Abstract
Dyspepsia or indigestion is known to be a symptom of several gastrointestinal issues including gastroesophageal reflux disease (GERD) and gastric cancer. With the ever-increasing popularity of traditional herbal medicines, the research selected rosemary, or Salvia rosmarinus, due to its historical ethnopharmacological applications and ease of cultivation. Potential targets of molecules from S. rosmarinus are explored for the molecular pathogenesis of functional dyspepsia. Through a network pharmacology approach, it has been shown that there is a significant interaction between the disease and the plant’s compounds. The pathways involving the target genes of S. rosmarinus that are related to functional dyspepsia were revealed to be implicated in the development of certain diseases like gastric cancer and fibrosis which are both precursors to dyspepsia. Moreover, through molecular docking, the results of the pathway analyses were computationally validated indicating that the compound luteolin has the most significant interaction against dyspepsia-related genes. It effectively promotes apoptosis in cancer pathways, reducing the chances of gastric cancer carcinogenesis. To further validate these findings, molecular dynamics simulation was performed to compare the stability and binding behavior of the top-performing compound against the reference compound. The results of this study could be a possible basis in developing pharmaceuticals against gastrointestinal diseases, specifically, dyspepsia.
1. Introduction
Dyspepsia, more commonly known as indigestion, is a condition that inflicts discomfort or even pain in the upper abdomen [1]. Dyspepsia is usually associated with gastrointestinal afflictions and affects 20% of the whole populace who got diagnosed. Some examples of gastrointestinal disorders are gastroesophageal reflux disease (GERD), ulcers, and gastric cancer involving Helicobacter pylori [2]. In the Philippines, the risk factor for dyspepsia was found to be around 34% for people in general, while 26.5% to 79.9% for patients who have already experienced dyspepsia. Moreover, the reported risk factor was also found to be associated with gastric cancer [3]. Dyspepsia is used as a medical term to refer to the occurrence of stomach lining irritation caused by gastric acid [4]. Dyspepsia on its own does not necessarily require immediate attention, but due to how it affects the quality of life of several people, it is important to address it promptly to alleviate discomfort and prevent potential complications.
There are organic and non-organic causes underlying dyspepsia. Organic causes tend to be more easily identified due to their nature, which results from medical conditions and what a person consumes. If there is an imbalance in the production of gastric acid or enzymes, or if the stomach lining is irritated, it can lead to symptoms such as pain, bloating, or discomfort [5]. Gastric acid is highly impacted by the proton pumps on the surface of the stomach wall. They primarily pump hydrogen ions from the parietal cells to the stomach lumen. This affects the production of HCl, a main component of gastric acid. Proton pump inhibitors are used to control the amount of gastric acid in treating related medical conditions [6]. Non-organic classifications tend to have no specific causes and may be linked to the brain [7]. The gut–brain axis is a communication system connecting the brain to the gastrointestinal tract and vice versa and involves the microbiota in the gut [8]. Several studies indicate that depression and anxiety also contribute to dyspepsia. Frequent visits should prompt medical practitioners to also test for possible psychiatric comorbidities [9]. Dyspepsia is easily prevented through a change of diet and avoiding certain food types such as fats, alcohol, and caffeine. Antacids are also commonly used to neutralize stomach acid for relief. In terms of inhibitors, the widely used proton pump inhibitors are omeprazole and lansoprazole [10].
Omeprazole is a known proton pump inhibitor and has been used for gastrointestinal issues for decades [11]. It inhibits the proton pumps namely the hydrogen (H+)/potassium (K+) ATPase enzymes which are located at the parietal cells of the stomach lining. The role of these enzymes is to exchange g K+ ions in the stomach lumen with H+ ions from the parietal cell cytoplasm. Specifically targeting these proton pumps effectively reduces the amount of gastric acid made, lessening the irritation in the stomach [12]. It has been used for GERD and ulcers especially in the presence of dyspepsia as a symptom [13,14]. Lansoprazole has similar functions as a proton pump inhibitor assisting in the relief of gastrointestinal afflictions [15]. However, lansoprazole may have differing effects depending on the etiology of dyspepsia [16].
Currently, the use of herbal medicines is becoming more popular due to several factors including dissatisfaction with conventional medicines, the perceived safety of natural herbs, and its rising number of scientific validations, [17,18,19]. A study from Alsukaibi et al. presented the assessment of natural products from dates and their bioactivity through in silico analysis [20]. Relatedly, some plants have been long used as traditional medicines due to their ability to synthesize a large variety of organic molecules which could have a significant pharmacological effect, thus, making them important in the field of drug discovery [21]. Several regions in the world have a history of using additional medicines including China, India, Japan, Egypt, and the Mediterranean region [22,23]. One of the herbs that has been traced to have originated in the Mediterranean was S. rosmarinus or rosemary [24]. This plant was known to grow in mild, wet winters and dry summers [25]. Despite the conditions necessary for its growth, rosemary was still able to be cultivated in several parts of the world if it is in the right conditions [26]. The medicinal properties of S. rosmarinus, however, show potential as an anti-inflammatory agent or as an antioxidant. Additionally, it can even be used in aromatherapy [27]. S. rosmarinus was also reported to have antibacterial, anti-inflammatory, and spasmolytic properties [28]. Furthermore, a study conducted by Sereiti et al., discussed the uses of rosemary extracts for choleretic, hepatoprotective, and antitumorigenic activities [29]. Alongside, the in silico method for analyzing the potentials of phytochemicals have been recently becoming prominent. Several studies have utilized this method to identify the possible mechanisms and drug capabilities of compounds [30]. Additionally, a study from Yadav et al. utilized the same specific method of molecular docking to identify the drug-likeliness of CK2 inhibitors as a potential drug for COVID-19 [31].
To further explore the potential uses of S. rosmarinus, an in silico analysis using bioinformatics will be conducted. Moreover, this study aims to explore the bioactive compounds of S. rosmarinus as well as their potential targets, and to assess their viability in the treatment of dyspepsia through network pharmacology analysis. This study also aims to evaluate the binding affinities of the bioactive compounds from S. rosmarinus against the targets to be traced related to dyspepsia in order to understand the potential efficacy of the compounds as therapeutic agents. Nevertheless, this study aims to highlight the key pathways involved in the development of functional dyspepsia and its precursors.
2. Materials and Methods
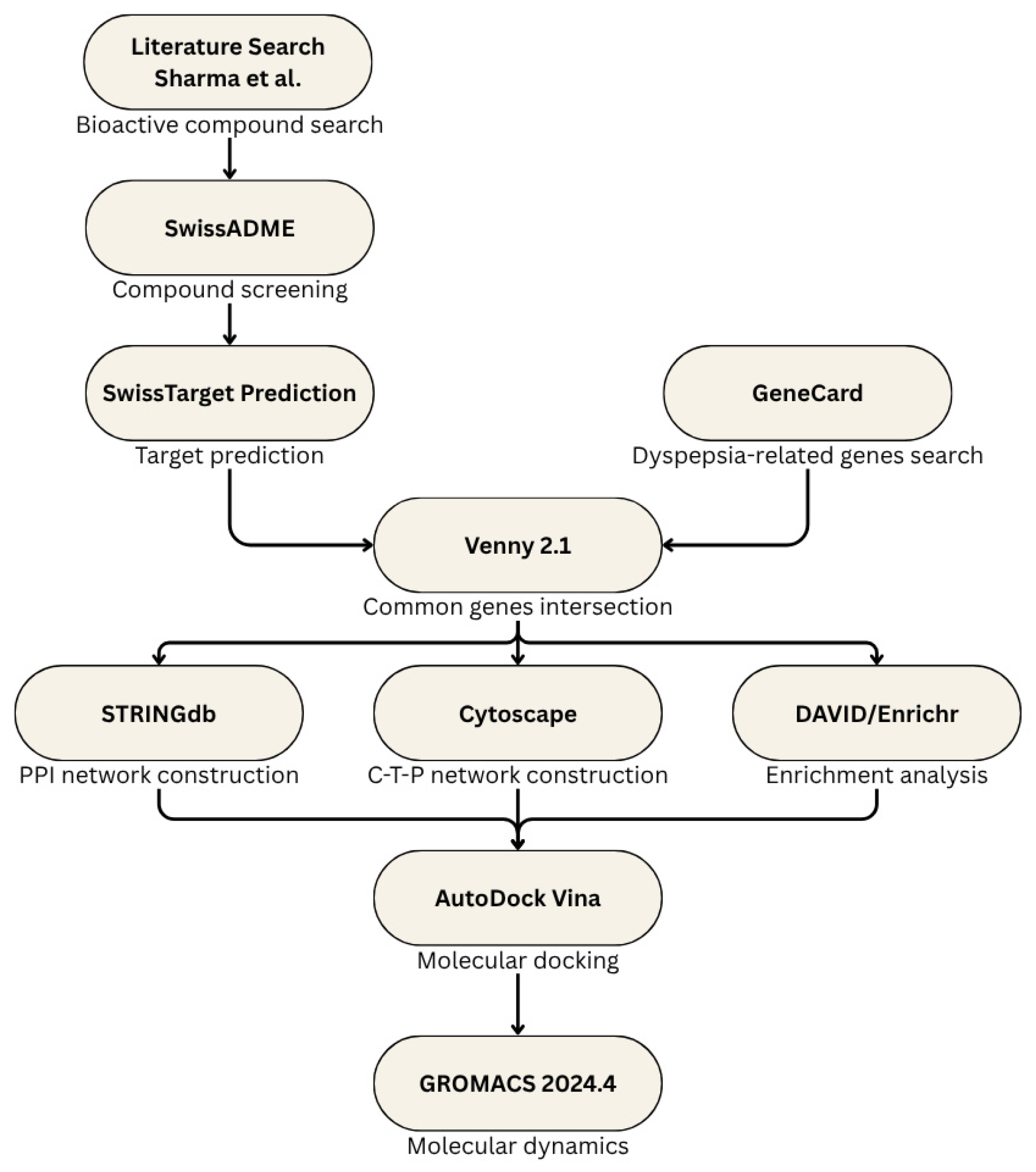
This study was conducted in silico and was divided into several parts: starting with data mining and literature search for the bioactive compounds of S. rosmarinus up to validation using molecular docking. The step-by-step process of this study can be seen in Figure 1.
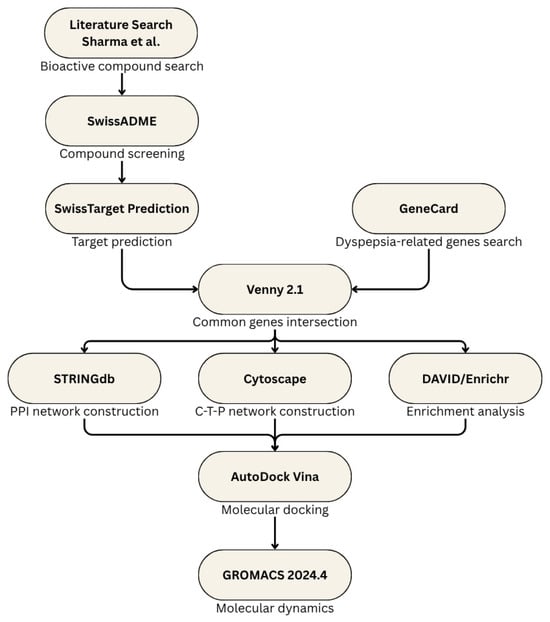
Figure 1.
Outline of the methodology.
2.1. Screening of Bioactive Compounds in S. rosmarinus
The bioactive compounds of S. rosmarinus were enlisted from the findings of Sharma et al. [32]. Here, the samples were analyzed by ultra-high-performance liquid chromatography, electrospray ionization coupled with quadrupole-time of flight mass spectrometry (UHPLC-ESI-QTOF-MS). The compounds were filtered according to their physicochemical properties which were mainly and drug-likeliness using SwissADME (http://www.swissadme.ch/index.php, accessed on 22 July 2024) [33]. The bioavailability threshold was set to ≥ 0.55, while the oral bioavailability was set to OB ≥ 30%, similar to the standards set by existing studies [30]. The established cutoff for both criteria may indicate a higher probability that a greater amount of compounds from S. rosmarinus reaches the systemic circulation. This may also indicate that the desired therapeutic effects are achieved. Consequently, the thresholds reflect a balance between the absorption and distribution of the compounds [34].
2.2. Target Prediction of Bioacidtive Compounds from S. rosmarinus
The bioactive compounds that have met the criteria for absorption, distribution, metabolism, and excretion (ADME) were subjected to target prediction using SwissTargetPrediction (http://www.swisstargetprediction.ch/, accessed on 22 July 2024) [35]. This web-based server is capable of predicting the target of small molecules. The canonical smiles of each compound were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/, accessed on 22 July 2024) [36]. Moreover, in predicting the targets of the compounds, the prediction was set to only “Homo sapiens” in order to narrow down the variation. The collected target genes were then filtered with a probability threshold ≥ 0.70.
2.3. Crossover of S. rosmarinus Targets and Dyspepsia-Related Genes
Using the GeneCard database (https://www.genecards.org/, accessed on 22 July 2024) [37], the genes related to dyspepsia were obtained. Here, the genes obtained were based of the relevance scores in the database. Identified disease-related genes were then filtered to top 25% by quantile using Python 3.12.4, and the Pandas 2.2.2 library. The top 25% results were selected to narrow down the set of genes involved and to ensure their relevance while having sufficient data. Consequently, the common genes from dyspepsia and S. rosmarinus were identified using the bioinformatics tool Venny 2.1 (https://www.biotools.fr/misc/venny, accessed on 22 July 2024) [38].
2.4. Construction of PPI Network
The protein–protein interaction network of the target genes from S. rosmarinus and dyspepsia was constructed using the STRINGdb (https://string-db.org/, accessed on 22 July 2024) [39]—a database for known protein to protein interaction. Here, the interaction was limited to “Homo sapiens”. The PPI networks were then sent to Cytoscape version 3.10.2 for further validation. The common genes were also filtered based on their maximal clique centrality (MCC), maximum neighborhood component (MNC), and degree using the Cytohubba plugin. This plugin is responsible for the identification of the important nodes in the network itself [40]. The same plugin was used to identify the core compound of S. rosmarinus based on its degree which is a metric to identify which node is the most biologically important. The analysis was set to the top five genes.
Moreover, the enrichment analysis of the network was done using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) (https://david.ncifcrf.gov/, accessed on 22 July 2024) [41]. This tool is capable of identifying several biological meanings given in a set of genes. Nevertheless, the analyses were limited to the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Using the same tool, the biological processes, cellular components, and molecular functions of the common genes were identified. Using Enrichr (https://maayanlab.cloud/Enrichr/enrich, accessed on 30 June 2025), the protein where the top-performing ligand was bound, was cross-checked to further analyze its GO biological processes [42].
