Abstract
Zornia brasiliensis Vogel (Leguminosae) is a species popularly known in Brazil as “urinária”, “urinana”, and “carrapicho”, it is popularly used as a diuretic and in the treatment of venereal diseases. A specific methodology to obtain a saponin-enriched fraction and high-performance liquid chromatography coupled with diode array detection, ion trap mass spectrometry, and TOF-MS (HPLC-DAD-ESI-MS/MS) was applied for the analysis of triterpene saponins. The MS and MS/MS experiments were carried out by ionization in negative mode. Molecular mass and fragmentation data were used to support the structural characterization of the saponins. Based on retention times, high-resolution mass determination and fragmentation, 35 oleanane-triterpene saponins were tentatively identified in Z. brasiliensis.
1. Introduction
The family Leguminosae consists of approximately 770 genera and 19,500 species. It previously included only three subfamilies, but a recent study reorganized the Leguminosae into six subfamilies [1]. These include the subfamily Papilionoideae, which contains the greatest number of genera, including the genus Zornia.
The genus Zornia contains approximately 80 species that are distributed throughout the world, mainly in tropical and subtropical regions. In Brazil, the genus is represented by 36 species, 15 of which are endemic [2,3]. Studies have investigated the pharmacological activities of compounds present in species of this genus, including cytotoxic, smooth muscle-relaxing, anticonvulsant, antibacterial, antitumor, anti-inflammatory, and antioxidant activities [4,5,6,7,8,9,10,11,12]. In turn, phytochemical studies have revealed the presence of isoflavonoids [13,14].
Zornia brasiliensis Vogel, a species popularly known in Brazil as “urinaria”, “urinana”, and “carrapicho”, is adopted in the alternative medicine as a diuretic and in the treatment of venereal diseases [15]. Z. brasiliensis is distributed in the North, Northeast, Center-West and Southeast regions of Brazil. It is associated with the Amazon, Caatinga, Cerrado, and Atlantic Forest phytogeographic domains [16], but is found mainly in the Brazilian Northeast [17] and Venezuela [18]. Two pharmacological studies of this species have reported the antinociceptive activity of 7-methoxyflavone isolated from its aerial parts [19] and the antitumor activity of its essential oil [20]. A recent phytochemical study of the aerial parts of this species resulted in the identification of 14 compounds, including flavones, flavanones, isoflavonoids, pterocarpans, chalcones, and a novel glycosylated dihydrochalcone—and this chalcone showed cytotoxic activity against HL-60 leukemic cells [21].
Hyphenated techniques, such as high-performance liquid chromatography (HPLC) coupled to mass spectrometry (MS) were used for the structural analysis and characterization of saponins [22,23,24,25,26,27,28,29]. This study represents the first contribution to describe the existence of triterpene saponins from the genus Zornia (Leguminosae). A total of 35 saponins were detected and tentatively identified by MS and MS/MS.
2. Results
2.1. Characterization of the Compounds by HPLC-ESI-MS/MS
Identification of triterpene saponins by high-performance liquid chromatography-electrospray ionization-tandem mass spectrometry (HPLC-ESI-MS/MS) with collision-induced dissociation (CID) under negative ESI-MS is a well-established and widely employed technique [24]. To obtain these MS/MS data, a low-resolution mass spectrometer was used; and the accurate mass was obtained by a high-resolution spectrometer, both using the same ionization parameters, and both coupled to a high-performance liquid chromatography system under the same chromatographic conditions. The 35 saponins that are present in the aerial parts of Z. brasiliensis and tentatively identified are shown on base peak chromatogram (Figure 1).
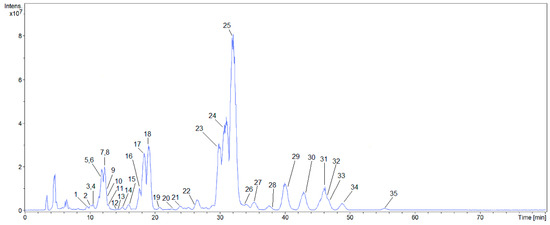
Figure 1.
Base peak chromatogram (BPC) of the concentrated fraction of saponins obtained by ESI-MS/MS in negative mode.
According to Pollier et al., after collision-induced dissociation (CID) under negative ESI-MS, the saponins undergo glycosidic cleavages, retaining the charge at the reducing end (e.g., aglycone containing fragment). In the MS/MS spectrum, daughter ions provide information on the sugar residues and the aglycone of the fragmented saponin. Sugar residues are mainly hexoses (e.g., glucose and galactose), 6-deoxyhexoses (e.g., rhamnose and fucose), pentoses (e.g., arabinose and xylose), and uronic acids (e.g., glucuronic acid and galacturonic).
The sugar residues and the aglycone could be identified from the MS/MS spectra. The typical losses represent water (−18 Da); and/or rhamnoside residue (−146 Da); and/or arabinose (−150 Da); and/or glucuronic acid (−158 Da). Therefore, the daughter aglycone ion, [Aglycone − H]− is based on this type or pattern of analysis. Based on this analysis, thirty-five saponins from Zornia braziliensis could be tentatively identified by mass spectrometry. The MS/MS data of the tentatively identified saponins are reported in Table 1.

Table 1.
Characterization of the saponins tentatively identified by HPLC-ESI-MS/MS in Z. brasiliensis.
