Markov Chain-Based Stochastic Modelling of HIV-1 Life Cycle in a CD4 T Cell
Abstract
1. Introduction
- Cell-to-cell variability in HIV progeny production;
- Multiplicity of single-cell infection;
- Global sensitivity of specific reaction steps on net virus production.
2. Results
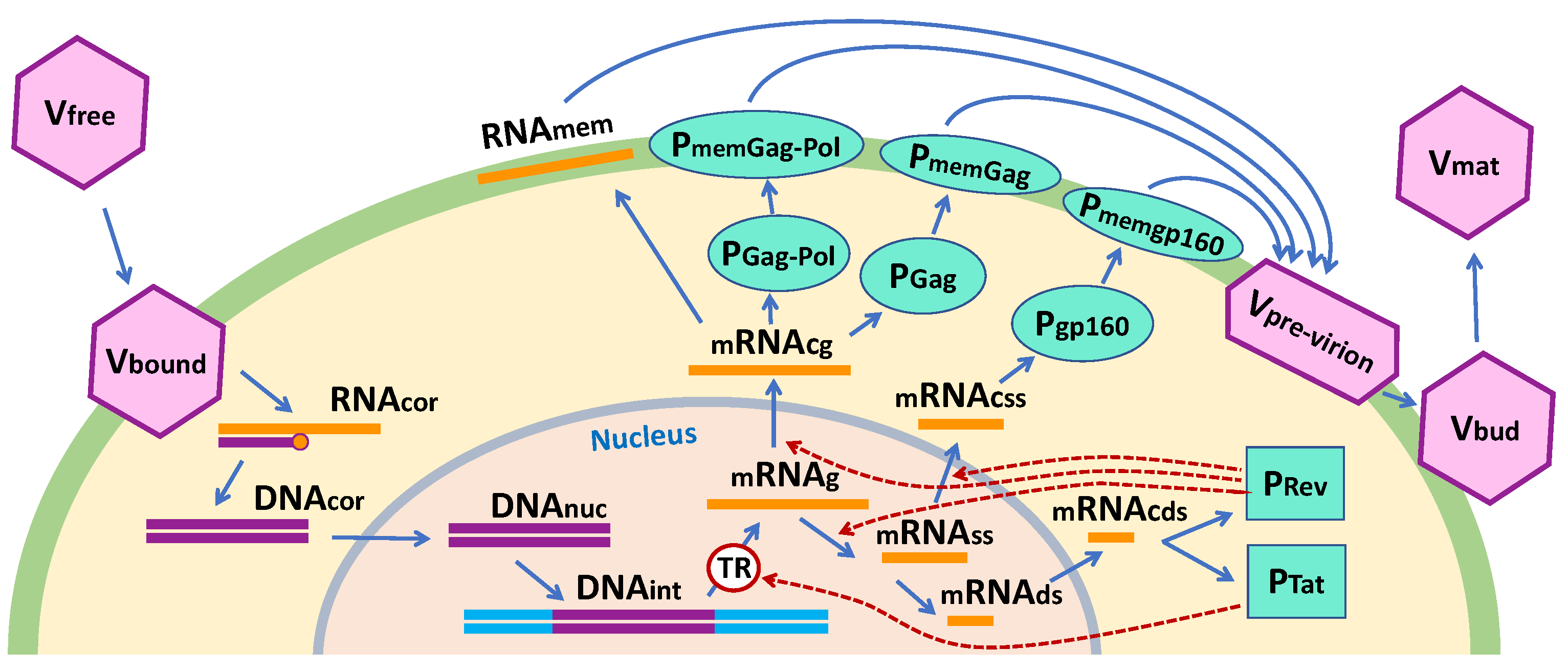
2.1. Governing Deterministic Equations
2.1.1. Virus Entry
- Virion binding to CD4 receptors (the viral glycoprotein gp120 binds to CD4 receptors on the T cell surface);
- Binding to the co-receptor CCR5 or CXCR4;
- Virus membrane and cell membrane fusion, i.e., the nucleocapsid is uncoated and the viral RNA is injected into the cell.
- is the number of free virions outside the cell;
- is the number of virions bound to CD4 and the co-receptor.
- h; h;
- h; h;
- represent the rate of virion binding to the CD4+ T cell membrane, the rate of virion fusion with the cell, the clearance rate of free mature virions, and the degradation rate of bound virions, respectively. Their reference values and admissible ranges are specified in [15].
2.1.2. Reverse Transcription
- Synthesis of minus-strand DNA from viral RNA;
- Synthesis of plus-strand DNA;
- Double-strand DNA formation.
- is the number of genomic RNA molecules in the cytoplasm;
- is the number of proviral DNA molecules synthesized by reverse transcription.
- h; h;
- h; h; represent the reverse transcription rate, the transport rate of DNA from cytoplasm to nucleus, the degradation rate of RNA in the cytoplasm and the degradation rate of DNA in the cytoplasm, respectively. Their reference values and admissible ranges are specified in [15].
2.1.3. Integration
- is the number of DNA molecules in the nucleus;
- is the number of integrated DNA.
- h; h; h; represent the integration rate, the degradation rate of DNA in the nucleus and the degradation rate of DNA integrated into the chromosome, respectively. Their reference values and admissible ranges are specified in [15].
2.1.4. Transcription
- is the number of HIV mRNA molecules in the nucleus: g for genomic or full-length;
- is the number of HIV singly spliced (ss) mRNA molecules in the nucleus;
- is the number of HIV doubly spliced (ds) mRNA molecules in the nucleus;
- is the number of HIV mRNA molecules in the cytoplasm: g for genomic or full-length;
- is the number of HIV singly spliced (ss) mRNA molecules in the cytoplasm;
- is the number of HIV doubly spliced (ds) mRNA molecules in the cytoplasm.
- ; ;
- ;
- .
