Abstract
Background/Objectives: Attention-deficit/hyperactivity disorder (ADHD) is a highly heritable neurodevelopmental condition, and pharmacogenetic studies aim to clarify interindividual variability in treatment responses and adverse effects. Despite increasing research, the field remains fragmented. This review provides a bibliometric analysis of ADHD pharmacogenetics (2005–2025), identifying its intellectual foundations, thematic structure, and global distribution. Methods: A bibliometric search was conducted in Scopus and Web of Science, retrieving 711 documents published between 2005 and July 2025. Data were analyzed with the Bibliometrix R package and Biblioshiny interface, applying bibliometric mapping, Bradford’s Law, co-word analysis, and thematic mapping. Only peer-reviewed journal articles, books, and book chapters were included to ensure scientific rigor. Results: The dataset shows a modest annual growth rate but strong impact, with an average of 29.6 citations per article. Highly cited works converge into four domains: (i) clinical guidelines and pharmacological treatments; (ii) cognitive heterogeneity and subtypes; (iii) neurodevelopmental and genetic mechanisms; (iv) environmental and health-related influences. Geographically, the United States leads with 24.8% of publications, followed by Brazil, China, and European countries. Keyword analysis reveals two main clusters: a clinical–therapeutic pole (methylphenidate, atomoxetine, child) and a genetic–molecular pole (dopamine transporter, SNPs, genotype). Conclusions: ADHD pharmacogenetics shows consolidation with strong clinical and genetic cores but limited integration of comorbidity, adult populations, and non-stimulant treatments. Future research should prioritize multi-center cohorts, multi-omic designs, and stronger international collaboration to advance precision medicine in ADHD.
1. Introduction
1.1. Epidemiology, Clinical Features, and Burden of ADHD
Attention-deficit/hyperactivity disorder (ADHD) is one of the most prevalent neurodevelopmental disorders in childhood, with a global prevalence range from 3% and 6% [1], reaching approximately 5% in children and adolescents [2] and 2.5% in adults [3]. These rates show regional variability, being slightly higher in North America and lower in Asia and Europe, although methodological differences partly account for this variation [4]. Clinically, ADHD is characterized by symptoms of inattention, hyperactivity, and impulsiveness. Its etiology is complex and multifactorial, involving genetic, environmental, and neurobiological components, which contribute to the marked interindividual variability in treatment response [1].
Longitudinal studies estimate that 60% and 70% of children diagnosed with ADHD continue to experience symptoms in adolescence, and around 50% present clinically significant manifestations into adulthood [5,6]. This persistence has a substantial impact on academic achievement, occupational functioning, and social relationships. Despite its high prevalence and long-term burden, ADHD remains underdiagnosed and undertreated in adults, particularly in low- and middle-income countries, due to stigma, limited access to mental health care, and insufficient professional training [7].
1.2. Pharmacological Treatment of ADHD and Neurobiological Basics
Against this background, pharmacogenetics, the discipline studying how genetic variability influences drug response, emerges as a promising tool to optimize ADHD management. Identifying genetic polymorphisms associated with the response to stimulant medications such as methylphenidate or amphetamines, as well as non-stimulants, could contribute to more personalized prescribing, reducing adverse effects and increasing therapeutic success rates [8].
In current pharmacological management, stimulants (methylphenidate and amphetamines) remain the first-line treatment due to their superior efficacy and acceptability; a network meta-analysis of 133 trials supports methylphenidate in children/adolescents and amphetamines in adults, consistent with AAP pediatric guidelines [9,10]. Among non-stimulants, norepinephrine reuptake inhibitors such as atomoxetine and extended-release viloxazine have shown clinical benefits, though with a slower onset of action and distinct side effect profiles; viloxazine has demonstrated efficacy in adults (phase III trial) as well as in pediatric populations [11]. Genetic and neurobiological studies have primarily informed pharmacogenetic hypotheses by implicating dopaminergic, noradrenergic, and serotonergic pathways in ADHD.
First, in the dopaminergic pathway, variable number tandem repeat (VNTR) polymorphisms in the dopamine transporter gene (SLC6A3/DAT1) have been shown to regulate gene expression, with the 10-repeat allele associated with altered transporter levels in both brain and lymphocytes, potentially affecting synaptic dopamine availability and contributing to ADHD risk [12,13]. Dopamine receptor gene variants, particularly in DRD4 and DRD5, are also associated with ADHD, influencing cognitive and behavioral phenotypes [13,14]. Functional studies further support the role of dopamine D2 receptors in modulating signal transduction and behavior relevant to ADHD [15].
Second, noradrenergic involvement is supported by associations between ADHD and polymorphisms in the norepinephrine transporter gene (SLC6A2) and the α2A-adrenergic receptor gene (ADRA2A), with specific variants such as rs28386840 in SLC6A2 linked to reduced promoter activity and altered noradrenergic signaling [16,17]. These genetic findings are consistent with the established role of noradrenergic dysfunction in attention regulation and executive function deficits in ADHD [18].
Finally, serotonergic system genes, including SLC6A4 and HTR2A, are implicated in ADHD, particularly in relation to impulsivity and aggression. Meta-analytic evidence supports associations between serotonergic gene variants and ADHD phenotypes, and specific HTR2A polymorphisms may predispose to aggressive traits [19,20].
Despite these advances, controversies remain regarding the strength and reproducibility of genetic associations, as many candidate gene studies have yielded inconsistent results across populations and sample sizes. Similarly, while polygenic risk scores and genome-wide association studies offer new perspectives, their clinical translation into pharmacogenetics-guided ADHD treatment is still debated. These divergences highlight the need for integrative approaches that combine genetics with environmental, developmental, and clinical factors to better explain variability in treatment response.
1.3. Genetic and Pharmacogenetic Evidence: Associations and Limitations
Pharmacogenetic research in ADHD has identified multiple genetic variants linked to disorder risk and medication response; however, most associations show small effect sizes and limited reproducibility. Despite growing interest in using genetic information to guide treatment, current evidence does not demonstrate clear clinical utility, and major guidelines do not recommend routine pharmacogenetic testing for ADHD.
Among the genes most frequently studied in attention-deficit/hyperactivity disorder (ADHD)—DAT1 (SLC6A3), DRD4, DRD5, SLC6A2, CYP2D6, ADRA2A, SLC6A4, and HTR2A—the variants showing the most consistent clinical associations with the disorder are DAT1, DRD4, DRD5, SLC6A4, and ADRA2A, although the magnitude of their effects remains small, with odds ratios typically ranging from 1.1 to 1.3 for the main risk alleles, reflecting only a modest increase in ADHD susceptibility [21,22]. The DAT1 10-repeat allele displays a statistically significant but weak association with childhood ADHD, mainly in European populations, and is more closely linked to the combined subtype [19,22]. The DRD4 exon 3 7-repeat allele represents one of the most replicated findings, with greater risk observed in combined and inattentive presentations (OR ≈ 1.3), while the DRD5 148 bp allele shows reproducible though subtype-nonspecific associations [21,23]. Serotonergic involvement is suggested by the SLC6A4 5-HTTLPR polymorphism, which exhibits modest effects and moderate reproducibility, including interaction patterns with ADRA2A, whose rs553668 variant increases risk particularly for inattentive symptoms and shows cumulative effects when combined with SLC6A4 alleles [19,24]. By contrast, CYP2D6, SLC6A2, and HTR2A do not demonstrate consistent or reproducible associations with ADHD risk; notably, CYP2D6 is clinically relevant primarily for atomoxetine metabolism rather than for core ADHD phenotypes [19,24,25]. Overall, the degree of reproducibility for these genetic associations is moderate, with consistent findings for DAT1, DRD4, and DRD5 across multiple meta-analyses and populations; however, effect sizes remain small and insufficient for clinical utility in diagnosis or treatment selection. The clinical phenotypes that appear most influenced by these genetic patterns are the combined and inattentive subtypes, with emerging evidence that gene–gene interactions may further modulate individual risk [22,24]. However, the limited effect sizes observed in risk-associated genes are mirrored in pharmacogenetic studies, where genetic markers have likewise shown insufficient predictive value for guiding treatment.
Pharmacogenetic-guided treatment for attention-deficit/hyperactivity disorder has not demonstrated clinically meaningful improvement in patient outcomes compared to standard care, based on the latest clinical trials and meta-analyses [26]. The most robust evidence concerns CYP2D6 for atomoxetine, where poor metabolizers experience higher drug exposure and increased risk of adverse effects, but genotype-guided dosing has not shown clear superiority in symptom control or overall clinical benefit [27]. For methylphenidate, meta-analyses and clinical trials have identified associations between response and variants in DAT1 (SLC6A3), DRD4, DRD5, SLC6A2, and ADRA2A, with some variants (e.g., SLC6A2 rs28386840, ADRA2A MspI) modestly predicting improved response or symptom domain-specific benefit, but effect sizes are small and not sufficient for clinical utility [8,28,29].
No gene among DAT1, DRD4, DRD5, SLC6A2, CYP2D6, ADRA2A, SLC6A4, or HTR2A has shown consistent, reproducible, and clinically actionable predictive value for ADHD medication selection, dosing, or outcome improvement in large, well-controlled studies [30,31]. The proportion of variance in treatment response explained by these genetic markers is low (typically <20%), and current pharmacogenetic tests do not outperform standard titration and clinical monitoring [27,30]. Thus, routine pharmacogenetic testing for ADHD is not supported by current evidence, does not improve outcomes over standard care, and its clinical relevance remains limited, a position consistently reflected in current clinical guidelines. While genetic variation in pharmacokinetic and pharmacodynamic genes (such as CYP2D6 for atomoxetine and CES1 for methylphenidate) can influence drug exposure and response, the evidence base does not support routine pharmacogenetic testing to guide ADHD medication selection or dosing in clinical practice [10,26,32,33].
1.4. Clinical Guidelines, Translational Gap, and Rationale for Bibliometric Analysis
The American Academy of Pediatrics explicitly states that pharmacogenetic tools are not recommended for ADHD treatment due to inconsistent findings, insufficient effect sizes, and lack of demonstrated impact on clinical outcomes or cost-effectiveness [10]. Although certain gene variants (e.g., CYP2D6 poor metabolizers for atomoxetine) can result in higher drug exposure and increased risk of adverse effects, actionable recommendations are limited to specific scenarios and are not broadly applicable to most patients [26,33]. For methylphenidate, candidate gene studies have identified associations with response variability, but these findings have not translated into clinically useful algorithms or guidelines [30,32,34].
Current evidence does not support the clinical utility of incorporating pharmacogenetic testing (whether based on polygenic risk scores (PRSs), single-gene markers, or multi-gene panels) into treatment decision-making for attention-deficit/hyperactivity disorder (ADHD). To date, no large-scale, prospective clinical trials or meta-analyses have demonstrated that such testing improves clinical outcomes compared with standard care [32,33,35]. Recent meta-analyses and systematic reviews consistently show that PRS explain only a small proportion of variance in ADHD diagnosis and symptom severity (typically 3–4%) and, although they correlate with certain clinical phenotypes, they do not reliably predict individual medication response or guide treatment selection in a clinically actionable manner [35,36,37,38]. Similarly, while candidate-gene and genome-wide studies have identified associations between specific genetic variants and ADHD medication response, these findings have not translated into improved treatment outcomes when used to inform clinical decision-making [32,36].
Despite extensive genetic research, only a limited subset of findings has translated into clinically actionable pharmacogenetic recommendations for ADHD treatment. Clinical guidelines and consensus statements from major professional societies, including the Clinical Pharmacogenetics Implementation Consortium (CPIC), likewise do not endorse the routine use of pharmacogenetic testing for ADHD medication selection or dosing due to insufficient evidence of clinical benefit [33,36]. The CPIC and Dutch Pharmacogenetics Working Group (DPWG) provide evidence-based, clinically actionable recommendations only for gene–drug pairs where genetic variation meaningfully alters pharmacokinetics or pharmacodynamics. Within ADHD pharmacotherapy, the most robust and actionable evidence pertains to atomoxetine, which is primarily metabolized by CYP2D6. Individuals who are CYP2D6 poor metabolizers exhibit markedly elevated plasma concentrations, prolonged drug exposure, and heightened risk of adverse effects; as a result, both CPIC and DPWG provide genotype-based dosing recommendations for atomoxetine [26,33]. In contrast, although emerging data suggests that CES1 variants may influence methylphenidate metabolism, current evidence remains insufficient to support guideline-level recommendations or routine clinical implementation [33].
From a clinical perspective, this bibliometric analysis aims to support psychiatrists and other clinicians by clarifying the current translational limits of pharmacogenetics in ADHD management. By systematically mapping the trends that have dominated the literature over the past two decades, and identifying areas that remain underexplored, this study enables clinicians to distinguish between well-established pharmacogenetic hypotheses and those lacking robust or reproducible evidence. Such an overview promotes realistic clinical expectations, discourages premature reliance on non-actionable genetic tests, and underscores the continued superiority of standard clinical titration and monitoring over genotype-guided decision-making. In light of recent advances and the absence of actualized comprehensive meta-analyses in this field [39,40,41], the present study undertakes a bibliometric analysis of literature published between 2005 and 2025 to characterize publication and citation trends, identify the most influential authors, sources, and institutions, and map the intellectual and thematic structure of pharmacogenetic research in ADHD. By synthesizing the current state of knowledge and delineating research gaps and emerging areas, this analysis provides critical guidance for future investigations and supports the responsible translation of pharmacogenetic findings into precision medicine strategies for ADHD management.
