Biocontainment Techniques and Applications for Yeast Biotechnology
Abstract
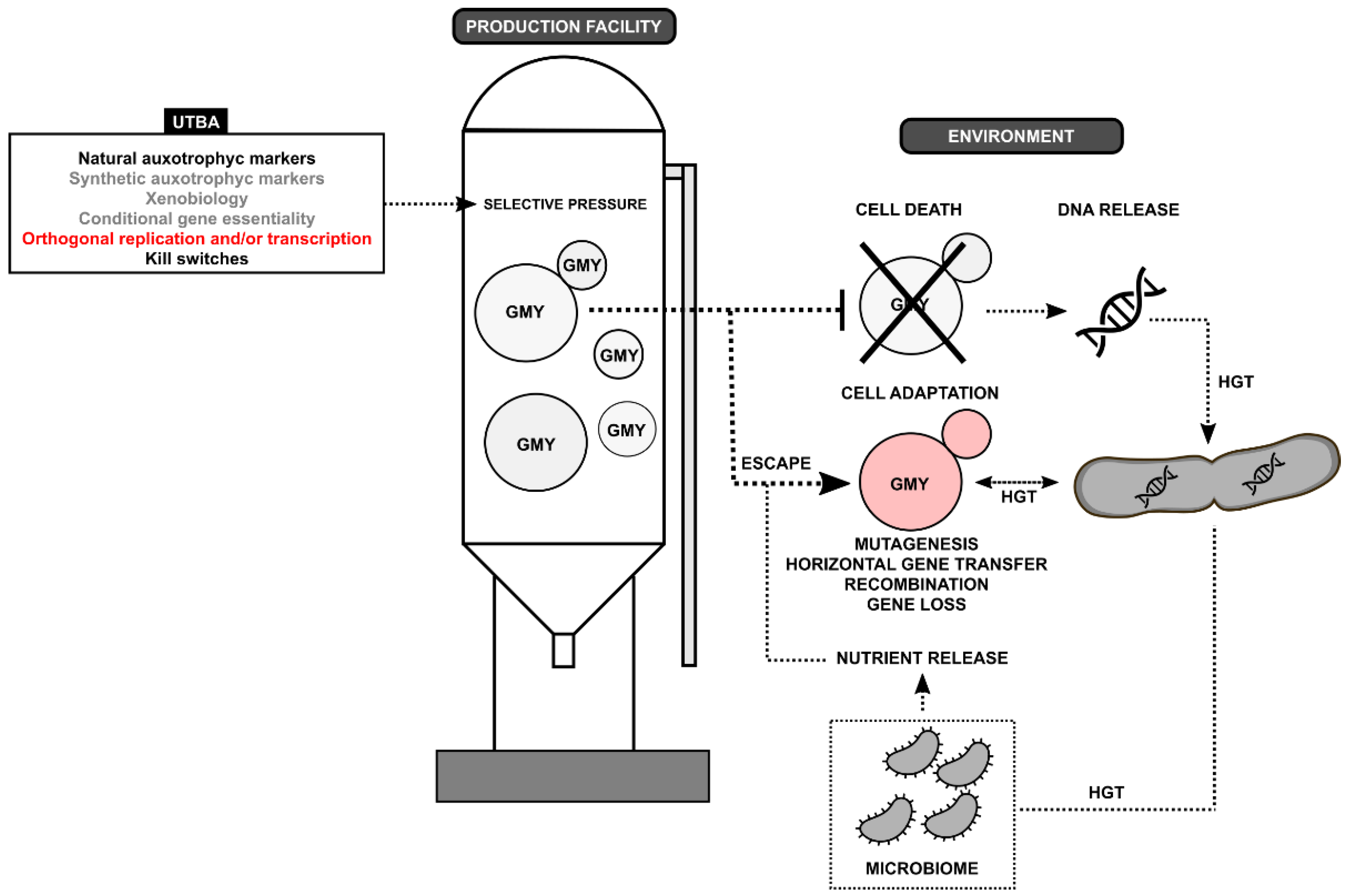
1. Introduction
2. Uniplex-Type Biocontainment Approaches
2.1. Natural Auxotrophic Markers
2.2. Xenobiology and Synthetic Auxotrophies
2.3. Conditional Gene Essentiality
2.4. Orthogonal DNA Replication and RNA Transcription
2.5. Nuclease-Based Kill Switches
2.6. Kill Switches Based on Type I and II Toxin–Antitoxin Systems
3. Multiplex-Type Biocontainment Approaches
4. Conclusions and Perspectives
Funding
Conflicts of Interest
References
- Hanlon, P.; Sewalt, V. GEMs: Genetically Engineered Microorganisms and the Regulatory Oversight of Their Uses in Modern Food Production. Crit. Rev. Food Sci. Nutr. 2021, 61, 959–970. [Google Scholar] [CrossRef] [PubMed]
- Olempska-Beer, Z.S.; Merker, R.I.; Ditto, M.D.; DiNovi, M.J. Food-Processing Enzymes from Recombinant Microorganisms—A Review. Regul. Toxicol. Pharmacol. 2006, 45, 144–158. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Tu, H.; Chen, T. Potential Application of Living Microorganisms in the Detoxification of Heavy Metals. Foods 2022, 11, 1905. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.K.; Rai, S.; Singh, D.; Upadhyay, R.S. Application of Soil Microorganisms for Agricultural and Environmental Sustainability: A Review. In Plant, Soil and Microbes in Tropical Ecosystems; Dubey, S.K., Verma, S.K., Eds.; Rhizosphere Biology; Springer: Singapore, 2021; pp. 151–175. ISBN 9789811633645. [Google Scholar]
- Shin, M.-K.; Yoo, H.S. Animal Vaccines Based on Orally Presented Yeast Recombinants. Vaccine 2013, 31, 4287–4292. [Google Scholar] [CrossRef] [PubMed]
- Adetunji, C.O.; Anani, O.A.; Olaniyan, O.T.; Bodunrinde, R.E.; Osemwegie, O.O.; Ubi, B.E. Chapter 14—Sustainability of Biofertilizers and Other Allied Products from Genetically Modified Microorganisms. In Biomass, Biofuels, Biochemicals; Varjani, S., Pandey, A., Bhaskar, T., Mohan, S.V., Tsang, D.C.W., Eds.; Elsevier: Amsterdam, The Netherlands, 2022; pp. 363–393. ISBN 978-0-323-89855-3. [Google Scholar]
- Ali, R.; Zulaykha, K.D.; Sajjad, N. Genetically Modified Microbes as Biofertilizers. In Bioremediation and Biotechnology, Vol 4: Techniques for Noxious Substances Remediation; Bhat, R.A., Hakeem, K.R., Eds.; Springer Nature: Cham, Switzerland, 2020; pp. 275–293. ISBN 978-3-030-48690-7. [Google Scholar]
- Kebede, G.; Abera, S.; Haregu, S.; Yeshitila, A.; Palanivel, H. Genetically Modified Bacteria for Alleviating Agrochemical Impact on the Environment. In Agrochemicals in Soil and Environment: Impacts and Remediation; Naeem, M., Bremont, J.F.J., Ansari, A.A., Gill, S.S., Eds.; Springer Nature: Singapore, 2022; pp. 565–583. ISBN 9789811693106. [Google Scholar]
- Ruchala, J.; Kurylenko, O.O.; Dmytruk, K.V.; Sibirny, A.A. Construction of Advanced Producers of First- and Second-Generation Ethanol in Saccharomyces cerevisiae and Selected Species of Non-Conventional Yeasts (Scheffersomyces stipitis, Ogataea polymorpha). J. Ind. Microbiol. Biotechnol. 2020, 47, 109–132. [Google Scholar] [CrossRef]
- Wang, G.; Huang, M.; Nielsen, J. Exploring the Potential of Saccharomyces cerevisiae for Biopharmaceutical Protein Production. Curr. Opin. Biotechnol. 2017, 48, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Kavšček, M.; Stražar, M.; Curk, T.; Natter, K.; Petrovič, U. Yeast as a Cell Factory: Current State and Perspectives. Microb. Cell Factories. 2015, 14, 94. [Google Scholar] [CrossRef]
- Budroni, M.; Zara, G.; Ciani, M.; Comitini, F. Saccharomyces and Non-Saccharomyces Starter Yeasts. In Brewing Technology; IntechOpen: London, UK, 2017; ISBN 978-953-51-3342-1. [Google Scholar] [CrossRef]
