Abstract
Background: The reproductive cycle of Octopus mimus is regulated by environmental and hormonal factors, with photoperiod playing a key role in spawning induction and reproductive maturation. Understanding its underlying molecular mechanisms is essential for developing strategies to enhance controlled reproduction in aquaculture. Methods: We analyzed the expression of genes involved in the photoperiod-activated spawning induction cascade in the optic lobe and its downstream effects on the oviducal gland by performing transcriptomic analyses on females exposed to continuous light (24:0), which inhibits reproductive development, and a natural photoperiod, which induces spawning. The mRNA sequencing (RNA-Seq), quality control, gene annotation, and differential expression analyses were conducted using edgeR. Results: Spawning was completely inhibited under constant light, while 80% of control females spawned. Expression profiling revealed 89 downregulated and 34 upregulated genes in the optic lobe, and 178 downregulated and 237 upregulated genes in the oviducal gland (FDR < 0.05, |log2FC| ≥ 2), including key orthologs such as FMRFamide and myomodulin. Conclusions: These results show that the optic lobe integrates photoperiodic cues that modulate reproductive activation via a neuroendocrine cascade and coordinates spawning regulation through the oviducal gland, providing insights for improving reproductive control in aquaculture systems.
1. Introduction
Octopus mimus (Gould, 1852), Gould octopus, is a species distributed in intertidal and subtidal rocky habitats along the southeastern Pacific Ocean, ranging from northern Peru to central Chile [1]. In recent years, its exploitation has increased due to high commercial demand. Combined with overfishing and the impacts of climatic events such as El Niño and La Niña, this has led to a drastic decline in recorded catches [2]. Under these circumstances, the implementation of sustainable aquaculture strategies has emerged as an alternative to ensure supply without compromising natural populations. One of the main challenges in this process is the domestication of the species, particularly the establishment of a viable broodstock, which requires an understanding of the mechanisms that regulate reproduction, especially the induction of spawning under controlled conditions.
Like other octopods, O. mimus is a semelparous species, meaning that females undergo a single reproductive event in their lifetime. Following spawning, female octopuses cease feeding and rely on stored energy reserves, triggering a senescence process that culminates in death [3,4]. During this period, females exhibit egg-caring behavior, tending to egg clusters until hatching occurs [1,3]. The optic gland, a structure present in all cephalopods except for nautiloids, plays a key role in regulating this process and has been functionally compared to the vertebrate pituitary gland [5]. Accordingly, ablation of the optic gland in O. vulgaris females suppresses senescent behavior and permits the resumption of feeding, suggesting that this structure is responsible for initiating the terminal phase of the reproductive cycle [4].
Environmental conditions directly influence cephalopod reproduction [5]. Factors such as temperature, light, and food availability modulate gonadal maturation and spawning [1,6]. In octopods, reproduction has been observed to occur preferentially under semi-dark conditions, while continuous light exposure inhibits these processes [1,7]. Thus, reproductive induction is largely mediated by the optic gland, although other structures, such as the subpedunculate lobe, olfactory lobe, and optic lobe, are also involved in regulating the reproductive cycle [8]. The subpedunculate and olfactory lobes work together to modulate optic gland activity through the release of neuropeptides such as FMRFamide and GnRH [9,10]. Studies in O. vulgaris have shown that the olfactory lobe also regulates optic gland activity, influencing the transition from growth to reproduction [11]. Additionally, the optic lobes have been reported to release FMRFamide-like neuropeptides that regulate a variety of functions in mollusks, including reproduction [12]. Furthermore, a correlation between reproductive senescence and cholesterol metabolism has been identified, with genes such as 7DHC and DAF36 implicated in the activation of the optic gland [12,13]. The signaling cascade initiated by the optic gland in turn promotes fertilization through the functional activation of the oviducal gland (OG), another key reproductive organ. The production of oct-GnRH may play a role in the optic gland–gonadal axis, analogous to the hypothalamic–pituitary axis in vertebrates [12,14].
Following gonadal maturation, the oviducal gland becomes functionally active, facilitating fertilization and oocyte encapsulation. During mating, sperm are stored in the oviducal gland and can remain viable for up to ten months before being reactivated for fertilization [3,4]. The oviducal gland also participates in the metabolism and secretion of steroid hormones that coordinate fertilization and oviposition [15]. Once fertilized, the chorion hardens to prevent polyspermy and to protect the embryo from mechanical damage and bacterial infection [16,17]. Finally, the eggs are transported by peristalsis and released in the form of an ovarian string, which the female affixes to a suitable substrate [17]. Although it has been proposed that structures such as the optic gland, subpedunculate lobe, olfactory lobe, and optic lobe participate in the regulation of the reproductive cycle [8], a direct relationship between each of these structures and specific environmental factors triggering their activation has yet to be established [18].
Nevertheless, considering the optic lobe’s specialization in the perception and interpretation of visual stimuli, it is hypothesized that this organ may be involved in the initial detection of photoperiod or light intensity, functioning as a key environmental sensor in reproductive regulation. Indeed, octopods possess an eye structure analogous to that of vertebrates [19], and extraocular photoreception mediated by skin-expressed opsins has also been demonstrated [20]. Together, these adaptations underscore the pivotal role of light in their evolutionary and environmental resilience [21]. Based on this evidence, the present study aims to identify differentially expressed genes in the optic lobe and oviducal gland of O. mimus females whose spawning induction was inhibited by constant light exposure (24:0), in comparison with a control group maintained under semi-dark conditions and in natural photoperiod (14:10). Given that the onset of spawning involves profound metabolic changes and is associated with self-destructive behavior, we propose that this process relies on the integration of internal and external cues processed by specialized organs. We hypothesize that the optic lobe, by interpreting visual signals such as photoperiod, provides specific input which, in concert with other factors, activates a neuroendocrine cascade involving the optic gland, olfactory gland, and supraesophageal structures, ultimately leading to fertilization, oviposition, and senescence in O. mimus females.
2. Materials and Methods
2.1. Experimental Design and Sample Collection
Thirty octopus individuals were captured using artisanal fishing techniques off the coast of Vila Vila in southern Peru (−18.1154453, −70.7242811) between October-November 2023. The specimens were transported in two 150 L tanks at a temperature of 15 °C with oxygen saturation to the experimental aquaculture facility in Vila Vila, southern Peru. Upon arrival, the octopuses were anesthetized with 1.5% magnesium chloride for initial weighing and sex determination. They were then distributed into 500 L circular tanks at a female-to-male ratio of 2:1, under a seawater flow-through system maintained in semi-darkness conditions (<100 lx), with a water temperature of 18 ± 1 °C and a salinity of 35 PSU. Each tank contained individual shelters made from PVC tubes (25 × 50 × 20 cm; length × width × diameter). The octopuses were fed ad libitum every two days with crab (Platyxanthus orbigny) and “muy muy” (Emerita analoga).
After 15 days, individuals were separated by sex, forming a total of 10 female-designated tanks, each containing two females along with their respective shelters. During this period, five of the tanks were exposed to artificial illumination using two LED tubes positioned above the water surface, emitting an intensity of 500 lx with a 24:0 (light:dark) photoperiod. The remaining five tanks served as the control group and were maintained under semi-darkness using a shading mesh, ensuring illumination levels remained below 100 lx under a natural photoperiod. Throughout the experiment, feeding procedures and physicochemical parameters remained constant, while monitoring was conducted to observe whether females ceased feeding or initiated spawning. Sampling was performed once, when at least half of the females in the control group had begun spawning, i.e., when egg clusters were present in the shelters [1,15].