2.5. Compound, Target, and Pathway (CTP) Network Construction
Cytoscape (version 3.10.2) [43] was then utilized to construct the CTP network. The compound to target interaction was first constructed by loading the targets that were screened by probability ≥ 0.70 into the software and was merged into the pathways network where the top scoring connected the genes were accounted. On the other hand, the top compounds from S. rosmarinus were identified using the degree algorithm from CytoHubba in the same software.
2.6. Molecular Docking of the Key Compounds from S. rosmarinus and Dyspepsia
To validate the results of the network pharmacology analyses of dyspepsia and S. rosmarinus, molecular docking was done for the bioactive compounds of the plant against the dyspepsia-related genes. The control compounds used were omeprazole and lansoprazole which are both proton pump inhibitors and has an efficient binding activity against dyspepsia-related genes [44]. They are used as a basis due to their established therapeutic effects on dyspepsia-related afflictions and not directly due to their specific mechanisms. The SDF structures of the bioactive compounds (ligands) were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/, accessed on 22 July 2024) [40] which were translated into PDB format using the chemistry toolbox Open Babel (https://openbabel.org/index.html, accessed on 22 July 2024) [45]. On the other hand, the structures of the dyspepsia proteins (target) were obtained from the Protein Data Bank (https://www.rcsb.org/, accessed on 22 July 2024) [46]. Consequently, the structures were prepared using the AutoDock tool where ions, waters, and hetero atoms were removed. Polar hydrogens were also added into the protein. Using the same software, the binding sites of the targets were also identified, specifically, by using the grid tool. Lastly, the molecular docking was done using the AutoDock Vina software (https://vina.scripps.edu/, accessed on 22 July 2024) [47]. The visualization of the protein-ligand complexes was done using the Biovia Discovery Studio (https://www.3ds.com/products/biovia, accessed on 22 July 2024) [48]. Prior to the visualization, the complexes were analyzed using UCSF ChimeraX (https://www.cgl.ucsf.edu/chimerax/, accessed on 22 July 2024) [49], where the proteins and ligands were loaded individually. Visualizing the complexes was able to give the results regarding their molecular interactions, which is also important to elucidate their possible mechanism of actions.
It is also worth noting that the protein target models were chosen according to the best resolution possible. Table 1 indicates the proteins’ respective IDs, as well as their resolution from the Protein Data Bank. Subsequently, the native ligands of the models were redocked to validate their RMSD. The same docking software was utilized as proteins and bioactive compounds. The RMSD of the native co-crystallized ligand and the redocked ligand were computed by aligning both the ligands in the PyMol version 2.5.8 software [50]. The RMSD of the redocked ligands are found in Table 2. Here, only four out of the six protein models were redocked with the native ligands as some only contain ions.

Table 1.
Compound targets and their respective IDs.

Table 2.
RMSD of the redocked ligands.
2.7. Molecular Dynamics Simulation
The molecular dynamics simulation was initiated through the preparation of the system using BioBB Python module v5.0.0 2024.2 [51]. It should be noted that the simulation workflow closely follows the protocol from a previously published study by Chen et al. [52]. A comparative dynamics simulation was done for the top-performing complex against the top performing control of the same protein which were KIT-luteolin and KIT-lansoprazole. The .pdb files of both complexes from the docking simulation were subjected to analysis, whereas the ligands and the proteins were isolated. Consequently, GAFF forcefield was applied to the ligands, while amber99sb-ildn forcefield was applied to the proteins. After the production of protein and ligan topologies, the complexes were remade and solvated using space as the water model. An octahedron 0.8 nm away from the system was then used as the solvent box, with Na and Cl ions acting as neutralizers for any protein charges present. Furthermore, the energy minimization was set to 5000 steps. The last step of the preparation involved the canonical ensemble (NVT) and isothermal-isobaric ensemble (NPT), having 5000 steps each at 10 ps.
Moreover, the simulation was conducted using Gromacs 2024.4 at 100 ns (50,000,000 steps). This software allows for the high-performance simulation of biomolecular systems such as proteins, lipids, and nucleic acids under physical forces [53]. The statistical analysis of the structural stability was done using the Kolmogorov-Smirnov test [54]. Here, the parameters that were considered include RMSD, radius of gyration, RMSF, and total number of hydrogen bonds. Lastly, the molecular dynamics simulation was validated through Cliff’s Delta analysis [55].
3. Results
3.1. Bioactive Compounds from S. rosmarinus
The screening of the 50 bioactive compounds from S. rosmarinus revealed only 10 compounds with the bioavailability score ≥ 0.55 and oral bioavailability ≥ 30%. Moreover, 8 out of the 10 compounds have also satisfied the Lipinski’s rule of five, indicating that these 8 compounds could possibly have good absorption occurrence due to the ratio of hydrogen bond donors to acceptors, molecular weight greater than 500, and a calculated partition coefficient greater than 5 [56]. The compounds were also screened according to other drug-likeliness scales including Ghose, Veber, Egan, and Muegge and was narrowed down to six compounds only which were luteolin, isorhamnetin, hesperitin, diosmetin, pectolinarigenin, and genkwanin. The following information on the compounds are found in Table 3.

Table 3.
Bioactive compounds of S. rosmarinus.
3.2. Predicted Dyspepsia-Related Genes and Target Genes of S. rosmarinus
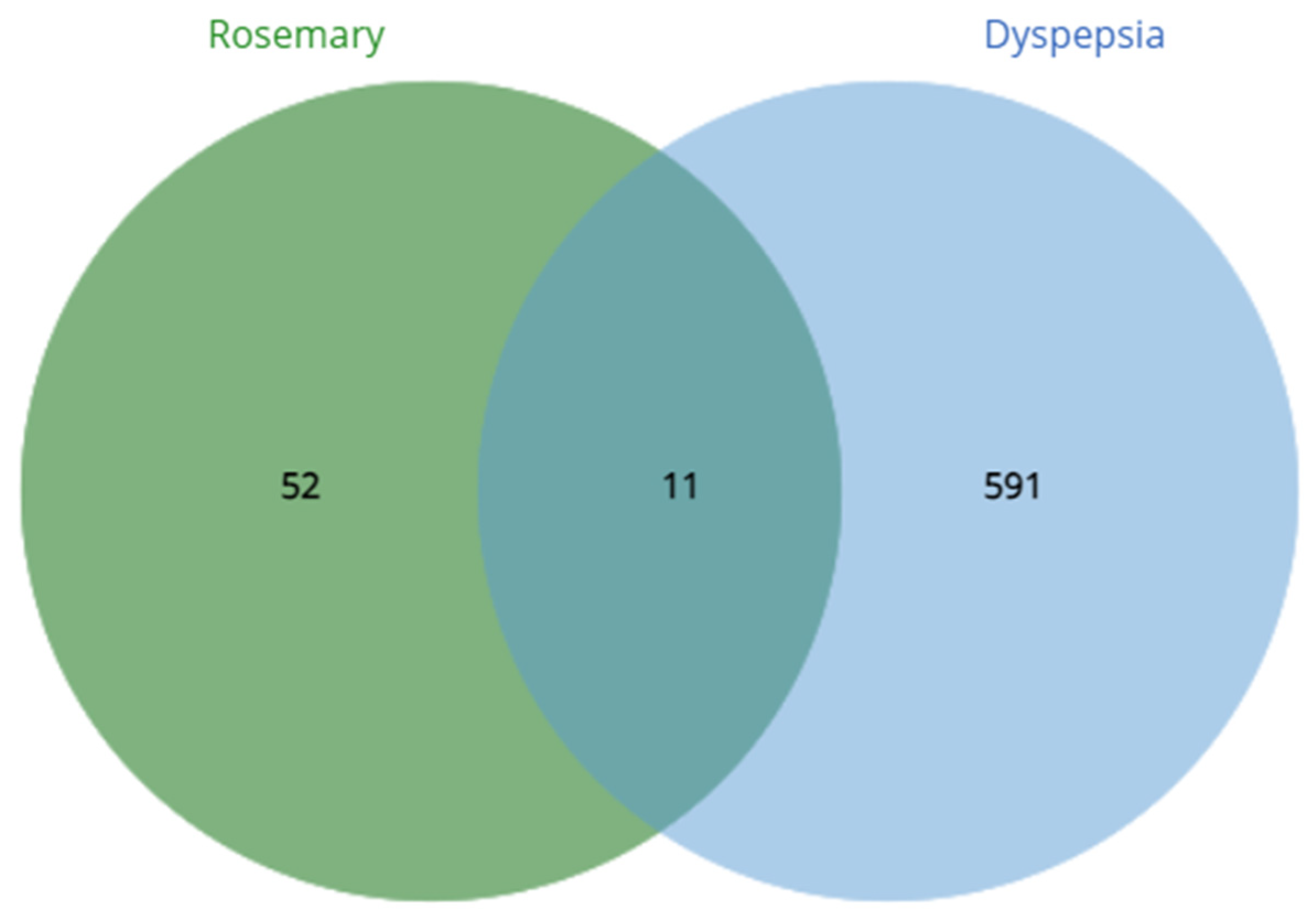
Using the GeneCard database, there were a total 2832 dyspepsia-related genes obtained and was limited to 602 genes after getting the top 25% by quantile. On the other hand, the target genes of S. rosmarinus had a total of 1000 which was narrowed down to 63 after getting the targets with probability ≥ 0.70. After the crossover, it was found that 11 common genes were found: proto-oncogene c-Kit, transthyretin, matrix metalloproteinase-9, polyunsaturated fatty acid 5-lipoxygenase, 72 kDa type IV collagenase, xanthine dehydrogenase/oxidase, broad substrate specificity ATP-binding cassette transporter, glycogen synthase kinase-3 beta, poly [ADP-ribose] polymerase 1, ADP-ribosyl cyclase/cyclic ADP-ribose hydrolase 1, and multidrug resistance-associated protein 1. The crossover of genes was done through a venn diagram and is visualized in Figure 2. Moreover, the known functions of each traced proteins can be seen in Table 4.
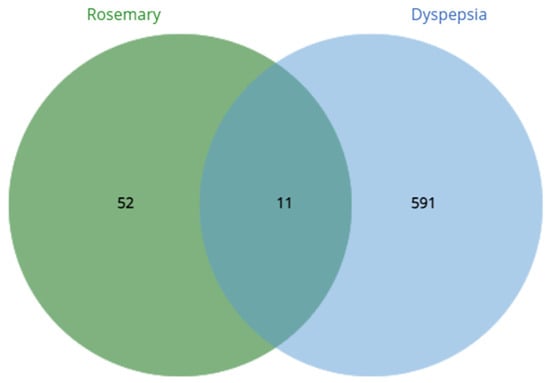
Figure 2.
Crossover of dyspepsia-related genes and S. rosmarinus targets.

Table 4.
Common genes of S. rosmarinus and dyspepsia.
3.3. PPI Network and Pathway Enrichment Analysis
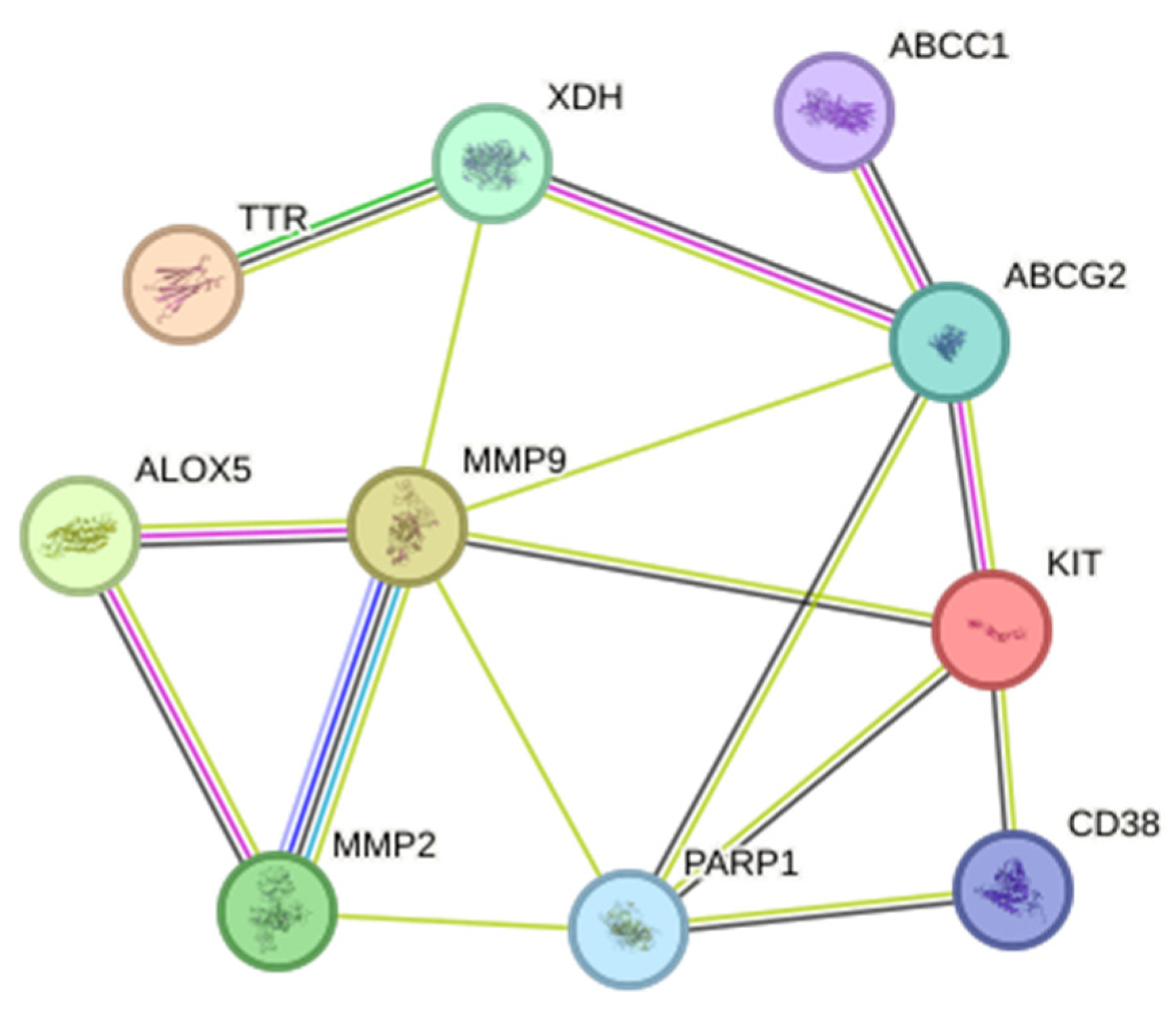
The PPI network constructed on the STRING database had a total of 11 nodes and 20 edges compared to the expected number of edges of 5. On the other hand, the average node degree was 3.64, while the average clustering coefficient was 0.715. The PPI enrichment p-value was , thus, indicating a higher protein interaction than what was expected. This also means that the proteins in the network are functionally related. On the other hand, since there were only 11 genes the application of different algorithms: MCC, MNC, and degree to filter out the hub genes was not necessary. The hub genes considered was that of the original 11 genes and can be seen in Figure 3.
