 2020, 3, 286.. [Google Scholar] [CrossRef]
- Mafi, A.; Purohit, R.; Vielmas, E.; Lauinger, A.R.; Lam, B.; Cheng, Y.S.; Zhang, T.; Huang, Y.; Kim, S.-K.; Goddard, W.A.; et al. Hedgehog proteins create a dynamic cholesterol interface. PLoS ONE 2021, 16, e0246814. [Google Scholar] [CrossRef] [PubMed]
- Faust, J.R.; Dice, J.F. Evidence for isopentenyl adenine modification of a cell cycle regulated protein. J. Biol. Chem. 1991, 266, 9961–9970. [Google Scholar] [CrossRef] [PubMed]
- Crowell, D.N.; Salaz, M.S. Inhibition of growth of cultures tobacco cells at low concentrations of lovastatin is reversed by cytokinin. Plant Physiol. 1992, 100, 2090–2095. [Google Scholar] [CrossRef] [PubMed]
- Hartig, K.; Beck, E. Assessment of lovastatin application as tool in probing cytokinin-mediated cell cycle regulation. Physiol. Plant. 2005, 125, 260–267. [Google Scholar] [CrossRef]
- Reid, T.S.; Terry, K.L.; Casey, P.J.; Beese, L.S. Crystallographic analysis of CaaX prenyltransferases complexed with substrates defines rules of protein substrate selectivity. J. Mol. Biol. 2004, 343, 417–433. [Google Scholar] [CrossRef]
- Ashok, S.; Hildebrandt, E.R.; Ruiz, C.S.; Hardgrove, D.S.; Coreno, D.W.; Schmidt, W.K.; Hougland, J.L. Protein farnesyltransferase catalyzes unanticipated farnesylation and geranylgeranylation of shortened target sequences. Biochemistry 2020, 59, 1149–1162. [Google Scholar] [CrossRef]
- Kim, J.H.; Hildebrandt, E.R.; Sarkar, A.; Yeung, W.; Waldon, L.R.A.; Kannan, N.; Schmidt, W.K. A comprehensive in vivo screen of yeast farnesyltransferase activity reveals broad reactivity across a majority of CXXX sequences. G3 2023, 13, jkad094. [Google Scholar] [CrossRef]
- Sarkar, A.; Hildebrandt, E.R.; Patel, K.V.; Mai, E.T.; Shah, S.A.; Kim, J.H.; Schmidt, W.K. Comprehensive analysis of CXXX sequence space reveals that Saccharomyces cerevisiae GGTase-I mainly relies on a2X substrate determinants. G3 2024, 14, jkae121. [Google Scholar] [CrossRef]
- Kim, H. Expanding Farnesyltransferase Specificity Beyond the Canonical CaaX Consensus Sequence. Ph.D. Thesis, University of Georgia, Athens, GA, USA, 2023. [Google Scholar]
- Dutilleul, C.; Ribeiro, I.; Blanc, N.; Nezames, C.D.; Deng, X.W.; Zglobicki, P.; Palacio Barrera, A.M.; Atehortùa, L.; Courtois, M.; Labas, V.; et al. ASG2 is a farnesylated DWD protein that acts as ABA negative regulator in Arabidopsis. Plant Cell Environ. 2016, 39, 185–198. [Google Scholar] [CrossRef]
- Ribeiro, I.; Ducos, E.; Giglioli-Guivarc’h, N.; Dutilleul, C. Tagging and capture of prenylated CaaX-proteins from plant cell cultures. Methods Mol. Biol. 2022, 2505, 241–248. [Google Scholar] [CrossRef]
- Blanden, M.J.; Suazo, K.F.; Hildebrandt, E.R.; Hardfrove, D.S.; Patel, M.; Saunders, P.; Distefano, M.D.; Schlidt, W.K.; Hougland, J.L. Efficient farnesylation of an extended C-terminal C(x)3X sequence motif expands the scope of the prenylated proteome. J. Biol. Chem. 2018, 293, 2770–2785. [Google Scholar] [CrossRef] [PubMed]
- Berger, B.M.; Kim, J.H.; Hildebrandt, E.R.; Davis, I.C.; Morgan, M.C.; Hougland, J.L.; Schmidt, W.K. Protein isoprenylation in yeast targets COOH-Terminal sequences not adhering to the CaaX consensus. Genetics 2018, 210, 1301–1316. [Google Scholar] [CrossRef] [PubMed]