For sampling, the octopuses were anesthetized in magnesium chloride (1.5% for 10 min and 3.5% for 15 min), after which dissections were conducted using entomological needles (Ento Sphinx inox 0.2 mm, Entomopraxis SCP, Lima, Peru). The optic lobe and oviducal gland were dissected in triplicate under sterile conditions, submerged in RNAlater (Invitrogen, Carlsbad, CA, USA), and stored at −80 °C until RNA extraction.
2.2. RNA Extraction and Sequencing
Total RNA was extracted from each tissue sample using the NZY Total RNA Isolation Kit (NZYtech, Lisbon, Portugal) following the manufacturer’s protocol. RNA concentration was measured with DeNovix DS-11 FS and RNA integrity was checked by running an aliquot of total RNA (~500 ng) on a 1% (w/v) agarose gel stained with GelRed™ nucleic acid stain (Biotium, Hayward, CA, USA). The RNA quality was evaluated by sample absorbance according to the A260/280 and A260/230 ratios using a BiotTek® Synergy HT microplate reader (Synergy HT, BioTek Instruments, Inc., Winooski, VT, USA). Strand-specific libraries were prepared at Novogen Inc. (Novagen, Darmstadt, Germany) using an insert size of 250–300 bp. Sequencing was performed on an Illumina NovaSeq 6000 platform (Illumina, San Diego, CA, USA), generating 150 bp paired-end reads.
2.3. Quality Control and Preprocessing
Raw sequencing reads were assessed for quality using FastQC v0.11.8 [22]. Illumina adapter sequences and low-quality bases were removed with Trimmomatic v0.38 [23] using the following parameters: LEADING:5 TRAILING:5 SLIDINGWINDOW:5:20 MINLEN:36. To correct sequencing errors, reads were processed with Rcorrector v1.0.3 [24]. Clean reads were subsequently re-evaluated using FastQC to ensure quality improvement.
2.4. Genome Annotation and Read Mapping
Since O. mimus has an available genome in NCBI (Genome assembly ASM2756897v1) but lacks annotation, we performed structural annotation using AUGUSTUS v3.4.0 [25]. AUGUSTUS was run using default parameters and the generic species model (--species = generic) in ab initio prediction mode, without external evidence such as RNA-Seq hints. Gene predictions were generated based solely on the input genome sequence, and the output was provided in GTF format, including only complete gene structures.
Prior to annotation, the integrity of the genome assembly was assessed with BUSCO v4.1.4, using the “eukaryota_odb9” dataset. The resulting GTF file was then converted to GFF format using AGAT v1.0.0 [26] for downstream integration and analysis. Then, the annotated genome was indexed with HISAT2 v2.2.1 [27] for efficient read alignment. Preprocessed RNA-Seq reads were mapped to the indexed genome using HISAT2 v2.2.1 with the following parameters: --max-intronlen 50,000 --rna-strandness RF --dta. Alignment statistics were assessed using SAMtools v1.9 [28] and read distribution across genomic features was verified.
Functional annotation of the predicted proteins from the genome and annotation datasets was performed using the AGAT toolkit [26]. General Feature Format (GFF3) files were generated from the initial GFF output files produced by AUGUSTUS, allowing for comprehensive structural analysis and preparation of data for downstream functional analyses. From the O. mimus genome, features including splice sites (SS), exons, coding sequences (CDS), and transcripts were extracted. To validate the functional annotation and obtain homology-based evidence, sequence similarity searches were conducted using DIAMOND [29]. Both BLASTp and BLASTn alignments were performed against major curated protein and nucleotide databases, including UniProt, Swiss-Prot (downloaded 18 February 2020; The UniProt Consortium, 2016) and NCBI non-redundant databases (downloaded 10 March 2024). Additionally, nucleotide and amino acid sequences for all genes, transcripts, and proteins were retrieved in FASTA format to facilitate further functional filtering.
2.5. Differential Gene Expression Analysis
Feature counts were generated using StringTie v.2.1.2 [30], and differential gene expression analysis was conducted using the edgeR package (v.3.26.8) [31] in R (v.3.6.1) to compare Light and Dark (control) photoperiod conditions in females for both tissues. For normalization across samples, the trimmed mean of M-values (TMM) method [32] was employed. Genes were retained for differential analysis if they showed at least 1 CPM (counts per million) in at least three biological replicates. Statistical significance for differential expression was determined using a False Discovery Rate (FDR)-adjusted p-value < 0.05 and a |log2 fold change| ≥ 2. Hierarchical heatmaps were generated using the Heatmapper Expression tool [33], applying the Average Linkage clustering method with Euclidean distance. Differential expression results were visualized using the EnhancedVolcano R package [34]. A multidimensional scaling plot (MDS) using the Glimma package and principal component analysis (PCA) using PCAtools [35] were also implemented to assess sample variance.
Raw sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject accession number PRJNA1416070, with individual libraries available under SRA accession numbers SRR37032714–SRR37032725.
2.6. Gene Ontology and KEGG Pathway Enrichment Analysis
For the identification of differentially expressed genes (DEGs) and proteins, filtering was conducted using seqtk subseq based on lists of gene IDs obtained from prior differential expression analyses. This enabled the isolation of sequences specific to each expression group for subsequent enrichment studies. Functional enrichment analysis of DEGs was carried out to identify overrepresented biological processes and metabolic pathways. Gene Ontology (GO) enrichment was performed using the g:GOSt module of the g:Profiler web tool [36]. The analysis included GO terms associated with biological processes (BP), molecular functions (MF), and cellular components (CC). Statistical significance of GO term enrichment was evaluated using Fisher’s exact test (p < 0.05) and multiple comparisons were corrected using the Benjamini–Hochberg FDR procedure. GO terms passing the adjusted significance threshold were considered significantly enriched.
For pathway enrichment analysis, DEGs were annotated using the Kyoto Encyclopedia of Genes and Genomes (KEGG) Automatic Annotation Server (KAAS) [37]. The single-directional best hit (SBH) method was selected for KO (KEGG Orthology) assignment, using the default set of representative eukaryotic species as reference. The resulting KO identifiers were mapped to the Octopus bimaculoides genome to reconstruct species-specific KEGG pathway maps. Visualization and further statistical analysis of KEGG pathway enrichment were performed in R (v4.2.0) using the clusterProfiler [38] and enrich KEGG functions. Pathways with an adjusted p-value (FDR < 0.05) were considered significantly enriched.
2.7. Statistical Analysis
To assess whether spawning incidence differed significantly between treatments (constant light vs. natural photoperiod), Fisher’s exact test was applied using R software (version 4.3.1). The test compared the proportion of spawning females between the two experimental conditions, using ten independent females per treatment (N = 20), to determine whether the observed difference was statistically significant (p < 0.05) [39].
3. Results
3.1. Experimental Female Induction
The females were maintained for approximately two months in the absence of males, with one group exposed to constant photoperiod (Light) and the control group kept under natural photoperiod conditions (Dark) until the first spawning was detected. A total of 8 out of 10 females (80%) in the control group initiated spawning, whereas none of the females (0%) exposed to continuous photoperiod spawned during the experimental period. This result indicates that the inhibition of spawning under constant photoperiod and light intensity conditions was statistically significant (Fisher’s exact test, p = 0.047, 95% CI = 1.02).