- h; h;
- ; ×; ;
- h; h; h;
- h; h; h;
- h; represent the cell intrinsic rate of basal transcription, the level of transcription induced by Tat transactivation, the inhibitory effect of Rev on the splicing rates implying their -fold reduction at the saturation level of Rev, the rate of splicing for full-length virus RNA, the rate of export from the nucleus, the rate of export from the nucleus, the rate of splicing for singly spliced virus RNA, the rate of export from the nucleus, the transport rate of to the cell membrane, and the degradation rates of , , respectively. Their reference values and admissible ranges are specified in [15].
2.1.5. Translation
- is the number of protein molecules: Gag-Pol;
- is the number of protein molecules: Gag;
- is the number of protein molecules: gp160;
- is the number of protein molecules: Tat;
- is the number of protein molecules: Rev.
- h;
- ; ; ; ; ;
- h; h; h;
- h; h; h;
- h; h; represent the rate of mRNA to proteins translation, stand for the fraction of coding , , , Gag, gp160, Tat, . Following them, the parameters define the rates of protein transport to membrane, , and the degradation rates of proteins Gag-Pol, Gag, gp160, Tat and Rev, respectively. Their reference values and admissible ranges are specified in [15].
2.1.6. Assembly, Budding and Maturation
- h; h; h; h;
- h; ; ; ; ; represent the rates of RNA and protein transport to the membrane, , Gag, , the incorporation rate of molecules into pre-virion complexes, the number of viral RNA transcripts in a new virion, the number of Gag-Pol molecules in a new virion, the number of Gag molecules in a new virion, and the number of gp160 molecules in a new virion, respectively. Their reference values and admissible ranges are specified in [15].
- h; h;
- h; h; represent the degradation rates of , the membrane-anchored protein Gag-Pol, the membrane-anchored protein Gag, and the membrane-associated gp160, respectively. Their reference values and admissible ranges are specified in [15].
- is the number of virions on the membrane;
- is the number of free viruses after budding from the cell;
- is the number of mature virions outside the cell.
- h; h; h;