2.1.1. Aglycones of Saponins of Z. Brasiliensis
In the MS/MS fragmentation spectra obtained for the 35 saponins, ions m/z 441–531 Da, characteristic of triterpene aglycones, were observed. Through the relationships of the deprotonated ions of the saponins—m/z 631.3856–999.5135 Da, and the respective fragment ions at the m/z 441–531 Da interval—10 aglycones were tentatively identified (Figure 2) [24]. Although these aglycones have been detected only as daughter ions with nominal m/z values, their molecular formulas can be predicted by the molecular formula of the exact mass of the parent ion and by the observed losses of the sugar residues [24]. Seven of these aglycones (olean-12-ene-3β,24-diol; soyasapogenol A, B, and E; kudzusapogenol A; wistariasapogenol A; and 22-O-acetate-soyasapogenol B) are of the oleanane type and have been previously reported in the family Leguminosae (Figure 2) [28,30,31,32,33].
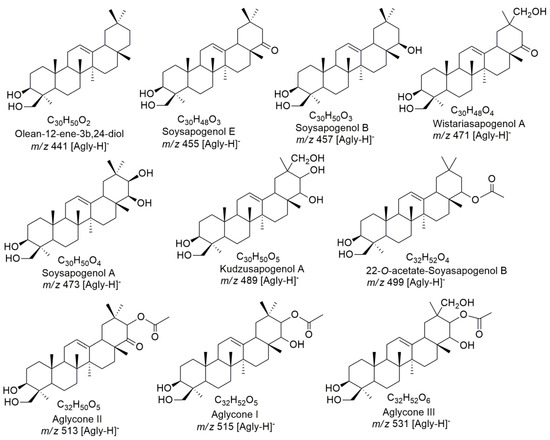
Figure 2.
Aglycones of Z. brasiliensis saponins tentatively identified by mass spectrometry.
No previous report of three of these sapogenins was found in the literature. Aglycone I has a fragment ion at m/z 515 [Agly − H]− and molecular formula C32H52O5, thus, we propose that aglycone would be an acetylation product of soysapogenin A (m/z 473), since it is possible to observe the loss of 42 Da (acetoxy) after subtraction of glycosidic residues, suggesting that acetoxy group is present in the sapogenin.
Aglycone II has a fragment ion m/z 513 [Agly − H]− and the molecular formula C32H50O5. The loss of 42 Da (acetoxy), similar to that observed in 21-acetoxy-soyasapogenol A (m/z 515), was observed. However, aglycone II has two fewer mass units, suggesting that C-22 was oxidized to generate a ketone [34]. The ion resulting from this loss of 42 Da, m/z 471, showed distinct fragmentation of wistariasapogenol A, a difference suggested by the absence of ion m/z 439 relative to the loss of 32 Da (CH4O) (Figure 2). Aglycone III has an m/z 531 [Agly − H]− and the molecular formula C32H52O6, it has been tentatively identified as an acetylated derivative of kudzusapogenol A, as evidenced by the 42 Da loss and the presence of the ion at m/z 489 (Figure 2).
This study has proposed the putative characterization of monodesmosidic saponins with the insertion of glycosides or its chains at the C-3 position, as observed in the saponin isolated and identified by NMR and supported by the literature which describes saponins present in the Leguminosae family [28,30,30,35]. The possibility of the presence of bidesmosidic saponins was removed due to the presence of fragments m/z 483 [M − H]− referring to the union of a dHex + Hex + HexA, m/z 453 [M − H]− dHex + Pen + HexA, m/z 337 [M − H]− Hex + HexA, and m/z 307 [M − H]− Pen + HexA, thus, demonstrating that the sugar chain is attached in the one position of the sapogenin, thus, excluding the possibility of bidesmosidic saponins.
2.1.2. Saponins of Z. brasiliensis
Peaks 1 and 2 showed ions at m/z 925.4814 and 779.4186, respectively, thus, they are isobars of compounds 27 and 28. However, they presented distinct fragmentation patterns that made it possible to propose that the difference between these compounds lies in their aglycones. In the tentative identification, compounds 1 and 2 have wistariasapogenol as an aglycone m/z 471 [Agly − H]−, this is proposed based on their fragmentation, since the fragment ion m/z 439 [Agly − H − CH4O]− is compatible with the loss of 32 Da corresponding to the loss of methanol. This suggests oxygenation in methyl C-30 rather than C-21, a ketone in the C-22 position similar to soyasapogenol E, and one other fragment ion m/z 407 [Agly − H − CH4O − CH4O]−, which is supposedly attributed to methanol at the C-23 position. Compound 1 was putatively identified as wistariasaponin A, which was previously isolated from a species of the family Leguminosae [35]; and compound 2 was tentatively identified as Pen-HexA-wistariasapogenol A.
Peaks 3 and 4 presented the fragment ion m/z 489 [Agly − H]− were tentatively identified as derivatives of kudzusapogenol A, a penta-oxygenated oleanane triterpene with oxygenations at C-21 and C-22, the oxygenation at C-29 is supported by the loss of 32 Da (methanol) observed in the fragment ion m/z 457 [Agly − H − CH4O]−. Compound 3 was putatively identified as Pen-HexA-kudzusapogenol A and compound 4 as subproside II, a saponin isolated from Sophorae subprostrata (Leguminosae) [36,43].
Group A soyasaponins are bidesmosidic, with insertion of the glycosidic chains in C-3 and C-22, and can be divided into acetylated and non-acetylated [22]. They have soyasapogenol A as the aglycone, which differs structurally from soyasapogenol B in the presence of a hydroxyl group at C-21 [44]. Five saponins that possess this aglycone were putatively identified (m/z 473). Compound 5 was identified as soyasaponin A3 [37], which was described by Curl et al. as a monodesmosidic derivative of soyasapogenol A. Compound 8, identified as Hex-HexA-soyasapogenol A, has already been described but not named [33], and compounds 6, 7, and 14 were putatively identified as dHex-Pen-HexA-soyasapogenol A, Pen-HexA-soyasapogenol A, and HexA-soyasapogenol A, respectively.