3.2. RNA Sequencing and Data Quality
A total of 63 gigabases (Gb) of sequencing data and approximately 330 million raw reads were generated from all samples derived from the optic lobe and oviducal gland of O. mimus. After quality control and preprocessing, a total of 59 gigabases (Gb) and approximately 320 million high-quality reads were retained. Over 97% of bases had a Phred score above Q30, indicating high sequencing accuracy (Table S1). Adapter trimming and error correction substantially enhanced read quality, as assessed by FastQC reports.
3.3. Genome Annotation and Read Mapping Outcomes
The BUSCO analysis of the Octopus mimus genome (Genome assembly ASM2756897v1) provided crucial insights into assembly completeness by assessing the presence of universal orthologous genes. The results indicated that 32.2% of the expected genes were identified as complete, all of which appeared as single-copy sequences with no duplicated genes detected. However, 36.1% of the genes were only partially present, appearing as fragmented sequences. Additionally, 31.7% of the expected genes were absent from the assembly, indicating that a substantial portion of the genetic information was not properly captured. The analysis encompassed a total of 255 BUSCO groups, representing the complete set of expected orthologous genes in eukaryotic genomes, based on the reference dataset rojecti_odb9.
In addition to the genome completeness assessment, the annotation process identified a total of 16,724 genes, out of approximately 23,000 genes expected based on related octopus species (O. vulgaris, O. bimaculoides), each with the same number of corresponding transcripts, reflecting an average of one transcript per gene. The genomic annotation revealed a total of 75,525 exons, of which 72,588 were located within coding sequences (CDS), 1216 were mapped to 5′ untranslated regions (5′ UTR), and 1105 to 3′ untranslated regions (3′ UTR). Regarding intronic regions, 55,864 introns were identified within CDS regions, while the total number of introns present across exonic regions reached 58,801.
Read alignment to the annotated genome yielded an average mapping rate of 65 ± 10% across all samples. This variation may be affected by factors such as differences in sequencing depth of 34–42 million raw reads, variable trimming losses (4–8%), GC content bias (37–45%) among tissues (Table S1). Structural annotation covered 39.9% of the genome by gene and transcript features. Coding sequences (CDS) and exons represented 0.7% and 0.8% of the genome, respectively. The longest predicted gene and transcript measured 1,052,705 bp, with the longest CDS spanning 47,952 bp.
3.4. Differential Gene Expression Analysis Identified
The expression patterns observed across the sample set demonstrated a strong clustering effect based on treatment conditions, with the highest degree of integration detected in tissues from females exposed to constant light (Figure S1). Furthermore, principal component analysis (PCA) of log-CPM values showed they were strictly clustered by tissue type and treatment, with no evidence of batch-based clustering (Figure S2).
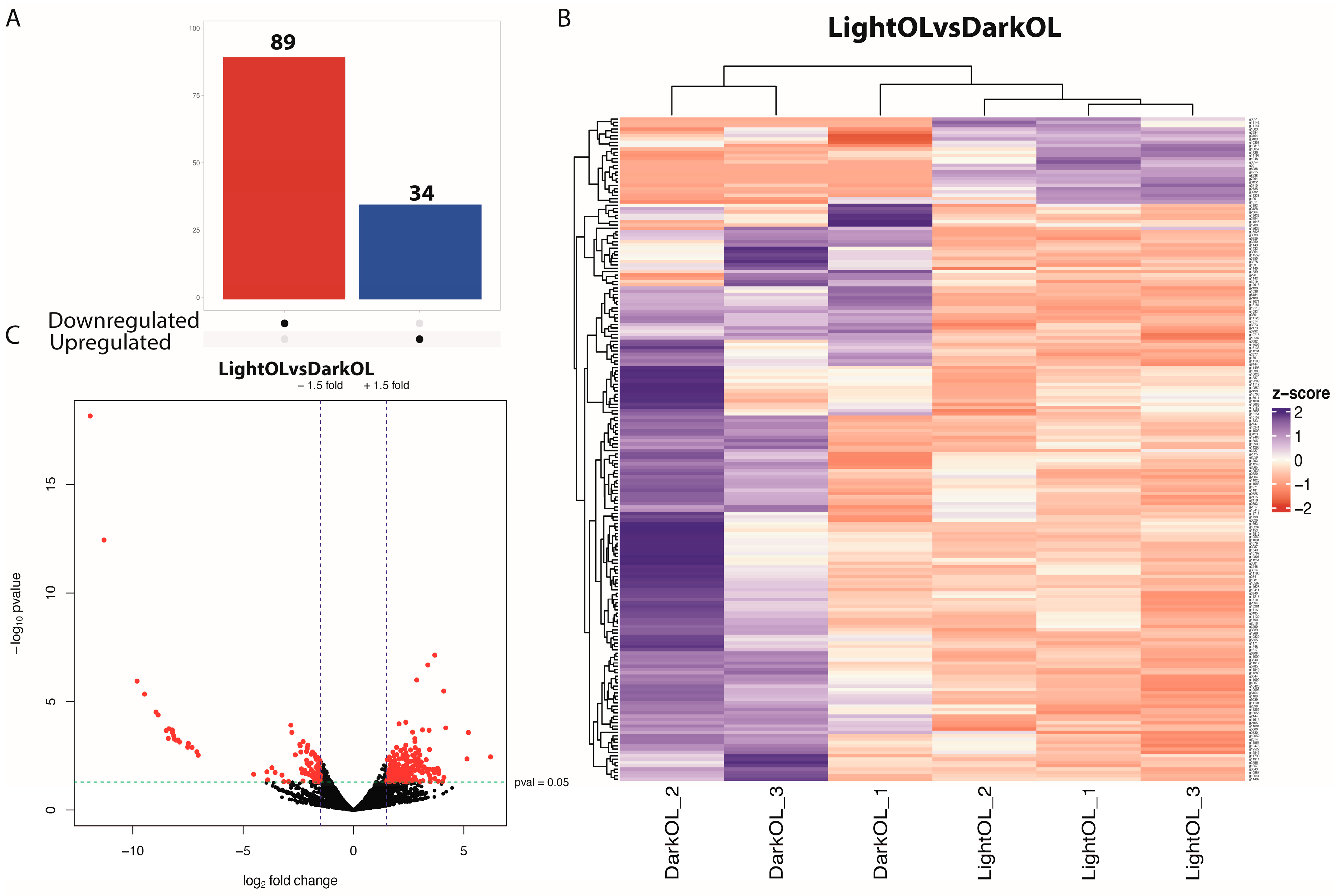
Of the 6578 genes that passed the threshold, differential gene expression (DGE) analysis in the optic lobe (OL) revealed 89 downregulated and 34 upregulated genes (adjusted p-value < 0.05, |log2FC| > 2; Figure 1A,B) in females exposed to a constant photoperiod compared to controls. The hierarchical clustering heat map analysis of OL DEGs was shown in Figure 1C. Hierarchical clustering revealed an over-representation of downregulated genes associated with control-related functions.
Figure 1.
Differential gene expression in the optic lobe of Octopus mimus females exposed to constant light conditions compared to those maintained under natural conditions (dark). (A), Differential expression analysis highlighting upregulated genes (blue) and downregulated genes (red) in the optic lobe under constant light (24:0) compared to natural photoperiod conditions (FDR < 0.05). (B), Heatmap and hierarchical clustering of differentially expressed genes based on normalized expression values. Columns represent individual optic lobe samples from females maintained under natural conditions (Dark OL 1–3) and constant light conditions (Light OL 1–3). Color hue and intensity indicate relative expression levels, showing clear separation by photoperiod treatment. (C), Volcano plot showing the distribution of gene expression changes, where the x-axis represents log2 fold change and the y-axis represents −log10 adjusted p-value. Genes on the left correspond to transcripts downregulated under constant light, whereas genes on the right represent transcripts upregulated under constant light.
