Abstract
Cytospora (Cytosporaceae, Diaporthales) is a fungal genus that usually inhabits plants as endophytes, saprobes, as well as pathogens. Species of this genus are characterized by possessing allantoid hyaline conidia and ascospores. Samples with typical Cytospora canker symptoms on Prunus davidiana, P. padus and Salix sp. were collected in Tibet and Xinjiang, China. Species were identified using both morphological and molecular approaches of combined loci of internal transcribed spacer region rDNA (ITS), the partial actin (act) region, RNA polymerase II second largest subunit (rpb2), the translation elongation factor 1-alpha (tef1) gene and the partial be-ta-tubulin (tub2) gene. Six isolates in the present study formed three distinct clades from previously known species. Cytospora hejingensis sp. nov. from Salix sp., C. jilongensis sp. nov. from P. davidiana and C. kunsensis from P. padus were proposed herein. The current study improves the understanding of species concept in Cytospora.
1. Introduction
Cytospora is a species-rich genus in family Cytosporaceae (order Diaporthales) and commonly inhabits plant tissues [1,2,3,4]. This genus was proposed in 1818 with four species, namely C. betulina, C. epimyces, C. resinae and C. ribis [5]. Another species C. chrysosperma was subsequently introduced [6] and later selected as the type species of this genus [7]. Cytospora can be different from the other diaporthalean genera by having allantoid hyaline conidia and ascospores [1,4,8,9,10].
Species of Cytospora were primarily identified and distinguished by their morphology and host [5,6,7]. However, recent studies employing molecular phylogeny revealed many cryptic species with similar morphology on the same host of known species of this genus [11,12,13,14,15]. For example, up to 28 Cytospora species were discovered from Eucalyptus spp. in South Africa with the help of DNA sequence evidence [2], eight from willow (Salix spp.) trees in China [16], six from Castanea mollissima in China [17], six from Populus hosts in China [18] and six from apple trees in Iran [19]. The taxonomy of Cytospora is currently more dependent on combined evidence of DNA sequence data, morphological features and ecology than species morphology and host associations [1,20].
Several species of Cytospora are reported to cause plant diseases including canker, wilt and dieback [21,22,23,24]. For example, C. carpobroti causes Carpobrotus edulis wilt disease in South Africa [21]; C. oleicola and C. olivarum are pathogenic to olives in the USA [22]; C. parasitica results in apple cankers in China [23]; and Cytospora pistaciae causes dieback and canker disease of pistachios in Italy [24]. There are still many cryptic species of Cytospora pathogenic to plants waiting for description.
In the present study, Cytospora canker symptoms were found from different tree hosts named Prunus davidiana, P. padus and Salix sp. in Tibet and Xinjiang, China. The aims of the present study were to identify the casual agents of the lesions, to introduce and describe new Cytospora species using both molecular and morphological approaches, and to discuss the species differences based on newly collected specimens.
2. Materials and Methods
2.1. Specimens and Strains
Investigations to collect fungal specimens were conducted in Tibet and Xinjiang during 2021 and 2022. During the surveys, dead and dying twigs and branches of tree hosts were checked manually, and then twigs and branches with obvious fungal fruiting bodies were recorded and collected. Samples were packed in paper bags and posted back for isolation.
Ascomata on branches of Prunus padus and Salix sp., and conidiomata on branches of P. davidiana were sectioned using sterile blades, and mucoid spore masses were removed and placed onto the surface of potato dextrose agar (PDA; potato, 200 g; glucose, 20 g; agar, 20 g; distilled water, to complete 1000 mL) media using sterile insect needles. Then, plates were incubated at 25 °C in darkness until spores germinated. Pieces of mycelium were cut and removed and placed onto a new PDA plate under a stereomicroscope to obtain the pure strains. Specimens and isolates were preserved in the China Forestry Culture Collection Center (CFCC; http://cfcc.caf.ac.cn/ (accessed on 2 January 2024)).
2.2. Morphological Observations
The Cytospora species observations were based on ascomata and conidiomata naturally formed on twigs and branches of Prunus davidiana, P. padus and Salix sp. The sexual and asexual fruiting bodies were sectioned using sterile blades and photographed using the Leica stereomicroscope (M205) (Leica Microsystems, Wetzlar, Germany). The asci, ascospores, conidiophores, conidiogenous cells and conidia were measured and photographed by a Nikon Eclipse 80i microscope (Nikon Corporation, Tokyo, Japan). The colony characteristics were observed and recorded on PDA plates at 25 °C in darkness.
2.3. DNA Extraction and Amplification
The total genomic DNA of Cytospora species were obtained from colonies growing on PDA plates by using the CTAB method [25]. The internal transcribed spacer region rDNA (ITS), the partial actin (act) region, RNA polymerase II second largest subunit (rpb2), the translation elongation factor 1-alpha (tef1) gene and the partial be-ta-tubulin (tub2) gene were amplified using primer pairs ITS1/ITS4, ACT512F/ACT783R, fRPB2-5f/fRPB2-7cR, 983F/2218R, Bt2a/Bt2b, respectively [26,27,28,29,30]. These regions were amplified as follows: an initial denaturation step of 5 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 50 s at 52 °C (ITS), 54°C (tef1 and tub2), 55 °C (rpb2) or 58 °C (act), and 1 min at 72 °C, and a final elongation step of 7 min at 72 °C. The polymerase chain reaction products were sequenced using an ABI PRISM 3730XL DNA Analyser with a BigDye Terminator Kit v.3.1 (Invitrogen, Waltham, MA, USA) at the Shanghai Invitrogen Biological Technology Company Limited (Beijing, China).
2.4. Molecular Phylogeny
Sequences obtained in the present study were preliminarily identified by the BLAST search to confirm their classification. The referenced sequences of Cytospora were collected from recent publications (Table 1) and downloaded [1,24,25]. Strain CBS 160.32 (species Diaporthe vaccinii) was selected as the outgroup taxon. The five individual loci ITS, act, rpb2, tef1 and tub2 were aligned using MAFFT v. 6.0 and manually adjusted using MEGA v. 6.0 [31,32]. Then, five loci were combined and analyzed based on maximum likelihood (ML) and Bayes methods in the CIPRES Science Gateway platform [33]. The GTR substitution model was employed and 1000 non-parametric bootstrap replicates were set for ML phylogenic analysis. Four simultaneous Markov Chain runs for 1,000,000 generations were set during Bayesian analysis. The resulting trees were visualized in FigTree v. 1.4.0 and edited using Adobe Illustrator 2020.

