Pragmatic Models for Detection of Hypertension Using Ballistocardiograph Signals and Machine Learning
Abstract
1. Introduction
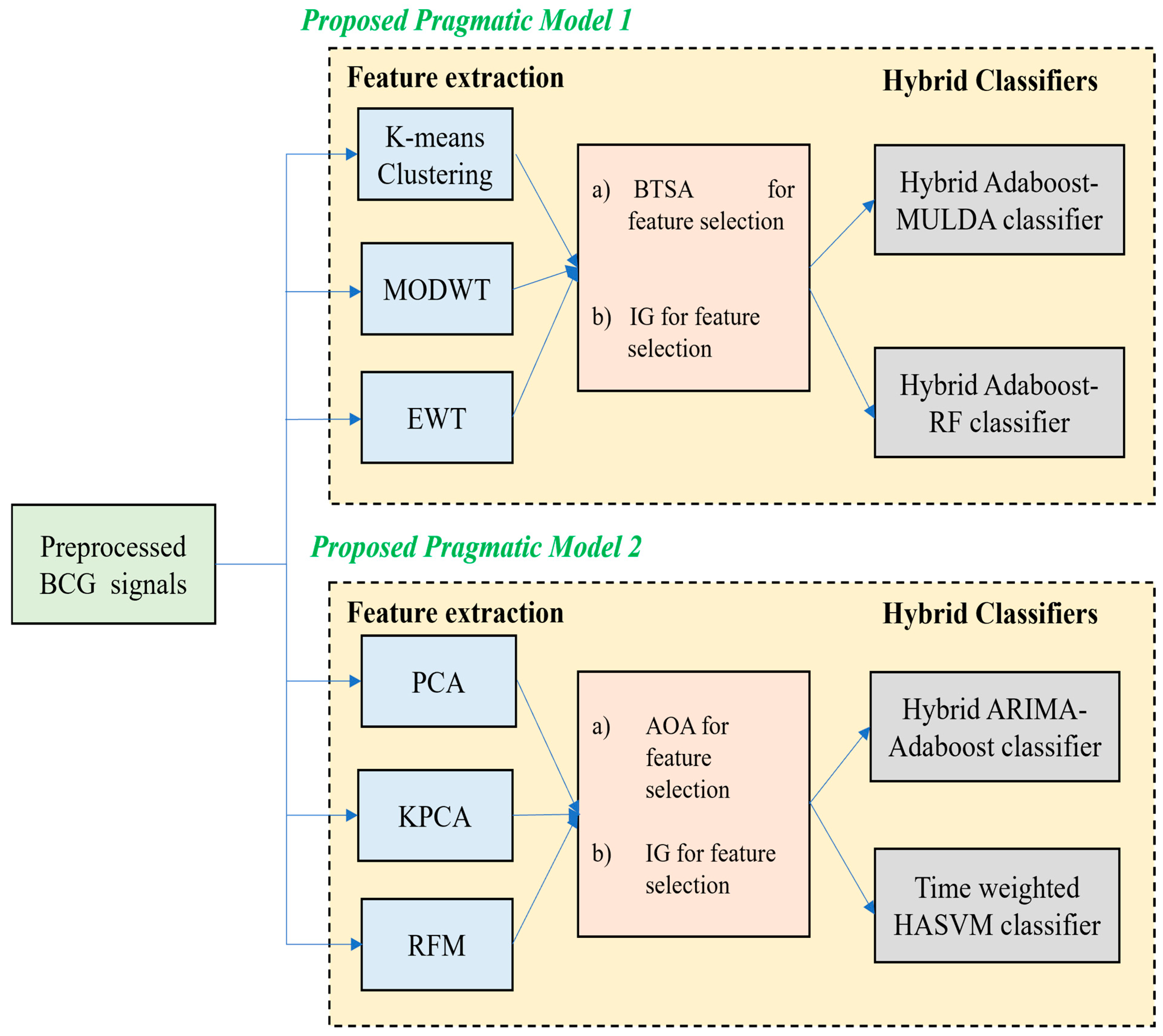
2. Proposed Pragmatic Model 1
2.1. Feature Extraction Using K-Means Clustering, MODWT and EWT
2.1.1. K-Means Clustering
2.1.2. Maximal Overlap Discrete Wavelet Transforms (MODWT)
2.1.3. Empirical Wavelet Transform (EWT)
2.2. Feature Selection Using Binary Tunicate Swarm Algorithm (BTSA)
2.3. Classification Using Two Hybrid Models
2.3.1. Hybrid AdaBoost–MULDA Classifier
2.3.2. Hybrid AdaBoost–RF Classifier
| Algorithm 1: AdaBoost Classification |
| Assume: Start For Weak classifier training phase using the distribution Obtain weak hypothesis Compute error Choose Updating all the parameters where denotes the normalization factors and it is presumed as Output the hypothesis |
| Algorithm 2: RF Classification |
| Input: Training samples Weight initialization by random values; Compute the sum of inputs : Multiply with weights for the hidden layer node Project the hidden output for hidden layer node The input for the output layer nodes is expressed as Project the output at output nodes The difference between the predicted output and actual output is computed so that the error rate is obtained The error rate is detected from hidden layer to output layer nodes The updating of the network weights is performed The final output is the difference between the predicted output and actual output |
| Algorithm 3: Hybrid AdaBoost–RF classification |
| Given Assign For Train the weak classifiers by using distribution Weak hypothesis is fetched Compute error Choose Increment the hypothesis if required Compute the sum of the inputs as Multiply with for the hidden layer nodes Compute the hidden output for hidden layer node Input for the output layer node is given as Compute the distribution as where the normalization factor is expressed as Output the final hypothesis |
3. Proposed Pragmatic Model 2
3.1. Feature Extraction Using PCA, KPCA and RFM Techniques
3.1.1. PCA and KPCA
3.1.2. Random Feature Mapping Technique
3.2. Feature Selection Using Information Gain (IG)
3.3. Feature Selection Using AOA
3.4. Classification Using Two Hybrid Models
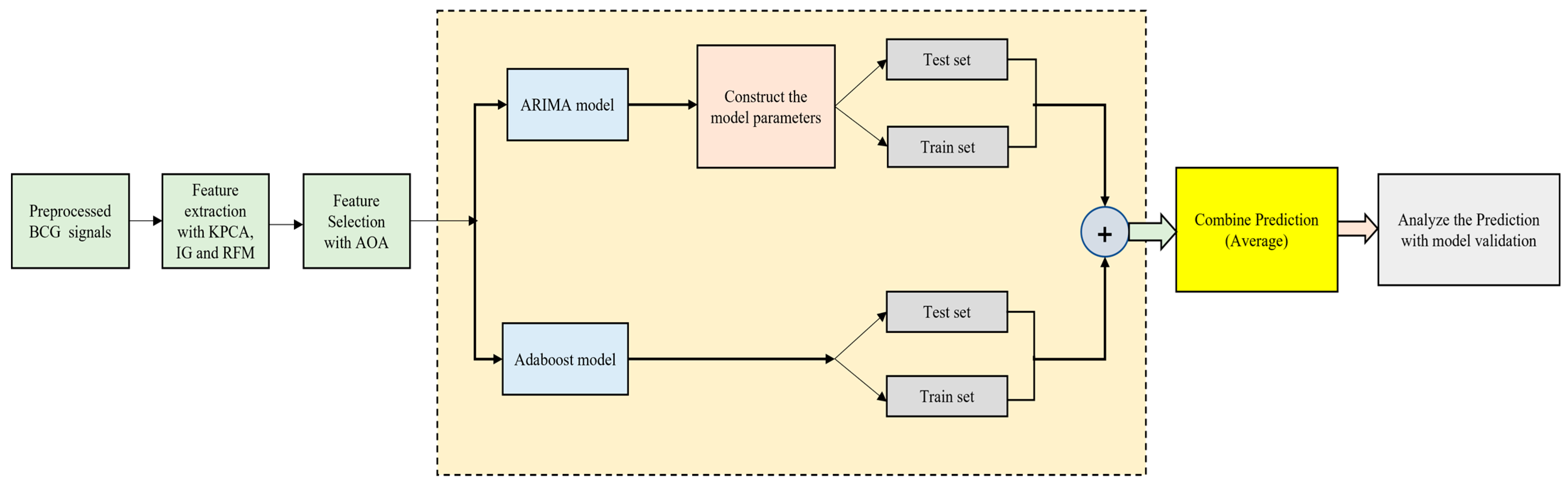
3.4.1. ARIMA–AdaBoost Hybrid Model
3.4.2. Time-Weighted Hybrid AdaBoost–Support Vector Machine (TW-HASVM) Model
4. Results and Discussion
4.1. Comparison with Previous Works
4.2. Limitations of This Work
5. Conclusions and Future Works
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Starr, I.; Schroeder, H.A. Ballistocardiogram. II. Normal standards, abnormalities commonly found in diseases of the heart and circulation, and their significance. J. Clin. Investig. 1940, 19, 437–450. [Google Scholar] [CrossRef]
