Abstract
Background: Prior studies illustrate the presence and clinical importance of detecting Aspergillus species in the airways of patients with chronic respiratory disease. Despite this, a low fungal biomass and the presence of PCR inhibitors limits the usefulness of quantitative PCR (qPCR) for accurate absolute quantification of Aspergillus in specimens from the human airway. Droplet digital PCR (ddPCR) however, presents an alternative methodology allowing higher sensitivity and accuracy of such quantification but remains to be evaluated in head-to-head fashion using specimens from the human airway. Here, we implement a standard duplex TaqMan PCR protocol, and assess if ddPCR is superior in quantifying airway Aspergillus when compared to standard qPCR. Methods: The molecular approaches of qPCR and ddPCR were applied to DNA fungal extracts in n = 20 sputum specimens obtained from non-diseased (n = 4), chronic obstructive pulmonary disease (COPD; n = 8) and non-cystic fibrosis bronchiectasis (n = 8) patients where Aspergillus status was known. DNA was extracted and qPCR and ddPCR performed on all specimens with appropriate controls and head-to-head comparisons performed. Results: Standard qPCR and ddPCR were both able to detect, even at low abundance, Aspergillus species (Aspergillus fumigatus - A. fumigatus and Aspergillus terreus - A. terreus) from specimens known to contain the respective fungi. Importantly, however, ddPCR was superior for the detection of A. terreus particularly when present at very low abundance and demonstrates greater resistance to PCR inhibition compared to qPCR. Conclusion: ddPCR has greater sensitivity for A. terreus detection from respiratory specimens, and is more resistant to PCR inhibition, important attributes considering the importance of A. terreus species in chronic respiratory disease states such as bronchiectasis.
1. Introduction
Quantitative PCR (qPCR) is an adaptation of standard PCR permitting the detection and real-time quantification of specific target amplification products. A target DNA sequence is selectively amplified using sequence-specific primers, a reporter and quencher labeled dual fluorochrome, an oligonucleotide hybridization probe and a Taq DNA polymerase enzyme. During amplification, the probe specifically hybridizes the accumulating product, and the endonuclease activity of the Taq DNA polymerase cleaves reporter-labeled nucleotides resulting in detectable fluorescence. Reactions are characterized by the time duration during the standard 40 cycles of qPCR amplification where a threshold of baseline fluorescence (Cqs) is exceeded. TaqMan qPCR is established as a useful method for the detection and identification of Aspergillus species in clinical samples including the airway [1,2,3,4,5]. Using these and other next-generation sequencing approaches, our group has demonstrated high levels of airway Aspergillus in patients with bronchiectasis where higher qPCR-derived A. fumigatus and A. terreus is associated with poorer clinical outcome [1,3,6]. Importantly, however, to determine absolute quantification of Aspergillus 18S rRNA, a serial dilution of plasmid containing 18S DNA is necessary for the generation of a standard curve on each plate, a time consuming and costly process limiting the specimens that can be studied. In addition, optimization of the employed standard curves is required, which in itself, demonstrates dynamic and differing ranges for the absolute quantification of Aspergillus species [7]. The results of even standard and test specimens may vary based on reaction efficiencies and differences in specimen content including the presence of inhibitors [8,9]. For all these reasons, an improved and alternative method may be beneficial.
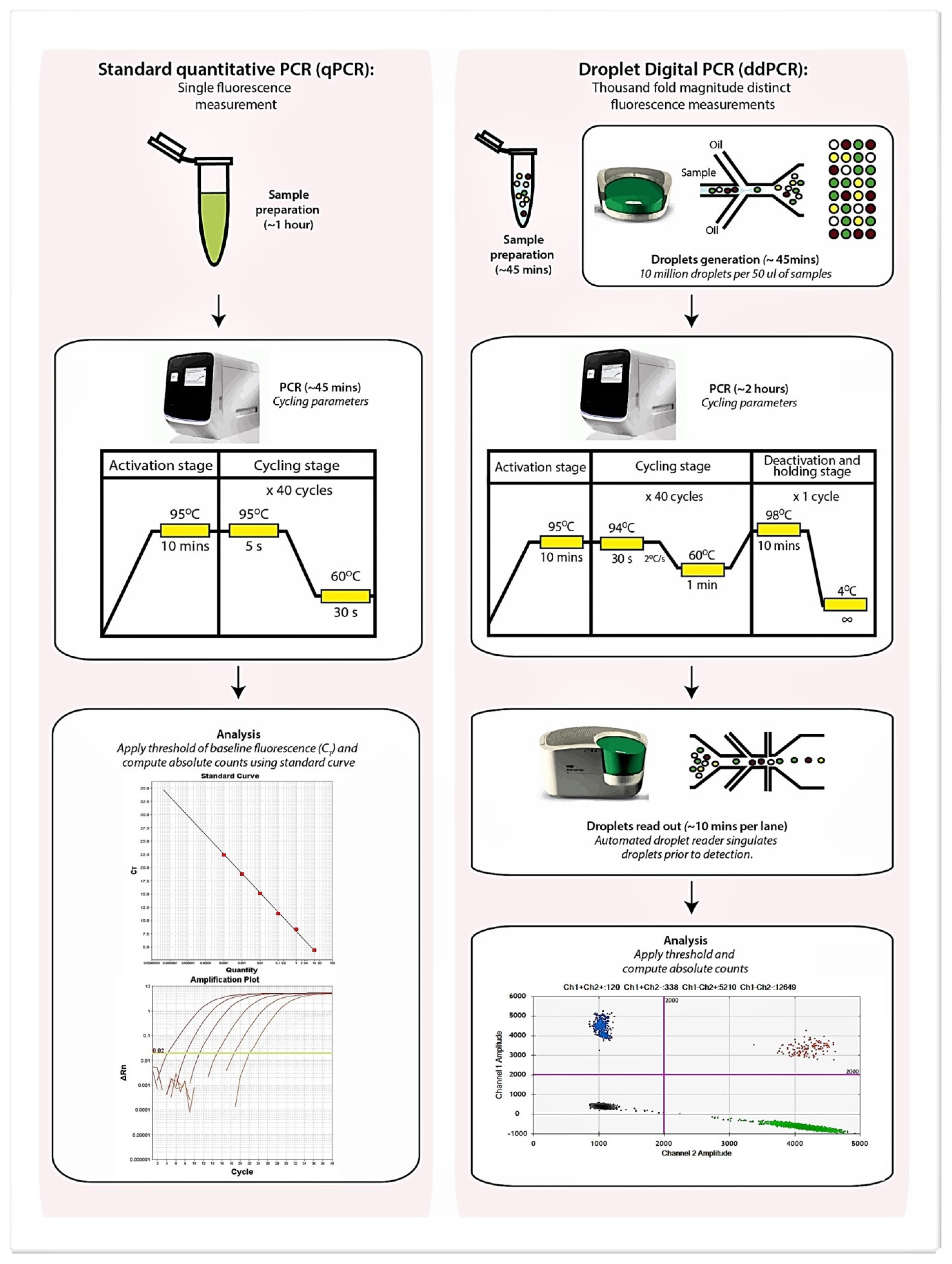
Recently, droplet digital PCR (ddPCR) has been developed and could potentially circumvent issues associated with qPCR [10,11,12]. This technique, based on partitioning the PCR reaction mix into a thousand-fold magnitudes smaller and segregated reaction droplets allows amplification of the respective target(s) within each individual droplet which is then quantified by a target-dependent fluorescence signal (Figure 1). The digital aspect of this approach relies on distributing the target gene into a significant number of partitions (or droplets) such that each receives a number of genes (i.e., 0, 1, 2, etc.). Performing PCR on such partitions results in the amplification being labeled positive (in those containing the target) or negative (no amplification). As positive readouts potentially contain more than a single gene copy of the target molecule, a simple summing of the number of positives will not yield the correct number of target molecules that may be present. Therefore, Poisson statistics are applied in ddPCR to estimate the total number of target molecules present within an interrogated specimen and avoids the need for reference to a standard curve [10,11,12]. As ddPCR represents an end-point PCR reaction, data are unaffected by variations to reaction efficiency and the absolute copy number of the target genes can be determined with confidence so long as the fluorescence readout is correctly partitioned to positive and negative droplets. The high precision and accuracy of ddPCR further reduces the need for technical replicates which improves experimental throughput, saves time, and effectively permits accurate quantification of targets in low volume human specimens such as that from the airway [10,11,12]. Table 1 summarizes the comparisons between qPCR and ddPCR.
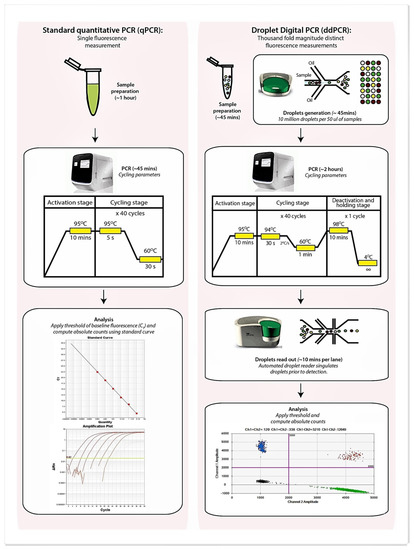
Figure 1.
Schematic diagram illustrating protocol related differences between qPCR and droplet digital PCR (ddPCR) including the estimated time required at each step. Sample preparation for both qPCR and ddPCR while comparable is slightly longer for qPCR due to a requirement for standard curve preparation to allow quantification and inclusion of an internal positive control to exclude PCR inhibition.

