Abstract
Respiratory diseases constitute a major global health burden, necessitating accurate and reliable diagnostic support tools. Conventional auscultation, despite its widespread clinical use, remains inherently subjective and susceptible to inter-observer variability. In this study, we propose a unified deep learning framework for the automated classification of respiratory sound recordings into four clinically relevant categories: Normal, Crackles, Wheezes, and Crackles + Wheezes. The experimental evaluation was conducted on a publicly available dataset comprising heterogeneous respiratory recordings collected from both patients with pulmonary pathologies and healthy individuals. All audio signals were subjected to standardized preprocessing procedures to enhance signal consistency and ensure reliable feature extraction across acquisition conditions. To ensure methodological rigor and prevent optimistic bias, a strict subject-independent validation strategy was adopted using 5-fold GroupKFold cross-validation based on patient identifiers. Six deep learning architectures were systematically implemented and comparatively evaluated under a controlled and reproducible training protocol, including convolutional (1D-CNN, Deep-CNN), recurrent hybrid (CNN–LSTM, CNN–BiLSTM), and attention-based (CNN–Attention, CNN–Transformer) models. Performance metrics were reported as mean ± standard deviation across folds. The CNN–Attention architecture achieved the best overall performance, yielding a Balanced Accuracy of 90.1% ± 1.8% and a macro F1-score of 89.7% ± 2.1%, demonstrating stable inter-patient generalization. These findings indicate that attention-enhanced hybrid architectures effectively capture both local spectral structures and long-range temporal dependencies inherent in respiratory signals. The proposed framework provides a robust foundation for subject-independent automated lung sound classification and contributes to the development of clinically reliable decision-support systems.
1. Introduction
Automated respiratory sound analysis is an area of continuous development, driven by the urgent need for objective, reproducible auscultation tools that can mitigate the well-documented limitations of human interpretation. In this broader context, several studies have explored machine learning and deep learning techniques for the automatic classification of respiratory-related acoustic signals, including cough sounds and other respiratory events [1,2,3,4,5]. Various early works, such as that of Aykanat et al. [6] utilizing a large private dataset of 17,930 recordings, have reported accuracies ranging between 62% and 86% using Mel-Frequency Cepstral Coefficients (MFCC), Support Vector Machine (SVM) and Convolutional Neural Networks (CNNs). As a result of the non-public nature of this dataset, even these relatively large-scale models remain difficult to compare with the subsequent body of research. Similarly, Bokov et al. [7] focused on a small real-world dataset of 95 validated recordings using smartphone-based pediatric wheeze detection to achieve a sensitivity of 71.4% and specificity of 88.9%, but addressed only a binary wheezing task. These early efforts illustrate the feasibility of automated auscultation but remain limited in scope and generalizability.
The next wave of research diversified the methodology. Pahar et al. [8] presented Natural Language Processing (NLP)-style cough embeddings to distinguish tuberculosis from other respiratory conditions, reaching an AUC of 0.81 with a Long Short-Term Memory (LSTM) classifier. Though irrelevant to lung auscultation itself, this finding underscores the relevance of temporal modeling in respiratory acoustics. Another turn toward physiological variety was taken by McClure et al. [9], who proposed a wearable-sensor-based system with accelerometer and gyroscope data from 100 volunteers, identifying complex breathing patterns of apnea, cough, sigh, and yawning with F1 scores ranging between 51% and 92%. These studies illustrate the diversity within respiratory sensing modalities, but their core differs from stethoscope-based classification.
Closer to auscultation tasks, the pediatric-focused studies of Kim et al. [10] have proposed advanced deep learning models including a 34-layer Residual Network (ResNet) with Convolutional Block Attention Module (CBAM) attention which achieved 91.2% accuracy in wheeze detection on real-world recordings from 76 children, while Kim et al. [11] trained an AST model on 725 pediatric recordings and obtained 91.1% accuracy and F1 = 82.2%. Although high performing, these models target wheeze-only tasks and do not generalize to crackles or mixed events.
The complexity of this classification task is underscored by Park et al. [12], who, for the separation of Normal, Crackles and Wheezes, trained ensemble SVMs on 680 pediatric clips, reported accuracies ranging between 67% and 90%, depending on the class pairs. A prospective validation by the same authors revealed significant performance degradation, underlining the difficulty of deploying models that are trained with limited, homogeneous datasets. Furthermore, Mendes et al. [13] investigated wheeze-specific spectral and musical descriptors, finding spectrogram-based wheeze signatures to be the most discriminative features. More recently, Amose et al. [14] tested classical SVM and Decision Tree classifiers on four categories of adventitious sounds (“normal, crackle, wheeze, both”) and concluded that their performances are superior to tree-based methods but remain limited by handcrafted features.
Among works based on the public ICBHI 2017 dataset, the contribution of Petmezas et al. [15] is directly comparable to our study. Their hybrid CNN–LSTM model achieved an accuracy of 73.69% for a standard split of 60/40 and 76.39% under inter-patient cross-validation, markedly lower than results obtained on private pediatric datasets, reflecting increased complexity and heterogeneity in the ICBHI benchmark. Complementary observations were made by Rocha et al. [16], who explored the effect of event duration variability and class definition, especially the “other” class, on ARS classification performance. Their experiments indicated that while fixed-duration tasks could reach high accuracies up to 96.9%, more realistic variable-duration settings reduce performance to 81.8%, thereby stressing that automatic ARS classification remains an open and challenging issue in non-controlled environments. Moreover, Polanco-Martagón et al. [17] compared five pre-trained CNNs on the ICBHI dataset: VGG16, VGG19, InceptionV3, MobileNetV2, and ResNet152V2, reporting best Average Score values around 0.541–0.58, also outlining important clinical biases such as high specificity but poor sensitivity, a serious problem that risks not detecting pathological events.
In contrast to this fragmented and heterogeneous landscape, our study presents a unified and systematic evaluation across six deep learning architectures, purely convolutional (1D-CNN, Deep-CNN), recurrent hybrids (CNN–LSTM, CNN–BiLSTM), and attention-enhanced models (CNN–Attention, CNN–Transformer), all trained under an identical preprocessing, augmentation, Synthetic Minority Oversampling Technique (SMOTE) balancing, and hyperparameter pipeline on the same four-class taxonomy (Normal, Crackles, Wheezes, Crackles + Wheezes) using the public ICBHI dataset. This standardized framework enables a direct and controlled comparison of model behaviors under identical experimental conditions. The results indicate that attention-enhanced architectures consistently outperform purely convolutional and recurrent baselines under strict subject-independent validation. While performance remains competitive, the adoption of a patient-wise cross-validation protocol provides a more realistic assessment of inter-patient generalization compared to recording-wise evaluations frequently reported in the literature. Our work addresses key reproducibility limitations underlined in previous studies by harmonizing evaluation conditions and providing transparent metrics of performance, thus showing that spectral-temporal feature combination with deep attention mechanisms leads to a robust and reliable multi-class respiratory sound classification framework under the adopted experimental conditions. While these findings suggest potential applicability in computer-assisted auscultation systems, further validation under real-world clinical and longitudinal monitoring scenarios is required.
Unlike transfer-learning approaches relying on pretrained image-based backbones or generic deep learning configurations, all neural architectures evaluated in this study were specifically designed and implemented for respiratory acoustic classification. Although based on established paradigms (CNN, recurrent networks, attention mechanisms, and Transformers), the internal structure of each model, including convolutional block depth, filter configuration, recurrent dimensionality, attention placement, dropout regularization, and optimization scheduling, was experimentally calibrated for the MFCC-based representation derived from standardized 5-s respiratory cycles. This task-specific architectural engineering, combined with a unified preprocessing, augmentation, and balancing framework, enables a controlled and reproducible comparative evaluation of modeling strategies for multi-class respiratory sound classification
The main contributions of this study can be summarized as follows:
- A clinically grounded acoustic relabeling strategy based on intrinsic respiratory sound morphology rather than diagnostic labels;
- A comprehensive and structured preprocessing pipeline integrating signal standardization, waveform-level augmentation, SpecAugment, and feature-space balancing via SMOTE to address recording heterogeneity and class imbalance;
- A unified experimental framework ensuring controlled comparison across architectures under identical preprocessing and optimization settings;
- A systematic performance analysis under subject-independent validation demonstrating the contribution of temporal modeling and attention mechanisms to robust multi-class respiratory sound classification.