- Vogt, E.; MacQuarrie, D.; Neary, J.P. Using ballistocardiography to measure cardiac performance: A brief review of its history and future significance. Clin. Physiol. Funct. Imaging 2012, 32, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Scarborough, W.R.; Talbot, S.A.; Braunstein, J.R.; Rappaport, M.B.; Dock, W.; Hamilton, W.; Smith, J.E.; Nickerson, J.L.; Starr, I. Proposals for ballistocardiographic nomenclature and conventions: Revised and extended. Circulation 1956, 14, 435–450. [Google Scholar] [CrossRef] [PubMed]
- Ashouri, H.; Hersek, S.; Inan, O.T. Universal pre-ejection period estimation using seismocardiography: Quantifying the effects of sensor placement and regression algorithms. IEEE Sensors J. 2017, 18, 1665–1674. [Google Scholar] [CrossRef]
- Javaid, A.Q.; Ashouri, H.; Dorier, A.; Etemadi, M.; Heller, J.A.; Roy, S.; Inan, O.T. Quantifying and reducing motion artifacts in wearable seismocardiogram measurements during walking to assess left ventricular health. IEEE Trans. Biomed. Eng. 2016, 64, 1277–1286. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.H.; Lee, H.J.; Yoon, H.N.; Jung, D.W.; Lee, Y.J.G.; Lee, Y.J.; Jeong, D.U.; Park, K.S. Unconstrained sleep apnea monitoring using polyvinylidene fluoride film-based sensor. IEEE Trans. Biomed. Eng. 2014, 61, 2125–2134. [Google Scholar] [CrossRef]
- Wang, F.; Zou, Y.; Tanaka, M.; Matsuda, T.; Chonan, S. Unconstrained cardiorespiratory monitor for premature infants. Int. J. Appl. Electromagn. Mech. 2007, 25, 469–475. [Google Scholar] [CrossRef]
- Kortelainen, J.M.; van Gils, M.; Pärkkä, J. Multichannel bed pressure sensor for sleep monitoring. In Proceedings of the 2012 Computing in Cardiology, Krakow, Poland, 9–12 September 2012; pp. 313–316. [Google Scholar]
- Katz, Y.; Karasik, R.; Shinar, Z. Contact-free piezo electric sensor used for real-time analysis of inter beat interval series. In Proceedings of the 2016 Computing in Cardiology Conference (CinC), Vancouver, BC, Canada, 11–14 September 2016; pp. 769–772. [Google Scholar]
- Junnila, S.; Akhbardeh, A.; Barna, L.C.; Defee, I.; Varri, A. A wireless ballistocardiographic chair. In Proceedings of the 2006 International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 5932–5935. [Google Scholar]
- Aubert, X.L.; Brauers, A. Estimation of vital signs in bed from a single unobtrusive mechanical sensor: Algorithms and real-life evaluation. In Proceedings of the 2008 30th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Vancouver, BC, Canada, 20–25 August 2008; pp. 4744–4747. [Google Scholar]
- Brüser, C.; Winter, S.; Leonhardt, S. Robust inter-beat interval estimation in cardiac vibration signals. Physiol. Meas. 2013, 34, 123–138. [Google Scholar] [CrossRef]