- Parapouli, M.; Vasileiadis, A.; Afendra, A.-S.; Hatziloukas, E. Saccharomyces cerevisiae and Its Industrial Applications. AIMS Microbiol. 2020, 6, 1–31. [Google Scholar] [CrossRef]
- Nielsen, J. Production of Biopharmaceutical Proteins by Yeast. Bioengineered 2013, 4, 207–211. [Google Scholar] [CrossRef]
- Petranovic, D.; Nielsen, J. Can Yeast Systems Biology Contribute to the Understanding of Human Disease? Trends Biotechnol. 2008, 26, 584–590. [Google Scholar] [CrossRef]
- Mustacchi, R.; Hohmann, S.; Nielsen, J. Yeast Systems Biology to Unravel the Network of Life. Yeast 2006, 23, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Andreu, C.; Zarnowski, R.; Del Olmo, M. Recent Developments in the Biology and Biotechnological Applications of Halotolerant Yeasts. World J. Microbiol. Biotechnol. 2022, 38, 27. [Google Scholar] [CrossRef]
- Strucko, T.; Andersen, N.L.; Mahler, M.R.; Martínez, J.L.; Mortensen, U.H. A CRISPR/Cas9 Method Facilitates Efficient Oligo-Mediated Gene Editing in Debaryomyces hansenii. Synth. Biol. 2021, 6, ysab031. [Google Scholar] [CrossRef] [PubMed]
- Potvin, G.; Ahmad, A.; Zhang, Z. Bioprocess Engineering Aspects of Heterologous Protein Production in Pichia pastoris: A Review. Biochem. Eng. J. 2012, 64, 91–105. [Google Scholar] [CrossRef]
- Yan, C.; Yu, W.; Zhai, X.; Yao, L.; Guo, X.; Gao, J.; Zhou, Y.J. Characterizing and Engineering Promoters for Metabolic Engineering of Ogataea polymorpha. Synth. Syst. Biotechnol. 2022, 7, 498–505. [Google Scholar] [CrossRef] [PubMed]
- Gellissen, G.; Kunze, G.; Gaillardin, C.; Cregg, J.; Berardi, E.; Veenhuis, M.; Vanderklei, I. New Yeast Expression Platforms Based on Methylotrophic Hansenula polymorpha and Pichia pastoris and on Dimorphic Arxula adeninivorans and Yarrowia lipolytica—A Comparison. FEMS Yeast Res. 2005, 5, 1079–1096. [Google Scholar] [CrossRef]
- Borneman, A.R.; Pretorius, I.S. Genomic Insights into the Saccharomyces Sensu Stricto Complex. Genetics 2015, 199, 281–291. [Google Scholar] [CrossRef]
- Hirschi, S.; Ward, T.R.; Meier, W.P.; Müller, D.J.; Fotiadis, D. Synthetic Biology: Bottom-Up Assembly of Molecular Systems. Chem. Rev. 2022, 122, 16294–16328. [Google Scholar] [CrossRef]
- Venter, J.C.; Glass, J.I.; Hutchison, C.A.; Vashee, S. Synthetic Chromosomes, Genomes, Viruses, and Cells. Cell 2022, 185, 2708–2724. [Google Scholar] [CrossRef]
- Pretorius, I.S.; Boeke, J.D. Yeast 2.0—Connecting the Dots in the Construction of the World’s First Functional Synthetic Eukaryotic Genome. FEMS Yeast Res. 2018, 18, foy032. [Google Scholar] [CrossRef]
- Arnolds, K.L.; Dahlin, L.R.; Ding, L.; Wu, C.; Yu, J.; Xiong, W.; Zuniga, C.; Suzuki, Y.; Zengler, K.; Linger, J.G.; et al. Biotechnology for Secure Biocontainment Designs in an Emerging Bioeconomy. Curr. Opin. Biotechnol. 2021, 71, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Agmon, N.; Choi, W.J.; Ubide, A.; Stracquadanio, G.; Caravelli, K.; Hao, H.; Bader, J.S.; Boeke, J.D. Intrinsic Biocontainment: Multiplex Genome Safeguards Combine Transcriptional and Recombinational Control of Essential Yeast Genes. Proc. Natl. Acad. Sci. USA 2015, 112, 1803–1808. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, P.; Gonçalves, C. Horizontal Gene Transfer in Yeasts. Curr. Opin. Genet. Dev. 2022, 76, 101950. [Google Scholar] [CrossRef]
- Schmidt, M.; de Lorenzo, V. Synthetic Bugs on the Loose: Containment Options for Deeply Engineered (Micro)organisms. Curr. Opin. Biotechnol. 2016, 38, 90–96. [Google Scholar] [CrossRef]
- NIH Guidelines for Research Involving Recombinant or Synthetic Nucleic Acid Molecules (NIH Guidelines)—April 2019. 2019. Available online: https://osp.od.nih.gov/wp-content/uploads/NIH_Guidelines.pdf (accessed on 28 February 2023).