- h; h; h; represent the incorporation rate of molecules into pre-virion complexes, the budding rate of new virions, the maturation rate, the degradation rate of the assembled pre-virion complex, the degradation rate for budded immature viral-like particles, and the clearance rate of free mature virions, respectively. Their reference values and admissible ranges are specified in [15].
2.2. Stochastic Markov Chain Modelling
2.2.1. Algorithm
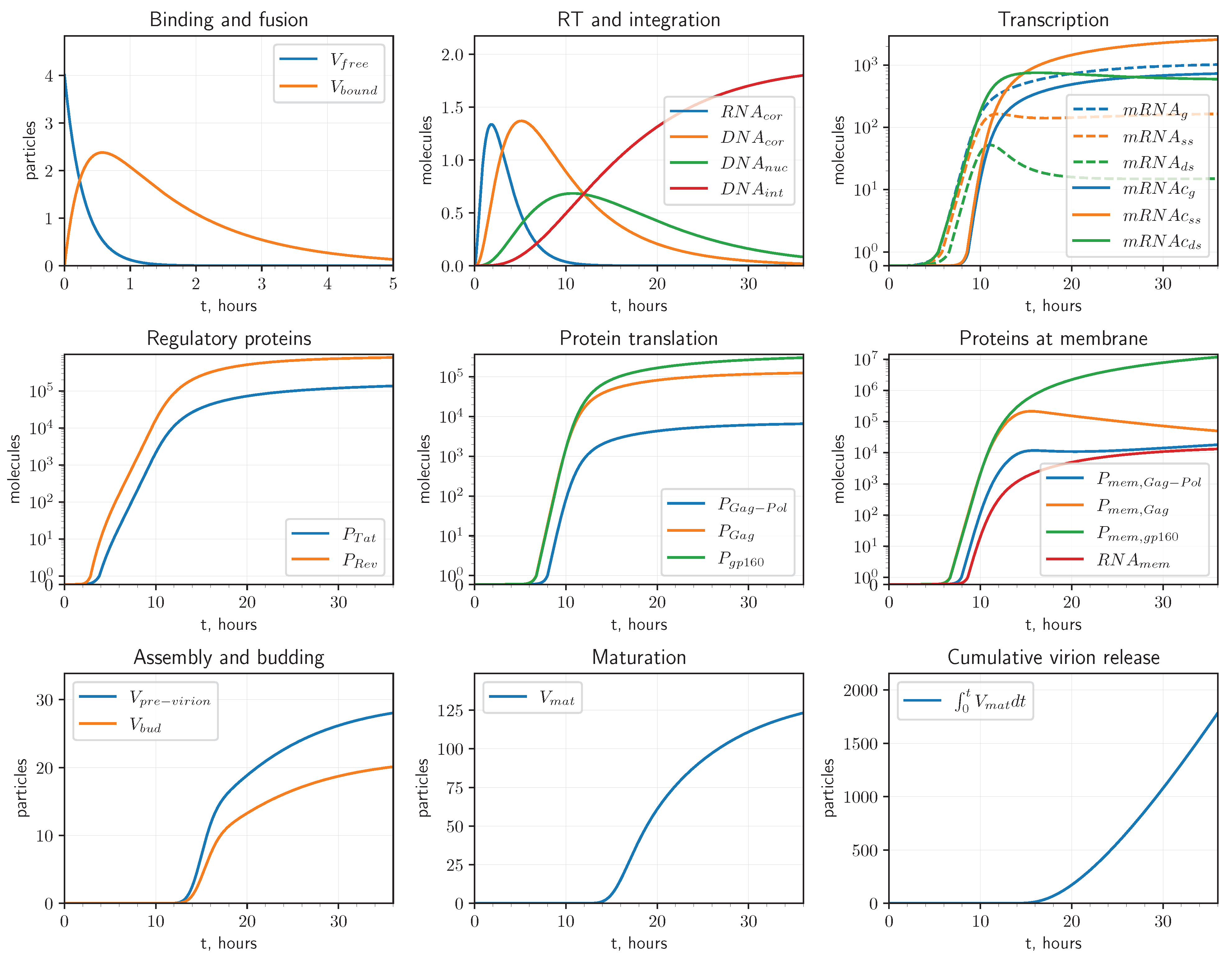
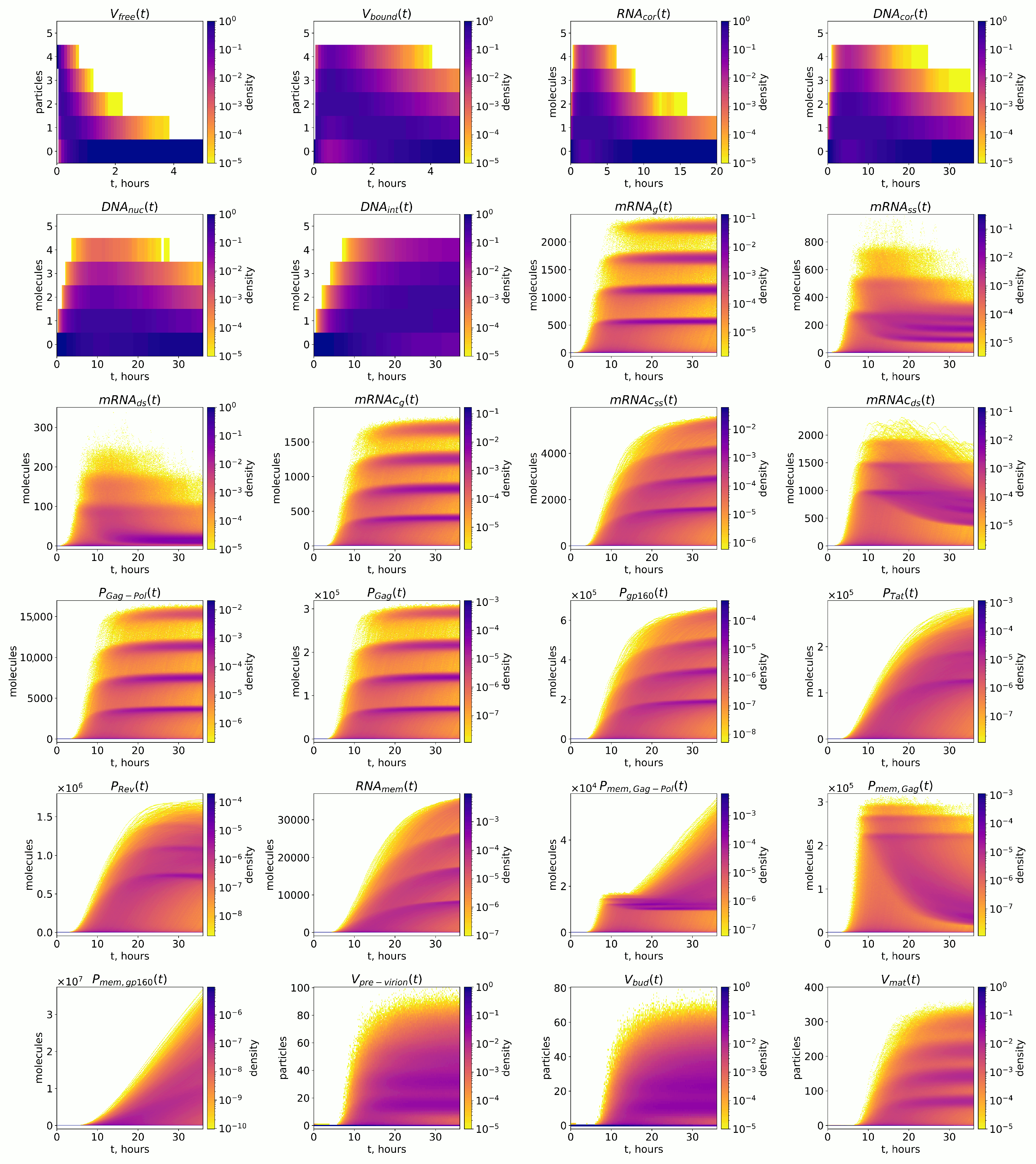
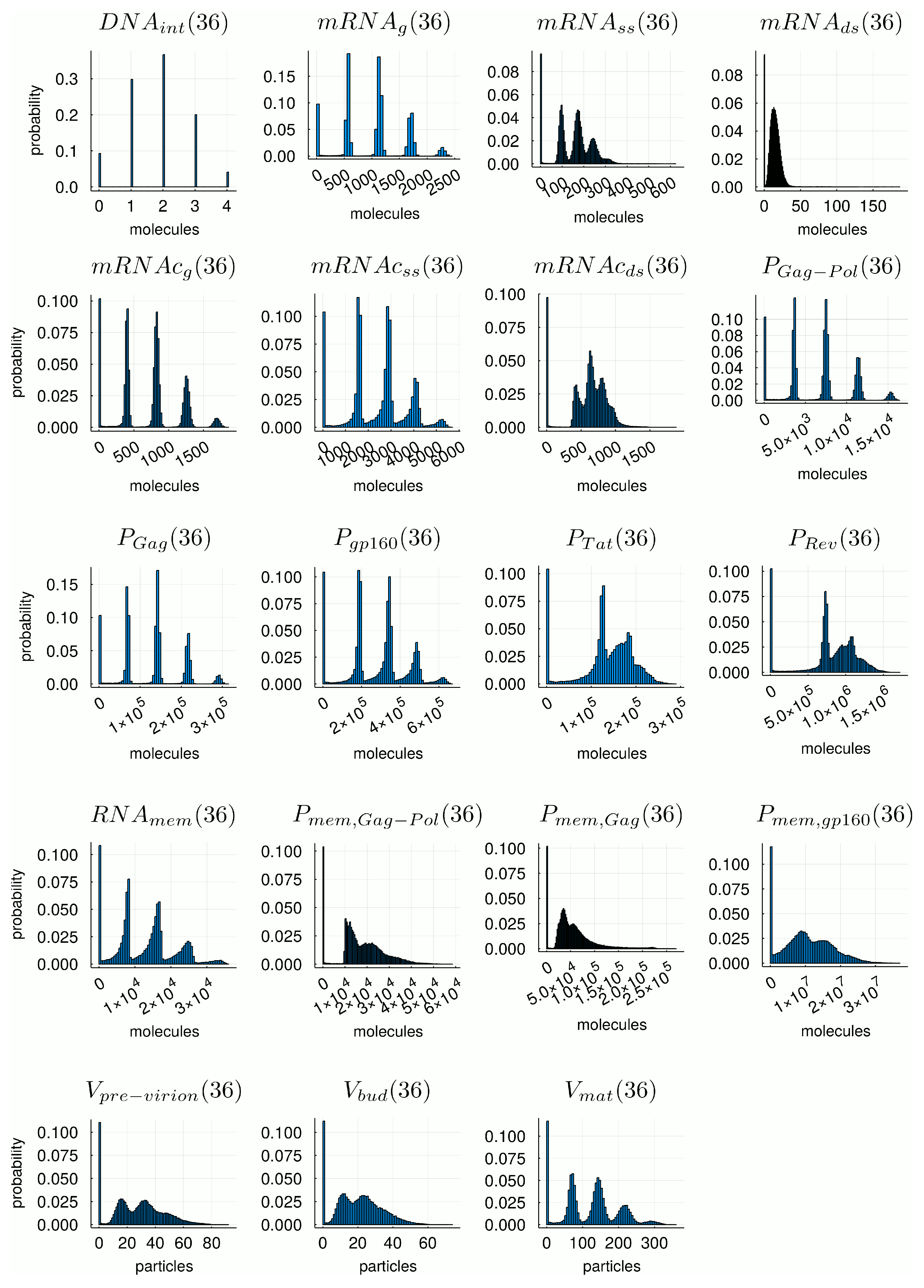
2.2.2. Stochastic Modelling Results
2.3. Sensitivity Analysis
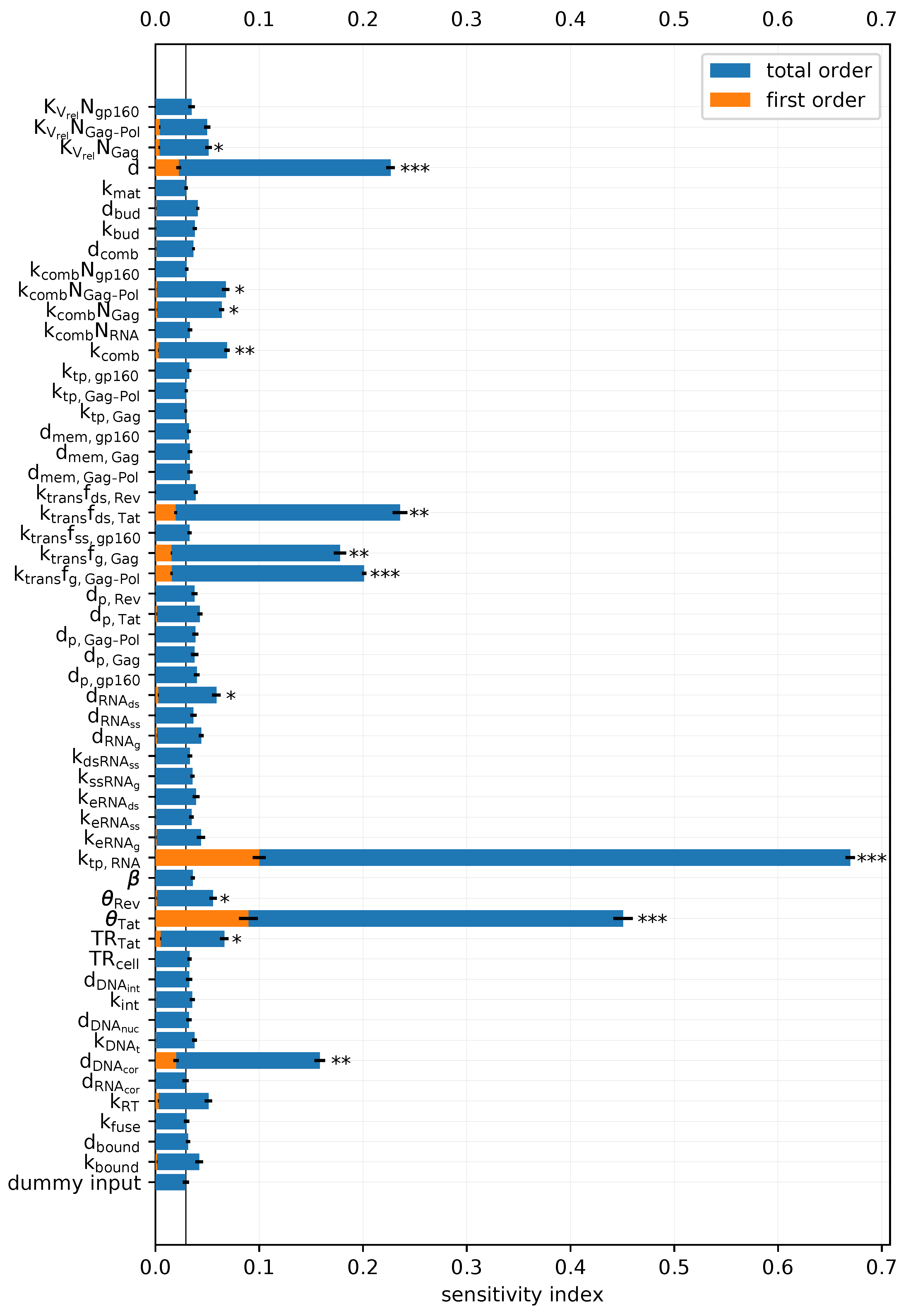
2.3.1. Sensitivity Analysis of the Deterministic Model towards the Cumulative Virion Release
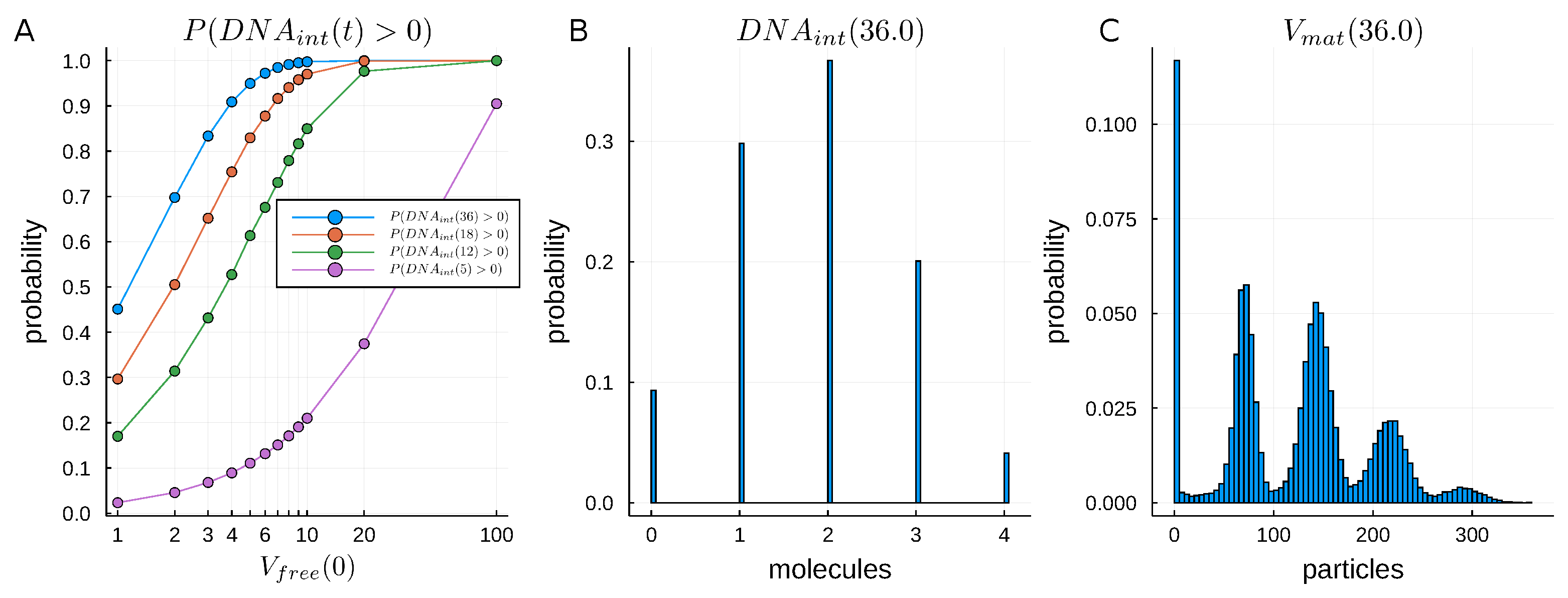
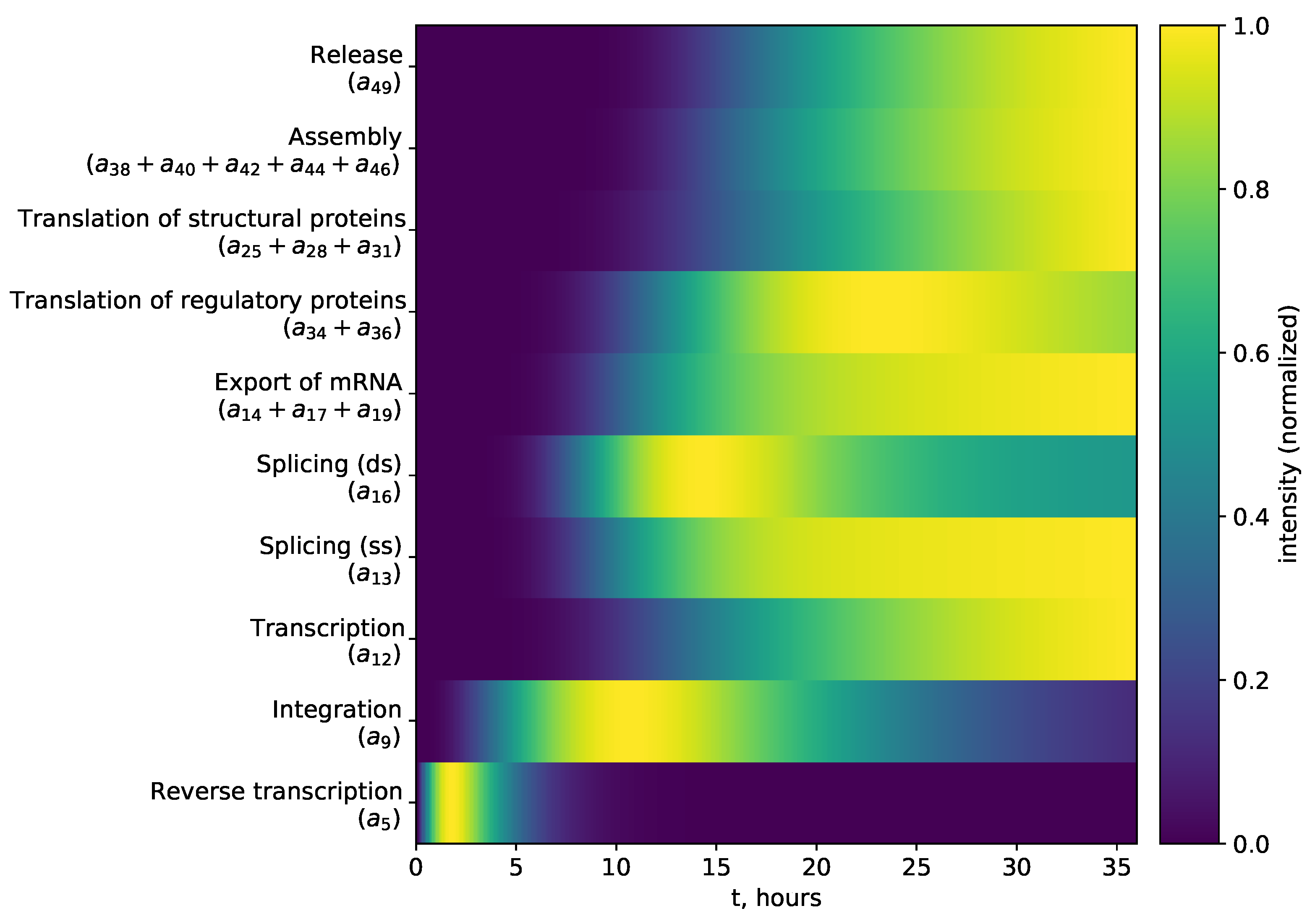
2.3.2. Sensitivity Analysis of Early Stages of Cell Infection Using the Stochastic Model
3. Discussion
- Transport of genomic mRNA to membranes;
- Tolerance of transcription activation to Tat-mediated regulation;
- Degradation of free and mature virions.
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| RT | Reverse Transcription |
| probability density function | |
| cdf | cumulative distribution function |
References
- Gao, Y.; McKay, P.; Mann, J. Advances in HIV-1 vaccine development. Viruses 2018, 10, 167. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Omer, S.B. Why and how vaccines work. Cell 2020, 183, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Gurdasani, D.; Iles, L.; Dillon, D.G.; Young, E.H.; Olson, A.D.; Naranbhai, V.; Fidler, S.; Gkrania-Klotsas, E.; Post, F.A.; Kellam, P.; et al. A systematic review of definitions of extreme phenotypes of HIV control and progression. AIDS 2014, 28, 149–162. [Google Scholar] [CrossRef]
- Siliciano, R.F.; Greene, W.C. HIV latency. Cold Spring Harb. Perspect. Med. 2011, 1, a007096. [Google Scholar] [CrossRef] [PubMed]
- Hahn, B.H.; Shaw, G.M.; Taylor, M.E.; Redfield, R.R.; Markham, P.D.; Salahuddin, S.Z.; Wong-Staal, F.; Gallo, R.C.; Parks, E.S.; Parks, W.P. Genetic variation in HTLV-III/LAV over time in patients with AIDS or at risk for AIDS. Science 1986, 232, 1548–1553. [Google Scholar] [CrossRef] [PubMed]