Peaks 9 and 10 showed the fragment ion m/z 531 [Agly − H]−, attributed to their aglycone and are here proposed as derivatives of aglycone III. It was also possible to observe the fragment ions m/z 489 and 457, which were attributed to the neutral loss of an acetoxy residue (42 Da) and a methanol (32 Da) of the aglycone. Peaks 9 and 10 were tentatively characterized as dHex-Pen-HexA-aglycone III and Pen-HexA-aglycone III, respectively.
Four saponins tentatively proposed possess aglycone II (m/z 513). These saponins were observed in peaks 11, 12, 13, and 15 and were proposed as dHex-Hex-HexA-aglycone II, Hex-HexA-aglycone II, dHex-Pen-HexA-aglycone II, and Pen-HexA-aglycone II, respectively.
Peak 20 was tentatively identified as Hex-HexA-22-O-acetate-soyasapogenol B. It presented fragment ion at m/z 499 [Agly − H]−, attributed to 22-O-acetate-soyasapogenol B; and fragment ion m/z 457, attributed to the neutral loss of the acetoxy residue (42 Da) in the aglycone.
Fragment ion at m/z 515 [Agly − H]−, attributed to this aglycone I, was observed in five compounds, 16, 17, 18, 19, and 22, all of which had the fragment ion at m/z 473 [Agly − H − C2H2O]−, compatible with the neutral loss of 42 Da relative to the acetoxy group present in the aglycone m/z 515 [Agly − H]−. These compounds were tentatively identified as dHex-Hex-HexA-aglycone I, dHex-Pen-HexA-aglycone I, Hex-HexA-aglycone I, Pen-HexA-aglycone I, and HexA-aglycone I, respectively.
Among soyasaponins A, B, and E, the soyasaponins of group E are the most restricted since they are considered photooxidation products of the soyasaponins of group B. The glycosidic chain at the C-3 position is the same for groups A, B, and E [41]. Peaks 21, 31, 32, 33, and 35 showed the same fragmentation pattern as that proposed for the olean-12-ene-3β,24-diol derivatives, however, the fragment ion attributed to the aglycone, m/z 455 [Agly − H]−, was proposed to be soyasapogenol E. Compound 21 was tentatively identified as soyasaponin Be [41], and 31, 32, and 33 were tentatively identified as soyasaponin Be', soyasaponin Bg, and soyasaponin Bg', respectively [44]. Compound 35 was tentatively characterized as hexA-soyasapogenol E.
The soyasaponins of group B have only one glycosidic chain at the C-3 position. They are commonly divided into two groups, one group has a conjugation in C-22 with 2,3-dihydro-2,5-dihydroxy-6-methyl-4-pyrone (DDMP) and is more abundant in soybean, and the members of the other group are DDMP-nonconjugated [22]. In this study, it was possible to observe peaks 23, 24, 25, 26, and 29, all of which had the fragment ion at m/z 457 [Agly – H]− attributed to soyasapogenol B. Compounds 23 and 25 were tentatively identified as soyasaponins I and II, respectively [44,45,46]. Compounds 24, 25, and 29 could be tentatively identified as soyasaponin III (or soyasaponin Bb'), soyasaponin IV (or soyasaponin Bc'), and soyasapogenol B monoglucuronide [28,38].
Peak 27 showed a precursor ion in m/z 925.5168 [M − H]− and its fragmentation, the ions at m/z 907 [M − H − H2O]−, m/z 863 [M − H − H2O − CO2]−, m/z 779 [M − H − dHex]−, m/z 717 [M − H − H2O − CO2 − dHex]−, m/z 599 [M − H − dHex − Hex]−, m/z 537 [M − H − H2O − CO2 − dHex − Hex]−, m/z 483 [M − H − Agly]−, m/z 441 [M − H − dHex − Hex − HexA]−, and m/z 439 [M − H − dHex − Hex − HexA]−. Thus, the uronic acid residue was observed, generating a loss of 158 and/or 160 Da (Figure 3). Peaks 28, 30, and 34 were tentatively identified as Hex-HexA-Olean-12-ene-3β,24-diol, dHex-Pen-HexA-Olean-12-ene-3β,24-diol, and Pen-HexA-Olean-12-ene-3β,24-diol, respectively (See proposed fragmentation pathways for all compounds in the Supplementary Materials).
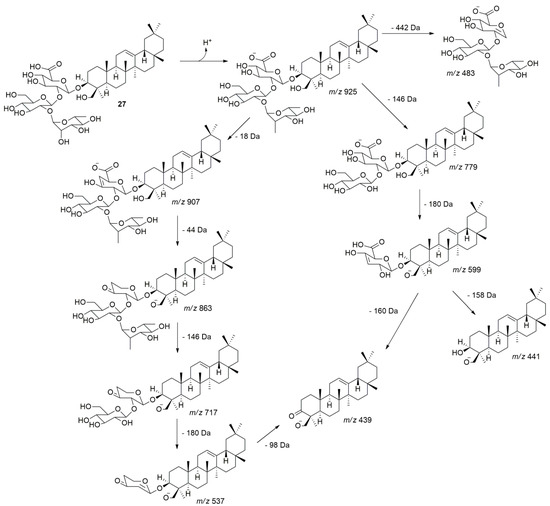
Figure 3.
Proposed fragmentation pathway for compound 27 in negative mode ESI-MS/MS. The pathway extends similarly to that for compounds 28, 30, and 34, generating the fragment ions m/z 441 and m/z 439 attributed to the aglycone olean-12-ene-3β,24-diol.
