Effects of Exploration Weight and Overtuned Kernel Parameters on Gaussian Process-Based Bayesian Optimization Search Performance
Abstract
1. Introduction
2. Gaussian Process-Based Bayesian Optimization
2.1. Surrogate Model
2.2. Tuning Kernel Parameters
2.3. Optimization Algorithm for Experiments
| Algorithm 1 Verification-targeted optimization algorithm |
Input: Initial observation dataset , maximum number of BO iterations , maximum number of GD iterations , GD learning rate , exploration weight , black-box function , observation noise , search space , initial KP Output: Solution and its value ................................................................................................................................................
................................................................................................................................................ Notes: · Lines 2–4: Tuning the KPs via GD; · Lines 6–8: Decisions regarding the next search point and observation; · Lines 10–11: Obtain an approximate solution. “GD”: gradient descent, “BO”: Bayesian optimization |
2.4. Indices
3. Experiments
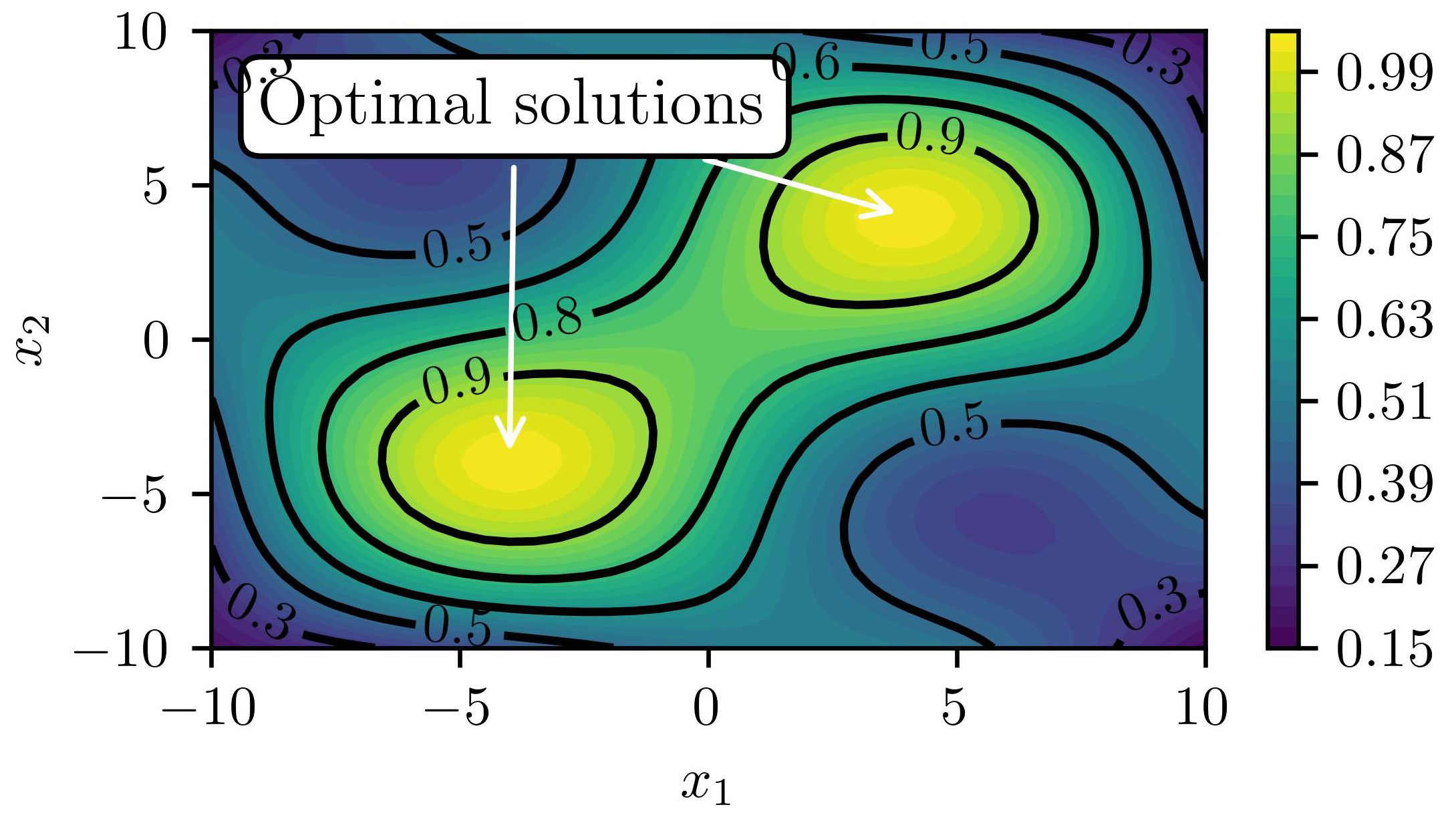
3.1. Objective and Outline
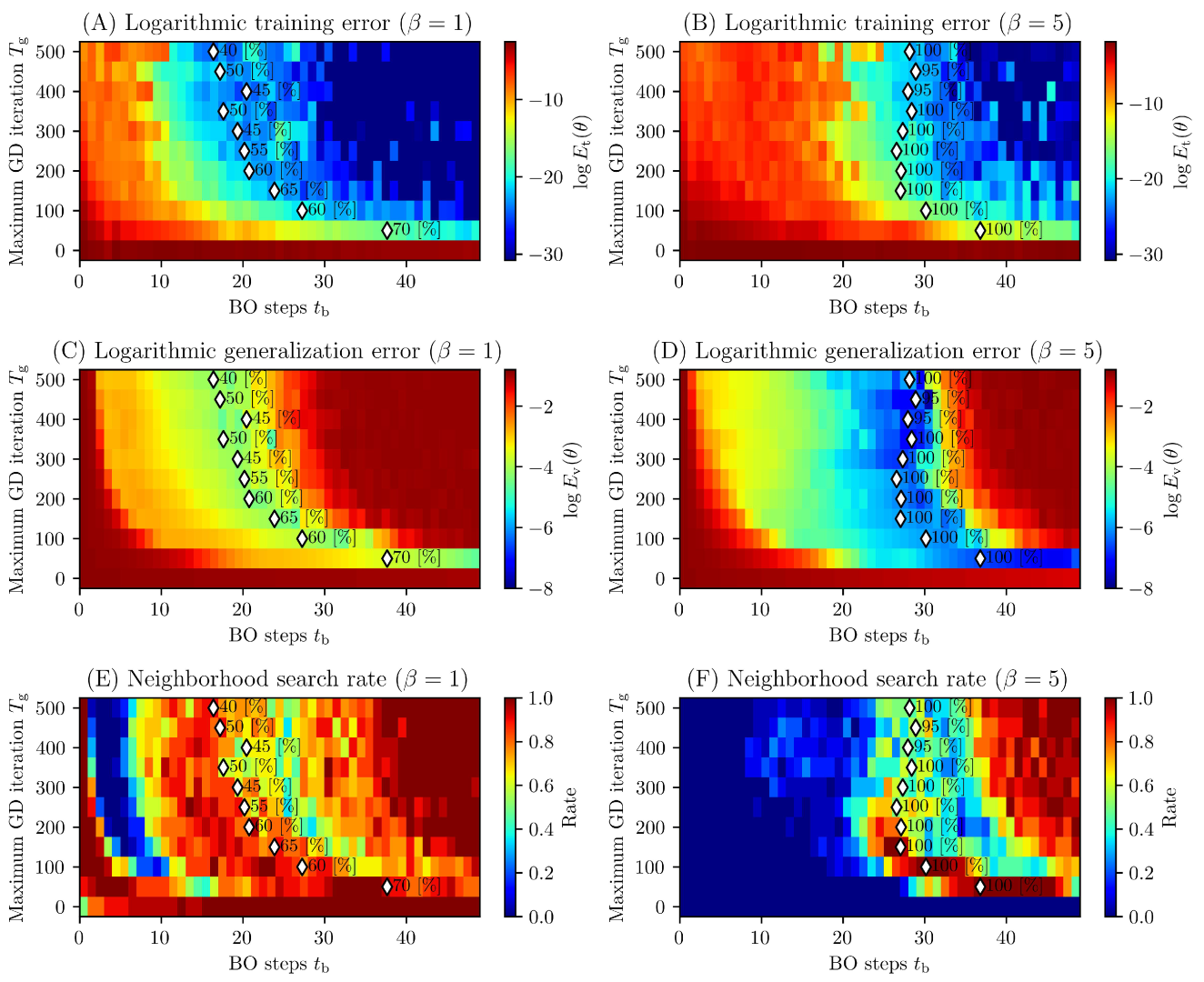
3.2. Results and discussions
4. Conclusions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Saleh, E.; Tarawneh, A.; Naser, M.Z.; Abedi, M.; Almasabha, G. You only design once (YODO): Gaussian Process-Batch Bayesian optimization framework for mixture design of ultra high performance concrete. Constr. Build. Mater. 2022, 330, 127270. [Google Scholar] [CrossRef]
- Mathern, A.; Steinholtz, O.S.; Sjöberg, A.; Önnheim, M.; Ek, K.; Rempling, R.; Gustavsson, E.; Jirstrand, M. Multi-objective constrained Bayesian optimization for structural design. Struct. Multidiscip. Optim. 2021, 63, 689–701. [Google Scholar] [CrossRef]
- Frazier, P.I.; Wang, J. Bayesian optimization for materials design. Springer Ser. Mater. Sci. 2015, 225, 45–75. [Google Scholar] [CrossRef]