- Schey, G.L.; Hildebrandt, E.R.; Wang, Y.; Diwan, S.; Passetti, H.A.; Potts, G.W.; Sprague-Getsy, A.M.; Leoni, E.R.; Kuebler, T.S.; Sham, Y.Y.; et al. Library screening, in vivo confirmation, and structural and bioinformatic analysis of pentapeptide sequences as substrates for protein farnesyltransferase. Int. J. Mol. Sci. 2024, 25, 5324. [Google Scholar] [CrossRef]
- Barghetti, A.; Sjögren, L.; Floris, M.; Paredes, E.B.; Wenkel, S.; Brodersen, P. Heat-shock protein 40 is the key farnesylation target in meristem size control, abscisic acid signaling, and drought resistance. Genes Dev. 2017, 31, 2282–2295. [Google Scholar] [CrossRef]
- Reiser, L.; Subramaniam, S.; Zhang, P.; Berardini, T. Using the Arabidopsis Information Resource (TAIR) to find information about Arabidopsis genes. Curr. Protoc. Bioinform. 2022, 2, e574. [Google Scholar] [CrossRef]
- Galichet, A.; Gruissem, W. Developmentally controlled farnesylation modulates AtNAP1; 1 function in cell proliferation and cell expansion during Arabidopsis leaf development. Plant Physiol. 2006, 142, 1412–1426. [Google Scholar] [CrossRef]
- Kho, Y.; Kim, S.C.; Jiang, C.; Barma, D.; Kwon, S.W.; Cheng, J.; Jaunbergs, J.; Weinbaum, C.; Tamanoi, F.; Falck, J.; et al. A tagging-via-substrate technology for detection and proteomics of farnesylated proteins. Proc. Natl. Acad. Sci. USA 2004, 101, 12479–12484. [Google Scholar] [CrossRef]
- Rikitake, Y. Regulation of the SNARE protein Ykt6 function by diprenylation and phosphorylation. J. Biochem. 2022, 172, 337–340. [Google Scholar] [CrossRef]
- McGrath, K.; Agarwal, S.; Tonelli, M.; Dergai, M.; Gaeta, A.L.; Shum, A.K.; Lacoste, J.; Zhang, Y.; Wen, W.; Chung, D.; et al. A conformational switch driven by phosphorylation regulates the activity of the evolutionarily conserved SNARE Ykt6. Proc. Natl. Acad. Sci. USA 2021, 118, e2016730118. [Google Scholar] [CrossRef]
- Wu, J.R.; Zohra, R.; Duong, N.K.T.; Yeh, C.H.; Lu, C.A.; Wu, S.J. A plant protein farnesylation system in prokaryotic cells reveals Arabidopsis AtJ3 produced and farnesylated in E. coli maintains its function of protecting proteins from heat inactivation. Plant Methods 2023, 19, 113. [Google Scholar] [CrossRef]
- Caldelari, D.; Sternberg, H.; Rodrıguez-Concepción, M.; Gruissem, W.; Yalovsky, S. Efficient prenylation by a plant geranylgeranyltransferase-I requires a functional CaaL box motif and a proximal polybasic domain. Plant Physiol. 2001, 126, 1416–1429. [Google Scholar] [CrossRef]
- Gao, J.; Zhu, Y.; Zhou, W.; Molinier, J.; Dong, A.; Shen, W.H. NAP1 family histone chaperones are required for somatic homologous recombination in Arabidopsis. Plant Cell 2012, 24, 1437–1447. [Google Scholar] [CrossRef]