2.2. NMR Identification of Compound 25
After analysis of the saponin-enriched fraction by HPLC-DAD, the major peak observed in the chromatogram was peak 25, enabling its isolation. The compound 25 (Figure 4) has an [M − H]− ion at m/z 911.5006 (m/z 911.5010, calcd.), compatible with the molecular formula C47H75O17. The MS2 experiment showed m/z 765 [M − H − Rha]−, m/z 615 [M − H − Rha − Ara]−, and m/z 457 [M − H − Rha − Ara − Glu]− fragment ions corresponding to the consecutive losses of 146, 150, and 158 Da, respectively. In the 13C NMR spectrum, 7 methyls were observed attributed to the triterpenic sapogenin [δC 22.3 (Me-23), 15.3 (Me-25), 16.7 (Me-26), 25.0 (Me-27), 20.4 (Me-28), 32.7 (Me-29), 28.4 (Me-30)], a signal in δC 61.6 (C-24) characteristic of oxygenated carbon was also observed, showing that, in this structure, a Me-24 is oxygenated. Two other signals were observed in δC 121.7 (C-12) and 144.1 (C-13), attributed to the sp2 carbons—C-12 (methine) and C-13 (nonhydrogenated), thus, this sapogenin is a triterpenic oleanane-type. The presence of a C-19 methylene carbon (δC 46.4) and C-20 (δC 30.3) suggests that this sapogenin is β-amyrin, which is oxygenated at the C-22 positions (δC 74.2 ) and C-3 (δC 90.0). In the 1H-NMR it was possible to observe the signals of 7 methyls [δH 1.08 (23-Me), 0.79 (25-Me), 0.88 (26-Me), 1.04 (Me-27), 0.74 (Me-28), 0.96 (Me-29), 0.96 (Me-30)], at the same time, a broad singlet at δH 5.16 with integral for one hydrogen, attributed to a sp2 methyl carbon. The signals at δH 3.80 (d, J = 11.0 Hz, 1H) and 3.04 (m, 1H) were assigned to Ha and Hb-24, respectively, δH 3.12 (m, 1H) to H-3, and δH3.23 (m, 1H) to H-22. In the 1H x 13C-HMBC spectrum, a correlation of the anomeric proton H-1′ [δH 4.13 (d, J = 7.5 Hz, 1H)] of β-glucuronic acid was observed with C-3 (δC 90.0), confirming the insertion of β-glucuronic acid in the C-3-position of the sapogenin. Another contour map between H-1′′′ [δH 4.97 (brs, 1H)] at three bonds distance with C-2′′ (δC 77.6) confirming the insertion of α-rhamnose into C-2′′ of β-arabinose. The HMBC also attributed a correlation of the proton H-2′ [δH 3.33 (m)] at two bonds distance with C-1′ (δC 103.9) and three bonds with C-1′′ (δC 100.7), thus confirming that β-arabinose is bonded to β-glucuronic acid and this insertion occurs at C-2′ glucuronic acid. Another contour map present in HMBC attributed Me-28 [δH 0.74 (s, 3H)] with C-22 (δC 74.2), C-16 (δC 28.0), C-17 (δC 37.0), and C-18 (δC 44.7) indicating that C-22 supports the hydroxyl. This information was corroborated by another contour map between H-16a [δH 1.33 (m)] and C-22 (δC 74.2). A cross-correlation was also observed between Me-29/30 [δH 0.96 (s, 6H)] with C-19 (δC 46.1), C-20 (δC 30.3), C-21 (δC 41.2), and C-29 (δC 32.7); H-19 [δH 1.67 (m) and 0.88 (m)] with C-20 (δC 30.3), C-29 (δC 32.7) and C-30 (δC 28.4) and H-21 [δH 1.30 (m)] with C-17 (δC 37.0), C-22 (δC 74.2) and C-30 (δC 28.4). It was also possible to observe, in HMBC, a three-bond correlation between H-24a [δH 3.80 (d, J = 11.0 Hz)], C-3 (δC 90.0) as well as Me-23 (δC 22.3). Thus, 25 corresponds to soyasaponin II (Figure 4). Comparison of the 1H and 13C-NMR spectral data of compound 25 with data from the literature corroborate this proposal. Thus, the m/z 457 fragment ion corresponds to the sapogenin soyasapogenol B (Figure 2) [44,45].
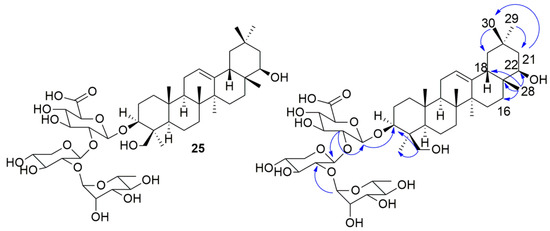
Figure 4.
Soyasaponin II (25) isolated from Z. brasiliensis. ( ) indicates some key HMBC correlations for compound 25.
) indicates some key HMBC correlations for compound 25.
 ) indicates some key HMBC correlations for compound 25.
) indicates some key HMBC correlations for compound 25.
Considering that the isolation, purification, and elucidation of triterpene saponins are difficult and time-consuming due to the high polarity and structural similarity of these compounds [23], the present study enabled us to obtain and chemically analyze a concentrated fraction of saponins of Z. brasiliensis for the first time. In this study, 35 triterpene saponins were tentatively identified by HPLC-ESI-MS/MS, suggesting a homologous series. In addition, saponins have a diversification of applications such as biological and pharmaceutical, for example, immunostimulating, hypocholesterolemic, and anticarcinogenic properties, as well as applications in the food, agriculture, and cosmetics industries. This fact led to the extraction and identification of saponins in numerous species [47] due to their relevance. This is the first report of the occurrence of saponins in the genus Zornia, placing the species Z. brasiliensis in the list of the bioproducing species of this important class of secondary metabolites that are the saponins.