- Meyerhans, A.; Cheynier, R.; Albert, J.; Seth, M.; Kwok, S.; Sninsky, J.; Morfeldt-Månson, L.; Asjö, B.; Wain-Hobson, S. Temporal fluctuations in HIV quasispecies in vivo are not reflected by sequential HIV isolations. Cell 1989, 58, 901–910. [Google Scholar] [CrossRef]
- Zanini, F.; Brodin, J.; Thebo, L.; Lanz, C.; Bratt, G.; Albert, J.; Neher, R.A. Population genomics of intrapatient HIV-1 evolution. eLife 2015, 4, e11282. [Google Scholar] [CrossRef]
- Josefsson, L.; King, M.S.; Makitalo, B.; Brännström, J.; Shao, W.; Maldarelli, F.; Kearney, M.F.; Hu, W.S.; Chen, J.; Gaines, H.; et al. Majority of CD4+ T cells from peripheral blood of HIV-1–infected individuals contain only one HIV DNA molecule. Proc. Natl. Acad. Sci. USA 2011, 108, 11199–11204. [Google Scholar] [CrossRef]
- Jung, A.; Maier, R.; Vartanian, J.P.; Bocharov, G.; Jung, V.; Fischer, U.; Meese, E.; Wain-Hobson, S.; Meyerhans, A. Multiply infected spleen cells in HIV patients. Nature 2002, 418, 144. [Google Scholar] [CrossRef]
- Schultz, A.; Sopper, S.; Sauermann, U.; Meyerhans, A.; Suspène, R. Stable multi-infection of splenocytes during SIV infection—The basis for continuous recombination. Retrovirology 2012, 9, 31. [Google Scholar] [CrossRef]
- Ito, Y.; Remion, A.; Tauzin, A.; Ejima, K.; Nakaoka, S.; Iwasa, Y.; Iwami, S.; Mammano, F. Number of infection events per cell during HIV-1 cell-free infection. Sci. Rep. 2017, 7, 6559. [Google Scholar] [CrossRef]
- Andreu-Moreno, I.; Bou, J.V.; Sanjuán, R. Cooperative nature of viral replication. Sci. Adv. 2020, 6, eabd4942. [Google Scholar] [CrossRef] [PubMed]
- Bocharov, G.; Chereshnev, V.; Gainova, I.; Bazhan, S.; Bachmetyev, B.; Argilaguet, J.; Martinez, J.; Meyerhans, A. Human immunodeficiency virus infection: From biological observations to mechanistic mathematical modelling. Math. Model. Nat. Phenom. 2012, 7, 78–104. [Google Scholar] [CrossRef]
- Bouchnita, A.; Bocharov, G.; Meyerhans, A.; Volpert, V. Towards a multiscale model of acute HIV infection. Computation 2017, 5, 6. [Google Scholar] [CrossRef]