Table 1.
Strains and their GenBank accession numbers used in this study.
3. Results
3.1. Phylogeny
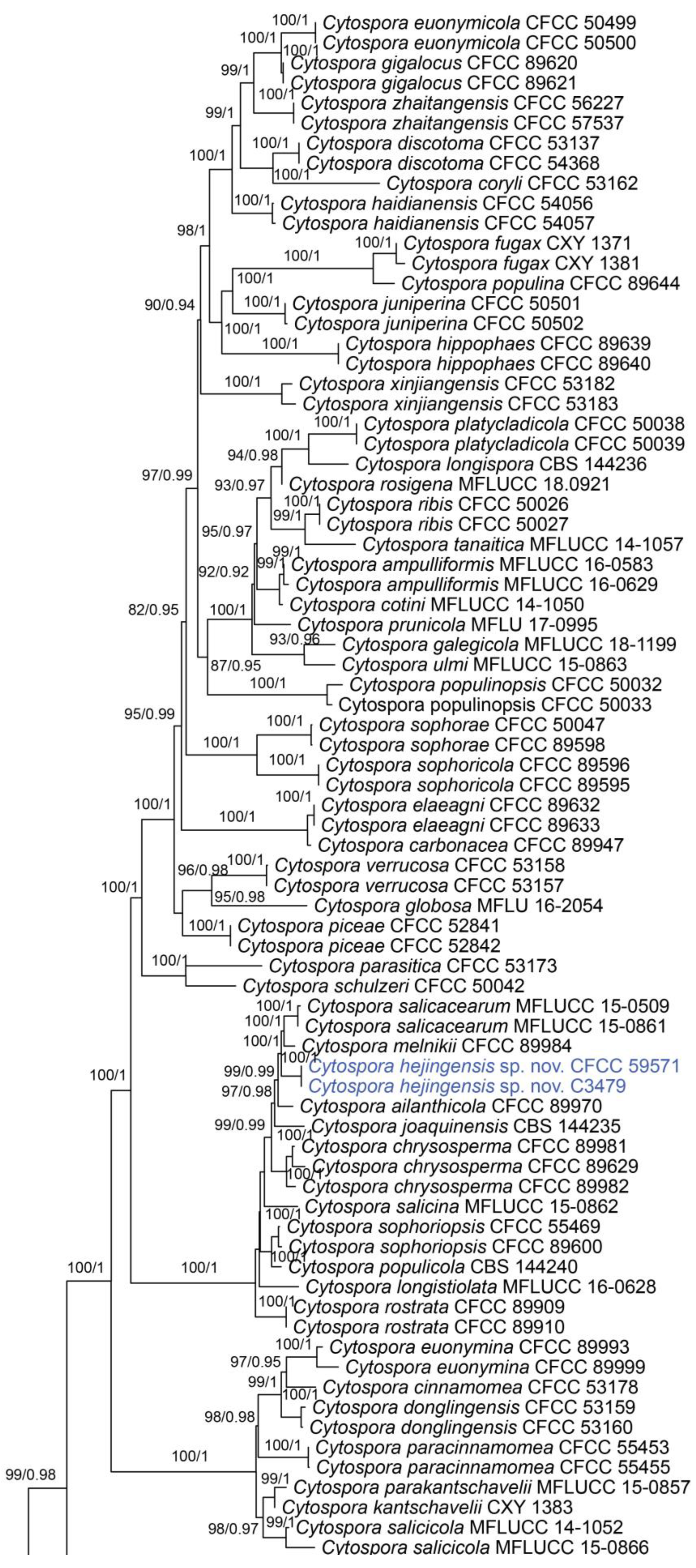
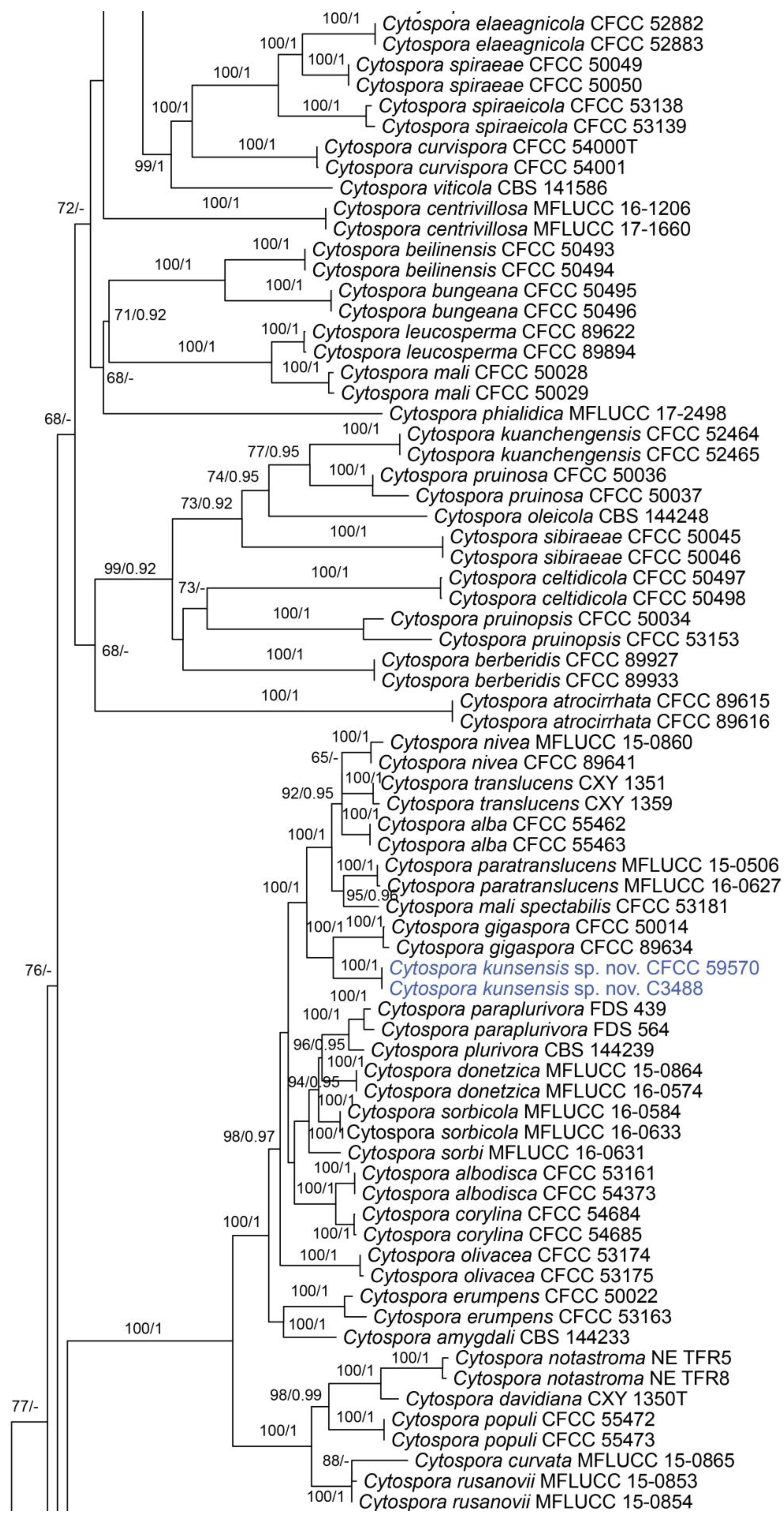
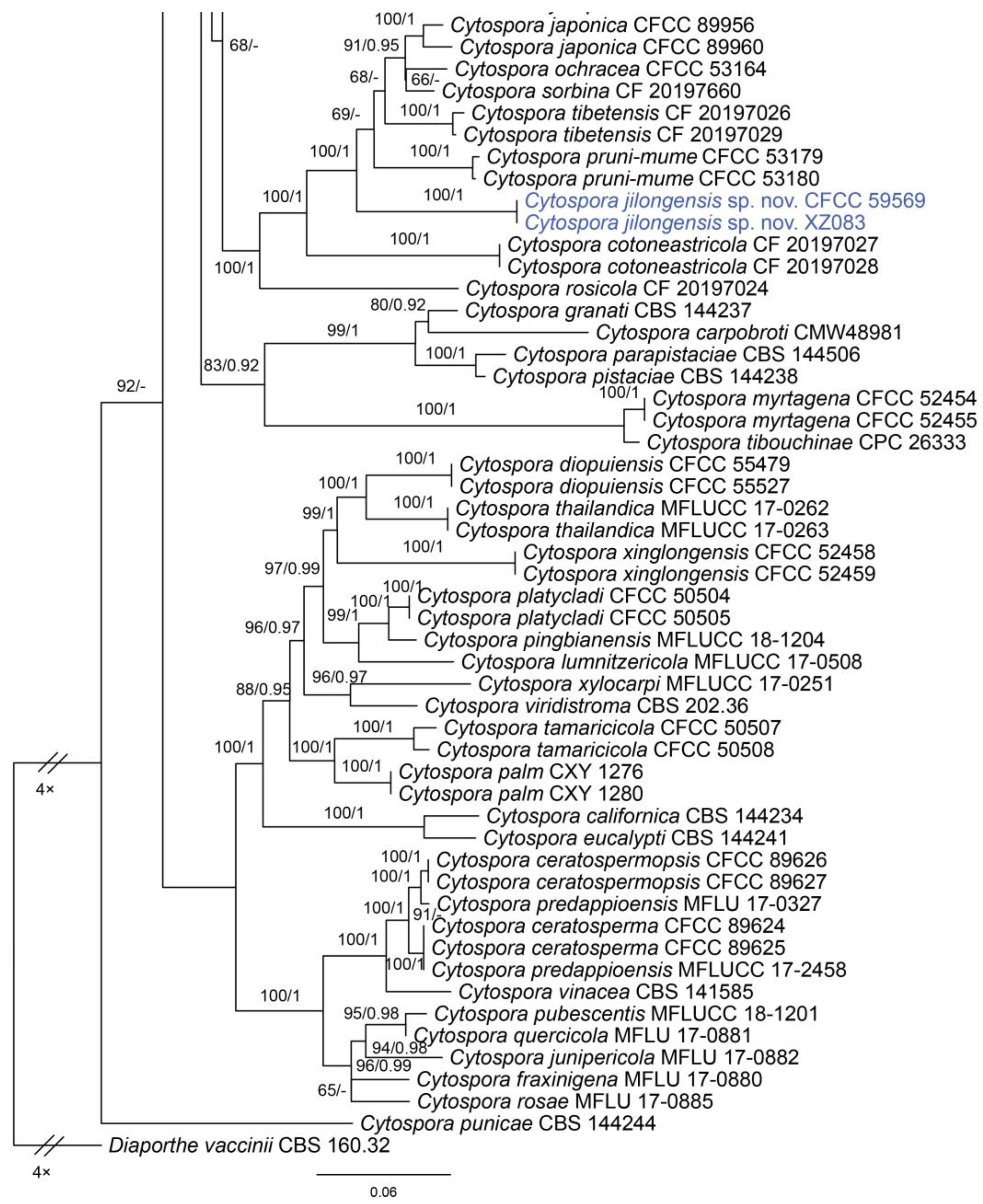
In the phylogenetic analysis, the combined dataset of ITS, act, rpb2, tef1 and tub2 consisted of 202 strains. The final alignment comprised 2561 characters including 588 characters in ITS, 211 characters in act, 617 characters in rpb2, 536 characters in tef1 and 609 characters tub2. The final ML optimization likelihood value of the best RAxML tree was −48,006.19, and the matrix had 1534 distinct alignment patterns, with 29.35% undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.242905, C = 0.286434, G = 0.242250 and T = 0.228411; substitution rates AC = 1.374316, AG = 3.693698, AT = 1.471034, CG = 0.981185, CT = 6.248472 and GT = 1.0; and gamma distribution shape parameter α = 0.321644. The topology of our phylogenetic tree is nearly identical to previous publications. The topology of isolates from the present study in the RAxML and Bayesian analyses were congruent. Isolates CFCC 59571 and C3479 formed a distinct clade to CFCC 89984 (C. melnikii), MFLUCC 15-0509 and MFLUCC 15-0861 (C. salicacearum) with high support values (BS = 100, BPP = 1). Isolates CFCC 59570 and C3488 formed a clade close to CFCC 50014 and CFCC 89634 (C. gigaspora) with full support values (BS = 100, BPP = 1). Isolates CFCC 59570 and C3488 clustered together with CFCC 89956 and CFCC 89960 (C. japonica), CFCC 53164 (C. ochracea), CF 20197660 (C. sorbina), CF 20197026 and CF 20197029 (C. tibetensis), and CFCC 53179 and CFCC 53180 (C. pruni-mume) supported by high values (BS = 100, BPP = 1). Hence, six isolates from the present study formed three new clades distinct from previously known species named Cytospora hejingensi sp. nov., C. jilongensis sp. nov. and C. kunsensis sp. nov. (Figure 1).