The predominance of downregulated genes suggests that light exposure Inhibits the molecular signaling cascade responsible for spawning induction. Genes associated with cholinergic neurons specific to the optic lobes were detected, such as cholinergic receptor nicotinic alpha 7 subunit (CHRNA7). Expression of FMRFamide receptor (FMRFaR), FMR1 autosomal homolog 1 (Fxr1), and hydroxysteroid (17-beta) dehydrogenase 4 (HSD17B4) were identified as upregulated and associated with reproduction in OL. Upregulated genes related to vision in orcokinin and octopaminergic neurons were identified, including rojectingn 5A (Sema5a) and protocadherins. Among the key downregulated genes were the transcription factor SRY-box 30 (SOX30) and beta-tubulin (SP-beta1), whereas upregulated genes included forkhead box P1 (FOXP1) and the hormone-inducible receptor Hr38.
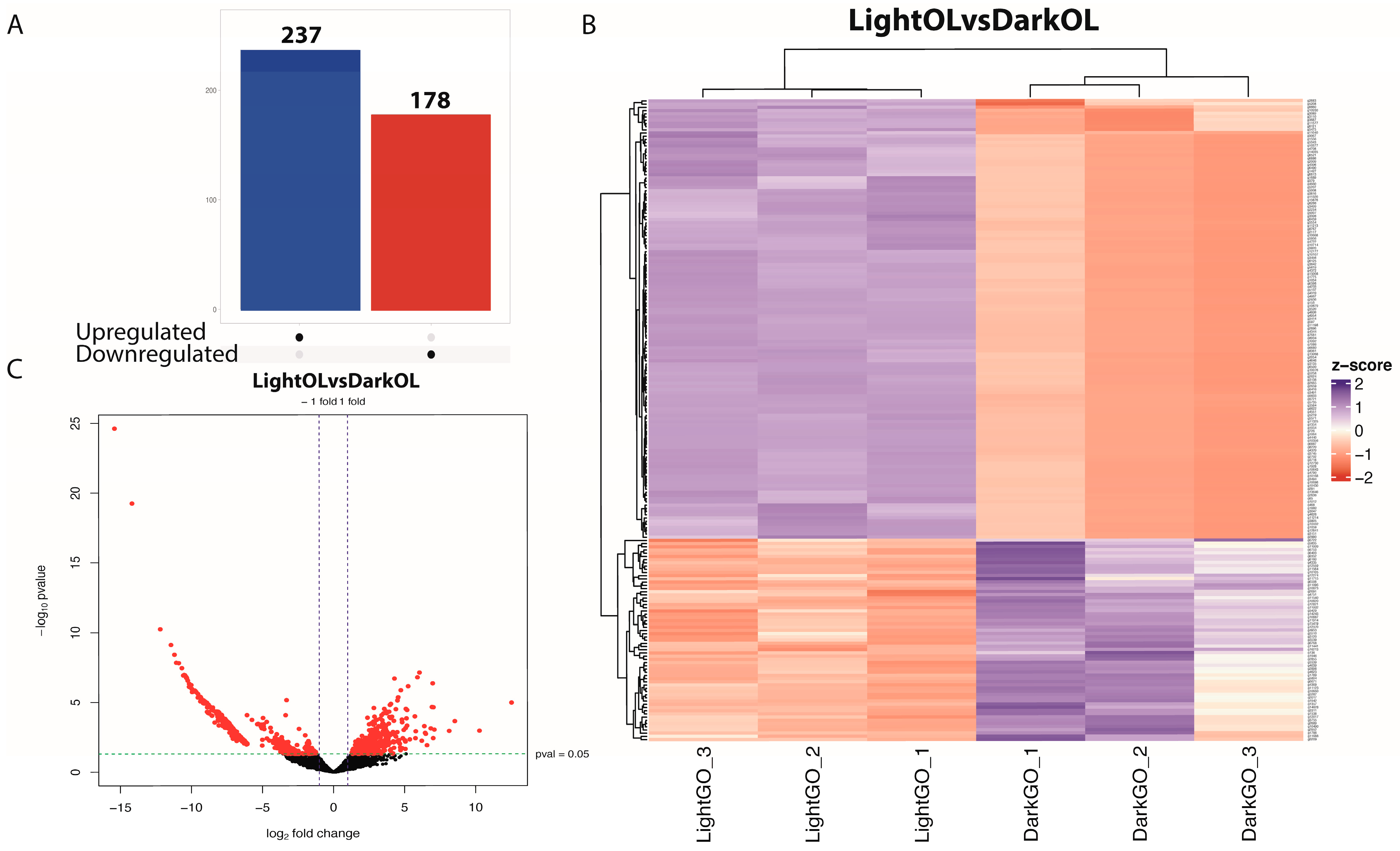
Of the 8025 genes that passed the initial filtering criteria, a total of 415 genes were identified as differentially expressed in the oviducal gland (adjusted p-value < 0.05, |log2FC| ≥ 2). Among these, 178 genes were downregulated and 237 were upregulated (Figure 2A,B). The remaining genes were excluded from downstream DEG-focused analyses. Key downregulated genes included forkhead box N3 (FOXN3) and TGFB-induced factor homeobox 2 (TGIF2), both associated with maturation induction. In contrast, overexpressed genes such as hormone-inducible receptor 39 (Hr39) and thyroid hormone receptor interactor 12 (TRIP12) were involved in hormonal regulation. Furthermore, stress response-related genes, such as heat shock protein 70 (HSP70) and ICE-like death-related caspase (Drice), were overexpressed in females exposed to constant light. Hierarchical clustering revealed n over-representation of genes downregulated under control conditions (Figure 2C). Signature genes for the oviducal gland were also detected, such as FMRFamide receptor (FMRFaR) and hydroxysteroid (17-beta) dehydrogenase 8 (HSD17B8), both related to fertilization, which were found in the clustering analysis of expression in spawning females. Other OG-specific genes, such as myomodulin, which function as a neurotransmitter in mollusks, were also identified.
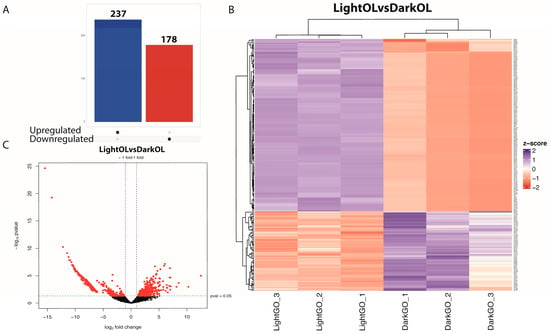
Figure 2.
Differential gene expression in the oviducal gland of Octopus mimus females exposed to constant light conditions compared to those maintained under natural photoperiod conditions (dark). (A), The panel shows the overall differences in gene expression, highlighting transcripts significantly upregulated (blue) or downregulated (red) under constant light (24:0) relative to natural conditions (FDR < 0.05). (B), Heatmap and hierarchical clustering of differentially expressed genes across individual oviducal gland samples. Columns correspond to females maintained under natural (Dark OG 1–3) and constant light (Light OG 1–3) conditions. Color intensity represents normalized relative expression, illustrating clear segregation between treatments. (C), Volcano plot depicting the distribution of gene expression changes. The x-axis represents log2 fold change, and the y-axis shows −log10 adjusted p-value. Genes on the left are downregulated in response to constant light, while those on the right are upregulated, reflecting photoperiod-dependent transcriptional modulation in the oviducal gland.