- Andersen, J.T.; Schäfer, T.; Jørgensen, P.L.; Møller, S. Using Inactivated Microbial Biomass as Fertilizer: The Fate of Antibiotic Resistance Genes in the Environment. Res. Microbiol. 2001, 152, 823–833. [Google Scholar] [CrossRef]
- Nielsen, K.M.; Johnsen, P.J.; Bensasson, D.; Daffonchio, D. Release and Persistence of Extracellular DNA in the Environment. Environ. Biosaf. Res. 2007, 6, 37–53. [Google Scholar] [CrossRef] [PubMed]
- Recorbet, G.; Picard, C.; Normand, P.; Simonet, P. Kinetics of the Persistence of Chromosomal DNA from Genetically Engineered Escherichia coli Introduced into Soil. Appl. Environ. Microbiol. 1993, 59, 4289–4294. [Google Scholar] [CrossRef]
- Gallagher, R.R.; Patel, J.R.; Interiano, A.L.; Rovner, A.J.; Isaacs, F.J. Multilayered Genetic Safeguards Limit Growth of Microorganisms to Defined Environments. Nucleic Acids Res. 2015, 43, 1945–1954. [Google Scholar] [CrossRef]
- Lee, J.W.; Chan, C.T.Y.; Slomovic, S.; Collins, J.J. Next-Generation Biocontainment Systems for Engineered Organisms. Nat. Chem. Biol. 2018, 14, 530–537. [Google Scholar] [CrossRef]
- Knudsen, S.M.; Karlström, O.H. Development of Efficient Suicide Mechanisms for Biological Containment of Bacteria. Appl. Environ. Microbiol. 1991, 57, 85–92. [Google Scholar] [CrossRef]
- Kong, W.; Wanda, S.-Y.; Zhang, X.; Bollen, W.; Tinge, S.A.; Roland, K.L.; Curtiss, R. Regulated Programmed Lysis of Recombinant Salmonella in Host Tissues to Release Protective Antigens and Confer Biological Containment. Proc. Natl. Acad. Sci. USA 2008, 105, 9361–9366. [Google Scholar] [CrossRef] [PubMed]
- Mandell, D.J.; Lajoie, M.J.; Mee, M.T.; Takeuchi, R.; Kuznetsov, G.; Norville, J.E.; Gregg, C.J.; Stoddard, B.L.; Church, G.M. Biocontainment of Genetically Modified Organisms by Synthetic Protein Design. Nature 2015, 518, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Pronk, J.T. Auxotrophic Yeast Strains in Fundamental and Applied Research. Appl. Environ. Microbiol. 2002, 68, 2095. [Google Scholar] [CrossRef] [PubMed]
- Kokina, A.; Ozolina, Z.; Liepins, J. Purine Auxotrophy: Possible Applications beyond Genetic Marker. Yeast 2019, 36, 649–656. [Google Scholar] [CrossRef]
- Ralser, M.; Kuhl, H.; Ralser, M.; Werber, M.; Lehrach, H.; Breitenbach, M.; Timmermann, B. The Saccharomyces cerevisiae W303-K6001 Cross-Platform Genome Sequence: Insights into Ancestry and Physiology of a Laboratory Mutt. Open Biol. 2012, 2, 120093. [Google Scholar] [CrossRef]
- Kokina, A.; Kibilds, J.; Liepins, J. Adenine Auxotrophy—Be Aware: Some Effects of Adenine Auxotrophy in Saccharomyces cerevisiae Strain W303-1A. FEMS Yeast Res. 2014, 14, 697–707. [Google Scholar] [CrossRef]
- Weng, Y.; Nickoloff, J.A. Nonselective URA3 Colony-Color Assay in Yeast Ade1 or Ade2 Mutants. BioTechniques 1997, 23, 237–242. [Google Scholar] [CrossRef]
- Bharathi, V.; Girdhar, A.; Prasad, A.; Verma, M.; Taneja, V.; Patel, B.K. Use of Ade1 and Ade2 Mutations for Development of a Versatile Red/White Colour Assay of Amyloid-Induced Oxidative Stress in Saccharomyces cerevisiae: Use of Ade1 and Ade2 Mutations for Development of Colour Assay. Yeast 2016, 33, 607–620. [Google Scholar] [CrossRef]
- Achilli, A.; Matmati, N.; Casalone, E.; Morpurgo, G.; Lucaccioni, A.; Pavlov, Y.I.; Babudri, N. The Exceptionally High Rate of Spontaneous Mutations in the Polymerase Delta Proofreading Exonuclease-Deficient Saccharomyces cerevisiae Strain Starved for Adenine. BMC Genet. 2004, 5, 34. [Google Scholar] [CrossRef]
- Heidenreich, E.; Wintersberger, U. Starvation for a Specific Amino Acid Induces High Frequencies of Rho− Mutants in Saccharomyces cerevisiae. Curr. Genet. 1997, 31, 408–413. [Google Scholar] [CrossRef]
- Hall, B.G. Selection-Induced Mutations Occur in Yeast. Proc. Natl. Acad. Sci. USA 1992, 89, 4300–4303. [Google Scholar] [CrossRef] [PubMed]
- Brachmann, C.B.; Davies, A.; Cost, G.J.; Caputo, E.; Li, J.; Hieter, P.; Boeke, J.D. Designer Deletion Strains Derived from Saccharomyces cerevisiae S288C: A Useful Set of Strains and Plasmids for PCR-Mediated Gene Disruption and Other Applications. Yeast 1998, 14, 115–132. [Google Scholar] [CrossRef]
- Heinisch, J.J.; Buchwald, U.; Gottschlich, A.; Heppeler, N.; Rodicio, R. A Tool Kit for Molecular Genetics of Kluyveromyces lactis Comprising a Congenic Strain Series and a Set of Versatile Vectors. FEMS Yeast Res. 2010, 10, 333–342. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Winkler, C.M.; Kolmbauer, M.; Pichler, H.; Schwab, H.; Emmerstorfer-Augustin, A. Pichia pastoris Protease-Deficient and Auxotrophic Strains Generated by a Novel, User-Friendly Vector Toolbox for Gene Deletion. Yeast 2019, 36, 557–570. [Google Scholar] [CrossRef]