Table 1.
Summary of comparison between qPCR and ddPCR.
In this study, we evaluate the accuracy and sensitivity of ddPCR (as compared to qPCR) in detecting Aspergillus species in the normal and diseased airway from respiratory specimens. To allow concurrent detection of A. fumigatus and A. terreus in our samples, we developed duplex primers and probes that allow interrogation of both species by ddPCR.
2. Results
2.1. Assessment for the Specificity of the A. fumigatus and A. terreus Duplex TaqMan Primer and Probe Set Used for ddPCR Evaluation
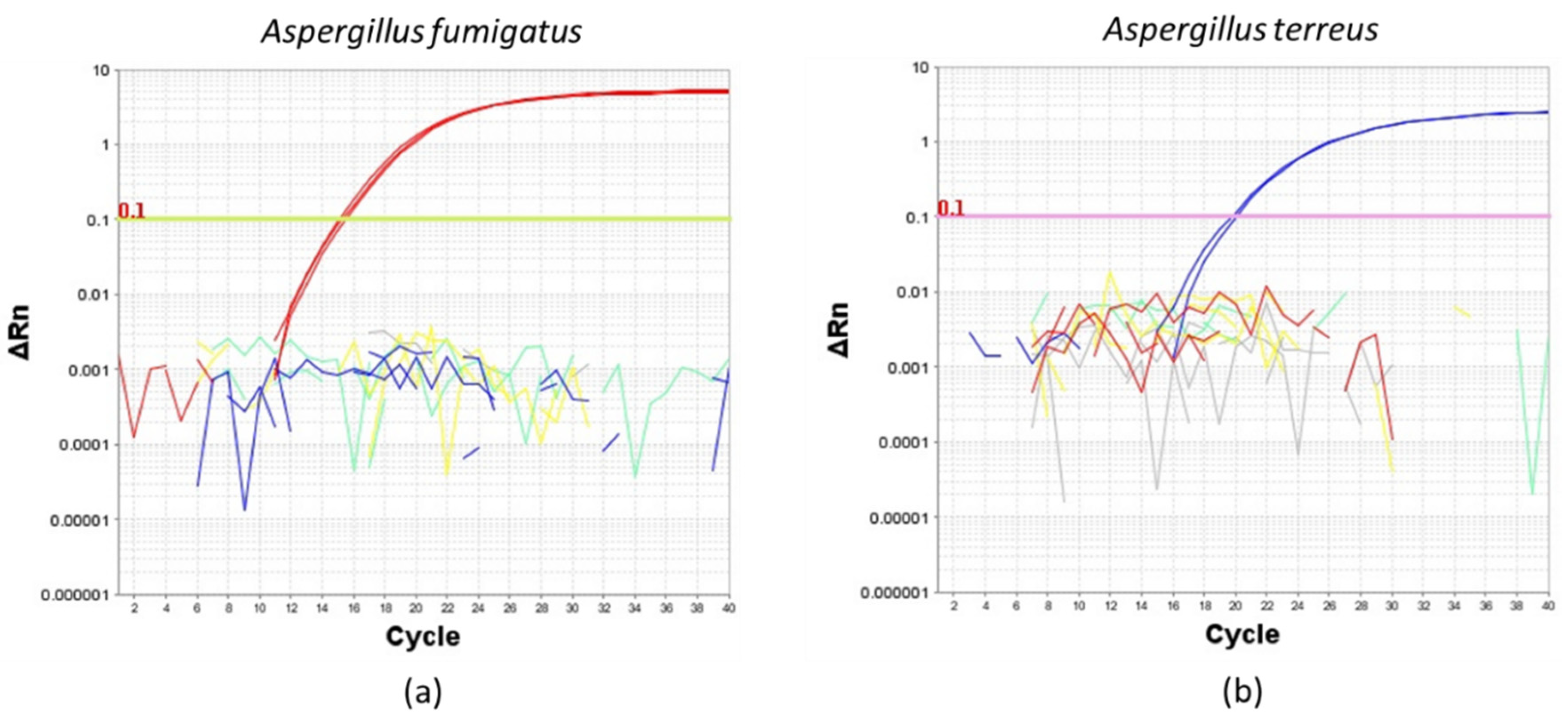
To determine the specificity of the TaqMan primers and probes, we first amplified A. fumigatus or A.terreus from conidial DNA extracted from four common Aspergillus species isolated from the human airway, namely A. fumigatus, A. terreus, A. flavus and A. niger (Figure 2). Importantly, A. fumigatus and A. terreus TaqMan primers and probes only amplified their respective DNA extracts and not extracts obtained from other species confirming the specificity for the primers and probes to be used for the evaluation of ddPCR (Figure 2).
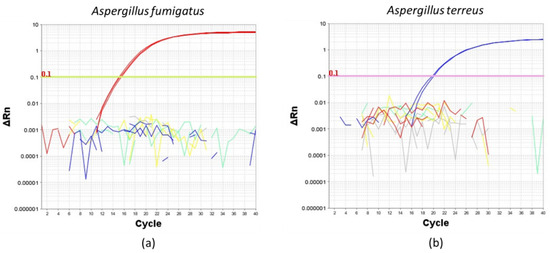
Figure 2.
Amplification plots illustrating the specificity of (a) Aspergillus fumigatus and (b) Aspergillus terreus TaqMan primers and probes using a reaction mix containing 0.1 ng of extracted DNA from pure cultures of (a) Aspergillus fumigatus (red line), (b) Aspergillus terreus (blue line), Aspergillus flavus (yellow line) and Aspergillus niger (green line). No template control is indicated by gray lines. ∆Rn: delta normalized reported value.
2.2. Evaluation of Limits for the Detection and Quantification of A. fumigatus and A. terreus Using TaqMan Duplex Sets by qPCR and ddPCR
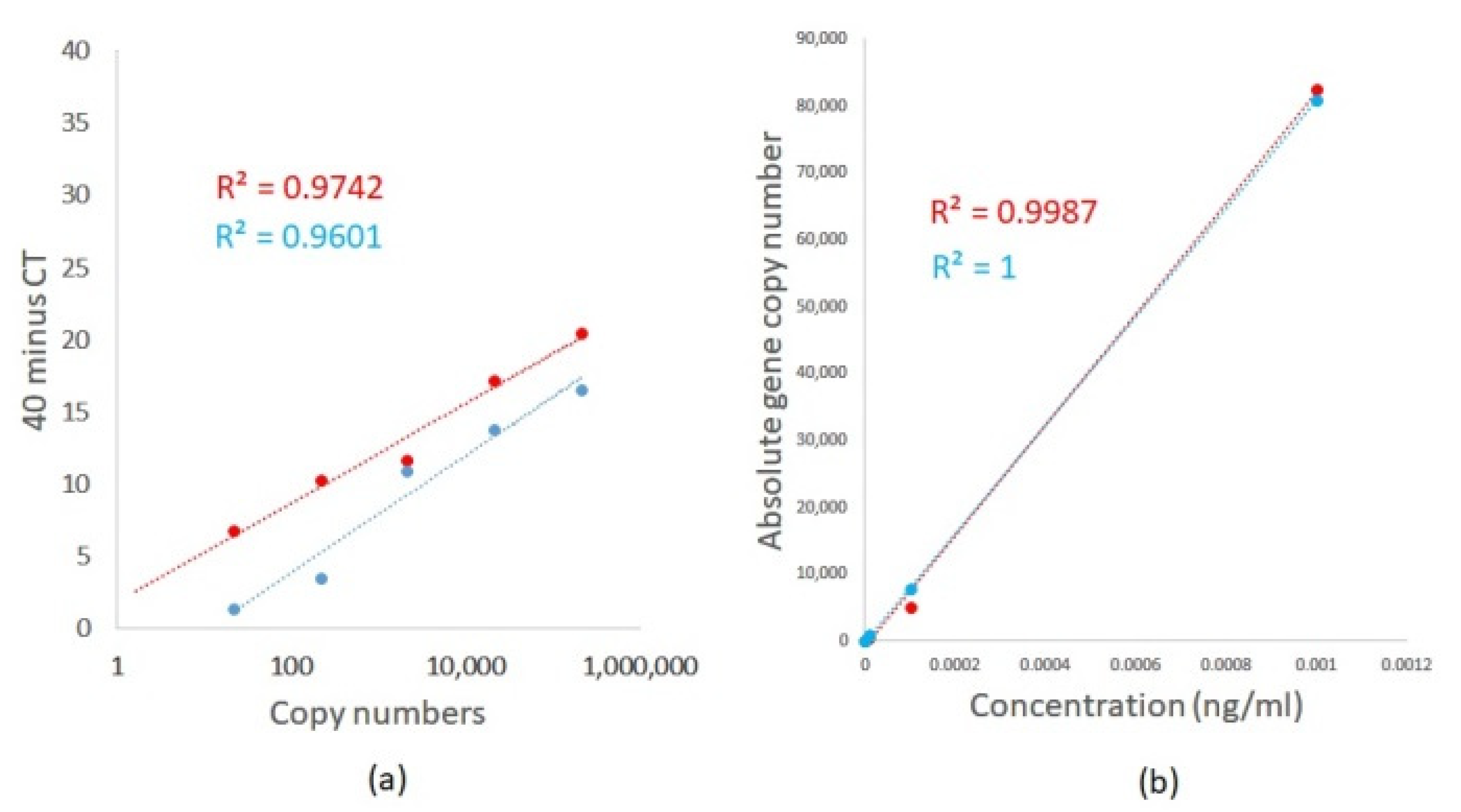
The lower limit for the detection and quantification of A. fumigatus and A. terreus was determined using TOPO plasmids containing the 18S ITS DNA sequence of each fungal species. Plasmid copy number was calculated based on plasmid concentration and sequence using the DNA/RNA copy number calculator (http://endmemo.com/bio/dnacopynum.php). TaqMan qPCR was first performed using a 10-fold serially diluted Aspergillus species 18S ITS DNA plasmid from 1 pg/mL to 0.1 fg/mL. Table 2 illustrates the results. We detected femtogram concentrations of plasmid DNA with an R2 of 0.9742 and 0.9601, respectively, for A. fumigatus and A. terreus using TaqMan qPCR (Figure 3a). Aspergillus TaqMan qPCR demonstrates amplification efficiencies of 102% and 96.5%, respectively, for A. fumigatus and A. terreus with a slope of 3.274 and 3.409, respectively (Figure 3a) [14]. Importantly, differing efficiencies between the respective primer and probe sets were observed (Figure 3a). For comparisons and utilizing the same primer and probe sets, ddPCR was similarly able to detect A. fumigatus and A. terreus, respectively, down to femtogram concentrations of plasmid DNA with superior linear regression R2 of 0.9987 and 1, respectively, for A. fumigatus and A. terreus (Figure 3b). For ddPCR assessment, absolute copy numbers were determined using negative and positive droplets based on a Poisson distribution statistical algorithm without the need for a standard curve (Table 3). Differences were obtained in derived fungal DNA copy numbers between qPCR and ddPCR, whereby qPCR illustrates significantly higher copy numbers (Table 2 and Table 3). Although both approaches have high sensitivity for A. fumigatus and A. terreus detection, a qPCR-based approach likely overestimates absolute fungal burden in comparison to ddPCR as DNA concentrations are estimated based on spectrophotometer measurements.