2. Materials and Methods
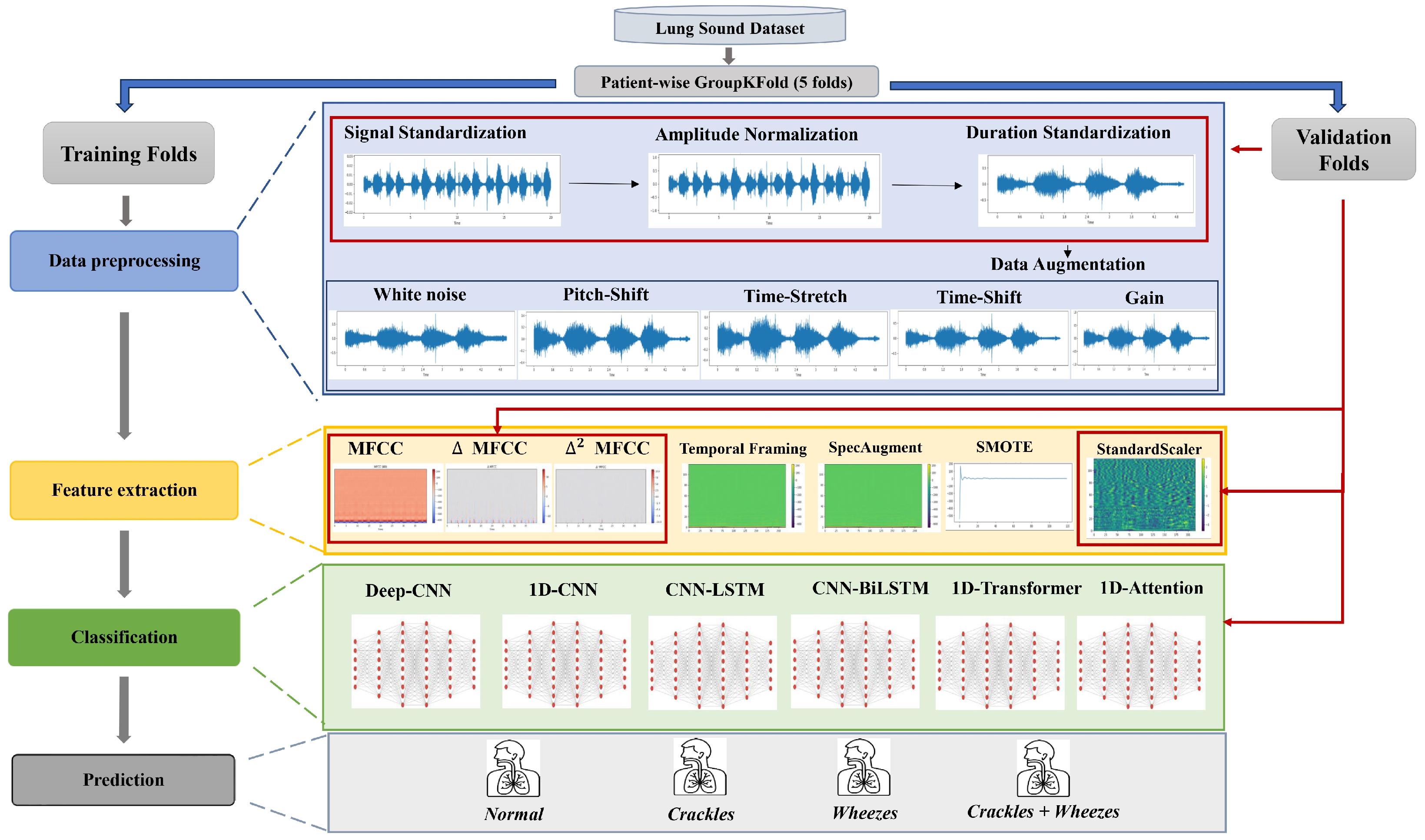
The methodological framework adopted in this study follows a structured and rigorously controlled multistage pipeline designed to ensure reproducibility and methodological transparency in respiratory sound classification. As illustrated in Figure 1, the entire workflow is embedded within a strict subject-independent validation protocol implemented using patient-wise 5-fold cross-validation. In each iteration, four folds containing mutually exclusive patient groups are used for model training, while the remaining fold serves as an independent validation subset, thereby ensuring complete inter-patient separation. Within each cross-validation iteration, raw respiratory recordings from the ICBHI 2017 dataset undergo deterministic preprocessing operations, including channel standardization, controlled resampling, band-pass filtering, amplitude normalization, and duration standardization. These operations are applied consistently across both training and validation folds to ensure uniform signal conditioning while avoiding information leakage. To address class imbalance and improve intra-class variability, stochastic data augmentation procedures are applied exclusively to the training folds. Feature extraction is subsequently performed using Mel-Frequency Cepstral Coefficients (MFCCs) and their first- and second-order temporal derivatives, computed over standardized 5-s segments to generate fixed-size time–frequency representations. Class imbalance within each training fold is mitigated using the Synthetic Minority Oversampling Technique (SMOTE), applied strictly within the training data to preserve fold independence. Feature scaling is performed using z-score normalization, where the mean and standard deviation parameters are estimated solely from the training data of each fold and subsequently applied to the corresponding validation fold. This fold-wise normalization strategy prevents statistical contamination between training and validation subsets. Finally, convolutional, recurrent hybrid, and attention-based deep neural network architectures are trained and evaluated independently within each fold to classify respiratory sounds into four acoustic categories: Normal, Crackles, Wheezes, and Crackles + Wheezes. Model performance is reported as mean ± standard deviation across the five folds, providing a statistically robust estimate of inter-patient generalization performance.
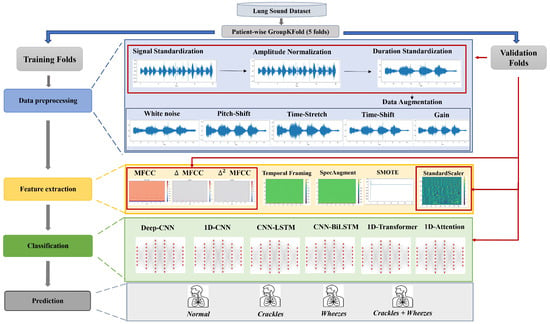
Figure 1.
Overall Processing Pipeline of the Proposed Respiratory Sound Classification Framework.
2.1. Dataset
This study uses the open-access ICBHI 2017 Respiratory Sound Database, originally introduced by [18], and released as part of the 1st International Conference on Biomedical and Health Informatics (ICBHI 2017) Challenge. It is widely used as a benchmark dataset for the validation and comparison of automatic lung sound classification techniques. The database consists of 920 audio recordings from 126 participants (both male and female, children up to elderly adults), including healthy subjects and patients diagnosed with respiratory conditions such as pneumonia, Chronic Obstructive Pulmonary Disease (COPD), asthma, bronchiectasis, bronchiolitis, Upper Respiratory Tract Infection (URTI), and Lower Respiratory Tract Infection (LRTI).
Recordings, whose length ranges from 0.5 to 90 s, were acquired in two different hospitals in Portugal and Greece with different electronic stethoscopes, including Littmann, Welch Allyn, and AKG-C417 L, with sampling rates ranging between 4 kHz and 44.1 kHz. Lung sounds were recorded over different thoracic regions from the trachea, the bases, the dorsal region, and the lateral regions. Recordings also include annotation files indicating the beginning and end of each respiratory cycle, including inspiration, expiration, and abnormal events occurring within each cycle. A total of about 5.5 h of recorded signals and 6898 annotated breath cycles are included in this dataset. Due to the heterogeneity in acquisition devices, recording environments, and signal durations, the dataset presents significant variability, thereby necessitating rigorous preprocessing prior to automated analysis.
Acoustic Relabeling and Class Balancing
- 1.
- Motivation and RationaleTo adapt the ICBHI 2017 Respiratory Sound Database to a strictly acoustic labeling framework, all 920 recordings were relabeled at the file level based on the respiratory sound events identified within the provided cycle-level annotations, rather than on their associated clinical diagnoses. This methodological decision is motivated by the observation that diagnostic labels such as asthma, COPD, or pneumonia often exhibit overlapping acoustic manifestations due to shared underlying physiological mechanisms. An acoustic-based taxonomy establishes a more objective and reproducible correspondence between auditory phenomena and classification targets. By focusing exclusively on the intrinsic morphological characteristics of respiratory sounds, the proposed relabeling strategy reduces inter-diagnostic ambiguity and allows the models to learn directly from spectral and temporal signal properties, rather than from diagnosis-dependent clinical metadata.
- 2.
- Relabeling Procedure & Class DistributionThe ICBHI dataset provides time-stamped annotations specifying the presence of Crackles and/or Wheezes within each audio recording. These annotations were aggregated at the file level, and each recording was assigned a single global acoustic label following a deterministic rule:In clinical auscultation, respiratory sounds are categorized into normal vesicular breathing and adventitious sounds. Crackles are brief, discontinuous, explosive sounds typically occurring during inspiration due to sudden airway opening, whereas wheezes are continuous, musical sounds generated by airflow limitation in narrowed bronchi. Recordings containing both acoustic events were grouped as “Crackles + Wheezes”. Although other pathological sounds such as stridor, rhonchi, or pleural friction rub exist, they are either not separately annotated or insufficiently represented in the ICBHI 2017 dataset. Therefore, the four-class taxonomy adopted in this work reflects both clinical relevance and dataset constraints.This approach effectively captures the major adventitious respiratory sound categories typically encountered in clinical auscultation. However, the resulting dataset remains highly imbalanced, containing 794 recordings labeled as Wheezes (86.3%), 55 recordings labeled as Crackles (6.0%), 58 recordings labeled as Normal (6.3%), and 13 recordings labeled as Crackles + Wheezes (1.4%). This imbalance reflects the higher clinical prevalence of wheezing in obstructive pulmonary conditions, alongside the underrepresentation of rarer acoustic events such as fine inspiratory crackles.
2.2. Data Preprocessing
2.2.1. Mono Conversion and Resampling
All respiratory sound recordings were initially converted to mono-channel signals to remove redundancy from stereo channels and ensure comparability across different acquisition devices. Subsequently, each file was resampled at a uniform frequency of 16 kHz, providing a consistent spectral resolution suitable for capturing the key frequency components of respiratory sounds. This harmonized resampling step ensures that all recordings share a common temporal and spectral scale, preventing distortion during feature extraction, particularly in CNN-based models, which are sensitive to inconsistencies in time–frequency representations.
2.2.2. Band-Pass Filtering
After resampling, a 10th-order Butterworth band-pass filter [19] with cutoff frequencies of 100 Hz and 2000 Hz was applied to each signal. This filtering stage removes low-frequency artifacts, such as heart sounds, body movements, and microphone handling noise, as well as high-frequency interference such as ambient or electronic noise. Mathematically, the filtering process can be expressed as:
where denotes the input respiratory acoustic signal prior to filtering, represents the filtered output signal, and are the Butterworth band-pass filter coefficients, and is the filter order. The selected 100–2000 Hz frequency range effectively encompasses the principal energy spectrum of respiratory acoustic phenomena, notably Crackles (200–1200 Hz) and Wheezes (400–1600 Hz), thereby isolating the most diagnostically informative spectral bands for subsequent analysis.