- Zhu, X.; Zhang, Z.; Jia, B.; Yuan, Y. Current Advances of Biocontainment Strategy in Synthetic Biology. Chin. J. Chem. Eng. 2023, 56, 141–151. [Google Scholar] [CrossRef]
- Kim, D.; Lee, J.W. Genetic Biocontainment Systems for the Safe Use of Engineered Microorganisms. Biotechnol. Bioprocess Eng. 2020, 25, 974–984. [Google Scholar] [CrossRef]
- Nieto-Domínguez, M.; Nikel, P.I. Intersecting Xenobiology and Neometabolism To Bring Novel Chemistries to Life. ChemBioChem 2020, 21, 2551–2571. [Google Scholar] [CrossRef]
- Odar, C.; Winkler, M.; Wiltschi, B. Fluoro Amino Acids: A Rarity in Nature, yet a Prospect for Protein Engineering. Biotechnol. J. 2015, 10, 427–446. [Google Scholar] [CrossRef]
- Handal-Marquez, P.; Anupama, A.; Pezo, V.; Marlière, P.; Herdewijn, P.; Pinheiro, V.B. Beneath the XNA World: Tools and Targets to Build Novel Biology. Curr. Opin. Syst. Biol. 2020, 24, 142–152. [Google Scholar] [CrossRef]
- Rovner, A.J.; Haimovich, A.D.; Katz, S.R.; Li, Z.; Grome, M.W.; Gassaway, B.M.; Amiram, M.; Patel, J.R.; Gallagher, R.R.; Rinehart, J.; et al. Recoded Organisms Engineered to Depend on Synthetic Amino Acids. Nature 2015, 518, 89–93. [Google Scholar] [CrossRef]
- de la Torre, D.; Chin, J.W. Reprogramming the Genetic Code. Nat. Rev. Genet. 2021, 22, 169–184. [Google Scholar] [CrossRef] [PubMed]
- Sanders, J.; Hoffmann, S.A.; Green, A.P.; Cai, Y. New Opportunities for Genetic Code Expansion in Synthetic Yeast. Curr. Opin. Biotechnol. 2022, 75, 102691. [Google Scholar] [CrossRef] [PubMed]
- Wan, W.; Tharp, J.M.; Liu, W.R. Pyrrolysyl-TRNA Synthetase: An Ordinary Enzyme but an Outstanding Genetic Code Expansion Tool. Biochim. Biophys. Acta 2014, 1844, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Hancock, S.M.; Uprety, R.; Deiters, A.; Chin, J.W. Expanding the Genetic Code of Yeast for Incorporation of Diverse Unnatural Amino Acids via a Pyrrolysyl-TRNA Synthetase/TRNA Pair. J. Am. Chem. Soc. 2010, 132, 14819–14824. [Google Scholar] [CrossRef] [PubMed]
- Wiltschi, B. Incorporation of Non-Canonical Amino Acids into Proteins in Yeast. Fungal Genet. Biol. 2016, 89, 137–156. [Google Scholar] [CrossRef]
- Chin, J.W.; Cropp, T.A.; Anderson, J.C.; Mukherji, M.; Zhang, Z.; Schultz, P.G. An Expanded Eukaryotic Genetic Code. Science 2003, 301, 964–967. [Google Scholar] [CrossRef]
- Budisa, N.; Wenger, W.; Wiltschi, B. Residue-Specific Global Fluorination of Candida antarctica Lipase B in Pichia pastoris. Mol. Biosyst. 2010, 6, 1630–1639. [Google Scholar] [CrossRef]
- Ai, H.-W.; Shen, W.; Brustad, E.; Schultz, P.G. Genetically Encoded Alkenes in Yeast. Angew. Chem. Int. Ed. 2010, 49, 935–937. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, L. New Methods Enabling Efficient Incorporation of Unnatural Amino Acids in Yeast. J. Am. Chem. Soc. 2008, 130, 6066–6067. [Google Scholar] [CrossRef]
- Chen, S.; Schultz, P.G.; Brock, A. An Improved System for the Generation and Analysis of Mutant Proteins Containing Unnatural Amino Acids in Saccharomyces cerevisiae. J. Mol. Biol. 2007, 371, 112–122. [Google Scholar] [CrossRef]
- Brustad, E.; Bushey, M.L.; Brock, A.; Chittuluru, J.; Schultz, P.G. A Promiscuous Aminoacyl-TRNA Synthetase That Incorporates Cysteine, Methionine, and Alanine Homologs into Proteins. Bioorg. Med. Chem. Lett. 2008, 18, 6004–6006. [Google Scholar] [CrossRef] [PubMed]
- Tippmann, E.M.; Schultz, P.G. A Genetically Encoded Metallocene Containing Amino Acid. Tetrahedron 2007, 63, 6182–6184. [Google Scholar] [CrossRef]
- Lemke, E.A.; Summerer, D.; Geierstanger, B.H.; Brittain, S.M.; Schultz, P.G. Control of Protein Phosphorylation with a Genetically Encoded Photocaged Amino Acid. Nat. Chem. Biol. 2007, 3, 769–772. [Google Scholar] [CrossRef] [PubMed]
- Summerer, D.; Chen, S.; Wu, N.; Deiters, A.; Chin, J.W.; Schultz, P.G. A Genetically Encoded Fluorescent Amino Acid. Proc. Natl. Acad. Sci. USA 2006, 103, 9785–9789. [Google Scholar] [CrossRef]
- Lee, H.S.; Guo, J.; Lemke, E.A.; Dimla, R.D.; Schultz, P.G. Genetic Incorporation of a Small, Environmentally Sensitive, Fluorescent Probe into Proteins in Saccharomyces cerevisiae. J. Am. Chem. Soc. 2009, 131, 12921–12923. [Google Scholar] [CrossRef]
- Young, T.S.; Ahmad, I.; Brock, A.; Schultz, P.G. Expanding the Genetic Repertoire of the Methylotrophic Yeast Pichia pastoris. Biochemistry 2009, 48, 2643–2653. [Google Scholar] [CrossRef]
- Himmelfarb, H.J.; Maicas, E.; Friesen, J.D. Isolation of the SUP45 Omnipotent Suppressor Gene of Saccharomyces cerevisiae and Characterization of Its Gene Product. Mol. Cell. Biol. 1985, 5, 816–822. [Google Scholar]
- Stansfield, I.; Jones, K.M.; Kushnirov, V.V.; Dagkesamanskaya, A.R.; Poznyakovski, A.I.; Paushkin, S.V.; Nierras, C.R.; Cox, B.S.; Ter-Avanesyan, M.D.; Tuite, M.F. The Products of the SUP45 (ERF1) and SUP35 Genes Interact to Mediate Translation Termination in Saccharomyces cerevisiae. EMBO J. 1995, 14, 4365–4373. [Google Scholar] [CrossRef]