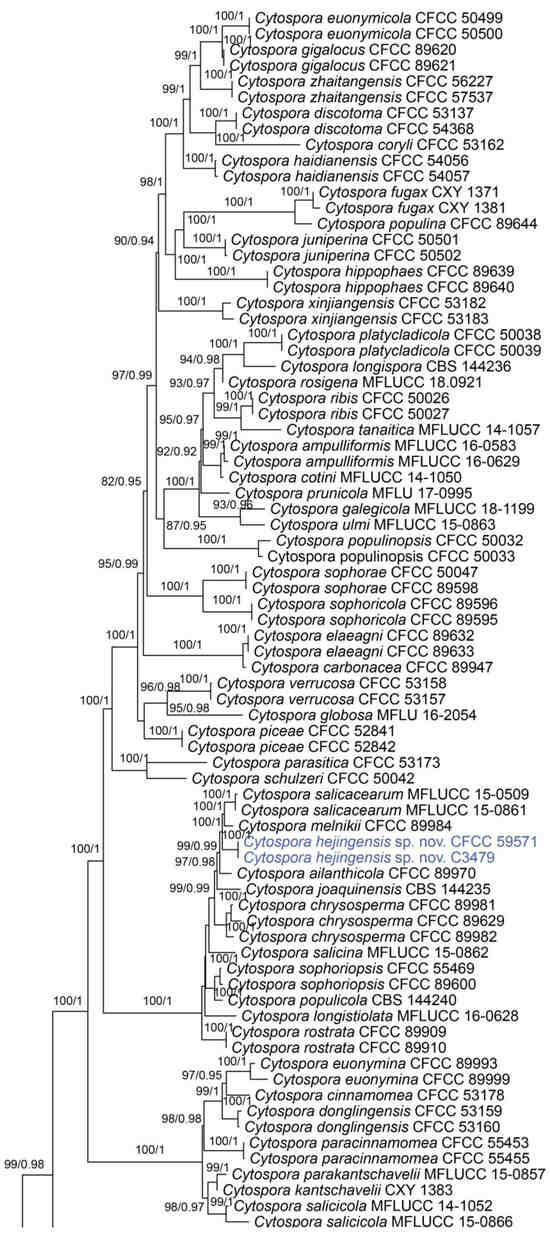

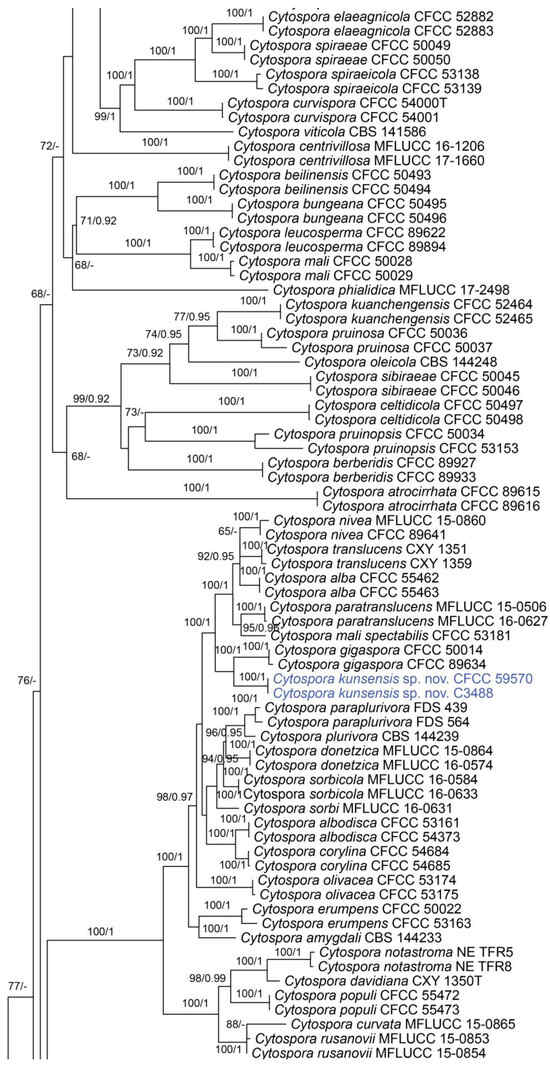
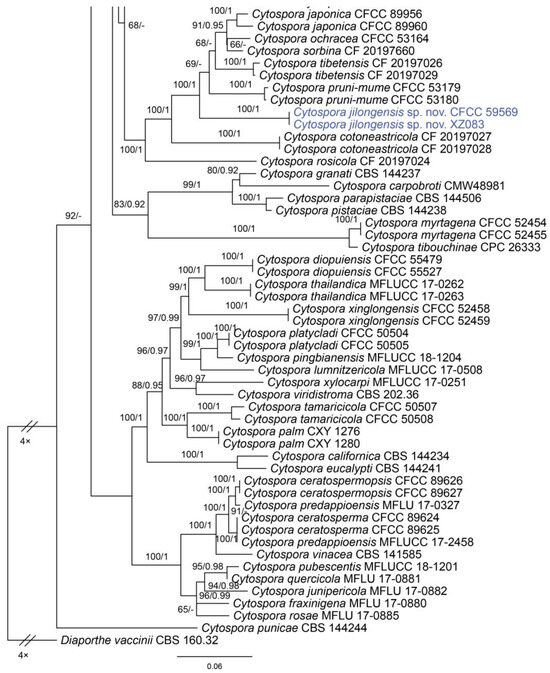
Figure 1.
Phylogram of Cytospora resulting from a maximum likelihood analysis, based on a combined matrix of ITS, act, rpb2, tef1 and tub2. Numbers above the branches indicate ML bootstraps (left, ML BS ≥ 50%) and Bayesian posterior probabilities (right, BPP ≥ 0.90). The tree is rooted with Diaporthe vaccinii (CBS 160.32). Isolates obtained from the present study are marked in blue.
