Abstract
Watermelon (Citrullus lanatus L.), which is an economically important cucurbit crop that is cultivated worldwide, is vulnerable to various adverse environmental conditions. Small heat shock protein 20s (HSP20s) are the most abundant plant HSPs and they play important roles in various biotic and abiotic stress responses. However, they have not been systematically investigated in watermelon. In this study, we identified 44 watermelon HSP20 genes and analyzed their gene structures, conserved domains, phylogenetic relationships, chromosomal distributions, and expression profiles. All of the watermelon HSP20 proteins have a conserved the α-crystallin (ACD) domain. Half of the ClHSP20s arose through gene duplication events. Plant HSP20s were grouped into 18 subfamiles and a new subfamily, nucleo-cytoplasmic XIII (CXIII), was identified in this study. Numerous stress- and hormone-responsive cis-elements were detected in the putative promoter regions of the watermelon HSP20 genes. Different from that in other species, half of the watermelon HSP20s were repressed by heat stress. Plant HSP20s displayed diverse responses to different virus infections and most of the ClHSP20s were generally repressed by Cucumber green mottle mosaic virus (CGMMV). Some ClHSP20s exhibited similar transcriptional responses to abscisic acid, melatonin, and CGMMV. Subcellular localization analyses of six selected HSP20- green fluorescence protein fusion proteins revealed diverse subcellular targeting. Some ClHSP20 proteins were affected by CGMMV, as reflected by changes in the size, number, and distribution of fluorescent granules. These systematic analyses provide a foundation for elucidating the physiological functions and biological roles of the watermelon HSP20 gene family.
1. Introduction
Heat shock proteins (HSPs), an important group of molecular chaperones, have been proven to participate in various plant stress responses and in regulating plant growth and development [1]. On the basis of their molecular weight, HSPs can be classified into six families, namely, HSP100, HSP90, HSP70, HSP60, HSP20 (or small heat shock protein), and ubiquitin. Of these groups, HSP20 is the largest family and the most well studied [2]. Numerous investigations have revealed the roles of HSP20 proteins in various abiotic stress responses. Most HSP20s are highly induced by heat stress and transgenic plants overexpressing HSP20s in Arabidopsis, rice, maize, wheat, and Chenopodium sp., by protecting proteins from irreversible denaturation, enhanced tolerance to heat stress [3,4,5,6,7]. Overexpression of OsHSP17.0 and OsHSP23.7 in rice decreases membrane damage, increases the abundance of protective molecules, and leads to improved seed vigor under osmotic and salt stress conditions [8]. Transgenic rice overexpressing OsHSP20s exhibited enhanced tolerance to ultraviolet-B radiation, salt, drought, and dehydration stresses [4,9]. Promoter analyses revealed that the expression of TaHSP26 in wheat may be induced by heat, cold, salt, and drought stresses [6,10]. Meanwhile, the ectopic expression of LimHSP16.45 from David Lily (Lilium davidii) in Arabidopsis increased the levels of superoxide dismutase and catalase activities and seed germination vigor under salt stress [11]. CaHsp22.5 in Capsicum annuum and AsHSP17 in Salix suchowensis were shown to regulate plant responses to salt stress via abscisic acid (ABA) signaling pathways [12,13].
Previous studies confirmed that HSP20s play significant roles in plant immunity [14,15]. In rice, some HSP20 genes were induced under Magnaporthe grisea infection, while other genes were repressed. Transgenic Arabidopsis with reduced HSP20 expression levels was more susceptible to pathogens [16]. Tomato HSP20, which interacts with the R protein I-2, was shown to improve resistance to Fusarium oxysporum [17]. Proteomic analyses indicated that the expression of tomato HSP17.6 protein was repressed by Rhizopus nigricans infection [18]. Tobacco HSP17.6 was found to participate in the immune response to Ralstonia solanacearum infection [16]. Potato StHSP17.8 could activate the Phytophthora infestans resistance gene WRKY1 only in resistant potato genotypes [19]. Two barley powdery mildew effector candidates, CSEP0105 and CSEP0162, which contributed to the success of Blumeria graminis f. sp. hordei infection, were found to interact with barley Hsp16.9 and Hsp17.5 [20]. GmHsp20s in soybean showed different response patterns between resistant and susceptible soybean genotypes under Meloidogyne javanica infection [21]. Microarray analyses have indicated that Arabidopsis HSP20s show various response patterns to different viral infections [22]. In rice, HSP20 was significantly affected by Rice stripe virus infection, and it was found to play an important role in RSV replication and movement via interacting with the RSV protein (RdRp) and the movement protein (NSVc4) [23,24].
Watermelon (Citrullus lanatus) is an important cucurbit crop that is consumed worldwide, but various adverse environmental conditions affect its development and decrease its yield and quality. Plants HSP20 proteins play important roles in the responses to various biotic and abiotic stresses. However, watermelon HSP20 genes have not been systematically investigated at the whole genome level in watermelon. In this study, all putative HSP20 elements in watermelon were identified in an in silico analysis. The classification, conserved domains, gene structures, chromosomal distributions, phylogenetic relationships, synteny relationships, and gene duplication events of watermelon HSP20 genes were predicted and analyzed in detail. Their subcellular localizations were also predicted and verified after the transient transformation of tobacco leaf epidermal cells with (or without) Cucumber green mottle mosaic virus (CGMMV) infection. The expression profiles of ClHSP20 genes were analyzed using quantitative real-time PCR analysis (qRT-PCR) to determine their responses to various stresses and plant hormone treatments. Our comprehensive analyses have elucidated the potential roles of ClHSP20s in stress and hormone responses. Our findings provide a framework for future studies to better understand the functions of watermelon HSP20 family genes in stress tolerance and hormone responses.
2. Results
2.1. Identification of HSP20 Genes in Watermelon and Cucumber
We conducted a genome-wide analysis of HSP20 gene family members in watermelon. Protein Basic Local Alignment Search Tool (BLASTP) searches were performed in the Cucurbit Genomics Database to identify putative HSP20s in watermelon. The search queries were 273 HSP20 protein sequences from Arabidopsis, rice, soybean, tomato, pepper, wheat, barley, and switchgrass. A total of 53 putative hits were identified in the watermelon genome database. Meanwhile, 41 sequences were annotated as being watermelon HSP20 genes after searches, with HMMER 3.0 using the global hidden Markov models (HMM) profile of the HSP20 characteristic domain. Redundant sequences by the above two methods were omitted to obtain unique putative HSP20 genes, and the remaining hits were further filtered using Conserved Domains Database (CDD) and Simple Modular Architecture Research Tool (SMART) according to the presence of structural characteristics and the conserved ACD domain. Finally, 44 HSP20 genes were identified in watermelon. The same methods were applied to identify 45 HSP20s in cucumber. HSP20 genes have been intensively studied in some model plant species and important crops. Among the plants studied to date, Arabidopsis has the smallest number of HSP20s and switchgrass has the largest number. The numbers of HSP20 genes in watermelon (44) and cucumber (45) were greater than those in all other plant species, except for soybean (51) and switchgrass (63) (Table 1).

Table 1.
Summary of the number of heat shock protein 20 (HSP20) genes in diverse plant species.
The amino acid sequences of all ClHSP20s ranged from 97 to 502 amino acids in length, corresponding to molecular weights ranging from 13.72 to 57.37 kDa (Table S1). The HSP20s in watermelon were named according to their protein molecular weight, as in previous studies [2,25,26]. Almost all of the ClHSP20s had a highly conserved α-crystallin domain (ACD) at the C-terminal end, as identified by SMART, as well as two conserved motifs (motif 1 and motif 2), as identified by MEME [30] (Figure 1 and Figure 2). Multiple sequence alignments indicated that the ACD domain contained two conserved regions with a β-sandwich of two antiparallel β-sheets domains (Figure S1 and Figure 2). These conserved regions are known to play important roles in the chaperone function of HSP20 proteins [31,32]. Seven ClHSP20 proteins (ClHSP18.9A, ClHSP21.8, ClHSP27.5, ClHSP38.8, ClHSP39.8, ClHSP42.6, and ClHSP50.3) had a transmembrane domain in the C- or N-terminal region, respectively.