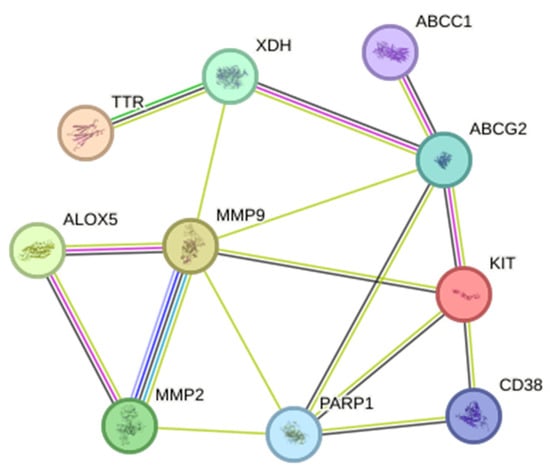
Figure 3.
Protein–protein network interaction of the common targets from S. rosmariunus and functional dyspepsia. The filled nodes indicate whether the 3D structures of the proteins are known or predicted.
3.4. Pathway and Process Enrichment Analysis
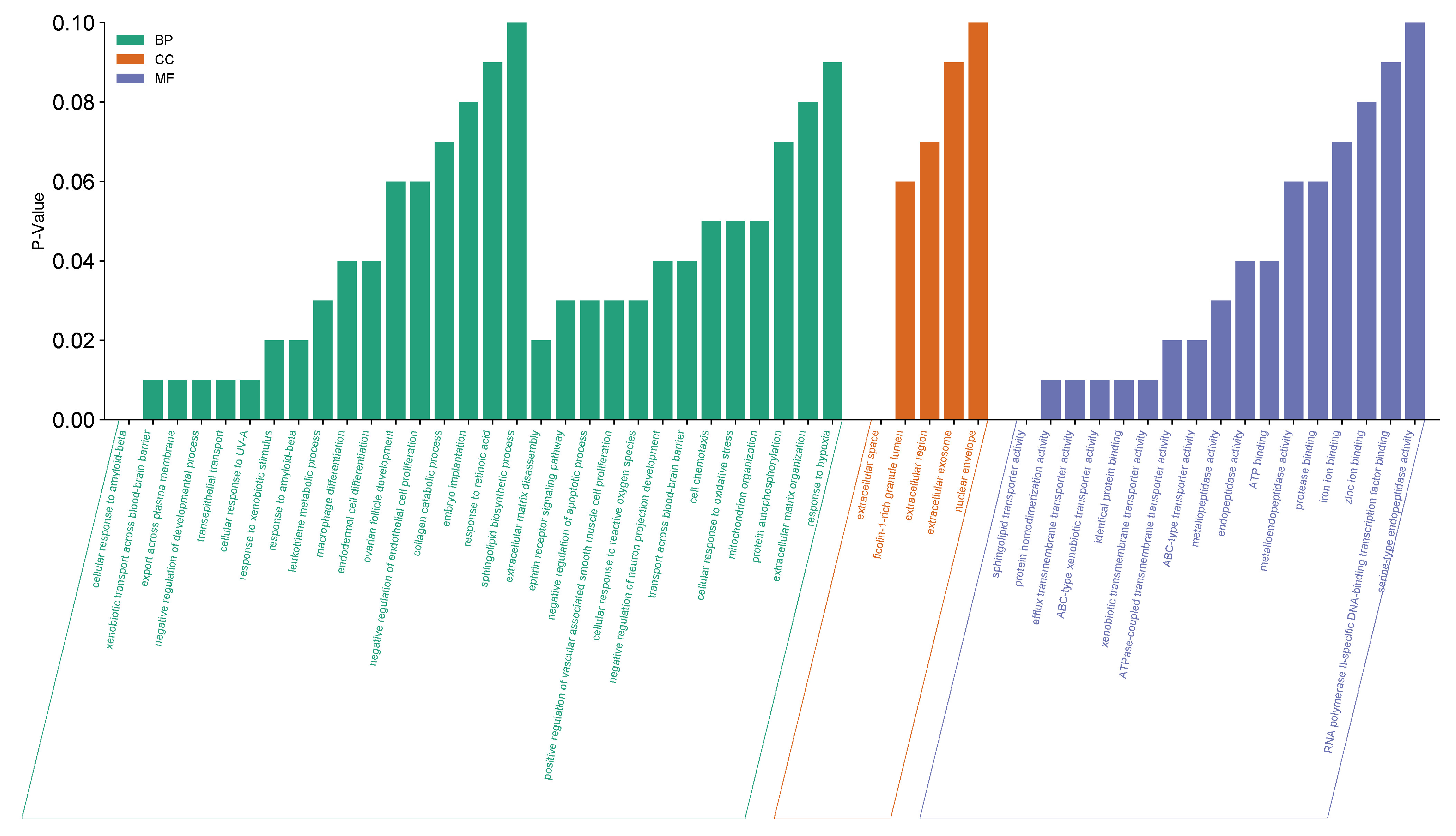
The enrichment analysis revealed 30 biological processes, 5 cellular components, and 17 molecular functions, and 13 KEGG pathways. Figure 4 shows that the biological processes are found to be concentrated on several immune responses to different triggers, as well as positive and negative regulations of several processes which indicate that S. rosmarinus can possibly control the triggers of functional dyspepsia, whether by up-regulation or down regulation of those that are involved in the development of the disease. On the other hand, the cellular components traced include most extracellular components as well as the nuclear envelope which may indicate that S. rosmarinus targets those that are outside of the cell components. Lastly, the molecular functions involved with the genes were very diverse—varying from transportation activities, enzymatic activities, and binding activities. Five of the 13 KEGG pathways traced from S. rosmarinus were found to be related in the functional dyspepsia. All of the pathways showed strong significance on the genes with a range of p-values from 0.025 to 0.075. A bar graph to visualize the terms and KEGG pathways can be seen in Figure 5. A cross-check analysis was conducted for KIT and its contribution to biological processes. Here, KIT was found to be highly significant in the Erythropoietin pathways, smooth muscle differentiation, dendritic and mast cells, sensory mechanotransduction, and megakaryocyte development. The biological processes where KIT is enriched are displayed in Figure 6.
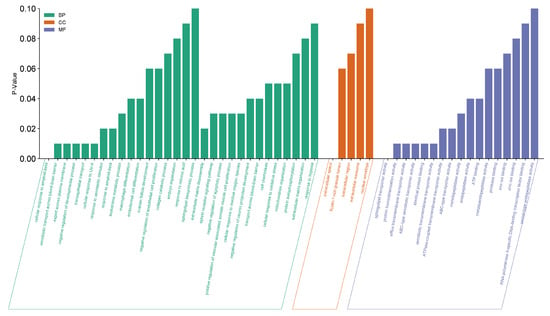
Figure 4.
Enrichment analysis of 11 hub genes from DAVID. Green bars represent the different biological processes (BP) involved with the gene. Orange bars represent cellular components (CC). While blue bars represent different molecular functions (MF).
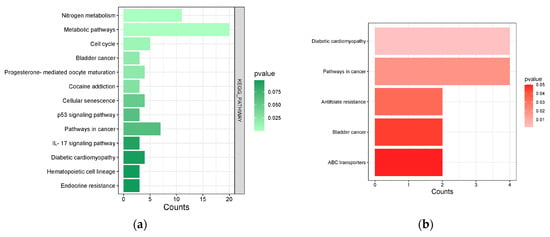
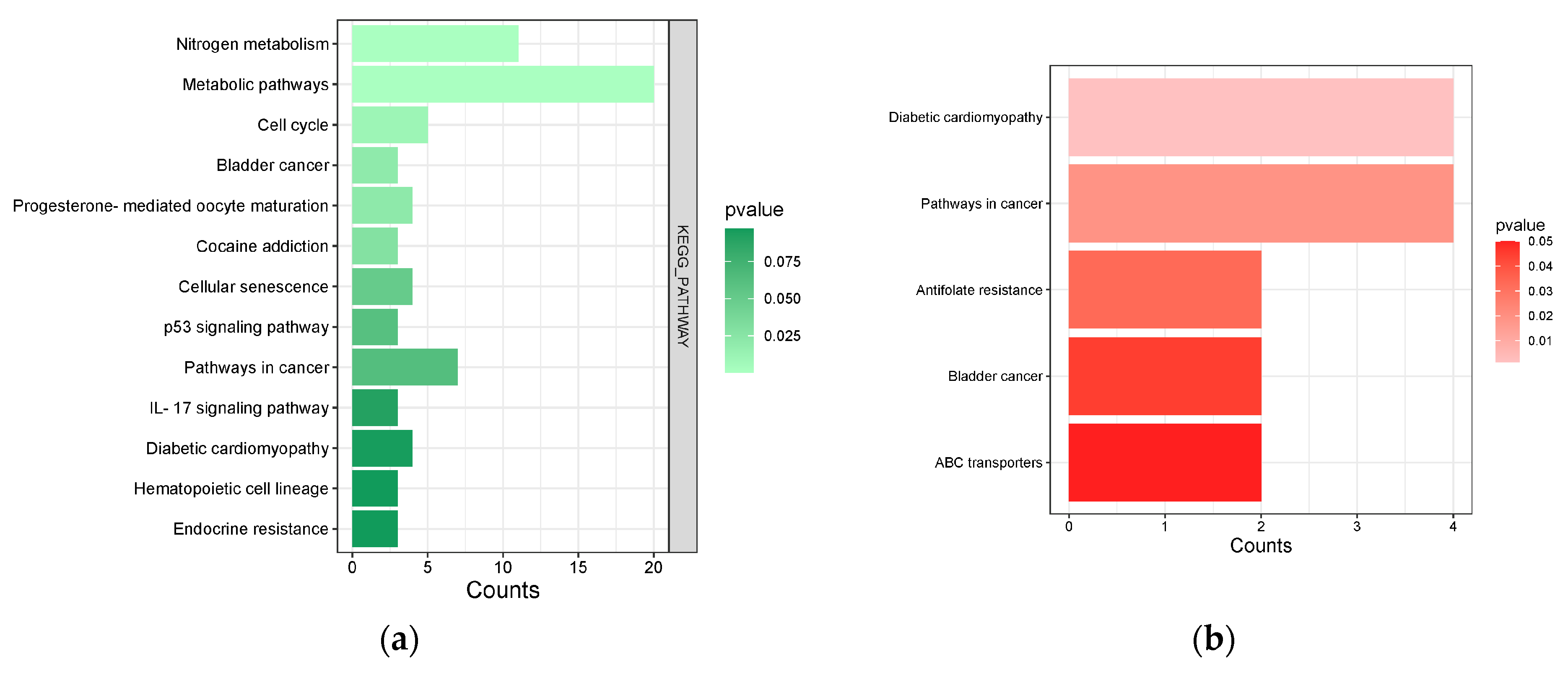
Figure 5.
(a) S. rosmarinus KEGG pathways from DAVID. Light green bars represent terms with smaller p-value, while dark green bars represent terms with higher p-value. (b) Common pathways involved in S. rosmarinus and functional dyspepsia. Light red bars represent terms with smaller p-value, while darker red bars represent terms with higher p-value.
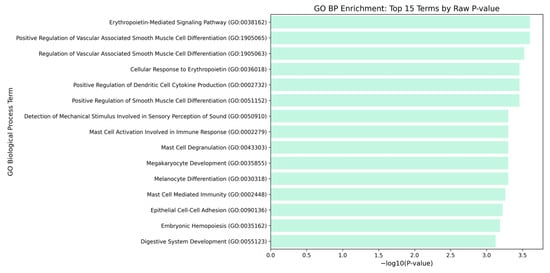
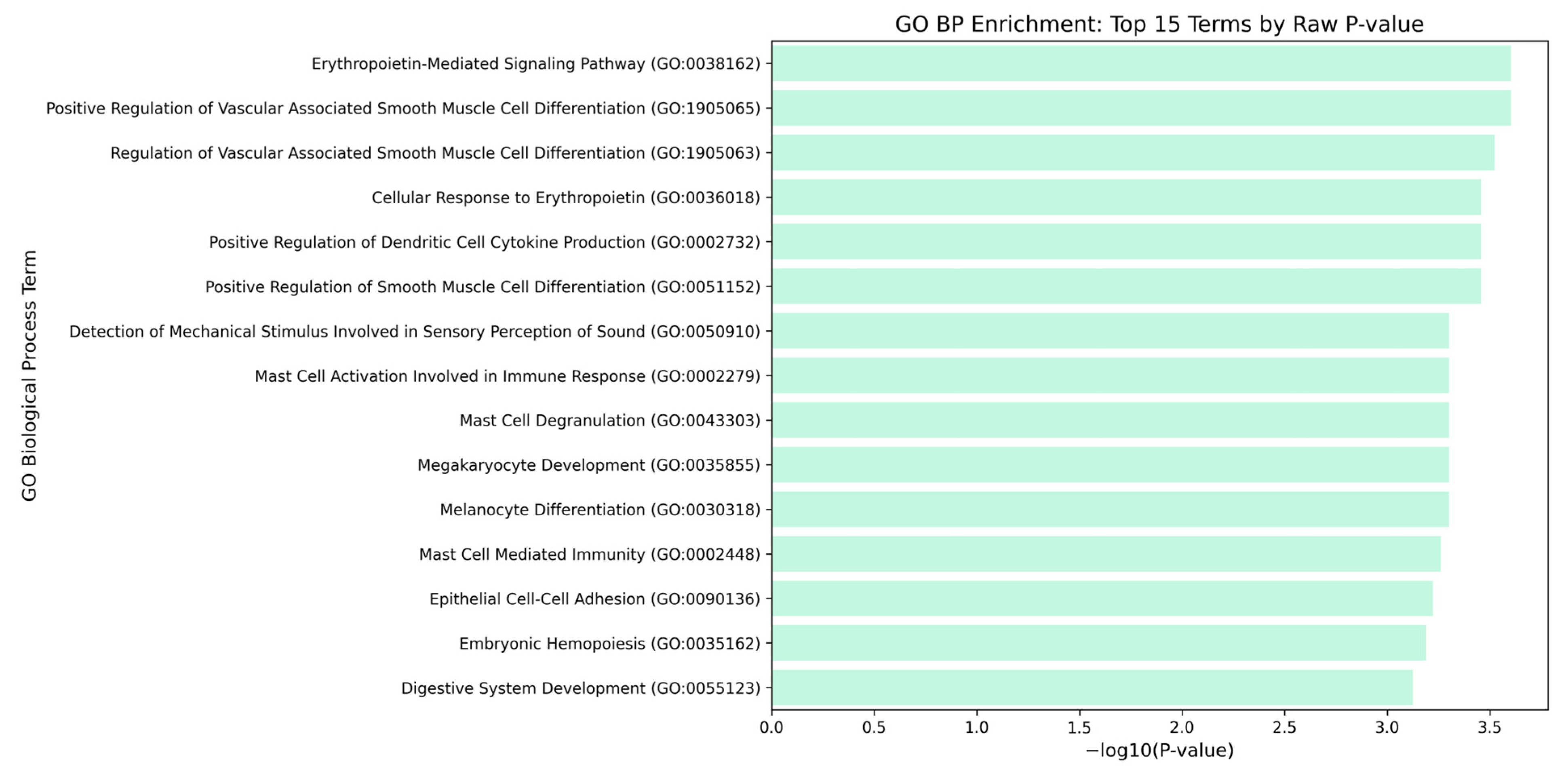
Figure 6.