- Zink, M.D.; Brüser, C.; Stüben, B.O.; Napp, A.; Stöhr, R.; Leonhardt, S.; Marx, N.; Mischke, K.; Schulz, J.B.; Schiefer, J. Unobtrusive nocturnal heartbeat monitoring by a ballistocardiographic sensor in patients with sleep disordered breathing. Sci. Rep. 2017, 7, 13175. [Google Scholar] [CrossRef]
- Watanabe, K.; Watanabe, T.; Watanabe, H.; Ando, H.; Ishikawa, T.; Kobayashi, K. Noninvasive measurement of heartbeat, respiration, snoring and body movements of a subject in bed via a pneumatic method. IEEE Trans. Biomed. Eng. 2005, 52, 2100–2107. [Google Scholar] [CrossRef]
- Pinheiro, E.; Postolache, O.; Girão, P. Theory and developments in an unobtrusive cardiovascular system representation: Ballistocardiography. Open Biomed. Eng. J. 2010, 4, 201. [Google Scholar] [CrossRef] [PubMed]
- Antonio, E.-Z. A Simple Ballistocardiographic System for a Medical Cardiovascular Physiology Course. Adv. Physiol. Educ. 2003, 27, 224–229. [Google Scholar] [CrossRef]
- Sadek, I.; Biswas, J. Nonintrusive heart rate measurement using ballistocardiogram signals: A comparative study. Signal Image Video Process. 2018, 13, 475–482. [Google Scholar] [CrossRef]
- Suliman, A.; Carlson, C.; Ade, C.J.; Warren, S.; Thompson, D.E. Performance comparison for ballistocardiogram peak detection methods. IEEE Access 2019, 7, 53945–53955. [Google Scholar] [CrossRef]
- Paalasmaa, J.; Ranta, M. Detecting heartbeats in the ballistocardiogram with clustering. In Proceedings of the ICML/UAI/COLT 2008 Workshop on Machine Learning for Health-Care Applications, Helsinki, Finland, 5–9 July 2008; Volume 9. [Google Scholar]
- Pinheiro, E.; Postolache, O.; Girão, P. Study on ballistocardiogram acquisition in a moving wheelchair with embedded sensors. Metrol. Meas. Syst. 2012, 19, 739–750. [Google Scholar] [CrossRef]
- Choe, S.T.; Cho, W.D. Simplified real-time heartbeat detection in ballistocardiography using a dispersion-maximum method. Biomed. Res. 2017, 28, 3974–3985. [Google Scholar]
- Inan, O.T.; Etemadi, M.; Wiard, R.M.; Giovangrandi, L.; Kovacs, G.T.A. Robust ballistocardiogram acquisition for home monitoring. Physiol. Meas. 2009, 30, 169–185. [Google Scholar] [CrossRef]
- Wiard, R.M.; Inan, O.T.; Argyres, B.; Etemadi, M.; Kovacs, G.T.A.; Giovangrandi, L. Automatic detection of motion artifacts in the ballistocardiogram measured on a modified bathroom scale. Med. Biol. Eng. Comput. 2010, 49, 213–220. [Google Scholar] [CrossRef]
- Bruser, C.; Stadlthanner, K.; de Waele, S.; Leonhardt, S. Adaptive beat-to-beat heart rate estimation in ballistocardiograms. IEEE Trans. Inf. Technol. Biomed. 2011, 15, 778–786. [Google Scholar] [CrossRef] [PubMed]
- Dziuda, Ł.; Skibniewski, F.W. A new approach to ballistocardiographic measurements using fibre Bragg grating-based sensors. Biocybern. Biomed. Eng. 2014, 34, 101–116. [Google Scholar] [CrossRef]