Table 2.
Table illustrating plasmid concentration, expected absolute plasmid copy number and corresponding 40 minus Cqs results from TaqMan qPCR. Cqs—quantification cycle, ng/mL—nanogram per milliliter, fg/mL—femtogram per milliliter.
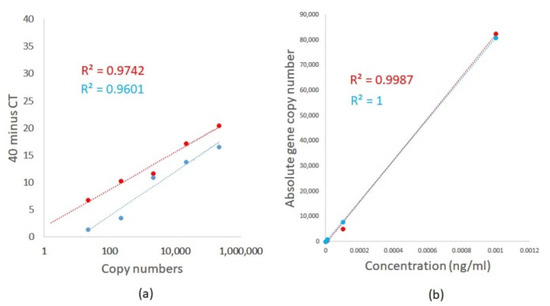
Figure 3.
Limit of detection and absolute quantification of A. fumigatus (red line) and A. terreus (blue line) using (a) TaqMan qPCR and (b) ddPCR. 18S ITS DNA sequences of A. fumigatus and A. terreus were cloned into a TOPO plasmid vector and qPCR and ddPCR performed using 10-fold serial dilutions of plasmid from 1 pg/mL to 0.1 fg/mL. Absolute copy numbers are computed by using known sequence and plasmid concentrations (for TaqMan qPCR) and by identifying negative and positive droplets based on a Poisson distribution statistical algorithm (for ddPCR).

Table 3.
Table illustrating absolute copy numbers of A. fumigatus and A. terreus derived by ddPCR.
2.3. Quantification of Airway A. fumigatus and A. terreus in Respiratory Specimens Obtained from Non-Diseased (Healthy) Individuals and Patients with Chronic Obstructive Pulmonary Disease (COPD) and Bronchiectasis by qPCR and ddPCR
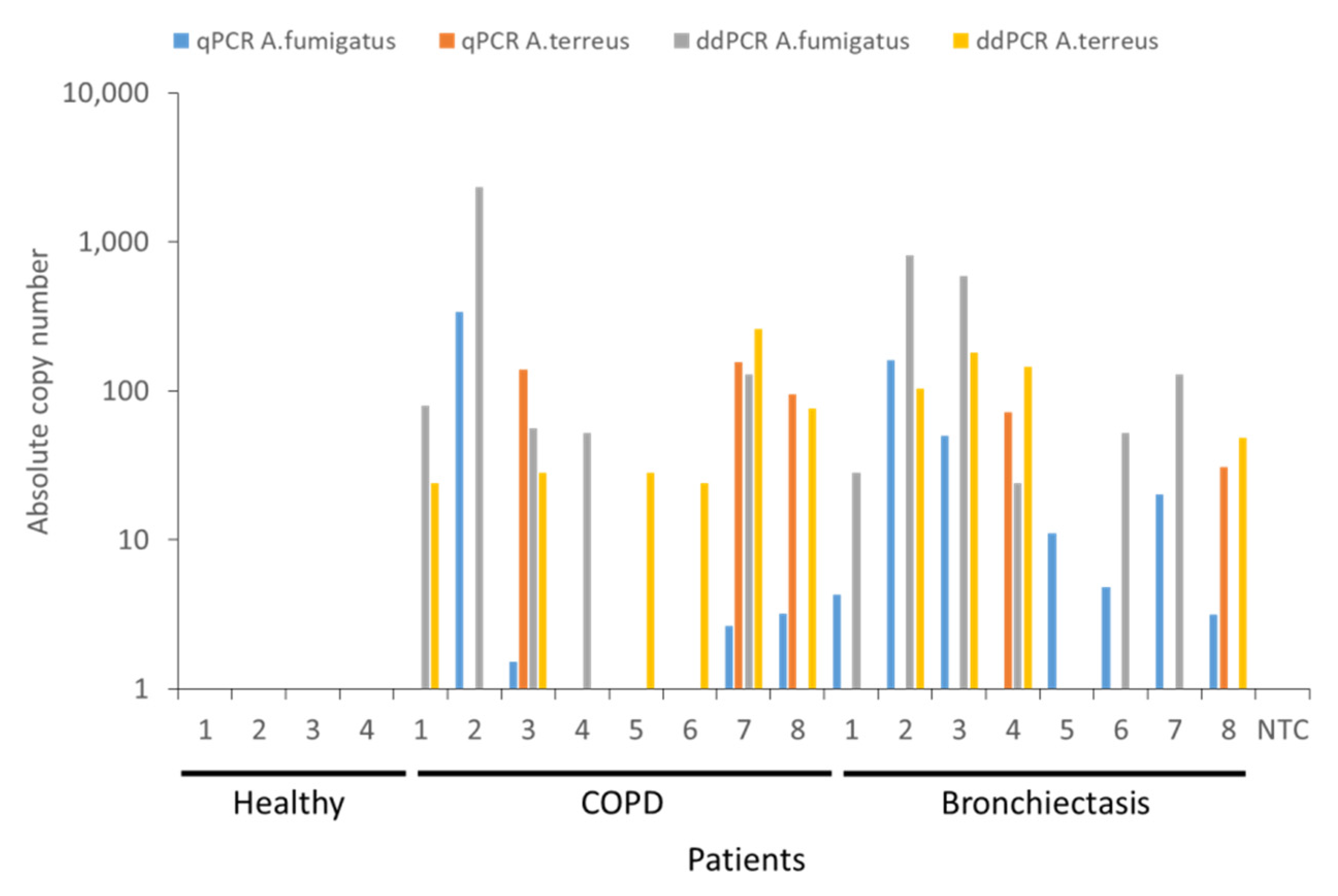
Having identified that ddPCR demonstrates greater sensitivity for the detection of A. fumigatus and A. terreus in earlier experiments, we next evaluated differences in detection and quantification ability using clinical specimens from healthy and diseased individuals. We prospectively recruited n = 20 individuals (n = 4 non-diseased (healthy), n = 8 with COPD and n = 8 with bronchiectasis), the latter diseased groups with known and previously detectable Aspergillus species based on 18S ITS mycobiome sequencing from our ongoing or previously published works [1,3,15]. While we did not detect any A. fumigatus or A. terreus in non-diseased (healthy) individuals, all patients with COPD and bronchiectasis demonstrated detectable A. fumigatus and/or A. terreus in their airway specimens by qPCR and/or ddPCR (Figure 4). All n = 4 healthy samples tested were known to be Aspergillus negative and the n = 16 clinical samples Aspergillus positive by 18S ITS sequencing. Using the qPCR and ddPCR approaches, we identified Aspergillus species in 13 of the 16 clinical samples (81.3%) by qPCR as opposed to 15 out of the 16 samples (93.8%) by ddPCR. Detection limits by qPCR were determined using known concentrations of plasmid containing A. fumigatus or A. terreus 18S sequences and ranged between ~2 and 500 copies while copy numbers for ddPCR ranged between 30 and 3000 copies of A. fumigatus and/or A. terreus, determined by Poisson distribution. No false-positives were detected for the healthy samples and false-positive rates could not be determined for the clinical specimens included as all were known to be Aspergillus positive prior to study entry. However, the false-negative rates were evaluated for clinical specimens and found to be 3 out of 16 (18.8%) for qPCR and only a single sample out of 16 (6.3%) for ddPCR. False-negative rates could not be determined for healthy specimens as all were known to be Aspergillus negative at study entry. Five of the eight COPD (62.5%) and six of the eight (75%) bronchiectasis patients, respectively, demonstrate A. fumigatus by both methods while A. terreus was detectable in 75% (n = 6) and 50% (n = 4) of COPD and bronchiectasis patients, respectively. Importantly, when A. fumigatus was considered alone and compared between qPCR and ddPCR approaches, no single approach was superior in its detection (i.e., COPD: four vs. five and bronchiectasis seven vs. six positive by qPCR and ddPCR, respectively) Importantly, for diseased patients demonstrating lowest airway A. terreus burdens, ddPCR was better than qPCR in identifying them with greater numbers deemed positive through ddPCR compared with the qPCR approach (Figure 4). ddPCR identified twice as many COPD and bronchiectasis patients as A. terreus positive, respectively, when compared with qPCR (i.e., COPD: three vs. six and bronchiectasis two vs. four positives by qPCR and ddPCR, respectively; Figure 4). Therefore, while both qPCR and ddPCR can detect the presence of A. fumigatus and A. terreus in respiratory specimens comparably, ddPCR demonstrated superiority in the detection of A. terreus especially when it was present in ultra-low burden.