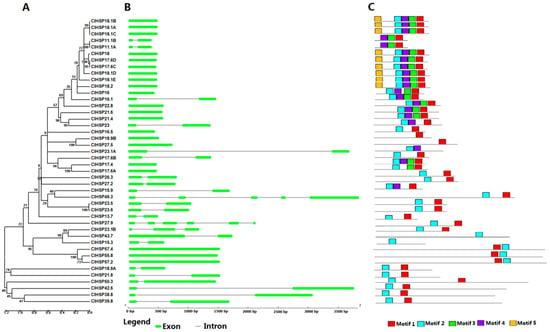
Figure 1.
Phylogenetic relationships, gene structures, and conserved motifs of all ClHSP20s in watermelon. (A) Unrooted phylogenetic tree was generated based on the amino acid sequences by the neighbor-joining (NJ) method using MEGA 5. Bootstrap supports from 1000 replicates are indicated at each branch. (B) Gene structure was analyzed using the Gene Structure Display Server online. Green boxes indicate exons, lines indicate introns. (C) Motif analysis was performed using MEME 4.0 software. Different coloured boxes represent different motifs in corresponding position of each ClHSP20 protein.
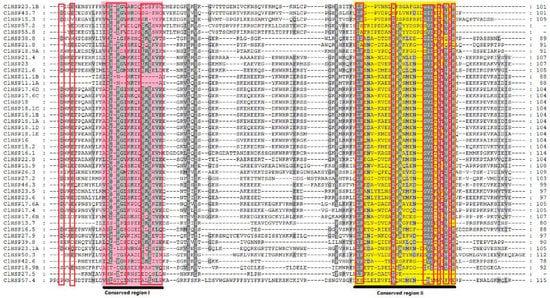
Figure 2.
Amino acid sequence alignment of the α-crystallin (ACD) domain from watermelon HSP20s. Sequences were aligned by Clustal X program. Conserved motifs are marked. Conserved region I and II were shown for red and yellow background, respectively, and the typical amino acid residues within these regions were indicated by red boxes.
According to the number of introns, the watermelon ClHSP20 genes could be divided into three types: Type I genes have no introns (21 ClHSP20s); type II genes have one intron (20 ClHSP20s); and, type III genes have more than one intron (ClHSP23.1B, ClHSP27.9, and ClHSP46.3) (Figure 1).
2.2. Phylogenetic Relationship of Plant HSP20 Members
To further analyze the phylogenetic relationships of HSP20 proteins, the amino acid sequences of HSP20 proteins from Arabidopsis, rice, tomato, soybean, switchgrass, cucumber, and watermelon were used to perform multiple alignments analyses and to construct a phylogenetic tree (Figure 3). The HSP20s were divided into 18 subfamilies. Notably, a new subfamily, nucleo-cytoplasmic XIII (CXIII), was identified in this study. This subfamily comprised only HSP20 proteins from watermelon and cucumber. Of the 18 subfamilies, 16 contained watermelon HSP20 proteins. Among all the watermelon HSP20 genes, 34 were nucleocytoplasmic (C) HSP20 genes (11 subfamilies); three were mitochondrial (M) HSP20 genes (two subfamilies); three were endoplasmic reticulum (ER) HSP20 genes; two were plastidic (P) HSP20 genes; one was a peroxisomal (Px) HSP20 gene; and, one was an orphan gene (ClHSP42.6). Watermelon had no HSP20 genes in the CVIII or CX subfamilies, since these two subfamilies were exclusively found in monocots. The CIV and CXIII subfamilies each had only one dicotyledon HSP20 gene. The largest subfamily was CI, which contained 12 ClHSP20 genes. Members of the CI subfamily were highly conserved. In fact, the gene pairs ClHSP11.1A and ClHSP11.1B, ClHSP17.6C and ClHSP17.6D, and ClHSP18.1A and ClHSP18.1D in CI subfamily shared identical amino acid sequences with each other (Table S2).
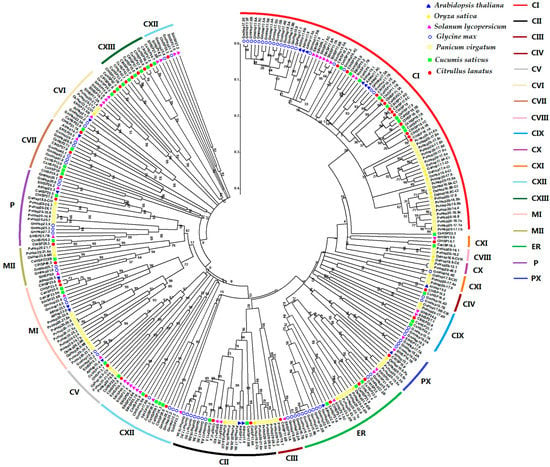
Figure 3.
Phylogenetic relationship of HSP20 proteins in Arabidopsis, rice, tomato, soybean, switchgrass, cucumber, and watermelon. Phylogenetic trees were constructed using neighbor-joining method with bootstrap tests by MEGA 5.0. The Bar represents the relative divergence of the sequences examined. The diverse HSP20 subgroups are indicated with different color arcs. The different colored symbols at the branch tips represent different species., C, cytoplasmic/nuclear; ER, endoplasmic reticulum; P, plastid; PX, peroxisome; M, mitochondria.
2.3. Genomic Distribution and Evolutionary Analysis of Watermelon HSP20s
All of the HSP20 genes in watermelon were non-randomly located on the 11 watermelon chromosomes (Chr), except for ClHSP18.1A and ClHSP27.2. Chr07 had the most ClHSP20 genes. Only one ClHSP20 was mapped to each of Chr01, Chr06, and Chr11 (Figure 4). Three tandem duplication clusters involving in 10 ClHSP20 genes were identified (Table 2). A duplicated gene pair (ClHSP23.1B and ClHSP43.7) was mapped on Chr02. This pair shared 54.2% amino acid sequence similarity (Table S2). The Ks value (i.e., synonymous substitution rate) for this duplicated pair was 1.7552, with the corresponding divergence time being 135.02 million years ago (Mya). ClHSP11.1A, ClHSP18.1B, ClHSP17.6C, and ClHSP18 were tandemly repeated on Chr 07 and shared 69.1% to 96.8% similarity with each other (Figure 4 and Table S2). The Ks values of these duplicates ranged from 0.0865 to 1.2543 corresponding to divergence times of 6.65 to 96.48 Mya (Table 2). ClHSP18.1D, ClHSP11.1B, ClHSP18.1C, and ClHSP17.6D were also distributed on Chr 07 and showed 70.1% to 90.6% amino acid similarity. The Ks values of these duplicated genes ranged from 0.3202 to 1.1953, corresponding to divergence times of 24.63 to 91.95 Mya. Interestingly, all of the eight tandem duplicates mapped on Chr 07 belonged to the CI subfamily, suggesting that the watermelon CI subfamily has probably expanded via tandem duplication. Thirteen gene pairs involving 14 ClHSP20s resulted from segment duplication (Table 2). Eight pairs of segment duplicates occurred between four ClHSP20s on Chr 07 and five genes that were scattered on Chr 03, Chr 04, Chr 10, and Chr 11. Two putative duplicated gene pairs (ClHSP16/ClHSP22.8 and ClHSP55.8/ClHSP57.2) were present on Chr 4 and Chr 10. ClHSP57.4 on Chr 02 was mapped to a duplicated region shared with ClHSP55.8 on Chr 04 and with ClHSP57.2 on Chr 10. Another duplicate pair was detected between ClHSP27.5 on Chr 05 and ClHSP18.9B on Chr 06 (Figure 4). The Ks vakues of these segment duplicated gene pairs ranged from 1.54 to 19.78, corresponding to divergence times of 118.42 to 1521.36 Mya. The Ka/Ks values of all tandem and segment duplicates were less than 1, indicating that they have undergone purifying selection.
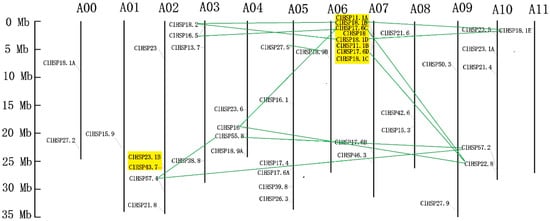
Figure 4.