- Nedoma, J.; Fajkus, M.; Martinek, R.; Kepak, S.; Cubik, J.; Zabka, S.; Vasinek, V. Comparison of bcg, pcg and ecg signals in application of heart rate monitoring of the human body. In Proceedings of the 40th International Conference on Telecommunications and Signal Processing (TSP), Barcelona, Spain, 5–7 July 2017; IEEE: New York, NY, USA, 2017; pp. 420–424. [Google Scholar]
- Song, Y.; Ni, H.; Zhou, X.; Zhao, W.; Wang, T. Extracting features for cardiovascular disease classification based on ballistocardiography. In Proceedings of the 2015 IEEE 12th Intl Conf on Ubiquitous Intelligence and Computing and 2015 IEEE 12th Intl Conf on Autonomic and Trusted Computing and 2015 IEEE 15th Intl Conf on Scalable Computing and Communications and its Associated Workshops (UIC-ATC-ScalCom), Beijing, China, 10–14 August 2015; IEEE: New York, NY, USA, 2015; pp. 1230–1235. [Google Scholar]
- Liu, F.; Zhou, X.; Wang, Z.; Wang, T.; Zhang, Y. Identification of hypertension by mining class association rules from multi-dimensional features. In Proceedings of the 2018 24th International Conference on Pattern Recognition (ICPR), Beijing, China, 20–24 August 2018; IEEE: New York, NY, USA, 2018; pp. 3114–3119. [Google Scholar]
- Liu, F.; Zhou, X.; Wang, Z.; Cao, J.; Wang, H.; Zhang, Y. Unobtrusive mattress-based identification of hypertension by integrating classification and association rule mining. Sensors 2019, 19, 1489. [Google Scholar] [CrossRef]
- Jain, P.; Gajbhiye, P.; Tripathy, R.; Acharya, U.R. A two-stage deep CNN architecture for the classification of low-risk and high-risk hypertension classes using multi-lead ECG signals. Informatics Med. Unlocked 2020, 21, 100479. [Google Scholar] [CrossRef]
- Rajput, J.S.; Sharma, M.; Kumbhani, D.; Acharya, U.R. Automated detection of hypertension using wavelet transform and nonlinear techniques with ballistocardiogram signals. Informatics Med. Unlocked 2021, 26, 100736. [Google Scholar] [CrossRef]
- Rajput, J.S.; Sharma, M.; Kumar, T.S.; Acharya, U.R. Automated detection of hypertension using continuous wavelet transform and a deep neural network with Ballistocardiography signals. Int. J. Environ. Res. Public Health 2022, 19, 4014. [Google Scholar] [CrossRef] [PubMed]
- Gupta, K.; Bajaj, V.; Ansari, I.A.; Acharya, U.R. Hyp-Net: Automated detection of hypertension using deep convolutional neural network and Gabor transform techniques with ballistocardiogram signals. Biocybern. Biomed. Eng. 2022, 42, 784–796. [Google Scholar] [CrossRef]
- Ge, B.; Yang, H.; Ma, P.; Guo, T.; Pan, J.; Wang, W. Detection of pulmonary hypertension associated with congenital heart disease based on time-frequency domain and deep learning features. Biomed. Signal Process. Control 2023, 81, 104316. [Google Scholar] [CrossRef]