2. Materials and Methods
This study adopts a bibliometric methodology organized into five clearly defined stages, as illustrated in Figure 1: (i) identification of the most relevant literature, focusing on peer-reviewed scientific publications; (ii) systematic collection and organization of bibliometric data to ensure consistency and reliability; (iii) analysis and processing of the data using statistical and bibliometric techniques to detect trends, patterns, and intellectual connections within the field; (iv) synthesis and visualization of the results, converting the findings into graphical representations to facilitate interpretation; (v) integration of the main outcomes to draw conclusions and provide critical insights that guide future research directions.

Figure 1.
Methodology phases for this bibliometric review.
In the first phase, relevant documents were retrieved from the Scopus and Web of Science databases. These sources were chosen for their rigorous indexing criteria and broad multidisciplinary coverage, both of which are critical for ensuring the reliability and reproducibility of bibliometric analyses [42,43,44]. Data collection took place in July 2025. The timespan was restricted to publications from 2005 to 2025, and only documents published in English were considered. The search terms were applied to titles, abstracts, and keywords using the following database-specific queries: in Scopus, TITLE-ABS-KEY (“pharmacogenetic*” AND “ADHD”); and in Web of Science, TS = (“pharmacogenetic*” AND “ADHD”). Here, the asterisk functions as a wildcard to capture variations in the root term. No additional database-level filters related to population (e.g., humans), study design (e.g., clinical trials), or publication status were applied, in order to avoid unnecessary restrictions and preserve the integrity of the bibliometric mapping. The complete metadata of the selected records were exported, including titles, author names, institutional affiliations, abstracts, keywords, citation counts, and reference lists.
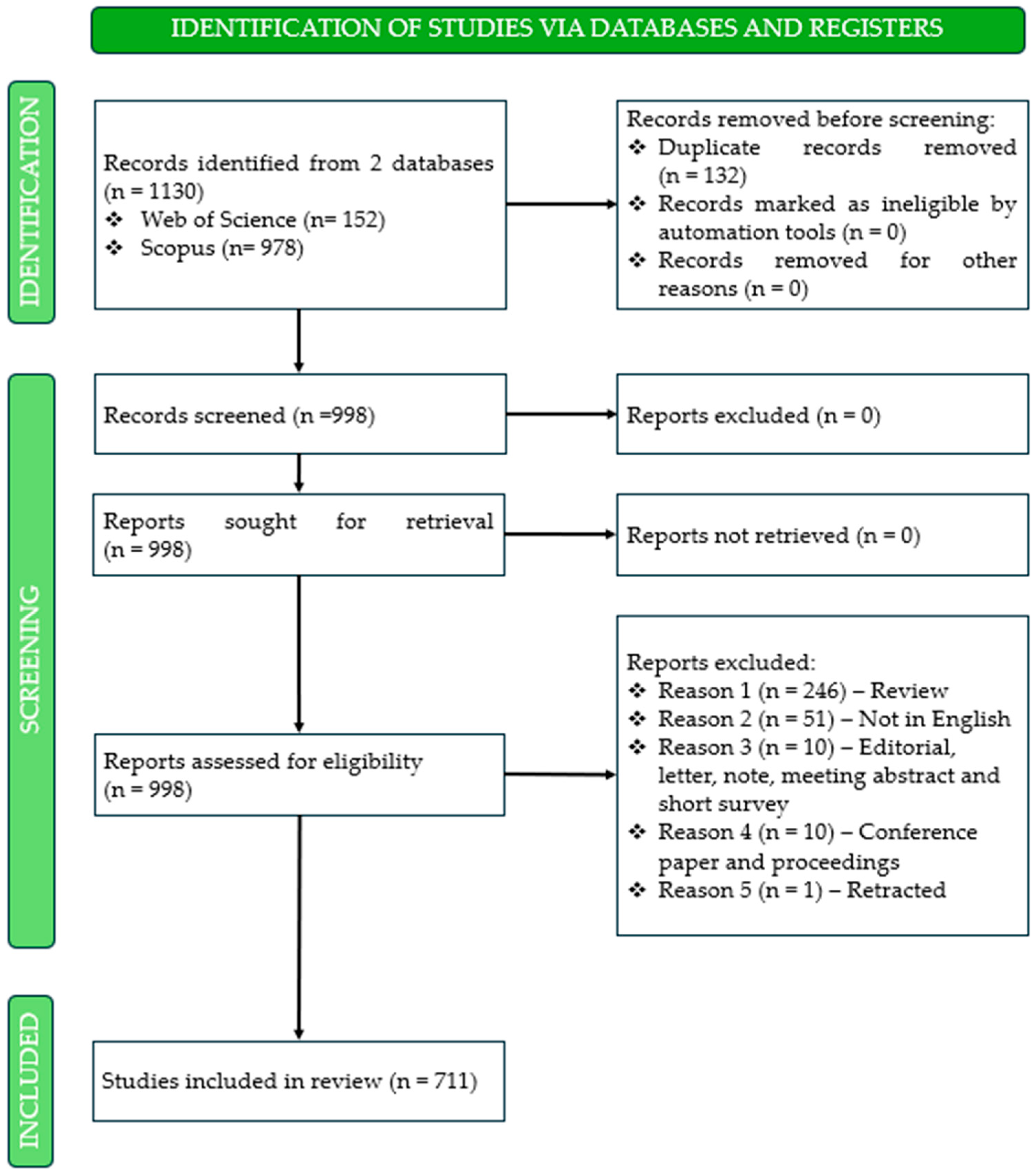
In the second phase, the PRISMA 2020 protocol was applied to ensure a rigorous selection and organization of research related to pharmacogenetic studies in ADHD, thereby preparing the dataset for subsequent analysis. The PRISMA 2020 protocol is a fundamental tool for bibliometric analysis, as it provides a structured and transparent guide for the elaboration of reviews [45]. As illustrated in Figure 2, the identification phase began with the retrieval of records from two databases, namely Web of Science (n = 152) and Scopus (n = 978), yielding a total of 1130 entries. Following the export and merging of datasets, 132 duplicate records were identified and removed, resulting in 998 unique entries for screening. During the screening phase, all records were evaluated to determine their eligibility. None were excluded at this stage, and all 998 reports were sought retrieval, with no records lost due to technical issues. Subsequently, the eligibility assessment was conducted, during which 318 reports were excluded based on predefined criteria: 246 review articles, 51 non-English publications, 10 editorials, notes, letters, meeting abstracts, or short surveys, 10 conference papers and proceedings, and 1 retracted publication. Here, only peer-reviewed scientific articles and books or book chapters were considered, ensuring the inclusion of high-quality sources for the subsequent bibliometric analysis. Upon completion of this comprehensive screening and eligibility assessment, a final set of 711 studies was retained as the core corpus for bibliometric analysis. Notably, no records were excluded due to automation-related ineligibility or retrieval errors, underscoring the rigor and reproducibility of the selection process.
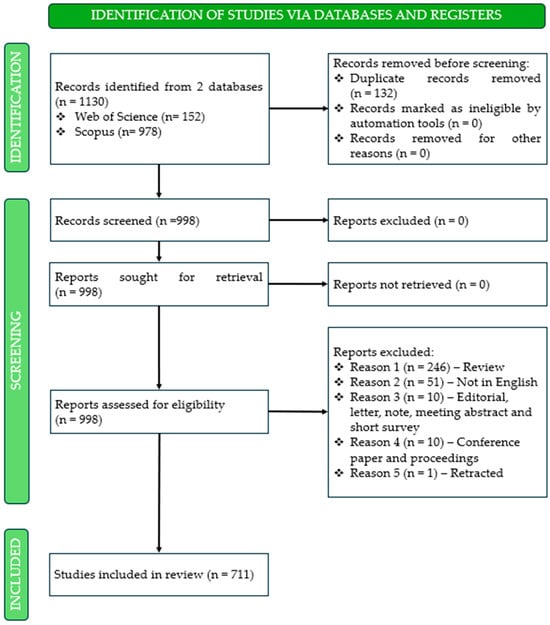
Figure 2.
PRISMA 2020 flow diagram illustrating the identification, screening, and inclusion of studies on pharmacogenetic research in attention-deficit/hyperactivity disorder.
In the third phase, bibliometric mapping was performed using the Bibliometrix R package (version 4.5.0), an open-source framework widely applied in scientometric and bibliometric research. Analyses were performed within the R statistical environment, and the Biblioshiny web interface was used to facilitate interactive data exploration and visualization [46]. To ensure data consistency and reproducibility, author names, institutional affiliations, and source titles were standardized using the built-in data cleaning and harmonization procedures of Bibliometrix, including automatic disambiguation and manual verification where necessary. Duplicate records were identified and removed prior to analysis, and all bibliometric parameters were applied consistently across datasets retrieved from Scopus and Web of Science. Core Bibliometrix functions (biblioAnalysis, summary, plot, networkPlot, thematicMap…) were applied to compute descriptive indicators, collaboration networks, co-occurrence structures, and thematic mappings. Network analyses were based on association strength normalization, and clustering was performed using standard algorithms implemented in Bibliometrix as a Louvain method for community detection.
The fourth phase was dedicated to the interpretation and integration of the bibliometric outputs, with an emphasis on exploring both the structural and thematic dimensions of the field. Co-word analysis, thematic clustering, and citation-based indicators (such as h-index, g-index, and total citation counts) were combined to provide a multidimensional perspective on the scientific landscape.
Finally, the fifth phase involved a critical assessment of the findings, highlighting their implications for both academic and practical domains. Particular attention was directed toward identifying research gaps, interdisciplinary synergies, and technological challenges linked to pharmacogenetic applications in ADHD, thereby outlining opportunities for future investigations.
3. Results
The results are organized into three parts: first, the general bibliometric characteristics of the dataset, including scope, evolution, and citation trends; second, the most productive and influential sources and authors; third, the institutional and geographical contributions that reflect the global distribution and impact of pharmacogenetic research in ADHD.
3.1. General Bibliometric Characteristics of the Dataset
Table 1 summarizes the main characteristics of the dataset analyzed in this bibliometric review in the last two decades, from 2005 to July 2025. A total of 711 documents were retrieved from Scopus and Web of Science, encompassing 350 different sources, including journals, books, and book chapters, that are the most impactful peer-reviewed documents. The dataset exhibits a low annual growth rate of 0.23%, suggesting a relatively stable but modest increase in the volume of research output over the years. The average age of the documents is 9.06 years, indicating that the field has a substantial foundation of mature literature, with a mix of both early and recent contributions.

Table 1.
Main characteristics of the dataset analyzed in this bibliometric review. ID: Keywords Plus (automatically generated index terms). DE: Author Keywords.
The average number of citations per document is 29.62, pointing to a moderate level of scholarly impact across the dataset. In terms of content, the dataset includes 4649 Keywords Plus (automatically generated index terms) and 1418 Author Keywords, revealing a rich diversity of thematic terms and research focus. This indicates a multidimensional and evolving field of study with varied conceptual approaches.
The dataset comprises contributions from 3578 unique authors. Of these, only 24 are associated with single-authored publications, while the vast majority are part of collaborative works. In total, there are 28 single-authored documents, reflecting a strong trend toward co-authorship in the field. On average, there are 7.37 co-authors per document, and 3.09% of the documents involve international collaborations. This relatively low percentage suggests that while collaboration is common, most research activity is conducted within national or institutional boundaries.
Regarding document types, the majority are journal articles (650), followed by book chapters (53), and a small number of books (6). This distribution highlights the predominance of peer-reviewed journal publications in disseminating research on pharmacogenetics in ADHD, while also reflecting contributions in academic books and edited volumes.