3.5. Functional Enrichment and Pathway Analysis
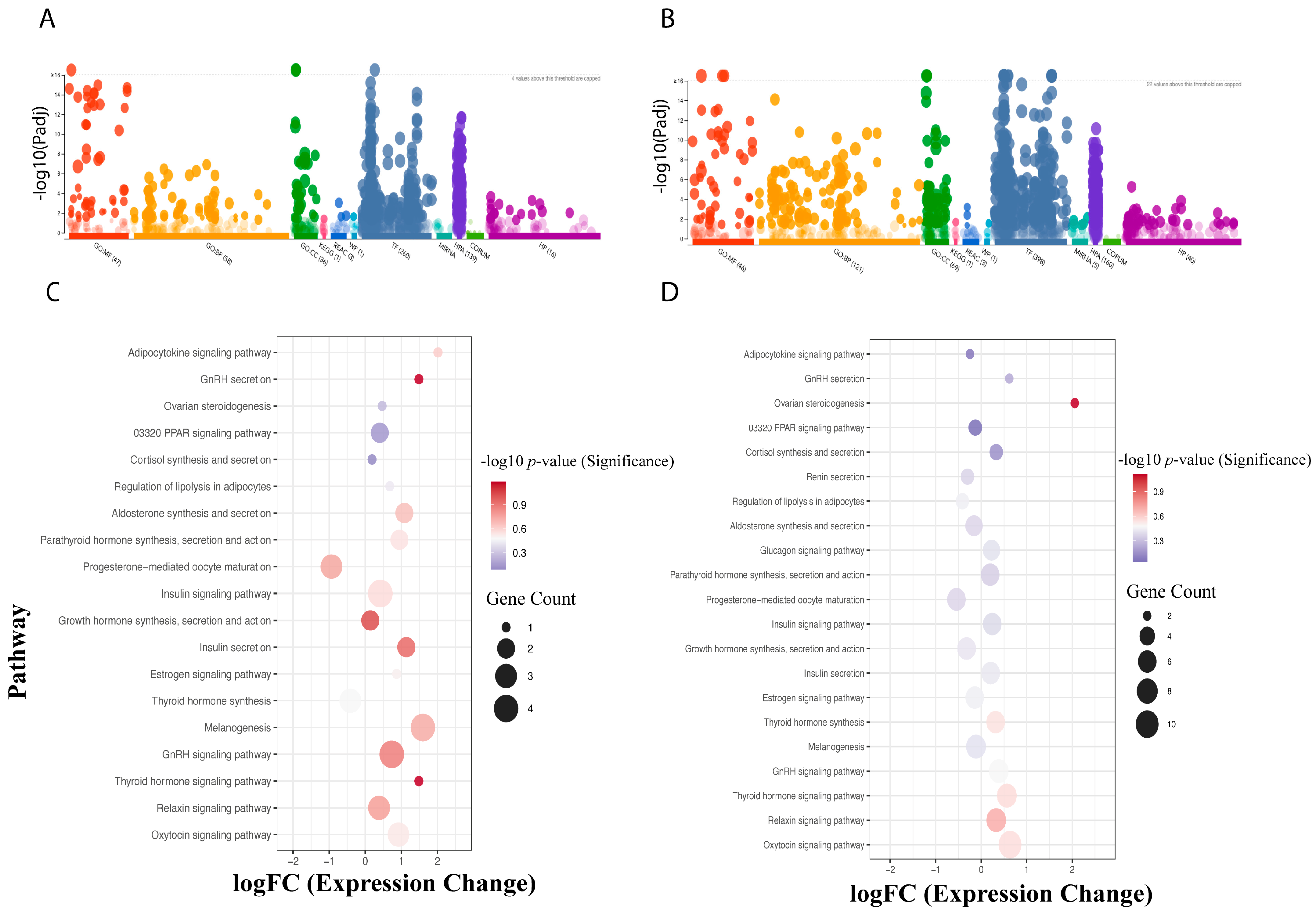
Functional annotation and Gene Ontology (GO) enrichment analysis were first performed for treatments under Light and Dark conditions. Differentially expressed genes (DEGs) were annotated into three primary GO categories: Molecular Function (MF), Biological Process (BP), and Cellular Component (CC). A higher number of DEGs accounted for the differential expression in the oviducal gland compared to the optic lobe (Figure 3A,B). A significant GO enrichment analysis was conducted across all DEGs, identifying 27 GO terms with significant enrichment, including catalytic activity, nucleoside phosphate binding, and hexosyltransferase activity (Table S2). Regarding BP, enriched terms included intracellular transport, macromolecule modification, cell cycle process, regulation of TOR signaling, and DNA metabolic process. For CC, significantly enriched terms included catalytic complex, intracellular protein-containing complex, and MutLalpha complex.
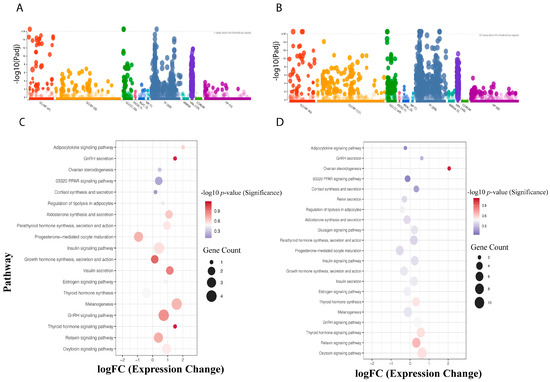
Figure 3.
Enriched functional annotations and endocrine pathways for differentially expressed genes (DEGs) in the optic lobe (A,C) and oviducal gland (B,D) of Octopus mimus females. Panels (A,B) illustrate significantly enriched Gene Ontology (GO) terms, including Molecular Function (MF, red), Biological Process (BP, orange), and Cellular Component (CC, green), together with pathway annotations from KEGG (pink), Reactome (blue), WikiPathways (turquoise), Transcription Factors (bluish-gray), microRNA (aqua green), Human Protein Atlas (violet), CORUM (light green), and Human Phenotype Ontology (magent). Panels (C,D) depict enriched KEGG endocrine pathways inferred through orthologous genes from Octopus bimaculoides. Comparison is between females exposed to constant light (24:0) with egg-laying inhibition and those under a normal (dark/light) photoperiod, highlighting photoperiod-dependent modulation of key neuroendocrine pathways.
The KEGG metabolic pathway database was used to compare and annotate all DEGs, resulting in a total of 336 genes associated with metabolic pathways. The comparison between Light and Control groups revealed that the optic lobe had a lower number of KEGG Orthologs (kOs) than the oviducal gland. However, in the results for endocrine-specific process assignments, gene–pathway coverage was less diverse in the optic lobe than in the oviducal gland (Figure 3C,D). Moreover, the enrichment analysis for endocrine-related KEGG pathways in the optic lobe was mainly associated with GnRH secretion, Thyroid hormone signaling pathway, GnRH signaling pathway, and Growth hormone synthesis, secretion, and action. In contrast, in the oviducal gland, we found pathways such as Ovarian steroidogenesis and Relaxin signaling pathway to be represented (Figure 3C,D).