- Jie, L.; Sanagi, M.; Luo, Y.; Maeda, H.; Fukao, Y.; Chiba, Y.; Yanagisawa, S.; Yamaguchi, J.; Takagi, J.; Sato, T. Histone chaperone NUCLEOSOME ASSEMBLY PROTEIN 1 proteins affect plant growth under nitrogen deficient conditions in Arabidopsis thaliana. Plant Biotech. 2023, 40, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Hougland, J.L.; Gangopadhyay, S.A.; Fierke, C.A. Expansion of protein farnesyltransferase specificity using “tunable” active site interactions: Development of bioengineered prenylation pathways. J. Biol. Chem. 2012, 287, 38090–38100. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Stroh, S.; Eisenhaber, F. Refinement and prediction of prenylation motifs. Genome Biol. 2005, 5, R55. [Google Scholar] [CrossRef]
- Maurer-Stroh, S.; Koranda, M.; Benetka, W.; Schneider, G.; Sirota, F.L.; Eisenhaber, F. Towards complete sets of farnesylated and geranylgeranylated proteins. PloS Comput. Biol. 2007, 3, 634–648. [Google Scholar] [CrossRef]
- Hussain, W.; Khan, Y.D.; Rasool, N.; Khan, S.A.; Chou, K.C. SPrenylC-PseAAC: A sequence-based model developed via Chou’s 5-step rule and general PseAAC for identifying S-prenylation sites in proteins. J. Theoretical Biol. 2019, 468, 1–11. [Google Scholar] [CrossRef]
- Berger, B.M.; Yeung, W.; Goyal, A.; Zhou, Z.; Hildebrandt, E.R.; Kannan, N.; Schmidt, W.K. Functional classification and validation of yeast prenylation motifs using machine learning and genetic reporters. PLoS ONE 2022, 17, e0270128. [Google Scholar] [CrossRef]
- Lavy, M.; Bracha-Drori, K.; Sternberg, H.; Yalovsky, S. A cell-specific, prenylation-independent mechanism regulates targeting of type II RACs. Plant Cell 2002, 14, 2431–2450. [Google Scholar] [CrossRef]
- Zhang, J.; Hill, D.R.; Sylvester, A.W. Diversification of the RAB guanosine triphosphatase family in dicots and monocots. J. Integr. Plant Biol. 2007, 49, 1129–1141. [Google Scholar] [CrossRef]
- Wilkins, J.A.; Kaasik, K.; Chalkley, R.J.; Burlingame, A.L. Characterization of prenylated C-terminal peptides using a thiopropyl-based capture technique and LC-MS/MS. Mol. Cell. Proteom. 2020, 19, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Chowdhury, S.M. Dual-Stage Neutral Loss Tandem Mass Spectrometric strategy for confident identification of protein prenylation. Anal. Chem. 2021, 93, 13169–13176. [Google Scholar] [CrossRef] [PubMed]
- Tsumagari, K.; Isobe, Y.; Imami, K.; Arita, M. Exploring protein lipidation by mass spectrometry-based proteomics. J. Biochem. 2024, 175, 225–233. [Google Scholar] [CrossRef]
- de Abreu-Neto, J.B.; Turchetto-Zolet, A.C.; de Oliveira, L.F.V.; Bodanese Zanettini, M.H.; Margis-Pinheiro, M. Heavy metal-associated isoprenylated plant protein (HIPP): Characterization of a family of proteins exclusive to plants. FEBS J. 2013, 280, 1604–1616. [Google Scholar] [CrossRef]
- Li, J.; Zhang, M.; Sun, J.; Mao, X.; Wang, J.; Liu, H.; Zheng, H.; Li, X.; Zhao, H.; Zou, D. Heavy metal stress-associated proteins in rice and Arabidopsis: Genome-wide identification, phylogenetics, duplication, and expression profiles analysis. Front. Genet. 2020, 11, 477. [Google Scholar] [CrossRef]
- Barth, O.; Vogt, S.; Uhlemann, R.; Zschiesche, W.; Humback, K. Stress induced and nuclear localized HIPP26 from Arabidopsis thaliana interacts via its heavy metal associated domain with the drought stress related zinc finger transcription factor ATHB29. Plant Mol. Biol. 2009, 69, 213–226. [Google Scholar] [CrossRef]