3.2. Description of Cytospora hejingensis sp. nov. from Salix sp.
Cytospora hejingensis R. Ma & Ning Jiang, sp. nov.
MycoBank: MB851771
Etymology: named after the collection site of the holotype, Hejing County.
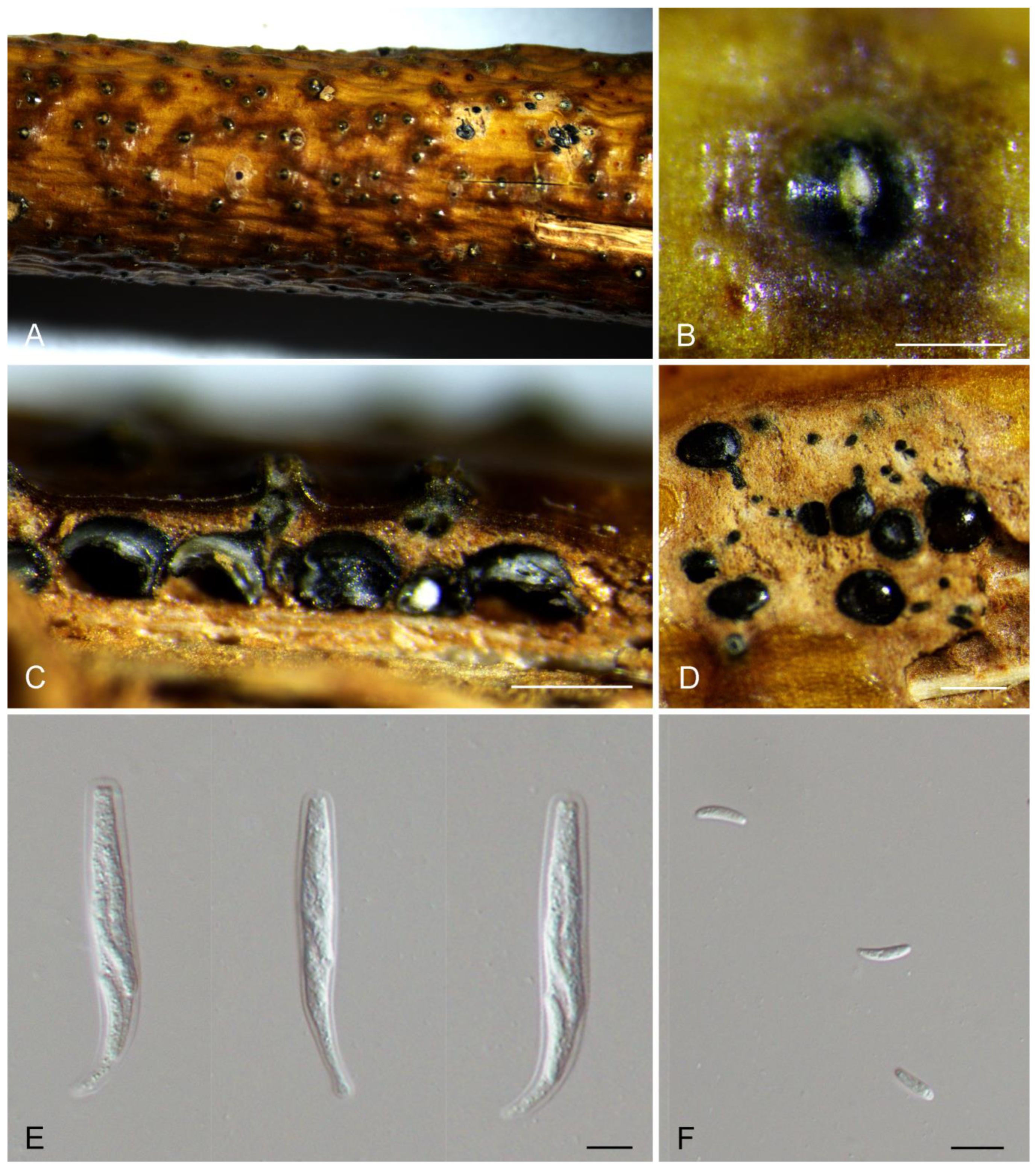
Description: Associated with branch and twig canker disease of Salix sp. Sexual morph: Ascostromata immersed in the bark, erumpent through the bark surface, scattered, (400–)650–900(–1250) μm diam., with 4–9 perithecia arranged irregularly. Conceptacle absent. Ectostromatic disc inconspicuous, usually surrounded by tightly aggregated ostiolar necks, (100–)150–250(–350) μm diam. Ostioles numerous, black, concentrated, arranged irregularly in a disc, (35–)50–65(–90) μm diam. Perithecia black, spherical, arranged circularly or irregularly, (120–)150–250(–300) μm diam. Asci free, clavate, (38–)45–70(–77) × (7–)8.5–10.5(–12.5) μm, 8-spored. Ascospores biseriate, allantoid, thin-walled, hyaline, aseptate, (6.5–)7–8(–9) × 2–2.5 μm. Asexual morph: undetermined.
Culture characteristics: colonies on PDA flat, spreading, with flocculent mycelium, initially white to grey, secreting a dark green to black pigment in culture medium after 10 days, reaching a 90 mm diameter after 15 days at 25 °C in the dark.
Materials examined: China, Xinjiang Uygur Autonomous Region, Bayingolin Mongol Autonomous Prefecture, Hejing County, Kunse Forest Park, on cankered twigs and branches of Salix sp., 24 July 2021, Rong Ma (XJAU 3488, holotype); ex-type culture CFCC 59571; ibid. (culture C3488).
Notes: Cytospora hejingensis from Salix sp. in China is phylogenetically close to C. melnikii from Malus domestica in Russia and C. salicacearum from Salix alba in Russia (Figure 1). C. hejingensis is only known in sexual morph, and the other two species in asexual morph. Hence, it is impossible to compare them in morphology. However, C. hejingensis differs from C. melnikii and C. salicacearum by sequence data (22/560 in ITS, 35/211 in act, 36/617 in rpb2 and 27/306 in tef1 from C. melnikii; 25/560 in ITS, 37/211 in act and 26/617 in rpb2 from C. salicacearum) [34].