- Ma, P.; Ge, B.; Yang, H.; Guo, T.; Pan, J.; Wang, W. Application of time-frequency domain and deep learning fusion feature in non-invasive diagnosis of congenital heart disease-related pulmonary arterial hypertension. MethodsX 2023, 10, 102032. [Google Scholar] [CrossRef]
- Sun, H.; Jiao, J.; Ren, Y.; Guo, Y.; Wang, Y. Multimodal fusion model for classifying placenta ultrasound imaging in pregnancies with hypertension disorders. Pregnancy Hypertens. 2023, 31, 46–53. [Google Scholar] [CrossRef]
- Dogan, S.; Barua, P.D.; Tuncer, T.; Acharya, U.R. An accurate hypertension detection model based on a new odd-even pattern using ballistocardiograph signals. Eng. Appl. Artif. Intell. 2024, 133, 108306. [Google Scholar] [CrossRef]
- Ozcelik, S.T.A.; Uyanık, H.; Deniz, E.; Sengur, A. Automated hypertension detection using ConvMixer and spectrogram techniques with ballistocardiograph signals. Diagnostics 2023, 13, 182. [Google Scholar] [CrossRef]
- Gupta, K.; Bajaj, V.; Ansari, I.A. A support system for automatic classification of hypertension using BCG signals. Expert Syst. Appl. 2022, 214, 119058. [Google Scholar] [CrossRef]
- Wahyuningrum, T.; Khomsah, S.; Suyanto, S.; Meliana, S.; Yunanto, P.E.; Al Maki, W.F. Improving Clustering Method Performance Using K-Means, Mini Batch K-Means, BIRCH and Spectral. In Proceedings of the 2021 4th International Seminar on Research of Information Technology and Intelligent Systems (ISRITI), Yogyakarta, Indonesia, 16–17 December 2021; pp. 206–210. [Google Scholar]
- Ali, H.H.S.M.; Sharif, S.M. Comparison Between Discrete Wavelet Transform and Maximal Overlap Discrete Wavelet Transform as an Analysis tool for H.264/AVC Video. In Proceedings of the 2018 International Conference on Computer, Control, Electrical, and Electronics Engineering (ICCCEEE), Khartoum, Sudan, 12–14 August 2018; pp. 1–5. [Google Scholar]
- Hidayat, R.; Kristomo, D.; Togarma, I. Feature extraction of the Indonesian phonemes using discrete wavelet and wavelet packet transform. In Proceedings of the 2016 8th International Conference on Information Technology and Electrical Engineering (ICITEE), Yogyakarta, Indonesia, 5–6 October 2016; pp. 1–6. [Google Scholar]
- Liu, W.; Cao, S.; Chen, Y. Seismic Time–Frequency Analysis via Empirical Wavelet Transform. IEEE Geosci. Remote Sens. Lett. 2015, 13, 28–32. [Google Scholar] [CrossRef]
- Prasad, K.D.; Khan, M.R.; Das, B. MUD in an Underwater Network Using Bioinspired Binary Tunicate Swarm Optimizer. In Proceedings of the 2025 3rd IEEE International Conference on Industrial Electronics: Developments & Applications (ICIDeA), Bhubaneswar, India, 21–22 February 2025; pp. 1–5. [Google Scholar]