- Shcherbatova, O.; Grebennikov, D.; Sazonov, I.; Meyerhans, A.; Bocharov, G. Modeling of the HIV-1 life cycle in productively infected cells to predict novel therapeutic targets. Pathogens 2020, 9, 255. [Google Scholar] [CrossRef] [PubMed]
- Weinberger, L.S.; Burnett, J.C.; Toettcher, J.E.; Arkin, A.P.; Schaffer, D.V. Stochastic gene expression in a Lentiviral positive-feedback Loop: HIV-1 Tat fluctuations drive phenotypic diversity. Cell 2005, 122, 169–182. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, D.T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 1977, 81, 2340–2361. [Google Scholar] [CrossRef]
- Hu, W.S.; Hughes, S. HIV-1 reverse transcription. Cold Spring Harb. Perspect. Med. 2012, 2, a006882. [Google Scholar] [CrossRef]
- Craigie, R.; Bushman, F. HIV DNA integration. Cold Spring Harb. Perspect. Med. 2012, 2, a006890. [Google Scholar] [CrossRef]
- Chereshnev, V.; Bocharov, G.; Bazhan, S.; Bachmetyev, B.; Gainova, I.; Likhoshvai, V.; Argilaguet, J.; Martinez, J.; Rump, J.; Mothe, B.; et al. Pathogenesis and treatment of HIV infection: The cellular, the immune system and the neuroendocrine systems perspective. Int. Rev. Immunol. 2013, 32, 1–25. [Google Scholar] [CrossRef]
- Kim, H.; Yin, J. Robust growth of human immunodeficiency virus type 1 (HIV-1). Biophys. J. 2005, 89, 2210–2221. [Google Scholar] [CrossRef] [PubMed]
- Likhoshvai, V.; Khlebodarova, T.; Bazhan, S.; Gainova, I.; Chereshnev, V.; Bocharov, G. Mathematical model of the Tat-Rev regulation of HIV-1 replication in an activated cell predicts the existence of oscillatory dynamics in the synthesis of viral components. BMC Genom. 2014, 15 (Suppl. 12), S1. [Google Scholar] [CrossRef] [PubMed]
- Reddy, B.; Yin, J. Quantitative intracellular kinetics of HIV type 1. AIDS Res. Hum. Retroviruses 1999, 15, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Freed, E. HIV-1 assembly, release and maturation. Nat. Rev. Microbiol. 2015, 13, 484–496. [Google Scholar] [CrossRef]
- Heldt, F.S.; Frensing, T.; Reichl, U. Modeling the intracellular dynamics of influenza virus replication to understand the control of viral RNA synthesis. J. Virol. 2012, 86, 7806–7817. [Google Scholar] [CrossRef] [PubMed]