- Gao, W.; Xiao, S.; Tsao, S.W.; Chye, M.L. Arabidopsis thaliana acyl-CoA-binding protein ACBP2 interacts with heavy-metal-binding farnesylated protein AtFP6. New Phytol. 2009, 181, 89–102. [Google Scholar] [CrossRef]
- Barr, Z.K.; Werner, T.; Tilsner, J. Heavy metal-associated isoprenylated plant proteins (HIPPs) at plasmodesmata: Exploring the link between localization and function. Plants 2023, 12, 3015. [Google Scholar] [CrossRef]
- Guo, T.; Weber, H.; Niemann, M.C.; Theisl, L.; Leonte, G.; Novák, O.; Werner, T. Arabidopsis HIPP proteins regulate endoplasmic reticulum-associated degradation of CKX proteins and cytokinin responses. Mol. Plant 2021, 14, 1918–1934. [Google Scholar] [CrossRef]
- Rono, J.K.; Sun, D.; Yang, Z.M. Metallochaperones: A critical regulator of metal homeostasis and beyond. Gene 2022, 822, 146352. [Google Scholar] [CrossRef]
- Lansing, H.; Doering, L.; Fischer, K.; Baune, M.-C.; von Schaewen, A. Analysis of potential redundancy among Arabidopsis 6-phosphogluconolactonase isoforms in peroxisomes. J. Exp. Bot. 2020, 71, 823–836. [Google Scholar] [CrossRef] [PubMed]
- Ku, Y.S.; Cheng, S.S.; Cheung, M.Y.; Law, C.H.; Lam, H.M. The re-localization of proteins to or away from membranes as an effective strategy for regulating stress tolerance in plants. Membranes 2022, 12, 1261. [Google Scholar] [CrossRef] [PubMed]
- Lam, P.Y.; Wang, L.; Lo, C.; Zhu, F.Y. Alternative splicing and its roles in plant metabolism. Int. J. Mol. Sci. 2022, 23, 7355. [Google Scholar] [CrossRef] [PubMed]
- Barff, T.; Sanchez Carrillo, I.B.; Parra Gutiérrez, V.P.; Plourde, M.B.; Joly, D.L.; Germain, H. Comprehensive mapping of Arabidopsis alternative splicing landscape reveals key insights into plant development and immunity. Plant Genome 2025, 18, e70022. [Google Scholar] [CrossRef]
- Endo, M.; Druso, J.E.; Cerione, R.A. The two splice variant forms of Cdc42 exert distinct and essential functions in neurogenesis. J. Biol. Chem. 2020, 295, 4498–4512. [Google Scholar] [CrossRef]
- Kim, M.C.; Chung, W.S.; Yun, D.J.; Cho, M.J. Calcium and calmodulin-mediated regulation of gene expression in plants. Mol. Plant 2009, 2, 13–21. [Google Scholar] [CrossRef]
- Agamasu, C.; Ghirlando, R.; Taylor, T.; Messing, S.; Tran, T.H.; Bindu, L.; Tonelli, M.; Nissley, D.V.; McCormick, F.; Stephen, A.G. KRAS prenylation is required for bivalent binding with calmodulin in a nucleotide-independent manner. Biophys. J. 2019, 116, 1049–1063. [Google Scholar] [CrossRef]
- Fivaz, M.; Meyer, T. Reversible intracellular translocation of KRas but not HRas in hippocampal neurons regulated by Ca2+/calmodulin. J. Cell Biol. 2005, 170, 429–441. [Google Scholar] [CrossRef]
- Rodríguez-Concepción, M.; Yalovsky, S.; Zik, M.; Fromm, H.; Gruissem, W. The prenylation status of a novel plant calmodulin directs plasma membrane or nuclear localization of the protein. EMBO J. 1999, 18, 1996–2007. [Google Scholar] [CrossRef]
- Jamshidiha, M.; Ishida, H.; Sutherland, C.; Gifford, J.L.; Walsh, M.P.; Vogel, H.J. Structural Analysis of a Calmodulin Variant from Rice: The C-terminal extension of OsCaM61 regulates its calcium binding and enzyme activation properties. J. Biol. Chem. 2013, 288, 32036–32049. [Google Scholar] [CrossRef]