Chromosomal distribution of HSP20 genes in watermelon. Chromosome number is indicated at the top of each chromosome. Tandemly duplicated genes are highlighted in yellow. Duplicated gene pairs are linked by green line.

Table 2.
Ka/Ks calculation and divergent time of the duplicated watermeolon HSP20 gene pairs.
There is a high degree of intergenomic homology between watermelon and cucumber, which belong to the Cucurbitaceae family [33]. Therefore, we assessed the syntenic relationships of HSP20 genes from watermelon and cucumber genomes using the Cucurbit Genomics Database to investigate cucurbit chromosomal evolution. A total of 42 syntenic gene pairs involving in 30 ClHSP20s that were located on all watermelon chromosomes, except for Chr 01, were identified between watermelon and cucumber. Four genes (ClHSP17.6C, ClHSP17.6D, ClHSP18.1B, and ClHSP18.1D) on watermelon Chr07 had the highest number of syntenic genes (10) in the cucumber genome (Figure 5).
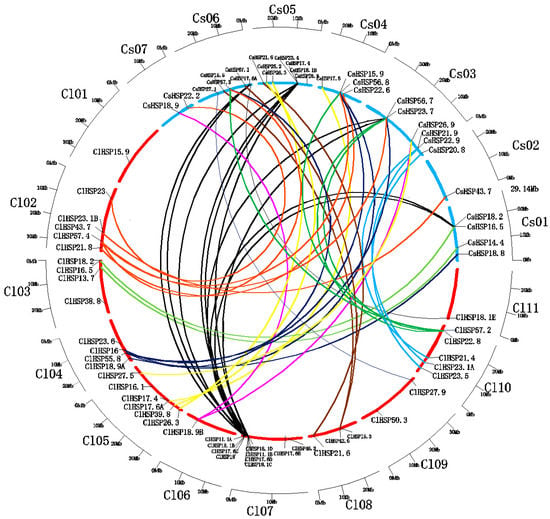
Figure 5.
Relationship diagrams of syntenic HSP20 genes in chromosomal synteny regions distributed in watermelon and cucumber. Chromosomes of watermelon and cucumber are represented by blue and red arcs according to their own sizes. Differently colored lines link representative syntenic HSP20 genes in watermelon and cucumber.
2.4. Analysis of Putative cis-Acting Elements in ClHSP20 Promoters
Heat shock elements (HSE) are common cis-acting elements in the promoters of HSP20s that confer heat shock-inducible expression. We identified and analyzed cis-acting elements in the putative promoter regions of the HSP20 genes in watermelon (Table S3). Numerous hormone- and stress-responsive elements were found in the ClHSP20 promoters. We detected MeJA-, GA-, salicylic acid (SA)-, and ABA- responsive elements in 23, 20, 20, and 10 HSP20 promoters, respectively. Of the 44 watermelon HSP20 genes, 27 had HSE motifs in their promoter regions. Drought- and low temperature- responsiveness motifs were identified in the promoters of 8 and 10 ClHSP20 genes, respectively.
2.5. Subcellular Localizations of HSP20 Proteins in Watermelon
The subcellular localizations of six watermelon HSP20 proteins from different subfamilies, namely ClHSP22.8, ClHSP15.9, ClHSP17.4, ClHSP18.9A, ClHSP18.9B, and ClHSP15.3, were analyzed by transiently expressing their green fluorescent protein (GFP) fusion proteins in tobacco leaf epidermal cells. The results showed that these six ClHSP20 proteins were localized to various subcellular compartments, the watermelon ClHSP20 proteins have diverse functions (Figure 6). Specifically, ClHSP15.9 was predicted to be located in peroxisomal according to the WoLF PSORT website, however, it was predominantly detected in cytoplasmic granules (Table S1 and Figure 6B,b). ClHSP22.8 and ClHSP15.3-GFP were predicted to be cytoplasmic and nuclear proteins, respectively, but their GFP fusion proteins were mainly agglomerated into granules in cytoplasm, and they were weakly detected in endoplasmic reticulum (ER) (Figure 6A,F,a,f). In contrast, ClHSP18.9A, ClHSP17.4, and ClHSP18.9B were predicted to be located in golgi, cytoplasm, and chloroplasts, respectively, but, in fact, the corresponding GFP fusion proteins were localized to the endoplasmic reticulum (Figure 6C–E,c–e). Additionally, ClHSP17.4-, ClHSP18.9A-, ClHSP18.9B-, and ClHSP15.3-GFP fusion proteins were also detected in nuclear (Figure S2). After CGMMV infections, the subcellular localizations of some ClHSP20s appeared to change (Figure 7). For example, ClHSP15.9-GFP fusion proteins accumulated into more vesicles and some newly formed vesicae in the cytoplasm, whereas ClHSP17.4-GFP began to agglomerate in relatively large granules along the endoplasmic reticulum, and ClHSP15.3-GFP appeared to no longer agglomerate in granules, but instead it was present at endoplasmic reticulum.
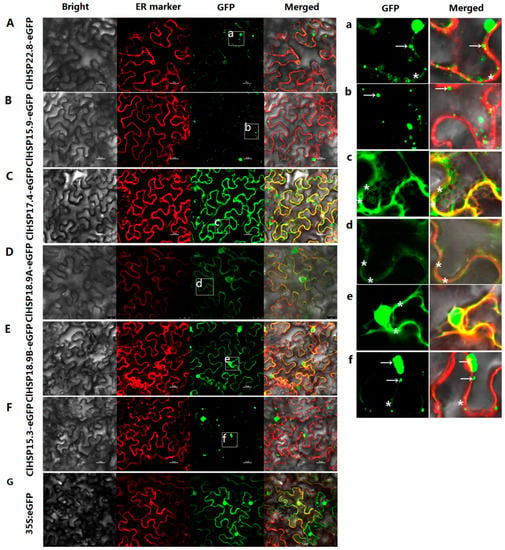
Figure 6.
Subcellular localizations of ClHSP20 proteins. Green fluorescent protein (GFP)-fusion proteins were transiently expressed in tobacco leaves. The results were observed after a 48 h-incubation. Bright-field, endoplasmic reticulum (ER) marker (red fluorescence), green fluorescence (GFP), and merged images (from left to right) of 35S:ClHSP22.8-EGFP (A); 35S:ClHSP15.9-EGFP (B); 35S:ClHSP17.4-EGFP (C); 35S:ClHSP18.9A-EGFP (D); 35S:ClHSP18.9B-EGFP (E); 35S:ClHSP15.3-EGFP (F); and, 35S:EGFP (G) in tobacco leaf epidermal cells. (a–f) displays enlarged images of the section delimited by a white square in A to F, respectively. Green fluorescence signal indicated the localizations of ClHSP20 fusion proteins. Red fluorescence signal indicated endoplasmic reticulum (ER) marker. White arrows indicated fluorescent granules. White stars labeled the GFP fluorescent signal located on endoplasmic reticulum. Scale bars, 25 μm.
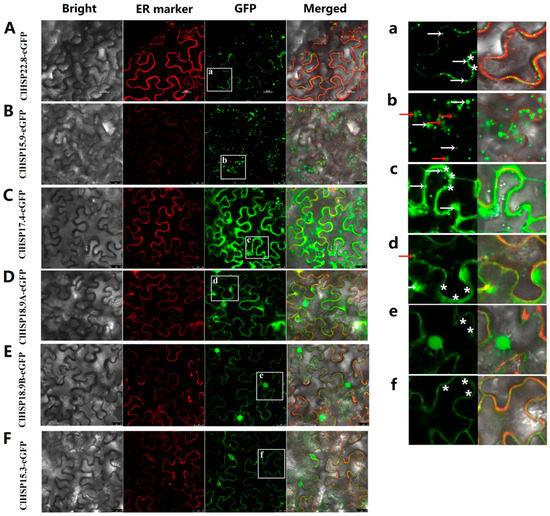
Figure 7.