3. Materials and Methods
3.1. Reagents and Materials
The NMR analyses were performed on a Varian NMR system 500-MHz spectrometer (Palo Alto, CA, USA) operated at 500 MHz for 1H-NMR and at 125 MHz for 13C-NMR; and on a Bruker Ascend 400 spectrometer (Bruker, Billerica, MA, USA) operated at 400 MHz for 1H-NMR and at 100 MHz for 13C NMR. The samples were prepared in deuterated dimethyl sulfoxide (DMSO-d6) (Cambridge Isotope Laboratories, Tewksbury, MA, USA) containing TMS as internal standard. Silica gel (ART 7734, Merck, Darmstadt, Germany; particle size 0.060–0.200 mm and 70–230 mesh) was used for column chromatography (CC). To obtain the crude extract and in chromatographic fractionation, the following solvents were used: Ethanol, hexane, dichloromethane, ethyl acetate, methanol, and n-butanol (Tedia, Fairfield, CT, USA). For the HPLC analyses, the following solvents were used as the mobile phase: HPLC-grade acetonitrile (Tedia), ultrapure water obtained using a Milli-Q purification system (Millipore, Burlington, NJ, USA), and formic acid (Qhemis, Indaiatuba, Brazil). To obtain mass spectra by direct injection, an Ion Trap-amaZonX spectrometer (Bruker, Billerica, MA, USA) was used for electrospray ionization mass spectrometry (ESIMS, low resolution), and a micrOTOF II spectrometer was used for high-resolution ESIMS (HRESIMS, Bruker, Billerica, MA, USA) at 3.5 kV capillary voltage, ESI in negative mode, 500 V end plate offset, 8.0 psi nebulizer, dry gas (N2) at a flow rate of 5.0 L/h, and a temperature of 200 °C. Spectra (m/z 50–1000) were recorded every 2.0 s. Collision-induced dissociation (CID) fragmentation was achieved in auto MS/MS mode using the advanced resolution mode for MS and MS/MS mode.
3.2. Plant Material
The aerial parts of Z. brasiliensis Vogel (Leguminosae) were collected in the municipality of Serra Branca (07°29′46′′S and 36°44′36′′W; altitude, 712 m) in the state of Paraíba, Brazil, in March 2016. Collection authorization No. 53894-1 was granted by the Chico Mendes Institute for Biodiversity Conservation (ICMBio, Brasília, Brazil) through the System of Authorization and Information on Biodiversity (SISBIO, Brasília, Brazil), and access registration in the National Management System of Genetic Patrimony and Associated Traditional Knowledge (SISGEN, Curitiba, Brazil) was obtained under number ADD107E. The species was identified by botanist Dr. José Iranildo Miranda de Melo of the Paraíba State University (UEPB, Campina Grande, Brazil). An exsiccatae was deposited at Arruda Câmara Herbarium (ACAM, Campina Grande, Brazil), Campus I, UEPB, under the number of registration 1862.
3.3. Extraction and Isolation of Z. brasiliensis Constituents
The dry and pulverized aerial parts of Z. brasiliensis (5 kg) were extracted with ethanol to obtain a crude ethanolic extract (CEE) which was partitioned with hexane, dichloromethane, ethyl acetate, ethyl acetate–methanol (90:10, v/v) and ethyl acetate–methanol (50:50, v/v)in order to obtain its respective fractions [21].
An aliquot (20.0 g) of the 50% ethyl acetate–methanol fraction was dissolved in 300 mL of distilled water, followed by partition with n-butanol (300 mL) three times. The extractive solution of the n-butanol phase was then concentrated on a rotary evaporator under reduced pressure at a temperature of 50 °C. The concentrated material was redissolved in methanol–ethyl acetate (1:5, v/v), and the saponins were precipitated by the addition of methanol. This material was decanted for 72 hours at room temperature. The pellet was then resuspended in methanol (200 mL) and concentrated on a rotary evaporator under reduced pressure at 40 °C. In this way, a concentrated fraction of saponins (1.1 g) was obtained [48].
3.4. HPLC-DAD Conditions
An analytical HPLC instrument (Shimadzu-Prominence, Kyoto, Japan) equipped with an LC-20AT pump, an SIL-20A HT auto injector, a CTO-20A oven, an SPD-M20A detector, and a CBM-20A controller unit was employed. The columns used were a Phenomenex Gemini® C18 (250.0 mm × 4.6 mm internal diameter [id], filled with 5.0-µm particles, Torrance, CA, USA) and a Security Guard Gemini® C18 precolumn (4.0 mm × 3.0 mm id, filled with 5.0-µm particles, Torrance, CA, USA). The elution method used water (0.1% formic acid) (A) and methanol (B). The gradient was 70.0% B (0.01 min) to 80.0% B (50 min), returning to 70.0% B (65 min), and remaining at 70.0% B until 80 min with a mobile phase flow rate of 0.6 mL/min, an injection volume of 20.0 µL, and a detection wavelength of 205 nm. The applied samples were filtered through 0.45-µm nylon membranes (Tedia, Fairfield, CT, USA).
Preparative chromatographic analyses were performed using an HPLC system (Shimadzu, Kyoto, Japan) equipped with an LC-6AD binary solvent pump, a Rheodyne injector, an SPD-M10A diode array detector, and an SCL-10A system controller. An ACE C18 column (250.0 mm × 21.2 mm id, 5.0-μm particle size) was used. The method used was similar to the method used in the analytical runs. The preparative scale injection volume was 100.0 µL, flow was conducted at 8.0 mL/min, and the detection wavelength was 205 nm. The applied samples were filtered through 0.45-µm nylon membranes (Tedia).