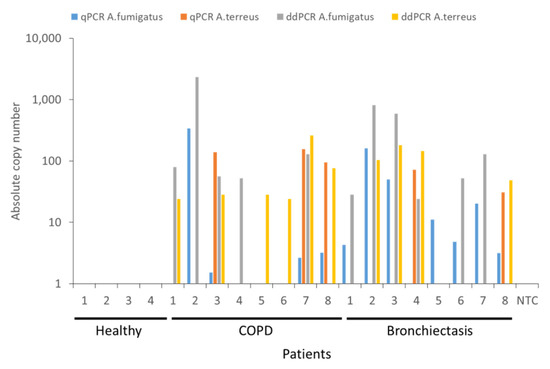
Figure 4.
Head-to-head comparison between qPCR and ddPCR for the detection of A. fumigatus and A. terreus burden in non-diseased (healthy) individuals (n = 4) and patients with COPD (n = 8) or bronchiectasis (B, n = 8). Quantification obtained from TaqMan qPCR is indicated in blue for A. fumigatus and orange for A. terreus, respectively. Quantification by ddPCR is indicated in gray for A. fumigatus) and yellow for A. terreus, respectively. NTC: no template control, Asp: Aspergillus, qPCR: quantitative polymerase chain reaction, ddPCR: droplet digital PCR, COPD: chronic obstructive pulmonary disease.
2.4. Quantification of Airway A. fumigatus by ddPCR is Resistant to PCR Inhibition
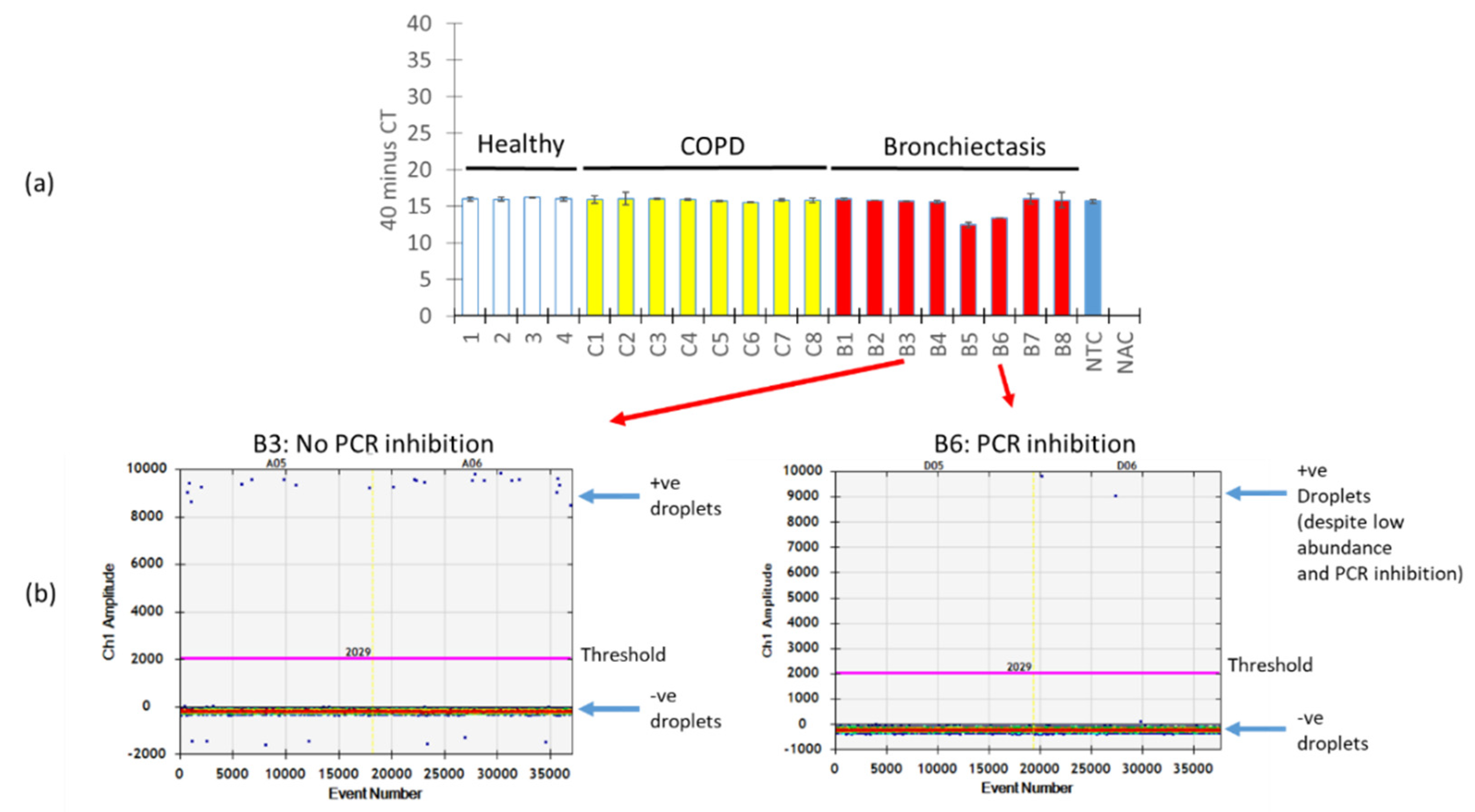
A key issue with PCR amplification using biological specimens is the presence of PCR inhibitors which, in turn, affect the quantification of the target PCR product. We next evaluated how the presence of PCR inhibitors in our respiratory specimens affects the quantification of A. fumigatus using ddPCR versus qPCR. To assess and identify PCR inhibition across our respiratory specimens, we “spiked in” an internal positive control into the mastermix followed by qPCR. Varying degrees of PCR inhibition across the respiratory specimens were noted, in particular for specimens B5 and B6, where measured absolute counts of A. fumigatus would be affected due to the detected Cqs differences translating to an almost two-fold change in expression (Figure 5a). Interestingly, however, when we examined two particular specimens: one demonstrating no PCR inhibition (B3) and the other PCR inhibition (B6), ddPCR was resistant to PCR inhibition, as long as segregation is achieved between its negative and positive fractions (Figure 5b).
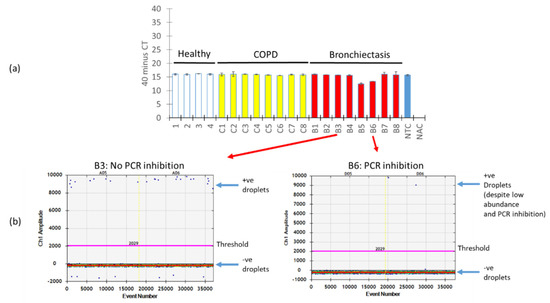
Figure 5.
ddPCR is resistant to PCR inhibition. (a) Respiratory specimens were “spiked” with an internal positive control and subjected to qPCR illustrating various degrees of PCR inhibition most prominent in specimens B5 and B6. (b) One-dimensional amplitude plot from ddPCR illustrating the gating of positive and negative populations (indicated by arrows) that permit an absolute quantification of A. fumigatus despite the presence of PCR inhibitors. Thresholds are indicated by a pink line separating positive and negative fractions from ddPCR. +ve: positive, −ve: negative, NTC: no template control, NAC: no amplification control, qPCR: quantitative polymerase chain reaction, ddPCR: droplet digital PCR, COPD: chronic obstructive pulmonary disease, B: bronchiectasis.
3. Discussion
The study of airway fungal communities, including next-generation sequencing of the pulmonary mycobiome, is gaining clinical and academic interest, particularly in the setting of chronic respiratory disease states such as asthma, COPD, cystic fibrosis and bronchiectasis [1,6,16,17,18,19,20,21,22,23]. Fungi, even in diseased states where dysbiosis is prevalent, demonstrate relatively low abundance at most body sites including the gastrointestinal and respiratory tracts where fungi typically make up less than 5% of the resident microbial community [1,3,24,25,26,27,28,29,30,31]. This low fungal abundance makes 18S ITS targeted amplicon sequencing and qPCR methodological approaches attractive, however, identifying and quantifying fungal load in this manner imposes a lower limit of detection, problematic in cases of ultra-low fungal burdens, and also makes accurate and reliable quantification challenging in the clinical setting. Here, for the first time, we evaluated ddPCR as an alternative to standard qPCR for the detection of Aspergillus species from the human airway. While both methods can detect and quantify Aspergillus species reliably, ddPCR demonstrates greater sensitivity for A. terreus detection and is more resistant to PCR inhibition making it an attractive alternative for the detection of microbes, such as fungi, that occur in low abundance in respiratory specimens but have clinical relevance.