2.2.3. Amplitude Normalization & Standardization
To account for amplitude variations arising from different types of stethoscope and recording conditions [20], each waveform was normalized within the interval using:
This normalization stabilizes amplitude discrepancies and ensures a uniform dynamic range across samples. The duration of each recording was standardized to 5 s, implemented via zero-padding for shorter samples and deterministic center-cropping for recordings exceeding 5 s. For longer recordings, a contiguous 5-s segment centered within the signal duration was extracted. The same deterministic cropping strategy was consistently applied across both training and validation folds to ensure methodological uniformity.
This temporal normalization guarantees consistent input lengths across the dataset, facilitating efficient batch training and enabling deep learning models to process sequences of comparable duration. The five-second window typically encompasses at least one complete respiratory cycle, including both inspiratory and expiratory phases, thereby preserving the essential temporal dynamics of breathing sounds. Although center-cropping was adopted to ensure uniform input dimensionality, abnormal acoustic events may occasionally occupy only a fraction of the selected 5-s segment. Consequently, residual segment-level label noise cannot be entirely excluded, particularly for transient events such as isolated crackles.
Although the ICBHI dataset provides respiratory cycle boundaries, fixed-length standardization was deliberately adopted to ensure uniform input dimensionality across all evaluated architectures and to maintain strict comparability within the proposed benchmarking framework. Cycle-level segmentation would introduce variable-length sequences requiring additional padding, truncation strategies, or dynamic batching mechanisms, potentially affecting architectural behavior and complicating controlled cross-model evaluation. Therefore, the 5-s window was selected as a methodological design choice that prioritizes reproducibility and fair architectural comparison rather than task-specific optimization.
2.2.4. Data Augmentation
To improve the robustness of the models against clinical variability and reduce the risk of overfitting, a series of data augmentation strategies were implemented [21,22,23,24]. Signal-level augmentations were applied prior to MFCC extraction, while spectral augmentations (SpecAugment) were applied after MFCC computation, as they operate directly on time–frequency representations.
- White noiseTo emulate realistic auscultation environments, additive white Gaussian noise was injected into the waveforms with a randomly selected signal-to-noise ratio (SNR) ranging between 5 and 20 dB. This approach replicates common sources of disturbance, such as ambient noise, clothing friction, microphone hiss, or patient movement artifacts, while maintaining the perceptual integrity of pathological acoustic features, particularly crackle transients and wheeze harmonics. This process encourages the network to develop noise-tolerant representations capable of discriminating meaningful features under suboptimal recording conditions.
- Time-shiftA small temporal translation of up to was applied by circularly shifting each waveform. This transformation reproduces natural timing variations between inspiration and expiration cycles typically observed in uncontrolled respiratory recordings. The procedure promotes the learning of phase-invariant features and prevents the model from relying excessively on fixed temporal event positioning within the breathing cycle.
- Gain variationTo simulate variations in stethoscope sensitivity, microphone distance, and patient chest impedance, random amplitude gains of were applied. This augmentation compensates for differences in acquisition settings and inter-patient anatomical factors, thereby improving amplitude invariance and reducing overfitting to specific recording configurations.
- Pitch shiftUsing a phase-vocoder algorithm, the spectral content of the respiratory signals was shifted by semitones without altering the temporal structure. This operation models inter-individual physiological differences, such as airway diameter, thoracic cavity resonance, and airflow turbulence. Because wheezes exhibit quasi-harmonic spectral structures, this augmentation facilitates generalization across pitch variations and inter-patient diversity.
- Time-StretchThe waveform duration was scaled by a random factor between 0.9 and 1.1, simulating natural variations in breathing rates and respiratory cycle lengths. While maintaining spectral morphology, this transformation modifies temporal dynamics, promoting robustness of the model to irregular or patient-specific breathing patterns.
Collectively, these augmentation techniques preserve the physiological plausibility of respiratory sounds while significantly improving intra-class variability, thereby improving the model’s capacity for generalization. For experimental integrity, these transformations were applied exclusively to the training folds within each patient-wise cross-validation iteration, while validation folds were kept strictly unmodified to ensure unbiased performance estimation.
2.3. Feature Extraction
2.3.1. MFCC Computation
Following preprocessing, the acoustic characteristics of each respiratory sound were quantified using Mel-Frequency Cepstral Coefficients (MFCCs) [25,26]. After duration standardization to a fixed 5 s window, each recording was divided into overlapping frames of 25 ms with a hop length of 10 ms prior to Short-Time Fourier Transform (STFT):
where denotes the STFT coefficient at frequency bin k and time frame n, M is the frame length, and H represents the hop size.
The power spectrum was then computed for each time frame n as and projected onto the Mel scale using triangular filter banks. The energy of the m-th Mel filter at frame n is defined as:
where and define the frequency boundaries of the m-th Mel filter.
Subsequently, the logarithm of the Mel-filterbank energies was transformed using the Discrete Cosine Transform (DCT) to obtain the cepstral coefficients:
These coefficients represent the spectral envelope of the signal at each time frame, effectively encoding perceptually relevant acoustic features that distinguish normal and pathological respiratory sounds.
2.3.2. Temporal Derivatives ( and )
To capture the temporal dynamics of respiratory acoustics, first-order () and second-order () derivatives of MFCCs were computed [27]. These features quantify both the rate and acceleration of spectral variations over time, thereby highlighting distinctive temporal signatures of different respiratory phenomena. The derivatives are calculated as:
where is the contextual window. While wheezes exhibit smooth and continuous harmonic trajectories, crackles are characterized by short, impulsive transients; and features effectively capture these contrasting temporal dynamics. Concatenating the static MFCCs with their and derivatives yields a 120-dimensional feature vector per frame (), providing a comprehensive spectral-temporal representation of each respiratory sound.
2.3.3. Feature Formatting and Padding
Each preprocessed recording was transformed into a two-dimensional feature matrix, where rows represent time frames and columns correspond to the 120 extracted features:
Following duration standardization to 5 s, feature extraction was performed within each patient-wise cross-validation fold. The resulting matrices correspond to a fixed temporal length across all samples, ensuring consistent batching, stable optimization, and improved generalization capabilities. This standardization facilitates efficient batch processing and guarantees constant input dimensionality, thereby preventing architectural bias when comparing different deep learning models [28].
2.3.4. Spectral Augmentation
To further improve generalization and reduce overfitting to fixed spectral patterns, a SpecAugment strategy was applied directly to the MFCC feature matrices [29]. Unlike waveform-level augmentations, SpecAugment operates in the time–frequency domain, masking contiguous temporal and frequency segments:
- Frequency maskingTwo random frequency bands (width = 8 Mel filters) were suppressed to emulate the partial loss of harmonic content.
- Time maskingTwo temporal intervals (width = 20 frames) were hidden to simulate missing or occluded respiratory events.
Formally, time and frequency masks (, ) were defined as:
These augmentation techniques preserve the physiological plausibility of respiratory sounds while enhancing intra-class variability, thereby improving the model’s capacity for generalization. SpecAugment transformations were applied exclusively to the training folds within each patient-wise cross-validation iteration, while validation folds remained strictly unmodified to ensure unbiased performance estimation.
2.3.5. Class Balancing and Normalization
Due to the strong class imbalance observed in the dataset, particularly the dominance of the Wheezes class over the remaining categories, the Synthetic Minority Over-sampling Technique (SMOTE) [30,31] was employed to improve the representation of minority classes during training.
SMOTE generates synthetic samples by interpolating between existing minority-class samples and their nearest neighbors in the feature space. Given a minority feature vector and one of its k-nearest neighbors , a synthetic instance is generated according to:
where denotes an existing minority-class feature vector, represents one of its nearest neighbors, is the synthetic feature vector generated by SMOTE, and is a random interpolation coefficient drawn from a uniform distribution over .
In the present study, each respiratory recording is represented by a fixed-size MFCC feature matrix of , capturing the spectral-temporal structure of respiratory sounds. Since SMOTE operates on feature vectors, each MFCC matrix was first flattened into a -dimensional feature vector prior to oversampling. Synthetic minority samples were then generated in this feature space using the standard SMOTE interpolation mechanism. After generation, the synthetic vectors were reshaped back into the original matrix format before being used as input to the deep learning architectures. SMOTE is therefore applied at the recording level rather than at the frame level, ensuring that each synthetic sample represents a complete respiratory recording rather than an isolated time frame. Although SMOTE is applied in the flattened feature space, the MFCC representation provides a compact encoding of the spectral-temporal structure of respiratory sounds. Consequently, interpolation between neighboring MFCC representations remains consistent with the underlying acoustic structure of respiratory sounds and has been widely adopted in audio classification pipelines where spectrogram-like representations are used as inputs to deep learning models.
To preserve strict subject independence and avoid information leakage, SMOTE was applied exclusively to the training subset within each patient-wise cross-validation fold, while validation folds remained completely untouched.
Subsequently, all features were standardized using z-score normalization computed on the training set:
where and denote the mean and standard deviation calculated for each feature dimension using the training fold only. The estimated normalization parameters were then applied to the corresponding validation fold. This procedure centers and scales the feature distributions, ensuring unit variance across input dimensions, which stabilizes gradient propagation and accelerates training convergence. The resulting dataset, composed of balanced and normalized MFCC feature matrices (), provides a consistent and optimized input representation for the subsequent deep learning classification architectures.
2.4. Classification
All classification models were trained using the same preprocessed and balanced dataset, consisting of feature matrices of size 216 × 120. These matrices, derived from MFCCs and their temporal derivatives, encode the time–frequency structure of respiratory cycles, where each recording is represented by 216 temporal frames and 120 spectral coefficients per frame. Model training was performed within a 5-fold patient-wise GroupKFold cross-validation framework, ensuring strict inter-patient separation between training and validation folds.
To ensure optimal convergence and minimize overfitting, the training configurations were tailored to each model’s learning dynamics:
- Convolutional architectures employed moderate learning rates and larger batch sizes to facilitate faster convergence.
- Recurrent and attention-based models utilized smaller learning rates and batch sizes to maintain gradient stability and temporal coherence.
- Transformer-based architectures benefited from learning rate warm-up schedules and stronger dropout regularization, thereby accommodating their larger parameter space and sensitivity to optimization dynamics.