Subcellular localizations of ClHSP20 proteins under Cucumber green mottle mosaic virus (CGMMV) infection. Green fluorescent protein (GFP)-fusion proteins were transiently expressed in tobacco leaves with CGMMV infectious clones. Fluorescence signal was detected after 48 h of incubation. Bright-field, endoplasmic reticulum (ER) marker (red fluorescence), green fluorescence (GFP), and merged images (from left to right) of 35S: ClHSP22.8-EGFP (A); 35S: ClHSP15.9-EGFP (B); 35S:ClHSP17.4-EGFP (C); 35S:ClHSP18.9A-EGFP (D); 35S:ClHSP18.9B-EGFP (E); and, 35S:ClHSP15.3-EGFP (F) in tobacco leaf epidermal cells. (a–f) displays enlarged images of the section delimited by a white square in A to F, respectively. Fluorescent granules were indicated by white arrows. Vesicae were marked by red arrows. White stars labeled the GFP fluorescent signal located on endoplasmic reticulum. Scale bars, 25 μm.
2.6. Expression Analyses of ClHSP20 Genes in Response to Abscisic Acid and Melatonin
The expression patterns of 30 randomly selected ClHSP20 genes in response to ABA and melatonin (MT) were analyzed in a qRT-PCR assay (Figure 8, Table S6). Most of the ClHSP20s could respond to ABA and MT. Under ABA treatment, the expression levels of 14 ClHSP20s were obviously downregulated from 4 h, while eight ClHSP20s exhibited upregulated expression. Of these genes, ClHSP18.1E, ClHSP18.2, and ClHSP23 expression levels increased by 70-, 40-, and 39-fold, respectively, at 4 h after the ABA treatment (Figure 8A, Table S6). Some ClHSP20s, such as ClHSP11.1A, ClHSP17.6B, and ClHSP21.6, were induced after repressed by ABA treatment. In contrast, MT downregulated the expression of most of the analyzed ClHSP20 genes, especially at 4 and 12 h after treatments. Moreover, the expression levels of some ClHSP20 genes, including ClHSP11.1A, ClHSP26.3, ClHSP21.6, and ClHSP23, were downregulated from 1 to 12 h after the MT treatment (Figure 8B). Meanwhile, MT upregulated the expression of 11 ClHSP20 genes, especially at 1 h after treatments. The ClHSP17.6A expression level increased by 145-fold at 12 h, whereas ClHSP22.8 expression was evidently downregulated at 1 h, but it increased by nearly three-fold at 4 h after the MT treatment.
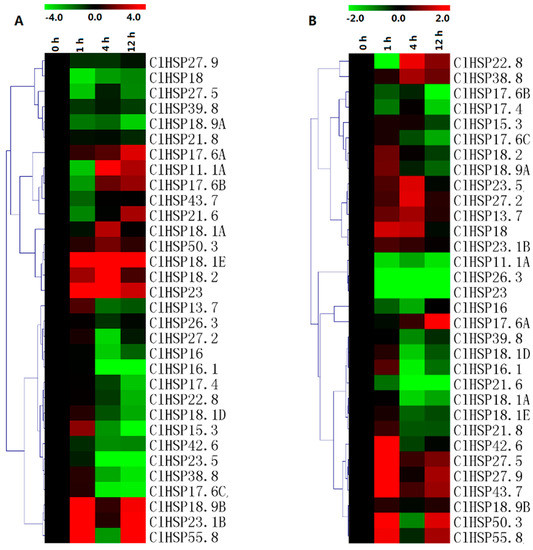

Figure 8.
Heat map representation of the responses of watermelon ClHSP20 genes to exogenous melatonin. (A) and abscisic acid (B). Watermelon leaves were collected at 0, 1, 4, and 12 h after 150 μM melatonin or 100 μM abscisic acid treatment. Heatmaps were constructed with the MeV4.8 program. Color scales representing the relative expression values are shown on the upper left of heatmap (log2 scale).
2.7. Expression Analyses of ClHSP20 Genes in Response to Heat and CGMMV
The qRT-PCR results indicated that the expression levels of only 11 of 30 analyzed ClHSP20 genes were upregulated by heat stress (Figure 9, Table S6). We observed that ClHSP11.1A, ClHSP50.3, and ClHSP17.4 expression levels considerably increased (>100-fold) in response to heat stress. Meanwhile, 11 genes exhibited the opposite trend following an exposure to heat, especially ClHSP27.5, ClHSP18.2, ClHSP15.3, ClHSP18.9B, and ClHSP23.1B. Infection by CGMMV induced eight watermelon HSP20s, especially ClHSP17.6A, and ClHSP21.5, which showed drastic increases in their transcript levels (145- and 25022-fold, respectively). Nevertheless, nine ClHSP20s (ClHSP11.1A, ClHSP16.1, ClHSP17.6B, ClHSP17.6C, ClHSP18.1A, ClHSP18.1D, ClHSP23, ClHSP21.6, and ClHSP43.7) were obviously repressed by CGMMV infection from 6 h. The ClHSP27.9 expression level was upregulated at 1 h, but it was downregulated at 6 and 48 h after the CGMMV infection. The other ClHSP20 genes appeared unresponsive to CGMMV.
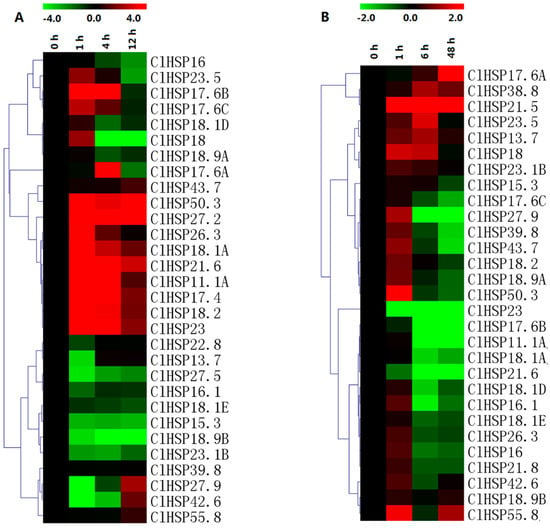
Figure 9.
Heat map representation of response patterns of watermelon HSP20 genes to heat stress (A) and CGMMV infection (B). For heat treatment, leaves were collected at 0, 1, 4, and 12 h after 42 °C treatment. Watermelon leaves were collected at 0, 1, 6, and 48 h after CGMMV infection. For other details, see Figure 7.
2.8. Gene Ontology Enrichment Analysis
To further explore their functions, the ClHSP20 genes underwent a Gene Ontology (GO) enrichment analysis. The enriched GO terms were grouped into the following three categories; biological process, molecular function, and cellular component (Figure 10 and Table S4). The significantly enriched molecular function GO term for molecular function was protein binding (GO:0005515). The ClHSP20 proteins were mainly located in the cytoplasm, plastid, and peroxisomal matrix. The main enriched biological process terms were hyperosmotic salinity response (GO:0042538), heat acclimation (GO:0010286), response to heat (GO:0009408), response to hydrogen peroxide (GO:0042542), and response to high light intensity (GO:0042538). The results of the GO enrichment analysis suggested that the ClHSP20 genes have significant roles in various stress responses.
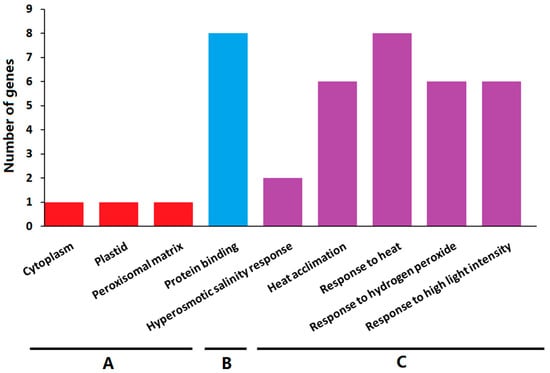
Figure 10.
The Gene Ontology (GO) analysis of HSP20 genes in watermelon. The ClHSP20 genes were categorized into three groups: cell component (A); biological process (B); and, molecular function (C).