- Lajoie, M.J.; Rovner, A.J.; Goodman, D.B.; Aerni, H.-R.; Haimovich, A.D.; Kuznetsov, G.; Mercer, J.A.; Wang, H.H.; Carr, P.A.; Mosberg, J.A.; et al. Genomically Recoded Organisms Expand Biological Functions. Science 2013, 342, 357–360. [Google Scholar] [CrossRef]
- Stieglitz, J.T.; Lahiri, P.; Stout, M.I.; Van Deventer, J.A. Exploration of Methanomethylophilus Alvus Pyrrolysyl-TRNA Synthetase Activity in Yeast. ACS Synth. Biol. 2022, 11, 1824–1834. [Google Scholar] [CrossRef]
- Zackin, M.T.; Stieglitz, J.T.; Van Deventer, J.A. Genome-Wide Screen for Enhanced Noncanonical Amino Acid Incorporation in Yeast. ACS Synth. Biol. 2022, 11, 3669–3680. [Google Scholar] [CrossRef]
- Stieglitz, J.T.; Van Deventer, J.A. High-Throughput Aminoacyl-TRNA Synthetase Engineering for Genetic Code Expansion in Yeast. ACS Synth. Biol. 2022, 11, 2284–2299. [Google Scholar] [CrossRef]
- Dunkelmann, D.L.; Oehm, S.B.; Beattie, A.T.; Chin, J.W. A 68-Codon Genetic Code to Incorporate Four Distinct Non-Canonical Amino Acids Enabled by Automated Orthogonal MRNA Design. Nat. Chem. 2021, 13, 1110–1117. [Google Scholar] [CrossRef] [PubMed]
- Hankore, E.D.; Zhang, L.; Chen, Y.; Liu, K.; Niu, W.; Guo, J. Genetic Incorporation of Noncanonical Amino Acids Using Two Mutually Orthogonal Quadruplet Codons. ACS Synth. Biol. 2019, 8, 1168–1174. [Google Scholar] [CrossRef] [PubMed]
- Niu, W.; Schultz, P.G.; Guo, J. An Expanded Genetic Code in Mammalian Cells with a Functional Quadruplet Codon. ACS Chem. Biol. 2013, 8, 1640–1645. [Google Scholar] [CrossRef] [PubMed]
- Fischer, E.C.; Hashimoto, K.; Zhang, Y.; Feldman, A.W.; Dien, V.T.; Karadeema, R.J.; Adhikary, R.; Ledbetter, M.P.; Krishnamurthy, R.; Romesberg, F.E. New Codons for Efficient Production of Unnatural Proteins in a Semi-Synthetic Organism. Nat. Chem. Biol. 2020, 16, 570–576. [Google Scholar] [CrossRef]
- Nie, P.; Bai, Y.; Mei, H. Synthetic Life with Alternative Nucleic Acids as Genetic Materials. Molecules 2020, 25, 3483. [Google Scholar] [CrossRef]
- Torres, L.; Krüger, A.; Csibra, E.; Gianni, E.; Pinheiro, V.B. Synthetic Biology Approaches to Biological Containment: Pre-Emptively Tackling Potential Risks. Essays Biochem. 2016, 60, 393–410. [Google Scholar] [CrossRef]
- Herdewijn, P.; Marlière, P. Toward Safe Genetically Modified Organisms through the Chemical Diversification of Nucleic Acids. Chem. Biodivers. 2009, 6, 791–808. [Google Scholar] [CrossRef]
- Dhami, K.; Malyshev, D.A.; Ordoukhanian, P.; Kubelka, T.; Hocek, M.; Romesberg, F.E. Systematic Exploration of a Class of Hydrophobic Unnatural Base Pairs Yields Multiple New Candidates for the Expansion of the Genetic Alphabet. Nucleic Acids Res. 2014, 42, 10235–10244. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Holliger, P. The XNA World: Progress towards Replication and Evolution of Synthetic Genetic Polymers. Curr. Opin. Chem. Biol. 2012, 16, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Hendrix, C.; Rosemeyer, H.; Verheggen, I.; Van Aerschot, A.; Seela, F.; Herdewijn, P. 1′, 5′-Anhydrohexitol Oligonucleotides: Synthesis, Base Pairing and Recognition by Regular Oligodeoxyribonucleotides and Oligoribonucleotides. Chem.–A Eur. J. 1997, 3, 110–120. [Google Scholar] [CrossRef]
- Schöning, K.U.; Scholz, P.; Guntha, S.; Wu, X.; Krishnamurthy, R.; Eschenmoser, A. Chemical Etiology of Nucleic Acid Structure: The α-Threofuranosyl-(3′→2′) Oligonucleotide System. Science 2000, 290, 1347–1351. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Verbeure, B.; Luyten, I.; Lescrinier, E.; Froeyen, M.; Hendrix, C.; Rosemeyer, H.; Seela, F.; Van Aerschot, A.; Herdewijn, P. Cyclohexene Nucleic Acids (CeNA): Serum Stable Oligonucleotides That Activate RNase H and Increase Duplex Stability with Complementary RNA. J. Am. Chem. Soc. 2000, 122, 8595–8602. [Google Scholar] [CrossRef]
- Noronha, A.M.; Wilds, C.J.; Lok, C.N.; Viazovkina, K.; Arion, D.; Parniak, M.A.; Damha, M.J. Synthesis and Biophysical Properties of Arabinonucleic Acids (ANA): Circular Dichroic Spectra, Melting Temperatures, and Ribonuclease H Susceptibility of ANA·RNA Hybrid Duplexes. Biochemistry 2000, 39, 7050–7062. [Google Scholar] [CrossRef]
- Kalota, A.; Karabon, L.; Swider, C.R.; Viazovkina, E.; Elzagheid, M.; Damha, M.J.; Gewirtz, A.M. 2′-Deoxy-2′-Fluoro-β-d-Arabinonucleic Acid (2′F-ANA) Modified Oligonucleotides (ON) Effect Highly Efficient, and Persistent, Gene Silencing. Nucleic Acids Res. 2006, 34, 451–461. [Google Scholar] [CrossRef]
- Zhang, L.; Peritz, A.; Meggers, E. A Simple Glycol Nucleic Acid. J. Am. Chem. Soc. 2005, 127, 4174–4175. [Google Scholar] [CrossRef]