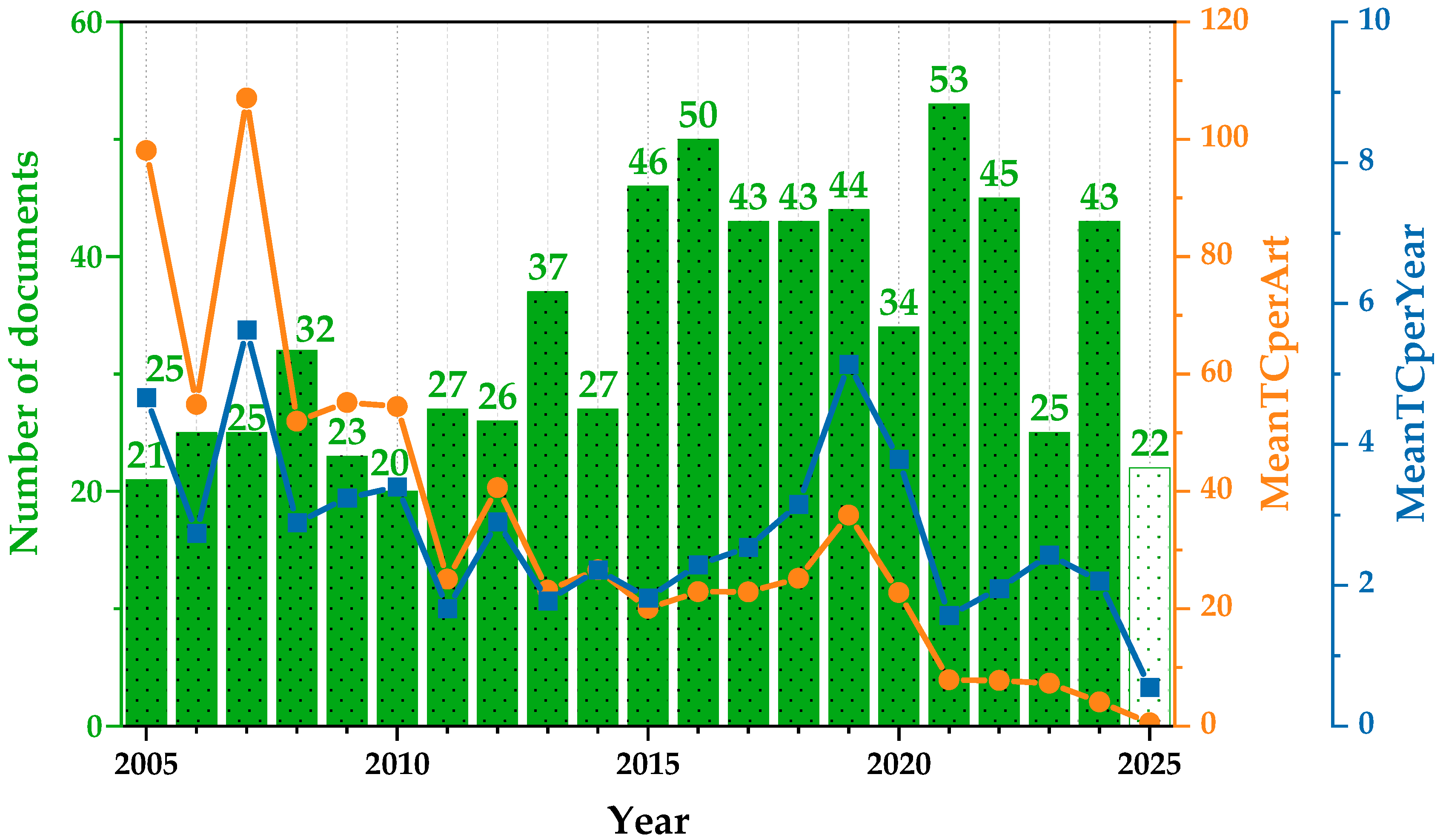
Figure 3 presents the annual evolution of research output and citation impact in the field in the last two decades. The green bars represent the number of documents published each year (N), while the orange and blue lines indicate the average number of total citations per article and the average number of citations per article per year, respectively.
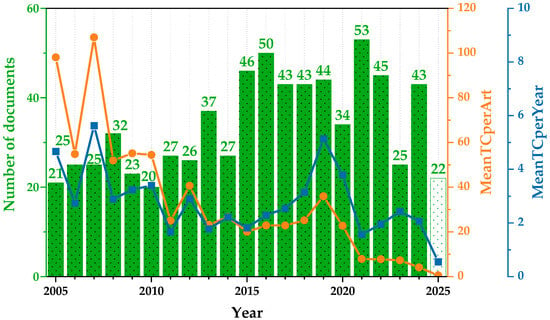
Figure 3.
Annual research production and citation growth: average TC per article and per year. Green bars represent the annual number of published documents. The orange line indicates the mean total citations per article (MeanTCperArt), and the blue line represents the mean total citations per article per year (MeanTCperYear). The white bar includes data available up to mid-2025.
The overall trend in scientific output reveals a progressive increase in the number of publications over the years, reaching its highest point in 2021, followed by a slight decline in subsequent years. However, this growth in volume has not been accompanied by a parallel increase in citation impact. The earliest publications within the dataset show notably high average citation counts, which likely correspond to foundational or widely cited studies that helped establish the field. Over time, the average number of citations per article has shown a gradual decline, with values stabilizing at moderate levels during the middle years and dropping more significantly in the most recent period. This pattern may be influenced by several factors, including the reduced time for newer publications to be cited, changes in research focus, or an overall diversification of topics within the field. Similarly, the average number of citations per article per year remains relatively consistent throughout much of the study period, with occasional fluctuations that may reflect the publication of particularly influential works. In the most recent years, however, a clear downward trend is evident, suggesting a decrease in short-term citation impact for new contributions.
3.2. Sources and Leading Authors
Table 2 presents the core journals identified through Bradford’s Law, which establishes that scientific knowledge in each field tends to be concentrated in a small group of highly productive journals (Zone 1), followed by progressively larger groups that contribute fewer publications. In this study, Zone 1 comprises ten journals that account for a substantial proportion of the research output. The Journal of Attention Disorders (23 articles), the American Journal of Medical Genetics, Part B: Neuropsychiatric Genetics (21 articles), and the Journal of Child and Adolescent Psychopharmacology (19 articles) stand out as the leading publication outlets. Other key contributors include Journal of Neural Transmission and Neuropsychopharmacology (15 articles each), and specialized outlets such as the Journal of the American Academy of Child and Adolescent Psychiatry (14 articles). Notably, multidisciplinary journals such as Scientific Reports and high-impact psychiatric journals like Molecular Psychiatry also appear in Zone 1, underscoring the field’s cross-disciplinary nature and its strong clinical orientation.

Table 2.
Core sources identified by Bradford’s Law (2005–2025).
Table 3 provides complementary insights by examining the scholarly impact of these journals using bibliometric indicators. The h-index reflects both productivity and citation impact; the g-index gives more weight to highly cited articles, offering a more comprehensive view of influence; the m-index normalizes the h-index by the number of years since the first publication, allowing comparisons across journals of different ages; and total citations capture the overall visibility of the research published in each journal. For example, while the Journal of Attention Disorders is the most productive outlet, it records a relatively modest citation performance (302 citations), suggesting that its articles, though numerous, are less frequently cited. In contrast, the Journal of the American Academy of Child and Adolescent Psychiatry accumulates 2385 citations with only 14 publications, highlighting its strong impact within the scientific community. Similarly, Neuropsychopharmacology combines high productivity with substantial citation counts (866), reflecting both output and influence.

Table 3.
Sources with the highest scholarly impact (2005–2025).
The analysis of Table 1 and Table 2 demonstrates that productivity and impact do not always coincide. On the one hand, a limited number of journals concentrate the majority of publications, shaping the thematic and disciplinary core of ADHD pharmacogenetics research. On the other hand, the journals exerting the greatest scholarly influence are those that combine specialization with broad recognition in psychiatry, neuroscience, and related biomedical domains. Thus, highly productive journals are key for mapping ongoing developments in the field, whereas high-impact journals offer greater visibility and influence for disseminating novel findings.
Table 4 lists the most influential authors in the field, evaluated through productivity and impact indicators. The results show that L. Rohde leads in terms of productivity, with 31 publications and the highest h-index (19), reflecting a consistent contribution to the field since 2007. J. Kim and S. Faraone also emerge as key figures, with 25 and 24 publications, respectively. Notably, Faraone registers the highest total citations (1162), underscoring the visibility and influence of his research despite producing fewer papers than Rohde. Similarly, M. Bellgrove and G. Polanczyk achieve strong citation counts (711 and 577, respectively), highlighting their relevance in shaping the thematic and methodological directions of the field. Another important observation is the geographical diversity reflected in the list of authors. Researchers from Brazil (e.g., Rohde, Hutz, Polanczyk, Genro, Grevet) appear prominently, suggesting the existence of a strong collaborative cluster in Latin America with significant international visibility. At the same time, contributions from other regions, particularly North America and Asia (e.g., Kim, Faraone, Hawi), emphasize the global dimension of this field.

Table 4.
Most influential authors (2005–2025).
3.3. Institutional and Geographical Contributions
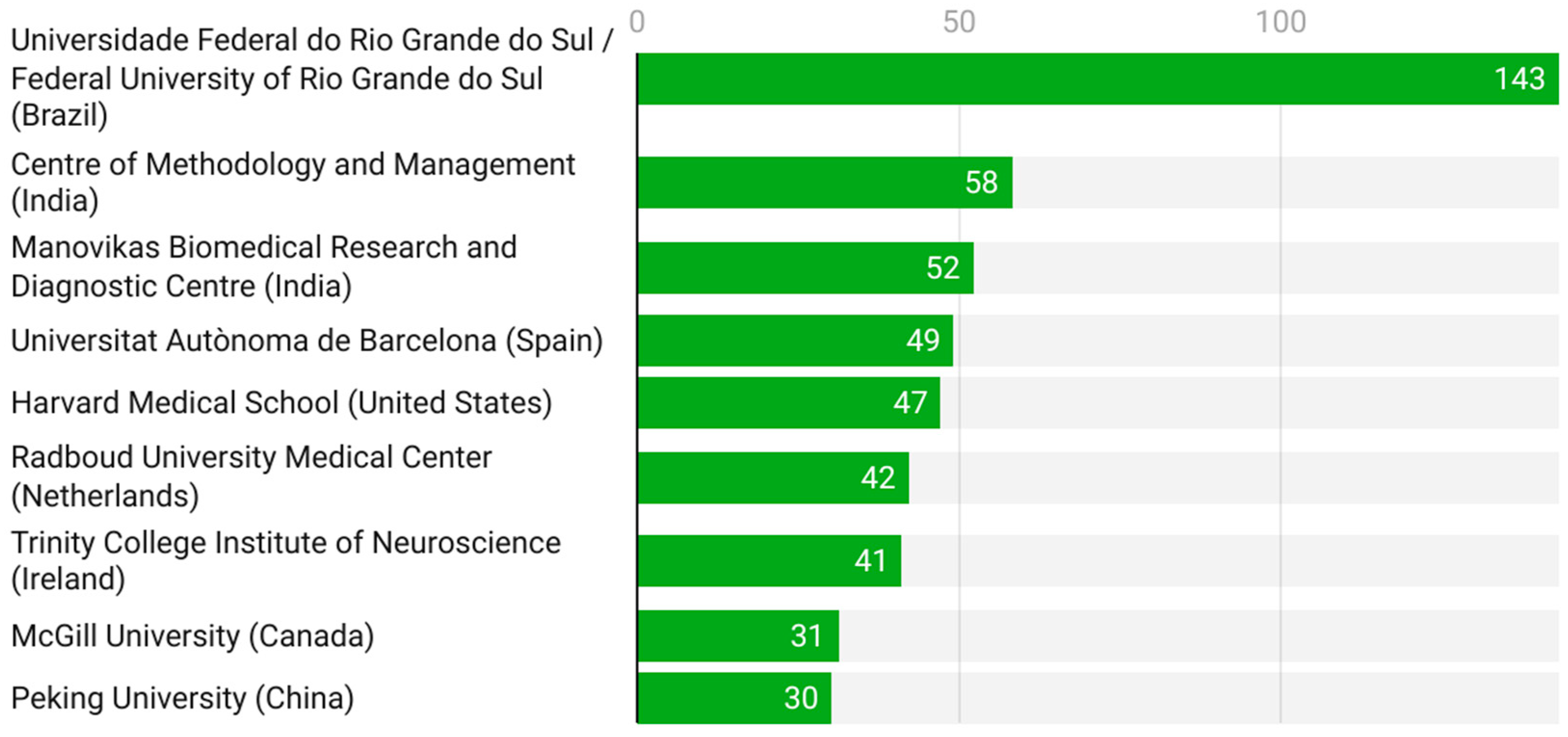
Figure 4 displays the most relevant institutional affiliations contributing to pharmacogenetic research in ADHD. The Universidade Federal do Rio Grande do Sul (Brazil) clearly dominates, with 143 publications, followed by institutions in India (Centre of Methodology and Management and Manovikas Biomedical Research and Diagnostic Centre) and Europe (Universitat Autònoma de Barcelona and Radboud University Medical Center). Prestigious institutions from North America (Harvard Medical School and McGill University) and Asia (Peking University) also rank among the top contributors. This distribution underscores the global and collaborative nature of ADHD pharmacogenetics, with strong research clusters in Brazil, India, and the United States.
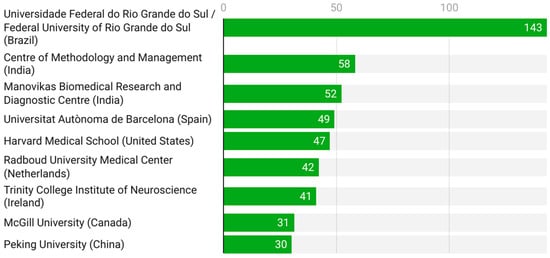
Figure 4.
Leading institutional affiliations (2005–2025).
Table 5 summarizes the distribution of publications by corresponding author’s country. The United States clearly leads with 176 publications (24.8%), followed by China (7.5%), Canada (5.2%), and Brazil (4.8%). European countries such as Germany, Spain, and the United Kingdom also contribute significantly, although with smaller percentages. The distinction between SCP (Single-Country Publications) and MCP (Multiple-Country Publications) provides insight into international collaboration. For instance, while the United States (169 SCP, 7 MCP) and China (53 SCP, 0 MCP) show a strong predominance of domestic research, Brazil demonstrates a more collaborative profile (29 SCP, 5 MCP), highlighting its role as a connector in international networks.

Table 5.
Distribution of publications by corresponding author’s country, including single-country and multi-country collaborations (2005–2025).