Proto-oncogene c-Kit GO Biological Processes. Bars represent the statistical significance of the −log10 (p values), where longer bars indicate a stronger statistical significance (smaller p values). All displayed terms were highly significant with p value < 0.05, and most p value = 0.001.
3.5. Compound, Target, and Pathway Network
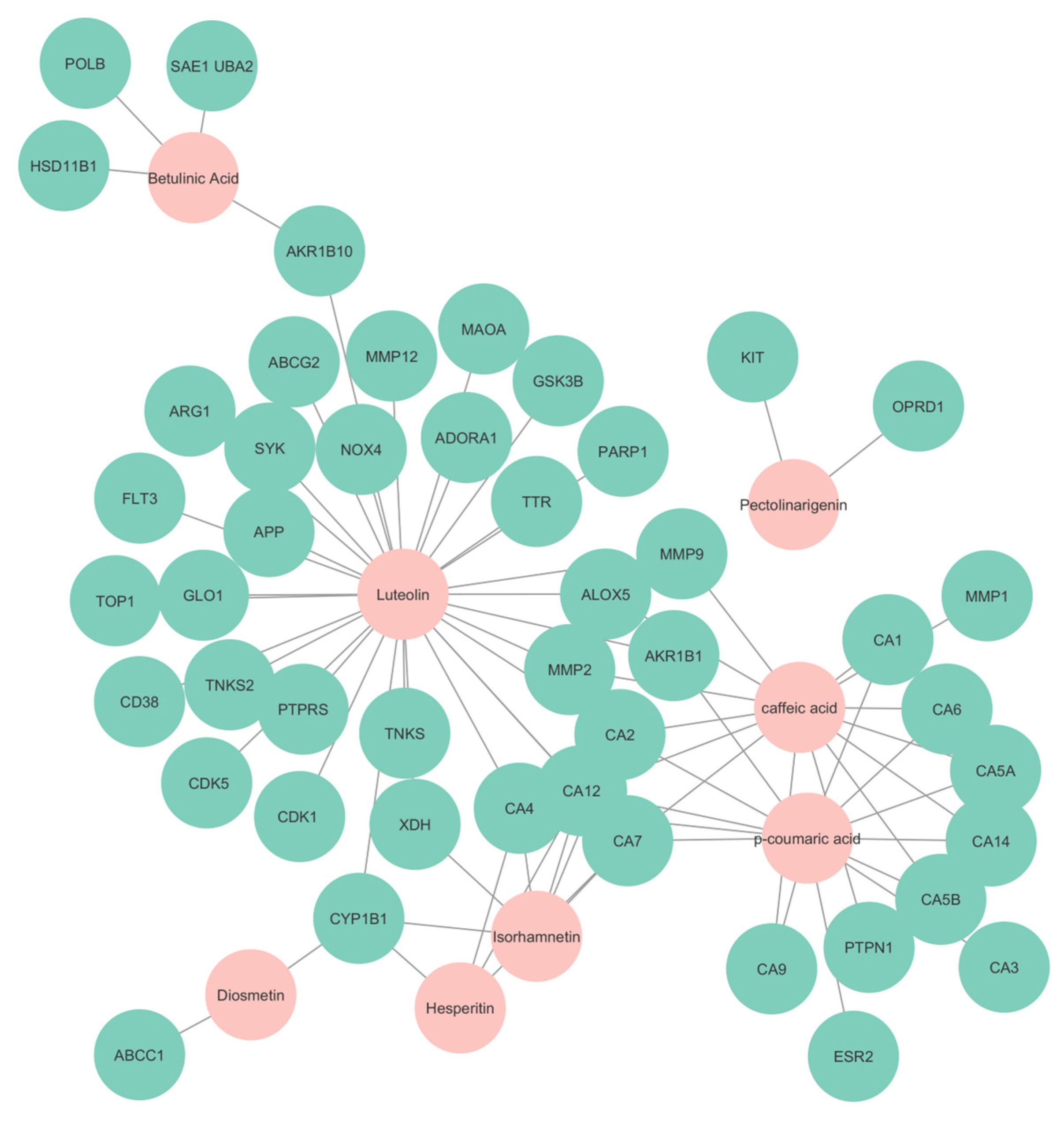
The CTP network revealed the necessary interactions within the bioactive compounds and the filtered targets. The compound and target network that was constructed can be seen in Figure 7. After the merging of the compound-target network with the pathway network, it was found that there were a total of 64 nodes and 96 edges, while the average number of neigbors was 3.082. Based on degree of association, it was revealed that the compounds with most significant interaction were luteolin, caffeic acid, p-coumaric acid, isorhamnetin, betulinic acid, hesperitin, pectolinarigenin, and diosmetin with degrees of 31, 14, 13, 6, 4, 1, and 1, respectively. These molecules are considered to be the core compounds of the plant as it indicates a greater interaction among others. It can also be suggested that these compounds will have a positive interaction with the genes related to dyspepsia.
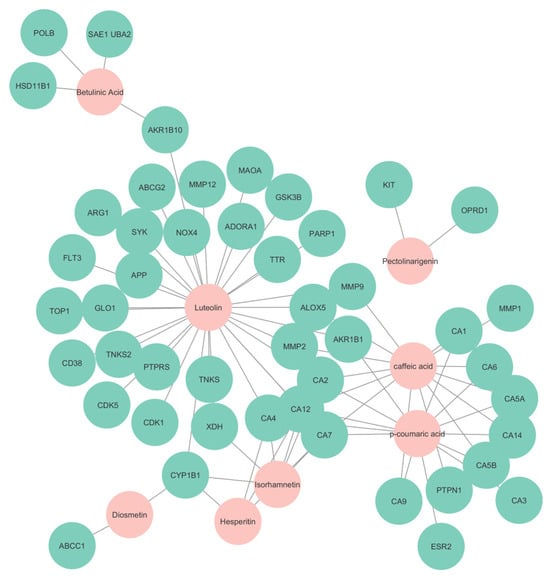
Figure 7.
Compound and target network. Nodes in cyan represent the targets predicted, while nodes in pink represent the compounds found in S. rosmarinus.
3.6. Molecular Docking of Core Compounds of S. rosmarinus Against Dyspepsia-Related Proteins
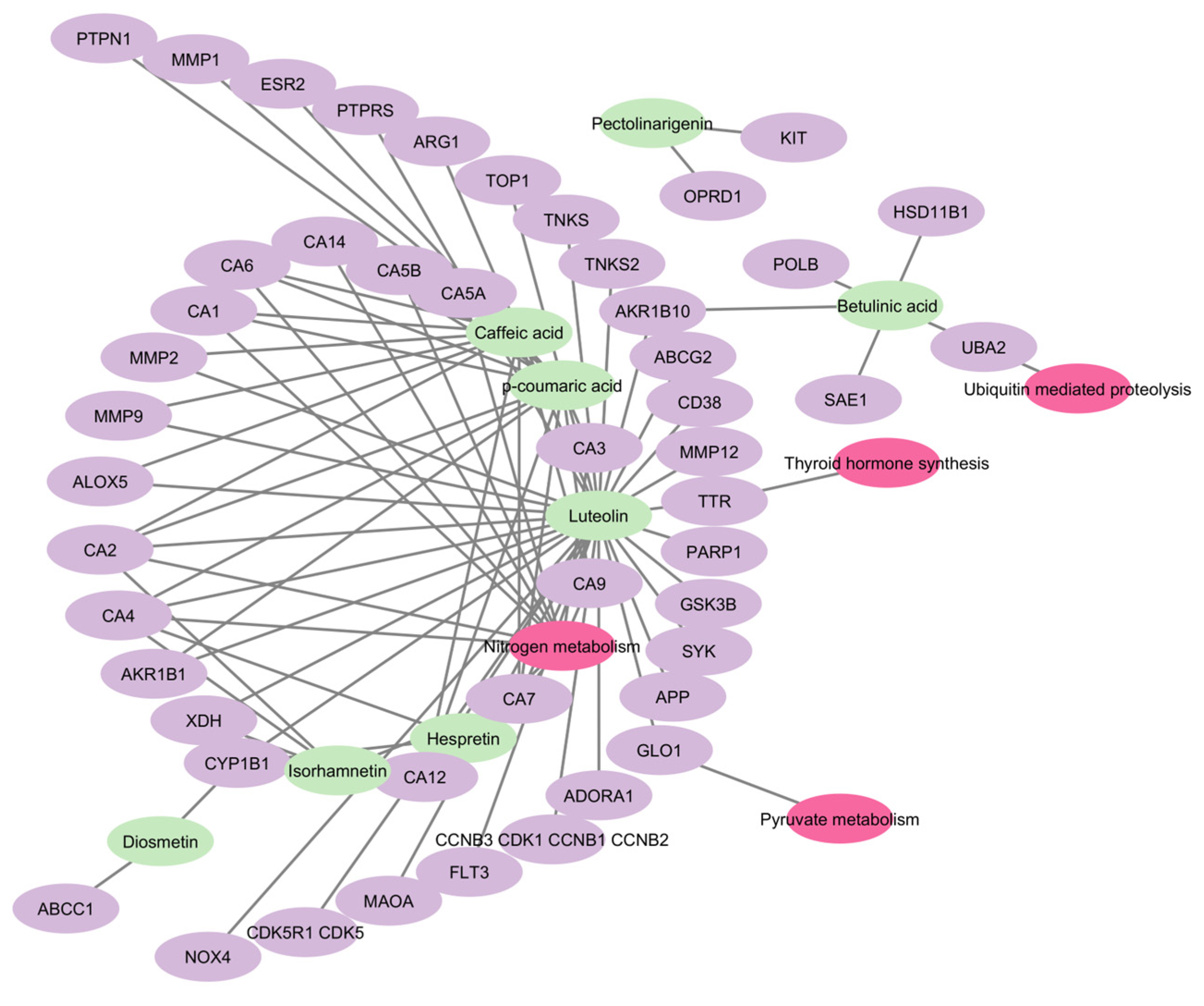
The pathways and network elucidation of S. rosmarinus with functional dyspepsia revealed the top interacting compounds within the gene set and were found to be luteolin, caffeic acid, p-coumaric acid, isorhamnetin, betulinic acid, hesperitin, pectolinarigenin, and diosmetin—indicating that these are the core compounds from the plant. Here, the complete compound, target, and pathway network is visualized in Figure 8. Out of the 8 interacting compounds, five have satisfied the drug-likeliness screening with 0 violations from all the criteria which were Lipinski, Ghose, Veber, Egan, and Muegge. Similarly, these compounds were found to have high GI absorption and have passed the screening for bioavailability and oral bioavailability. The following compounds were luteolin, isorhamnetin, hesperitin, and pectolinarigenin, and diosmentin. In relation, an additional bioactive compound, genkwanin, satisfied the said criteria and drug-likeliness. These compounds were analyzed according to their binding affinity against the hub genes.
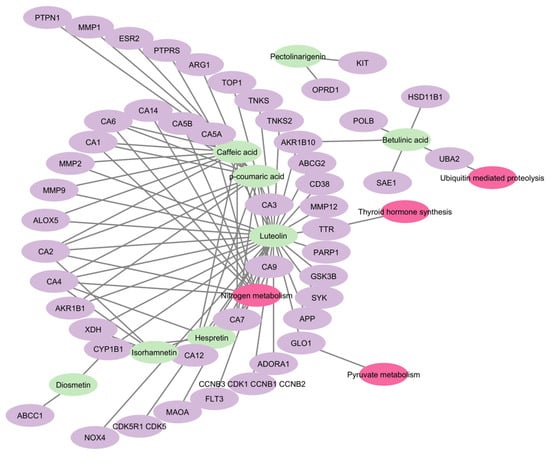
Figure 8.
Combined network of compound to target and target pathway. Green nodes represent the bioactive compounds found in S. rosmarinus, purple nodes represent the targets of the respective compounds, whole pink nodes represent the pathways in which the targets are enriched.