- Ohno, H. Empirical studies of Gaussian process based Bayesian optimization using evolutionary computation for materials informatics. Expert Syst. Appl. 2018, 96, 25–48. [Google Scholar] [CrossRef]
- Ueno, T.; Rhone, T.D.; Hou, Z.; Mizoguchi, T.; Tsuda, K. COMBO: An efficient Bayesian optimization library for materials science. Mater. Discov. 2016, 4, 18–21. [Google Scholar] [CrossRef]
- Elsayad, A.M.; Nassef, A.M.; Al-Dhaifallah, M. Bayesian optimization of multiclass SVM for efficient diagnosis of erythemato-squamous diseases. Biomed. Signal Process. Control 2022, 71, 103223. [Google Scholar] [CrossRef]
- Agrawal, A.K.; Chakraborty, G. On the use of acquisition function-based Bayesian optimization method to efficiently tune SVM hyperparameters for structural damage detection. Struct. Control. Health Monit. 2021, 28, e2693. [Google Scholar] [CrossRef]
- Xie, W.; Nie, W.; Saffari, P.; Robledo, L.F.; Descote, P.Y.; Jian, W. Landslide hazard assessment based on Bayesian optimization–support vector machine in Nanping City, China. Nat. Hazards 2021, 109, 931–948. [Google Scholar] [CrossRef]
- Wu, J.; Chen, X.Y.; Zhang, H.; Xiong, L.D.; Lei, H.; Deng, S.H. Hyperparameter Optimization for Machine Learning Models Based on Bayesian Optimization. J. Electron. Sci. Technol. 2019, 17, 26–40. [Google Scholar] [CrossRef]
- Kumar, P.; Nair, G.G. An efficient classification framework for breast cancer using hyper parameter tuned Random Decision Forest Classifier and Bayesian Optimization. Biomed. Signal Process. Control 2021, 68, 102682. [Google Scholar] [CrossRef]
- Snoek, J.; Larochelle, H.; Adams, R.P. Practical Bayesian Optimization of Machine Learning Algorithms. In Proceedings of the 25th International Conference on Neural Information Processing Systems, Lake Tahoe, NV, USA, 3–6 December 2012. [Google Scholar]
- Kolar, D.; Lisjak, D.; Pajak, M.; Gudlin, M. Intelligent Fault Diagnosis of Rotary Machinery by Convolutional Neural Network with Automatic Hyper-Parameters Tuning Using Bayesian Optimization. Sensors 2021, 21, 2411. [Google Scholar] [CrossRef] [PubMed]