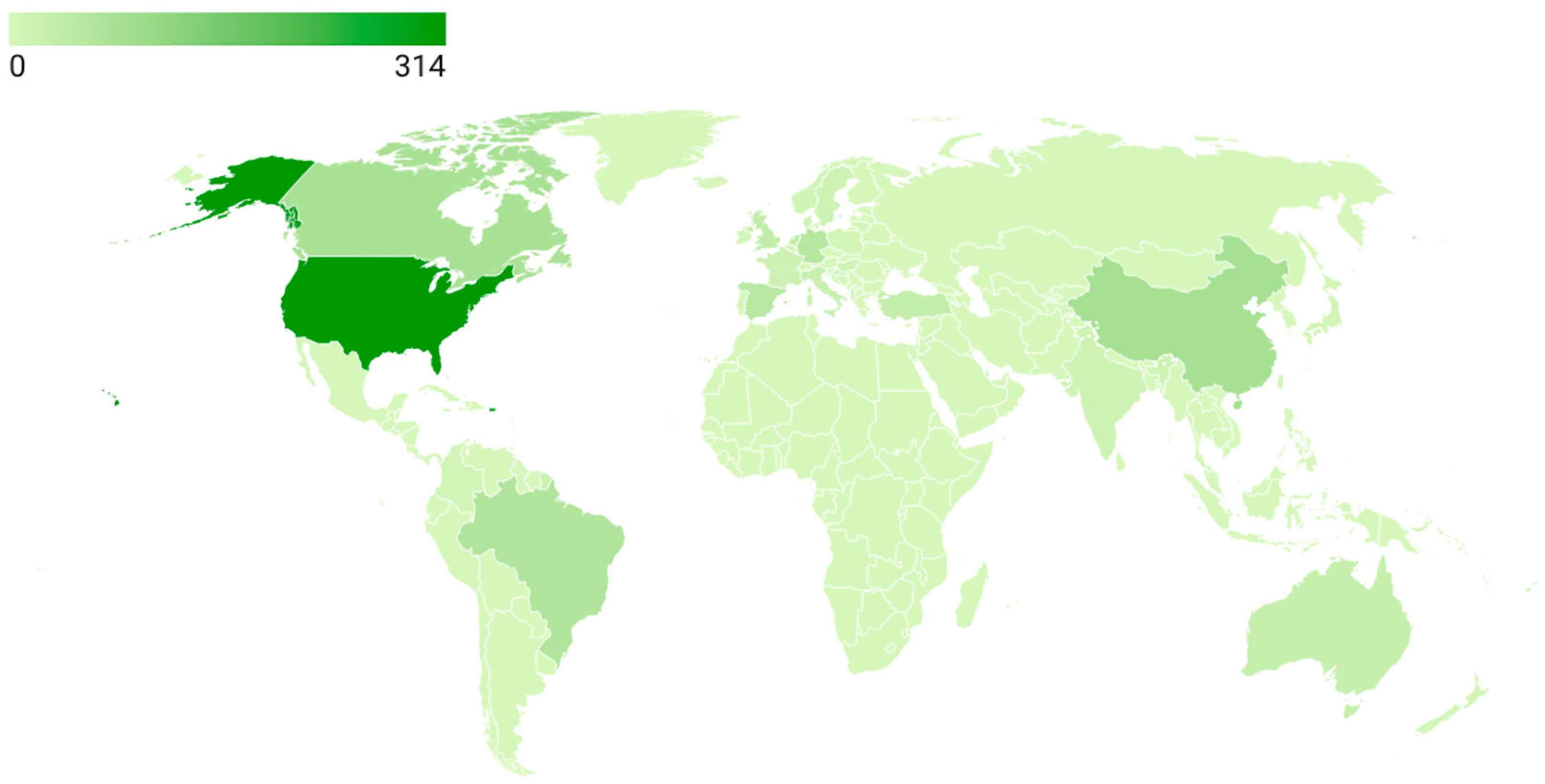
Figure 5 illustrates the global distribution of scientific production by country based on the number of documents included in the dataset. The United States clearly leads the field with 314 publications, accounting for the largest share of research output in ADHD pharmacogenetics. This is followed by China (78 documents), Canada (74), Brazil (65), Germany (56), and Spain (48), which together form a second tier of highly productive countries. A substantial contribution is also observed from several European nations, including the United Kingdom (36), Turkey (33), Netherlands (40), Italy (24), France (19), Sweden (16), and Denmark (15). Additional, though more moderate, contributions come from countries such as India (14), Hungary (12), Norway (11), Japan (8), New Zealand (7), and Poland (7), reflecting a broader but less concentrated global engagement. However, the strong presence of Canada, Germany, and China indicates that pharmacogenetic studies in ADHD have expanded beyond North America to Europe and Asia. Meanwhile, the contributions from Brazil and Spain reinforce the increasing participation of Latin America and Southern Europe, consolidating more global research.
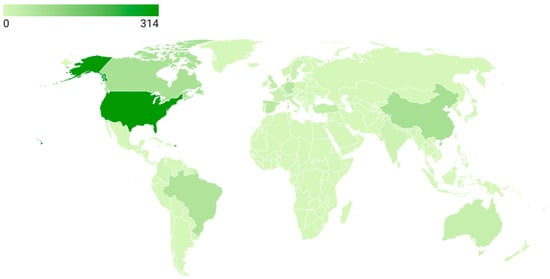
Figure 5.
Global scientific production (2005–2025).
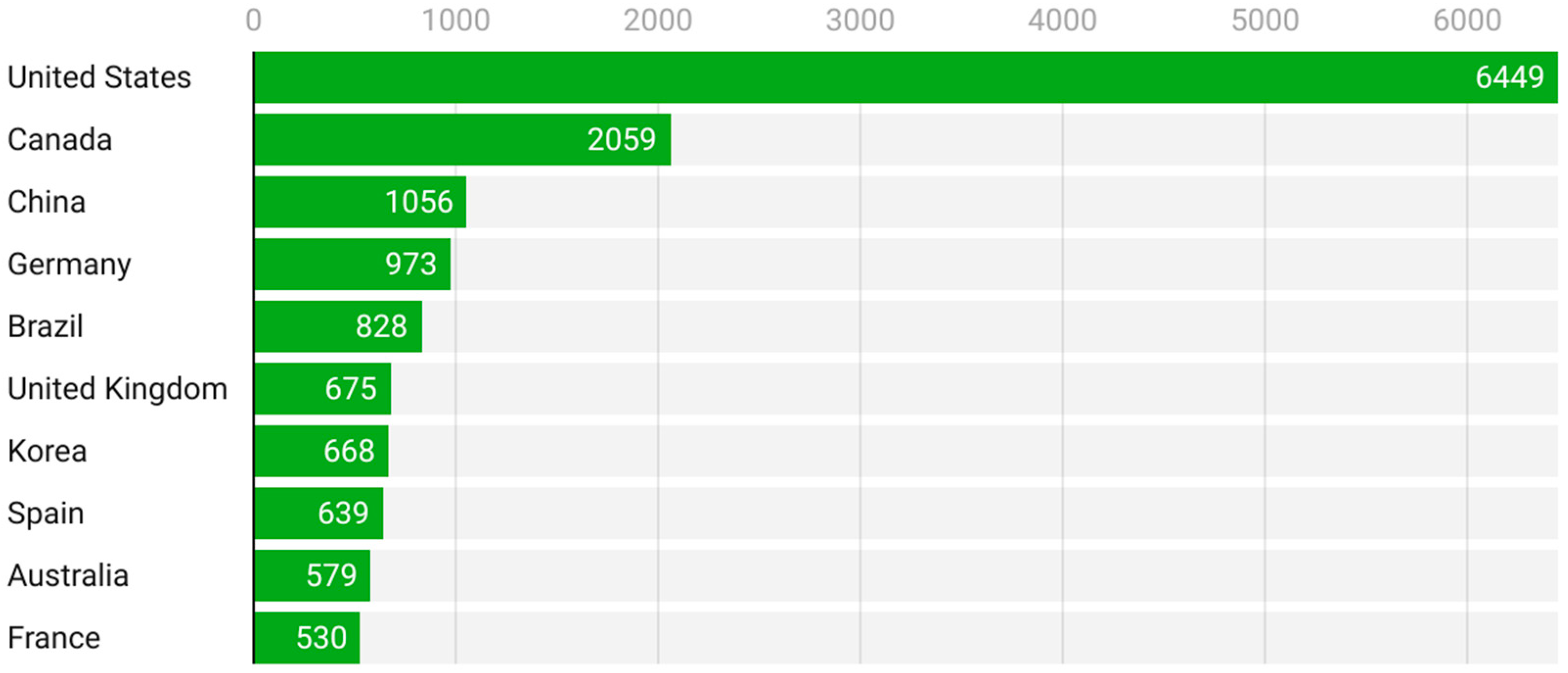
Figure 6 presents the most cited countries, offering insight into scholarly impact. The United States again dominates with 6449 citations, followed at a distance by Canada (2059), China (1056), Germany (973), and Brazil (828). Interestingly, although countries like Spain and France produce fewer publications, their citation counts indicate a relatively high impact per article, pointing to strong visibility and recognition of their contributions.
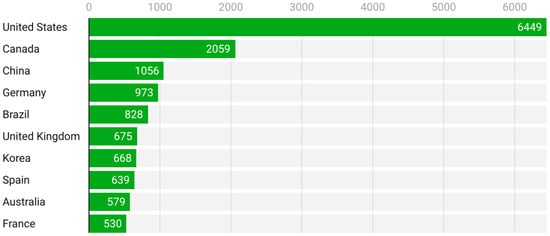
Figure 6.
Most cited countries (2005–2025).
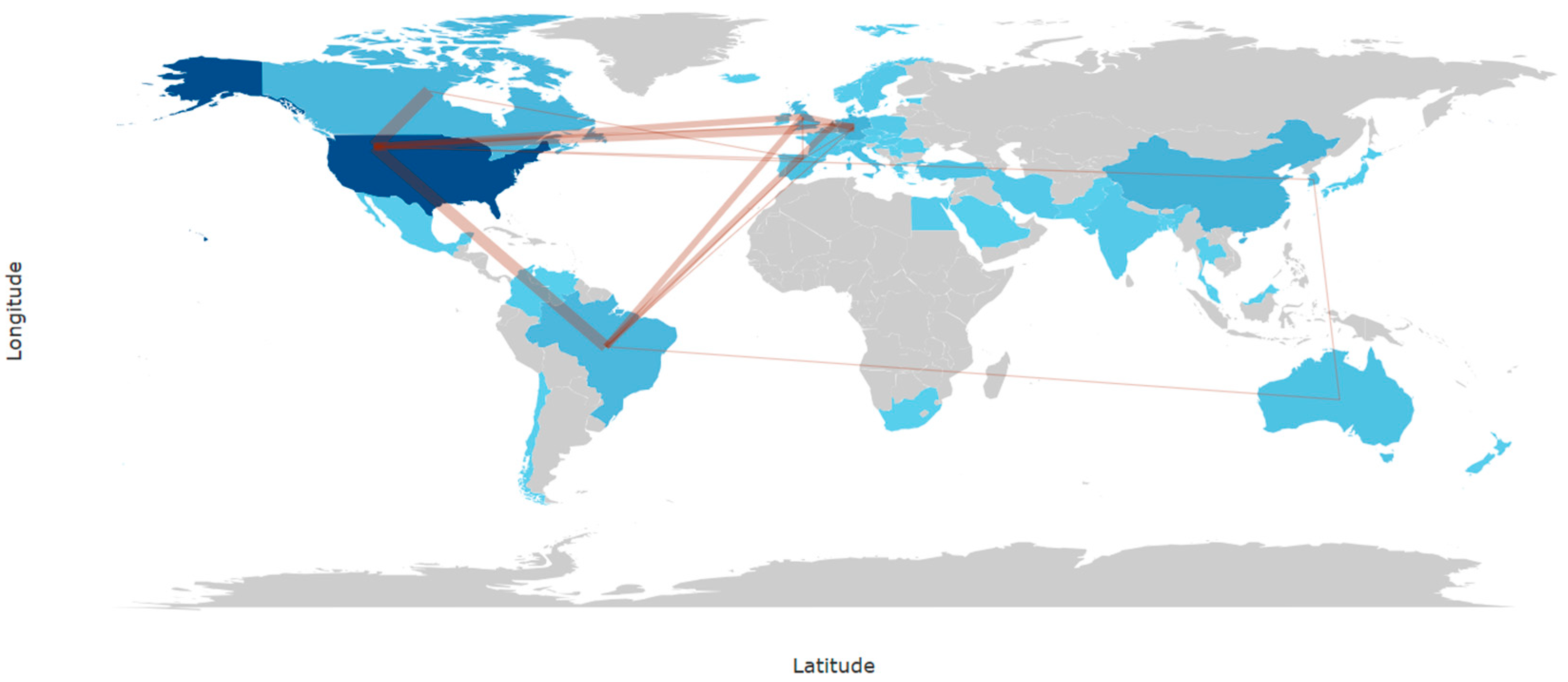
Figure 7 depicts the global pattern of international collaborations the field between 2005 and 2025. The collaboration network is highly centralized, with the United States clearly emerging as the main hub, maintaining the strongest and most numerous collaborative links with European countries, Canada, and selected regions of Latin America and Asia. These dense transatlantic connections highlight the coordinating role of North American institutions in multinational research efforts. In contrast, collaborations among countries outside North America and Western Europe are comparatively limited, with countries such as Brazil, China, and Australia participating mainly through partnerships mediated by the United States or major European collaborators rather than through autonomous regional networks. Despite the increasing geographical diversification of research output, international remains unevenly distributed and strongly centralized, reinforcing the importance of expanding balanced, multi-country collaborations to support large-scale and reproducible research in this field.