3. Discussion
The HSP20 gene family has been identified in Arabidopsis, rice, soybean, wheat, barley, tomato, pepper, and switchgrass, respectively [2,22,25,27,28,29,34,35]. The number of HSP20 gene family members ranges from 13 (barley) to 63 (soybean). In this study, we identified 44 ClHSP20s in watermelon, fewer than in switchgrass (63), soybean (51), and cucumber (45), but more than in the other species (Table 1). Although the genome size of watermelon (425 Mb) is obviously bigger than in Arabidopsis (125 Mb) and cucumber (367 Mb), the sizes of many gene families in watermelon are similar to those in cucumber but slightly smaller than those in Arabidopsis [36,37,38]. This is probably because watermelon avoided more recent wholegenome duplication events, except for the core-eudicot common hexaploidization (ECH) event [39,40]. However, the number of HSP20s in watermelon is twice than that in Arabidopsis in this study, which appears to contradict the above findings [39], but is consistent with the latest research that a cucurbit-common tetraploidization (CCT, 90–102 Mya) event occurred in the Cucurbitaceae shortly after the ECH event (115–130 Mya) [33]. The 13 tandemly duplicated gene pairs of ClHSP20s arose 6.65 to 135.02 Mya, probably as a result of gene losses and chromosomal rearrangements after the CCT event, However, there are relatively low rates of preserved CCT colinear genes in the watermelon genome (2.5–5.5%), which explains why many other watermelon gene families are quite small [36,37,38,39]. Watermelon has retained high genomic homology and it shares more colinear genes with cucumber [33]. Therefore, we analyzed the linear relationships between HSP20 genes in watermelon and cucumber in this study. Forty-two colinear gene pairs between the two species were identified (Figure 5). Some HSP20s in the watermelon genome had more than one syntenic gene in the cucumber genome. These watermelon colinear genes were distributed on all of the chromosomes, except for Chr 01, with the largest number of colinear genes on Chr 07. The existence of so many colinear genes and colinear blocks between the watermelon and cucumber genomes was probably due to another round of genomic repatterning after the CCT event [33].
The HSP20 genes are important for responses to various stresses [3,5,6,14,15]. In this study, GO enrichment results indicated that ClHSP20 genes were mainly enriched in the heat, salinity, hydrogen peroxide, and high light signal response pathway. The HSP20 genes are characterized by high and rapid upregulation in response to heat stress. Almost all of HSP20s from soybean, pepper, and switchgrass exhibit upregulated expression in response to heat stress [21,28,29]. In watermelon, all HSP20s could respond to heat stress, except for ClHSP39.8, ClHSP43.7, and ClHSP55.8. However, the expression levels of approximately half of the ClHSP20 genes are repressed by heat shock, especially ClHSP27.5, ClHSP18.1E, ClHSP16.1, ClHSP18.9B, ClHSP15.3, and ClHSP23.1B. The induced HSP20s could enhance heat tolerance by protecting proteins from irreversible denaturation, but it is necessary to elucidate the roles of these downregulated genes [41]. Infections by CGMMV are devastating for cucurbit crops, and they have been responsible for considerable watermelon yield losses [42]. However, little is known about the molecular mechanism underlying the CGMMV-induced watermelon disease. In response to CGMMV infection, most watermelon HSP20s were obviously repressed from 6 h after being slightly induced at 1 h in this study. Just few reports indicated plant HSP20s involved in viral response. Microarray technology analysis indicated that Arabidopsis HSP17.4 was induced under five plant viruses infection and it was speculated to have a common mechanism in response to viral infection, but its homologous gene, ClHSP18.1E in watermelon, was repressed by CGMMV infection [43]. The RNA-Seq data analysis found that two HSP20s from ER subfamily, namely, ClHSP23 and ClHSP21.6, in CGMMV-inoculated watermelon fruits, were significantly repressed, which had similar responses in leaves in this study [44]. The results suggested that ER subfamily genes in watermelon probably played negative roles in response to CGMMV.
Recently, studies found that melatonin widely participated in plant biotic and abiotic stress responses, even it has begun to be considered by some experts in plant hormones [45]. HSP20s have been to proved to participate in stress responses via the MT signaling pathway, and the expression levels of tomato HSP20s increased under heat and cadmium stress after melatonin treatment [46,47]. In this study, the qRT-PCR results indicated that most of the watermelon HSP20 genes were induced by MT at 1 h but obviously repressed at 4 and 12 h. Similarly, most of the watermelon HSP20s were also generally repressed by ABA treatment, as well as the expression levels of nearly all switchgrass HSP20 genes are downregulated in response to an ABA treatment [29]. Notably, more ClHSP20s has similar response patterns to ABA, MT, and CGMMV, as they were generally repressed by the three treatments. Take the newly identified CXIII subfamily gene ClHSP55.8 and ClHSP15.3 as examples, following ABA and MT treatments, the ClHSP55.8 transcript abundance dramatically increased at 1 h, decreased at 4 h, and then increased to high levels at 12 h. Similarly, ClHSP55.8 expression was induced by CGMMV at 1 and 48 h, but was repressed at 6 h. In contrast, ClHSP15.3 expression was repressed by ABA, MT, and CGMMV, except for a transient increase at 1 h in response to ABA. These findings imply that these ClHSP20 genes probably participate in a common signaling pathway in response to MT, ABA, and CGMMV.
Plant HSP20 proteins have been detected in various cellular locations, including the cytosol, nucleus, chloroplast, endoplasmic reticulum, mitochondrion, and peroxisome [25]. This diversity in subcellular localization is probably due to the multiple functions for these proteins. To analyze the subcellular localization and distribution pattern of watermelon HSP20 proteins, six ClHSP20s fused with GFP at their N terminus were constructed and introduced into tobacco epidermal cells. The ClHSP20s displayed different subcellular localizations that were expressed alone in tobacco epidermal cells. The fluorescence signal of ClHSP18.9A-, ClHSP17.4-, and ClHSP18.9B-GFP proteins were mainly detected in endoplasmic reticulum and nuclear (Figure 6, Figure S2). ClHSP22.8, ClHSP15.9, and ClHSP15.3 formed different fluorescent granules in cytoplasm. These granules probably were large oligomers that were formed in vitro by HSP20 proteins as previous reports (Figure 6) [48]. Recently, a study found that the Rice stripe virus could alter the sub-cellular distribution of HSP20s in rice and tobacco [23]. In this study, some ClHSP20-GFP fusion proteins, including ClHSP15.9-GFP, ClHSP17.4-GFP, and ClHSP15.3-GFP, were affected by CGMMV, especially regarding the size, number, and distribution of the granules that they formed (Figure 7). The ClHSP15.9-GFP and ClHSP17.4-GFP proteins primarily accumulated in granules. Notably, ClHSP15.9-GFP proteins formed new cytoplasmic vesicles. In contrast, the fluorescence of ClHSP15.3-GFP proteins that was initially detected in granules was subsequently mainly observed in the membrane. These granules or vesicles detected in this study were similar to stress granules, and were identified in animal cells in response to virus infection. The formation of stress granules probably was an anti-viral response [23,49]. A more thorough characterization of these granules will likely clarify the mechanism underlying ClHSP20 responses to viral infections.
4. Materials and Methods
4.1. Identification of HSP20s in Watermelon and Cucumber
Protein sequences of all known HSP20 genes, that is, 19, 23, 42, 27, 13, 35, 51, and 63 members of the HSP20 family in Arabidopsis, rice, tomato, wheat, barely, pepper, soybean, and switchgrass, respectively, were downloaded from Phytozome (http://phytozome.jgi.doe.gov/pz/portal.html). These sequences were then used as queries to carry out BLASTP searches with E-value of 1 × 10−5 as the threshold. Meanwhile, Hidden Markov Model (HMM) profiles were generated using HSP20 conserved domain sequences from Pfam (http://pfam.janelia.org/). The HMM profiles were used to identify the putative watermelon and cucumber HSP20 proteins using HMMER 3.0 software (http:// hmmer.janelia.org/) with a default E-value. Finally, redundant sequences were omitted to retain unique HSP20 genes. ExPASy (http://web.expasy.org/compute_pi/) was used to calculate the molecular weights and isoelectric points (PIs) of putative watermelon HSP20 proteins. Subcellular localizations were predicted using WoLF PSORT website (https://wolfpsort.hgc.jp/).