- Snelson, E.; Ghahramani, Z. Local and global sparse Gaussian process approximations. In Proceedings of the Eleventh International Conference on Artificial Intelligence and Statistics, San Juan, Puerto Rico, 21–24 March 2007; pp. 524–531. [Google Scholar]
- Snelson, E.; Ghahramani, Z. Sparse Gaussian Processes using Pseudo-inputs. In Proceedings of the 18th International Conference on Neural Information Processing Systems, Vancouver, BC, Canada 5–8 December 2005. [Google Scholar]
- Csató, L.; Opper, M. Sparse On-Line Gaussian Processes. Neural Comput. 2002, 14, 641–668. [Google Scholar] [CrossRef] [PubMed]
- Seeger, M.W.; Williams, C.K.I.; Lawrence, N.D. Fast Forward Selection to Speed Up Sparse Gaussian Process Regression. In Proceedings of the Ninth International Workshop on Artificial Intelligence and Statistics, Key West, FL, USA, 3–6 January 2003; pp. 254–261. [Google Scholar]
- Chen, H.; Zheng, L.; Kontar, R.A.; Raskutti, G. Gaussian Process Parameter Estimation Using Mini-batch Stochastic Gradient Descent: Convergence Guarantees and Empirical Benefits. J. Mach. Learn. Res. 2022, 23, 1–59. [Google Scholar]
- Martino, L.; Laparra, V.; Camps-Valls, G. Probabilistic cross-validation estimators for Gaussian Process regression. In Proceedings of the 25th European Signal Processing Conference, EUSIPCO 2017, Kos, Greece, 28 August–2 September 2017; pp. 823–827. [Google Scholar] [CrossRef]
- Zhang, R.; Zhao, X. Inverse Method of Centrifugal Pump Blade Based on Gaussian Process Regression. Math. Probl. Eng. 2020, 2020, 4605625. [Google Scholar] [CrossRef]
- Senanayake, R.; O’callaghan, S.; Ramos, F. Predicting Spatio-Temporal Propagation of Seasonal Influenza Using Variational Gaussian Process Regression. Proc. AAAI Conf. Artif. Intell. 2016, 30, 3901–3907. [Google Scholar] [CrossRef]