The clinical relevance of low abundance microbes in the airway is well demonstrated in chronic respiratory disease states complicated by the infection such as cystic fibrosis (CF) and bronchiectasis [20,32,33,34,35]. In these settings, the acquisition of a new organism or confirmation that an eradication regime has been effective are clinically critical features for which standard microbiological approaches are lacking. While standard qPCR is useful, it lacks sensitivity below its lower limit of detectable thresholds for individual organisms. In CF and bronchiectasis, the airway ecology is a complex milieu of multiple co-existing organisms from a variety of kingdoms, and hence the ability to detect organisms at ultra-low abundance has value. A clinical example is the identification of allergic bronchopulmonary aspergillosis (ABPA) in CF where likely ultra-low amounts of fungi exist in the airway but are currently either missed or undetected through standard approaches. ddPCR may, therefore, offer an attractive alternative to qPCR in selected clinical settings [20,33,34,35,36,37,38].
One key advantage of ddPCR over traditional TaqMan qPCR is the direct quantification of a target microbe without the need for a standard curve or controls. This leads to improved reproducibility and better accuracy by eliminating reliance on quantitative reference materials whose quantification, source, batch, storage and handling conditions can all influence qPCR results for biological specimens [39]. “Known” quantities of the target, used in TaqMan qPCR standards, as measured by UV spectrophotometry using Nanodrop, employs an indirect quantification approach which potentially affects reliability and accuracy because of quantification assumption uncertainty (i.e., gene copy number per cell and conversions of measured absorption to copy numbers, etc.) [40]. Spectrophotometry quantifies all nucleic acids that absorb at 260 nm which include quantifying DNA and RNA impurities within the standard itself which, when used to quantify the target in samples of unknown quantity, may lead inadvertently to an overestimation.
Our detected higher precision for quantifying a target of ultra-low abundances such as A. terreus and the higher run-to-run reproducibility observed with ddPCR is consistent with the binary nature of digital PCR quantification and the findings of others [10,39,40,41,42]. ddPCR quantifies by counting frequencies of positive endpoint PCRs based on a Poisson distribution, whose quantification is not dependent on variability in PCR amplification efficiency, an issue with TaqMan qPCR. Substrate competition and PCR amplification efficiency are likely explanations for the observed variability between qPCR and ddPCR results for A. fumigatus and A. terreus, respectively, where no superiority of one technique was present for A. fumigatus but the latter (ddPCR) better for A. terreus. A combined effect of the target number, abundance and amplicon length on reaction mix consumption is also greater for multiplex compared to single-plex reactions. Therefore, if any reaction component is limiting, multiplex reactions can show either significant Cqs delays or even total loss of PCR products particularly for targets of lowest abundance. The improved precision attained through ddPCR is a key consideration for respiratory specimens of ultra-low concentrations or where the target is undetectable by TaqMan qPCR.
Our detected tolerance to PCR inhibitors with ddPCR is consistent with its binary nature [43]. PCR inhibitors function either through DNA sequestration or by reducing PCR amplification efficiency, both of which increase Cqs values and lead to underestimation with a TaqMan qPCR approach. Despite reduced amplification (i.e., lower fluorescence intensity due to PCR inhibition), quantification by ddPCR depends on end-point droplet fluorescence which remains higher than the background fluorescence readings. Biological samples including respiratory specimens are often complex and contain PCR inhibitors; therefore, the robust and resistant nature of ddPCR against such inhibitors may be useful in the clinical setting alleviating cost and inconsistent recovery efficiency associated with DNA purification procedures. ddPCR, however, is not fully immune to PCR inhibition, and where severe inhibition exists, ddPCR can experience “total molecular drop-out” where the target remains unamplified [44].
ddPCR permits multiplexing, an additional advantage where clinical material is scarce.
The majority of available Aspergillus-related diagnostic kits do not specifically detect A. fumigatus and A. terreus in biological samples and usually consist of primers and probes that employ a pan- Aspergillus approach which cross-react across several Aspergillus species. While duplexing is more challenging in a TaqMan qPCR approach, largely due to substrate competition and the need for internal positive controls, simultaneous measurements of A. fumigatus and A. terreus are possible with ddPCR due to the individual amplification approach of the generated droplets. ddPCR duplexing while reducing labor, improving logistic arrangements and optimizing data quality through limiting accumulated pipetting errors and the lack of a need for standards does come with the added costs of reagents and consumables unique to ddPCR.
One key drawback, however, of ddPCR is its poor performance with high abundance samples containing >105 gene copy numbers. This is due to the partitioning aspect of this technology, the number of droplets generated and the Poisson distribution algorithm employed for the accurate determination of the number of DNA copies per sample [11]. Therefore, in such circumstances, sample dilution may be required, which poses an additional experimental variability, or one can simply revert to TaqMan qPCR. Sample processing, droplet generation, thermal cycling and droplet analysis further adds additional processing time (~2 h) to the overall process compared to qPCR. The balance between achieving good sensitivity and high accuracy for any test is critically important. Sensitivity is the proportion of true positives that are correctly identified by a diagnostic test while accuracy is the overall proportion of true results, whether true positive or true negative. In our work, sensitivity was high for both qPCR and ddPCR approaches but highest using the latter. Accuracy also remained high for both the healthy and diseased specimens tested in this work however again was highest using the ddPCR approach. Therefore, ddPCR appears to be both highly sensitive and accurate, likely explained by the reaction being performed at the individual droplet level as compared to qPCR where competition for the substrate reagents occurs particularly in duplex assays and these reactions remain subject to PCR inhibition.
While several groups internationally, including ours, have developed qPCR protocols to detect Aspergillus species in biological samples, emerging technologies such as ddPCR offer an alternate, highly sensitive and accurate quantification for samples with ultra-low microbial (fungal) burdens such as that of the airway where detection, even in small amounts has an important clinical consequence and may be potentially missed by qPCR [1,2,3,4,5,7,45]. Attributes such as that offered by ddPCR may be useful in the current era, where microbiomes including the fungal mycobiome, are taking on greater importance and relevance in understanding pathogenesis, disease course and their consequence across a range of respiratory disease states [1,3,27,30,41,42,46,47]. Going beyond fungi, the promise of ddPCR will likely extend across a wide range of microorganisms, organ systems and human disease if it is appropriately applied to the right specimen, in the right setting and to resolve a specific clinical question.
4. Materials and Methods
4.1. Design of Primers and Probes:
Aspergillus
Species-specific assays were designed employing the NCBI primer-blast software using the 18S ITS sequence of Aspergillus fumigatus (accession number NR_121481) and Aspergillus terreus (accession number NR_131276). Primers and probes annealing to sequences of the respective target Aspergillus species were designed to ensure selectivity for no other fungal genera. To ensure that no non-specific interactions between primers and probes occur (for duplex PCR), the Multiple Primer Analyzer (https://www.thermofisher.com) software was employed to exclude the presence of primer-dimers. Cloning of Aspergillus ITS sequences was performed using pan-Aspergillus PCR primers. Primer sequences were derived by homology alignment of the conserved regions between the 18S rRNA genes of the 4 most clinically isolated Aspergillus species (A. fumigatus, A. terreus, A. flavus and A. niger). All primers and probes were synthesized by Bio Basic Asia Pacific Pte Ltd. (Singapore). All sequences of the primers and probes used in this study are shown in Table 4 and specific locations of RT-PCR assay targets in Appendix A.

Table 4.
Summary of all primers and probes used in this study. 6-FAM: 6-Carboxyfluorescein, BHQ1: Black Hole Quencher-1, ITS: internal transcribed spacer, Mod: modification.
4.2. Growth and Harvesting of Fungal Cultures
Fungal conidia were isolated from A. fumigatus and A. terreus, respectively, after growth on Sabouraud agar for 2–7 days, dependent on growth rate. Conidia were harvested by washing the culture with phosphate-buffered saline (PBS) containing 0.05% Tween 20. Conidia were then separated from hyphal fungal elements by filtration through sterile gauze. Following centrifugation at 2000× g for 2 min, conidia were washed twice (with sterile PBS) and resuspended in PBS. Conidia were then used for direct DNA extraction as positive-control material and as sources for cloning experiments.
4.3. Cloning of A. Fumigatus and A. Terreus ITS
A. fumigatus and A. terreus ITS plasmids were employed for absolute quantification and cloned from the conidial extracted DNA using the designed pan-Aspergillus ITS primers (Table 4). PCR was carried out using the KAPA HiFi HotStart ReadyMix (Roche, Basel, Switzerland) which contained 12.5 µL of 2X KAPA HiFi HotStart ReadyMix, 0.5 µM of each primer and 10 ng of Aspergillus DNA in a final volume of 25 µL. Cycling conditions were as follows: 1 cycle at 95 °C for 3 min, 30 cycles at 98 °C for 20 s, 60 °C for 15 s and 72 °C for 15 s with a final cycle at 72 °C for 1 min. PCR products were then gel purified using the Qiagen gel purification kit according to the manufacturer’s instructions. Adenosine overhangs were next added to the PCR product using the 2× PCR Master Mix (Thermo Fisher Scientific, Waltham, MA, United States) with a final volume of 10 µL and cycling at 72 °C for 15 min. PCR products were next cloned using the TOPO™ TA Cloning™ Kit for Sequencing (Invitrogen, Waltham, MA, United States) with One Shot™ TOP10 Chemically Competent E. coli according to the manufacturer’s instructions. Plasmids were then extracted with the Qiagen miniprep kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions and plasmid sequences verified through sequencing (Bio Basic Asia Pacific Pte Ltd., Singapore).