- Singh, S.K.; Koshkin, A.A.; Wengel, J.; Nielsen, P. LNA (Locked Nucleic Acids): Synthesis and High-Affinity Nucleic Acid Recognition. Chem. Commun. 1998, 4, 455–456. [Google Scholar] [CrossRef]
- Liu, C.; Cozens, C.; Jaziri, F.; Rozenski, J.; Maréchal, A.; Dumbre, S.; Pezo, V.; Marlière, P.; Pinheiro, V.B.; Groaz, E.; et al. Phosphonomethyl Oligonucleotides as Backbone-Modified Artificial Genetic Polymers. J. Am. Chem. Soc. 2018, 140, 6690–6699. [Google Scholar] [CrossRef]
- Diwo, C.; Budisa, N. Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation. Genes 2019, 10, 17. [Google Scholar] [CrossRef]
- Lopez, G.; Anderson, J.C. Synthetic Auxotrophs with Ligand-Dependent Essential Genes for a BL21(DE3) Biosafety Strain. ACS Synth. Biol. 2015, 4, 1279–1286. [Google Scholar] [CrossRef] [PubMed]
- Motomura, K.; Sano, K.; Watanabe, S.; Kanbara, A.; Gamal Nasser, A.-H.; Ikeda, T.; Ishida, T.; Funabashi, H.; Kuroda, A.; Hirota, R. Synthetic Phosphorus Metabolic Pathway for Biosafety and Contamination Management of Cyanobacterial Cultivation. ACS Synth. Biol. 2018, 7, 2189–2198. [Google Scholar] [CrossRef] [PubMed]
- Asin-Garcia, E.; Batianis, C.; Li, Y.; Fawcett, J.D.; de Jong, I.; dos Santos, V.A.P.M. Phosphite Synthetic Auxotrophy as an Effective Biocontainment Strategy for the Industrial Chassis Pseudomonas putida. Microb. Cell Factories 2022, 21, 156. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.R.; Steingesser, M.G.; Weems, A.D.; Khan, A.; Gladfelter, A.; Bertin, A.; McMurray, M.A. Guanidine Hydrochloride Reactivates an Ancient Septin Hetero-Oligomer Assembly Pathway in Budding Yeast. eLife 2020, 9, e54355. [Google Scholar] [CrossRef]
- Hassell, D.S.; Steingesser, M.G.; Denney, A.S.; Johnson, C.R.; McMurray, M.A. Chemical Rescue of Mutant Proteins in Living Saccharomyces cerevisiae Cells by Naturally Occurring Small Molecules. G3 GenesGenomesGenetics 2021, 11, jkab252. [Google Scholar] [CrossRef]
- Agmon, N.; Tang, Z.; Yang, K.; Sutter, B.; Ikushima, S.; Cai, Y.; Caravelli, K.; Martin, J.A.; Sun, X.; Choi, W.J.; et al. Low Escape-Rate Genome Safeguards with Minimal Molecular Perturbation of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2017, 114, E1470–E1479. [Google Scholar] [CrossRef]
- Lindstrom, D.L.; Gottschling, D.E. The Mother Enrichment Program: A Genetic System for Facile Replicative Life Span Analysis in Saccharomyces cerevisiae. Genetics 2009, 183, 413–422. [Google Scholar] [CrossRef]
- Arzumanyan, G.A.; Gabriel, K.N.; Ravikumar, A.; Javanpour, A.A.; Liu, C.C. Mutually Orthogonal DNA Replication Systems In Vivo. ACS Synth. Biol. 2018, 7, 1722–1729. [Google Scholar] [CrossRef]
- Ravikumar, A.; Arzumanyan, G.A.; Obadi, M.K.A.; Javanpour, A.A.; Liu, C.C. Scalable, Continuous Evolution of Genes at Mutation Rates above Genomic Error Thresholds. Cell 2018, 175, 1946–1957.e13. [Google Scholar] [CrossRef]
- Strak, M.J.R.; Boyd, A.; Mileham, A.J.; Ramonos, M.A. The Plasmid-Encoded Killer System of Kluyveromyces lactis: A Review. Yeast 1990, 6, 1–29. [Google Scholar] [CrossRef]
- van Nies, P.; Westerlaken, I.; Blanken, D.; Salas, M.; Mencía, M.; Danelon, C. Self-Replication of DNA by Its Encoded Proteins in Liposome-Based Synthetic Cells. Nat. Commun. 2018, 9, 1583. [Google Scholar] [CrossRef] [PubMed]
- Sakatani, Y.; Ichihashi, N.; Kazuta, Y.; Yomo, T. A Transcription and Translation-Coupled DNA Replication System Using Rolling-Circle Replication. Sci. Rep. 2015, 5, 10404. [Google Scholar] [CrossRef] [PubMed]
- Libicher, K.; Hornberger, R.; Heymann, M.; Mutschler, H. In Vitro Self-Replication and Multicistronic Expression of Large Synthetic Genomes. Nat. Commun. 2020, 11, 904. [Google Scholar] [CrossRef]
- Medina, E.; Yik, E.J.; Herdewijn, P.; Chaput, J.C. Functional Comparison of Laboratory-Evolved XNA Polymerases for Synthetic Biology. ACS Synth. Biol. 2021, 10, 1429–1437. [Google Scholar] [CrossRef] [PubMed]
- Kar, S.; Ellington, A.D. Construction of Synthetic T7 RNA Polymerase Expression Systems. Methods 2018, 143, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Morse, N.J.; Wagner, J.M.; Reed, K.B.; Gopal, M.R.; Lauffer, L.H.; Alper, H.S. T7 Polymerase Expression of Guide RNAs In Vivo Allows Exportable CRISPR-Cas9 Editing in Multiple Yeast Hosts. ACS Synth. Biol. 2018, 7, 1075–1084. [Google Scholar] [CrossRef]
- Stirling, F.; Bitzan, L.; O’Keefe, S.; Redfield, E.; Oliver, J.W.K.; Way, J.; Silver, P.A. Rational Design of Evolutionarily Stable Microbial Kill Switches. Mol. Cell 2017, 68, 686–697.e3. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.T.Y.; Lee, J.W.; Cameron, D.E.; Bashor, C.J.; Collins, J.J. “Deadman” and “Passcode” Microbial Kill Switches for Bacterial Containment. Nat. Chem. Biol. 2016, 12, 82–86. [Google Scholar] [CrossRef]