- Thomaz, C.E.; Gillies, D.F. A Maximum Uncertainty LDA-Based Approach for Limited Sample Size Problems—With Application to Face Recognition. In Proceedings of the XVIII Brazilian Symposium on Computer Graphics and Image Processing (SIBGRAPI’05), Natal, Brazil, 9–12 October 2005; pp. 89–96. [Google Scholar]
- Wu, S.; Nagahashi, H. Parameterized AdaBoost: Introducing a Parameter to Speed Up the Training of Real AdaBoost. IEEE Signal Process. Lett. 2014, 21, 687–691. [Google Scholar] [CrossRef]
- Martins, W.; Bagesteiro, L.B.; Weber, T.O.; Balbinot, A. FPGA-based Implementation of Random Forest Classifier for sEMG Signal Classification. In Proceedings of the 2024 46th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 15–19 July 2024; pp. 1–4. [Google Scholar]
- Ge, W.; Hongzhe, X.; Weibin, Z.; Weilu, Z.; Baiyang, F. Multi-kernel PCA based high-dimensional images feature reduction. In Proceedings of the 2011 International Conference on Electric Information and Control Engineering, Wuhan, China, 15–17 April 2011; pp. 5966–5969. [Google Scholar]
- Chung, A.G.; Shafiee, M.J.; Wong, A. Random feature maps via a Layered Random Projection (LARP) framework for object classification. In Proceedings of the 2016 IEEE International Conference on Image Processing (ICIP), Phoenix, AZ, USA, 25–28 September 2016; pp. 246–250. [Google Scholar]
- Yang, Y.Z.; Liu, P.Y.; Zhu, Z.F.; Qiu, Y. The research of an improved information gain method using distribution information of terms. In Proceedings of the 2009 IEEE International Symposium on IT in Medicine & Education, Jinan, China, 14–16 August 2009; pp. 938–941. [Google Scholar]
- Li, X.; Ma, Y.; Li, Y.; Li, H.; Zuo, H.; Sun, W. Improved Aquila Optimizer Optimization Algorithm Based on Multi-strategy Fusion. In Proceedings of the 2022 2nd International Conference on Computer Science, Electronic Information Engineering and Intelligent Control Technology (CEI), Nanjing, China, 23–25 September 2022; pp. 757–760. [Google Scholar]
- Fei, W.; Bai, L. Auto-regressive models of non-stationary time series with finite length. Tsinghua Sci. Technol. 2005, 10, 162–168. [Google Scholar] [CrossRef]
- Shang, Y. Finite-Time Weighted Average Consensus and Generalized Consensus Over a Subset. IEEE Access 2016, 4, 2615–2620. [Google Scholar] [CrossRef]
- Bernardini, M.; Romeo, L.; Misericordia, P.; Frontoni, E. Discovering the Type 2 Diabetes in Electronic Health Records Using the Sparse Balanced Support Vector Machine. IEEE J. Biomed. Health Inform. 2019, 24, 235–246. [Google Scholar] [CrossRef]
- Li, W.; Wang, R.; Huang, D. Assessment of Micro-movement Sensitive Mattress Sleep Monitoring System (RS611) in the detection of obstructive sleep apnea hypopnea syndrome. Chin. J. Gerontol. 2015, 35, 1160–1162. [Google Scholar]
- Park, S.H.; Reyes, J.A.; Gilbert, D.R. Prediction of protein-protein interaction types using association rule-based classification. BMC Bioinform. 2009, 10, 36. [Google Scholar] [CrossRef]




