Scale-Up of the Fermentation Process for the Production and Purification of Serratiopeptidase Using Silkworm Pupae as a Substrate
Abstract
1. Introduction
2. Materials and Methods
2.1. Initial Culture Conditions
2.2. Proteolytic Activity and Total Protein Assay
2.3. Plackett-Burman and Taguchi Designs
2.4. The Fermentation Process Was Scaled Up
2.5. Optimization of Enzyme Purification
3. Results
3.1. Plackett–Burman Design
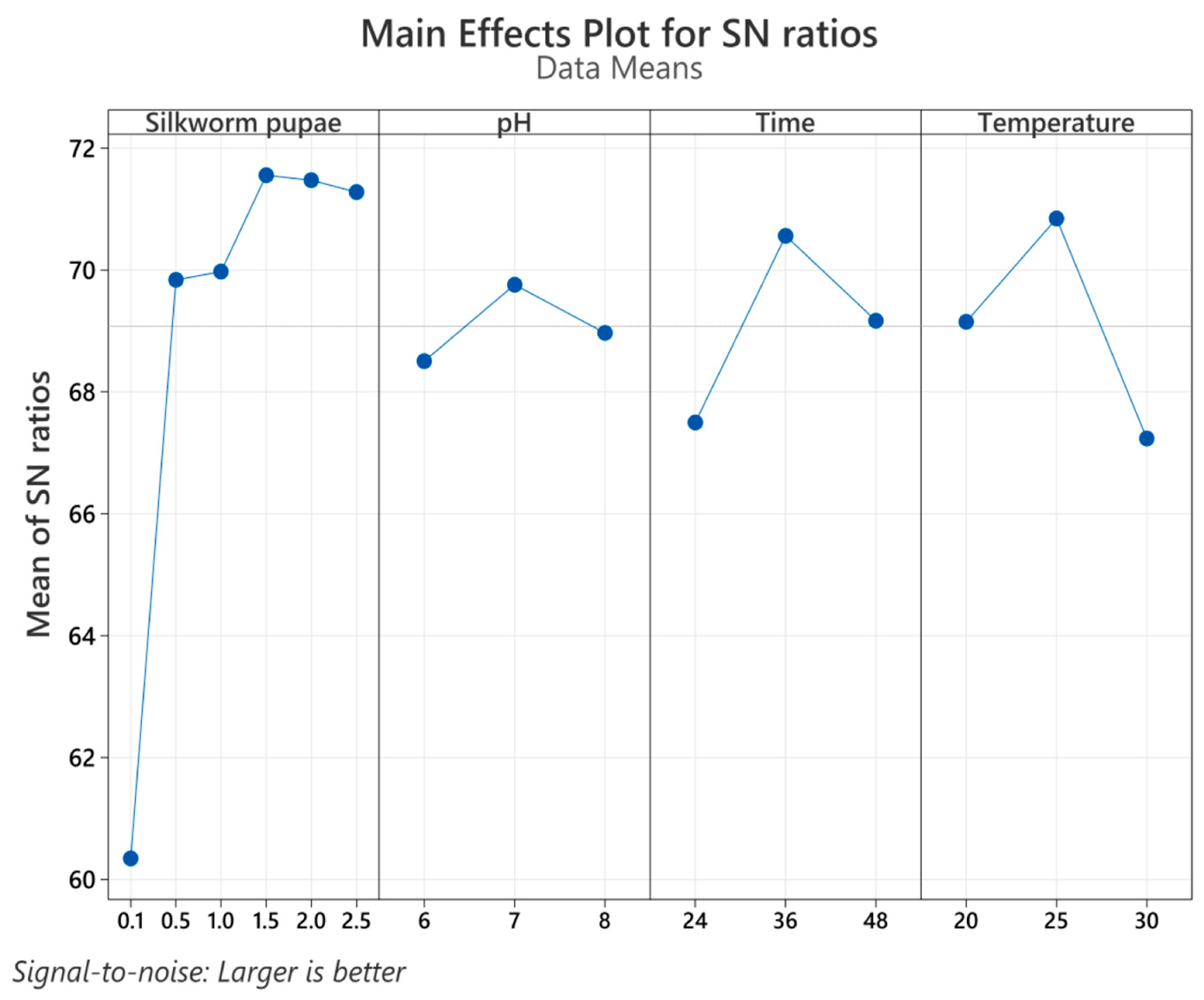
3.2. Taguchi Design
3.3. The Fermentation Process Was Scaled Up
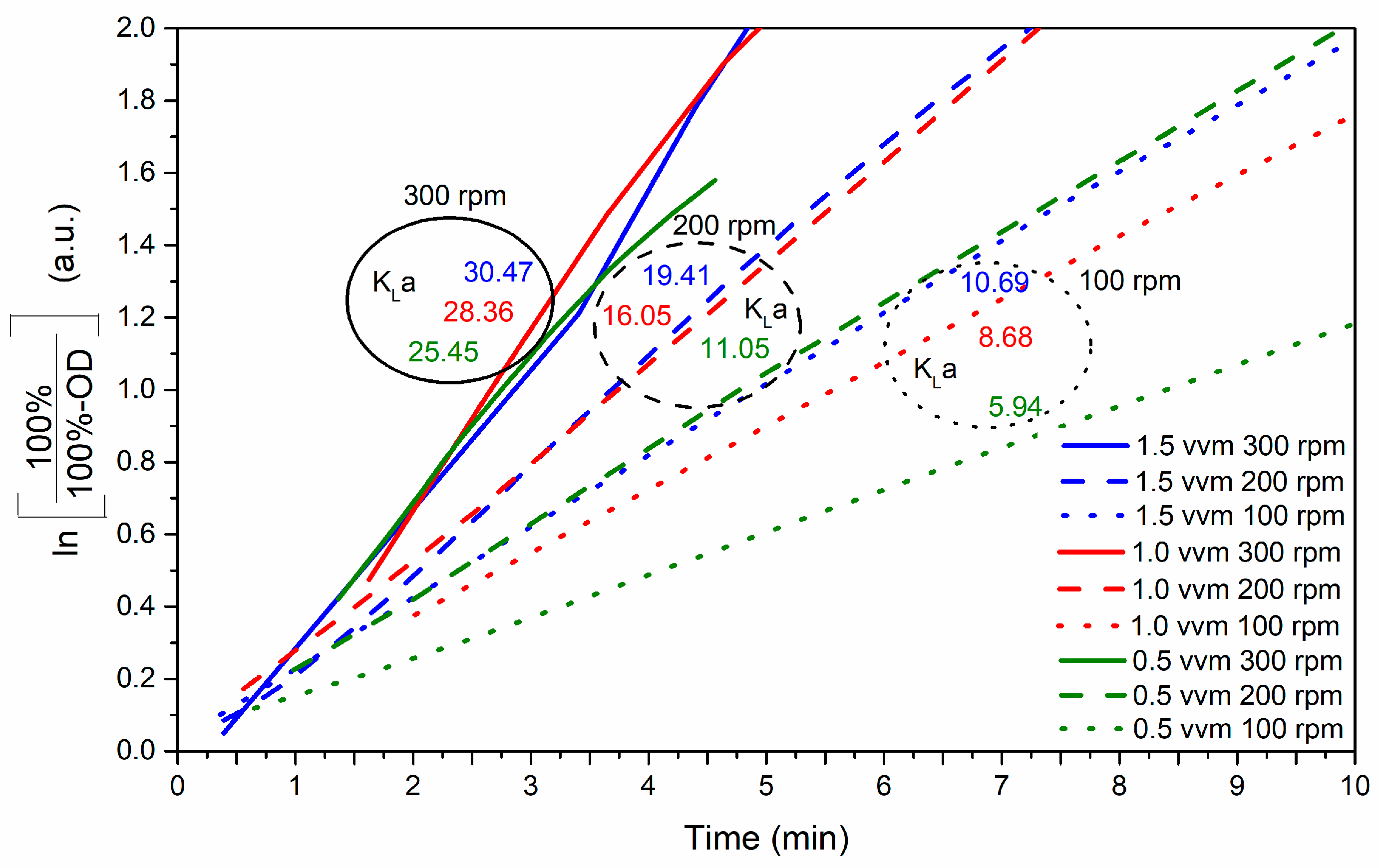
3.3.1. Determination of the Volumetric Oxygen Transfer Coefficient (kLa)
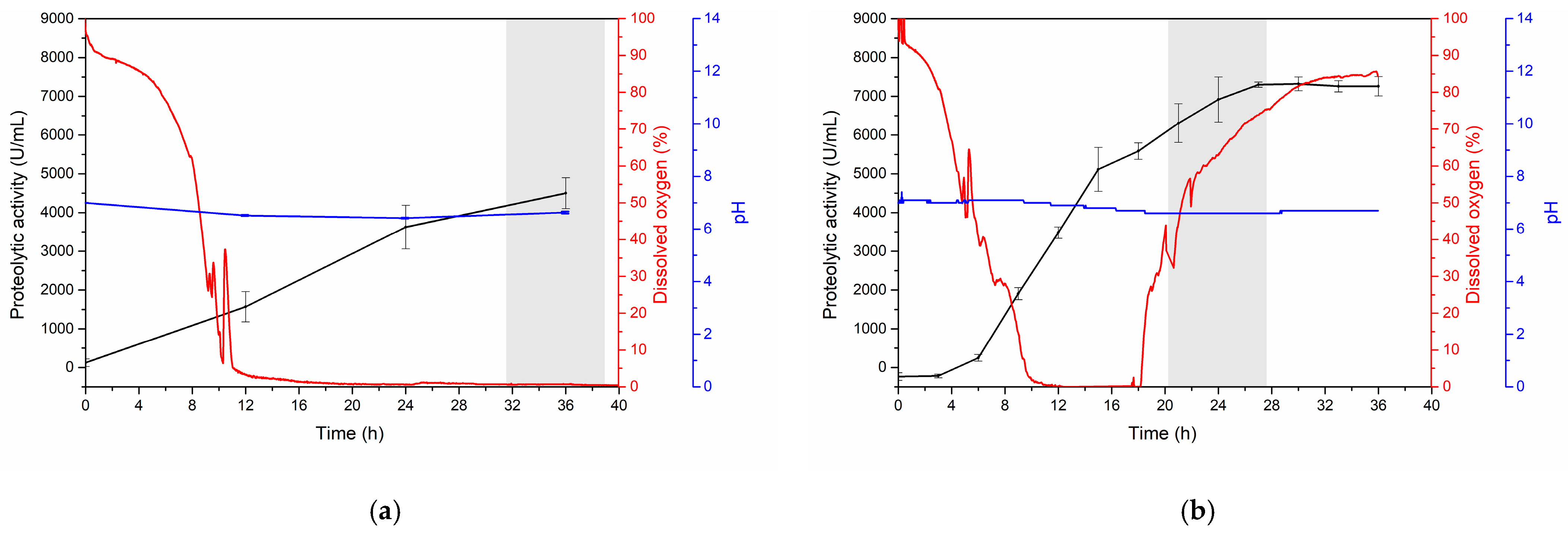
3.3.2. Kinetics of Serratiopeptidase Production at the Bioreactor Level
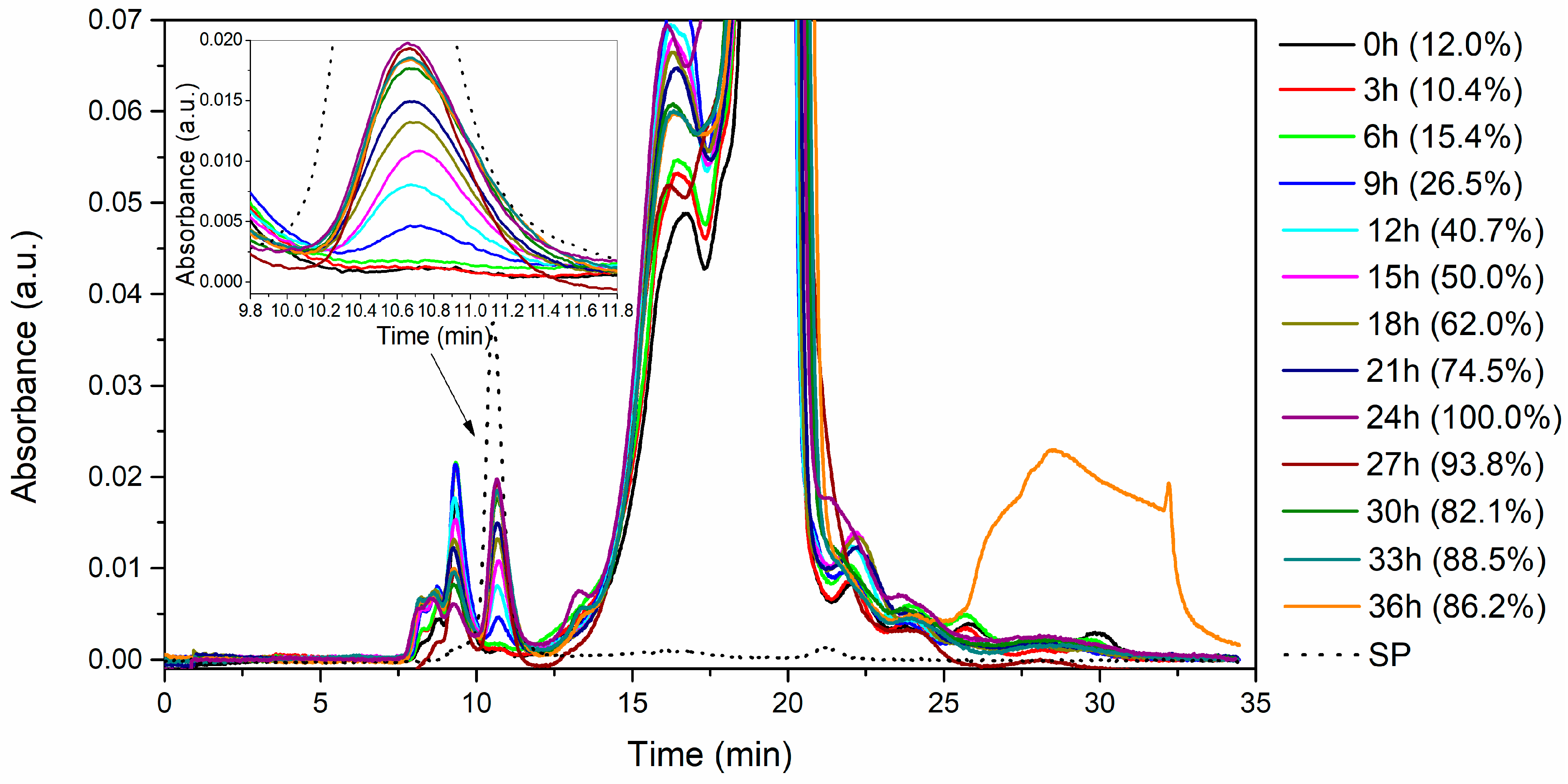
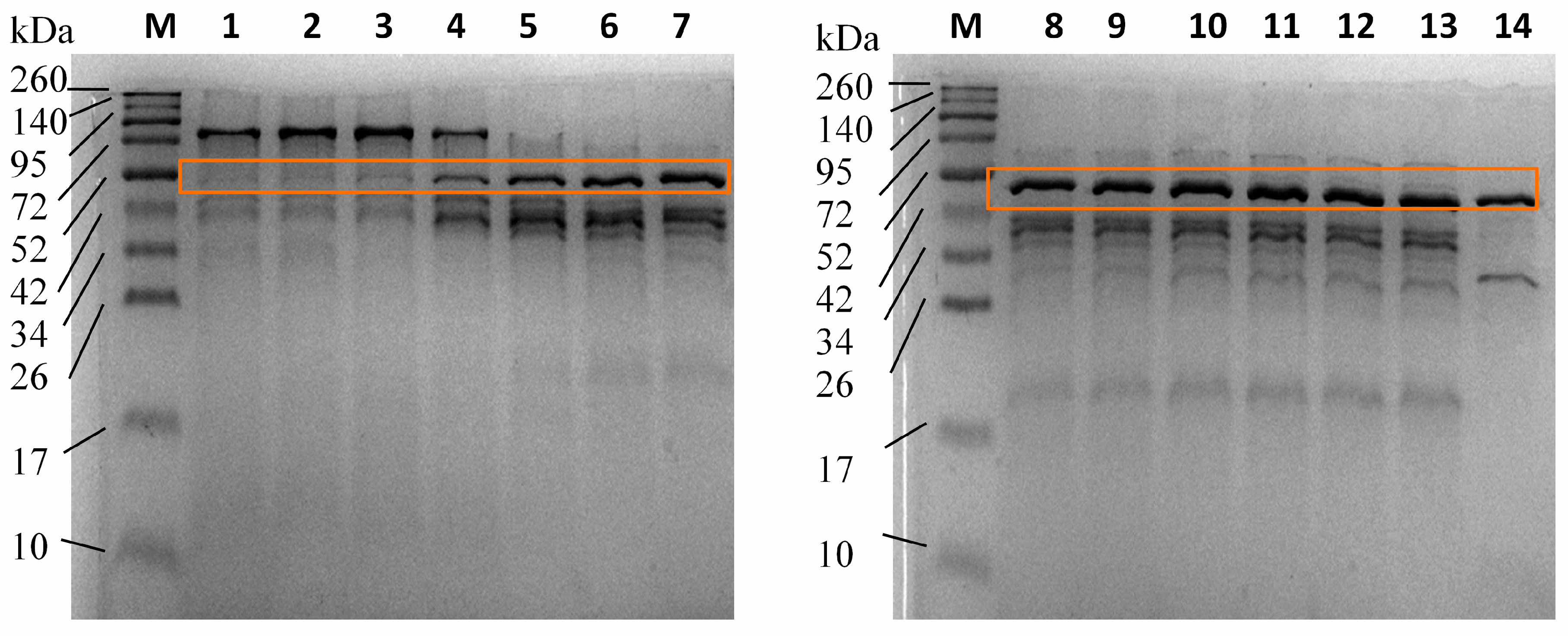
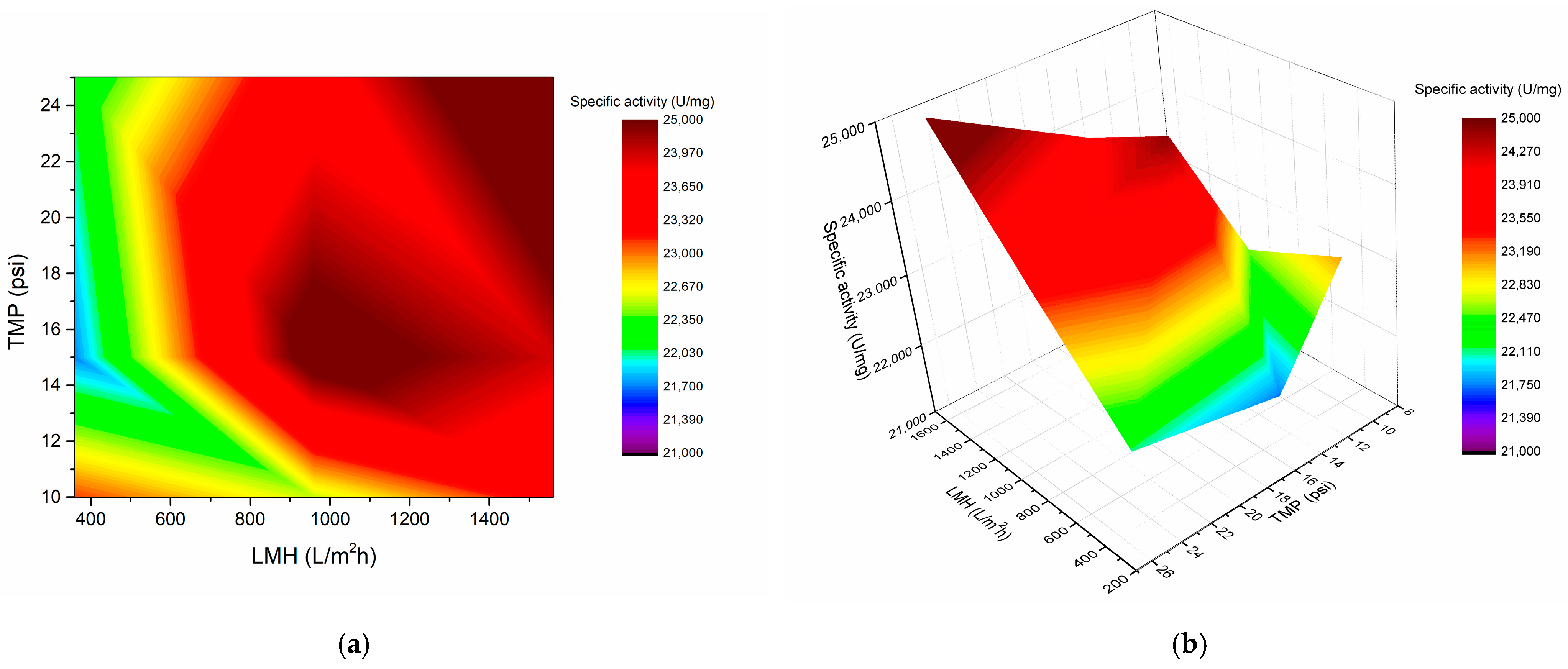
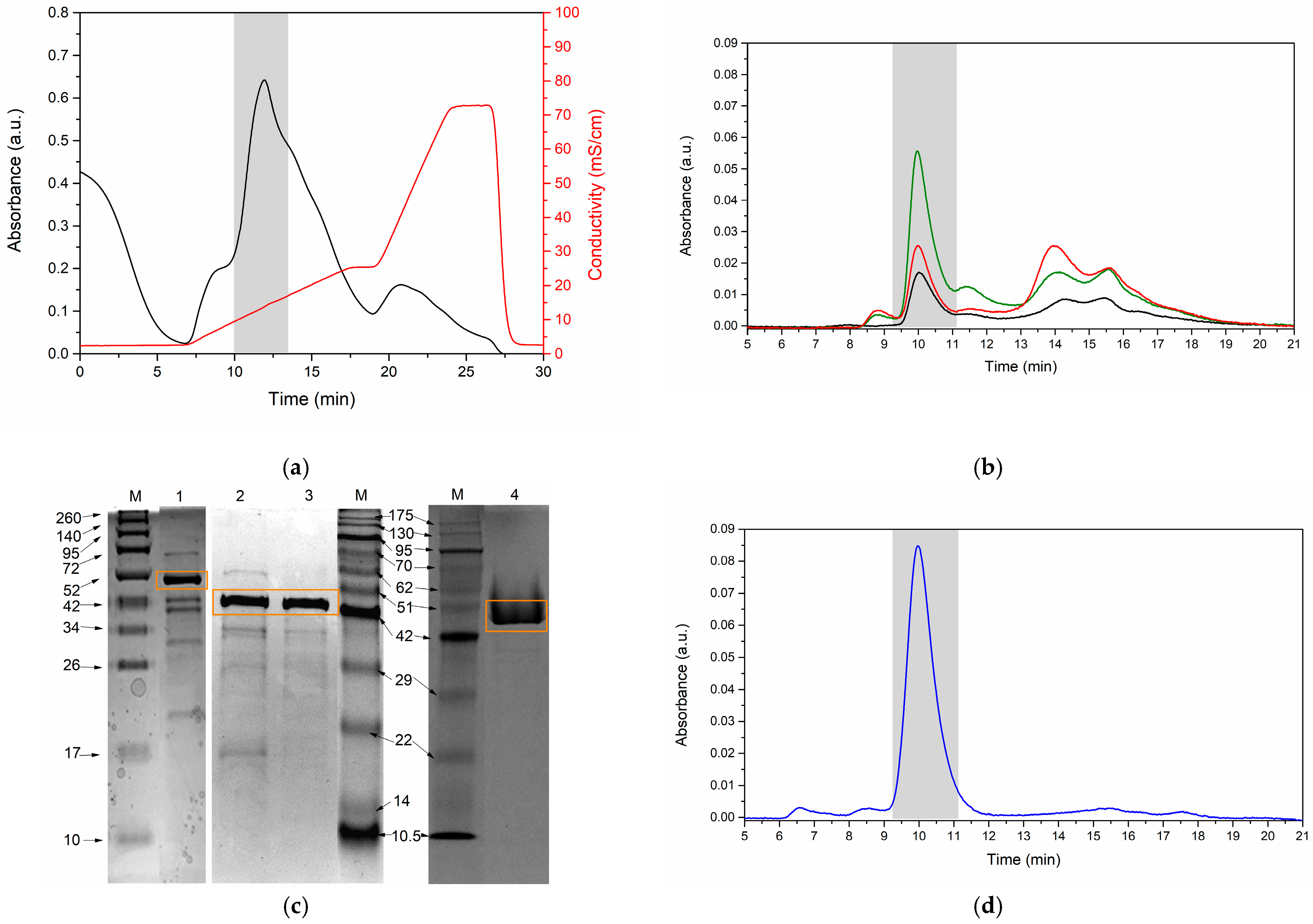
3.4. Enzyme Purification
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tarafdar, A.; Sirohi, R.; Gaur, V.K.; Kumar, S.; Sharma, P.; Varjani, S.; Pandey, H.O.; Sindhu, R.; Madhavan, A.; Rajasekharan, R.; et al. Engineering Interventions in Enzyme Production: Lab to Industrial Scale. Bioresour. Technol. 2021, 326, 124771. [Google Scholar] [CrossRef]
- Kumar, D.; Dev, P.; Kumar, V.R. Biomedical Applications of Natural Proteins: An Emerging Era in Biomedical Sciences; Kumar, D., Dev, P., Kumar, V.R., Eds.; Springer: New Delhi, India, 2015. [Google Scholar]
- Delepelaire, P.; Wandersman, C. Protein Secretion in Gram-Negative Bacteria. J. Biol. Chem. 1990, 265, 17118–17125. [Google Scholar] [CrossRef]
- Al-Dhabi, N.A.; Esmail, G.A.; Ghilan, A.K.M.; Arasu, M.V. Isolation and Screening of Streptomyces sp. Al-Dhabi-49 from the Environment of Saudi Arabia with Concomitant Production of Lipase and Protease in Submerged Fermentation. Saudi. J. Biol. Sci. 2020, 27, 474–479. [Google Scholar] [CrossRef]
- Dhiman, S.; Mukherjee, G. Application of OFAT Approach for Optimizing Tannase Production under Submerged Fermentation. Mater. Today Proc. 2023. [Google Scholar] [CrossRef]
- Singh, V.; Haque, S.; Niwas, R.; Srivastava, A.; Pasupuleti, M.; Tripathi, C.K.M. Strategies for Fermentation Medium Optimization: An in-Depth Review. Front. Microbiol. 2017, 7, 2087. [Google Scholar] [CrossRef]