- Halvorsen, T.M.; Ricci, D.P.; Park, D.M.; Jiao, Y.; Yung, M.C. Comparison of Kill Switch Toxins in Plant-Beneficial Pseudomonas fluorescens Reveals Drivers of Lethality, Stability, and Escape. ACS Synth. Biol. 2022, 11, 3785–3796. [Google Scholar] [CrossRef]
- Wright, O.; Delmans, M.; Stan, G.-B.; Ellis, T. GeneGuard: A Modular Plasmid System Designed for Biosafety. ACS Synth. Biol. 2015, 4, 307–316. [Google Scholar] [CrossRef]
- Whitford, C.M.; Dymek, S.; Kerkhoff, D.; März, C.; Schmidt, O.; Edich, M.; Droste, J.; Pucker, B.; Rückert, C.; Kalinowski, J. Auxotrophy to Xeno-DNA: An Exploration of Combinatorial Mechanisms for a High-Fidelity Biosafety System for Synthetic Biology Applications. J. Biol. Eng. 2018, 12, 13. [Google Scholar] [CrossRef] [PubMed]
- Ahrenholtz, I.; Lorenz, M.G.; Wackernagel, W. A Conditional Suicide System in Escherichia coli Based on the Intracellular Degradation of DNA. Appl. Environ. Microbiol. 1994, 60, 3746–3751. [Google Scholar] [CrossRef] [PubMed]
- Balan, A.; Schenberg, A.C.G. A Conditional Suicide System for Saccharomyces cerevisiae Relying on the Intracellular Production of the Serratia marcescens Nuclease. Yeast 2005, 22, 203–212. [Google Scholar] [CrossRef]
- Lewis, L.K.; Kirchner, J.M.; Resnick, M.A. Requirement for End-Joining and Checkpoint Functions, but Not RAD52-Mediated Recombination, after EcoRI Endonuclease Cleavage of Saccharomyces cerevisiae DNA. Mol. Cell. Biol. 1998, 18, 1891–1902. [Google Scholar] [CrossRef]
- Barnes, G.; Rine, J. Regulated Expression of Endonuclease EcoRI in Saccharomyces cerevisiae: Nuclear Entry and Biological Consequences. Proc. Natl. Acad. Sci. USA 1985, 82, 1354–1358. [Google Scholar] [CrossRef] [PubMed]
- Westmoreland, J.W.; Summers, J.A.; Holland, C.L.; Resnick, M.A.; Lewis, L.K. Blunt-Ended DNA Double-Strand Breaks Induced by Endonucleases PvuII and EcoRV Are Poor Substrates for Repair in Saccharomyces cerevisiae. DNA Repair 2010, 9, 617–626. [Google Scholar] [CrossRef] [PubMed]
- Caliando, B.J.; Voigt, C.A. Targeted DNA Degradation Using a CRISPR Device Stably Carried in the Host Genome. Nat. Commun. 2015, 6, 6989. [Google Scholar] [CrossRef]
- Rottinghaus, A.G.; Ferreiro, A.; Fishbein, S.R.S.; Dantas, G.; Moon, T.S. Genetically Stable CRISPR-Based Kill Switches for Engineered Microbes. Nat. Commun. 2022, 13, 672. [Google Scholar] [CrossRef]
- Rottinghaus, A.G.; Vo, S.; Moon, T.S. Computational Design of CRISPR Guide RNAs to Enable Strain-Specific Control of Microbial Consortia. Proc. Natl. Acad. Sci. USA 2023, 120, e2213154120. [Google Scholar] [CrossRef]
- Pantoja Angles, A.; Valle-Pérez, A.U.; Hauser, C.; Mahfouz, M.M. Microbial Biocontainment Systems for Clinical, Agricultural, and Industrial Applications. Front. Bioeng. Biotechnol. 2022, 10, 830200. [Google Scholar] [CrossRef]
- Yeo, C.C.; Abu Bakar, F.; Chan, W.T.; Espinosa, M.; Harikrishna, J.A. Heterologous Expression of Toxins from Bacterial Toxin-Antitoxin Systems in Eukaryotic Cells: Strategies and Applications. Toxins 2016, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, Y.; Park, J.-H.; Inouye, M. Toxin-Antitoxin Systems in Bacteria and Archaea. Annu. Rev. Genet. 2011, 45, 61–79. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yao, J.; Sun, Y.-C.; Wood, T.K. Type VII Toxin/Antitoxin Classification System for Antitoxins That Enzymatically Neutralize Toxins. Trends Microbiol. 2021, 29, 388–393. [Google Scholar] [CrossRef] [PubMed]
- Kamruzzaman, M.; Wu, A.Y.; Iredell, J.R. Biological Functions of Type II Toxin-Antitoxin Systems in Bacteria. Microorganisms 2021, 9, 1276. [Google Scholar] [CrossRef]
- Fineran, P.C.; Blower, T.R.; Foulds, I.J.; Humphreys, D.P.; Lilley, K.S.; Salmond, G.P.C. The Phage Abortive Infection System, ToxIN, Functions as a Protein-RNA Toxin-Antitoxin Pair. Proc. Natl. Acad. Sci. USA 2009, 106, 894–899. [Google Scholar] [CrossRef]
- Masuda, H.; Tan, Q.; Awano, N.; Wu, K.-P.; Inouye, M. YeeU Enhances the Bundling of Cytoskeletal Polymers of MreB and FtsZ, Antagonizing the CbtA (YeeV) Toxicity in Escherichia coli. Mol. Microbiol. 2012, 84, 979–989. [Google Scholar] [CrossRef]
- Wang, X.; Lord, D.M.; Cheng, H.-Y.; Osbourne, D.O.; Hong, S.H.; Sanchez-Torres, V.; Quiroga, C.; Zheng, K.; Herrmann, T.; Peti, W.; et al. A New Type V Toxin-Antitoxin System Where MRNA for Toxin GhoT Is Cleaved by Antitoxin GhoS. Nat. Chem. Biol. 2012, 8, 855–861. [Google Scholar] [CrossRef]
- Aakre, C.D.; Phung, T.N.; Huang, D.; Laub, M.T. A Bacterial Toxin Inhibits DNA Replication Elongation through a Direct Interaction with the β Sliding Clamp. Mol. Cell 2013, 52, 617–628. [Google Scholar] [CrossRef]