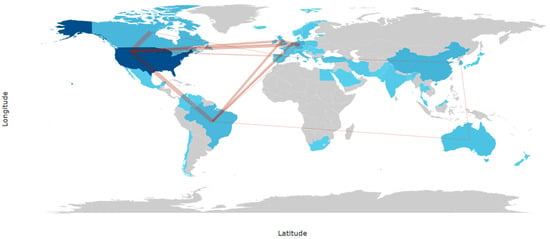
Figure 7.
Countries’ collaboration world map (2005–2025).
The study of affiliations and countries shows that research is concentrated in a small number of leading institutions and countries, while at the same time reflecting an increasing global scope. North America, particularly the United States, emerges as the main driver of scientific production and impact, supported by contributions from Canada and leading European countries such as Germany, Spain, and the United Kingdom. Brazil plays a distinctive role by combining high productivity with strong international collaboration, positioning Latin America as an active participant in the field. Asian countries, especially China and Korea, have also increased their presence, although their influence is still consolidating in terms of global impact. The geographical and institutional distribution highlights a research landscape characterized by North American leadership, growing European and Asian participation, and the notable emergence of Latin American institutions, which together contribute to shaping a diverse and internationally connected scientific community.
4. Discussion
4.1. Intellectual Foundations of ADHD Pharmacogenetics from the Most Cited Articles
Table 6 summarizes the most cited articles in pharmacogenetic research on ADHD. The top-ranked study by Pliszka et al. has accumulated 1409 citations, reflecting its status as a seminal contribution that continues to inform clinical and pharmacogenetic approaches to ADHD management. Wolraich et al. stand out with the highest citation rate per year (132.86), underscoring the rapid uptake and influence of their work. Other key studies, such as those by Greenhill et al. and Bouchard et al., show sustained relevance over time, while normalized citation indicators confirm their enduring impact beyond the initial years of publication. The list also highlights the multidisciplinary character of the field. Diamond and Shaw contributed developmental and neurobiological perspectives, Cortese examined comorbidities and risk factors, while Prehn-Kristensen provided a translational approach with notable annual impact (25 citations per year). This variety reflects how pharmacogenetics in ADHD draws on clinical, pharmacological, neurodevelopmental, and translational research traditions.

Table 6.
Most cited articles ranked by total citations and citation impact indicators.
The analysis of the ten most cited articles (Table 6) highlights the seminal contributions that have shaped pharmacogenetic research in ADHD, which converge into four major thematic domains: (i) clinical guidelines and pharmacological treatments; (ii) cognitive heterogeneity and subtypes; (iii) neurodevelopmental and genetic mechanisms; (iv) environmental and health-related influences.
The first domain is anchored in clinical guidelines and therapeutic evidence. The practice parameter by Pliszka et al. [47] and the updated recommendations of the American Academy of Pediatrics by Wolraich et al. [10] are among the most highly cited contributions, providing standardized frameworks for the diagnosis and treatment of ADHD. Both documents are among the most highly cited works in the dataset, reflecting their strong influence on the clinical and conceptual framing of ADHD treatment within pharmacogenetic literature. These guidelines also integrate safety considerations, including cardiovascular risk and growth monitoring, reflecting the field’s growing attention to long-term outcomes. Complementary clinical trials by Greenhill et al. [48] in preschool children and Spencer et al. [51] in adults demonstrated the efficacy and tolerability of methylphenidate across the lifespan. Importantly, both trials confirmed that treatment response depends critically on dosage optimization and careful monitoring of adverse events, reinforcing the principle of individualized pharmacotherapy.
A second influential domain relates to cognitive heterogeneity, where ADHD is conceptualized as a multidimensional syndrome rather than a unitary disorder. Diamond [50] proposed that the inattentive subtype is distinguished by deficits in working memory and hypoarousal, in contrast to the impulsivity-driven deficits of the combined subtype. This conceptual distinction has had a lasting impact on the field, encouraging a more nuanced clinical characterization of ADHD. Similarly, MacDonald et al. [52] introduced the concept of intra-individual variability in performance as a potential cognitive marker of ADHD, showing that attentional fluctuations reflect unstable dopaminergic regulation in prefrontal networks. The recognition of such cognitive signatures supports the development of endophenotypes that may ultimately bridge genetic vulnerability and clinical manifestation, opening opportunities for more personalized diagnostic and therapeutic approaches.
The third domain highlights the neurodevelopmental and genetic underpinnings of ADHD, particularly through neuroimaging-genetic integration. Shaw et al. [54] demonstrated that children carrying the DRD4-7R allele exhibit thinner cortical structures in prefrontal and parietal regions, but with delayed normalization during adolescence. These findings reinforced the “developmental delay” hypothesis of ADHD and suggested that specific genotypes may influence not only risk but also long-term prognosis. More broadly, this line of work underscores that ADHD should be understood as a dynamic disorder of brain maturation, where neurobiological differences are most pronounced in childhood and tend to diminish over time.
Finally, the fourth domain encompasses environmental and systemic health-related influences. Bouchard et al. [49] provided epidemiological evidence linking pesticide exposure to increased ADHD risk, raising concerns about modifiable environmental contributors. Cortese et al. [53] documented a robust association between ADHD and obesity, later confirmed by meta-analyses, suggesting shared mechanisms of impulsivity, reward dysfunction, and metabolic dysregulation. Prehn-Kristensen et al. [55] introduced the novel perspective of gut–brain interactions, identifying alterations in microbial composition in children with ADHD. Thus, ADHD is increasingly conceptualized as a brain-based disorder and as a condition influenced by systemic and environmental factors, which broadens the scope of pharmacogenetic research to include public health and lifestyle dimensions.
In summary, the most influential studies show that impact in ADHD pharmacogenetics stems from novelty, clinical relevance, and translational value rather than sheer volume. Clinical guidelines and trials shape practice, cognitive and neurodevelopmental research refine theoretical models, and environmental studies expand the etiological perspective. Together, these patterns illustrate how citation impact in ADHD pharmacogenetics is driven by clinical relevance, conceptual innovation, and translational framing within the literature.
4.2. Keywords Analysis
The keyword analysis offers information about the thematic structure of pharmacogenetic research in ADHD and its evolution over the last two decades. By examining the most recurrent terms and their interconnections, it is possible to identify the central topics that have guided research and the conceptual bridges between clinical and genetic dimensions of the field.
Table 7 summarizes the most frequently occurring keywords from 2005 to 2025. Population-related descriptors such as male (408 occurrences), female (358), and child (302) are highly prevalent, indicating that research has primarily been conducted in pediatric cohorts, with a marked interest in sex-related differences. The frequent appearance of adolescent and humans further reflects the predominance of developmental and human subject studies over animal models. Diagnostic terms such as attention deficit disorder (379), attention deficit disorder with hyperactivity (375), and ADHD (269) highlight the central role of clinical categorization in structuring research questions. The presence of major clinical study (248) and controlled study (297) also points to a strong emphasis on clinical trials and observational human studies, underscoring the translational orientation of the field.

Table 7.
Most frequently occurring keywords (2005–2025).
From a therapeutic perspective, methylphenidate (225 occurrences) and atomoxetine (77) stand out as the most prominent pharmacological agents. This reflects the continued reliance on methylphenidate as the gold-standard stimulant for ADHD treatment, while atomoxetine represents the most extensively studied non-stimulant alternative. Additional pharmacological descriptors such as central nervous system stimulants (103) and central stimulant agent (85) further consolidate the pharmacogenetic emphasis on treatment efficacy and variability of response.
At the molecular level, highly recurrent terms such as genetics (225), genotype (180), single nucleotide polymorphism (112), allele (88), and dopamine transporter (85) reveal the importance of genetic association studies. The emphasis on polymorphisms in dopamine-related genes reflects a persistent interest in the catecholaminergic system, given its central role in ADHD pathophysiology and stimulant drug mechanisms. The prominence of comorbidity (76) also suggests that many pharmacogenetic studies integrate broader psychiatric or medical contexts, examining ADHD alongside conditions such as anxiety, mood disorders, or obesity.
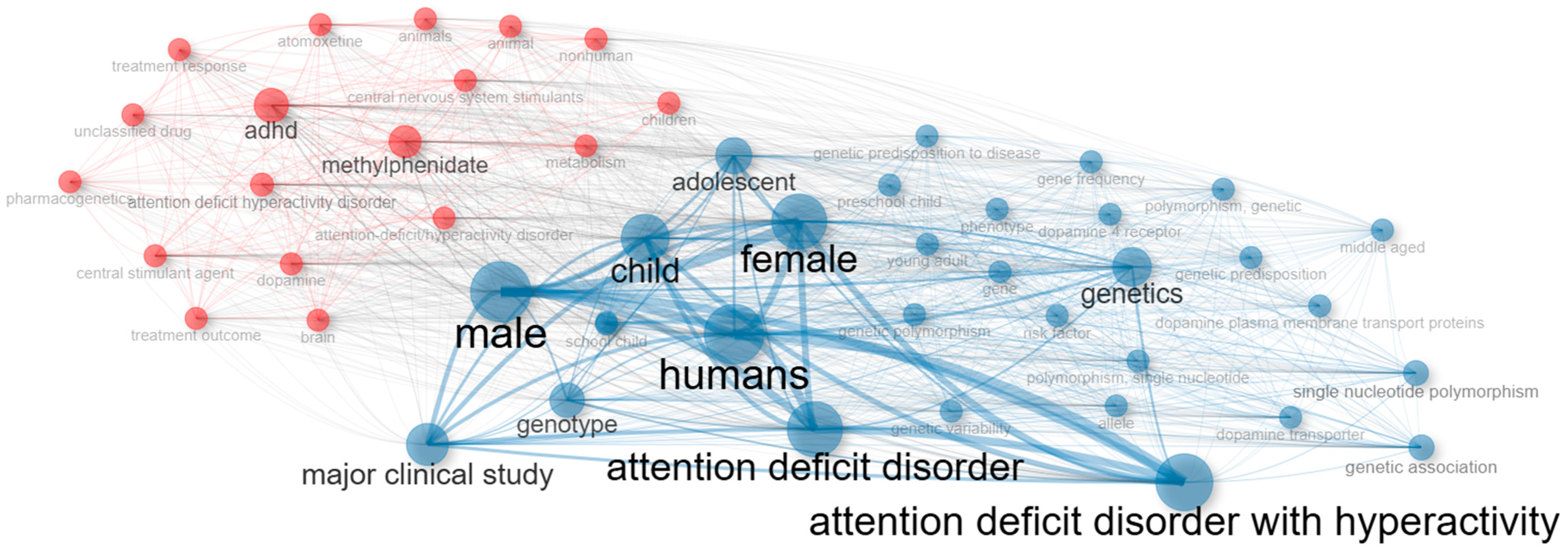
Figure 8 visualizes the co-occurrence network and clearly reveals the existence of two major clusters. The red cluster groups terms associated with the clinical–therapeutic dimension, such as ADHD, methylphenidate, stimulants, and drug therapy, often linked to descriptors of study populations (child, male, female). This cluster represents the translational clinical research domain, where pharmacological treatments are evaluated in relation to demographic variables and diagnostic categories. In contrast, the blue cluster captures the genetic–molecular dimension, dominated by terms such as genetics, genotype, single nucleotide polymorphism, and dopamine receptor. These keywords reflect the candidate gene approach that has historically dominated ADHD pharmacogenetics, with a strong emphasis on dopaminergic mechanisms. The nodes at the interface, including child, humans, and disorder, play a bridging role, suggesting that clinical samples are frequently used to explore genetic variability, thereby connecting therapeutic outcomes with underlying molecular mechanisms.
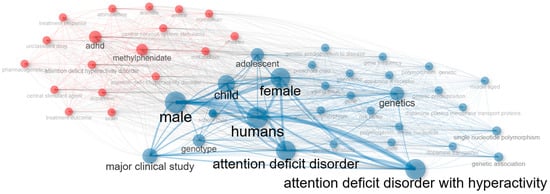
Figure 8.
Co-occurrence network of keywords showing clinical–therapeutic (red) and genetic–molecular (blue) clusters.