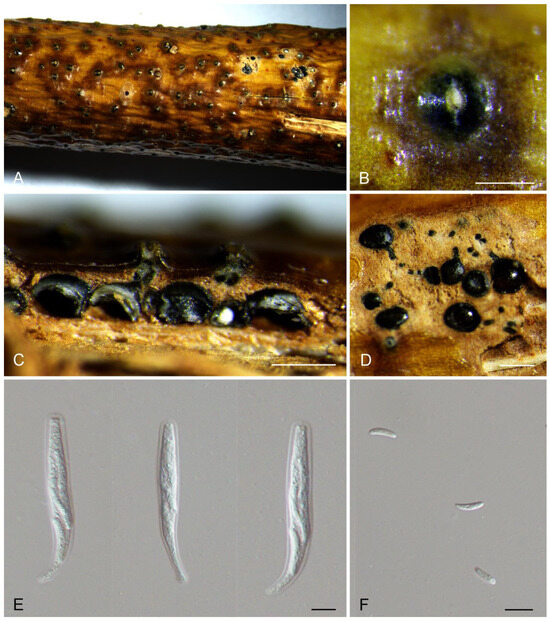
Figure 2.
Morphology of Cytospora hejingensis from Salix sp. (A,B) Ascomata formed on branches. (C) Longitudinal section through the ascomata. (D) Transverse section of ascomata. (E) Asci. (F) Ascospores. Scale bars: (B) = 500 μm; (C) = 200 μm; (D) = 300 μm; (E,F) = 10 μm.
3.3. Description of Cytospora jilongensis sp. nov. from Prunus davidiana
Cytospora jilongensis R. Ma & Ning Jiang, sp. nov.
MycoBank: MB851772
Etymology: named after the collection site of the holotype, Jilong County.
Description: Associated with branch canker disease of Prunus davidiana. Sexual morph: undetermined. Asexual morph: Pycnidial stromata ostiolated, semi-immersed in the host bark, scattered, pulvinate, with multiple locules. Conceptacle dark brown, circular surrounded stromata. Ectostromatic grey, circular to ovoid, (100–)180–240(–370) μm diam., with one ostiole per disc. Ostioles dark, at the same level as the disc, (30–)50–75(–95) μm diam. Locule numerous, arranged circularly or elliptically with independent walls, (250–)400–500(–750) μm diam. Peridium comprising few layers of cells of textura angularis, brown to dark brown. Conidiophores hyaline, branched, thin-walled, filamentous. Conidiogenous cells enteroblastic polyphialidic, 7.5–18.5 × 1.5–2.5 μm. Conidia hyaline, allantoid, smooth, aseptate, thin-walled, (9.3–)10.2–11.6(–12.5) × 2.6–3.2 μm.
Culture characteristics: colonies on PDA flat, spreading, with moderate flocculent mycelium, initially white, becoming orange after 10 days, reaching a 90 mm diameter after 25 days at 25 °C in the dark.
Materials examined: China, Tibet Tibetan Autonomous Region, Shigatse City, Jilong County, Jilong Town, on cankered branches of Prunus davidiana, 12 August 2022, Jin Peng, Jiang Ning and Liu Min (CAF800087, holotype); ex-type culture CFCC 59569; ibid. (culture XZ083).
Notes: Cytospora jilongensis from Prunus davidiana is phylogenetically close to C. japonica from P. cerasifera and P. persica, C. ochracea from Cotoneaster sp., C. sorbina from Sorbus tianschanica, C. tibetensis from Cotoneaster sp. and C. pruni-mume from Prunus mume (Figure 1). However, C. jilongensis (10.2–11.6 × 2.6–3.2 μm) differs from C. japonica (6.5–8.5 × 1.5–2 μm), C. ochracea (8.5–9.0 × 1.5–2.5 μm), C. sorbina (4.5–5.5 × 1–1.5 μm), C. tibetensis (5.0–5.5 × 1.5–2 μm) and C. pruni-mume (5.5–6.5 × 1.5–2 μm) in conidial size and hosts [1,14].
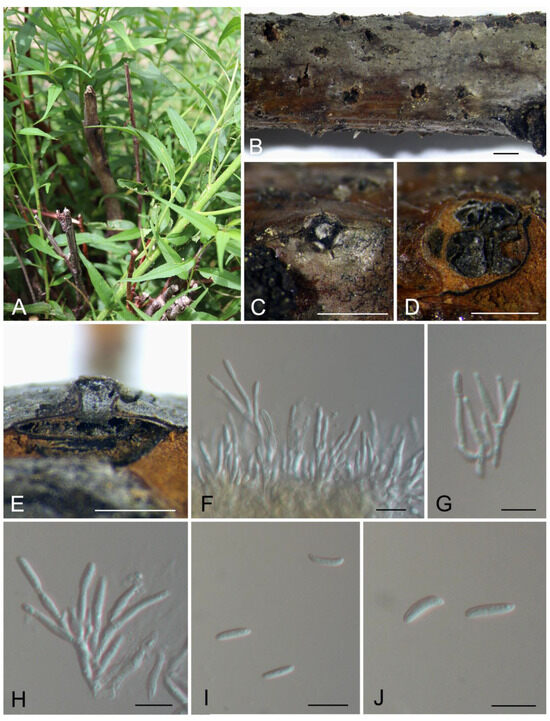

Figure 3.
Morphology of Cytospora jilongensis from Prunus davidiana. (A) Symptoms of canker disease on the host. (B,C) Conidiomata formed on branches. (D) Transverse section through the conidioma. (E) Longitudinal section through the conidioma. (F–H) Conidiophores and conidiogenous cells. (I,J) Conidia. Scale bars: (B) = 2 mm; (C,D) = 1 mm; (E) = 800 μm; (F–J) = 10 μm.
3.4. Description of Cytospora kunsensis sp. nov. from Prunus padus
Cytospora kunsensis R. Ma & Ning Jiang, sp. nov.
MycoBank: MB851773
Etymology: named after the collection site of the holotype, Kunse Forest Park.