3.5. HPLC-ESI-MS/MS Conditions
A Shimadzu (Prominence, Kyoto, Japan) HPLC instrument equipped with an LC-20AT binary solvent pump, an SIL-20A auto injector, a DGU-20A degassing system, an SPD-M20A diode array detector, and a CBM-20A controller was used. A Phenomenex Gemini® C18 column (250.0 mm × 4.6 mm id, filled with 5.0-µm particles) and a SecurityGuard Gemini® C18 precolumn (4.0 mm × 3.0 mm id, filled with 5.0-µm particles) were used. The chromatographic method was the same as the method developed for analytical HPLC. The applied samples were filtered through 0.45-µm nylon membranes (Tedia). This HPLC instrument was coupled to an Ion Trap-amaZonX low-resolution mass spectrometer (Bruker) that used the electrospray ionization technique. The analysis parameters of the Ion-Trap spectrometer were 4.5 kV capillary, ESI in negative ion mode, a 500 V end plate offset, a 35.0 psi nebulizer, a dry gas (N2) flow rate of 8.0 L/h, and a temperature of 300 °C. CID fragmentation was achieved in auto MS/MS mode with the advanced resolution mode for MS and MS/MS mode. Spectra (m/z 200–1200) were recorded every 2.0 s.
For high-resolution HPLC-ESI-MS analyses, we used the HPLC instrument described above with the same column and the same chromatographic conditions. The instrument was coupled to a micrOTOF II high-resolution mass spectrometer (Bruker) that used the electrospray ionization technique. The micrOTOF II spectrometer analysis parameters were 4.0 kV capillary, ESI in negative ion mode, a 500 V end plate offset, a 35.0 psi nebulizer, a dry gas (N2) flow rate of 8.0 L/h, and a temperature of 300 °C. Spectra (m/z 50–1000) were recorded every 2.0 s. The sodium formate was used as the internal calibrator during the chromatographic run. The equipment resolution is ~15,000, however, when operating in HPLC-HRESIMS mode it drops to ~11,000.
Supplementary Materials
Supplementary data (Figures S1–S27) associated with this article, including HRESIMS, MS/MS, 1D, and 2D-NMR spectra data and the proposed fragmentation pathways of the tentatively identified compounds can be found in the online version.
Author Contributions
Y.M.N., L.S.A., R.L.L. and V.C.O.C. performed the experiments; J.I.M.d.M., botanic material collection; R.B.-F., M.S.S. and J.F.T. analyzed the data; Y.M.N., M.S.S. and J.F.T. wrote the paper.
Funding
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior-Brasil (CAPES)-Finance Code 001.
Acknowledgments
We thank the Graduate Program in Natural and Synthetic Bioactive Products for their support.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Legume Phylogeny Working Group. A new subfamily classification of the Leguminosae based on a taxonomically comprehensive phylogeny. Taxon 2017, 66, 44–77. [Google Scholar] [CrossRef]
- Perez, A.P.F.; Lewis, G.P.; Queiroz, R.T.; Silva, J.S.; De Azevedo Tozzi, A.M.G.; Rodrigues, K.F. Fruit as diagnostic characteristic to recognize Brazilian species of Zornia (Leguminosae, Papilionoideae). Phytotaxa 2015, 219, 27–42. [Google Scholar] [CrossRef]
- Fortuna-Perez, A.P. O gênero Zornia J.F. Gmel. (Leguminosae, Papilionoideae, Dalbergieae): Revisão taxonômica das espécies ocorrentes no Brasil e filogenia. Ph.D. Thesis, Universidade Estadual de Campinas—Campinas-SP, Campinas, Brazil, 2009. [Google Scholar]
- Belcavello, L.; Cunha, M.R.H.; Andrade, M.A.; Batitucci, M.C.P. Citotoxicidade e danos ao DNA induzidos pelo extrato de Zornia diphylla, uma planta medicinal. Nat. Line 2012, 10, 140–145. [Google Scholar]
- Rojas, A.; Bah, M.; Rojas, J.I.; Serrano, V.; Pacheco, S. Spasmolytic activity of some plants used by the otomi indians of Queretaro (Mexico) for the treatment of gastrointestinal disorders. Phytomedicine 1999, 6, 367–371. [Google Scholar] [CrossRef]
- Greetha, K.M.; Shilpa, S.; Murugan, V. Anticonvulsant Activity of the Methanolic Extract of Whole Plant of Zornia diphylla (Linn) Pers. J. Pharm. Res. 2012, 5, 3670–3672. [Google Scholar]
- Arunkumar, R.; Nair, S.A.; Subramoniam, A. Induction of cell-specific apoptosis and protection of mice from cancer challenge by a steroid positive compound from Zornia diphylla (L.) Pers. J. Pharmacol. Pharmacother 2012, 3, 233–241. [Google Scholar]
- Obi, C.L.; Ramalivhana, J.; Samie, A.; Igumbor, E.O. Prevalence, pathogenesis, antbiotic susceptibility profiles, and in-vitro activity of selected medicinal plants against Aeromonas isolated from stool samples of patients in the Venda region of South Africa. J. Health Popul. Nutr. 2007, 25, 428–435. [Google Scholar] [PubMed]