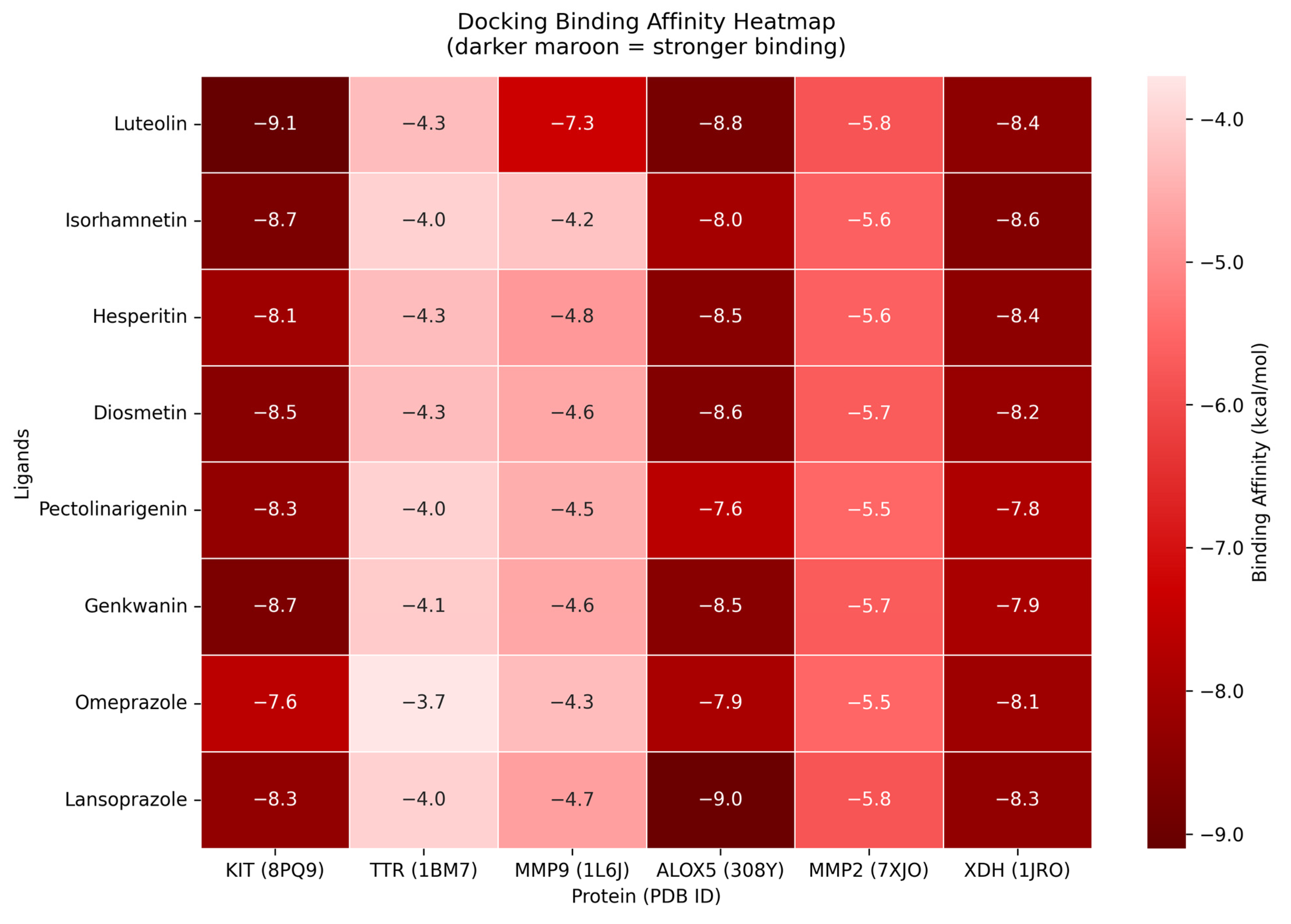
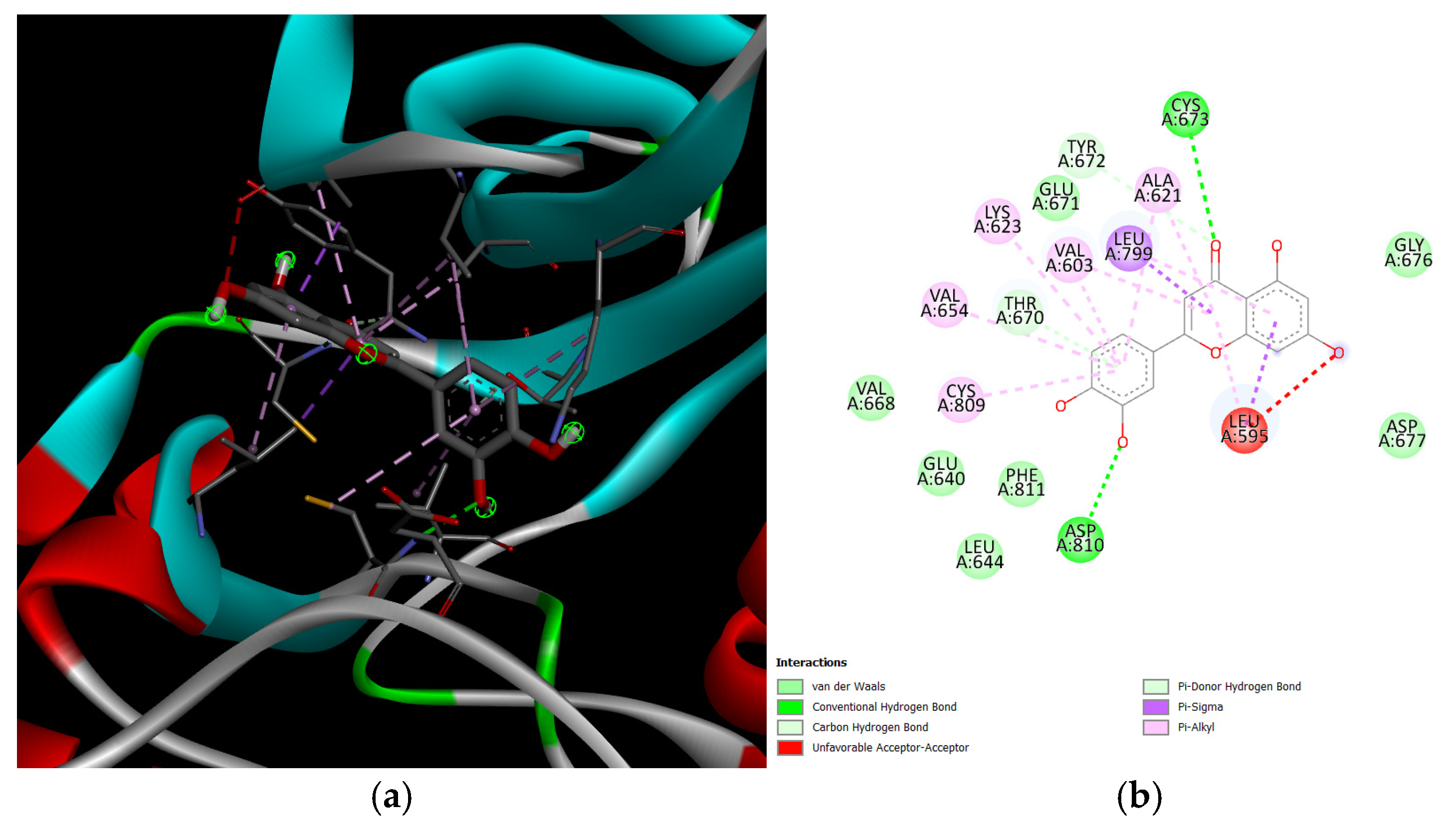
The results of the molecular docking showed that the compound with the highest binding affinity was luteolin and proto-oncogene c-Kit, with a docking score of −9.1 kcal/mol. Compared to the two controls, omeprazole and lansoprazole, luteolin exhibited a significantly higher docking score. The complete docking scores for all the protein candidates are visualized through a heatmap and can be seen in Figure 9. Moreover, the result of the docking indicates that luteolin may bind to the proteins in the pathways of dyspepsia, specifically, that of KIT. Another point of interest are the intermolecular forces acting upon the complexes. As seen in Figure 10, luteolin and KIT exhibited unfavorable acceptor-acceptor interaction, which nevertheless demonstrated key stabilizing interactions such as van der Waals, conventional hydrogen bond, and carbon hydrogen bond. These interactions suggest that the ligand retains the potential to bind within the binding sites despite the structural limitations. The models and diagram for the remaining complexes are found in Appendix A.
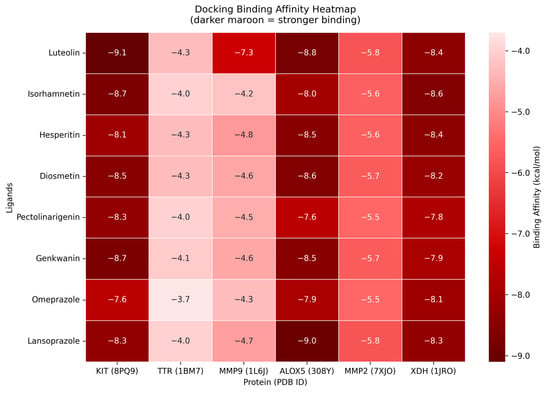
Figure 9.
Heatmap of molecular docking binding affinities. Y–axis represents the core compounds together with the compounds that have passed the drug-likeliness criteria. The x–axis represents the hub genes traced from the crossover of bioactive-targets and dyspepsia-related genes.
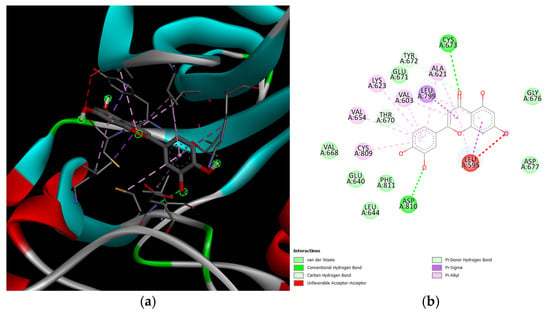
Figure 10.
KIT and luteolin complex. The leftmost figure (a) shows the interactions between the two proteins (KIT and luteolin). The rightmost figure (b) shows the 2D diagram of interactions between the two proteins, as well as the intermolecular forces acting upon the complex.
3.7. Molecular Dynamics Simulation Results
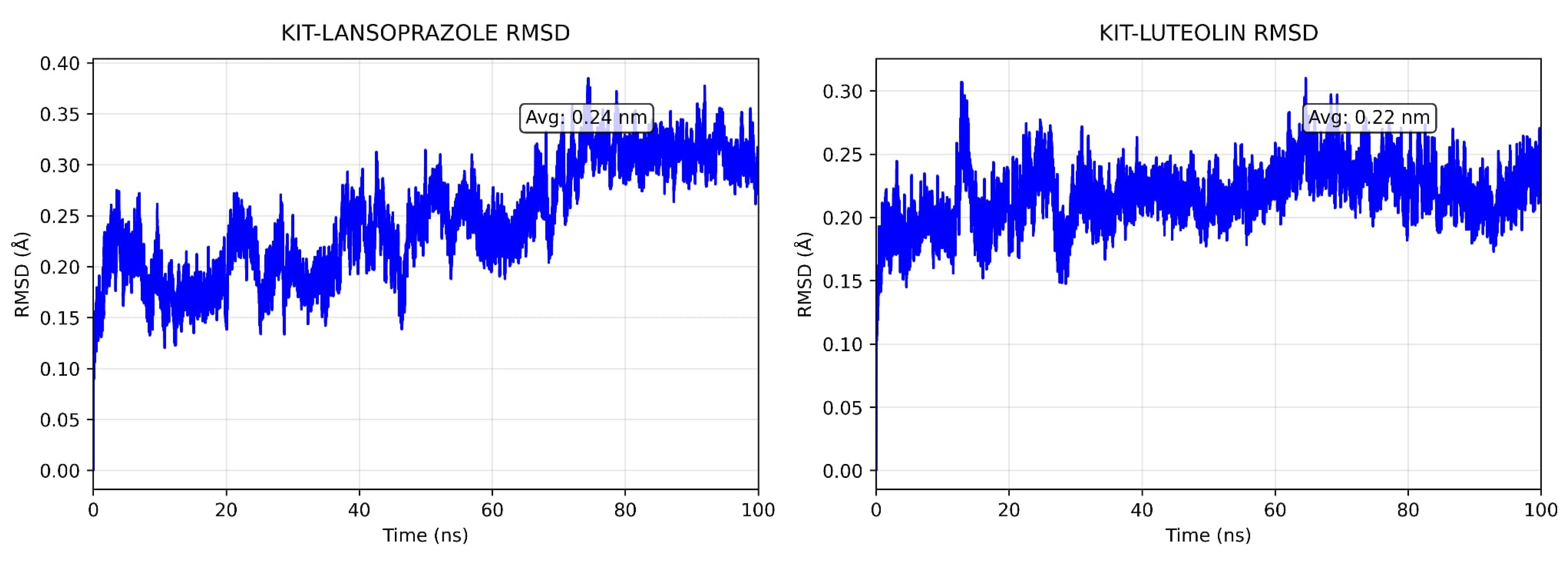
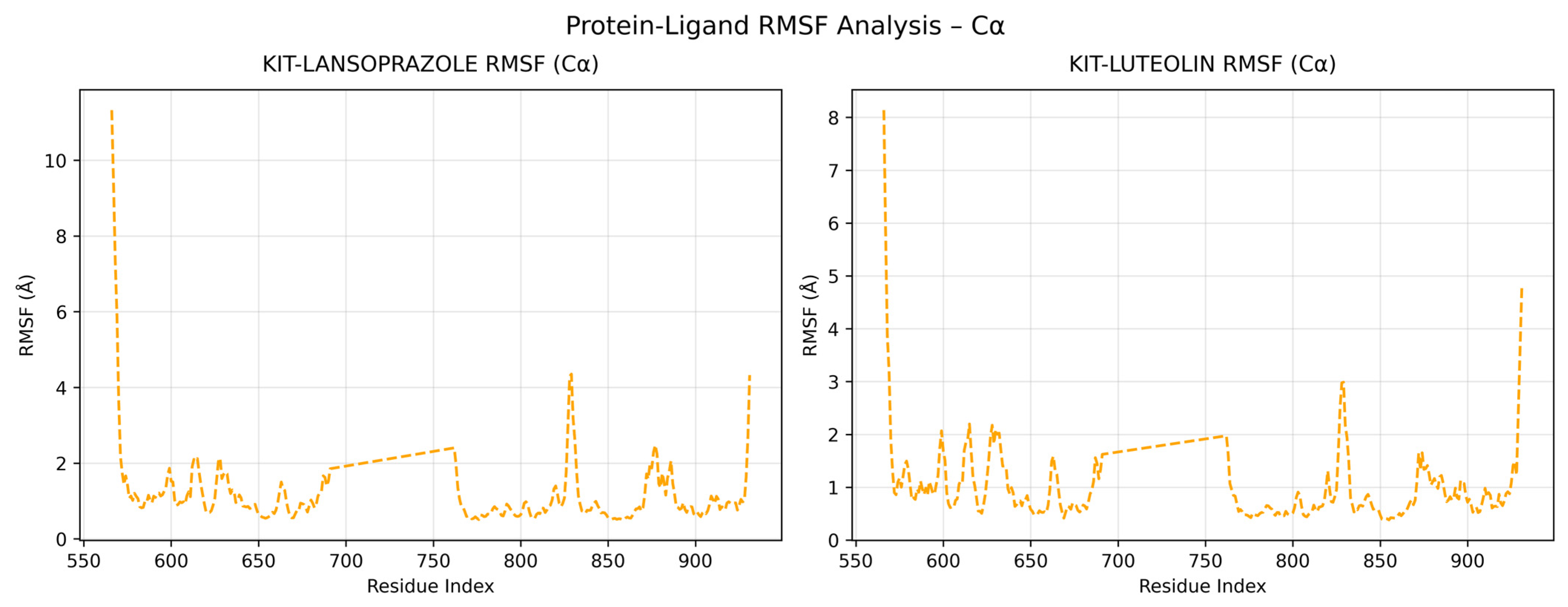
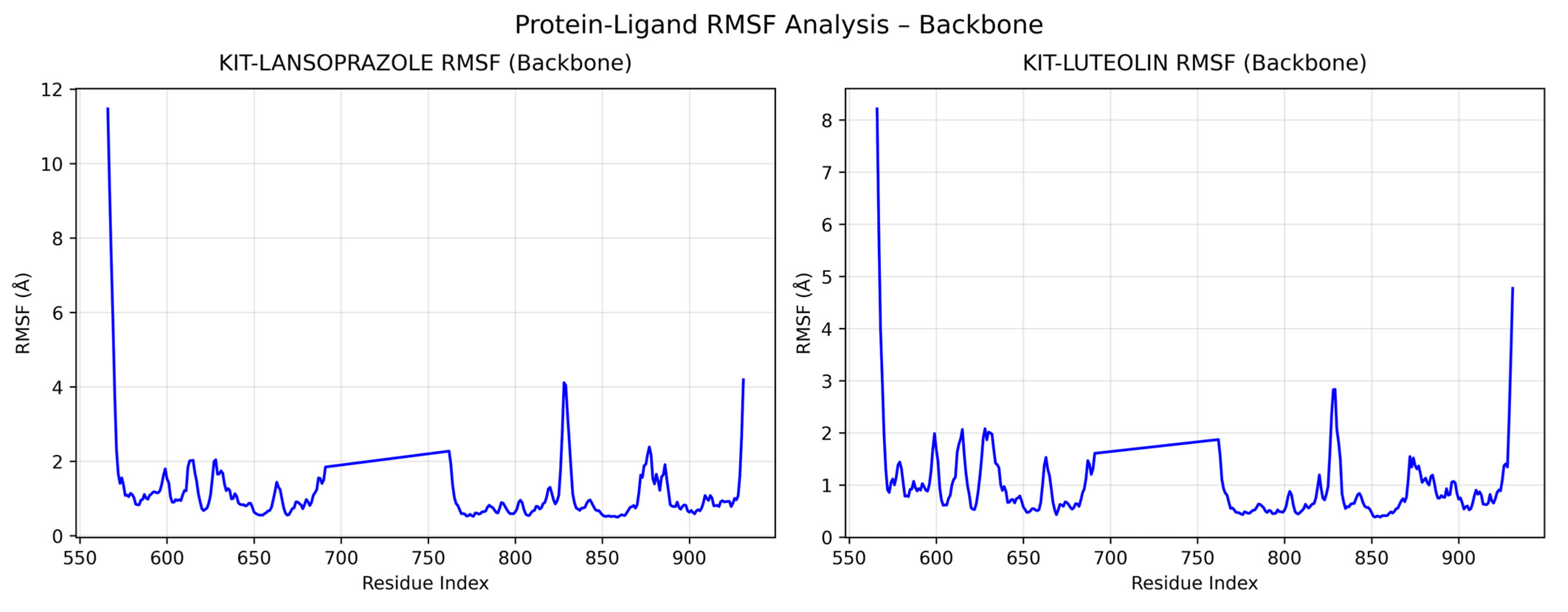
A comparative analysis on molecular dynamics of the KIT-luteolin (test) complex against the KIT-lansoprazole (control) complex was done to verify the nature of stability of the top-binding compound at 100 ns simulation. The results of the simulation revealed that there is a slight difference between the control complex and the test complex in terms of their stability. As seen in Figure 11 the KIT-lansoprazole complex exhibited a slightly less stable RMSD of 0.24 nm, while KIT-luteolin complex exhibited a slightly more stable RMSD of 0.22 after 20 ns. The protein-ligand residue fluctuations analysis (RMSF) on the other hand, revealed that the both complexes have localized flexibility, particularly at the N-terminal region for both alpha carbon (Cα) atoms only, and all backbone atoms considered. However, the KIT-lansoprazole complex showed a higher peak RMSF value as seen in Figure 9 and Figure 10, indicating that luteolin binding conferred higher restrains. Equally important is the radius of gyration analysis in which both complexes exhibited a closely related compactness where the KIT-lansoprazole yielded an average radius of gyration equal to 19.96 nm, while the KIT-luteolin yielded an average radius of gyration equal to 19.56 nm. Another factor is to take into account is the hydrogen bonding patterns. Throughout the 100 ns simulation, the KIT-luteolin complex consistently maintained a significantly higher number of hydrogen bonds, with an average of 2.26 and a maximum of 6 hydrogen bonds. Conversely, the KIT-lansoprazole complex formed virtually no stable hydrogen bonds, with an average of 0.00 and not exceeding a maximum of 1. This underscores the superiority of luteolin’s binding affinity and stability which is also facilitated by multiple and sustained intermolecular forces within the KIT binding pockets. All things considered, the molecular dynamics data point to Luteolin being a more favorable ligand for KIT, as compared to lansoprazole. The comparison of alpha carbons and all backbone residues for the KIT and lansoprazole, as well as KIT and luteolin complexes are visualized in Figure 12 and Figure 13, respectively.
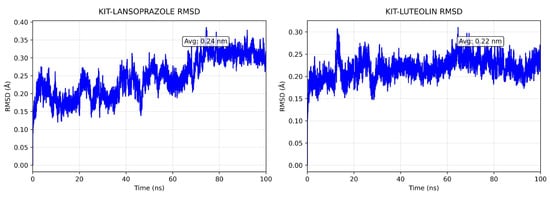
Figure 11.
Comparison of backbone RMSD for KIT-Lansoprazole and KIT-luteolin complexes.
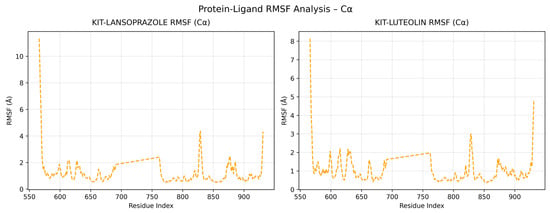
Figure 12.
Comparison of alpha carbon residue fluctuations for KIT-Lansoprazole and KIT-luteolin complexes.
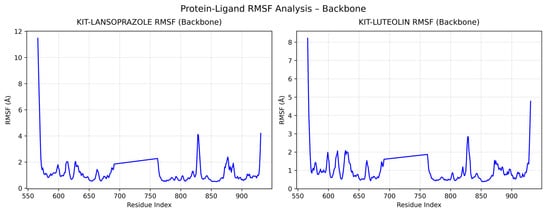
Figure 13.
Comparison of all backbone residue fluctuations for KIT-Lansoprazole and KIT-luteolin complexes.
It is also worth mentioning that Cliff’s Delta was applied to furhter evaluate the results of the comparative molecular dynamics between the test complext and control complex. The three key parameters were RMSD, radius of gyration, and hydrogen bonds. As seen in Table 5, the RMSD values were not large enough to imply a significant difference in te overall backbone stability between the two complexes. Despite this result, the radius of gyration and the hydrogen binds shows a strong favor to KIT-luteolin complex, with an enhanced overall compactness and a considerable higher amount of hydrogen bonds. This indicates that luteolin binding may lead to a greater conformational stability.

Table 5.
Cliff’s delta analysis summary.
4. Discussion
Functional dyspepsia is a type of digestive disorder which can be persistent and will cause discomfort in the upper stomach without any cause [69]. Although the mechanism of development of dyspepsia is yet to be known until now, several studies have related the central factors that might trigger the occurrence of functional dyspepsia including stress and anxiety and is a precursor to gastroesophageal reflux diseases, peptic ulcer, gastric cancer, and gastroparesis [70,71]. Several drugs have been developed to treat dyspepsia and its related diseases which are mainly proton-pump inhibitors. Some of these include omeprazole and lansoprazole and are commercially available [72]. These drugs can both provide short reliefs to gastric symptoms of the stomach and can possibly pose side effects which is not efficient to persistent dyspepsia. With the rising opportunities for herbal medicines through several scientific validations, the molecules from S. rosmarinus have been analyzed for their possible drug-likeliness and interactions against functional dyspepsia.