- Petelin, D.; Filipic, B.; Kocijan, J. Optimization of Gaussian process models with evolutionary algorithms. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Berlin/Heidelberg, Germany, 2011; Volume 6593, pp. 420–429. [Google Scholar] [CrossRef]
- Ouyang, Z.L.; Zou, Z.J. Nonparametric modeling of ship maneuvering motion based on Gaussian process regression optimized by genetic algorithm. Ocean Eng. 2021, 238, 109699. [Google Scholar] [CrossRef]
- Cheng, L.; Ramchandran, S.; Vatanen, T.; Lietzen, N.; Lahesmaa, R.; Vehtari, A.; Lähdesmäki, H. LonGP: An additive Gaussian process regression model for longitudinal study designs. bioRxiv 2018, 259564. [Google Scholar] [CrossRef]
- Israelsen, B.; Ahmed, N.; Center, K.; Green, R.; Bennett, W., Jr. Adaptive Simulation-Based Training of Artificial-Intelligence Decision Makers Using Bayesian Optimization. J. Aerosp. Comput. Inf. Commun. 2018, 15, 38–56. [Google Scholar] [CrossRef]
- Rasmussen, C.E.; Williams, C.K.I. Gaussian Processes for Machine Learning; MIT Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Deringer, V.L.; Bartók, A.P.; Bernstein, N.; Wilkins, D.M.; Ceriotti, M.; Csányi, G. Gaussian Process Regression for Materials and Molecules. Chem. Rev. 2021, 121, 10073–10141. [Google Scholar] [CrossRef]
- Oliveira, R.; Ott, L.; Ramos, F. Bayesian optimisation under uncertain inputs. In Proceedings of the Twenty-Second International Conference on Artificial Intelligence and Statistics, Naha, Japan, 16–18 April 2019; Volume 89, pp. 1177–1184. [Google Scholar]
- Ath, G.D.; Everson, R.M.; Rahat, A.A.M.; Fieldsend, J.E. Greed is Good: Exploration and Exploitation Trade-offs in Bayesian Optimisation. ACM Trans. Evol. Learn. Optim. 2021, 1, 1–22. [Google Scholar] [CrossRef]
- Mochihashi, D.; Oba, S. Gaussian Process and Machine Learning; Kodansha Scientific: Tokyo, Japan, 2019. [Google Scholar]
- Blonigen, B.A.; Knittel, C.R.; Soderbery, A. Keeping it Fresh: Strategic Product Redesigns and Welfare. Int. J. Ind. Organ. 2017, 53, 170–214. [Google Scholar] [CrossRef]