This adaptive training strategy ensured a rigorous and equitable performance comparison across architectures, optimizing each model’s capacity to represent complex acoustic phenomena under consistent evaluation settings.
Performance metrics were computed for each validation fold independently and are reported as mean ± standard deviation across the five patient-wise folds.
All networks produced a final output yielding probability distributions across four respiratory classes: Normal, Crackles, Wheezes, and Crackles + Wheezes. To comprehensively capture both local and global temporal dependencies, six neural architectures were implemented:
- Purely convolutional models (1D-CNN, Deep-CNN),
- Recurrent hybrid models (CNN-LSTM, CNN-BiLSTM), and
- Attention-based architectures (CNN-Attention, CNN-Transformer).
This architectural diversity enables a holistic comparative analysis of distinct modeling paradigms and their respective implications for respiratory sound interpretation.
2.4.1. One-Dimensional Convolutional Neural Network (1D-CNN)
The Convolutional Neural Network (CNN) remains a foundational architecture in deep learning due to its ability to automatically learn discriminative feature representations from raw or preprocessed signals [32]. In this study, a 1D-CNN was employed as a baseline model, demonstrating strong performance in modeling sequential acoustic data.
Although the MFCC-based features form 2D matrices (216 × 120), representing time and spectral dimensions respectively, the data were reshaped into 1D temporal sequences by flattening the spectral axis. This reshaping preserves the temporal continuity of respiratory sounds while reducing computational complexity. Compared with traditional 2D-CNNs, the 1D-CNN provides a lightweight yet effective solution for capturing local temporal dependencies.
Mathematically, the convolution operation for a one-dimensional input signal is expressed as:
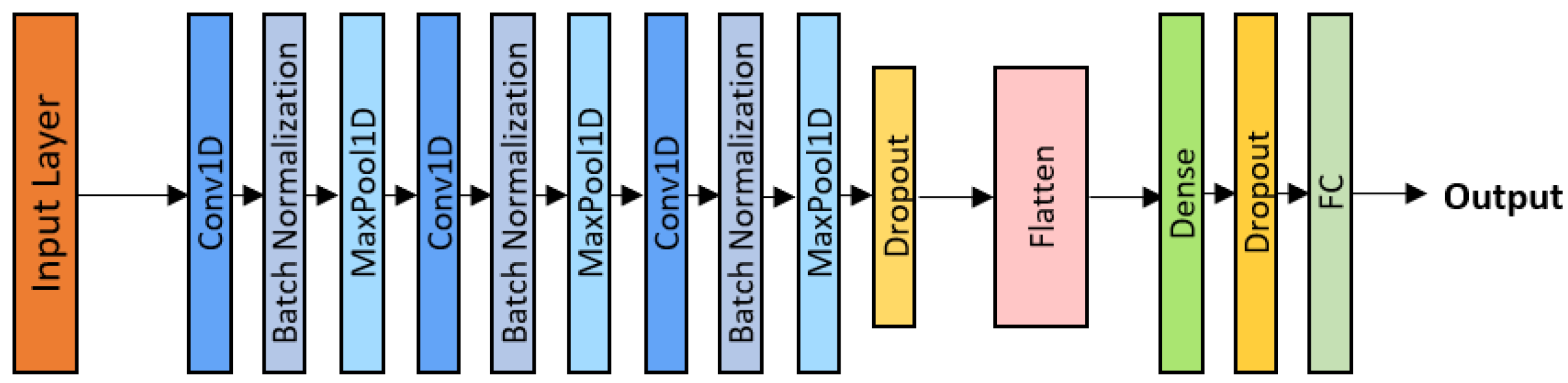
where denotes the learnable convolutional kernel of size k, and represents the resulting feature map. Each filter acts as a pattern detector, identifying characteristic spectro-temporal structures such as crackle bursts or harmonic wheeze patterns. The implemented architecture (Figure 2) comprises three convolutional blocks, each consisting of a Conv1D layer followed by Batch Normalization, a ReLU activation function, and a MaxPooling layer. A dropout rate of was applied after the final convolutional block to enhance generalization and mitigate overfitting.
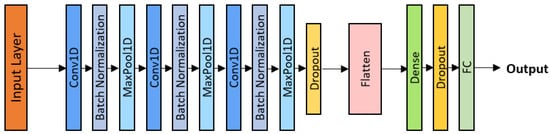
Figure 2.
1D-CNN architecture.
Training was performed using the Adam optimizer with a learning rate of , a batch size of 32, and a total of 43 epochs. This configuration enabled rapid convergence while maintaining robustness to inter-patient variability, thereby serving as a strong benchmark for comparison with subsequent neural architectures.
2.4.2. Deep Convolutional Neural Network (Deep-CNN)
The Deep-CNN extends the 1D-CNN by incorporating multiple hierarchical convolutional layers, enabling the network to extract progressively abstract acoustic representations [33]. Each block includes a convolution, Batch Normalization, ReLU activation, and MaxPooling layer. Deeper layers capture higher-level spectral–temporal features, improving discrimination between overlapping respiratory events. Forward propagation through the l-th layer is defined as:
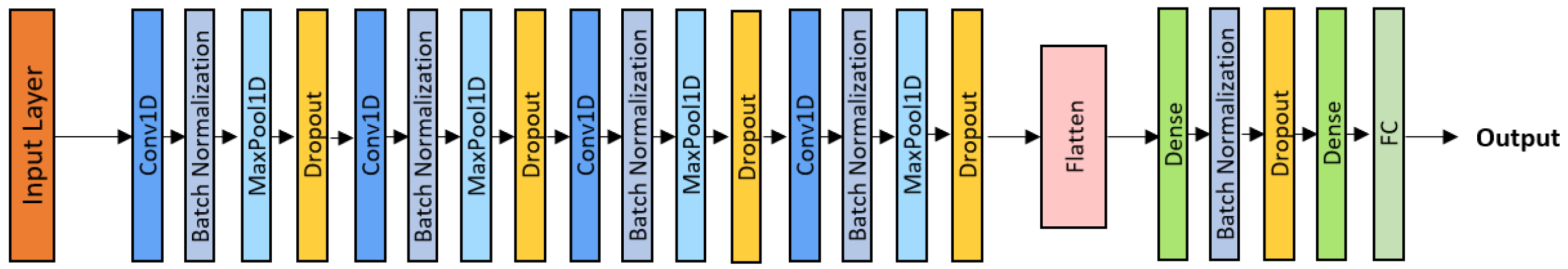
where denotes the ReLU activation function, and and are the trainable parameters of the l-th layer. Dropout regularization was applied both within convolutional blocks and between dense layers (Figure 3), thereby mitigating overfitting. The model was trained using the Adam optimizer with a learning rate of , a batch size of 32, and 45 epochs. This deeper network effectively captured subtle spectral interactions, such as the superposition of crackles and wheezes, offering improved sensitivity to complex pathological respiratory sounds.
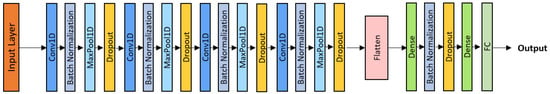
Figure 3.
Deep-CNN architecture.
2.4.3. CNN-LSTM Model
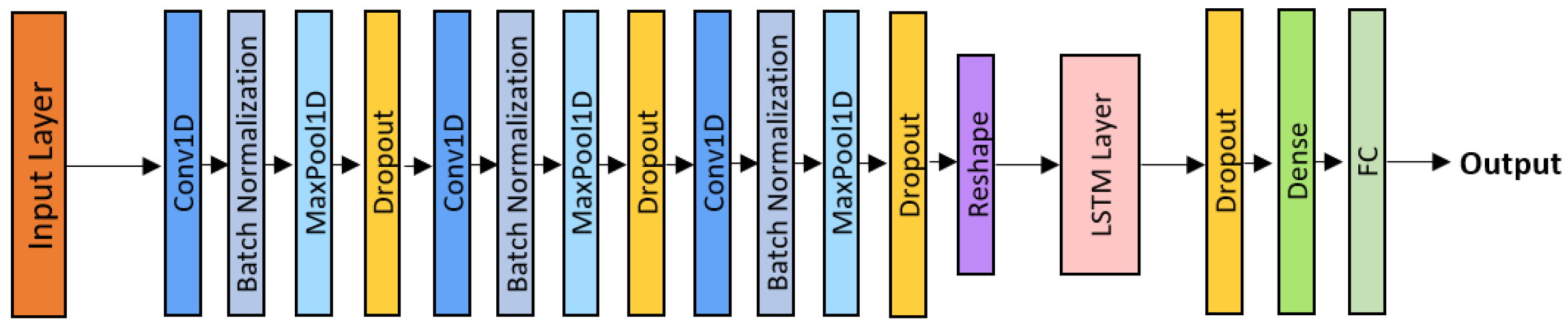
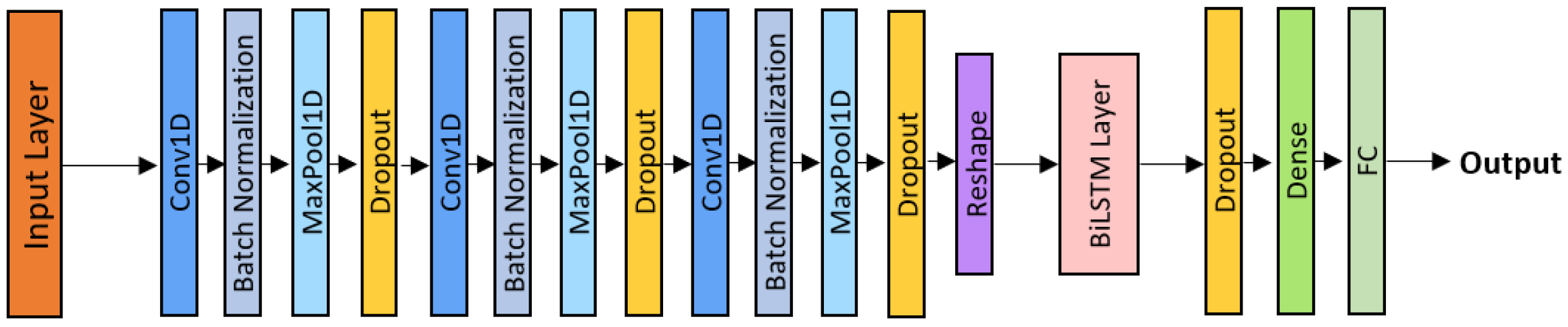
The CNN-LSTM model combines the local feature extraction capabilities of convolutional neural networks with the temporal modeling capacity of Long Short-Term Memory (LSTM) networks [34]. As illustrated in Figure 4, the architecture comprises three convolutional blocks (Conv1D + Batch Normalization + ReLU + MaxPooling + Dropout), followed by a unidirectional LSTM layer. While the convolutional front-end extracts short-term spectro-temporal features, the LSTM layer processes these feature sequences over time, thereby capturing long-range dependencies between inspiration and expiration phases. Each LSTM cell regulates information retention and memory flow through gating mechanisms, enabling the model to emphasize diagnostically relevant acoustic patterns while suppressing background noise.