- Ranty, B.; Aldon, D.; Galaud, J.-P. Plant calmodulins and calmodulin-related proteins: Multifaceted relays to decode calcium signals. Plant Signal. Behav. 2006, 1, 96–104. [Google Scholar] [CrossRef]
- Kushwaha, R.; Singh, A.; Chattopadhyay, S. Calmodulin7 plays an important role as transcriptional regulator in Arabidopsis seedling development. Plant Cell 2008, 20, 1747–1759. [Google Scholar] [CrossRef]
- Dutta, S.; Basu, R.; Pal, A.; Kunalika, M.H.; Chattopadhyay, S. The homeostasis of AtMYB4 is maintained by ARA4, HY5, and CAM7 during Arabidopsis seedling development. Plant J. 2024, 120, 2515–2535. [Google Scholar] [CrossRef]
- González-Mendoza, V.; Zurita-Silva, A.; Sánchez-Calderón, L.; Sánchez-Sandoval, M.E.; Oropeza-Aburto, A.; Gutiérrez-Alanís, D.; Alatorre-Cobos, F.; Herrera-Estrella, L. APSR1, a novel gene required for meristem maintenance, is negatively regulated by low phosphate availability. Plant Sci. 2013, 205, 2–12. [Google Scholar] [CrossRef]
- Pantoja-Guerra, M.; Valero-Valero, N.; Ramírez, C.A. Total auxin level in the soil-plant system as a modulating factor for the effectiveness of PGPR inocula: A review. Chem. Biol. Technol. Agric. 2023, 10, 6. [Google Scholar] [CrossRef]
- Li, H.; Hu, J.; Pang, J.; Zhao, L.; Yang, B.; Kang, X.; Wang, A.; Xu, T.; Yang, Z. Rho GTPase ROP1 interactome analysis reveals novel ROP1-associated pathways for pollen tube polar growth in Arabidopsis. Int. J. Mol. Sci. 2020, 21, 7033. [Google Scholar] [CrossRef] [PubMed]
- Ohad, N.; Shichrur, K.; Yalovsky, S. The analysis of protein-protein interactions in plants by bimolecular fluorescence complementation. Plant Physiol. 2007, 145, 1090–1099. [Google Scholar] [CrossRef] [PubMed]
- Surana, K.R.; Pawar, R.B.; Khairnar, R.A.; Mahajan, S.K. Protein prenylation and their applications. In Modifications in Biomacromolecules; Zhan, X., Jabbari, A., Eds.; IntechOpen: London, UK, 2024. [Google Scholar] [CrossRef]
- Venkatachalapathy, S.; Lichtenfels, C.; Wagner, C.R.; Distefano, M.D. Engineering protein prenylation: An emerging tool for selective protein modification. Biochem. Soc. Trans. 2025, 53, 1129–1149. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chevalier, Q.; Debié, P.; Huchelmann, A.; Hemmerlin, A. Protein Prenylation Makeovers in Plants: Insights into Substrate Diversification. Int. J. Mol. Sci. 2025, 26, 10638. https://doi.org/10.3390/ijms262110638
Chevalier Q, Debié P, Huchelmann A, Hemmerlin A. Protein Prenylation Makeovers in Plants: Insights into Substrate Diversification. International Journal of Molecular Sciences. 2025; 26(21):10638. https://doi.org/10.3390/ijms262110638
Chicago/Turabian StyleChevalier, Quentin, Pauline Debié, Alexandre Huchelmann, and Andréa Hemmerlin. 2025. "Protein Prenylation Makeovers in Plants: Insights into Substrate Diversification" International Journal of Molecular Sciences 26, no. 21: 10638. https://doi.org/10.3390/ijms262110638
APA StyleChevalier, Q., Debié, P., Huchelmann, A., & Hemmerlin, A. (2025). Protein Prenylation Makeovers in Plants: Insights into Substrate Diversification. International Journal of Molecular Sciences, 26(21), 10638. https://doi.org/10.3390/ijms262110638