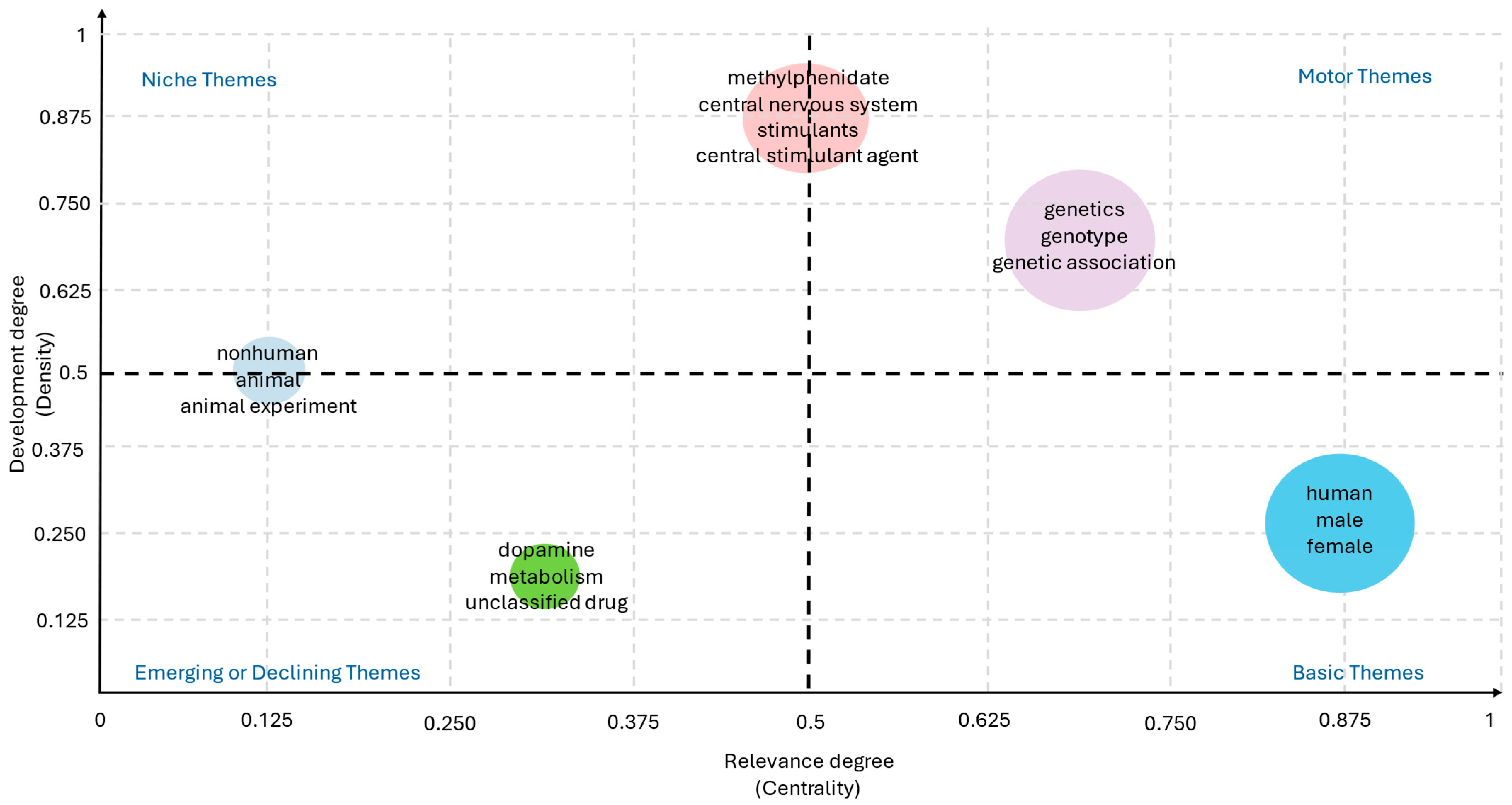
Figure 9 highlights the relative weight and interconnections of the most relevant keywords. In bibliometric co-occurrence mapping, each node represents a keyword, with its size reflecting the frequency of occurrence, while the thickness of the edges indicates the strength of co-occurrence between pairs of terms. Beyond frequency, the positioning of terms in the thematic map allows their classification according to two dimensions: centrality, or the degree to which a theme connects with other themes, and density, or the internal cohesion of a theme cluster.
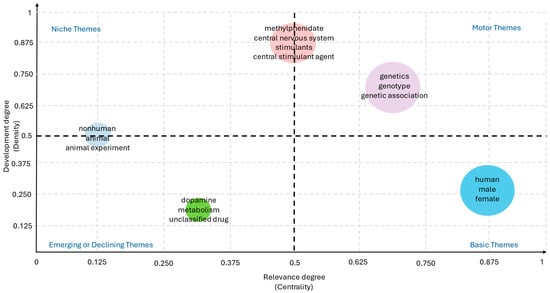
Figure 9.
Strategic mapping of keywords by centrality and density, highlighting motor themes and thematic structure of the field (2005–2025).
Motor themes, located in the upper-right quadrant of the map, are characterized by high centrality (≈0.65–0.75) and high density (≈0.60–0.70), indicating that they are well developed and strongly connected to the rest of the network. In ADHD pharmacogenetics, terms such as ADHD, methylphenidate, and genetics clearly function as motor themes, consolidating the conceptual backbone of the field. Indeed, the majority of pharmacogenetic studies to date have focused on the genetic underpinnings of ADHD and the clinical response to methylphenidate, underscoring the pivotal role of these terms in structuring the literature [56].
By contrast, basic or transversal themes, located in the lower-right quadrant, exhibit very high centrality (≈0.85–0.90) but low density (≈0.20–0.30). These themes are widely used across studies but do not form specialized research niches. Demographic descriptors such as male, female, and child fall into this category, ranking among the most frequent keywords in ADHD-related publication. Their prevalence reflects the fact that ADHD pharmacogenetic research is predominantly conducted in pediatric populations, where sex-based differences and developmental trajectories are critical variables in study design [57,58]. Moreover, the recurrent use of these terms underscores the enduring emphasis on early-life manifestations of ADHD and the need to expand analyses toward longitudinal and lifespan perspectives. Incorporating broader demographic diversity—including adult cohorts, cross-cultural populations, and sex-specific pharmacogenetic pathways—could enhance the translational relevance of future studies, particularly considering growing evidence that age, sex, and developmental stage may modulate both genetic associations and drug efficacy [59,60].
Niche themes, positioned in the upper-left quadrant, are defined by high density (≈0.50–0.85) and low to moderate centrality (≈0.10–0.50). These themes represent well-developed research areas with strong internal cohesion but limited integration into the broader literature. In this field, dopamine transporter and single nucleotide polymorphism exemplify niche themes, reflecting focused investigations on specific molecular targets. The prominence of the dopamine transporter (DAT1/SLC6A3) reflects its central role in the neurobiology of ADHD, as variations in this gene have been consistently associated with differences in dopamine reuptake efficiency and, consequently, with variability in stimulant response [61,62,63]. Similarly, studies centered on single nucleotide polymorphisms (SNPs) have provided valuable insights into the genetic architecture underlying ADHD pharmacogenetics, identifying specific variants in dopaminergic, serotonergic, and noradrenergic pathways that may predict treatment outcomes [64,65,66,67]. However, the classification of these topics as niche themes indicates that, while internally cohesive and methodologically robust, such lines of research remain relatively isolated from the broader clinical and translational discourse of the field. This partial disconnection suggests that molecular findings have not yet been fully integrated into clinical practice or multidisciplinary models, limiting their impact on the development of personalized therapeutic strategies.
Finally, emerging or declining themes, located in the lower-left quadrant, are characterized by low centrality (≈0.30–0.40) and low density (≈0.10–0.20). Terms such as atomoxetine and comorbidity illustrate this category. Their peripheral position suggests either the early stages of development—topics that may gain relevance in the near future—or declining prominence due to a shift in research focus. In the case of atomoxetine, the relatively limited number of pharmacogenetic studies compared with methylphenidate reflects its secondary role in clinical practice, despite its importance as a non-stimulant alternative targeting the noradrenergic system [68]. Research investigating genetic predictors of atomoxetine response, including polymorphisms in CYP2D6 and other metabolic pathways, has shown promising but heterogeneous results, which may explain its current marginalization in the thematic map [69,70]. Similarly, comorbidity—a highly relevant clinical dimension given the frequent overlap of ADHD with disorders such as anxiety, depression, and substance use—remains underexplored from a pharmacogenetic perspective [71,72].
In conclusion, the keyword analysis reflects a research field that has reached a degree of conceptual consolidation while still leaving ample room for future development. The strong interplay between clinical and genetic dimensions highlights the translational orientation of ADHD pharmacogenetics, yet the presence of peripheral and underdeveloped themes suggests that important questions remain open. Moving forward, the field is likely to benefit from expanding beyond its current focus to embrace more diverse pharmacological targets, broader genomic and multi-omics approaches, and greater consideration of developmental, demographic, and comorbidity factors. Such directions would deepen the understanding of ADHD’s biological underpinnings while simultaneously accelerating the transition toward precision medicine frameworks, where genetic insights are fully integrated into individualized treatment strategies.
4.3. Methodological and Clinical Challenges
Despite notable progress, pharmacogenetic research in ADHD continues to face significant methodological and clinical challenges that limit the translation of findings into routine practice. A major barrier lies in the predominance of small and heterogeneous samples, which restricts statistical power and complicates the replication of results across independent cohorts. Given the multifactorial etiology of ADHD, single-gene association studies often yield inconsistent findings, underscoring the need for larger, multi-center investigations and the adoption of genome-wide or multi-omics approaches that can capture the polygenic and dynamic nature of the disorder [73,74].
Clinical heterogeneity also poses a considerable challenge. ADHD encompasses diverse presentations that vary by age, sex, developmental trajectory, and comorbidities, all of which may modulate genetic associations and treatment responses. Current pharmacogenetic studies remain heavily focused on pediatric populations, with limited inclusion of adolescents, adults, and elderly patients, thereby narrowing the scope of representation within the published literature [75]. Furthermore, comorbid conditions—such as anxiety, mood disorders, and substance use—are often underrepresented, despite their potential to influence both pharmacodynamics and pharmacokinetics [76]. Addressing these gaps requires more inclusive study designs that reflect the complexity of real-world clinical practice.
Another limitation concerns the lack of standardized phenotyping and outcome measures. Variability in diagnostic criteria, cognitive assessments, and treatment endpoints hampers comparability across studies and slows the accumulation of robust evidence. Harmonization of protocols, along with the integration of neuroimaging, cognitive, and biomarker-based endpoints, could help refine the identification of pharmacogenetic predictors and strengthen their interpretability and comparability across studies [77].
Finally, there are translational hurdles to overcome. Although several candidate polymorphisms and pathways are recurrent in the literature, bibliometric mapping shows limited progression toward widespread guideline integration. Bridging this gap will require not only methodological improvements but also stronger collaborations between clinicians, geneticists, and bioinformaticians. Such interdisciplinary approaches are crucial for moving beyond proof-of-concept studies toward precision medicine frameworks where pharmacogenetic insights are systematically incorporated into individualized treatment strategies.
5. Conclusions
This bibliometric review synthesized 711 studies published between 2005 and 2025, retrieved from Scopus and Web of Science, representing the most comprehensive mapping of pharmacogenetic research in ADHD to date. The field is characterized by conceptual consolidation rather than rapid expansion, as reflected by a modest growth rate but a consistently high citation impact, indicating that scientific influence is driven by a relatively small number of foundational and highly cited studies.
In terms of scholarly output, a limited number of journals, authors and countries concentrate most of the productivity and impact. The Journal of Attention Disorders and the American Journal of Medical Genetics, Part B stand out as the most productive sources, while highly cited contributions often appear in Neuropsychopharmacology and the Journal of the American Academy of Child and Adolescent Psychiatry. Similarly, research groups led by L. Rohde (Brazil), J. Kim (Korea), and S. Faraone (USA) dominate authorship, with Brazil, North America, and Europe consolidating themselves as central hubs.
From a thematic perspective, the field is structured around four interconnected domains: (i) clinical guidelines and pharmacological treatments, where stimulant medications (notably methylphenidate) remain the central focus; (ii) cognitive heterogeneity and subtypes, advancing understanding of ADHD as a multidimensional syndrome; (iii) neurodevelopmental and genetic mechanisms, emphasizing candidate-gene studies of dopaminergic pathways and single nucleotide polymorphisms (SNPs); (iv) environmental and health-related influences, including pesticide exposure, obesity, and microbiome alterations. Together, these domains reflect the multifactorial nature of ADHD and the translational orientation of pharmacogenetic research.
Beyond the bibliometric structure, the findings underscore the clinical and translational significance of ADHD pharmacogenetics. Research has consistently centered on stimulant medications, particularly methylphenidate, and their genetic underpinnings, highlighting both efficacy and variability in treatment response. At the same time, genetic association studies of dopamine-related genes and SNPs provide mechanistic insights into ADHD pathophysiology, while newer explorations—such as atomoxetine pharmacogenetics, comorbidity analyses, and microbiome influences—point to the diversification of approaches.
Finally, thematic evolution reveals the coexistence of well-established core topics and underdeveloped areas, highlighting both maturity and persistent gaps in the field. Key limitations include the predominance of pediatric populations, limited representation of adults and comorbidities, and continued reliance on single-gene approaches. Addressing these gaps through multi-omic strategies, more diverse study populations, and stronger international collaboration will be critical for improving the clinical translation of ADHD pharmacogenetics.