- Heldt, F.S.; Kupke, S.Y.; Dorl, S.; Reichl, U.; Frensing, T. Single-cell analysis and stochastic modelling unveil large cell-to-cell variability in influenza A virus infection. Nat. Commun. 2015, 6, 8938. [Google Scholar] [CrossRef]
- Gibson, M.A.; Bruck, J. Efficient exact stochastic simulation of chemical systems with many species and many channels. J. Phys. Chem. A 2000, 104, 1876–1889. [Google Scholar] [CrossRef]
- Marchetti, L.; Priami, C.; Thanh, V.H. Simulation Algorithms for Computational Systems Biology; Texts in Theoretical Computer Science. An EATCS Series; Springer: Berlin/Heidelberg, Germany, 2017. [Google Scholar] [CrossRef]
- Sazonov, I.; Kelbert, M.; Gravenor, M.B. A two-stage model for the SIR outbreak: Accounting for the discrete and stochastic nature of the epidemic at the initial contamination stage. Math. Biosci. 2011, 234, 108–117. [Google Scholar] [CrossRef]
- Safta, C.; Sargsyan, K.; Debusschere, B.; Najm, H.N. Hybrid discrete/continuum algorithms for stochastic reaction networks. J. Comput. Phys. 2015, 281, 177–198. [Google Scholar] [CrossRef]
- Sazonov, I.; Grebennikov, D.; Kelbert, M.; Bocharov, G. Modelling stochastic and deterministic behaviours in virus infection dynamics. Math. Model. Nat. Phenom. 2017, 12, 63–77. [Google Scholar] [CrossRef][Green Version]
- Sazonov, I.; Grebennikov, D.; Kelbert, M.; Meyerhans, A.; Bocharov, G. Viral infection dynamics model based on a Markov process with time delay between cell infection and progeny production. Mathematics 2020, 8, 1207. [Google Scholar] [CrossRef]
- Mohammadi, P.; Desfarges, S.; Bartha, I.; Joos, B.; Zangger, N.; Muñoz, M.; Günthard, H.F.; Beerenwinkel, N.; Telenti, A.; Ciuffi, A. 24 h in the life of HIV-1 in a T cell line. PLoS Pathog. 2013, 9, e1003161. [Google Scholar] [CrossRef]
- Sobol, I.M. Sensitivity analysis for non-linear mathematical models. Math. Model. Comput. Exp. 1993, 1, 407–414. [Google Scholar]
- Sobol, I. Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Math. Comput. Simul. 2001, 55, 271–280. [Google Scholar] [CrossRef]
- Saltelli, A.; Tarantola, S.; Chan, K.P.S. A quantitative model-independent method for global sensitivity analysis of model output. Technometrics 1999, 41, 39–56. [Google Scholar] [CrossRef]