4. Discussion
The results of this study demonstrate that continuous light exposure induces distinct phenotypic and transcriptomic alterations in Octopus mimus females. Continuous light exposure completely inhibited spawning in O. mimus females, as none of the individuals under a 24:0 photoperiod spawned, while 80% of females under a 14:10 light:dark regime successfully initiated spawning [1,15]. Moreover, the multidimensional scaling (MDS) analysis revealed that transcriptomic profiles of both the optic lobe and the oviducal gland in the constant light group were highly homogeneous, whereas profiles from the control group showed greater dispersion (Figures S1 and S2). This robust segregation in gene expression suggests that the optic lobe plays a critical role in interpreting photoperiodic signals, as evidenced by the low dispersion in gene expression of among samples exposed to constant photoperiod.
In the optic lobe, differential gene expression analysis revealed a predominance of negatively regulated genes related to general control functions under constant light. However, key genes involved in visual processing and sensory integration, including the alpha-7 subunit of the nicotinic cholinergic receptor (CHRNA7), rojectingn 5A (Sema5a), and several protocadherins, were differentially expressed, confirming that the optic lobe actively participates in processing photic information [40]. The optic lobe was included as the primary sensory node because it integrates visual input and mediates neuroendocrine signaling in cephalopods, as previously reported in O. vulgaris [8,12]. This role is supported by our transcriptomic data, which reveal differential expression of genes involved in photic perception (e.g., CHRNA7, Sema5a, protocadherins) under constant light conditions (Figures S1 and S2). We further propose a timing function for the optic lobe, whereby it interprets photoperiodic changes and conveys this information to downstream neuroendocrine centers, including the subesophageal lobe and optic gland, thereby coordinating the onset of spawning. This extends previous models that primarily described the optic lobe as a visual processing center, providing new insights into temporal regulation of reproduction. Differential expression of neuroendocrine-related genes, such as FMRFamide receptors in both the optic and oviducal glands, supports this timing function, indicating that the optic lobe not only senses photoperiod but also initiates downstream reproductive pathways leading to spawning.
Furthermore, we identified the expression of FMRFamide in the optic lobe, which has been implicated in fertilization in Sepiella japonica [40] and is also expressed in the optic gland of O. vulgaris [8]. Additionally, the expression of FMRFamide, together with dopamine, was verified in the common octopus (O. vulgaris) as a modulator for retinal light adaptation and an inhibitor of sexual maturation, making it a key gene for the relationship between light and spawning inhibition [1,12]. Consistent with this identification, FMRFamide receptors (FMRFaR) were found in the oviducal gland, where their activation in the presence of FMRFamide is linked to the passage of oocytes through the oviducal canal by causing canal contraction [7,41]. Furthermore, myomodulin, a neurotransmitter associated with the cephalopod peduncle complex, was differentially expressed in the oviducal gland [42]. This peduncle complex, located along the optic tract between the optic lobe and the central brain, includes the peduncle lobe (visomotor regulation), the olfactory lobe (olfactory input processing), and the optic gland (sexual maturation and maternal death control) [12,13]. Based on our transcriptomic data, we propose a sequential framework in which photic information is first integrated by the optic lobe and then transmitted to downstream neuroendocrine centers, including the peduncle and olfactory lobes, leading to optic gland activation or inhibition. The oviducal gland subsequently acts as the terminal effector coordinating fertilization and oviposition. These results refine current models by defining a functional sequence linking sensory perception, neuroendocrine integration, and reproductive output in female octopuses [42].
However, although our data support a putative regulatory network linking the optic lobe, optic gland, and oviducal gland, the observed gene expression changes (e.g., in FMRFamide, CHRNA7, and HSD17B8) appear to be functionally related. Nevertheless, the coordinated and tissue-specific regulation of these genes under constant light conditions suggests a structured neuroendocrine response rather than a stochastic transcriptional effect. In particular, the modulation of cholinergic signaling (CHRNA7), neuropeptidergic pathways (FMRFamide), and steroid metabolism (HSD17B8) has been previously implicated in mechanisms controlling reproductive inhibition and maturation in cephalopods [13,14,42]. However, specific functional studies, such as gene knockout or receptor assays, will be required to validate their causal roles in photoperiod-mediated spawning. Moreover, hierarchical clustering of the transcriptomes from the oviducal gland revealed that, in general, the reproductive state of females is reflected at the molecular level. In control females, expression profiles indicated active pathways related to gamete maturation and fertilization, whereas constant light exposure was associated with the suppression of genes involved in these processes. Our results confirm this pattern and provide additional transcriptomic evidence that photoperiod regulates oviducal gland activity. Notably, the differential expression of markers such as the forkhead box N3 (FOXN3), the TGFB-induced homeobox 2 factor (TGIF2), and the hormone-inducible receptor 39 (Hr39) underscores that the activity of the oviducal gland is closely tied to the initiation of spawning in Octopus sinensis and Octopus minor, serving as an effective indicator of the overall reproductive response [43,44].
Genes identified in both the optic lobe and oviducal gland, such as hydroxysteroid (17-beta) dehydrogenase 8 (HSD17B8), which is involved in steroid formation for reproduction [12,45], were also detected in the clustering analysis of expression in reproductive females. Thus, the activity of HSD17B8 in the white body during reproduction in Octopus maya females [14] aligns with the activity detected in the optic lobe in this study.
Gene Ontology (GO) enrichment analysis supported the concept of cellular recycling during reproductive inhibition. Genes associated with catalytic activity, apoptosis, and exocytosis were significantly enriched, suggesting a process of reallocation of cellular resources initially destined for gamete production. These findings were complemented by KEGG pathway analysis, which showed enrichment in pathways related to apoptosis, metabolic reprogramming, and autophagy, a pattern that aligns with reports indicating that oocytes and components of accessory glands are reabsorbed under inhibitory conditions [46,47]. Our enrichment analysis results show that KEGG pathways involved in female self-destruction [13], such as apoptosis, autophagy, and necroptosis, are activated, consistent with oocyte resorption and glandular remodeling at the onset of spawning.
According to our findings, endocrine-related KEGG pathways in the optic lobe were mainly associated with GnRH secretion and the GnRH signaling pathway, a reproduction-related pathway linked to the olfactory lobe in the common octopus (Octopus vulgaris) [7,9]. This activity in the optic lobe indicates synergistic interactions between the different lobes and glands coordinating sensory input, neuroendocrine signaling, and reproductive activation, such as octopus reproduction. Additionally, other relevant pathways included the thyroid hormone signaling pathway, identified in the Pacific oyster Crassostrea gigas and associated with calcium-related developmental and reproductive processes with effects on development and reproduction [48]. Within the oviducal gland, we detected pathways such as ovarian steroidogenesis, which plays a key role in the conversion of cholesterol into hormones in humans, with evidence in octopuses of steroidogenesis activity in cholesterol metabolism during senescence [13].