- Bej, A.K.; Perlin, M.H.; Atlas, R.M. Model Suicide Vector for Containment of Genetically Engineered Microorganisms. Appl. Environ. Microbiol. 1988, 54, 2472–2477. [Google Scholar] [CrossRef]
- Schweder, T.; Schmidt, I.; Herrmann, H.; Neubauer, P.; Hecker, M.; Hofmann, K. An Expression Vector System Providing Plasmid Stability and Conditional Suicide of Plasmid-Containing Cells. Appl. Microbiol. Biotechnol. 1992, 38, 91–93. [Google Scholar] [CrossRef]
- Kroll, J.; Klinter, S.; Schneider, C.; Voss, I.; Steinbüchel, A. Plasmid Addiction Systems: Perspectives and Applications in Biotechnology. Microb. Biotechnol. 2010, 3, 634–657. [Google Scholar] [CrossRef]
- Kristoffersen, P.; Jensen, G.B.; Gerdes, K.; Piškur, J. Bacterial Toxin-Antitoxin Gene System as Containment Control in Yeast Cells. Appl. Environ. Microbiol. 2000, 66, 5524–5526. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, K.; Zavialov, A.V.; Pavlov, M.Y.; Elf, J.; Gerdes, K.; Ehrenberg, M. The Bacterial Toxin RelE Displays Codon-Specific Cleavage of MRNAs in the Ribosomal A Site. Cell 2003, 112, 131–140. [Google Scholar] [CrossRef]
- Li, G.-Y.; Zhang, Y.; Inouye, M.; Ikura, M. Inhibitory Mechanism of Escherichia coli RelE-RelB Toxin-Antitoxin Module Involves a Helix Displacement Near an MRNA Interferase Active Site. J. Biol. Chem. 2009, 284, 14628–14636. [Google Scholar] [CrossRef] [PubMed]
- Monti, M.C.; Hernández-Arriaga, A.M.; Kamphuis, M.B.; López-Villarejo, J.; Heck, A.J.R.; Boelens, R.; Díaz-Orejas, R.; van den Heuvel, R.H.H. Interactions of Kid–Kis Toxin–Antitoxin Complexes with the ParD Operator-Promoter Region of Plasmid R1 Are Piloted by the Kis Antitoxin and Tuned by the Stoichiometry of Kid–Kis Oligomers. Nucleic Acids Res. 2007, 35, 1737–1749. [Google Scholar] [CrossRef]
- de la Cueva-Méndez, G.; Mills, A.D.; Clay-Farrace, L.; Díaz-Orejas, R.; Laskey, R.A. Regulatable Killing of Eukaryotic Cells by the Prokaryotic Proteins Kid and Kis. EMBO J. 2003, 22, 246–251. [Google Scholar] [CrossRef]
- Zielenkiewicz, U.; Cegłowski, P. The Toxin-Antitoxin System of the Streptococcal Plasmid PSM19035. J. Bacteriol. 2005, 187, 6094–6105. [Google Scholar] [CrossRef] [PubMed]
- Zielenkiewicz, U.; Kowalewska, M.; Kaczor, C.; Cegłowski, P. In Vivo Interactions between Toxin-Antitoxin Proteins Epsilon and Zeta of Streptococcal Plasmid PSM19035 in Saccharomyces cerevisiae. J. Bacteriol. 2009, 191, 3677–3684. [Google Scholar] [CrossRef] [PubMed]
- Picardeau, M.; Le Dantec, C.; Richard, G.-F.; Saint Girons, I. The Spirochetal ChpK-Chromosomal Toxin–Antitoxin Locus Induces Growth Inhibition of Yeast and Mycobacteria. FEMS Microbiol. Lett. 2003, 229, 277–281. [Google Scholar] [CrossRef]
- Picardeau, M.; Ren, S.; Saint Girons, I. Killing Effect and Antitoxic Activity of The Leptospira Interrogans Toxin-Antitoxin System In Escherichia coli. J. Bacteriol. 2001, 183, 6494–6497. [Google Scholar] [CrossRef]
- Engelberg-Kulka, H.; Hazan, R.; Amitai, S. MazEF: A Chromosomal Toxin-Antitoxin Module That Triggers Programmed Cell Death in Bacteria. J. Cell Sci. 2005, 118, 4327–4332. [Google Scholar] [CrossRef]
- Yang, J.; Jiang, W.; Yang, S. MazF as a Counter-Selectable Marker for Unmarked Genetic Modification of Pichia pastoris. FEMS Yeast Res. 2009, 9, 600–609. [Google Scholar] [CrossRef]
- Knudsen, S.; Saadbye, P.; Hansen, L.H.; Collier, A.; Jacobsen, B.L.; Schlundt, J.; Karlström, O.H. Development and Testing of Improved Suicide Functions for Biological Containment of Bacteria. Appl. Environ. Microbiol. 1995, 61, 985–991. [Google Scholar] [CrossRef]
- Moser, F.; Broers, N.J.; Hartmans, S.; Tamsir, A.; Kerkman, R.; Roubos, J.A.; Bovenberg, R.; Voigt, C.A. Genetic Circuit Performance under Conditions Relevant for Industrial Bioreactors. ACS Synth. Biol. 2012, 1, 555–564. [Google Scholar] [CrossRef]
| Disadvantages | Advantages | Biocontainment |
|---|---|---|
|
| Uniplex-type approach (UTBA) |
|
| Multiplex-type approach (MTBA) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pavão, G.; Sfalcin, I.; Bonatto, D. Biocontainment Techniques and Applications for Yeast Biotechnology. Fermentation 2023, 9, 341. https://doi.org/10.3390/fermentation9040341
Pavão G, Sfalcin I, Bonatto D. Biocontainment Techniques and Applications for Yeast Biotechnology. Fermentation. 2023; 9(4):341. https://doi.org/10.3390/fermentation9040341
Chicago/Turabian StylePavão, Guilherme, Isabela Sfalcin, and Diego Bonatto. 2023. "Biocontainment Techniques and Applications for Yeast Biotechnology" Fermentation 9, no. 4: 341. https://doi.org/10.3390/fermentation9040341
APA StylePavão, G., Sfalcin, I., & Bonatto, D. (2023). Biocontainment Techniques and Applications for Yeast Biotechnology. Fermentation, 9(4), 341. https://doi.org/10.3390/fermentation9040341