Description: Associated with branch and twig canker disease of Prunus padus. Sexual morph: Ascostromata immersed in the bark, erumpent through the bark surface, scattered, (750–)950–1100(–1350) μm diam., with 5–11 perithecia arranged circularly. Conceptacle absent. Ectostromatic disc white, surrounded by tightly aggregated ostiolar necks, (100–)150–300(–350) μm diam. Ostioles numerous, black, concentrated, arranged circularly in a disc, (40–)50–75(–90) μm diam. Perithecia black, spherical, arranged circularly or irregularly, (180–)250–350(–420) μm diam. Asci free, clavate, (38–)48–80(–86) × (7.5–)9–12(–13.5) μm, eight-spored. Ascospores biseriate, allantoid, thin-walled, hyaline, aseptate, (10–)12.5–17(–19.5) × 2–2.5 μm. Asexual morph: undetermined.
Culture characteristics: colonies on PDA flat, spreading, with flocculent mycelium, white, with a dark grey color in the center, fast growing, reaching a 90 mm diameter after 7 days and forming abundant black ascomata after 25 days at 25 °C.
Materials examined: China, Xinjiang Uygur Autonomous Region, Bayingolin Mongol Autonomous Prefecture, Hejing County, Kunse Forest Park, on cankered twigs and branches of Prunus padus, 24 July 2021, Rong Ma (XJAU 3479, holotype); ex-type culture CFCC 59570; ibid. (culture C3479).
Notes: Cytospora kunsensis from Prunus padus is phylogenetically close to C. gigaspora from Salix psammophila (Figure 1). However, C. kunsensis can be distinguished from C. gigaspora by sequence data (19/548 in ITS, 32/211 in act, 56/617 in rpb2, 36/303 in tef1 and 42/421 in tub2) [11].
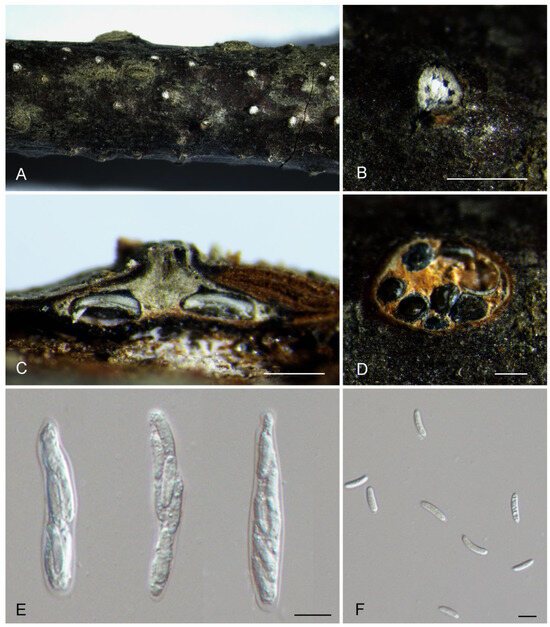

Figure 4.
Morphology of Cytospora kunsensis from Prunus padus. (A,B) Ascomata formed on branches. (C) Longitudinal section through the ascomata. (D) Transverse section of ascomata. (E) Asci. (F) Ascospores. Scale bars: (B) = 500 μm; (C,D) = 300 μm; (E,F) = 10 μm.
4. Discussion
In the present study, samples of Cytospora with fruiting bodies were collected from Xinjiang and Tibet, and identified based on both morphological and phylogenetical approaches of combined ITS, act, rpb2, tef1 and tub2 loci. We proposed three new species, i.e., Cytospora hejingensis sp. nov. from Salix sp., C. jilongensis sp. nov. from P. davidiana and C. kunsensis from P. padus.
Of the new species introduced in the current study, two taxa (C. jilongensis and C. kunsensis) were isolated from the plant genus Prunus. Hence, a total of nine species of Cytospora were found in host genus Prunus, where the previous seven species are C. cinnamomea, C. erumpens, C. japonica, C. leucostoma, C. olivacea, C. populinopsis and C. pruni-mume [14]. C. kunsensis is distinguished from C. populinopsis in eight-spored asci, and these two species are only known in sexual morph [1]. The other seven species are known in asexual species with similar conidial morphology but different sequence data of ITS, act, rpb2, tef1 and tub2 loci. The example of Cytospora species from Prunus implies that DNA sequence data are necessary to separate species during pathogen identifications.
Another example is the Cytospora species from the host genus Salix. Until now, over 10 species of Cytospora were discovered from the host genus Salix, including one species Cytospora hejingensis introduced in the current study [35]. Most of them are confirmed to be pathogens associated with canker diseases [35]. The new species from the present study needs a pathogenicity test to evaluate its virulence to willow trees in the future.
In the traditional classification and identification of species in Cytospora, spore morphology and host information are the most important evidence to identify Cytospora species [5,6,7]. However, by using the molecular data, many cryptic species with the same hosts and similar spore morphology were recently revealed [1,14,15,35]. The molecular classification system for Cytospora based on morphology, phylogeny and host information is more scientific than that mainly based on morphology before.