4.2. Phylogenetic Analysis, Gene Structure Construction, and Motif Analysis
The similarity of HSP20s from Arabidopsis, watermelon, and cucumber was calculated using DNAStar software (Madison, WI, USA). Phylogenetic analysis based on full-length protein sequences was performed using the MEGA 5.0 program by the neighbor-joining (NJ) method with 1000 bootstrap replicates [50]. The structures of all watermelon HSP20 genes were analyzed via the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/). Motif analysis and annotation of conserved motifs in HSP20 proteins were conducted using MEME (http://meme.nbcr.net/meme/cgi-bin/meme.cgi). The predicted peptide sequences of conserved domains in HSP20 proteins were confirmed using CDD (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi) and SMART databases (http://smart.emblheidelberg.de/). Multiple-sequence alignment of predicted peptide sequences of the conserved α-crystallin (ACD) domain was carried out using Clustal X v1.81 with default parameters [51].
4.3. Chromosomal Localization, Gene Duplication, and Evolutionary Analysis
All HSP20 genes were assigned to corresponding watermelon chromosomes based on the Cucurbit Genomics Database. Gene pairs that were separated by fewer than five intervening genes and sharing ≥40% amino acid sequence similarity were considered to have undergone a tandem duplication event [52]. The synonymous (Ks) and non-synonymous (Ka) substitution rates were estimated, as described by Tang et al., 2008 [53]. Synteny analysis was performed with the Cucurbit Genomics Database. CLUSTALW (http://www.genome.jp/tools/clustalw/) was used to align the amino acid sequences and corresponding CDS sequences of HSP20 elements, and then Ks and Ka values were calculated using the Codeml procedure of the PAML (http://www.bork.embl.de/pal2nal/). The divergence time of each duplicated gene pair was estimated using the synonymous mutation rate of substitutions per synonymous site per year, as follows: T = Ks/2x (x = 6.56 × 10−9) [54]. Gene Ontology (GO) enrichment analysis was performed while using AgBase (http://agbase.arizona.edu/cgi-bin/tools/index.cgi).
4.4. Analysis of cis-Acting Elements in HSP20 Putative Promoter Regions in Watermelon
To identify the cis-elements in the promoter sequences of HSP20 genes in watermelon, the 1.5-kb upstream regions were obtained from the Cucurbit Genomics Database, and analyzed using the online tools at the PlantCARE website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/search_CARE.html).
4.5. Watermelon Plant Growth and Treatments
The watermelon advanced inbred line ‘JJZ-M’ was used for expression analyses. The plants were grown in a growth chamber in temperature-controlled greenhouses under day/night temperatures of 28/22 ± 1 °C, light intensity of 200 μmol·m−2·s−1, and a 16-h light/8-h dark photoperiod. Three-week-old watermelon seedlings were used for stress and exogenous hormone treatments. For MT and ABA treatments, the seedlings were sprayed with 150 μM MT and 100 μM ABA, respectively [38,55]. The second true leaf on each plant was sampled at 0 (control), 1, 4, and 12 h after treatment. In the heat treatment, watermelon plants were subjected to heat treatment at 42 °C and the leaves were collected at 0 (control), 1, 4, and 12 h. For CGMMV infection, transformed Agrobacterium cells containing the infectious full-length cDNA of the CGMMV genome were incubated at 28 °C for two days. The cells were then resuspended in infiltration solution (500 mM acetosyringone, 10 mM MES, pH 5.8, and 10 mM MgCl2) and injected into the abaxial side of watermelon leaves, as described by Voinnet et al., 2003 [56]. Infiltrated leaves were collected at 0 (control), 1, 6, and 48 h after infection. All treatments were repeated three times and each treatment contained 20 seedlings. All materials were frozen at −75 °C until RNA isolation.
4.6. RNA Isolation and qRT–PCR
The total RNA was extracted from samples using TRIZOL reagent (Invitrogen, Germany), according to the manufacturer’s protocol. First-strand cDNA was generated from 1 μg of total RNA using the PrimeScript RT reagent kit (Takara, Japan), according to the manufacturer’s instructions. Specific primers that were used in the qRT-PCR were designed using Primer 5 software, and each primer was searched in the watermelon database to ensure its specificity. The qRT-PCR reactions (reaction volume, 15 μL) were performed on a CFX96 Real Time System machine (Bio-RAD, USA), programmed to heat for 30 s at 95 °C, followed by 40 cycles of 5 s at 95 °C and 45 s at 55 °C, and at the end, one cycle of 1 min at 95 °C, 30 s at 50 °C, and 30 s at 95 °C. Two biological and three technical replicates for each sample were analyzed using the SYBR Premix Ex Taq kit (TOYOBO, Japan). Watermelon β-actin (Cla007792) was selected as an internal control [57]. The relative gene expression level was calculated using the 2−ΔΔCt method. The heatmap was generated from relative gene expression data using Multiple Array Viewer.
4.7. Subcellular Localization Analyses
The CDS sequences of ClHSP22.8, ClHSP15.9, ClHSP17.4, ClHSP18.9A, ClHSP18.9B, and ClHSP15.3 were amplified using gene-specific primers and they were cloned into the pFGC-eGFP plasmid via the Xba I and BamH I restriction sites (Table S5). Recombinant expression vectors of ClHSP20s fused to the CDS of enhanced green fluorescent protein (eGFP) protein were constructed. These plasmids were transformed into Agrobacteriumt tumefaciens GV3101 and transiently expressed in tobacco leaf cells with (or without) transformed Agrobacterium containing the infectious full-length cDNA clone of the CGMMV genome. The pFGC:eGFP empty vector served as the positive control. Images were acquired at 48 h using a Leica DMLE camera (Leica, Wetzlar, Germany).
5. Conclusions
HSP20s are the most abundant HSPs in plants and they play important roles in various biotic and abiotic stresses. However, HSP20 genes in watermelon had not been systematically analyzed. Here, 44 HSP20 genes in watermelon were identified and their gene structure, conserved domains, phylogenetic relationships, chromosome evolution, expression profiles, and subcellular localizations were analyzed. All of the watermelon HSP20 proteins contained a conserved α-crystallin (ACD) domain. Plant HSP20s could be divided into 18 subfamilies and a new subfamily, nucleo-cytoplasmic XIII (CXIII) was identified in this study. Of the 26 duplicated gene pairs in the watermelon genome, 13 arose by tandem duplication and 13 arose by segmental duplication. Numerous stress- and hormone-responsive cis-elements were detected in the putative promoter regions of the watermelon HSP20 genes. Almost all HSP20s from soybean, pepper, and switchgrass induced by heat stress, but about half of the watermelon HSP20s were repressed by heat stress via qRT-PCR analyse. Plant HSP20s displayed diverse responses to different virus infections in previous studies and most of the ClHSP20s were generally repressed by Cucumber green mottle mosaic virus (CGMMV). Several ClHSP20s showed similar response patterns to ABA, MT, and CGMMV. Subcellular localization analyses of six selected HSP20-GFP fusion proteins revealed diverse subcellular targeting in the epidermal cells of N. benthamiana. The subcellular localization and distribution patterns of ClHSP20 proteins, especially the size, number, and distribution patterns of the granules formed by ClHSP20-GFP proteins, were markedly affected by CGMMV infection. This systematic analysis will provide a foundation for elucidating the physiological functions and biological roles of the HSP20 family.
Supplementary Materials
Supplementary materials can be found at http://www.mdpi.com/1422-0067/20/1/12/s1.
Author Contributions
Y.H. performed the experiments, analyzed the data, and drafted the manuscript. L.L. participated in qRT-PCR experiments and data analysis. Y.S. and L.L. collected the public dataset and assisted with data analysis, M.F. prepared the watermelon samples, conceived the study and its design, and assisted with revisions to the manuscript. All authors read and consented to the final version of the manuscript.
Funding
This research was funded by the National Key Research & Development Program of China (2018YFD0100703), Natural Science Foundation of Zhejiang Province (LQ18C150003), National Natural Science Foundation of China (31572145 and 31772332), and Zhejiang Major Agricultural Science and Technology Projects for New Varieties Breeding (2016C02051-4-2).