| Number | 61 |
| Gender (male) | 33 |
| Gender (female) | 38 |
| Age group | 55.6 ± 7.9 |
| Body Mass Index (BMI) | 24.3 ± 3.6 |
| Heart rate (bpm) | 77.1 ± 9.2 |
| Systolic blood pressure (mmHg) | 155.6 ± 11.2 |
| Diastolic blood pressure (mmHg) | 103.6 ± 8.2 |
| Number | 67 |
| Gender (male) | 35 |
| Gender (female) | 32 |
| Age group | 53.2 ± 9.2 |
| Body Mass Index (BMI) | 23.4 ± 3.3 |
| Heart rate (bpm) | 73.6 ± 8.3 |
| Systolic blood pressure (mmHg) | 112.1 ± 15.7 |
| Diastolic blood pressure (mmHg) | 74.4 ± 6.3 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 89.45 | 89.58 | 89.05 | 91.45 | 95.51 | 95.45 |
| MODWT | 91.67 | 91.14 | 90.64 | 92.24 | 96.48 | 95.48 |
| EWT | 90.98 | 90.57 | 89.87 | 90.69 | 94.37 | 93.54 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 88.34 | 87.21 | 88.09 | 90.22 | 93.09 | 91.98 |
| MODWT | 90.56 | 89.36 | 88.97 | 91.24 | 93.87 | 92.23 |
| EWT | 88.63 | 88.78 | 87.45 | 90.57 | 92.42 | 91.58 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 85.09 | 82.21 | 83.87 | 84.33 | 86.21 | 85.02 |
| MODWT | 84.97 | 83.24 | 84.74 | 81.37 | 87.36 | 86.34 |
| EWT | 83.58 | 84.67 | 85.35 | 83.89 | 86.98 | 85.67 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 84.23 | 80.09 | 84.21 | 86.09 | 88.11 | 88.05 |
| MODWT | 83.45 | 81.87 | 82.25 | 83.98 | 89.26 | 89.04 |
| EWT | 83.68 | 82.34 | 84.78 | 85.42 | 89.90 | 87.37 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 86.32 | 84.89 | 82.21 | 84.09 | 90.21 | 89.33 |
| MODWT | 87.66 | 85.97 | 83.25 | 86.98 | 91.23 | 90.46 |
| EWT | 88.78 | 85.55 | 82.78 | 85.77 | 90.58 | 88.78 |
| RF | DT | NBC | AdaBoost | Hybrid AdaBoost–MULDA | Hybrid AdaBoost–RF | |
|---|---|---|---|---|---|---|
| K-means | 85.21 | 84.03 | 85.21 | 87.03 | 89.21 | 87.01 |
| MODWT | 85.33 | 86.45 | 83.23 | 84.33 | 89.67 | 88.21 |
| EWT | 85.48 | 86.68 | 82.69 | 85.59 | 87.99 | 87.08 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
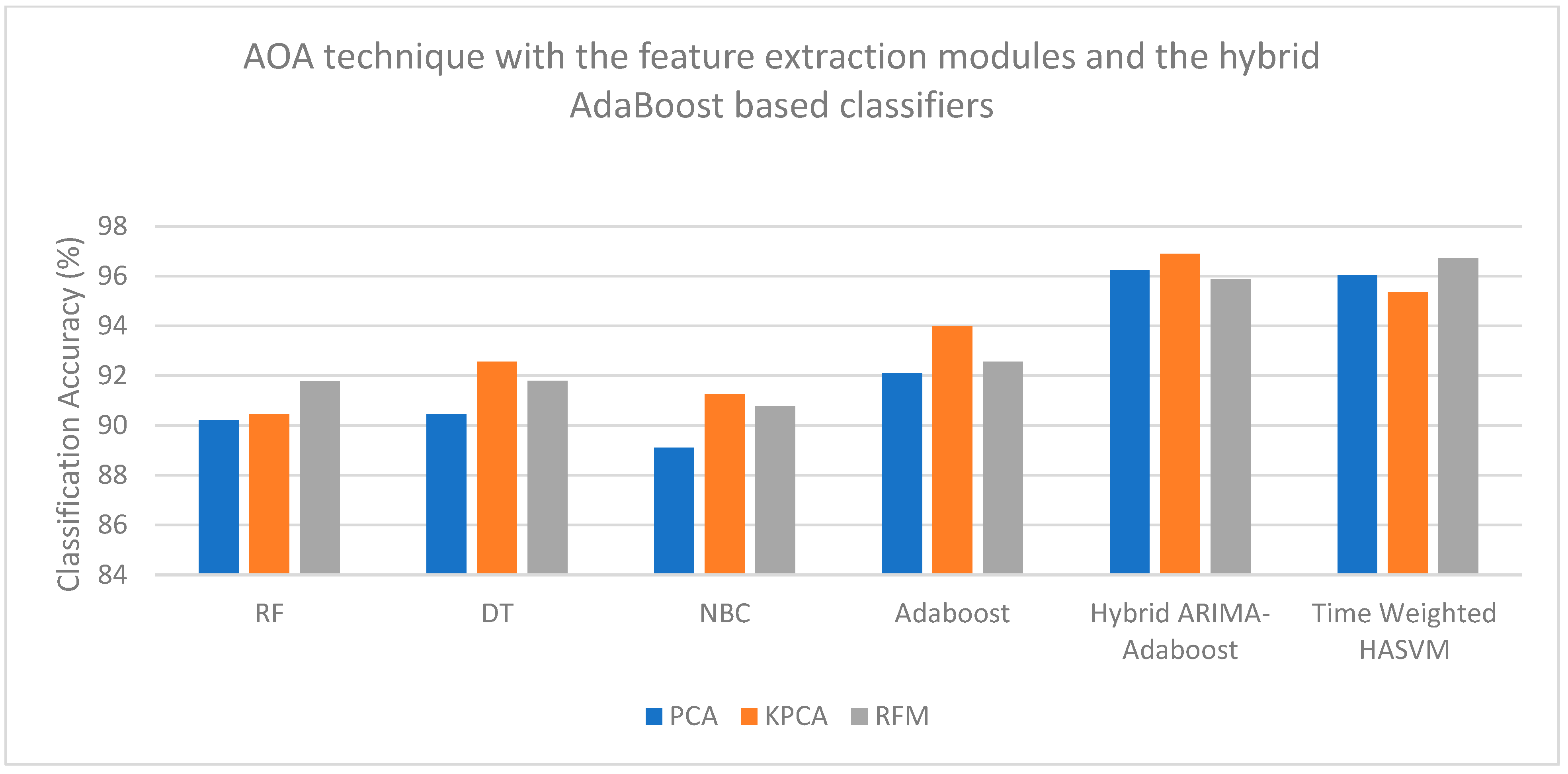
| PCA | 90.21 | 90.44 | 89.11 | 92.09 | 96.23 | 96.02 |
| KPCA | 90.45 | 92.56 | 91.24 | 93.98 | 96.89 | 95.34 |
| RFM | 91.78 | 91.79 | 90.78 | 92.56 | 95.89 | 96.71 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
| PCA | 87.23 | 86.21 | 89.98 | 90.02 | 92.11 | 92.12 |
| KPCA | 89.44 | 88.22 | 88.87 | 91.34 | 91.26 | 92.37 |
| RFM | 87.57 | 87.57 | 89.58 | 91.68 | 91.87 | 91.98 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
| PCA | 84.21 | 80.78 | 84.09 | 85.21 | 87.22 | 84.22 |
| KPCA | 83.22 | 81.98 | 85.87 | 82.35 | 88.37 | 85.37 |
| RFM | 81.46 | 82.36 | 83.36 | 84.78 | 86.65 | 89.30 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
| PCA | 82.89 | 82.12 | 82.09 | 84.21 | 86.87 | 87.09 |
| KPCA | 80.44 | 80.34 | 81.84 | 85.25 | 85.77 | 88.88 |