4.4. Study Population
Stable patients (age 21 years or above) with COPD and bronchiectasis were recruited from Singapore General Hospital between March 2016 and July 2018 (Table 1). Stability was defined as the absence of acute symptomatic deterioration (or exacerbation) and/or infection in the preceding four-week period prior to study recruitment. Chronic respiratory disease states were diagnosed based on established international guidelines and are further described below. A separate cohort of non-diseased subjects aged >21 years old with no underlying respiratory disease or other medical history, with normal pulmonary function were recruited through an established program at Nanyang Technological University, Singapore. COPD patients were diagnosed based on the Global Initiative for COPD (GOLD) guidelines and were all > 40 years old, had a smoking history of > 10 packs a year and abnormal lung function defined as forced expiratory volume in one second (FEV1)/forced vital capacity (FVC) ratio < 0.7 and FEV1 < 80% predicted with symptoms defined as presence of cough and/or sputum or breathlessness [48]. Bronchiectasis was confirmed by high-resolution computed tomography (HRCT) and the presence of two or more clinical symptoms of cough, mucopurulent sputum production and shortness of breath and the exclusion of other predominant chronic respiratory disease states [49,50,51]. Patients were excluded if they were pregnant or breastfeeding, had active mycobacterial disease, or were on chemotherapy in any form. Table 5 summarizes the patient demographics.

Table 5.
Demographics of the study population. Data are presented as median and interquartile ranges (IQR) or n (percentage; %). COPD: chronic obstructive pulmonary disease, Post BD: post bronchodilator; BMI: body mass index; BSI: Bronchiectasis Severity Index.
4.5. Ethical Approval
All subjects gave their informed consent for study inclusion. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of NTU and all participating hospitals as follows: CIRB 2016/2715, 2016/2628 and CIRB 2016/2073 (Singapore General Hospital), IRB-2017-03-013, IRB-2016-10-057 and IRB-2016-01-031 (Nanyang Technological University, Singapore).
4.6. Specimen (Sputum) Collection and DNA Extraction
Spontaneously expectorated “representative” sputum from a deep cough was collected and examined [52]. After weighing, the sputum was added to an equal volume of Sputasol (Thermo Fisher Scientific, Waltham, MA, United States) containing 0.1% dithiothreitol and incubated for 15 min at 37 °C. Two volumes of RNAlater (Sigma-Aldrich, St. Louis, MO, United States) were added and samples homogenized before DNA extraction [35]. Sputum DNA was extracted using methods as previously described [35]. Briefly, homogenized sputum in RNAlater were centrifuged (13,000 rpm for 10 min) and pellets resuspended in 500 µL sterile PBS (GE Lifesciences, Marlborough, MA, United States). After transfer to sterile bead mill tubes (VWR) containing acid washed 1 mm sterile glass beads (Sigma-Aldrich, St. Louis, MO, United States), DNA extraction was performed using the Roche High Pure polymerase chain reaction (PCR) Template Preparation Kit (Roche, Basel, Switzerland) according to the manufacturer’s instructions. The ratio of absorbance at 260 nm and 280 nm by Nanodrop was used to assess the purity of DNA. A ratio of ~1.8 is generally accepted as “pure” DNA.
4.7. Quantitative-PCR (qPCR) and Digital Droplet PCR (ddPCR) Detection of Aspergillus Species
The presence of A. fumigatus and A. terreus from human airway specimens were assessed by a probe-based qPCR assay as previously described [1]. An A. fumigatus probe (tagged with a 5′ 6-carboxyfluorescein (FAM) reporter dye and a 3′ Black Hole Quencher 1 (BHQ1) quencher) and A. terreus probe (tagged with a 5′ HEX reporter dye and a 3′ Black Hole Quencher 1 quencher, Bio Basic Asia Pacific, Singapore) were used. qPCR assays with Cqs values of <40 were considered positive. The Cqs cut-off of 40 was used in line with our prior published work with A. fumigatus and terreus species which is previously validated through microbial sequencing approaches [1]. Each TaqMan reaction mix contained 1× PCR master mix (PrimeTime Gene Expression Mastermix, Integrated DNA Technologies Pte. Ltd., Singapore), 900nM of forward and reverse primers each, 250 nM of probe, 1 µL of DNA template; and 2.5 µL of internal positive control (Applied Biosystems® TaqMan® Exogenous Internal Positive Control) to make a final volume of 25 µL. Inclusion of the internal positive control allowed assessment of PCR inhibitors in the DNA extract for each respective airway specimen including the no template control (NTC). Reactions were performed with the Applied Biosystems® MicroAmp® Fast Optical 96-Well Reaction Plate on a StepOne™ Real-Time PCR System (Applied Biosystems, Foster City, CA, United States) under the following conditions: 95 °C for 3 min, followed by 40 cycles of denaturation at 95 °C for 5 s and annealing/extension at 60 °C for 30 s. The same assay was adapted for ddPCR (Bio-Rad QX200, Hercules, CA, United States) excluding the IPS control. For ddPCR, each 25 µL reaction setup contained droplet PCR Supermix (Bio-Rad, Hercules, CA, United States), 900 nmol of each primer, 250 nmol of the probe and 1 µL of sample DNA. The reaction volume was then mixed with droplet generation oil (20 µL mixture with 70 µL oil) via microfluidics using the Bio-Rad Automated Droplet Generator (Bio-Rad, Hercules, CA, United States). The water-in-oil droplets were transferred to 96-well PCR plates and heat-sealed to be run on the Bio-Rad C1000 (Bio-Rad, Hercules, CA, United States) thermocycler (ramping speed at 2 °C/s) for PCR amplification using the following conditions: activation for 10 min at 95 °C, followed by 40 cycles of denaturation for 30 s at 94 °C and annealing/extension for 60 s at 60 °C, followed by a 10 min hold step at 98 °C. On completion, automated measurements of fluorescence for each droplet were determined using the QX200 Droplet Reader (Bio-Rad, Hercules, CA, United States) with the RED (rare event detection) setting. No standard curve was required for ddPCR experiments and the droplets quantified using the Bio-Rad QuantiSoftTM software version 1.7 (Bio-Rad, Hercules, CA, United States). Two replicates per sample were performed and a threshold cut-off of 20,000 employed (based on optimization experiments which accurately separated positive from negative droplets). For both protocols, no template controls comprising DNase- and RNase-free water were included. To ensure a thorough assessment of false-negative results from either qPCR or ddPCR for all included clinical specimens (n = 20), any samples showing no amplification or an amplification discrepancy between qPCR and ddPCR was re-tested for confirmation. All included healthy controls (n = 4) were known to be Aspergillus negative and all included samples from patients with chronic respiratory disease (n = 16) known to be Aspergillus positive by 18S ITS sequencing hence a detailed evaluation of false-positive rates was limited. All results from qPCR and ddPCR assays were compared to these samples of known Aspergillus status. In compliance with established guidelines, supplemental information (Supplementary Table S1) is provided in regards to methodologies used in this study.
4.8. Data Analysis (qPCR)
For TaqMan qPCR, data was analyzed using StepOneTM software (version 2.3, ABI Biosystems, Foster City, CA, United States). Fluorescence thresholds for A. fumigatus and A. terreus were set at 0.1 to obtain Cqs values. Quantification of fungal burden in clinical specimens was based on the derived standard curves.
4.9. Data Analysis (ddPCR)
ddPCR data was analyzed using QuantaSoft™ software (version 1.7 Bio-Rad, Hercules, CA, United States) following the manufacturer’s recommendations. Total droplet number was determined and only wells containing > 10,000 droplets accepted for analysis. Fluorescence thresholds were set to approximately 2SDs (~2000 fluorescence units) above background fluorescence measurement of negative droplets from NTC wells. Target concentrations in copy number per ml reaction were then automatically calculated by QuantaSoft software. Wells with >1 positive droplet were considered positive. The copy number per µL reaction (x) was then determined by multiplication by a factor of 40 as the final reaction mix was 40 µL.
5. Conclusions
ddPCR represents an alternate methodology for the detection and quantification of Aspergillus species in the human airway. Here, we evaluate its sensitivity for quantifying airway Aspergillus burden, for A. fumigatus and A. terreus against a standard TaqMan qPCR approach employing human airway specimens from patients with chronic respiratory disease. While both approaches (standard TaqMan qPCR and ddPCR) can detect these Aspergilli (A. fumigatus and A. terreus), ddPCR importantly demonstrates better sensitivity and greater resistance to PCR inhibition for A. terreus particularly when present at low abundance. Considering the emerging importance of A. terreus species in chronic respiratory disease states such as bronchiectasis, ddPCR represents a useful, viable and reliable alternative to qPCR in such patients.