- Mareček, R.; Řĺha, P.; Bartoňová, M.; Kojan, M.; Lamoš, M.; Gajdoš, M.; Vojtĺšek, L.; Mikl, M.; Bartoň, M.; Doležalová, I.; et al. Automated fusion of multimodal imaging data for identifying epileptogenic lesions in patients with inconclusive magnetic resonance imaging. Hum. Brain Mapp. 2021, 42, 2921–2930. [Google Scholar] [CrossRef] [PubMed]
- Che, K.; Chen, X.; Guo, M.; Wang, C.; Liu, X. Genetic Variants Detection Based on Weighted Sparse Group Lasso. Front. Genet. 2020, 11, 155. [Google Scholar] [CrossRef] [PubMed]
- Surjanovic, S.; Bingham, D. Beale Function. 2013. Available online: https://www.sfu.ca/~ssurjano/beale.html (accessed on 27 June 2023).
- Surjanovic, S.; Bingham, D. Goldstein-Price Function. 2013. Available online: https://www.sfu.ca/~ssurjano/goldpr.html (accessed on 27 June 2023).




| Parameters | Value(s) |
|---|---|
| Maximum number of GD iterations | |
| GD learning rate | |
| Maximum number of BO iterations | 50 |
| Exploration weight | |
| Initial KP | |
| Search space | Equation (25) |
| Black-box function | Equation (25) |
| Observation noise | 0 |
| Initial observation dataset | Four points randomly selected in |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Omae, Y. Effects of Exploration Weight and Overtuned Kernel Parameters on Gaussian Process-Based Bayesian Optimization Search Performance. Mathematics 2023, 11, 3067. https://doi.org/10.3390/math11143067
Omae Y. Effects of Exploration Weight and Overtuned Kernel Parameters on Gaussian Process-Based Bayesian Optimization Search Performance. Mathematics. 2023; 11(14):3067. https://doi.org/10.3390/math11143067
Chicago/Turabian StyleOmae, Yuto. 2023. "Effects of Exploration Weight and Overtuned Kernel Parameters on Gaussian Process-Based Bayesian Optimization Search Performance" Mathematics 11, no. 14: 3067. https://doi.org/10.3390/math11143067
APA StyleOmae, Y. (2023). Effects of Exploration Weight and Overtuned Kernel Parameters on Gaussian Process-Based Bayesian Optimization Search Performance. Mathematics, 11(14), 3067. https://doi.org/10.3390/math11143067