- Bhargavi, P.L.; Prakasham, R.S. Agro-Industrial Wastes Utilization for the Generation of Fibrinolytic Metalloprotease by Serratia marcescens RSPB11. Biocatal. Agric Biotechnol. 2017, 9, 201–208. [Google Scholar] [CrossRef]
- Ghorbel-Bellaaj, O.; Hmidet, N.; Jellouli, K.; Younes, I.; Maâlej, H.; Hachicha, R.; Nasri, M. Shrimp Waste Fermentation with Pseudomonas aeruginosa A2: Optimization of Chitin Extraction Conditions through Plackett-Burman and Response Surface Methodology Approaches. Int. J. Biol. Macromol. 2011, 48, 596–602. [Google Scholar] [CrossRef] [PubMed]
- Plackett, R.L.; Burman, J.P. The Design of Optimum Multifactorial Experiments. Biometrika 1946, 33, 305–325. [Google Scholar] [CrossRef]
- Majumdar, S.; Mandal, T.; Mandal, D.D. Chitosan Based Micro and Nano-Particulate Delivery Systems for Bacterial Prodigiosin: Optimization and Toxicity in Animal Model System. Int. J. Biol. Macromol. 2022, 222, 2966–2976. [Google Scholar] [CrossRef] [PubMed]
- Prakasham, R.S.; Rao, C.S.; Rao, R.S.; Rajesham, S.; Sarma, A.P.N. Optimization of Alkaline Protease Production by Bacillus sp. Using Taguchi Methodology. Appl. Biochem. Biotechnol. 2005, 120, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Mechri, S.; Bouacem, K.; Chalbi, T.; Khaled, M.; Allala, F.; Bouanane-Darenfed, A.; Hacene, H.; Jaouadi, B. A Taguchi Design Approach for the Enhancement of a Detergent-biocompatible Alkaline Thermostable Protease Production by Streptomyces mutabilis Strain TN-X30. J. Surfactants Deterg. 2022, 25, 487–504. [Google Scholar] [CrossRef]
- Tripathi, N.K.; Shrivastava, A. Recent Developments in Bioprocessing of Recombinant Proteins: Expression Hosts and Process Development. Front. Bioeng. Biotechnol. 2019, 7, 420. [Google Scholar] [CrossRef] [PubMed]
- Sosa-Martínez, J.D.; Montañez, J.; Contreras-Esquivel, J.C.; Balagurusamy, N.; Gadi, S.K.; Morales-Oyervides, L. Agroindustrial and Food Processing Residues Valorization for Solid-State Fermentation Processes: A Case for Optimizing the Co-Production of Hydrolytic Enzymes. J. Environ. Manag. 2023, 347, 119067. [Google Scholar] [CrossRef]
- Shin, W.-S.; Lee, D.; Kim, S.; Jeong, Y.-S. Application of Scale-Up Criterion of Constant Oxygen Mass Transfer Coefficient (KLa) for Production of Itaconic Acid in a 50 L Pilot-Scale Fermentor by Fungal Cells of Aspergillus Terreus. J. Microbiol. Biotechnol. 2013, 23, 1445–1453. [Google Scholar] [CrossRef]