Supplementary Materials
The following are available online at https://www.mdpi.com/1422-0067/21/9/3043/s1, Table S1: MIQE Guidelines: Essential Information Checklist
Author Contributions
Conceptualization, T.Y.P., N.A.B.M.A. and L.L.Y.C.; methodology, T.Y.P.; formal analysis, T.Y.P.; investigation, P.Y.T. and T.Y.P.; resources, P.Y.T. and S.H.C.; data curation, P.Y.T. and T.Y.P.; writing—original draft preparation, T.Y.P., P.Y.T., N.A.B.M.A. and L.L.Y.C.; S.H.C. writing—review and editing, T.Y.P. and S.H.C.; visualization, T.Y.P. and N.A.B.M.A.; supervision, S.H.C.; project administration, T.Y.P.; funding acquisition, S.H.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research is supported by the Singapore Ministry of Health’s National Medical Research Council under its Clinician-Scientist Individual Research Grant (MOH-000141, S.H.C.) and the NTU Integrated Medical, Biological and Environmental Life Sciences (NIMBELS), Nanyang Technological University, Singapore [NIM/03/2018] (S.H.C.). P.Y.T. is supported by Singapore Ministry of Health’s National Medical Research Council under its Research Training Fellowship (NMRC/Fellowship/0049/2017). L.L.Y.C. is supported by Lee Kong Chian School of Medicine Dean’s Postdoctoral Fellowship & Wong Peng Onn Postdoctoral Fellowship (2019).
Acknowledgments
The authors would like to acknowledge The Academic Respiratory Initiative for Pulmonary Health (TARIPH) for collaboration support.
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Abbreviations
| % | percentage |
| +ve | positive |
| < | less than |
| > | greater than |
| ± | plus/minus |
| ∆Rn | delta normalized reporter value |
| °C | degrees Celsius |
| µL | microliter |
| µM | micro-molar |
| 6-FAM | 6-carboxyfluorescein |
| Asp | Aspergillus |
| BHQ1 | Black Hole Quencher 1 |
| BLAST | Basic Local Alignment Search Tool |
| BMI | body mass index |
| B | bronchiectasis |
| BSI | Bronchiectasis Severity Index |
| CAT | COPD assessment test |
| COPDCF | chronic obstructive pulmonary diseaseCystic Fibrosis |
| Cqs | Quantification cycle |
| ddPCR | droplet digital PCR |
| DNA | deoxyribonucleic acid |
| FEV1 | forced expiratory volume in the first second |
| Fg | femtogram |
| FVC | forced vital capacity |
| g | gravitational force |
| HRCT | high-resolution computed tomography |
| IPC | internal positive control |
| IQR | interquartile range |
| ITS | internal transcribed spacer |
| mins | Minutes |
| Mod | modification |
| N | sample size |
| next-gen | next-generation |
| ng | nanogram |
| nM | nano-molar |
| No. | Number |
| PBS | phosphate-buffered saline |
| pg | picogram |
| Post BD | post bronchodilator |
| Pte Ltd. | Private Limited |
| qPCR | quantitative polymerase chain reaction |
| RNAi | RNA interference |
| rpm | rounds per minute |
| rRNA | ribosomal ribonucleic acid |
| Secs | Seconds |
| siRNA | small interfering ribonucleic acid |
| SNP | single nucleotide polymorphism |
| Taq | Thermus aquaticus |
| -ve | Negative |
Appendix A
Alignment of four clinically important Aspergillus species. TaqMan primers (blue) and probes (red) are shown by their respective coloration. Pan-Aspergillus primers are indicated by boxes.


References
- Mac Aogain, M.; Chandrasekaran, R.; Lim, A.Y.H.; Low, T.B.; Tan, G.L.; Hassan, T.; Ong, T.H.; Ng, A.H.Q.; Bertrand, D.; Koh, J.Y.; et al. Immunological corollary of the pulmonary mycobiome in bronchiectasis: The CAMEB study. Eur. Respir. J. 2018, 52, 1800766. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.J.; Wissel, M.C.; Grantham, K.J.; Petraitiene, R.; Petraitis, V.; Kasai, M.; Francesconi, A.; Cotton, M.P.; Hughes, J.E.; Greene, L.; et al. Molecular detection and species-specific identification of medically important Aspergillus species by real-time PCR in experimental invasive pulmonary aspergillosis. J. Clin. Microbiol. 2011, 49, 4150–4157. [Google Scholar] [CrossRef] [PubMed]
- Poh, T.Y.; Tiew, P.Y.; Lim, A.Y.H.; Thng, K.X.; Ali, N.A.B.M.; Narayana, J.K.; Aogáin, M.M.; Tien, Z.; Chew, W.M.; Chan, A.K.W.; et al. Increased chitotriosidase is associated with Aspergillus and frequent exacerbations in south-east asians with bronchiectasis. CHEST 2020. [Google Scholar] [CrossRef]
- White, P.L.; Barnes, R.A.; Springer, J.; Klingspor, L.; Cuenca-Estrella, M.; Morton, C.O.; Lagrou, K.; Bretagne, S.; Melchers, W.J.; Mengoli, C.; et al. Clinical Performance of Aspergillus PCR for Testing Serum and Plasma: A Study by the European Aspergillus PCR Initiative. J. Clin. Microbiol. 2015, 53, 2832–2837. [Google Scholar] [CrossRef] [PubMed]
- Buess, M.; Cathomas, G.; Halter, J.; Junker, L.; Grendelmeier, P.; Tamm, M.; Stolz, D. Aspergillus-PCR in bronchoalveolar lavage for detection of invasive pulmonary aspergillosis in immunocompromised patients. BMC Infect. Dis. 2012, 12, 237. [Google Scholar] [CrossRef]
- Mac Aogain, M.; Tiew, P.Y.; Lim, A.Y.H.; Low, T.B.; Tan, G.L.; Hassan, T.; Ong, T.H.; Pang, S.L.; Lee, Z.Y.; Gwee, X.W.; et al. Distinct “Immunoallertypes” of Disease and High Frequencies of Sensitization in Non-Cystic Fibrosis Bronchiectasis. Am. J. Respir. Crit. Care Med. 2019, 199, 842–853. [Google Scholar] [CrossRef]
- Johnson, G.L.; Bibby, D.F.; Wong, S.; Agrawal, S.G.; Bustin, S.A. A MIQE-compliant real-time PCR assay for Aspergillus detection. PLoS ONE 2012, 7, e40022. [Google Scholar] [CrossRef]
- Brankatschk, R.; Bodenhausen, N.; Zeyer, J.; Burgmann, H. Simple absolute quantification method correcting for quantitative PCR efficiency variations for microbial community samples. Appl. Environ. Microbiol. 2012, 78, 4481–4489. [Google Scholar] [CrossRef]
- Bustin, S.A.; Nolan, T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech. 2004, 15, 155–166. [Google Scholar]
- Hindson, B.J.; Ness, K.D.; Masquelier, D.A.; Belgrader, P.; Heredia, N.J.; Makarewicz, A.J.; Bright, I.J.; Lucero, M.Y.; Hiddessen, A.L.; Legler, T.C.; et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 2011, 83, 8604–8610. [Google Scholar] [CrossRef]
- Pinheiro, L.B.; Coleman, V.A.; Hindson, C.M.; Herrmann, J.; Hindson, B.J.; Bhat, S.; Emslie, K.R. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal. Chem. 2012, 84, 1003–1011. [Google Scholar] [CrossRef]
- Mazaika, E.; Homsy, J. Digital Droplet PCR: CNV Analysis and Other Applications. Curr. Protoc. Hum. Genet. 2014, 82, 7.24.1–7.24.13. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Q.; Bhattacharya, S.; Kotsopoulos, S.; Olson, J.; Taly, V.; Griffiths, A.D.; Link, D.R.; Larson, J.W. Multiplex digital PCR: Breaking the one target per color barrier of quantitative PCR. Lab Chip 2011, 11, 2167–2174. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Ali, N.; Mac Aogain, M.; Morales, R.F.; Tiew, P.Y.; Chotirmall, S.H. Optimisation and Benchmarking of Targeted Amplicon Sequencing for Mycobiome Analysis of Respiratory Specimens. Int. J. Mol. Sci. 2019, 20, 4991. [Google Scholar] [CrossRef]
- Cui, L.; Morris, A.; Ghedin, E. The human mycobiome in health and disease. Genome Med. 2013, 5, 63. [Google Scholar] [CrossRef]
- El-Jurdi, N.; Ghannoum, M.A. The Mycobiome: Impact on Health and Disease States. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- Suhr, M.J.; Hallen-Adams, H.E. The human gut mycobiome: Pitfalls and potentials--a mycologist’s perspective. Mycologia 2015, 107, 1057–1073. [Google Scholar] [CrossRef]
- Tiew, P.Y.; Mac Aogain, M.; Ali, N.; Thng, K.X.; Goh, K.; Lau, K.J.X.; Chotirmall, S.H. The Mycobiome in Health and Disease: Emerging Concepts, Methodologies and Challenges. Mycopathologia 2020. [Google Scholar] [CrossRef]
- Bell, S.C.; Mall, M.A.; Gutierrez, H.; Macek, M.; Madge, S.; Davies, J.C.; Burgel, P.R.; Tullis, E.; Castanos, C.; Castellani, C.; et al. The future of cystic fibrosis care: A global perspective. Lancet Respir. Med. 2020, 8, 65–124. [Google Scholar] [CrossRef]