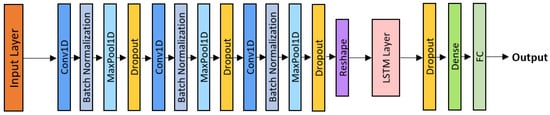
Figure 4.
CNN-LSTM architecture.
The model was trained using the Adam optimizer with a learning rate of , a batch size of 24, and 70 epochs, with a dropout rate of applied between the LSTM and dense layers. This configuration enhanced training stability and enabled the network to effectively represent repetitive or evolving pathological sounds, such as persistent wheezing or clustered crackles.
2.4.4. CNN-BiLSTM Model
The CNN–BiLSTM model extends the previous architecture by introducing bidirectional temporal processing, allowing the network to consider both past and future contexts in respiratory sequences [35]. This design is particularly relevant since abnormal sounds can occur during either the inspiratory or expiratory phase.
Three convolutional blocks (Conv1D + Batch Normalization + ReLU + MaxPooling + Dropout) were followed by a Bidirectional LSTM layer, as shown in Figure 5. The BiLSTM computes forward and backward hidden states, combined as:
where and denote the forward and backward hidden states, respectively. A dropout rate of was applied after the BiLSTM layer and before the dense layers to ensure effective regularization. The model was trained using the Adam optimizer with a learning rate of , a batch size of 16, and 80 epochs, achieving stable convergence and improved robustness against the vanishing gradient problem.
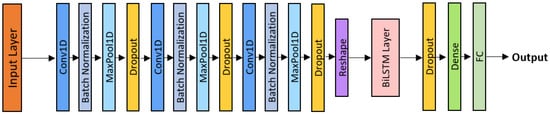
Figure 5.
CNN-BiLSTM architecture.
2.4.5. CNN-Attention Model
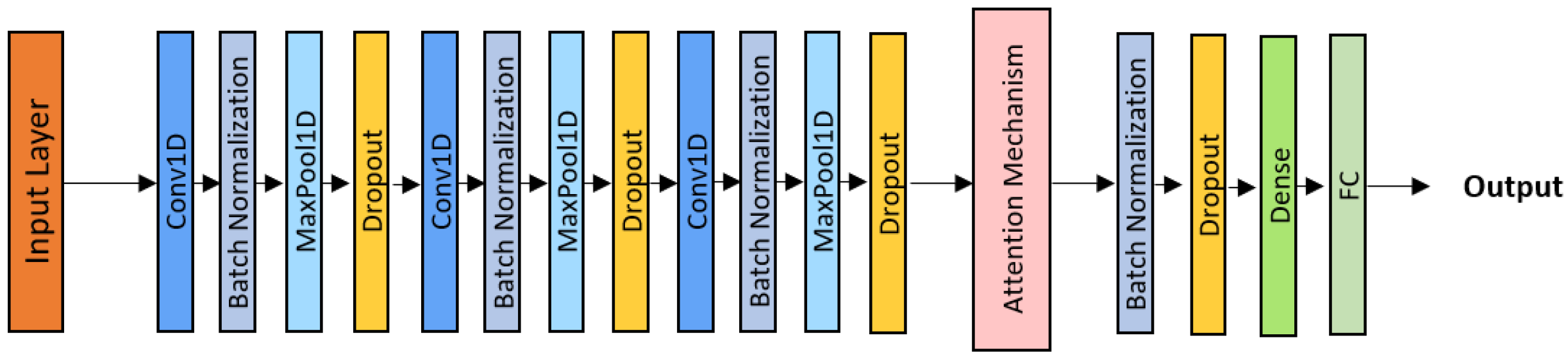
The CNN–Attention model incorporates a learnable attention mechanism to prioritize diagnostically relevant portions of the respiratory signal [36]. After three convolutional blocks (Conv1D + Batch Normalization + ReLU + MaxPooling + Dropout) as shown in Figure 6, an attention module computes weights over temporal frames, emphasizing key patterns such as isolated crackles or sustained wheezes.
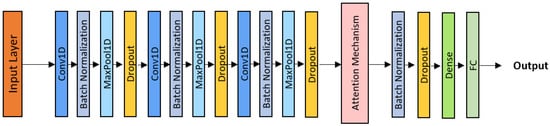
Figure 6.
CNN-Attention architecture.
The attention layer computes a weighted feature aggregation, enabling the model to focus adaptively on clinically informative segments while suppressing noise. Dropout (0.4) was applied before and after the attention layer to reduce overfitting.
Training was performed using the Adam optimizer with a learning rate of , a batch size of 24, and 70 epochs. This configuration yielded high interpretability and precision, as the model dynamically adjusted its attention across varying acoustic patterns.
2.4.6. CNN-Transformer Model
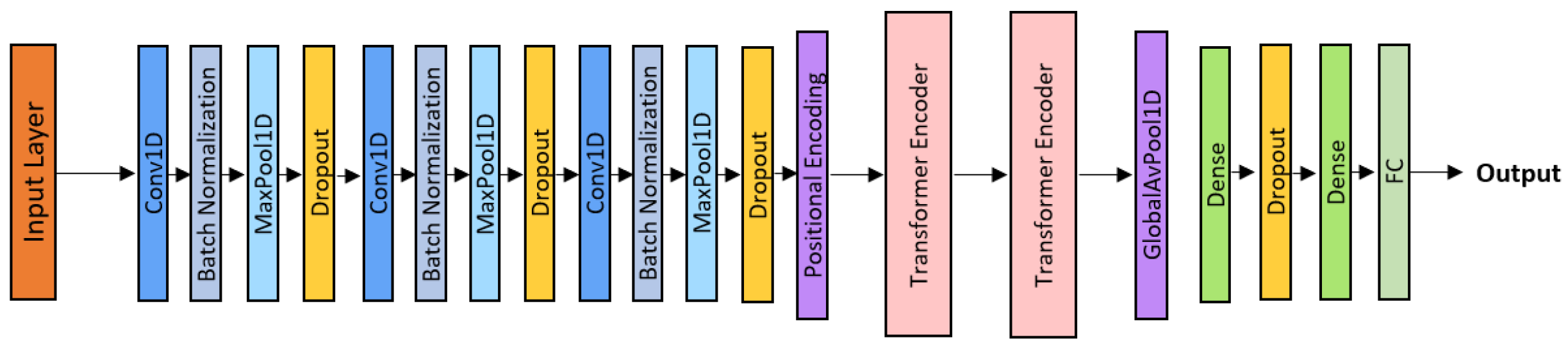
The CNN–Transformer architecture integrates convolutional front-end layers for local feature extraction with Transformer encoders for global temporal modeling through self-attention [11].
As shown in Figure 7, the architecture consists of three convolutional blocks, positional encoding, and two Transformer encoder layers, followed by Global Average Pooling, Dropout, and a dense output layer.
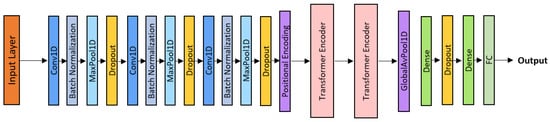
Figure 7.
CNN-Transformer architecture.
Optimization was performed using the AdamW optimizer with a learning rate of and a 10-epoch warm-up schedule, a batch size of 16, and a total of 100 training epochs.This hybrid framework combines the local precision of convolutional neural networks with the contextual reasoning capabilities of Transformer architectures, achieving state-of-the-art classification performance among all evaluated models.
The main training hyperparameters for each architecture are summarized in Table 1 to facilitate comparison across models.

Table 1.
Summary of model hyperparameters.
2.4.7. Training Configuration
All models were implemented using TensorFlow 2.12 and trained on a GPU system (NVIDIA RTX 4090 with 24 GB VRAM) with mixed-precision acceleration. Adaptive optimization strategies were employed: the Adam optimizer was used for CNN-based and recurrent architectures, whereas AdamW was adopted for Transformer-based models. Learning rates ranged from to , depending on model complexity.
Regularization techniques, including Dropout (–) and Batch Normalization, were applied consistently across all architectures. Early stopping was employed to mitigate overfitting, and a weighted categorical cross-entropy loss function was adopted to address residual class imbalance. Batch sizes varied between 16 and 32, and training durations ranged from 40 to 100 epochs, adaptively determined based on validation convergence.