- Marino, S.; Hogue, I.B.; Ray, C.J.; Kirschner, D.E. A methodology for performing global uncertainty and sensitivity analysis in systems biology. J. Theor. Biol. 2008, 254, 178–196. [Google Scholar] [CrossRef] [PubMed]
- Degasperi, A.; Gilmore, S. Sensitivity analysis of stochastic models of bistable biochemical reactions. In Formal Methods for Computational Systems Biology; Lecture Notes in Computer Science; Bernardo, M., Degano, P., Zavattaro, G., Eds.; Springer: Berlin/Heidelberg, Germany, 2008; pp. 1–20. [Google Scholar] [CrossRef]
- Morris, M.D. Factorial sampling plans for preliminary computational experiments. Technometrics 1991, 33, 161–174. [Google Scholar] [CrossRef]
- Saltelli, A.; Ratto, M.; Andres, T.; Campolongo, F.; Cariboni, J.; Gatelli, D.; Saisana, M.; Tarantola, S. Global Sensitivity Analysis. The Primer; Google-Books-ID: WAssmt2vumgC; John Wiley & Sons: Hoboken, NJ, USA, 2008. [Google Scholar] [CrossRef]
- Cheng, Z.; Hoffmann, A. A stochastic spatio-temporal (SST) model to study cell-to-cell variability in HIV-1 infection. J. Theor. Biol. 2016, 395, 87. [Google Scholar] [CrossRef] [PubMed]
| m | Transition | Propensity, | m | Transition | Propensity, |
|---|---|---|---|---|---|
| 1 | 27 | ||||
| 2 | 28 | ||||
| 3 | 29 | ||||
| 4 | 30 | ||||
| 5 | 31 | ||||
| 6 | 32 | ||||
| 7 | 33 | ||||
| 8 | 34 | ||||
| 9 | 35 | ||||
| 10 | 36 | ||||
| 11 | 37 | ||||
| 12 | 38 | ||||
| 13 | 39 | ||||
| 14 | 40 | ||||
| 15 | 41 | ||||
| 16 | 42 | ||||
| 17 | 43 | ||||
| 18 | 44 | ||||
| 19 | 45 | ||||
| 20 | 46 | ||||
| 21 | 47 | ||||
| 22 | 48 | ||||
| 23 | 49 | ||||
| 24 | 50 | ||||
| 25 | 51 | ||||
| 26 |
| Parameter | Description | Sensitivity Indices | |
|---|---|---|---|
| First Order | Total Order | ||
| Transport of genomic mRNA to membrane | 0.099 ± 0.006 | 0.669 ± 0.004 | |
| Tolerance of transcription activation to Tat-mediated regulation | 0.089 ± 0.009 | 0.450 ± 0.009 | |
| Translation of Tat molecules | 0.019 ± 0.001 | 0.235 ± 0.001 | |
| d | Degradation of free and mature virions | 0.022 ± 0.002 | 0.226 ± 0.004 |
| Translation of Gag-Pol molecules | 0.0155 ± 0.0009 | 0.201 ± 0.002 | |