The network pharmacology results revealed that the bioactive compounds of S. rosmarinus can interact with dyspepsia-related genes. Through the compound-target network, it was found that the compound with the highest degree of interaction was luteolin. Consequently, the molecular docking revealed that among all protein-ligand complexes, luteolin showed the most activity with a good binding affinity against most of the dyspepsia-related genes. Several compounds including isorhamnetin, hesperitin, diosmetin, pectolinarigenin, and genkwanin have also exhibited plausible interactions with spontaneous binding affinities against the disease, indicating that the herb can possibly treat dyspepsia. KIT-luteolin complex indicates the highest affinity even surpassing omeprazole and lansoprazole. This gives the possibility of further testing into possible applications for treating dyspepsia. Despite the presence of the unfavorable acceptor-acceptor bond, it is still worth highlighting the consistency of the other intermolecular forces including van de Waals, conventional hydrogen bond, and carbon hydrogen bond. Another point of interest is the molecular dynamics simulation conducted to validate the results of the molecular docking. Here, KIT-Luteolin complex was analyzed in comparison to KIT-Lansoprazole complex which is the control with highest binding affinity to KIT. The simulation revealed that Luteolin is a more favorable ligand, as opposed to Lansoprazole. The bioactive compound exhibited a more consistent hydrogen bonding, and more compact protein-ligand complex. The molecular dynamics simulation was then validated by Cliff’s Delta analysis where a small positive effect in RMSD, a highly negative effect in terms of radius of gyration, and a substantial amount of hydrogen bonds. These results further substantiate the potential of Luteolin as a ligand compared to Lansoprazole.
Luteolin, isorhamnetin, hesperetin, diosmetin, pectolinarigenin, and genkwanin are flavonoids which were studied to have potential therapeutic effects. Several studies discussed the various pharmacological activities of luteolin. One of which is its capability to produce anti-inflammatory effects through the inhibition of nuclear factor-kꞵ (NF-kꞵ) to reduce interleukin-8 mRNA expression in human intestinal Caco-2 cells, oxidative stress inhibition to activate the nuclear factor erythroid 2-related factor 2 (Nrf2) pathway, and regulating intestinal flora [73,74,75]. Moreover, isorhamnetin was found to exhibit various biological activities including anti-inflammatory and anti-gastric cancer effects [76]. In a study by Ricardo et. al., isorhamnetin was found to exhibit inhibitory effects on Tumor Necrosis Factor-ɑ (TNF-ɑ) and interleukin-6 (IL-6) [77]. Similarly, lipopolysaccharide (LPS)-induced models for inflammation studies revealed that hesperetin showed anti-inflammatory effects during treatments. A study by Yang et. al., presented that hesperetin metabolites are capable of downregulating LPS-induced prostaglandin E2 (PGE2) which is an important inflammatory product of COX-2 activity in smooth muscle cells. Additionally, hesperetin metabolites also exhibited suppression of NF-kꞵ in the same model for raw cells [78]. This indicates that hesperetin might also act as an inhibitor for the dyspepsia-related genes. Research indicates the use of diosmetin for treatment shows 90% reduction of growth due to inhibiting H. pylori by interfering with its biological functions [79]. In ethnopharmacology, pectolinarigenin has been known to have inhibitory effects which effectively treat various human ailments such as ulcers, stomach aches, and several more [78,80]. Genkwanin shows antagonist properties in inflammation which could possibly inhibit dyspepsia providing relief from the discomfort and pain [81,82]. The majority of the mechanisms of the six compounds were found to be mostly inhibition in terms of other diseases. This suggests that these compounds can possibly have inhibitory effects against dyspepsia-related genes in relieving intestinal discomfort.
The KEGG pathway analysis revealed that the predicted targets of S. rosmarinus are involved in several pathways. One of which is the metabolic pathway which is responsible for producing metabolites from breaking down of molecules [83]. In the context of functional dyspepsia, several metabolic pathways were found to be altered in an individual with functional dyspepsia. These metabolic pathways were said to be involved with energy metabolism, amino acid, gut microbiota, and host-co metabolism [84]. Furthermore, the gastrointestinal microbiota is related to the metabolic process in the human body, whereas, intestinal contents along with endogenous mucus secreted in the intestinal epithelium are used to produce metabolites including short-chain fatty acids (ScFAs), cholic acid, choline metabolites, phenols, lipids, and carbohydrates which can either be harmful or beneficial to the human body [85]. Thus, alterations in the pathway can contribute to the development of functional dyspepsia. Interestingly, this pathway exhibited a relationship with incredibly high number of genes with the lowest p-value compared to other pathways that were traced. There were also a plausible number of genes related to the nitrogen metabolism, most of which were carbonic anhydrases. This enzyme was known to form carbonic acid which can neutralize intracellular hydroxyl ions from luminal acid secretion [86]. Another pathway that was traced was the IL-17 pathway which is responsible in many inflammatory diseases [87]. Although the p-value of the genes in the pathway was the highest, however, a low p-value still suggests significance. The same pathway was also found to be associated with H. pylori infection where dyspepsia is once again a precursor by triggering gastrointestinal inflammation [88,89]. Furthermore, increased levels of IL-17 were also correlated with the symptoms of some gastrointestinal disorders including dyspepsia [90]. Nevertheless, all pathways that were traced were still significantly related to S. rosmarinus with smaller p-value, indicating that they are statistically significant despite the small count of genes involved [91].
Among the 13 pathways that were traced, five pathways were found related to dyspepsia-related genes. The top three pathways with lowest p-values were diabetic cardiomyopathy, pathways in cancer, and antifolate resistance. The diabetic cardiomyopathy pathway involves the matrix metalloproteinase enzyme which is involved in the growth factor beta (TGF-β) synthesis which is a master regulator of fibrosis. In relation, several studies suggested that the pathogenesis of dyspepsia can be found in the gastric mucosa in chronic gastritis and fibrosis [92]. Additionally, poly [ADP-ribose] polymerase 1 is involved in the inhibition of glyceraldehyde 3-phosphate dehydrogenase which is known for its primarily known for its role in glycolysis. This phenomenon suggests that GADPH might play a role in the regulatory functions of dyspepsia.