- Madeira, J.V.; Contesini, F.J.; Calzado, F.; Rubio, M.V.; Zubieta, M.P.; Lopes, D.B.; Rodrigues, R. Agro-Industrial Residues and Microbial Enzymes: An Overview on the Eco-Friendly Bioconversion into High Value-Added Products. In Biotechnology of Microbial Enzymes; Elsevier Inc.: Chennai, India, 2017; pp. 476–481. [Google Scholar]
- Pakhale, S.V.; Bhagwat, S.S. Purification of Serratiopeptidase from Serratia marcescens NRRL B 23112 Using Ultrasound Assisted Three Phase Partitioning. Ultrason. Sonochem. 2016, 31, 532–538. [Google Scholar] [CrossRef]
- Wu, D.; Ran, T.; Wang, W.; Xu, D. Structure of a Thermostable Serralysin from Serratia sp. FS14 at 1.1 A ° Resolution. Acta Cryst. 2016, F72, 10–15. [Google Scholar]
- Nair, S.R.; Subathra Devi, C. Serratiopeptidase: An Integrated View of Multifaceted Therapeutic Enzyme. Biomolecules 2022, 12, 1468. [Google Scholar] [CrossRef]
- Bhagat, S.; Agarwal, M.; Roy, V. Serratiopeptidase: A Systematic Review of the Existing Evidence. Int. J. Surg. 2013, 11, 209–217. [Google Scholar] [CrossRef]
- Meng, Y.; Yang, M.; Liu, W.; Li, J. Cell-Free Expression of a Therapeutic Protein Serratiopeptidase. Molecules 2023, 28, 3132. [Google Scholar] [CrossRef]
- Rouhani, M.; Valizadeh, V.; Bakhshandeh, H.; Hosseinzadeh, S.A.; Molasalehi, S.; Atyabi, S.M.; Norouzian, D. Improved Anti-Biofilm Activity and Long-Lasting Effects of Novel Serratiopeptidase Immobilized on Cellulose Nanofibers. Appl. Microbiol. Biotechnol. 2023, 107, 6487–6496. [Google Scholar] [CrossRef]
- Yadav, V.; Sharma, S.; Kumar, A.; Singh, S.; Ravichandiran, V. Serratiopeptidase Attenuates Lipopolysaccharide-Induced Vascular Inflammation by Inhibiting the Expression of Monocyte Chemoattractant Protein-1. Curr. Issues Mol. Biol. 2023, 45, 2201–2212. [Google Scholar] [CrossRef] [PubMed]
- Artini, M.; Vrenna, G.; Trecca, M.; Tuccio Guarna Assanti, V.; Fiscarelli, E.V.; Papa, R.; Selan, L. Serratiopeptidase Affects the Physiology of Pseudomonas aeruginosa Isolates from Cystic Fibrosis Patients. Int. J. Mol. Sci. 2022, 23, 12645. [Google Scholar] [CrossRef] [PubMed]
- Jadhav, S.B.; Shah, N.; Rathi, A.; Rathi, V.; Rathi, A. Serratiopeptidase: Insights into the Therapeutic Applications. Biotechnol. Rep. 2020, 28, e00544. [Google Scholar] [CrossRef]