- Chalmers, J.D.; Chotirmall, S.H. Bronchiectasis: New therapies and new perspectives. Lancet Respir. Med. 2018, 6, 715–726. [Google Scholar] [CrossRef]
- Goh, K.J.; Yii, A.C.A.; Lapperre, T.S.; Chan, A.K.; Chew, F.T.; Chotirmall, S.H.; Koh, M.S. Sensitization to Aspergillus species is associated with frequent exacerbations in severe asthma. J. Asthma Allergy 2017, 10, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Chotirmall, S.H.; Gellatly, S.L.; Budden, K.F.; Mac Aogain, M.; Shukla, S.D.; Wood, D.L.; Hugenholtz, P.; Pethe, K.; Hansbro, P.M. Microbiomes in respiratory health and disease: An Asia-Pacific perspective. Respirology 2017, 22, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Auchtung, T.A.; Fofanova, T.Y.; Stewart, C.J.; Nash, A.K.; Wong, M.C.; Gesell, J.R.; Auchtung, J.M.; Ajami, N.J.; Petrosino, J.F. Investigating Colonization of the Healthy Adult Gastrointestinal Tract by Fungi. mSphere 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Tipton, L.; Ghedin, E.; Morris, A. The lung mycobiome in the next-generation sequencing era. Virulence 2017, 8, 334–341. [Google Scholar] [CrossRef]
- Nguyen, L.D.; Viscogliosi, E.; Delhaes, L. The lung mycobiome: An emerging field of the human respiratory microbiome. Front. Microbiol. 2015, 6, 89. [Google Scholar] [CrossRef]
- Cui, L.; Lucht, L.; Tipton, L.; Rogers, M.B.; Fitch, A.; Kessinger, C.; Camp, D.; Kingsley, L.; Leo, N.; Greenblatt, R.M.; et al. Topographic diversity of the respiratory tract mycobiome and alteration in HIV and lung disease. Am. J. Respir. Crit. Care Med. 2015, 191, 932–942. [Google Scholar] [CrossRef]
- Hager, C.L.; Ghannoum, M.A. The mycobiome in HIV. Curr. Opin. HIV AIDS 2018, 13, 69–72. [Google Scholar] [CrossRef]
- McTaggart, L.R.; Copeland, J.K.; Surendra, A.; Wang, P.W.; Husain, S.; Coburn, B.; Guttman, D.S.; Kus, J.V. Mycobiome Sequencing and Analysis Applied to Fungal Community Profiling of the Lower Respiratory Tract During Fungal Pathogenesis. Front. Microbiol. 2019, 10, 512. [Google Scholar] [CrossRef]
- Chotirmall, S.H.; Al-Alawi, M.; Mirkovic, B.; Lavelle, G.; Logan, P.M.; Greene, C.M.; McElvaney, N.G. Aspergillus-associated airway disease, inflammation, and the innate immune response. Biomed. Res. Int. 2013, 2013, 723129. [Google Scholar] [CrossRef]
- Budden, K.F.; Shukla, S.D.; Rehman, S.F.; Bowerman, K.L.; Keely, S.; Hugenholtz, P.; Armstrong-James, D.P.H.; Adcock, I.M.; Chotirmall, S.H.; Chung, K.F.; et al. Functional effects of the microbiota in chronic respiratory disease. Lancet Respir. Med. 2019, 7, 907–920. [Google Scholar] [CrossRef]
- Chotirmall, S.H.; Martin-Gomez, M.T. Aspergillus Species in Bronchiectasis: Challenges in the Cystic Fibrosis and Non-cystic Fibrosis Airways. Mycopathologia 2018, 183, 45–59. [Google Scholar] [CrossRef] [PubMed]
- Chotirmall, S.H.; McElvaney, N.G. Fungi in the cystic fibrosis lung: Bystanders or pathogens? Int. J. Biochem. Cell Biol. 2014, 52, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Coughlan, C.A.; Chotirmall, S.H.; Renwick, J.; Hassan, T.; Low, T.B.; Bergsson, G.; Eshwika, A.; Bennett, K.; Dunne, K.; Greene, C.M.; et al. The effect of Aspergillus fumigatus infection on vitamin D receptor expression in cystic fibrosis. Am. J. Respir. Crit. Care Med. 2012, 186, 999–1007. [Google Scholar] [CrossRef]
- Leung, J.M.; Tiew, P.Y.; Mac Aogain, M.; Budden, K.F.; Yong, V.F.; Thomas, S.S.; Pethe, K.; Hansbro, P.M.; Chotirmall, S.H. The role of acute and chronic respiratory colonization and infections in the pathogenesis of COPD. Respirology 2017, 22, 634–650. [Google Scholar] [CrossRef]
- Yii, A.C.; Koh, M.S.; Lapperre, T.S.; Tan, G.L.; Chotirmall, S.H. The emergence of Aspergillus species in chronic respiratory disease. Front. Biosci. 2017, 9, 127–138. [Google Scholar] [CrossRef]
- Chotirmall, S.H.; Branagan, P.; Gunaratnam, C.; McElvaney, N.G. Aspergillus/allergic bronchopulmonary aspergillosis in an Irish cystic fibrosis population: A diagnostically challenging entity. Respir. Care 2008, 53, 1035–1041. [Google Scholar]
- Hindson, C.M.; Chevillet, J.R.; Briggs, H.A.; Gallichotte, E.N.; Ruf, I.K.; Hindson, B.J.; Vessella, R.L.; Tewari, M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat. Methods 2013, 10, 1003–1005. [Google Scholar] [CrossRef]
- Sanders, R.; Huggett, J.F.; Bushell, C.A.; Cowen, S.; Scott, D.J.; Foy, C.A. Evaluation of digital PCR for absolute DNA quantification. Anal. Chem. 2011, 83, 6474–6484. [Google Scholar] [CrossRef]
- Hijano, D.R.; Brazelton de Cardenas, J.; Maron, G.; Garner, C.D.; Ferrolino, J.A.; Dallas, R.H.; Gu, Z.; Hayden, R.T. Clinical correlation of influenza and respiratory syncytial virus load measured by digital PCR. PLoS ONE 2019, 14, e0220908. [Google Scholar] [CrossRef] [PubMed]
- Sze, M.A.; Abbasi, M.; Hogg, J.C.; Sin, D.D. A comparison between droplet digital and quantitative PCR in the analysis of bacterial 16S load in lung tissue samples from control and COPD GOLD 2. PLoS ONE 2014, 9, e110351. [Google Scholar] [CrossRef] [PubMed]
- Huggett, J.F.; Foy, C.A.; Benes, V.; Emslie, K.; Garson, J.A.; Haynes, R.; Hellemans, J.; Kubista, M.; Mueller, R.D.; Nolan, T.; et al. The digital MIQE guidelines: Minimum Information for Publication of Quantitative Digital PCR Experiments. Clin. Chem. 2013, 59, 892–902. [Google Scholar] [CrossRef] [PubMed]
- Nixon, G.; Garson, J.A.; Grant, P.; Nastouli, E.; Foy, C.A.; Huggett, J.F. Comparative study of sensitivity, linearity, and resistance to inhibition of digital and nondigital polymerase chain reaction and loop mediated isothermal amplification assays for quantification of human cytomegalovirus. Anal. Chem. 2014, 86, 4387–4394. [Google Scholar] [CrossRef]
- Imbert, S.; Meyer, I.; Palous, M.; Brossas, J.Y.; Uzunov, M.; Touafek, F.; Gay, F.; Trosini-Desert, V.; Fekkar, A. Aspergillus PCR in Bronchoalveolar Lavage Fluid for the Diagnosis and Prognosis of Aspergillosis in Patients With Hematological and Non-hematological Conditions. Front. Microbiol. 2018, 9, 1877. [Google Scholar] [CrossRef]
- Fonceca, A.M.; Chopra, A.; Levy, A.; Noakes, P.S.; Poh, M.W.; Bear, N.L.; Prescott, S.; Everard, M.L. Infective respiratory syncytial virus is present in human cord blood samples and most prevalent during winter months. PLoS ONE 2017, 12, e0173738. [Google Scholar] [CrossRef]
- Schwartz, S.L.; Lowen, A.C. Droplet digital PCR: A novel method for detection of influenza virus defective interfering particles. J. Virol. Methods 2016, 237, 159–165. [Google Scholar] [CrossRef]
- Vogelmeier, C.F.; Criner, G.J.; Martinez, F.J.; Anzueto, A.; Barnes, P.J.; Bourbeau, J.; Celli, B.R.; Chen, R.; Decramer, M.; Fabbri, L.M.; et al. Global Strategy for the Diagnosis, Management, and Prevention of Chronic Obstructive Lung Disease 2017 Report. GOLD Executive Summary. Am. J. Respir. Crit. Care Med. 2017, 195, 557–582. [Google Scholar] [CrossRef]
- Chalmers, J.D.; Goeminne, P.; Aliberti, S.; McDonnell, M.J.; Lonni, S.; Davidson, J.; Poppelwell, L.; Salih, W.; Pesci, A.; Dupont, L.J.; et al. The bronchiectasis severity index. An international derivation and validation study. Am. J. Respir. Crit. Care Med. 2014, 189, 576–585. [Google Scholar] [CrossRef]
- Pasteur, M.C.; Bilton, D.; Hill, A.T. British Thoracic Society guideline for non-CFbronchiectasis. Thorax 2010, 65, i1–i58. [Google Scholar] [CrossRef]
- Naidich, D.P.; McCauley, D.I.; Khouri, N.F.; Stitik, F.P.; Siegelman, S.S. Computed tomography of bronchiectasis. J. Comput. Assist. Tomogr. 1982, 6, 437–444. [Google Scholar] [CrossRef]
- Chotirmall, S.H.; O’Donoghue, E.; Bennett, K.; Gunaratnam, C.; O’Neill, S.J.; McElvaney, N.G. Sputum Candida albicans presages FEV(1) decline and hospital-treated exacerbations in cystic fibrosis. Chest 2010, 138, 1186–1195. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).