Author Contributions
Conceptualization, S.W. and R.M.; methodology, N.J.; software, S.W.; validation, S.W., N.J. and R.M.; formal analysis, S.W., N.J. and R.M.; investigation, S.W., N.J. and R.M.; resources, S.W., N.J. and R.M.; data curation, S.W., N.J. and R.M.; writing—original draft preparation, S.W., N.J. and R.M.; writing—review and editing, S.W., N.J. and R.M.; visualization, S.W., N.J. and R.M.; supervision, S.W., N.J. and R.M.; project administration, R.M.; funding acquisition, R.M. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Natural Science Foundation of China, grant number 31960316.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All sequence data are available in NCBI GenBank (Table 1).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Fan, X.L.; Bezerra, J.D.P.; Tian, C.M.; Crous, P.W. Cytospora (Diaporthales) in China. Persoonia 2020, 45, 1–45. [Google Scholar] [CrossRef]
- Adams, G.C.; Wingfield, M.J.; Common, R.; Roux, J. Phylogenetic relationships and morphology of Cytospora species and related teleomorphs (Ascomycota, Diaporthales, Valsaceae) from Eucalyptus. Stud. Mycol. 2005, 52, 1–144. [Google Scholar]
- Norphanphoun, C.; Raspé, O.; Jeewon, R.; Wen, T.C.; Hyde, K.D. Morphological and phylogenetic characterisation of novel Cytospora species associated with mangroves. MycoKeys 2018, 38, 93–120. [Google Scholar] [CrossRef] [PubMed]
- Senanayake, I.C.; Crous, P.W.; Groenewald, J.Z.; Maharachchikumbura, S.S.; Jeewon, R.; Phillips, A.J.; Bhat, J.D.; Perera, R.H.; Li, Q.R.; Li, W.J.; et al. Families of Diaporthales based on morphological and phylogenetic evidence. Stud. Mycol. 2017, 86, 217–296. [Google Scholar] [CrossRef] [PubMed]
- Ehrenberg, C.G. Sylvae Mycologicae Berolinenses; Formis Theophili Bruschcke: Berlin, Germany, 1818. [Google Scholar]
- Fries, E.M. Systema Mycologicum; Ex Officina Berlingiana: Lund, Sweden, 1823; Volume 2, pp. 275–620. [Google Scholar]
- Donk, M.A. Nomina conservanda proposita 1. Proposals Fungi Deuteromycetes Regnum Veg. 1964, 34, 7–15. [Google Scholar]
- Jiang, N.; Fan, X.L.; Crous, P.W.; Tian, C.M. Species of Dendrostoma (Erythrogloeaceae, Diaporthales) associated with chestnut and oak canker diseases in China. MycoKeys 2019, 48, 67–96. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Du, Z.; Bezerra, J.D.; Tian, C. Taxonomic circumscription of melanconis-like fungi causing canker disease in China. MycoKeys 2018, 42, 89–124. [Google Scholar] [CrossRef]
- Jiang, N.; Fan, X.L.; Tian, C.M. Identification and pathogenicity of Cryphonectriaceae species associated with chestnut canker in China. Plant Pathol. 2019, 68, 1132–1145. [Google Scholar] [CrossRef]
- Fan, X.L.; Hyde, K.D.; Yang, Q.; Liang, Y.M.; Ma, R.; Tian, C.M. Cytospora species associated with canker disease of three anti-desertification plants in northwestern China. Phytotaxa 2015, 1974, 227–244. [Google Scholar] [CrossRef]
- Yang, Q.; Fan, X.L.; Crous, P.W.; Liang, Y.M.; Tian, C.M. Cytospora from Ulmus pumila in Northern China. Mycol. Prog. 2015, 14, 74. [Google Scholar] [CrossRef]
- Zhu, H.Y.; Pan, M.; Bonthond, G.; Tian, C.M.; Fan, X.L. Diaporthalean fungi associated with canker and dieback of trees from Mount Dongling in Beijing, China. MycoKeys 2019, 59, 67–94. [Google Scholar] [CrossRef] [PubMed]
- Pan, M.; Zhu, H.; Bonthond, G.; Tian, C.M.; Fan, X.L. High diversity of Cytospora associated with canker and dieback of Rosaceae in China, with 10 new species described. Front. Plant Sci. 2020, 11, 690. [Google Scholar] [CrossRef]
- Pan, M.; Zhu, H.; Tian, C.M.; Huang, M.; Fan, X.L. Assessment of Cytospora isolates from conifer cankers in China, with the descriptions of four new Cytospora species. Front. Plant Sci. 2021, 12, 636460. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Pan, M.; Gao, H.; Tian, C.; Fan, X. The potential fungal pathogens of Euonymus japonicus in Beijing, China. J. Fungi 2023, 9, 271. [Google Scholar] [CrossRef] [PubMed]
- Jiang, N.; Yang, Q.; Fan, X.L.; Tian, C.M. Identification of six Cytospora species on Chinese chestnut in China. MycoKeys 2020, 62, 1–25. [Google Scholar] [CrossRef]
- Lin, L.; Pan, M.; Bezerra, J.D.; Tian, C.; Fan, X. Re-evaluation of the fungal diversity and pathogenicity of Cytospora species from Populus in China. Plant Dis. 2023, 107, 83–96. [Google Scholar] [CrossRef] [PubMed]
- Mehrabi, M.E.; Mohammadi, G.E.; Fotouhifar, K.B. Studies on Cytospora canker disease of apple trees in Semirom region of Iran. J. Agric. Technol. 2011, 7, 967–982. [Google Scholar]
- Shang, Q.J.; Hyde, K.D.; Camporesi, E.; Maharachchikumbura, S.S.N.; Norphanphoun, C.; Brooks, S.; Liu, J.K. Additions to the genus Cytospora with sexual morph in Cytosporaceae. Mycosphere 2020, 11, 189–224. [Google Scholar] [CrossRef]
- Jami, F.; Marincowitz, S.; Crous, P.W.; Jacobsohn, A.; Wingfield, M.J. A new Cytospora species pathogenic on Carpobrotus edulis in its native habitat. Fungal Syst. Evol. 2018, 2, 37–43. [Google Scholar] [CrossRef]
- Úrbez-Torres, J.R.; Lawrence, D.P.; Hand, F.P.; Trouillas, F.P. Olive twig and branch dieback in California caused by Cytospora oleicola and the newly described species Cytospora olivarum sp. nov. Plant Dis. 2020, 104, 1908–1917. [Google Scholar] [CrossRef]
- Ma, R.; Liu, Y.M.; Yin, Y.X.; Tian, C.M. A canker disease of apple caused by Cytospora parasitica recorded in China. For. Pathol. 2018, 48, e12416. [Google Scholar] [CrossRef]
- Aiello, D.; Polizzi, G.; Gusella, G.; Fiorenza, A.; Guarnaccia, V. Characterization of Eutypa lata and Cytospora pistaciae causing die-back and canker of pistachio in Italy. Phytopathol. Mediterr. 2019, 58, 699–706. [Google Scholar]
- Doyle, J.J. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 3, 553–556. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among Ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef] [PubMed]
- Rehner, S.A. Primers for Elongation Factor 1-Alpha (EF1-Alpha). 2001. Available online: http://ocid.nacse.org/research/deephyphae/EF1primer.pdf (accessed on 13 May 2023).
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microb. 1995, 61, 1323–1330. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 304, 772–780. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop, GCE 2010, New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Norphanphoun, C.; Wen, T.C.; Hyde, K.D.; Doilom, M.; Daranagama, D.A.; Phookamsak, R.; Bulgakov, T.S. Revisiting the genus Cytospora and allied species. Mycosphere 2017, 8, 51–97. [Google Scholar] [CrossRef]
- Lin, L.; Pan, M.; Tian, C.; Fan, X. Fungal richness of Cytospora species associated with willow canker disease in China. J. Fungi 2022, 8, 377. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).