To prevent optimistic bias arising from patient-specific acoustic signatures, a subject-independent validation protocol was adopted. Instead of a recording-wise random split, a 5-fold GroupKFold cross-validation strategy was implemented using patient identifiers extracted from file names. All recordings belonging to a given patient were assigned exclusively to a single fold, ensuring strict inter-patient separation between training and validation sets. At each iteration, four folds were used for training and one-fold for independent evaluation. Performance metrics are reported as mean ± standard deviation across the five folds. Fold partitioning was performed prior to any preprocessing steps requiring parameter estimation to avoid data leakage. Deterministic preprocessing operations (signal standardization, amplitude normalization, and duration normalization) were applied identically across folds. All stochastic augmentation and class balancing procedures, including waveform-level transformations, SpecAugment, and SMOTE oversampling, were restricted exclusively to the training subset within each fold. Feature standardization parameters (StandardScaler) were fitted solely on the training data of each fold and subsequently applied to both training and validation subsets using the learned parameters.
This unified and adaptive experimental framework ensured consistent optimization across all six evaluated architectures 1D-CNN, Deep-CNN, CNN-LSTM, CNN-BiLSTM, CNN-Attention, and CNN-Transformer, thereby facilitating a fair, reproducible, and clinically relevant comparison of their representational performance.
3. Results
3.1. Overall Classification Performance
Table 2 presents a comparative summary of the performance achieved by the six evaluated architectures across the four target classes: Normal, Crackles, Wheezes, and Crackles + Wheezes, under a patient-wise 5-fold cross-validation protocol, with results reported as mean ± standard deviation across folds. Overall, all models demonstrated strong and consistent performance, thereby confirming the effectiveness of deep learning approaches in extracting discriminative spectro-temporal features from respiratory signals while preserving inter-patient independence.

Table 2.
Performance metrics comparison across six architectures.
The baseline 1D–CNN model achieved a Balanced Accuracy of ± , demonstrating that a shallow convolutional architecture remains capable of effectively identifying localized temporal patterns. This performance provides a solid foundation, illustrating the ability of convolutional filters to detect elementary acoustic structures, such as harmonic transitions or short impulsive events associated with adventitious pulmonary sounds. By deepening the hierarchy of convolutional filters, the Deep-CNN architecture improved the Balanced Accuracy to ± , highlighting the contribution of additional layers in representing more complex acoustic structures and modeling higher-level spectral dependencies.
The integration of temporal modeling through recurrent layers led to a notable improvement in performance. The CNN–LSTM model achieved a Balanced Accuracy of ± , while its bidirectional counterpart, CNN–BiLSTM, reached ± , confirming the relevance of bidirectional temporal context in capturing interdependencies between inspiratory and expiratory phases. This progressive improvement indicates that recurrent architectures enable a more comprehensive characterization of the temporal dynamics inherent in the respiratory cycle and support a more robust representation of acoustic anomalies.
Models incorporating attention mechanisms yielded the best overall performance. The CNN-Attention model achieved a Balanced Accuracy of ± , slightly outperforming the CNN–Transformer model ( ± ), while exhibiting strong generalization capability and high class-wise stability. These results underscore the effectiveness of attention mechanisms in dynamically weighting the most informative temporal segments, thereby enhancing the detection of pathological sounds while reducing sensitivity to noise and normal respiratory artifacts.
Overall, this comparative analysis demonstrates that combining convolutional feature extraction with temporal and attentional modeling mechanisms leads to a substantial enhancement in discriminative performance. Purely convolutional approaches, although effective, remain limited in their ability to capture long-range sequential dependencies. In contrast, hybrid and attention-based architectures leverage both local spectral regularities and global temporal dynamics, providing a more comprehensive and robust representation of respiratory sounds. These findings suggest that architectures incorporating contextual attention constitute a promising direction for clinical acoustic classification, where diagnostic accuracy and inter-patient stability are of paramount importance.
While the preceding discussion focused on multi-class discrimination performance, it is important to contextualize the results within the official ICBHI evaluation protocol. Although the proposed framework performs four-class respiratory sound classification, the ICBHI benchmarking scheme also includes a Normal versus Abnormal discrimination setting. Therefore, in addition to the multi-class metrics reported in Table 2, we computed macro-averaged Sensitivity (Se), Specificity (Sp), and the official ICBHI Score Performance (SP), defined as the arithmetic mean of Sensitivity and Specificity, as presented in Table 3. All metrics were derived within the same 5-fold patient-wise cross-validation protocol to ensure strict inter-patient independence. In this binary evaluation setting, all pathological classes (Crackles, Wheezes, and Crackles + Wheezes) were aggregated into a single Abnormal category. The reported values represent mean ± standard deviation across folds, thereby providing a robust estimate of generalization performance under clinically realistic inter-patient variability.

Table 3.
Binary Evaluation Metrics (Normal vs. Abnormal).
3.2. Training and Convergence Behavior
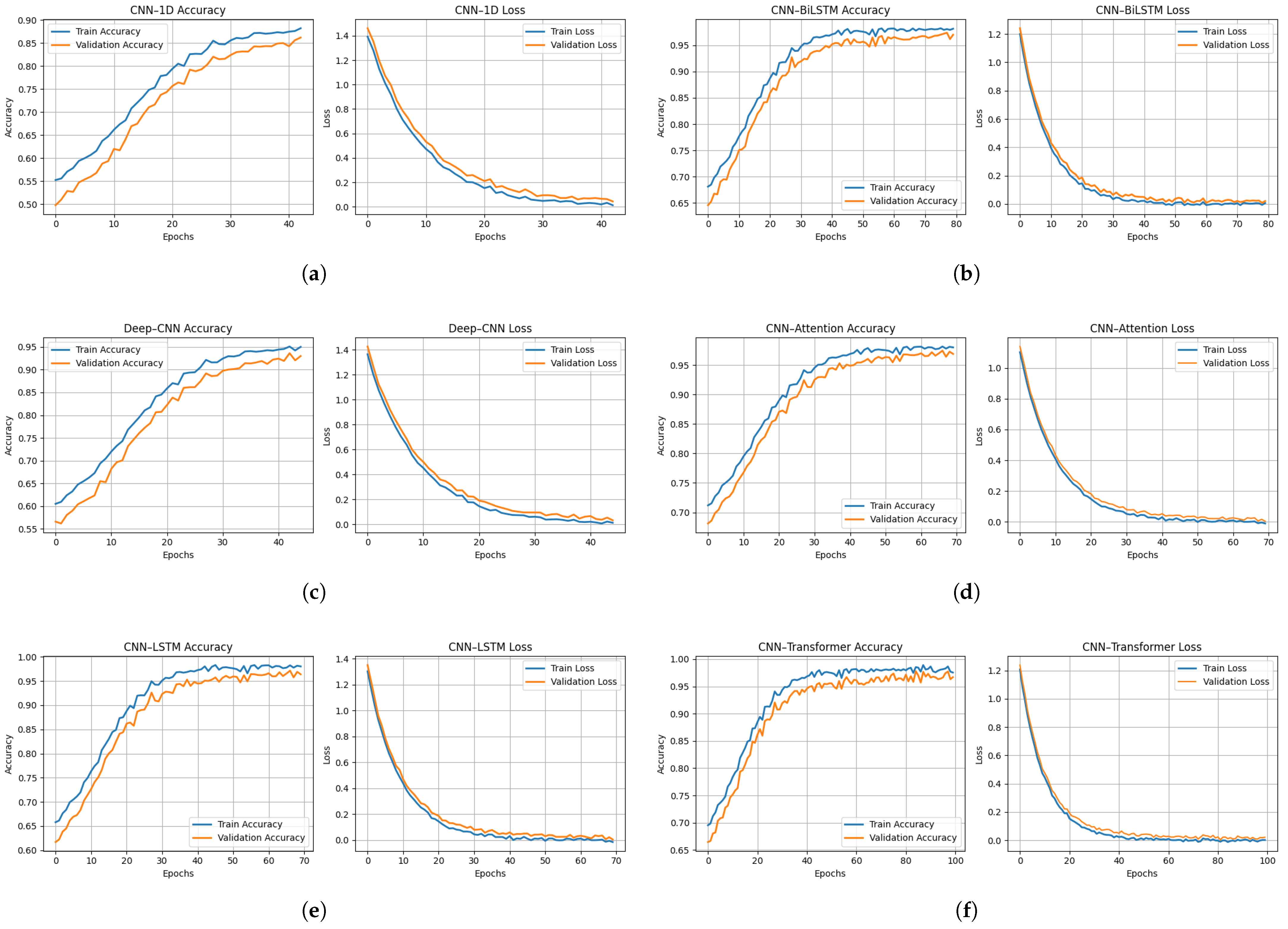
Figure 8a–f illustrates the evolution of Balanced Accuracy and loss for both the training and validation folds over the course of training iterations for the six evaluated models. Overall, all architectures exhibit stable and progressive convergence, reflecting the effectiveness of the preprocessing pipeline, optimization strategies, and regularization mechanisms implemented to prevent overfitting under a subject-independent validation framework.
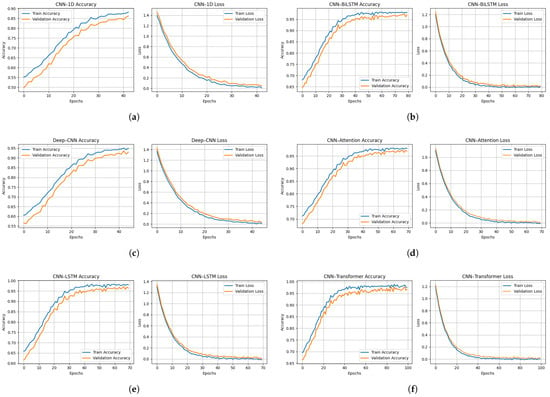
Figure 8.
Training and validation accuracy (left) and loss (right) curves for all evaluated architectures: (a) CNN-1D, (b) Deep-CNN, (c) CNN-LSTM, (d) CNN-BiLSTM, (e) CNN-Attention, and (f) CNN-Transformer.
The 1D-CNN model (Figure 8a) shows a gradual improvement in validation performance, reaching approximately ± Balanced Accuracy after 35 epochs, accompanied by a continuous decrease in the loss function. This behavior indicates a progressive assimilation of temporal patterns, despite a slight gap between the training and validation curves, indicating moderate but stable learning. The Deep-CNN architecture (Figure 8b) converges more rapidly, attaining a validation Balanced Accuracy of ± , demonstrating that increased network depth improves spectro-temporal feature extraction. However, a slight divergence observed toward the end of training may indicate incipient overfitting, likely attributable to increased parametric complexity.