- Vélez-Gómez, J.M.; Melchor-Moncada, J.J.; Veloza, L.A.; Sepúlveda-Arias, J.C. Purification and Characterization of a Metalloprotease Produced by the C8 Isolate of Serratia marcescens Using Silkworm Pupae or Casein as a Protein Source. Int. J. Biol. Macromol. 2019, 135, 97–105. [Google Scholar] [CrossRef]
- Luthra, U.; Babu, P.; Patel, Y.; Ramesh, J.V.; Sharma, M.; Majeed, I.; Subbiah, S.K.; Pandiyan, R. Serratiopeptidase: A Statistical Approach toward Enhancement of Fermentation and Biomass Product Recovery. Biomass Convers. Biorefin. 2022, 1, 1–8. [Google Scholar] [CrossRef]
- Sriram, K.; Uma Maheswari, P.; Meera Sheriffa Begum, K.M.; Arthanareeswaran, G. Functionalized Chitosan with Super Paramagnetic Hybrid Nanocarrier for Targeted Drug Delivery of Curcumin. Iran. Polym. J. (Engl. Ed.) 2018, 27, 469–482. [Google Scholar] [CrossRef]
- Michelin, M.; de Oliveira Mota, A.M.; de Moraes Polizeli, M.d.L.T.; da Silva, D.P.; Vicente, A.A.; Teixeira, J.A. Influence of Volumetric Oxygen Transfer Coefficient (KLa) on Xylanases Batch Production by Aspergillus niger van Tieghem in Stirred Tank and Internal-Loop Airlift Bioreactors. Biochem. Eng. J. 2013, 80, 19–26. [Google Scholar] [CrossRef]
- Casarin, F.; Cladera-Olivera, F.; Brandelli, A. Use of Poultry Byproduct for Production of Keratinolytic Enzymes. Food Bioproc. Technol. 2008, 1, 301–305. [Google Scholar] [CrossRef]
- Vachher, M.; Sen, A.; Kapila, R.; Nigam, A. Microbial Therapeutic Enzymes: A Promising Area of Biopharmaceuticals. Curr. Res. Biotechnol. 2021, 3, 195–208. [Google Scholar] [CrossRef]
- El-Abd, M.A.; Ibrahim, E.A. Production and One-Step Purification of Serratiopeptidase Enzyme from Serratia marcescens with Potent Anti-Inflammatory and Antioxidant Power. Egypt. Pharm. J. 2020, 19, 238–243. [Google Scholar] [CrossRef]
- Zhu, L.; Xu, B.; Wu, X.; Lei, J.; Hacker, D.L.; Liang, X.; Wurm, F.M. Analysis of Volumetric Mass Transfer Coefficient (KLa) in Small- (250 mL) to Large-Scale (2500 L) Orbitally Shaken Bioreactors. 3 Biotech 2020, 10, 397. [Google Scholar] [CrossRef]
- Pansuriya, R.C.; Singhal, R.S. Effects of Dissolved Oxygen and Agitation on Production of Serratiopeptidase by Serratia marcescens NRRL B-23112 in Stirred Tank Bioreactor and Its Kinetic Modeling. J. Microbiol. Biotechnol. 2011, 21, 430–437. [Google Scholar] [CrossRef]