- Laxane, S.N.; Swarnkar, S.K.; Setty, M.M. Antioxidant studies on the ethanolic extract of Zornia gibbosa. Pharmacol. Online 2008, 1, 319–330. [Google Scholar]
- Laxane, S.N.; Swarnkar, S.K.; Zanwar, S.B.; Setty, M.M. Anti-inflammatory studies of the alcoholic extract of Zornia gibbosa. Pharmacol. Online 2011, 1, 67–76. [Google Scholar]
- Brahmachari, G.; Ghosh, S.; Mondal, S.; Jash, S.K.; Mandal, L.C.; Mondal, A. Cyclic voltammetric studies with plant extracts of some traditionally used Indian medicinal plants to evaluate their antioxidant potential. Biochem. Indian J. 2009, 3, 32–35. [Google Scholar]
- Arunkumar, R.; Nair, S.A.; Rameshkumar, K.B.; Subramoniam, A. The essential oil constituents of Zornia diphylla (L.) Pers., and anti-inflammatory and antimicrobial activities of the oil (Article). Rec. Nat. Prod. 2014, 8, 385–393. [Google Scholar]
- Leuner, O.; Havlik, J.; Prokudina, E.; Novy, P.; Kokoska, L. Distribution of isoflavones and coumestrol in neglected tropical and subtropical legumes. J. Sci. Food Agric. 2013, 93, 575–579. [Google Scholar] [CrossRef] [PubMed]
- Ren, F.Z.; Gao, Y.Q.; Cheng, X.X.; Li, L.H.; Chen, S.H.; Zhang, Y.L. Study on Chemical Constituents of Zornia diphylla. Chin. Pharm. J. 2012, 47, 179–181. [Google Scholar]
- Agra, M.F.; Freitas, P.F.; Barbosa-Filho, J.M. Synopsis of the plants know as medicinal and poisonous in Northeast of Brazil. Rev. Bras. Farmacogn. 2007, 17, 114–140. [Google Scholar] [CrossRef]
- BFG—The Brazil Flora Group. Growing knowledge: An overview of Seed Plantdiversity in Brazil. Rodriguésia 2015, 6, 1085–1113. [Google Scholar]
- Mohlenbrock, R.H. A monograph of the leguminous genus Zornia. Webbia 1961, 16, 1–141. [Google Scholar] [CrossRef]
- Missouri Botanical Garden. 2017. Available online: http://www.tropicos.org/Name/13035232 (accessed on 22 October 2018).
- Da Silva, A.D.S.; Cavalcante-Silva, L.H.A.; Da Matta, C.B.B.; Silva, D.D.F.; Araújo, M.V.D.; Tavares, J.F.; Alexandre-Moreira, M.S. Antinociceptive effect of 7-methoxyflavone isolated from Zornia brasiliensis. Nat. Prod. Res. 2013, 27, 1695–1699. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.V.; Menezes, L.R.; Rocha, S.L.; Baliza, I.R.; Dias, R.B.; Rocha, C.A.; Soares, M.B.; Bezerra, D.P. Antitumor properties of the leaf essential oil of Zornia brasiliensis. Planta Med. 2015, 81, 563–567. [Google Scholar] [CrossRef]
- Nascimento, Y.M.; Abreu, L.S.; Lima, R.L.; Silva, A.D.S.; Costa, V.C.O.; Melo, J.I.M. Zornioside, a dihydrochalcone C-glycoside, and other compounds from Zornia brasiliensis. Rev. Bras. Farmacogn. 2017, 28, 192–197. [Google Scholar] [CrossRef]
- Huhman, D.V.; Sumner, L.W. Metabolic profiling of saponins in Medicago sativa and Medicago truncatula using HPLC coupled to an electrospray ion-trap mass spectrometer. Phytochemistry 2002, 59, 347–360. [Google Scholar] [CrossRef]
- Ling, Y.; Lin, Z.; Zha, W.; Lian, T.; You, S. Rapid Detection and Characterisation of Triterpene Saponins from the Root of Pulsatilla chinensis (Bunge) Regel by HPLC-ESI-QTOFMS/MS. Phytochem. Anal. 2016, 27, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Pollier, J.; Morreel, K.; Geelen, D.; Goossens, A. Metabolite Profiling of Triterpene Saponins in Medicago truncatula Hairy Roots by Liquid Chromatography Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. J. Nat. Prod. 2011, 74, 1462–1476. [Google Scholar] [CrossRef] [PubMed]
- Negri, G.; Tabach, R. Saponins, tannins and flavonols found in hydroethanolic extract from Periandra dulcis roots. Rev. Bras. Farmacogn. 2013, 23, 851–860. [Google Scholar] [CrossRef]
- Perret, C.; Wolfender, J.L.; Hostettmann, K. LC/ES-MS Analysis of Triterpene Glycosides: Rapid Estimation of the Saponin Content of Dried Berries of Phytolacca dodecandra. Phytochem. Anal. 1999, 10, 272–278. [Google Scholar] [CrossRef]
- Zhang, W.; Popovich, D.G. Chemical and Biological Characterization of Oleanane Triterpenoids from Soy. Molecules 2009, 14, 2959–2975. [Google Scholar] [CrossRef]
- Berhow, M.A.; Kong, S.B.; Vermillion, K.E.; Duval, S.M. Complete Quantification of Group A and Group B Soyasaponins in Soybeans. J. Agric. Food Chem. 2006, 54, 2035–2044. [Google Scholar] [CrossRef]
- Jin, M.; Yang, Y.; Su, B.; Ren, Q. Rapid quantification and characterization of soyasaponins by high-performance liquid chromatography coupled with electrospray mass spectrometry. J. Chromatogr. A 2006, 1108, 31–37. [Google Scholar] [CrossRef]
- Takada, Y.; Sasama, H.; Sayama, T.; Kikuchi, A.; Kato, S.; Ishimoto, M.; Tsukamoto, C. Genetic and chemical analysis of a key biosynthetic step for soyasapogenol A, an aglycone of group A saponins that influence soymilk flavor. Theor. Appl. Genet. 2013, 126, 721–731. [Google Scholar] [CrossRef]