5. Conclusions
Overall, our results illustrate a sophisticated interaction between environmental signals and neuroendocrine regulation in O. mimus. The optic lobe provides strong evidence of acting as a key integrator of photoperiodic information, likely as part of a complex with the optic gland, olfactory lobe, and supraesophageal structures, which were not included in the present analysis. The expression of genes related to image interpretation, retinal modulation, and neurochemical signaling suggests a dual sensory and modulatory function for this structure, which could cooperate with the optic gland, olfactory gland, or subesophageal lobes in the induction or inhibition of spawning onset. Moreover, in the oviducal gland, specific genes and neuropeptide receptors such as FMRFamide and myomodulin, previously described in the pedunculated system, were identified, indicating a potential functional connection between neurosensory centers and effector tissues. Collectively, this study highlights the dual role of the optic lobe in photoperiod interpretation and suggests a mechanism for spawning inhibition through changes in gene expression in the final effector of fertilization, the oviducal gland. The optic lobe becomes a crucial environmental sensor, integrating light signals and initiating a neuroendocrine cascade that culminates in spawning. However, future targeted studies involving other organs, such as the olfactory gland or optic gland, should be conducted at specific times during the pre- and post-spawning phase to assess how captivity conditions, population density, and reproductive behaviors influence these molecular pathways. Additionally, future research should examine how photoperiod and light intensity affect other physiological traits, including stress responses and nutritional status, to provide a more comprehensive understanding of female condition under aquaculture conditions. Yet, our results could be useful for implementing aquaculture for this species and incorporating biomarkers that allow for identifying the maturation periods of females and the start of spawning.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/aquacj6010007/s1. Figure S1. Distribution of genes for each female Octopus mimus subjected to different photoperiods (constant (24:0) vs. natural (14:10)) during the spawning season. Panels (A,B) show the analysis of principal components for the optic lobe and oviductal gland, respectively, while panels (C,D) represent the data through MDS (Multidimensional Scaling) analysis; Figure S2. Principal component analysis (PCA) of log-CPM values for optic lobe and oviductal gland samples under constant light and semi-dark treatments; Table S1. Statistics of RNA-seq data; Table S2. Functional enrichment of biological processes for the Octopus mimus females optic lobe and oviductal gland subjected to constant photoperiod contracted with the natural one. Molecular Function (GO:MF), Biological Process (GO:BP), and Cellular Component (GO:CC). Each entry lists the term name, unique identifier (term_id), and the adjusted p-value, summarizing the key functional aspects of the analyzed gene set.
Author Contributions
Conceptualization, C.Q.-P. and M.N.; Formal analysis, D.O., R.R.-P., A.G.d.S., L.F.C.C. and M.N.; Funding acquisition, C.Q.-P., F.W.D.-C., K.A.B.M.-S.-C., C.P.F.-C. and M.N.; Investigation, I.F., F.W.D.-C., K.A.B.M.-S.-C., A.G.d.S., L.F.C.C. and M.N.; Methodology, C.Q.-P., I.F., D.O., R.R.-P., F.W.D.-C., K.A.B.M.-S.-C., C.P.F.-C., A.G.d.S., L.F.C.C. and M.N.; Project administration, C.Q.-P., F.W.D.-C., K.A.B.M.-S.-C., C.P.F.-C., L.F.C.C. and M.N.; Resources, K.A.B.M.-S.-C., L.F.C.C. and M.N.; Supervision, K.A.B.M.-S.-C., C.P.F.-C., L.F.C.C. and M.N.; Technical support, F.E.T.-A., L.F.C.C. and M.N.; Validation, F.W.D.-C., A.G.d.S., L.F.C.C. and M.N.; Writing—review and editing, C.Q.-P., I.F., D.O., R.R.-P., F.W.D.-C., K.A.B.M.-S.-C., C.P.F.-C., A.G.d.S., L.F.C.C. and M.N. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by the Universidad Nacional Jorge Basadre Grohmann through the program “Fondos del canon, sobrecanon y regalías mineras”, as approved by Resolución Rectoral No. 11174-2023-UNJBG. Therojectt supported under this funding was “Plan de mejora biotecnológica del manejo de reproductores de pulpo (Octopus mimus), alimentación, control de la reproducción y calidad de las puestas”. Additional support was provided by the Fundação para a Ciência e a Tecnologia (FCT) for AGS under grant 2023.07625.CEECIND/CP2848/CT0010.
Institutional Review Board Statement
The experimental design was conducted in accordance with the principles of the 3Rs (Replacement, Reduction, and Refinement) as outlined by [49], employing the minimum number of animals necessary to achieve scientific validity. For all handling and sampling procedures, animals were first anesthetized with 1.5% magnesium chloride for 15 min and then immersed in a 3.5% solution for an additional 20 min. Subsequently, the vena cava was severed to ensure humane euthanasia prior to tissue collection. All procedures were reviewed and approved by the Ethics Committee of the Universidad Nacional Jorge Basadre Grohmann under registration number Version 1.1. (The protocol code is 2023-012-CEIUNJBG and the date of approval is 30 January 2023).
Data Availability Statement
The data presented in this study are openly available in the NCBI Sequence Read Archive (SRA) under BioProject accession number PRJNA1416070. Individual libraries are available under SRA accession numbers SRR37032714–SRR37032725.
Conflicts of Interest
The authors declare no competing interests (financial, professional, or personal relationships) that could influence the work reported here.
References
- Zúñiga, O.; Olivares, A.; Ossadón, L. Influencia de la luz en la maduración sexual de hembras Octopus mimus. Estud. Ocean. 1995, 14, 75–76. [Google Scholar]
- Jiménez, J.; Louis, V.; Yamashiro, C.; Kochzius, M. Genetics for Conservation and Fisheries: Population Structure and Connectivity of Octopus mimus Along the Peruvian Coast. In Book of Abstracts–VLIZ Marine Science Day; VLIZ Special Publication: Bredene, Belgium, 2019; Available online: https://www.vliz.be/imisdocs/publications/326178.pdf#page=129 (accessed on 15 November 2025).