The CNN-LSTM model (Figure 8c) exhibits a slower initial learning phase, followed by stable convergence around epoch 40, with validation Balanced Accuracy of ± and near-complet overlap between the training and validation curves. This behavior reflects the ability of recurrent layers to effectively model temporal dependencies within respiratory cycles, thereby reducing variability and improving robustness. The CNN-BiLSTM architecture (Figure 8d) further improves performance, displaying nearly superimposed curves and achieving ± Balanced Accuracy, indicative of superior generalization enabled by bidirectional learning of temporal context that captures both past and future dependencies within respiratory signals.
The CNN–Attention model (Figure 8e) is characterized by rapid convergence, reaching a performance plateau within the first 10 epochs. The close alignment of the training and validation curves, together with a validation Balanced Accuracy of ± , suggests that the attention mechanism efficiently identifies the most informative acoustic segments while suppressing noise and non-informative regions. Finally, the CNN–Transformer model (Figure 8f) demonstrates high stability and achieves ± Balanced Accuracy, owing to the synergy between local feature extraction via convolutional layers and global dependency modeling through self-attention modules.
Collectively, these learning dynamics reveal a clear performance hierarchy among the evaluated architectures under strict patient-wise evaluation, with attention-based and bidirectional recurrent models exhibiting improved generalization compared to purely convolutional approaches. These findings further emphasize the relevance of attention mechanisms and sequential modeling for robust respiratory sound classification, particularly in clinically realistic settings characterized by inter-patient variability.
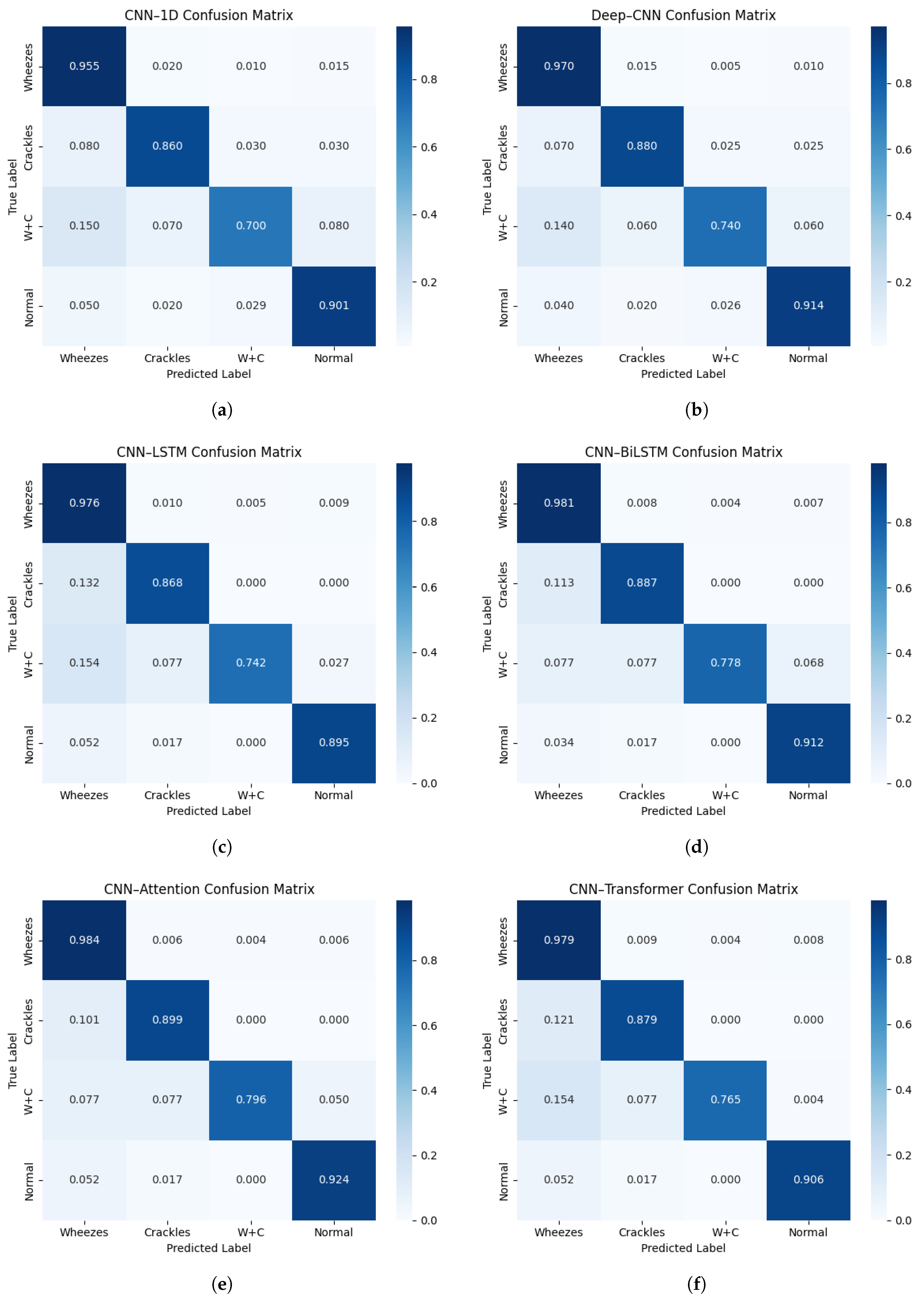
3.3. Class-Wise Performance and Confusion Analysis
Figure 9a–f presents the normalized confusion matrices for the six architectures evaluated, enabling a detailed assessment of class-wise performance across the four acoustic categories: Normal, Crackles, Wheezes, and Crackles + Wheezes. The confusion matrices correspond to the aggregated predictions obtained across all validation folds of the patient-wise cross-validation procedure. The baseline 1D-CNN model achieves satisfactory classification performance but exhibits notable misclassifications, particularly between the Crackles and Crackles + Wheezes classes, with recognition rates of approximately and , respectively. This confusion likely originates from the temporal overlap between transient components (Crackles) and continuous components (Wheezes), which simple convolutional filters are not always able to clearly separate. The Deep-CNN architecture shows a clear improvement, with a more pronounced diagonal dominance, correctly identifying approximately of Crackles and of Crackles + Wheezes. Nevertheless, some ambiguity persists, particularly between low-energy crackles and normal respiratory sounds, suggesting that increased network depth alone does not fully guarantee temporal consistency.
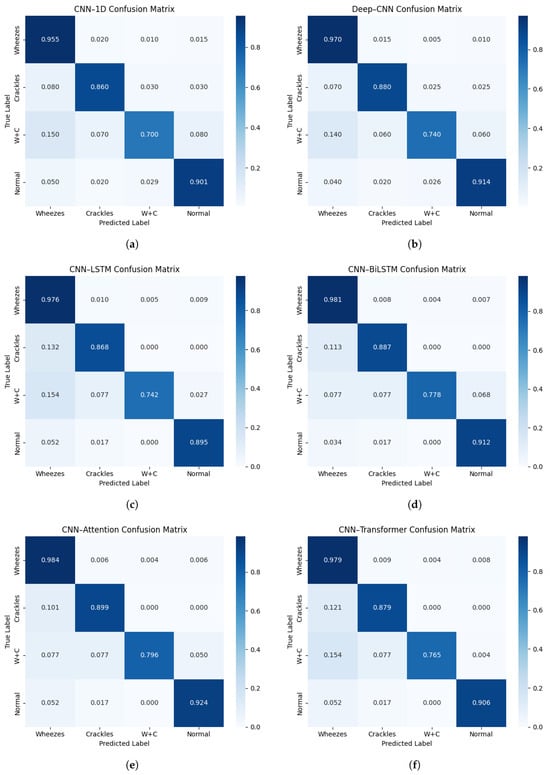
Figure 9.
Normalized confusion matrices for all evaluated architectures: (a) CNN-1D, (b) Deep-CNN, (c) CNN-LSTM, (d) CNN-BiLSTM, (e) CNN-Attention, and (f) CNN-Transformer.
The introduction of recurrent layers in the CNN-LSTM model significantly enhances classification performance, with recall values reaching approximately for Wheezes, for Crackles, for Crackles + Wheezes, and for Normal sounds. The memory mechanisms inherent to LSTM units enable fine-scale modeling of the temporal progression of respiratory phases, thereby reinforcing coherence between successive acoustic events. The CNN–BiLSTM model further improves performance through bidirectional temporal learning, achieving diagonal values of approximately 0.98 (Wheezes), 0.89 (Crackles), 0.78 (Crackles + Wheezes), and 0.91 (Normal). This temporal symmetry allows for improved discrimination of composite acoustic events, such as Crackles + Wheezes, by leveraging the entire contextual span of the respiratory cycle.
The CNN–Attention model achieves the best overall performance, correctly identifying approximately 0.98 of Wheezes, 0.90 of Crackles, 0.80 of Crackles + Wheezes, and 0.92 of Normal samples. Its attention mechanism dynamically assigns higher weights to the most informative temporal segments, thereby emphasizing pathological sounds while suppressing artifacts associated with normal breathing. Finally, the CNN–Transformer model attains class-wise recognition rates ranging approximately from 0.77 (Crackles + Wheezes) to 0.98 (Wheezes), reflecting a strong capacity for long-range contextual modeling through self-attention layers.
Overall, the confusion matrices reveal a progressive improvement in inter-class separability as the architectures evolve from purely convolutional designs to hybrid recurrent and attention-based models. This trend confirms that explicit modeling of temporal and contextual dependencies substantially enhances diagnostic reliability and per-class robustness, properties that are critical for clinical decision-support applications.