| RFM | 80.57 | 81.78 | 83.22 | 84.67 | 87.63 | 87.34 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
| PCA | 87.21 | 86.87 | 85.09 | 88.23 | 90.77 | 89.21 |
| KPCA | 88.22 | 87.75 | 86.87 | 85.45 | 90.63 | 91.36 |
| RFM | 89.36 | 87.49 | 84.34 | 87.61 | 91.48 | 89.89 |
| RF | DT | NBC | AdaBoost | Hybrid ARIMA–AdaBoost | TW-HASVM | |
|---|---|---|---|---|---|---|
| PCA | 83.21 | 85.09 | 82.23 | 85.09 | 88.24 | 89.09 |
| KPCA | 82.34 | 84.36 | 84.67 | 86.97 | 88.67 | 87.88 |
| RFM | 84.98 | 83.78 | 85.33 | 84.45 | 89.84 | 89.21 |
| Year | Authors | Methods | Performance Metrics |
|---|---|---|---|
| 2018 | Liu et al. [28] | Mining class association from multi-dimensional features | Acc = 85.20% |
| 2019 | Liu et al. [29] | Integrated classification and association rule mining | Acc = 84.40% |
| 2021 | Rajput et al. [31] | EMD, wavelet decomposition and nonlinear techniques | Acc = 89.00% |
| 2022 | Rajput et al. [32] | Continuous wavelet transform combined with deep neural networks | Acc = 86.14% |
| 2022 | Gupta et al. [33] | Gabor transform and deep convolutional neural network | Acc = 97.65% |
| 2023 | Gupta et al. [39] | Integrated tunable Q-factor wavelet transform and multiverse optimization concept | Acc = 92.21% |
| 2023 | Ozcelik et al. [38] | Spectrogram tasks + ConvMixer | Acc = 97.69% |
| 2024 | Dogan et al. [37] | Odd–even pattern with KNN algorithm and majority voting | Acc = 97.78% |
| 2025 | Proposed works |
| Acc = 95.51% Acc = 96.48% Acc = 94.37% |
| Acc = 93.09% Acc = 93.87% Acc = 92.42% | ||
| Acc = 96.23% Acc = 96.89% Acc = 95.89% | ||
| Acc = 92.12% Acc = 92.37% Acc = 91.98% |
| Year | Authors | Methods | Computational Time |
|---|---|---|---|
| 2025 | Proposed works |
| 10.563 s 11.342 s 11.291 s |
| 10.247 s 9.298 s 9.037 s | ||
| 6.567 s 5.112 s 6.012 s | ||
| 8.378 s 9.001 s 8.981 s |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Prabhakar, S.K.; Won, D.-O. Pragmatic Models for Detection of Hypertension Using Ballistocardiograph Signals and Machine Learning. Bioengineering 2026, 13, 43. https://doi.org/10.3390/bioengineering13010043
Prabhakar SK, Won D-O. Pragmatic Models for Detection of Hypertension Using Ballistocardiograph Signals and Machine Learning. Bioengineering. 2026; 13(1):43. https://doi.org/10.3390/bioengineering13010043
Chicago/Turabian StylePrabhakar, Sunil Kumar, and Dong-Ok Won. 2026. "Pragmatic Models for Detection of Hypertension Using Ballistocardiograph Signals and Machine Learning" Bioengineering 13, no. 1: 43. https://doi.org/10.3390/bioengineering13010043
APA StylePrabhakar, S. K., & Won, D.-O. (2026). Pragmatic Models for Detection of Hypertension Using Ballistocardiograph Signals and Machine Learning. Bioengineering, 13(1), 43. https://doi.org/10.3390/bioengineering13010043