- Konoshima, T.; Kozuka, M.; Haruna, M.; Ito, K.; Kimura, T. The structures of New Triterpenoids from Wistaria brachybotrys Sieb. et Zucc. Chem. Pharm. Bull. 1989, 37, 1550–1553. [Google Scholar] [CrossRef]
- Takeshita, T.; Hamada, S.; Nohara, T. New triterpenoid sapogenols from Abrus cantoniensis (I). Chem. Pharm. Bull. 1989, 37, 846–848. [Google Scholar] [CrossRef]
- Arao, T.; Idzu, T.; Kinjo, J.; Nohara, T.; Isobe, R. Oleanene-type triterpene glycosides from puerariae radix. III. Three new saponins from Pueraria thomsonii. Chem. Pharm. Bull. 1996, 44, 1970–1972. [Google Scholar] [CrossRef]
- Kitagawa, I.; Wang, H.K.; Taniyama, T.; Yoshikawa, M. Reinvestigation of the Structures os Soyasapogenols A, B and E, Oleanene-Sapogenols from Soybean. Structures od Soyasaponins I, II, e III. Chem. Pharm. Bull. 1988, 36, 153–161. [Google Scholar] [CrossRef]
- Price, K.R.; Johnson, I.T.; Fenwick, G.R.; Malinow, M.R. The chemistry and biological significance of saponins in foods and feedingstuffs. Crit. Rev. Food Sci. Nutr. 1987, 26, 27–135. [Google Scholar] [CrossRef] [PubMed]
- Konoshima, T.; Kozuka, M.; Haruna, M.; Ito, K.; Kimura, T.; Tokuda, H. Studies on the constituents of leguminous plants. XII. The structures of new triterpenoid saponins from Wistaria brachybotrys Sieb. et Zucc. Chem. Pharm. Bull. 1989, 37, 2731–2735. [Google Scholar] [CrossRef][Green Version]
- Ding, Y.; Takeshita, T.; Yokoyama, K.; Kinjo, J.; Nohara, T. Triterpenoid Glycosides from Sophorae Subprostratae Radix. Chem. Pharm. Bull. 1992, 40, 139–142. [Google Scholar] [CrossRef]
- Curl, C.L.; Price, K.R.; Fenwick, G.R. Soyasaponin A3, a New Monodesmosidic Saponin Isolated from the Seeds of Glycine max. J. Nat. Prod. 1988, 51, 122–124. [Google Scholar] [CrossRef]
- Woldemichael, G.M.; Montenegro, G.; Timmermann, B.N. Triterpenoidal lupin saponins from the Chilean legume Lupinus oreophilus Phil. Phytochemistry 2003, 63, 853–857. [Google Scholar] [CrossRef]
- Giuliana, B.; Raffaella, P.; Cecilia, F.C.; Maria, A.A.; Cataldi, T.R.; Philippe, S.K.; Alessandro, B.; Daniela, R.; Luigi, M. Determination of soyasaponins in Fagioli di Sarconi beans (Phaseolus vulgaris L.) by LC-ESI-FTICR-MS and evaluation of their hypoglycemic activity. Anal. Bioanal. Chem. 2018, 410, 1561–1569. [Google Scholar]
- Decroos, K.; Vincken, J.P.; Heng, L.; Bakker, R.; Gruppen, H.; Verstraete, W. Simultaneous quantification of differently glycosylated, acetylated, and 2,3-dihydro-2,5-dihydroxy-6-methyl-4H-pyran-4-one-conjugated soyasaponins using reversed-phase high-performance liquid chromatography with evaporative light scattering detection. J. Chromatogr. A 2005, 1072, 185–193. [Google Scholar] [CrossRef]
- Udayama, M.; Ohkawa, M.; Yoshida, N.; Kinjo, J.; Nohara, T. Structures of three new oleanene glucuronides isolated from Lathyrus palustres var. Pilosus and hepatoprotective activity. Chem. Pharm. Bull. 1998, 46, 1412–1415. [Google Scholar] [CrossRef][Green Version]
- Sakamoto, S.; Kuroyanagi, M.; Ueno, A.; Sekuta, S. Triterpenoid saponins from Sophora subprostrata. Phytochemistry 1992, 31, 1339–1342. [Google Scholar] [CrossRef]
- Rupasinghe, H.V.; Jackson, C.J.C.; Poysa, V.; Di Berardo, C.; Bewley, J.D.; Jenkinson, J. Soyasapogenol A and B Distribution in Soybean (Glycine max L. Merr.) in Relation to Seed Physiology; Genetic Variability, and Growing Location. J. Agric. Food Chem. 2003, 51, 5888–5894. [Google Scholar] [CrossRef] [PubMed]
- Kudou, S.; Tonomura, M.; Tsukamoto, C.; Uchida, T.; Sakabe, T.; Tamura, N.; Okubo, K. Isolation and Structural Elucidation of DDMP-Conjugated Soyasaponins as Genuine Saponins from Soybean Seeds. Biosci. Biotechnol. Biochem. 1993, 57, 546–550. [Google Scholar] [CrossRef]
- Dalluge, J.J.; Eliason, E.; Frazer, S. Simultaneous Identification of Soyasaponins and Isoflavones and Quantification of Soyasaponin B in Soy Products, Using Liquid Chromatography/Electrospray Ionization-Mass Spectrometry. J. Agric. Food Chem. 2003, 51, 3520–3524. [Google Scholar] [CrossRef] [PubMed]
- Güçlü-Ustündağ, O.; Mazza, G. Saponins: Properties, applications and processing. Crit. Rev. Food Sci. Nutr. 2007, 47, 231–258. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, A.P.; Silva, B.P.; Parente, J.P. A new complex triterpenoid saponin from Samanea saman with haemolytic activity and adjuvant effect. Phytochem. Lett. 2012, 5, 626–631. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).