- Lv, P.J.; Qiang, S.; Liu, L.; Hu, C.Y.; Meng, Y.H. Dissolved-Oxygen Feedback Control Fermentation for Enhancing β-Carotene in Engineered Yarrowia lipolytica. Sci. Rep. 2020, 10, 17114. [Google Scholar] [CrossRef]
- Li, Z.; Zhao, S.; Xin, X.; Zhang, B.; Thomas, A.; Charles, A.; Lee, K.S.; Jin, B.R.; Gui, Z. Purification and Characterization of a Novel Immunomodulatory Hexapeptide from Alcalase Hydrolysate of Ultramicro-Pretreated Silkworm (Bombyx mori) Pupa Protein. J. Asia Pac. Entomol. 2019, 22, 633–637. [Google Scholar] [CrossRef]
- Dhillon, G.S.; Kaur, S. Industrial Enzymes: Recovery and Purification Challenges. In Agro-Industrial Wastes as Feedstock for Enzyme Production; Nikki Levy: Cambridge, MA, USA, 2016; pp. 110–125. ISBN 978-0-12-802392-1. [Google Scholar]
- Sharma, C.; Jha, N.K.; Meeran, M.F.N.; Patil, C.R.; Goyal, S.N.; Ojha, S. Serratiopeptidase, A Serine Protease Anti-Inflammatory, Fibrinolytic, and Mucolytic Drug, Can Be a Useful Adjuvant for Management in COVID-19. Front Pharmacol. 2021, 12, 603997. [Google Scholar] [CrossRef]
- Katsipis, G.; Avgoulas, D.I.; Geromichalos, G.D.; Petala, M.; Pantazaki, A.A. In Vitro and in Silico Evaluation of the Serrapeptase Effect on Biofilm and Amyloids of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2023, 107, 7269–7285. [Google Scholar] [CrossRef]
- Mei, J.F.; Cai, S.F.; Yi, Y.; Wang, X.D.; Ying, G.Q. Study of the Fibrinolytic Activity of Serrapeptase and Its in Vitro Thrombolytic Effects. Braz. J. Pharm. Sci. 2022, 58, 1–9. [Google Scholar] [CrossRef]
- Kumar, D.; Verma, D.; Abbot, V. A Review on Pharmaceutical, Pharmacological and Chemical Aspects of Serratiopeptidase as Anti-Inflammatory Agent. Mater. Today Proc. 2023. [Google Scholar] [CrossRef]
- Yeruva, T.; Jayaram, H.; Aurade, R.; Shunmugam, M.M.; Shinde, V.S.; Baragi Venkatesharao, S.R.; Azhiyakathu, M.J. Profiling of Nutrients and Bioactive Compounds in the Pupae of Silkworm, Bombyx mori. Food Chem. Adv. 2023, 3, 100382. [Google Scholar] [CrossRef]
- Patil, S.R.; Amena, S.; Vikas, A.; Rahul, P.; Jagadeesh, K.; Praveen, K. Utilization of Silkworm Litter and Pupal Waste-an Eco-Friendly Approach for Mass Production of Bacillus thuringiensis. Bioresour. Technol. 2013, 131, 545–547. [Google Scholar] [CrossRef]