Acknowledgments
We thank Jennifer Smith, PhD, from Liwen Bianji, Edanz Group China (www.liwenbianji. cn/ac), for editing the English text of a draft of this manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Asea, A.A.A.; Kaur, P.; Calderwood, S.K. Heat Shock Proteins and Plants; Springer International Publishing: Cham, Switzerland, 2016. [Google Scholar] [CrossRef]
- Sarkar, N.K.; Kim, Y.K.; Grover, A. Rice sHSP genes: Genomic organization and expression profiling under stress and development. BMC Genom. 2009, 10, 393. [Google Scholar] [CrossRef] [PubMed]
- Charng, Y.Y.; Liu, H.C.; Liu, N.Y.; Hsu, F.C.; Ko, S.S. Arabidopsis Hsa32, a novel heat shock protein, is essential for acquired thermotolerance during long recovery after acclimation. Plant Physiol. 2006, 140, 1297–1305. [Google Scholar] [CrossRef] [PubMed]
- Murakami, T.; Matsuba, S.; Funatsuki, H.; Kawaguchi, K.; Saruyama, H.; Tanida, M.; Sato, Y. Overexpression of a small heat shock protein, sHSP17.7, confers both heat tolerance and UV-B resistance to rice plants. Mol. Breed. 2004, 13, 165–175. [Google Scholar] [CrossRef]
- Sun, L.P.; Liu, Y.; Kong, X.P.; Zhang, D.; Pan, J.W.; Zhou, Y.; Wang, L.; Li, D.Q.; Yang, X.H. ZmHSP16.9, a cytosolic class I small heat shock protein in maize (Zea mays), confers heat tolerance in transgenic tobacco. Plant Cell Rep. 2012, 31, 1473–1484. [Google Scholar] [CrossRef] [PubMed]
- Khurana, N.; Chauhan, H.; Khurana, P. Wheat chloroplast targeted sHSP26 promoter confers heat and abiotic stress inducible expression in transgenic Arabidopsis plants. PLoS ONE 2013, 8, e54418. [Google Scholar] [CrossRef] [PubMed]
- Shakeel, S.N.; Heckathorn, S.; Luthe, D.S. Analysis of gene sequences indicates that quantity not quality of chloroplast small HSPs improves thermotolerance in C4 and CAM plants. Plant Cell Rep. 2012, 31, 1943–1957. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Liu, C.F.; Liu, A.L.; Zou, D.; Chen, X.B. Overexpression of OsHsp17.0 and OsHsp23.7 enhances drought and salt tolerance in rice. J. Plant Physiol. 2012, 169, 628–635. [Google Scholar] [CrossRef] [PubMed]
- Kaur, H.; Petla, B.P.; Kamble, N.U.; Singh, A.; Rao, V.; Salvi, P.; Ghosh, S.; Majee, M. Differentially expressed seed aging responsive heat shock protein OsHSP18.2 implicates in seed vigor, longevity and improves germination and seedling establishment under abiotic stress. Front. Plant Sci. 2015, 6, 713. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, H.; Khurana, N.; Nijhavan, A.; Khurana, J.P.; Khurana, P. The wheat chloroplastic small heat shock protein (sHSP26) is involved in seed maturation and germination and imparts tolerance to heat stress. Plant Cell Environ. 2012, 35, 1912–1931. [Google Scholar] [CrossRef] [PubMed]
- Mu, C.J.; Zhang, S.J.; Yu, G.Z.; Chen, N.; Li, X.F.; Liu, H. Overexpression of small heat shock protein LimHSP16.45 in Arabidopsis enhances tolerance to abiotic stresses. PLoS ONE 2013, 8, e82264. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ji, L.; Jia, Z.; Yang, X.; Meng, Q.; Guo, S. Constitutive expression of CaHsp22.5 enhances chilling tolerance in transgenic tobacco by promoting the activity of antioxidative enzymes. Funct. Plant Biol. 2018, 45, 575–585. [Google Scholar] [CrossRef]
- Sun, X.; Sun, C.; Li, Z.; Hu, Q.; Han, L.; Luo, H. AsHSP17, a creeping bentgrass small heat shock protein modulates plant photosynthesis and ABA-dependent and independent signalling to attenuate plant response to abiotic stress. Plant Cell Environ. 2016, 39, 1320–1337. [Google Scholar] [CrossRef] [PubMed]
- Park, C.J.; Seo, Y.S. Heat shock proteins: A review of the molecular chaperones for plant immunity. Plant Pathol. J. 2015, 31, 323–333. [Google Scholar] [CrossRef] [PubMed]
- Mandadi, K.K.; Scholthof, K.B.G. Plant immune responses against viruses: How does a virus cause disease? Plant Cell 2013, 25, 1489–1505. [Google Scholar] [CrossRef] [PubMed]
- Maimbo, M.; Ohnishi, K.; Hikichi, Y.; Yoshioka, H.; Kiba, A. Induction of a small heat shock protein and its functional roles in Nicotiana plants in the defense response against Ralstonia solanacearum. Plant Physiol. 2007, 145, 1588–1599. [Google Scholar] [CrossRef] [PubMed]
- Van Ooijen, G.; Lukasik, E.; Van Den Burg, H.A.; Vossen, J.H.; Cornelissen, B.J.C.; Takken, F.L.W. The small heat shock protein 20 RSI2 interacts with and is required for stability and function of tomato resistance protein I-2. Plant J. 2010, 63, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.Q.; Zhu, B.Z.; Luo, Y.B.; Fu, D.Q. Unraveling the protein network of tomato fruit in response to necrotrophic phytopathogenic Rhizopus nigricans. PLoS ONE 2013, 8, e73034. [Google Scholar] [CrossRef]
- Yogendra, K.N.; Kumar, A.; Sarkar, K.; Li, Y.L.; Pushpa, D.; Mosa, K.A.; Duggavathi, R.; Kushalappa, A.C. Transcription factor StWRKY1 regulates phenylpropanoid metabolites conferring late blight resistance in potato. J. Exp. Bot. 2015, 66, 7377–7389. [Google Scholar] [CrossRef]
- Ahmed, A.A.; Pedersen, C.; Schultz-Larsen, T.; Kawaaitaal, M.; Jørgensen, H.J.L.; Thrdal-Christensen, H. The barley powdery mildew candidate secreted effector protein csep0105 inhibits the chaperone activity of a small heat shock protein. Plant Physiol. 2015, 168, 321–333. [Google Scholar] [CrossRef]
- Lopes-Caitar, V.S.; de Carvalho, M.C.; Darben, L.M.; Kuwahara, M.K.; Nepomuceno, A.L.; Dias, W.P.; Abdelnoor, R.V.; Marcelino-Guimarães, F.C. Genome-wide analysis of the Hsp20 gene family in soybean: Comprehensive sequence, genomic organization and expression profile analysis under abiotic and biotic stresses. BMC Genom. 2013, 14, 577. [Google Scholar] [CrossRef]
- Bricchi, I.; Bertea, C.M.; Occhipinti, A.; Paponov, I.A.; Maffei, M.E. Dynamics of membrane potential variation and gene expression induced by Spodoptera littoralis, Myzus persicae, and Pseudomonas syringae in Arabidopsis. PLoS ONE 2012, 7, e46673. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xiang, C.Y.; Yang, J.; Chen, J.P.; Zhang, H.M. Interaction of HSP20 with a viral RdRp changes its sub-cellular localization and distribution pattern in plants. Sci. Rep. UK 2015, 5, 14016. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Du, Z.; Qin, M.; Wang, P.; Lan, H.; Niu, X.; Jia, D.; Xie, L.; Lin, Q.; Xie, L.; et al. Pc4, a putative movement protein of Rice stripe virus, interacts with a type I DnaJ protein and a small Hsp of rice. Virus Genes 2009, 38, 320–327. [Google Scholar] [CrossRef] [PubMed]
- Scharf, K.D.; Siddique, M.; Vierling, E. The expanding family of Arabidopsis thaliana small heat stress proteins and a new family of proteins containing α-crystallin domains (ACD proteins). Cell Stress Chaperones 2001, 6, 225–237. [Google Scholar] [CrossRef]
- Yu, J.H.; Cheng, Y.; Feng, K.; Ruan, M.Y.; Ye, Q.J.; Wang, R.Q.; Li, Z.M.; Zhou, G.Z.; Yao, Z.P.; Yang, Y.J.; et al. Genome-wide identification and expression profiling of tomato HSP20 gene family in response to biotic and abiotic stresses. Front Plant Sci. 2016, 7, 1215. [Google Scholar] [CrossRef] [PubMed]