| Translation of Gag molecules | 0.0153 ± 0.0006 | 0.177 ± 0.006 | |
| Degradation of DNA during RT | 0.019 ± 0.002 | 0.158 ± 0.005 | |
| Assembly of pre-virion complexes | 0.0028 ± 0.0001 | 0.068 ± 0.002 | |
| Tat-induced transcription rate | 0.0049 ± 0.0004 | 0.066 ± 0.003 | |
| Gag contribution to pre-virion assembly | 0.0014 ± 0.0001 | 0.063 ± 0.002 | |
| Gag-Pol contribution to pre-virion assembly | 0.00125 ± 0.00005 | 0.063 ± 0.002 | |
| Degradation of doubly-spliced mRNA | 0.0027 ± 0.0003 | 0.058 ± 0.004 | |
| Tolerance of pre-virion assembly to Gag availability on membrane | 0.0036 ± 0.0006 | 0.051 ± 0.003 | |
| Parameter | Description | Sensitivity |
|---|---|---|
| Binding rate of free virions to CD4+ T cell membrane | 0.498 | |
| d | Degradation rate of HIV particles | 0.752 |
| Rate of virion fusion into the cell | 0.988 | |
| Degradation rate of virions bound to membrane | 1.080 | |
| Rate of reverse transcription | 0.804 | |
| Degradation rate of viral RNA in cytoplasm | 0.712 | |
| Rate of viral DNA transfer to nucleus | 0.738 | |
| Degradation rate of viral DNA in cytoplasm | 0.464 | |
| Rate of viral DNA integration into host chromosome | 0.446 | |
| Degradation rate of viral DNA in nucleus | 0.346 | |
| Degradation rate of DNA integrated into chromosome | 0.582 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sazonov, I.; Grebennikov, D.; Meyerhans, A.; Bocharov, G. Markov Chain-Based Stochastic Modelling of HIV-1 Life Cycle in a CD4 T Cell. Mathematics 2021, 9, 2025. https://doi.org/10.3390/math9172025
Sazonov I, Grebennikov D, Meyerhans A, Bocharov G. Markov Chain-Based Stochastic Modelling of HIV-1 Life Cycle in a CD4 T Cell. Mathematics. 2021; 9(17):2025. https://doi.org/10.3390/math9172025
Chicago/Turabian StyleSazonov, Igor, Dmitry Grebennikov, Andreas Meyerhans, and Gennady Bocharov. 2021. "Markov Chain-Based Stochastic Modelling of HIV-1 Life Cycle in a CD4 T Cell" Mathematics 9, no. 17: 2025. https://doi.org/10.3390/math9172025
APA StyleSazonov, I., Grebennikov, D., Meyerhans, A., & Bocharov, G. (2021). Markov Chain-Based Stochastic Modelling of HIV-1 Life Cycle in a CD4 T Cell. Mathematics, 9(17), 2025. https://doi.org/10.3390/math9172025