- Prakasham, R.S.; Rao, C.S.; Sarma, P.N. Green Gram Husk-an Inexpensive Substrate for Alkaline Protease Production by Bacillus sp. in Solid-State Fermentation. Bioresour. Technol. 2006, 97, 1449–1454. [Google Scholar] [CrossRef]
- Moula Ali, A.M.; Bavisetty, S.C.B. Purification, Physicochemical Properties, and Statistical Optimization of Fibrinolytic Enzymes Especially from Fermented Foods: A Comprehensive Review. Int. J. Biol. Macromol. 2020, 163, 1498–1517. [Google Scholar] [CrossRef]
- Srimathi, U.; Virivinti, N. DFT-Guided Sustainable Extractive Fermentation and Partial Purification of Fibrinolytic Protease from Agro-Industrial Waste. Sustain. Chem. Pharm. 2024, 37, 101446. [Google Scholar] [CrossRef]
- Fahmy, N.M.; El-Deeb, B. Optimization, Partial Purification, and Characterization of a Novel High Molecular Weight Alkaline Protease Produced by Halobacillus sp. HAL1 Using Fish Wastes as a Substrate. J. Genet. Eng. Biotechnol. 2023, 21, 48. [Google Scholar] [CrossRef] [PubMed]
- Singh Dhillon, G.; Kaur, S. Apply and Exploit the Emerging and Valuable Use Options of Waste Biomass. In Agro-Industrial Wastes as Feedstock for Enzyme Production, 1st ed.; Singh Dhillon, G., Kaur, S., Eds.; Nikki Levy: Cambridge, MA, USA, 2016; ISBN 978-0-12-802392-1. [Google Scholar]
- Pansuriya, R.C.; Singhal, R.S. Evolutionary Operation (EVOP) to Optimize Whey-Independent Serratiopeptidase Production from Serratia marcescens NRRL B-23112. J. Microbiol. Biotechnol. 2010, 20, 950–957. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.M.; Kumar, R.; Panwar, S.; Kumar, A. Microbial Alkaline Proteases: Optimization of Production Parameters and Their Properties. J. Genet. Eng. Biotechnol. 2017, 15, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Bach, E.; Sant’Anna, V.; Daroit, D.J.; Corrêa, A.P.F.; Segalin, J.; Brandelli, A. Production, One-Step Purification, and Characterization of a Keratinolytic Protease from Serratia marcescens P3. Process Biochem. 2012, 47, 2455–2462. [Google Scholar] [CrossRef][Green Version]
- Gaviria, G.Y.S.; Zapata, M.J.E. Optimization and Scale up of the Enzymatic Hydrolysis of Californian Red Worm Protein (Eisenia foetida). Heliyon 2023, 9, e16165. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.; Roy, S. A Comparative Study on Diauxic Growth of Microorganisms of Pseudomonas Species. Mater. Today Proc 2023, 72, 471–476. [Google Scholar] [CrossRef]