- Pandey, B.; Kaur, A.; Gupta, O.P.; Sharma, I.; Sharma, P. Identification of HSP20 gene family in wheat and barley and their differential expression profiling under heat stress. Appl. Biochem. Biotechnol. 2015, 175, 2427–2446. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Liu, J.H.; Lu, J.P.; Zhai, Y.F.; Wang, H.; Gong, Z.H.; Wang, S.B.; Lu, M.H. Genome-wide analysis of the CaHsp20 gene family in pepper: Comprehensive sequence and expression profile analysis under heat stress. Front Plant Sci. 2015, 6, 806. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Zhang, A.; Chen, J.; He, X.; Xu, B.; Xie, G.; Miao, Z.; Zhang, X.; Huang, L. Genome-Wide Analysis of the PvHsp20 Family in Switchgrass: Motif, Genomic Organization, and Identification of Stress or Developmental-Related Hsp20s. Front. Plant Sci. 2017, 8, 1024. [Google Scholar] [CrossRef]
- De Jong, W.W.; Caspers, G.J.; Leunissen, J.A. Genealogy of the acrystallin–small heat-shock protein superfamily. Int. J. Biol. Macromol. 1998, 22, 151–162. [Google Scholar] [CrossRef]
- Bondino, H.G.; Valle, E.M.; Ten, H.A. Evolution and functional diversification of the small heat shock protein/α-crystallin family in higher plants. Planta 2012, 235, 1299–1313. [Google Scholar] [CrossRef]
- Waters, E.R. The evolution, function, structure, and expression of the plant sHSPs. J. Exp. Bot. 2013, 64, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Sun, P.; Li, Y.; Liu, Y.; Yang, N.; Yu, J.; Ma, X.; Sun, S.; Xia, R.; Liu, X.; et al. An overlooked paleo-tetraploidization in Cucurbitaceae. Mol. Biol. Evol. 2018, 35, 16. [Google Scholar] [CrossRef] [PubMed]
- Waters, E.R.; Aevermann, B.D.; Sanders-Reed, Z. Comparative analysis of the small heat shock proteins in three angiosperm genomes identifies new subfamilies and reveals diverse evolutionary patterns. Cell Stress Chaperones 2008, 13, 127–142. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, Y.; Chen, J.; Xie, W.; Wang, L.; Zhang, Q. Comprehensive sequence and expression profile analysis of Hsp20 gene family in rice. Plant Mol. Biol. 2009, 70, 341–357. [Google Scholar] [CrossRef]
- Yu, Y.J.; Liang, Y.; Lv, M.L.; Wu, J.; Lu, G.; Cao, J.S. Genome-wide identification and characterization of polygalacturonase genes in Cucumis sativus and Citrullus lanatus. Plant Physiol. Biochem. 2014, 74, 263–275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Huang, X.; Bao, Y.; Wang, B.; Zeng, H.; Cheng, W.; Tang, M.; Li, Y.; Ren, J.; Sun, Y. Genome-wide identification of SAUR genes in watermelon (Citrullus lanatus). Physiol. Mol. Biol. Plants Int. J. Funct. Plant Biol. 2017, 23, 619. [Google Scholar] [CrossRef]
- He, Y.; Liu, X.; Zou, T.; Pan, C.; Qin, L.; Chen, L.; Lu, G. Genome-Wide Identification of Two-Component System Genes in Cucurbitaceae Crops and Expression Profiling Analyses in Cucumber. Front. Plant Sci. 2016, 7, 899. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Zhang, J.; Sun, H.; Salse, J.; Lucas, W.J.; Zhang, H.; Zheng, Y.; Mao, L.; Ren, Y.; Wang, Z.; et al. The draft genome of watermelon (Citrullus lanatus) and resequencing of 20 diverse accessions. Nat. Genet. 2013, 45, 51–58. [Google Scholar] [CrossRef]
- Huang, S.; Li, R.; Zhang, Z.; Li, L.; Gu, X.; Fan, W.; Lucas, W.J.; Wang, X.; Xie, B.; Ni, P.; et al. The genome of the cucumber, Cucumis sativus L. Nat. Genet. 2009, 41, 1275–1281. [Google Scholar] [CrossRef]
- Sun, W.; Van Montagu, M.; Verbruggen, N. Small heat shock proteins and stress tolerance in plants. Biochim. Biophys. Acta 2002, 1577, 1–9. [Google Scholar] [CrossRef]
- Dombrovsky, A.; Tran-Nguyen, L.T.T.; Jones, R.A.C. Cucumber green mottle mosaic virus: Rapidly increasing global distribution, etiology, epidemiology, and management. Annu. Rev. Phytopathol. 2017, 55, 231–256. [Google Scholar] [CrossRef] [PubMed]
- Whitham, S.A.; Quan, S.; Chang, H.S.; Cooper, B.; Zhu, T.; Wang, X.; Hou, Y. Diverse RNA viruses elicit the expression of common sets of genes in susceptible Arabidopsis thaliana plants. Plant J. 2003, 33, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; An, M.; Xia, Z.; Bai, X.; Wu, Y. Transcriptome analysis of watermelon (Citrullus lanatus) fruits in response to Cucumber green mottle mosaic virus (CGMMV) infection. Sci. Rep. UK 2017, 7, 16747. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Functions of melatonin in plants: A review. J. Pineal Res. 2015, 59, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.Y.; Zhang, Y.; Xu, Y.P.; Qi, Z.Y.; Li, M.Q.; Ahammed, G.J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Reiter, R.J.; Yu, J.Q.; Zhou, J. HsfA1a upregulates melatonin biosynthesis to confer cadmium tolerance in tomato plants. J. Pineal Res. 2017, 62, e12387. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Cai, S.; Zhang, Y.; Wang, Y.; Ahammed, G.J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Yu, J.Q.; Reiter, R.J.; Zhou, J. Melatonin enhances thermotolerance by promoting cellular protein protection in tomato plants. J. Pineal Res. 2016, 61, 457–469. [Google Scholar] [CrossRef]
- Basha, E.; O Neill, H.; Vierling, E. Small heat shock proteins and a-crystallins: Dynamic proteins with flexible functions. Trends Biochem. Sci. 2012, 37, 106–117. [Google Scholar] [CrossRef]
- Piotrowska, J.; Hansen, S.J.; Park, N.; Jamka, K.; Sarnow, P.; Gustin, K.E. Stable formation of compositionally unique stress granules in virus-infected cells. J. Virol. 2010, 84, 3654–3665. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef]
- Hu, L.F.; Liu, S.Q. Genome-wide analysis of the MADS-box gene family in cucumber. Genet. Mol. Res. 2013, 12, 4317–4331. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Bowers, J.E.; Wang, X.; Ming, R.; Alam, M.; Paterson, A.H. Synteny and collinearity in plant genomes. Science 2008, 320, 486–488. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Feng, L.; Zhu, Y.; Li, Y.; Yan, H.; Xiang, Y. Comparative genomic analysis of the WRKY III gene family in populus, grape, Arabidopsis, and rice. Biol. Direct 2015, 10, 48. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Chang, J.; Zheng, J.; Dong, Y.; Liu, Q.; Yang, X.; Wei, C.; Zhang, Y.; Ma, J.; Zhang, X. Local melatonin application induces cold tolerance in distant organs of Citrullus lanatus L. via long distance transport. Sci. Rep. UK 2017, 7, 40858. [Google Scholar] [CrossRef] [PubMed]
- Voinnet, O.; Rivas, S.; Mestre, P.; Baulcombe, D. An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 2003, 33, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Yuan, J.; Gao, L.Y.; Zhao, S.; Jiang, W.; Huang, Y.; Bie, Z. Identification of suitable reference genes for gene expression normalization in qRT-PCR analysis in watermelon. PLoS ONE 2014, 9, e90612. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).