- Cortez, T.; Castro, B.G.; Guerra, A. Reproduction and condition of female Octopus mimus (Mollusca: Cephalopoda). Mar. Biol. 1995, 123, 505–510. [Google Scholar] [CrossRef]
- Wodinsky, J. Hormonal inhibition of feeding and death in octopus: Control by the optic gland. Science 1977, 198, 948–951. [Google Scholar] [CrossRef] [PubMed]
- Wells, M.J.; Wells, J. Pituitary-like function of the octopus optic gland. J. Exp. Biol. 1959, 36, 1–33. [Google Scholar] [CrossRef]
- Sweedler, J.V.; Li, L.; Floyd, P.; Gilly, W. Mass spectrometric survey of peptides in cephalopods with an emphasis on the FMRFamide-related peptides. J. Exp. Biol. 2000, 203, 3565–3573. [Google Scholar] [CrossRef]
- Di Cosmo, A.; Di Cristo, C. GnRH and FMRFamide immunoreactivity in the olfactory lobe of Octopus vulgaris. J. Comp. Neurol. 1998, 398, 1–12. [Google Scholar] [CrossRef]
- Di Cristo, C. Nervous control of reproduction in Octopus vulgaris: A new model. Invertebr. Neurosci. 2013, 13, 27–34. [Google Scholar] [CrossRef]
- Iwakoshi-Ukena, E.; Ukena, K.; Takuwa-Kuroda, K.; Furukawa, K.; Oumi, T.; Fujita, T.; Minakata, H.; Nomoto, K.; Kawada, T.; Tsutsui, K. Isolation and characterization of a novel octopus GnRH-like peptide. Biochem. Biophys. Res. Commun. 2004, 318, 565–575. [Google Scholar]
- Zatylny-Gaudin, C.; Favrel, P. Diversity of the RFamide peptide family in mollusks. Front. Endocrinol. 2014, 5, 178. [Google Scholar] [CrossRef]
- Polese, G.; Bertapelle, C.; Di Cosmo, A. Role of olfactory organs in reproduction of Octopus vulgaris: New insights from electrical stimulation experiments. J. Exp. Biol. 2015, 218, 1740–1747. [Google Scholar]
- Wang, Z.Y.; Ragsdale, C.W. Multiple optic gland signaling pathways implicated in octopus maternal behaviors and death. J. Exp. Biol. 2018, 221, jeb185751. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Pergande, M.R.; Ragsdale, C.W.; Cologna, S.M. Steroid hormones of the octopus self-destruct system. Curr. Biol. 2022, 32, 2572–2579. [Google Scholar] [CrossRef]
- Juárez, O.E.; López-Galindo, L.; Pérez-Carrasco, L.; Lago-Lestón, A.; Rosas, C.; Di Cosmo, A.; Galindo-Sánchez, C.E. Octopus maya white body show sex-specific transcriptomic profiles during the reproductive phase, with high differentiation in signaling pathways. PLoS ONE 2019, 14, e0216982. [Google Scholar]
- Zúñiga, O.; Olivares, A.; Rosas, C. Octopus mimus. In Cephalopod Culture; Iglesias, J., Fuentes, L., Villanueva, R., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 397–413. [Google Scholar] [CrossRef]
- Sato, N. A review of sperm storage methods and post-copulatory sexual selection in the Cephalopoda. Biol. J. Linn. Soc. 2021, 134, 285–302. [Google Scholar] [CrossRef]
- Forsythe, J.W.; Hanlon, R.T. Aspects of egg development, post-hatching behavior, growth and reproductive biology of Octopus burryi Voss, 1950 (Mollusca: Cephalopoda). Vie Milieu 1985, 35, 273–282. [Google Scholar]
- Songco-Casey, J.O.; Coffing, G.C.; Piscopo, D.M.; Pungor, J.R.; Kern, A.D.; Miller, A.C.; Niell, C.M. Cell types and molecular architecture of the Octopus bimaculoides visual system. Curr. Biol. 2022, 32, 5031–5044.e4. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, D.E.; Johnsen, S.; Warrant, E. Cephalopod versus vertebrate eyes. Curr. Biol. 2023, 33, R1100–R1105. [Google Scholar] [CrossRef]
- Kingston, A.C.; Kuzirian, A.M.; Hanlon, R.T.; Cronin, T.W. Visual phototransduction components in cephalopod chromatophores suggest dermal photoreception. J. Exp. Biol. 2015, 218, 1596–1602. [Google Scholar] [CrossRef] [PubMed]
- McElroy, K.E.; Audino, J.A.; Serb, J.M. Molluscan genomes reveal extensive differences in photopigment evolution across the phylum. Mol. Biol. Evol. 2023, 40, msad263. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 15 November 2025).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible read trimming tool for Illumina NGS data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Song, L.; Florea, L. Rcorrector: Efficient and accurate error correction for Illumina RNA-seq reads. GigaScience 2015, 4, 48. [Google Scholar] [CrossRef]
- Stanke, M.; Morgenstern, B. AUGUSTUS: A web server for gene prediction in eukaryotes. Nucleic Acids Res. 2005, 33, W465–W467. [Google Scholar] [CrossRef] [PubMed]
- Dainat, J.; Hereñú, D.; Pucholt, P. AGAT: Another Gff Analysis Toolkit to Handle Annotations in any GTF/GFF Format. 2020. Available online: https://www.doi.org/10.5281/zenodo.3552717 (accessed on 15 November 2025).
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef]
- Babicki, S.; Arndt, D.; Marcu, A.; Liang, Y.; Grant, J.R.; Maciejewski, A.; Wishart, D.S. Heatmapper: Web-enabled heat mapping for all. Nucleic Acids Res. 2016, 44, W147–W153. [Google Scholar] [CrossRef] [PubMed]
- Blighe, K.; Rana, S.; Lewis, M. EnhancedVolcano: Publication-Ready Volcano Plots with Enhanced Colouring and Labeling, version 1.2.0; R Package; 2019. Available online: https://bioconductor.org/packages/devel/bioc/vignettes/EnhancedVolcano/inst/doc/EnhancedVolcano.html (accessed on 15 November 2025).
- Blighe, K.; Lewis, M.; Lun, A.; Blighe, M.K. PCAtools: Everything Principal Components Analysis. R Package, Version 2.0.0. 2019. Available online: https://github.com/kevinblighe/PCAtools (accessed on 15 November 2025).
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019, 47, W191–W198. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Sato, Y.; Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 2016, 428, 726–731. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Zar, J.H. Biostatistical Analysis, 4th ed.; Prentice Hall: Upper Saddle River, NJ, USA, 1999. [Google Scholar]
- Cao, H.M.; Wu, J.H.; Li, S.; Zhou, X.; Zheng, L.B.; Chi, C.F. A Na+ channel receptor of FMRFamide in the cephalopod Sepiella japonica: Identification, characterization, and expression profiling during different stages of gonadal development. Neuropeptides 2024, 106, 102437. [Google Scholar] [CrossRef]
- Domínguez-Estrada, A.; Galindo-Sánchez, C.E.; Ventura-López, C.; Rosas, C.; Juárez, O.E. Response of optic gland pathways to thermal stress in the reproductive phase of female Octopus maya. J. Molluscan Stud. 2022, 88, eyac018. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Ragsdale, C.W. Signaling ligand heterogeneities in the peduncle complex of the cephalopod mollusc Octopus bimaculoides. Brain Behav. Evol. 2024, 99, 158–170. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Chen, S.; Zhang, T.; Pan, L.; Liu, C.; Bian, L. Gonadal transcriptome sequencing analysis reveals the candidate sex-related genes and signaling pathways in the East Asian common octopus, Octopus sinensis. Genes 2024, 15, 682. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Wang, H.; Liu, C.; Chen, S.; Zhao, Q.; Ge, J.; Liu, Y. Screening of sex-special genes in common long-arm octopus Octopus minor based on gonadal transcriptome sequencing. Isr. J. Aquac.-Bamidgeh 2024, 76, 173–184. [Google Scholar] [CrossRef]
- Croll, R.P.; Wang, C. Possible Roles of Sex Steroids in the Control of Reproduction in Bivalve Molluscs. Aquaculture 2007, 272, 76–86. [Google Scholar] [CrossRef]
- Arreola-Meráz, E. Reabsorption of oocytes and components of accessory glands under reproductive inhibition conditions in cephalopods. Mar. Biol. 2018, 165, 1–13. [Google Scholar]
- Budelmann, B.U. Autophagy in octopus. S. Afr. J. Mar. Sci. 1998, 20, 101–108. [Google Scholar] [CrossRef]
- Huang, W.; Xu, F.; Qu, T.; Zhang, R.; Li, L.; Que, H.; Zhang, G. Identification of thyroid hormones and functional characterization of thyroid hormone receptor in the pacific oyster Crassostrea gigas provide insight into evolution of the thyroid hormone system. PLoS ONE 2015, 10, e0144991. [Google Scholar] [CrossRef] [PubMed]
- Fiorito, G.; Affuso, A.; Anderson, D.B.; Basil, J.; Bonnaud, L.; Botta, G.; Andrews, P. Cephalopods in neuroscience: Regulations, research and the 3Rs. Invertebr. Neurosci. 2014, 14, 13–36. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.