- Chander, D.; Khousla, J.K.; Koul, D.; Hossain, M.M.; Dar, M.J.; Chaubey, A. Purification and Characterization of Thermoactive Serratiopeptidase from Serratia marcescens AD-W2. AMB Express 2021, 11, 53. [Google Scholar] [CrossRef]
- Doshi, P.; Bhargava, P.; Singh, V.; Pathak, C.; Joshi, C.; Joshi, M. Escherichia coli Strain Engineering for Enhanced Production of Serratiopeptidase for Therapeutic Applications. Int. J. Biol. Macromol. 2020, 160, 1050–1060. [Google Scholar] [CrossRef] [PubMed]
- Doshi, P.; Dantroliya, S.; Modi, A.; Shukla, A.; Patel, D.; Joshi, C.; Joshi, M. Enhanced Production Process of Recombinant Mature Serratiopeptidase in Escherichia coli Using Fed-Batch Culture by Self-Proteolytic Activity of Fusion Protein. Fermentation 2022, 8, 307. [Google Scholar] [CrossRef]
- Koul, D.; Chander, D.; Manhas, R.S.; Chaubey, A. Isolation and Characterization of Serratiopeptidase Producing Bacteria from Mulberry Phyllosphere. Curr. Microbiol. 2021, 78, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, V.; Mishra, S.; Chaudhuri, T.K. Enhanced Production of Recombinant Serratiopeptidase in Escherichia coli and Its Characterization as a Potential Biosimilar to Native Biotherapeutic Counterpart. Microb Cell Fact. 2019, 18, 215. [Google Scholar] [CrossRef]
- Rouhani, M.; Valizadeh, V.; Molasalehi, S.; No-Rouzian, D. Production and Expression Optimization of Heterologous Serra-Tiopeptidase. Iran. J. Public Health 2020, 49, 931. [Google Scholar]
- Nageswara, S.; Guntuku, G.; Yakkali, B.L. Purification, Characterization, and Structural Elucidation of Serralysin-like Alkaline Metalloprotease from a Novel Source. J. Genet. Eng. Biotechnol. 2019, 17, 1. [Google Scholar] [CrossRef]

| Factor | Silkworm Pupae (%) | Casein (%) | Soy Oil (%) | (NH4)2HPO4 (%) | ZnCl2 (%) | CaCl2·2H2O (%) |
|---|---|---|---|---|---|---|
| High | 2.50 | 2.50 | 2.00 | 2.00 | 0.20 | 0.20 |
| Low | 0.00 | 0.10 | 0.10 | 0.50 | 0.01 | 0.01 |
| Trial Number * | pH | Temperature (°C) | Time (h) | Silkworm Pupae (%w/v) |
|---|---|---|---|---|
| 1 | 6 | 20 | 24 | 0.1 |
| 2 | 7 | 25 | 36 | 0.1 |
| 3 | 8 | 30 | 48 | 0.1 |
| 4 | 6 | 25 | 24 | 0.5 |
| 5 | 7 | 30 | 36 | 0.5 |
| 6 | 8 | 20 | 48 | 0.5 |
| 7 | 6 | 20 | 36 | 1.0 |