Moreover, the proto-oncogene receptor tyrosine kinase (KIT) gene was found to be involved in the MAPK signaling pathway. This pathway is responsible for the downregulation of Glial Cell Line-Derived Neutrophic Factor (GDNF)—resulting to the overexpression of microRNA-211 the expression of which in dyspepsia was found to be significantly increased [93]. The matrix metalloproteinases were found to be related to the sustained angiogenesis, which is closely associated with chronic inflammation due to H. pylori infection. Here, the upregulation of pro-angiogenic factors can lead to changes in the gastric mucosa, which can lead to dyspepsia [94]. The enrichment analysis of KIT through gene ontology biological processes also provides a valuable insight into functional dyspepsia. Among the most notable pathways are those associated with rythropoietin signaling, smooth muscle cell differentiation, immune cell activation, and mechanosensory perception which are all interconnected in the disruption of gut motility, immune regulation, and visceral sensitivity.
5. Conclusions
In light of these findings, it can be concluded that the bioactive compounds of S. rosmarinus can potentially treat functional dyspepsia. Results have shown that luteolin, isorhamnetin, hesperitin, diosmetin, pectolinarigenin, and genkwanin might pose possible pose possible effects on the dyspepsia-related proteins: KIT, TTR, MMP9, ALOX5, MMP2, and XDH. The pathway analysis revealed five pathways related to both the bioactive compounds of S. rosmarinus and dyspepsia-related genes. The top three pathways were found to be diabetic cardiomyopathy, pathways in cancer, and antifolate resistance. These path-ways were also found to be involved in the upregulation and downregulation of sever-al proteins involved in the pathogenesis of dyspepsia. The simulations also confirmed that luteolin that exhibited low binding energy requirement through molecular docking, validated by the strong interaction to KIT through molecular dynamics. Luteolin outperformed the controls, omeprazole and lansoprazole in most metrics, suggesting that it has promising potential as a natural therapeutic agent for treating functional dyspepsia. Substantiated by the bioavailability and oral bioavailability of the compound, luteolin may pose less risk and side effects than that of the existing drugs. While this study focused on the six bioactive compounds of S. rosmarinus, ABCG2, GSK-3-β, PARP1, CD38, and ABCC1 were also identified but not subjected to in-depth analysis. Nevertheless, this does not diminish the value of the present findings, rather, can be subjected to future studies to explore the full therapeutic potential of S. rosmarinus in functional dyspepsia.
Author Contributions
Conceptualization, G.M.C.D.; methodology, G.M.C.D.; software, G.M.C.D.; validation, G.M.C.D.; formal analysis, G.M.C.D.; investigation, G.M.C.D. and E.J.G.V.C.; resources, G.M.C.D. and E.J.G.V.C.; data curation, G.M.C.D.; writing—original draft preparation, G.M.C.D. and E.J.G.V.C.; writing—review and editing, G.M.C.D. and E.J.G.V.C.; visualization, G.M.C.D.; supervision, H.S.C.; project administration, H.S.C.; funding acquisition, H.S.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
Appendix A
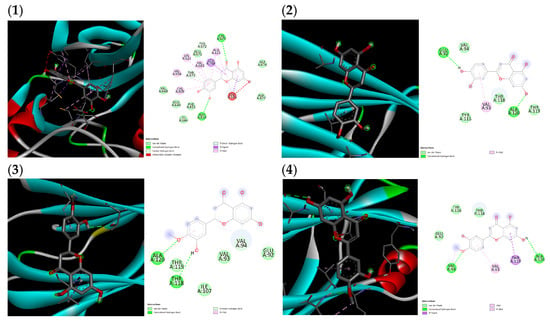
Figure A1.
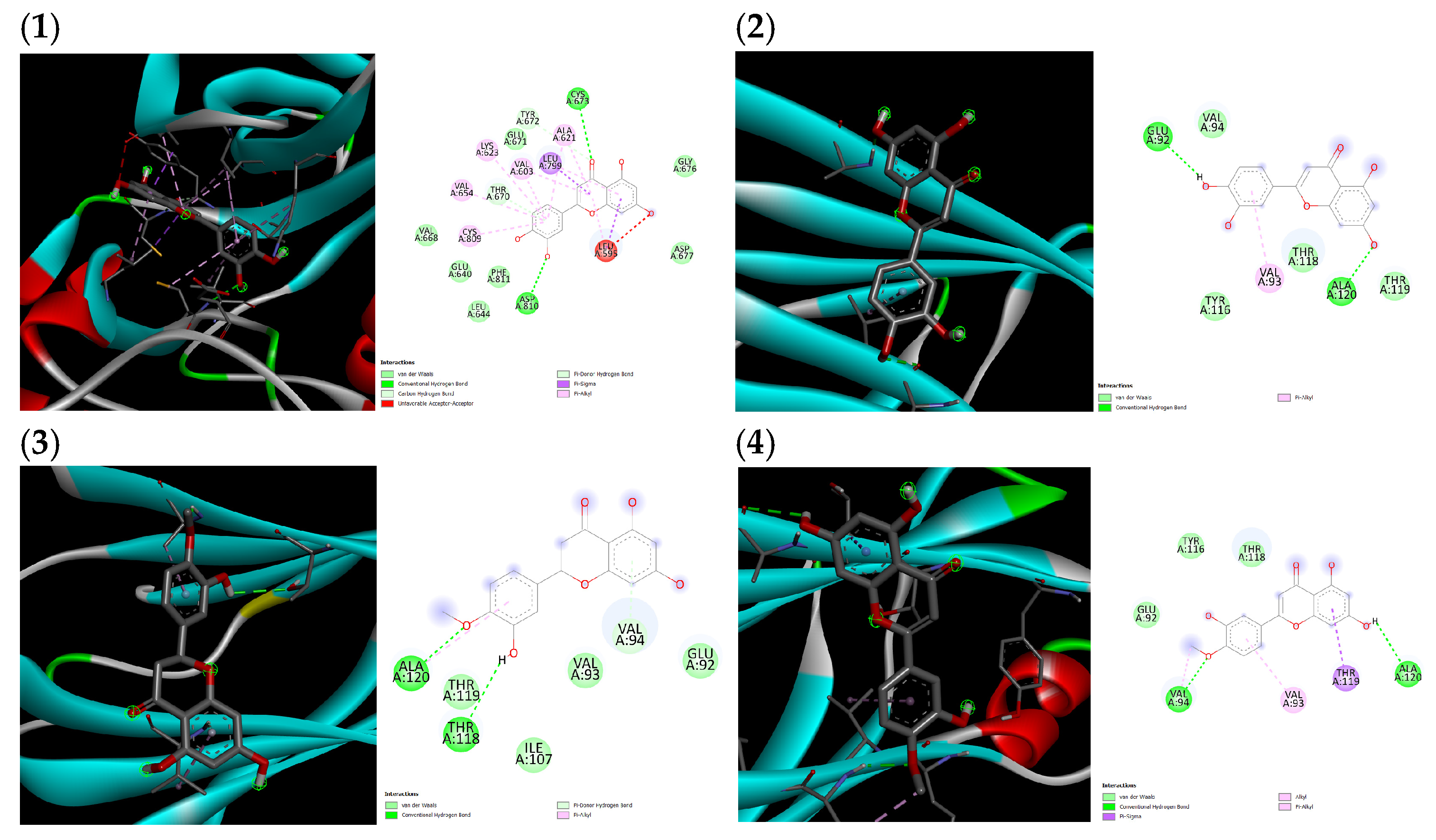
(1) Proto-oncogene c-Kit and luteolin complex, (2) transthyretin and luteolin complex, (3) transthyretin and hesperitin complex, (4) transthyretin and diosmetin complex.
Figure A1.
(1) Proto-oncogene c-Kit and luteolin complex, (2) transthyretin and luteolin complex, (3) transthyretin and hesperitin complex, (4) transthyretin and diosmetin complex.
References
- Francis, P.; Zavala, S.R. Functional Dyspepsia. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2025. [Google Scholar]
- Newman, T. What to know about indigestion or dyspepsia. Medical News Today, 11 January 2024. [Google Scholar]
- Quebral, E.P.B.; Badua, C.L.D.; Tantengco, O.A.G. Helicobacter pylori infection and the risk of gastric cancer in the Philippines. Lancet Reg. Health West. Pac. 2022, 23, 100475. [Google Scholar] [CrossRef]
- King, L.M. Indigestion (Dyspepsia): Symptoms, Causes, and Treatments; Web MD: Newark, NJ, USA, 2024. [Google Scholar]
- National Institute of Diabetes and Digestive and Kidney Diseases. Abdominal Adhesions; National Institute of Diabetes and Digestive and Kidney Diseases: Bethesda, MD, USA, 2016.
- Phillips, M.M. Gastroesophageal reflux disease. In Medical Encyclopedia; National Library of Medicine: Bethesda, MD, USA, 2024. [Google Scholar]
- Cleveland Clinic. Functional Dyspepsia: Symptoms, Diet, Treatment & Living with; Cleveland Clinic: Cleveland, OH, USA, 2025. [Google Scholar]
- Mayer, E.A.; Nance, K.; Chen, S. The Gut–Brain Axis. Annu. Rev. Med. 2022, 73, 439–453. [Google Scholar] [CrossRef]
- Mak, A.D.; Wu, J.C.; Chan, Y.; Chan, F.K.; Sung, J.J.; Lee, S. Dyspepsia is strongly associated with major depression and generalised anxiety disorder - a community study. Aliment. Pharmacol. Ther. 2012, 36, 800–810. [Google Scholar] [CrossRef]
- National Institute of Diabetes and Digestive and Kidney Diseases. Treatment of Indigestion. NIDDK; National Institute of Diabetes and Digestive and Kidney Diseases: Bethesda, MD, USA, 2025.
- Talley, N.J.; Schmidt, M.; Paré, P.; Duckworth, M.; Räisänen, P.; Pap, A.; Kordecki, H.; Schmid, V. Efficacy of omeprazole in functional dyspepsia: Double-blind, randomized, placebo-controlled trials (the Bond and Opera studies). Aliment. Pharmacol. Ther. 1998, 12, 1055–1065. [Google Scholar] [CrossRef]
- Waller, D.G.; Sampson, A.P. Dyspepsia and peptic ulcer disease. In Medical Pharmacology and Therapeutics; Elsevier: Amsterdam, The Netherlands, 2018; pp. 401–410. [Google Scholar] [CrossRef]
- MedlinePlus. Omeprazole: MedlinePlus Drug Information; MedlinePlus: Bethesda, MD, USA, 2023. [Google Scholar]
- Mayo Clinic. Omeprazole (Oral Route) Precautions; Mayo Clinic: Rochester, MN, USA, 2025. [Google Scholar]
- Suzuki, H.; Kusunoki, H.; Kamiya, T.; Futagami, S.; Yamaguchi, Y.; Nishizawa, T.; Iwasaki, E.; Matsuzaki, J.; Takahashi, S.; Sakamoto, C.; et al. Effect of lansoprazole on the epigastric symptoms of functional dyspepsia (ELF study): A multicentre, prospective, randomized, double-blind, placebo-controlled clinical trial. United Eur. Gastroenterol. J. 2013, 1, 445–452. [Google Scholar] [CrossRef]
- A Peura, D.; Kovacs, T.O.; Metz, D.C.; Siepman, N.; Pilmer, B.L.; Talley, N.J. Lansoprazole in the treatment of functional dyspepsia: Two Double-Blind, Randomized, Placebo-Controlled trials. Am. J. Med. 2004, 116, 740–748. [Google Scholar] [CrossRef]
- Welz, A.N.; Emberger-Klein, A.; Menrad, K. Why people use herbal medicine: Insights from a focus-group study in Germany. BMC Complement. Altern. Med. 2018, 18, 92. [Google Scholar] [CrossRef] [PubMed]
- Benzie, I.F.F.; Wachtel-Galor, S. Herbal Medicine: Biomolecular and Clinical Aspects; CRC Press: Boca Raton, FL, USA, 2011. [Google Scholar]
- Mount Sinai—New York. Herbal Medicine Information; Mount Sinai Health System: New York, NY, USA, 2024. [Google Scholar]
- Alsukaibi, A.K.D.; Alenezi, K.M.; Haque, A.; Saeed, M.; Verma, M.; Ansari, I.A.; Hsieh, M.-F. Chemical, biological and in silico assessment of date (P. dactylifera L.) fruits grown in Ha’il region. Front. Chem. 2023, 11, 1138057. [Google Scholar] [CrossRef] [PubMed]
- Farnsworth, N.R. Screening Plants for New Medicines; National Academies Press: Washington, DC, USA, 1988. [Google Scholar]
- Wang, C.; Meng, Q. Global Research Trends of Herbal Medicine for Pain in Three Decades (1990–2019): A Bibliometric Analysis. J. Pain Res. 2021, 14, 1611–1626. [Google Scholar] [CrossRef] [PubMed]
- Leonti, M.; Verpoorte, R. Traditional Mediterranean and European herbal medicines. J. Ethnopharmacol. 2017, 199, 161–167. [Google Scholar] [CrossRef]
- Edward, S. Rosemary: Growing Tips, Varieties & Uses; Seasol: Boronia, Australia, 2024; Available online: https://www.seasol.com.au/tomatoes-herbs-and-vegetables/herbs/rosemary/ (accessed on 22 June 2024).
- De Guzman, C.C.; Siemonsmaeditors, J.S. (Eds.) Plant Resources of South-East Asia, No.13. Spices; Backhuys Publishers: Leiden, The Netherlands, 1999. [Google Scholar]
- Nordqvist, J. Rosemary: Health Benefits, Precautions, and Drug Interactions. The Medical News Today. 2017. Available online: https://www.medicalnewstoday.com/articles/266370 (accessed on 22 June 2024).
- Degner, S.C.; Papoutsis, A.J.; Romagnolo, D.F. Health Benefits of Traditional Culinary and Medicinal Mediterranean Plants. In Complementary and Alternative Therapies and the Aging Population; Elsevier: Amsterdam, The Netherlands, 2009; pp. 541–562. [Google Scholar] [CrossRef]
- Drugs.com, “Rosemary”. Drugs.com. 2025. Available online: https://www.drugs.com/npp/rosemary.html (accessed on 1 August 2025).