Author Contributions
Conceptualization, A.C., M.S.-G. and Z.V.; methodology, A.C. and A.d.B.; software, A.C. and A.d.B.; formal analysis, A.C.; investigation, A.C. and A.d.B.; resources, A.d.B.; data curation, A.C.; writing—original draft preparation, A.C. and A.d.B.; writing—review and editing, M.S.-G. and Z.V.; supervision, A.C., M.S.-G. and Z.V. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding authors.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders; American Psychiatric Association: Washington, DC, USA, 2013. [Google Scholar] [CrossRef]
- Song, P.; Zha, M.; Yang, Q.; Zhang, Y.; Li, X.; Rudan, I. The Prevalence of Adult Attention-Deficit Hyperactivity Disorder: A Global Systematic Review and Meta-Analysis. J. Glob. Health 2021, 11, 04009. [Google Scholar] [CrossRef]
- Simon, V.; Czobor, P.; Bálint, S.; Mészáros, Á.; Bitter, I. Prevalence and Correlates of Adult Attention-Deficit Hyperactivity Disorder: Meta-Analysis. Br. J. Psychiatry 2009, 194, 204–211. [Google Scholar] [CrossRef] [PubMed]
- Polanczyk, G.V.; Salum, G.A.; Sugaya, L.S.; Caye, A.; Rohde, L.A. Annual Research Review: A Meta-Analysis of the Worldwide Prevalence of Mental Disorders in Children and Adolescents. J. Child Psychol. Psychiatry 2015, 56, 345–365. [Google Scholar] [CrossRef] [PubMed]
- Biederman, J.; Petty, C.R.; Clarke, A.; Lomedico, A.; Faraone, S.V. Predictors of Persistent ADHD: An 11-Year Follow-up Study. J. Psychiatr. Res. 2010, 45, 150. [Google Scholar] [CrossRef]
- Biederman, J.; Petty, C.R.; Woodworth, K.Y.; Lomedico, A.; Hyder, L.L.; Faraone, S.V. Adult Outcome of Attention-Deficit/Hyperactivity Disorder: A Controlled 16-Year Follow-up Study. J. Clin. Psychiatry 2012, 73, 941–950. [Google Scholar] [CrossRef]
- Hodgkins, P.; Montejano, L.; Sasané, R.; Huse, D. Risk of Injury Associated with Attention Deficit/Hyperactivity Disorder in Adults Enrolled in Employer-Sponsored Health Plans: A Retrospective Analysis. Prim. Care Companion J. Clin. Psychiatry 2011, 13, e1–e12. [Google Scholar] [CrossRef]
- Myer, N.M.; Boland, J.R.; Faraone, S.V. Pharmacogenetics Predictors of Methylphenidate Efficacy in Childhood ADHD. Mol. Psychiatry 2018, 23, 1929–1936. [Google Scholar] [CrossRef]
- Cortese, S.; Adamo, N.; Del Giovane, C.; Mohr-Jensen, C.; Hayes, A.J.; Carucci, S.; Atkinson, L.Z.; Tessari, L.; Banaschewski, T.; Coghill, D.; et al. Comparative Efficacy and Tolerability of Medications for Attention-Deficit Hyperactivity Disorder in Children, Adolescents, and Adults: A Systematic Review and Network Meta-Analysis. Lancet Psychiatry 2018, 5, 727. [Google Scholar] [CrossRef]
- Wolraich, M.L.; Hagan, J.F.; Allan, C.; Chan, E.; Davison, D.; Earls, M.; Evans, S.W.; Flinn, S.K.; Froehlich, T.; Frost, J.; et al. Clinical Practice Guideline for the Diagnosis, Evaluation, and Treatment of Attention-Deficit/Hyperactivity Disorder in Children and Adolescents. Pediatrics 2019, 144, e20192528. [Google Scholar] [CrossRef] [PubMed]
- Nasser, A.; Hull, J.T.; Chaturvedi, S.A.; Liranso, T.; Odebo, O.; Kosheleff, A.R.; Fry, N.; Cutler, A.J.; Rubin, J.; Schwabe, S.; et al. A Phase III, Randomized, Double-Blind, Placebo-Controlled Trial Assessing the Efficacy and Safety of Viloxazine Extended-Release Capsules in Adults with Attention-Deficit/Hyperactivity Disorder. CNS Drugs 2022, 36, 897–915. [Google Scholar] [CrossRef]
- Mill, J.; Asherson, P.; Browes, C.; D’Souza, U.; Craig, I. Expression of the Dopamine Transporter Gene Is Regulated by the 3′ UTR VNTR: Evidence from Brain and Lymphocytes Using Quantitative RT-PCR. Am. J. Med. Genet.-Neuropsychiatr. Genet. 2002, 114, 975–979. [Google Scholar] [CrossRef]
- Brookes, K.J.; Neale, B.M.; Sugden, K.; Khan, N.; Asherson, P.; D’Souza, U.M. Relationship between VNTR Polymorphisms of the Human Dopamine Transporter Gene and Expression in Post-Mortem Midbrain Tissue. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2007, 144, 1070–1078. [Google Scholar] [CrossRef]
- Wu, J.; Xiao, H.; Sun, H.; Zou, L.; Zhu, L.Q. Role of Dopamine Receptors in ADHD: A Systematic Meta-Analysis. Mol. Neurobiol. 2012, 45, 605–620. [Google Scholar] [CrossRef]
- Picetti, R.; Saiardi, A.; Samad, T.A.; Bozzi, Y.; Baik, J.H.; Borrelli, E. Dopamine D2 Receptors in Signal Transduction and Behavior. Crit. Rev. Neurobiol. 1997, 11, 121–142. [Google Scholar] [CrossRef]
- Park, L.; Nigg, J.T.; Waldman, I.D.; Nummy, K.A.; Huang-Pollock, C.; Rappley, M.; Friderici, K.H. Association and Linkage of α-2A Adrenergic Receptor Gene Polymorphisms with Childhood ADHD. Mol. Psychiatry 2005, 10, 572–580. [Google Scholar] [CrossRef]
- Kim, B.N.; Kim, J.W.; Hong, S.B.; Cho, S.C.; Shin, M.S.; Yoo, H.J. Possible Association of Norepinephrine Transporter -3081(A/T) Polymorphism with Methylphenidate Response in Attention Deficit Hyperactivity Disorder. Behav. Brain Funct. 2010, 6, 57. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, M.; Denardin, D.; Silva, T.L.; Pianca, T.; Roman, T.; Hutz, M.H.; Faraone, S.V.; Rohde, L.A. Association Between Alpha-2a-Adrenergic Receptor Gene and ADHD Inattentive Type. Biol. Psychiatry 2006, 60, 1028–1033. [Google Scholar] [CrossRef]
- Gizer, I.R.; Ficks, C.; Waldman, I.D. Candidate Gene Studies of ADHD: A Meta-Analytic Review. Hum. Genet. 2009, 126, 51–90. [Google Scholar] [CrossRef] [PubMed]
- Banlaki, Z.; Elek, Z.; Nanasi, T.; Szekely, A.; Nemoda, Z.; Sasvari-Szekely, M.; Ronai, Z. Polymorphism in the Serotonin Receptor 2a (HTR2A) Gene as Possible Predisposal Factor for Aggressive Traits. PLoS ONE 2015, 10, e0117792. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Sham, P.C.; Owen, M.J.; He, L. Meta-Analysis Shows Significant Association between Dopamine System Genes and Attention Deficit Hyperactivity Disorder (ADHD). Hum. Mol. Genet. 2006, 15, 2276–2284. [Google Scholar] [CrossRef]
- Grünblatt, E.; Werling, A.M.; Roth, A.; Romanos, M.; Walitza, S. Association Study and a Systematic Meta-Analysis of the VNTR Polymorphism in the 3′-UTR of Dopamine Transporter Gene and Attention-Deficit Hyperactivity Disorder. J. Neural. Transm. 2019, 126, 517–529. [Google Scholar] [CrossRef] [PubMed]
- Kustanovich, V.; Ishii, J.; Crawford, L.; Yang, M.; McGough, J.J.; McCracken, J.T.; Smalley, S.L.; Nelson, S.F. Transmission Disequilibrium Testing of Dopamine-Related Candidate Gene Polymorphisms in ADHD: Confirmation of Association of ADHD with DRD4 and DRD5. Mol. Psychiatry 2004, 9, 711–717. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, T.; Du, Y.; Hu, D.; Zhang, Y.; Li, H.; Pei, W. Polygenic Risk of Genes Involved in the Catecholamine and Serotonin Pathways for ADHD in Children. Neurosci. Lett. 2021, 760, 136086. [Google Scholar] [CrossRef]
- Hawi, Z.; Cummins, T.D.R.; Tong, J.; Johnson, B.; Lau, R.; Samarrai, W.; Bellgrove, M.A. The Molecular Genetic Architecture of Attention Deficit Hyperactivity Disorder. Mol. Psychiatry 2015, 20, 289–297. [Google Scholar] [CrossRef]
- Brown, J.T. The Pharmacogenetic Impact on the Pharmacokinetics of ADHD Medications. Methods Mol. Biol. 2022, 2547, 427–436. [Google Scholar] [CrossRef]
- Bishop, J.R.; Zhou, C.; Gaedigk, A.; Krone, B.; Kittles, R.; Cook, E.H.; Newcorn, J.H.; Stein, M.A. Dopamine Transporter and CYP2D6 Gene Relationships with Attention-Deficit/Hyperactivity Disorder Treatment Response in the Methylphenidate and Atomoxetine Crossover Study. J. Child Adolesc. Psychopharmacol. 2024, 34, 458–469. [Google Scholar] [CrossRef]
- Froehlich, T.E.; Epstein, J.N.; Nick, T.G.; Melguizo Castro, M.S.; Stein, M.A.; Brinkman, W.B.; Graham, A.J.; Langberg, J.M.; Kahn, R.S. Pharmacogenetic Predictors of Methylphenidate Dose-Response in Attention-Deficit/Hyperactivity Disorder. J. Am. Acad. Child Adolesc. Psychiatry 2011, 50, 1129–1139.e2. [Google Scholar] [CrossRef]
- Yuan, D.; Zhang, M.; Huang, Y.; Wang, X.; Jiao, J.; Huang, Y. Noradrenergic Genes Polymorphisms and Response to Methylphenidate in Children with ADHD: A Systematic Review and Meta-Analysis. Medicine 2021, 100, E27858. [Google Scholar] [CrossRef] [PubMed]
- Balogh, L.; Pulay, A.J.; Réthelyi, J.M. Genetics in the ADHD Clinic: How Can Genetic Testing Support the Current Clinical Practice? Front. Psychol. 2022, 13, 751041. [Google Scholar] [CrossRef] [PubMed]
- Kranz, T.M.; Grimm, O. Update on Genetics of Attention Deficit/Hyperactivity Disorder: Current Status 2023. Curr Opin Psychiatry 2023, 36, 257–262. [Google Scholar] [CrossRef]
- Elsayed, N.A.; Yamamoto, K.M.; Froehlich, T.E. Genetic Influence on Efficacy of Pharmacotherapy for Pediatric Attention-Deficit/Hyperactivity Disorder: Overview and Current Status of Research. CNS Drugs 2020, 34, 389–414. [Google Scholar] [CrossRef]
- Malik, S.; Verma, P.; Ruaño, G.; Al Siaghy, A.; Dilawar, A.; Bishop, J.R.; Strawn, J.R.; Namerow, L.B. Pharmacogenetics in Child and Adolescent Psychiatry: Background and Evidence-Based Clinical Applications. J. Child Adolesc. Psychopharmacol. 2024, 34, 4–20. [Google Scholar] [CrossRef]
- Pagerols, M.; Richarte, V.; Sánchez-Mora, C.; Rovira, P.; Soler Artigas, M.; Garcia-Martínez, I.; Calvo-Sánchez, E.; Corrales, M.; Da Silva, B.S.; Mota, N.R.; et al. Integrative Genomic Analysis of Methylphenidate Response in Attention-Deficit/Hyperactivity Disorder. Sci. Rep. 2018, 8, 1881. [Google Scholar] [CrossRef]
- Li, J.J.; He, Q. Polygenic Scores for ADHD: A Meta-Analysis. Res. Child Adolesc. Psychopathol. 2021, 49, 297–310. [Google Scholar] [CrossRef]
- Cabana-Domínguez, J.; Llonga, N.; Arribas, L.; Alemany, S.; Vilar-Ribó, L.; Demontis, D.; Fadeuilhe, C.; Corrales, M.; Richarte, V.; Børglum, A.D.; et al. Transcriptomic Risk Scores for Attention Deficit/Hyperactivity Disorder. Mol. Psychiatry 2023, 28, 3493–3502. [Google Scholar] [CrossRef]
- Zhong, Y.; Yang, B.; Su, Y.; Qian, Y.; Cao, Q.; Chang, S.; Wang, Y.; Yang, L. The Association with Quantitative Response to Attention-Deficit/Hyperactivity Disorder Medication of the Previously Identified Neurodevelopmental Network Genes. J. Child Adolesc. Psychopharmacol. 2020, 30, 348–354. [Google Scholar] [CrossRef]
- Vuijk, P.J.; Martin, J.; Braaten, E.B.; Genovese, G.; Capawana, M.R.; O’Keefe, S.M.; Lee, B.A.; Lind, H.S.; Smoller, J.W.; Faraone, S.V.; et al. Translating Discoveries in Attention-Deficit/Hyperactivity Disorder Genomics to an Outpatient Child and Adolescent Psychiatric Cohort. J. Am. Acad. Child Adolesc. Psychiatry 2020, 59, 964–977. [Google Scholar] [CrossRef]