| 8 | 7 | 25 | 48 | 1.0 |
| 9 | 8 | 30 | 24 | 1.0 |
| 10 | 6 | 30 | 48 | 1.5 |
| 11 | 7 | 20 | 24 | 1.5 |
| 12 | 8 | 25 | 36 | 1.5 |
| 13 | 6 | 30 | 36 | 2.0 |
| 14 | 7 | 20 | 48 | 2.0 |
| 15 | 8 | 25 | 24 | 2.0 |
| 16 | 6 | 25 | 48 | 2.5 |
| 17 | 7 | 30 | 24 | 2.5 |
| 18 | 8 | 20 | 36 | 2.5 |
| Sample | Protein (mg) | Specific Activity (U/mg) | Total Activity (U) | Recovery (%) | Purification Fold |
|---|---|---|---|---|---|
| Crude extract | 70.09 ± 2.77 | 22,983.32 ± 1561.35 | 1,611,728.39 ± 142,346.87 | 100.00 | 1.00 |
| Ultrafiltration TFF 10 kDa | 63.79 ± 1.53 | 24,325.81 ± 1515.69 | 1,550,370.37 ± 67,709.37 | 96.60 ± 8.15 | 1.06 ± 0.07 |
| Diafiltration TFF 10 kDa | 33.33 ± 1.30 | 27,758.61 ± 879.48 | 924,333.33 ± 6555.55 | 57.62 ± 4.50 | 1.21 ± 0.11 |
| Strong anion exchange | 31.94 ± 0.61 | 27,269.88 ± 1845.97 | 870,296.30 ± 44,954.06 | 54.42 ± 7.21 | 1.19 ± 0.15 |
| Ultrafiltration 10 kDa | 8.86 ± 0.61 | 36,152.12 ± 1708.81 | 318,474.07 ± 23,735.51 | 19.94 ± 3.07 | 1.59 ± 0.31 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Melchor-Moncada, J.J.; García-Barco, A.; Zuluaga-Vélez, A.; Veloza, L.A.; Sepúlveda-Arias, J.C. Scale-Up of the Fermentation Process for the Production and Purification of Serratiopeptidase Using Silkworm Pupae as a Substrate. Methods Protoc. 2024, 7, 19. https://doi.org/10.3390/mps7020019
Melchor-Moncada JJ, García-Barco A, Zuluaga-Vélez A, Veloza LA, Sepúlveda-Arias JC. Scale-Up of the Fermentation Process for the Production and Purification of Serratiopeptidase Using Silkworm Pupae as a Substrate. Methods and Protocols. 2024; 7(2):19. https://doi.org/10.3390/mps7020019
Chicago/Turabian StyleMelchor-Moncada, Jhon Jairo, Alejandra García-Barco, Augusto Zuluaga-Vélez, Luz Angela Veloza, and Juan Carlos Sepúlveda-Arias. 2024. "Scale-Up of the Fermentation Process for the Production and Purification of Serratiopeptidase Using Silkworm Pupae as a Substrate" Methods and Protocols 7, no. 2: 19. https://doi.org/10.3390/mps7020019
APA StyleMelchor-Moncada, J. J., García-Barco, A., Zuluaga-Vélez, A., Veloza, L. A., & Sepúlveda-Arias, J. C. (2024). Scale-Up of the Fermentation Process for the Production and Purification of Serratiopeptidase Using Silkworm Pupae as a Substrate. Methods and Protocols, 7(2), 19. https://doi.org/10.3390/mps7020019