- Al-Sereiti, M.R.; Abu-Amer, K.M.; Sena, P. Pharmacology of rosemary (Rosmarinus officinalis Linn.) and its therapeutic potentials. Indian. J. Exp. Biol. 1999, 37, 124–130. [Google Scholar]
- Sintos, A.M.L.; Cabrera, H.S. Network Pharmacology Reveals Curcuma aeruginosa Roxb. Regulates MAPK and HIF-1 Pathways to Treat Androgenetic Alopecia. Biology 2024, 13, 497. [Google Scholar] [CrossRef]
- SYadav, S.; Ahamad, S.; Gupta, D.; Mathur, P. Lead optimization, pharmacophore development and scaffold design of protein kinase CK2 inhibitors as potential COVID-19 therapeutics. J. Biomol. Struct. Dyn. 2023, 41, 1811–1827. [Google Scholar] [CrossRef]
- Sharma, Y.; Velamuri, R.; Fagan, J.; Schaefer, J. Full-Spectrum Analysis of Bioactive Compounds in Rosemary (Rosmarinus officinalis L.) as Influenced by Different Extraction Methods. Molecules 2020, 25, 4599. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Stielow, M.; Witczyńska, A.; Kubryń, N.; Fijałkowski, Ł.; Nowaczyk, J.; Nowaczyk, A. The Bioavailability of Drugs—The Current State of Knowledge. Molecules 2023, 28, 8038. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019, 47, W357–W364. [Google Scholar] [CrossRef]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; A Shoemaker, B.; A Thiessen, P.; Yu, B.; et al. PubChem 2023 update. Nucleic Acids Res. 2023, 51, D1373–D1380. [Google Scholar] [CrossRef] [PubMed]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Stein, T.I.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr. Protoc. Bioinform. 2016, 54. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.J.; Lillard, J.W. A Comprehensive Review of Bioinformatics Tools for Genomic Biomarker Discovery Driving Precision Oncology. Genes 2024, 15, 1036. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Nastou, K.; Koutrouli, M.; Kirsch, R.; Mehryary, F.; Hachilif, R.; Hu, D.; E Peluso, M.; Huang, Q.; Fang, T.; et al. The STRING database in 2025: Protein networks with directionality of regulation. Nucleic Acids Res. 2024, 53, D730–D737. [Google Scholar] [CrossRef]
- Chin, C.-H.; Chen, S.-H.; Wu, H.-H.; Ho, C.-W.; Ko, M.-T.; Lin, C.-Y. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 2014, 8 (Suppl. S4), S11. [Google Scholar] [CrossRef] [PubMed]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Chen, E.Y.; Tan, C.M.; Kou, Y.; Duan, Q.; Wang, Z.; Meirelles, G.V.; Clark, N.R.; Ma’Ayan, A. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 2013, 14, 128. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Cartee, N.M.P.; Wang, M.M.; Ganesan, A. Binding of omeprazole to protein targets identified by monoclonal antibodies. PLoS ONE 2020, 15, e0239464. [Google Scholar] [CrossRef]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J. Cheminform 2011, 3, 33. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Dassault Systèmes BIOVIA. Discovery Studio Visualizer (Version 2024); Dassault Systèmes: San Diego, CA, USA, 2024. [Google Scholar]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Schrödinger, L. The PyMOL Molecular Graphics System, version 2.5; Schrödinger, LLC: New York, NY, USA, 2021. [Google Scholar]
- Python Software Foundation. Python, v5.0.0 2024.2; Python Software Foundation: Beaverton, OR, USA, 2024. [Google Scholar]
- Chen, Y.-H.; Hsieh, C.-Y.; Chiou, C.-T.; Caro, E.J.G.V.; Tayo, L.L.; Tsai, P.-W. In Vitro and In Silico Studies on the Anti-H1N1 Activity of Bioactive Compounds from Marine-Derived Streptomyces ardesiacus. Mar. Drugs 2025, 23, 149. [Google Scholar] [CrossRef] [PubMed]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- “Kolmogorov–Smirnov Test”. In The Concise Encyclopedia of Statistics; Springer: New York, NY, USA, 2008; pp. 283–287. [CrossRef]
- Meissel, K.; Yao, E.S. Using Cliff’s Delta as a Non-Parametric Effect Size Measure: An Accessible Web App and R Tutorial. Pract. Assess. Res. Eval. 2024, 29, 2. [Google Scholar]
- Varma, M.V.; Perumal, O.P.; Panchagnula, R. Functional role of P-glycoprotein in limiting peroral drug absorption: Optimizing drug delivery. Curr. Opin. Chem. Biol. 2006, 10, 367–373. [Google Scholar] [CrossRef] [PubMed]
- NCBI. KIT KIT Proto-Oncogene, Receptor Tyrosine Kinase [Homo Sapiens (Human)]; NCBI: Bethesda, MD, USA, 2025. [Google Scholar]
- Liz, M.A.; Coelho, T.; Bellotti, V.; Fernandez-Arias, M.I.; Mallaina, P.; Obici, L. A Narrative Review of the Role of Transthyretin in Health and Disease. Neurol. Ther. 2020, 9, 395–402. [Google Scholar] [CrossRef]
- Nagase, H.; Visse, R.; Murphy, G. Structure and function of matrix metalloproteinases and TIMPs. Cardiovasc. Res. 2006, 69, 562–573. [Google Scholar] [CrossRef] [PubMed]
- Rådmark, O.; Samuelsson, B. 5-Lipoxygenase: Mechanisms of regulation. J. Lipid Res. 2009, 50, S40–S45. [Google Scholar] [CrossRef]
- UniProt, “MMP2–Matrix Metalloproteinase-2–Homo Sapiens (Human),” UniProtKB, 2025. Available online: https://www.uniprot.org/uniprotkb/P08253/entry (accessed on 1 August 2025).
- Madamanchi, N.R.; Runge, M.S. Redox signaling in cardiovascular health and disease. Free Radic. Biol. Med. 2013, 61, 473–501. [Google Scholar] [CrossRef]
- Tanaka, K.J.; Song, S.; Mason, K.; Pinkett, H.W. Selective substrate uptake: The role of ATP-binding cassette (ABC) importers in pathogenesis. Biochim. Biophys. Acta (BBA) Biomembr. 2018, 1860, 868–877. [Google Scholar] [CrossRef]
- Mishra, R. Glycogen synthase kinase 3 beta: Can it be a target for oral cance”. Mol. Cancer 2010, 9, 144. [Google Scholar] [CrossRef]
- Jubin, T.; Kadam, A.; Jariwala, M.; Bhatt, S.; Sutariya, S.; Gani, A.; Gautam, S.; Begum, R. The PARP family: Insights into functional aspects of poly (ADP-ribose) polymerase-1 in cell growth and survival. Cell Prolif. 2016, 49, 421–437. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Sasaki, M.; Suzuki, T.; Nishie, H.; Kataoka, H. Combination of talaporfin photodynamic therapy and Poly (ADP-Ribose) polymerase (PARP) inhibitor in gastric cancer. Biochem. Biophys. Res. Commun. 2021, 539, 1–7. [Google Scholar] [CrossRef]
- Lund, F.E.; Cockayne, D.A.; Randall, T.D.; Solvason, N.; Schuber, F.; Howard, M.C. CD38: A new paradigm in lymphocyte activation and signal transduction. Immunol. Rev. 1998, 161, 79–93. [Google Scholar] [CrossRef]
- Stefan, S.M.; Wiese, M. Small-molecule inhibitors of multidrug resistance-associated protein 1 and related processes: A historic approach and recent advances. Med. Res. Rev. 2019, 39, 176–264. [Google Scholar] [CrossRef]
- Mayo Clinic. Functional dyspepsia—Symptoms and Causes; Mayo Clinic: Rochester, MN, USA, 2025. [Google Scholar]
- Amerikanou, C.; Kleftaki, S.-A.; Valsamidou, E.; Chroni, E.; Biagki, T.; Sigala, D.; Koutoulogenis, K.; Anapliotis, P.; Gioxari, A.; Kaliora, A.C. Food, Dietary Patterns, or Is Eating Behavior to Blame? Analyzing the Nutritional Aspects of Functional Dyspepsia. Nutrients 2023, 15, 1544. [Google Scholar] [CrossRef]
- Schmidt-Martin, D.; Quigley, E.M.M. Structural Causes of Dyspepsia. In Dyspepsia in Clinical Practice; Springer: New York, NY, USA, 2011; pp. 29–41. [Google Scholar] [CrossRef]
- Jones, R.; Crouch, S.L. Low-dose lansoprazole provides greater relief of heartburn and epigastric pain than low-dose omeprazole in patients with acid-related dyspepsia. Aliment. Pharmacol. Ther. 1999, 13, 413–419. [Google Scholar] [CrossRef]
- Nishitani, Y.; Yamamoto, K.; Yoshida, M.; Azuma, T.; Kanazawa, K.; Hashimoto, T.; Mizuno, M. Intestinal anti-inflammatory activity of luteolin: Role of the aglycone in NF-κB inactivation in macrophages co-cultured with intestinal epithelial cells. BioFactors 2013, 39, 522–533. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.; Fan, H.; Ding, J.; Han, C.; Guan, Y.; Zhu, F.; Wu, H.; Liu, Y.; Zhang, W.; Hou, X.; et al. ROS-responsive nanoparticles for oral delivery of luteolin and targeted therapy of ulcerative colitis by regulating pathological microenvironment. Mater. Today Bio 2022, 14, 100246. [Google Scholar] [CrossRef]
- Li, Y.; Shen, L.; Luo, H. Luteolin ameliorates dextran sulfate sodium-induced colitis in mice possibly through activation of the Nrf2 signaling pathway. Int. Immunopharmacol. 2016, 40, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, A.K.; Kim, J.; Lee, Y.; Balasubramanian, P.K.; Kim, Y. Isorhamnetin Has Potential for the Treatment of Escherichia coli-Induced Sepsis. Molecules 2019, 24, 3984. [Google Scholar] [CrossRef]
- Antunes-Ricardo, M.; Gutiérrez-Uribe, J.A.; Martínez-Vitela, C.; Serna-Saldívar, S.O. Topical Anti-Inflammatory Effects of Isorhamnetin Glycosides Isolated from Opuntia ficus-indica. Biomed. Res. Int. 2015, 2015, 847320. [Google Scholar] [CrossRef]
- Cheriet, T.; Ben-Bachir, B.; Thamri, O.; Seghiri, R.; Mancini, I. Isolation and Biological Properties of the Natural Flavonoids Pectolinarin and Pectolinarigenin—A Review. Antibiotics 2020, 9, 417. [Google Scholar] [CrossRef] [PubMed]
- Rahman, L.; Khalil, A.T.; Shahid, S.A.; Shinwari, Z.K.; Almarhoon, Z.M.; Alalmaie, A.; Sharifi-Rad, J.; Calina, D. Diosmin: A promising phytochemical for functional foods, nutraceuticals and cancer therapy. Food Sci. Nutr. 2024, 12, 6070–6092. [Google Scholar] [CrossRef]
- Ghisalberti, E.L. Lantana camara L. (Verbenaceae). Fitoterapia 2000, 71, 467–486. [Google Scholar] [CrossRef]
- El Menyiy, N.; Aboulaghras, S.; Bakrim, S.; Moubachir, R.; Taha, D.; Khalid, A.; Abdalla, A.N.; Algarni, A.S.; Hermansyah, A.; Ming, L.C.; et al. Genkwanin: An emerging natural compound with multifaceted pharmacological effects. Biomed. Pharmacother. 2023, 165, 115159. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; He, Y.; Hu, F.; Li, M.; Yao, Y. Genkwanin Alleviates Mitochondrial Dysfunction and Oxidative Stress in a Murine Model of Experimental Colitis: The Participation of Sirt1. Ann. Clin. Lab. Sci. 2022, 52, 301–313. [Google Scholar]
- Energy and Metabolism—Metabolic Pathways. In Biology LibreTexts; University of California: Davis, CA, USA, 2018.
- Zhang, S.; Xu, Z.; Cao, X.; Xie, Y.; Lin, L.; Zhang, X.; Zou, B.; Liu, D.; Cai, Y.; Liao, Q.; et al. Shenling Baizhu San improves functional dyspepsia in rats as revealed by 1 H-NMR based metabolomics. Anal. Methods 2020, 12, 2363–2375. [Google Scholar] [CrossRef]
- Zhou, L.; Zeng, Y.; Zhang, H.; Ma, Y. The Role of Gastrointestinal Microbiota in Functional Dyspepsia: A Review. Front. Physiol. 2022, 13, 910568. [Google Scholar] [CrossRef]
- A Liu, K.; Lashinger, L.M.; Rasmussen, A.J.; Hursting, S.D. Leucine supplementation differentially enhances pancreatic cancer growth in lean and overweight mice. Cancer Metab. 2014, 2, 6. [Google Scholar] [CrossRef]
- Monin, L.; Gaffen, S.L. Interleukin 17 Family Cytokines: Signaling Mechanisms, Biological Activities, and Therapeutic Implications. Cold Spring Harb. Perspect. Biol. 2018, 10, a028522. [Google Scholar] [CrossRef] [PubMed]
- Dewayani, A.; Fauzia, K.A.; Alfaray, R.I.; Waskito, L.A.; Doohan, D.; Rezkitha, Y.A.A.; Abdurachman, A.; Kobayashi, T.; I’tishom, R.; Yamaoka, Y.; et al. The Roles of IL-17, IL-21, and IL-23 in the Helicobacter pylori Infection and Gastrointestinal Inflammation: A Review. Toxins 2021, 13, 315. [Google Scholar] [CrossRef] [PubMed]
- Arachchi, P.S.; Fernando, N.; Weerasekera, M.M.; Senevirathna, B.; Weerasekera, D.D.; Gunasekara, C.P. Proinflammatory Cytokine IL-17 Shows a Significant Association with Helicobacter pylori Infection and Disease Severity. Gastroenterol. Res. Pract. 2017, 2017, 6265150. [Google Scholar] [CrossRef] [PubMed]
- Winarta, J.; Waleleng, B.J.; Wenas, N.T.; Fujiyanto; Miguna, O.; Rahardja, M. Correlation between Interleukin-17, High Sensitivity C-Reactive Protein and Pepsinogen in Helicobacter pylori Infected Gastritis. Gastroenterol. Insights 2024, 15, 32–41. [Google Scholar] [CrossRef]
- Buchan, I. P-Values England: StatsDirect. 2024. Available online: https://www.statsdirect.com/help/basics/p_values.htm (accessed on 1 July 2024).
- Khurana, V.; Singh, T. Gastric mucosal fibrosis: A novel explanation for dyspepsia. Med. Hypotheses 2003, 61, 513–516. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Gu, S.; Qin, B.; Granito, A. Overexpression of microRNA-211 in Functional Dyspepsia via Downregulation of the Glial Cell Line-Derived Neurotrophic Factor (GDNF) by Increasing Phosphorylation of p38 MAPK Pathway. Can. J. Gastroenterol. Hepatol. 2022, 2022, 9394381. [Google Scholar] [CrossRef]
- Malespín-Bendaña, W.; Alpízar-Alpízar, W.; Figueroa-Protti, L.; Reyes, L.; Molina-Castro, S.; Une, C.; Ramírez-Mayorga, V. Helicobacter pylori infection induces gastric precancerous lesions and persistent expression of Angpt2, Vegf-A and Tnf-A in a mouse model. Front. Oncol. 2023, 13, 1072802. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).