- Contini, V.Ô.; Rovaris, D.L.; Victor, M.M.; Grevet, E.H.; Rohde, L.A.; Bau, C.H.D. Pharmacogenetics of Response to Methylphenidate in Adult Patients with Attention-Deficit/Hyperactivity Disorder (ADHD): A Systematic Review. Eur. Neuropsychopharmacol. 2013, 23, 555–560. [Google Scholar] [CrossRef] [PubMed]
- Bruxel, E.M.; Akutagava-Martins, G.C.; Salatino-Oliveira, A.; Contini, V.; Kieling, C.; Hutz, M.H.; Rohde, L.A. ADHD Pharmacogenetics across the Life Cycle: New Findings and Perspectives. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2014, 165, 263–282. [Google Scholar] [CrossRef]
- Cortese, S.; Sabé, M.; Chen, C.; Perroud, N.; Solmi, M. Half a Century of Research on Attention-Deficit/Hyperactivity Disorder: A Scientometric Study. Neurosci. Biobehav. Rev. 2022, 140, 104769. [Google Scholar] [CrossRef] [PubMed]
- Vergara, D.; del Bosque, A.; Lampropoulos, G.; Fernández-Arias, P. Trends and Applications of Artificial Intelligence in Project Management. Electronics 2025, 14, 800. [Google Scholar] [CrossRef]
- del Bosque, A.; Lampropoulos, G.; Vergara, D. Nanocomposites for Multifunctional Sensors: A Comprehensive Bibliometric Exploration. Nanomaterials 2024, 15, 34. [Google Scholar] [CrossRef]
- del Bosque, A.; Vergara, D.; Fernández-Arias, P. An Overview of Smart Composites for the Aerospace Sector. Appl. Sci. 2025, 15, 2986. [Google Scholar] [CrossRef]
- Moher, D.; Stewart, L.; Shekelle, P. Implementing PRISMA-P: Recommendations for Prospective Authors. Syst. Rev. 2016, 5, 15. [Google Scholar] [CrossRef]
- Aria, M.; Cuccurullo, C. Bibliometrix: An R-Tool for Comprehensive Science Mapping Analysis. J. Informetr. 2017, 11, 959–975. [Google Scholar] [CrossRef]
- Pliszka, S. Practice Parameter for the Assessment and Treatment of Children and Adolescents with Attention-Deficit/Hyperactivity Disorder. J. Am. Acad. Child Adolesc. Psychiatry 2007, 46, 894–921. [Google Scholar] [CrossRef] [PubMed]
- Greenhill, L.; Kollins, S.; Abikoff, H.; McCracken, J.; Riddle, M.; Swanson, J.; McGough, J.; Wigal, S.; Wigal, T.; Vitiello, B.; et al. Efficacy and Safety of Immediate-Release Methylphenidate Treatment for Preschoolers with ADHD. J. Am. Acad. Child Adolesc. Psychiatry 2006, 45, 1284–1293. [Google Scholar] [CrossRef]
- Bouchard, M.F.; Bellinger, D.C.; Wright, R.O.; Weisskopf, M.G. Attention-Deficit/Hyperactivity Disorder and Urinary Metabolites of Organophosphate Pesticides. Pediatrics 2010, 125, e1270–e1277. [Google Scholar] [CrossRef] [PubMed]
- Diamond, A. Attention-Deficit Disorder (Attention-Deficit/ Hyperactivity Disorder without Hyperactivity): A Neurobiologically and Behaviorally Distinct Disorder from Attention-Deficit/Hyperactivity Disorder (with Hyperactivity). Dev. Psychopathol. 2005, 17, 807–825. [Google Scholar] [CrossRef]
- Spencer, T.; Biederman, J.; Wilens, T.; Doyle, R.; Surman, C.; Prince, J.; Mick, E.; Aleardi, M.; Herzig, K.; Faraone, S. A Large, Double-Blind, Randomized Clinical Trial of Methylphenidate in the Treatment of Adults with Attention-Deficit/Hyperactivity Disorder. Biol. Psychiatry 2005, 57, 456–463. [Google Scholar] [CrossRef]
- MacDonald, S.W.S.; Li, S.C.; Bäckman, L. Neural Underpinnings of Within-Person Variability in Cognitive Functioning. Psychol. Aging 2009, 24, 792–808. [Google Scholar] [CrossRef]
- Cortese, S.; Angriman, M.; Maffeis, C.; Isnard, P.; Konofal, E.; Lecendreux, M.; Purper-Ouakil, D.; Vincenzi, B.; Dalla Bernardina, B.; Mouren, M.C. Attention-Deficit/Hyperactivity Disorder (ADHD) and Obesity: A Systematic Review of the Literature. Crit. Rev. Food Sci. Nutr. 2008, 48, 524–537. [Google Scholar] [CrossRef]
- Shaw, P.; Gornick, M.; Lerch, J.; Addington, A.; Seal, J.; Greenstein, D.; Sharp, W.; Evans, A.; Giedd, J.N.; Castellanos, F.X.; et al. Polymorphisms of the Dopamine D4 Receptor, Clinical Outcome, and Cortical Structure in Attention-Deficit/Hyperactivity Disorder. Arch. Gen. Psychiatry 2007, 64, 921–931. [Google Scholar] [CrossRef]
- Prehn-Kristensen, A.; Zimmermann, A.; Tittmann, L.; Lieb, W.; Schreiber, S.; Baving, L.; Fischer, A. Reduced Microbiome Alpha Diversity in Young Patients with ADHD. PLoS ONE 2018, 13, e0200728. [Google Scholar] [CrossRef] [PubMed]
- Froehlich, T.E.; McGough, J.J.; Stein, M.A. Progress and Promise of Attention-Deficit Hyperactivity Disorder Pharmacogenetics. CNS Drugs 2010, 24, 99–117. [Google Scholar] [CrossRef] [PubMed]
- Murray, A.L.; Booth, T.; Eisner, M.; Auyeung, B.; Murray, G.; Ribeaud, D. Sex Differences in ADHD Trajectories across Childhood and Adolescence. Dev. Sci. 2019, 22, e12721. [Google Scholar] [CrossRef]
- Mechler, K.; Banaschewski, T.; Hohmann, S.; Häge, A. Evidence-Based Pharmacological Treatment Options for ADHD in Children and Adolescents. Pharmacol. Ther. 2022, 230, 107940. [Google Scholar] [CrossRef] [PubMed]
- Henning, C.T.; Summerfeldt, L.J.; Parker, J.D.A. Longitudinal Associations Between Symptoms of ADHD and Life Success: From Emerging Adulthood to Early Middle Adulthood. J. Atten. Disord. 2024, 28, 1139–1151. [Google Scholar] [CrossRef]
- Franke, B.; Michelini, G.; Asherson, P.; Banaschewski, T.; Bilbow, A.; Buitelaar, J.K.; Cormand, B.; Faraone, S.V.; Ginsberg, Y.; Haavik, J.; et al. Live Fast, Die Young? A Review on the Developmental Trajectories of ADHD across the Lifespan. Eur. Neuropsychopharmacol. 2018, 28, 1059–1088. [Google Scholar] [CrossRef]
- Salatino-Oliveira, A.; Rohde, L.A.; Hutz, M.H. The Dopamine Transporter Role in Psychiatric Phenotypes. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2018, 177, 211–231. [Google Scholar] [CrossRef]
- Brown, A.B.; Biederman, J.; Valera, E.M.; Doyle, A.E.; Bush, G.; Spencer, T.; Monuteaux, M.C.; Mick, E.; Whitfield-Gabrieli, S.; Makris, N.; et al. Effect of Dopamine Transporter Gene (SLC6A3) Variation on Dorsal Anterior Cingulate Function in Attention-Deficit/Hyperactivity Disorder. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2010, 153B, 365–375. [Google Scholar] [CrossRef]
- Waldie, K.E.; Cornforth, C.M.; Webb, R.E.; Thompson, J.M.D.; Murphy, R.; Moreau, D.; Slykerman, R.; Morgan, A.R.; Ferguson, L.R.; Mitchell, E.A. Dopamine Transporter (DAT1/SLC6A3) Polymorphism and the Association between Being Born Small for Gestational Age and Symptoms of ADHD. Behav. Brain Res. 2017, 333, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.K.; Bhat, A.A.; Hashem, S.; Nisar, S.; Kamal, M.; Syed, N.; Temanni, M.R.; Gupta, R.K.; Kamran, S.; Azeem, M.W.; et al. Genetic Variations Influence Brain Changes in Patients with Attention-Deficit Hyperactivity Disorder. Transl. Psychiatry 2021, 11, 349. [Google Scholar] [CrossRef]
- Kuś, J.; Saramowicz, K.; Czerniawska, M.; Wiese, W.; Siwecka, N.; Rozpędek-Kamińska, W.; Kucharska-Lusina, A.; Strzelecki, D.; Majsterek, I. Molecular Mechanisms Underlying NMDARs Dysfunction and Their Role in ADHD Pathogenesis. Int. J. Mol. Sci. 2023, 24, 12983. [Google Scholar] [CrossRef]
- Stein, M.A.; McGough, J.J. The Pharmacogenomic Era: Promise for Personalizing Attention Deficit Hyperactivity Disorder Therapy. Child Adolesc. Psychiatr. Clin. N. Am. 2008, 17, 475–490. [Google Scholar] [CrossRef][Green Version]
- Turic, D.; Swanson, J.; Sonuga-Barke, E. DRD4 And DAT1 in ADHD: Functional Neurobiology to Pharmacogenetics. Pharmgenomics Pers. Med. 2010, 3, 61–78. [Google Scholar] [CrossRef]
- Guo, H.L.; Huang, J.; Wang, J.; Fan, L.; Li, Y.; Wu, D.D.; Liu, Q.Q.; Chen, F. Precision Pharmacotherapy of Atomoxetine in Children with ADHD: How to Ensure the Right Dose for the Right Person? Front. Pharmacol. 2024, 15, 1484512. [Google Scholar] [CrossRef]
- Kole, I.H.; Vural, P.; Yurdacan, B.; Alemdar, A.; Mutlu, C. Evaluation of SLC6A2 and CYP2D6 Polymorphisms’ Effects on Atomoxetine Treatment in Attention Deficit and Hyperactivity Disorder. Eur. J. Clin. Pharmacol 2024, 80, 1773–1785. [Google Scholar] [CrossRef]
- Michelson, D.; Read, H.A.; Ruff, D.D.; Witcher, J.; Zhang, S.; McCracken, J. CYP2D6 and Clinical Response to Atomoxetine in Children and Adolescents with ADHD. J. Am. Acad. Child Adolesc. Psychiatry 2007, 46, 242–251. [Google Scholar] [CrossRef]
- Fu, X.; Wu, W.; Wu, Y.; Liu, X.; Liang, W.; Wu, R.; Li, Y. Adult ADHD and Comorbid Anxiety and Depressive Disorders: A Review of Etiology and Treatment. Front. Psychiatry 2025, 16, 1597559. [Google Scholar] [CrossRef]
- Lewandrowski, K.U.; Blum, K.; Sharafshah, A.; Thanos, K.Z.; Thanos, P.K.; Zirath, R.; Pinhasov, A.; Bowirrat, A.; Jafari, N.; Zeine, F.; et al. Genetic and Regulatory Mechanisms of Comorbidity of Anxiety, Depression and ADHD: A GWAS Meta-Meta-Analysis Through the Lens of a System Biological and Pharmacogenomic Perspective in 18.5 M Subjects. J. Pers. Med. 2025, 15, 103. [Google Scholar] [CrossRef]
- Franke, B.; Faraone, S.V.; Asherson, P.; Buitelaar, J.; Bau, C.H.D.; Ramos-Quiroga, J.A.; Mick, E.; Grevet, E.H.; Johansson, S.; Haavik, J.; et al. The Genetics of Attention Deficit/Hyperactivity Disorder in Adults, a Review. Mol. Psychiatry 2012, 17, 960–987. [Google Scholar] [CrossRef] [PubMed]
- Demontis, D.; Walters, R.K.; Martin, J.; Mattheisen, M.; Als, T.D.; Agerbo, E.; Baldursson, G.; Belliveau, R.; Bybjerg-Grauholm, J.; Bækvad-Hansen, M.; et al. Discovery of the First Genome-Wide Significant Risk Loci for Attention Deficit/Hyperactivity Disorder. Nat. Genet. 2019, 51, 63–75. [Google Scholar] [CrossRef] [PubMed]
- Cortese, S. The Neurobiology and Genetics of Attention-Deficit/Hyperactivity Disorder (ADHD): What Every Clinician Should Know. Eur. J. Paediatr. Neurol. 2012, 16, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Kooij, S.J.J.; Bejerot, S.; Blackwell, A.; Caci, H.; Casas-Brugué, M.; Carpentier, P.J.; Edvinsson, D.; Fayyad, J.; Foeken, K.; Fitzgerald, M.; et al. European Consensus Statement on Diagnosis and Treatment of Adult ADHD: The European Network Adult ADHD. BMC Psychiatry 2010, 10, 67. [Google Scholar] [CrossRef]
- Shaw, P.; Eckstrand, K.; Sharp, W.; Blumenthal, J.; Lerch, J.P.; Greenstein, D.; Clasen, L.; Evans, A.; Giedd, J.; Rapoport, J.L. Attention-Deficit/Hyperactivity Disorder Is Characterized by a Delay in Cortical Maturation. Proc. Natl. Acad. Sci. USA 2007, 104, 19649–19654. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.