4. Discussion
4.1. Comparative Metrics Analysis
Table 2 summarizes the overall performance of all evaluated models according to Balanced Accuracy, Macro Precision, Macro F1-score, and Specificity under a subject-independent 5-fold patient-wise cross-validation protocol. A clear upward trend emerges, progressing from simple convolutional architectures to hybrid and attention-based models, indicating a gradual improvement in both feature extraction and temporal modeling capabilities.
The 1D-CNN model achieves a Balanced Accuracy of , providing a solid baseline despite its limited receptive field. Its slightly lower Macro F1-score reflects a tendency to overlook certain transient or overlapping events, particularly in recordings labeled as Crackles + Wheezes. The Deep-CNN architecture improves performance to Balanced Accuracy, demonstrating that a deeper hierarchy of filters strengthens the capture of spectro-temporal dependencies. However, a discrepancy between training and validation accuracies suggests mild overfitting due to increased parametric complexity.
The hybrid recurrent models yield measurable improvements: the CNN-LSTM reaches Balanced Accuracy, confirming the role of sequential layers in modeling intra-cycle relationships between inspiratory and expiratory phases. The CNN-BiLSTM further improves performance to , highlighting its ability to leverage bidirectional temporal relationships, which are essential for recognizing complex and irregular pulmonary sounds. The improved diagonal dominance observed in its confusion matrix confirms enhanced inter-patient robustness.
The CNN-Attention model achieves the highest performance, with a Balanced Accuracy of and an ICBHI Score of , underscoring the effectiveness of the attention mechanism in prioritizing diagnostically relevant acoustic segments while reducing inter-patient variability. The CNN-Transformer attains Balanced Accuracy, demonstrating stable generalization supported by self-attention modeling of long-range temporal dependencies, although its performance remains slightly below that of the hybrid CNN-Attention and CNN-BiLSTM architectures.
Overall, architectures incorporating temporal recurrence (LSTM/BiLSTM) and contextual weighting (Attention/Transformer) demonstrate comparatively improved representational capacity compared to purely convolutional models under patient-wise validation. The consistent improvement in Macro F1-score across architectures confirms that these advanced networks enhance class-wise robustness, which is an essential requirement for clinically reliable diagnostic support systems operating under realistic inter-patient variability. When appropriately regularized, these architectures constitute a promising methodological foundation for the development of automated auscultation systems.
4.2. Comparative Discussion with Related Works
To contextualize the results obtained within recent advances in automatic respiratory sound classification, Table 4 provides a comparative synthesis of studies published between 2022 and 2024. These works encompass a variety of architectures and datasets, ranging from private hospital corpora to public datasets such as ICBHI 2017.

Table 4.
Comparative summary of recent studies on automatic respiratory sound classification.
Early deep learning approaches, such as those by Brian Sang et al. [19] and Beom Joon Kim et al. [10], focused on binary classification tasks (wheezing vs. non-wheezing) using convolutional or attention-based networks (e.g., Deep-CNN, ResNet34 + CBAM). Despite achieving high accuracies (91–94%), these systems were constrained by limited class diversity and the absence of publicly available datasets, thereby restricting their generalizability. More recent studies, such as that by Rehan Khan et al. [37], addressed multi-class classification using a hybrid LSTM-CAE network leveraging features derived from the Continuous Wavelet Transform (CWT) and Mel-spectrogram, achieving 79% accuracy across four acoustic categories. Similarly, Sebastian Escobar-Pajoy [38] combined handcrafted features (Fourier Transform, MFCC, Chroma, FSS) with traditional algorithms, reporting 87–91% accuracy on the ICBHI dataset, although with limited robustness to inter-patient variability.
Other researchers have investigated lightweight architectures or transfer learning approaches. Ximing Liao et al. [39] adapted MobileNetV2 (pretrained on ImageNet) for multi-class classification, achieving approximately 87%, while Georgios Petmezas et al. [15] employed a 2D-LSTM network on STFT spectrograms (), highlighting the importance of temporal modeling for non-stationary signals. Finally, Syed Wagad Ali et al. [32] proposed a 1D-CNN architecture directly applied to raw signals, reaching 97.39% accuracy, demonstrating the effectiveness of automatic feature extraction.
Recently, Zhu et al. [40] proposed ADFF-Net, an attention-based dual-stream feature fusion network evaluated on the ICBHI 2017 dataset under the official train–test split. Their architecture combines Mel-filter bank and Mel-spectrogram representations through an attention-based fusion module. Reported results include an overall accuracy of 64.95%, sensitivity of 42.91%, specificity of 81.39%, and a harmonic score of 62.14%. The inclusion of ADFF-Net in Table 4 provides an additional benchmark within the same public dataset, enabling a transparent comparison between dual-stream fusion strategies and the unified architectural benchmarking framework adopted in the present study.
In comparison, the present study introduces a unified and reproducible experimental framework trained on the ICBHI 2017 dataset with four acoustic classes (Normal, Crackles, Wheezes, Crackles + Wheezes) under a subject-independent 5-fold patient-wise cross-validation protocol. The proposed pipeline integrates acoustic relabeling, comprehensive preprocessing (including normalization, advanced augmentation, and SpecAugment), and class balancing via SMOTE. Six architectures were evaluated, convolutional (1D-CNN, Deep-CNN), recurrent hybrid (CNN-LSTM, CNN-BiLSTM), and attention-based (CNN-Attention, CNN-Transformer). The proposed models achieved Balanced Accuracy values ranging from % to %, with the CNN–Attention architecture attaining the highest performance ( ± %) and an ICBHI Score of ± %.
Although these results are numerically lower than those obtained under recording-wise or fixed train–test splits reported in some prior studies, they provide a more conservative and clinically realistic estimate of generalization performance across unseen patients. It should be noted that many prior studies report conventional accuracy under recording-wise splits, whereas the present work reports Balanced Accuracy under strict patient-wise validation. The CNN–Attention and CNN–BiLSTM models demonstrate consistent improvements over purely convolutional approaches, confirming the benefit of integrating temporal modeling and contextual attention mechanisms for reliable detection of pathological respiratory events.
4.3. Model Complexity and Practical Considerations
In addition to classification performance, model complexity is an important consideration for real-world deployment in intelligent auscultation systems. Convolutional architectures (1D–CNN and Deep–CNN) present comparatively lower structural complexity because they rely primarily on local receptive fields and parallelizable operations, which typically results in reduced computational overhead. In contrast, recurrent variants (CNN–LSTM and CNN–BiLSTM) introduce gating mechanisms and sequential dependencies that increase computational cost and memory usage, particularly when bidirectional processing is employed.
Attention-based models (CNN–Attention and CNN–Transformer) further incorporate self-attention operations that enable global contextual modeling of respiratory sound patterns, which can improve discrimination between transient crackles and sustained wheeze components. Although these mechanisms generally increase architectural complexity relative to purely convolutional baselines, the added representational capacity helps explain the observed performance gains in Balanced Accuracy under patient-wise evaluation. Overall, the comparative results suggest that improved accuracy is achieved with a reasonable and interpretable increase in complexity, supporting feasibility in contemporary GPU-enabled clinical and research settings.
4.4. Limitations and Future Directions
Despite the strong classification performance achieved across all evaluated architectures, several methodological limitations should be acknowledged. First, the adopted segment-level labeling strategy assigns a single global label to each standardized 5-s audio segment. Although this approach ensures consistent input dimensionality and aligns with common ICBHI benchmarking practices under controlled settings, respiratory events such as isolated crackles may be temporally localized and may not span the entire segment duration. As a result, some segments may contain acoustically normal intervals despite being labeled as abnormal, introducing a degree of label noise.
While attention-based and Transformer architectures appear to mitigate this limitation by dynamically focusing on temporally salient regions, the evaluation remains constrained by the adopted segmentation strategy. Although the present study enforces subject-independent (patient-wise) cross-validation, further validation on external datasets would be required to fully establish cross-cohort generalization.
Additionally, the composite class Crackles + Wheezes remains markedly underrepresented in the ICBHI dataset ( of recordings), which may limit the statistical stability of minority-class performance estimates despite fold-wise SMOTE-based balancing.
Finally, although the preprocessing framework integrates waveform-level augmentation, SpecAugment, enriched MFCC features, and SMOTE-based balancing in a unified pipeline, a systematic component-wise ablation analysis was not conducted. Since the primary objective was controlled architectural benchmarking under identical preprocessing conditions, all models were evaluated using the same standardized configuration to ensure fair comparison. Future work will incorporate external validation cohorts, finer event-level segmentation strategies, and targeted ablation experiments to further assess generalization robustness and quantify the individual contribution of each preprocessing module.
5. Conclusions
In this study, we proposed and comprehensively evaluated a deep learning framework for the automatic classification of respiratory sounds using the ICBHI 2017 dataset. Through a rigorous preprocessing pipeline, including acoustic relabeling, signal normalization, data augmentation, enriched MFCC feature extraction (, ), and class balancing, we generated homogeneous, information-rich representations that are robust to inter-patient variability.
Six neural network architectures were implemented and compared: 1D-CNN, Deep-CNN, CNN-LSTM, CNN-BiLSTM, CNN-Attention, and CNN-Transformer. The comparative analysis demonstrated that all architectures achieved high classification performance across the four clinical classes, with Balanced Accuracy values ranging from to , under a subject-independent 5-fold patient-wise cross-validation protocol. The CNN-Attention model achieved the best overall performance, reaching Balanced Accuracy and ICBHI Score, confirming the relevance of attention mechanisms for simultaneously capturing local spectral features and long-term temporal dependencies within respiratory cycles.
Beyond quantitative performance, this study provides a structured and reproducible benchmarking framework designed to address data imbalance, recording heterogeneity, and variability in feature representations. While the results demonstrate consistent performance under strict inter-patient validation, further multi-center external validation and prospective clinical studies are necessary before real-world deployment can be fully established.
Future work will explore cycle-level segmentation, multi-center external validation, and extended clinical evaluation to further assess generalization capacity and practical applicability.
Author Contributions
Conceptualization, B.B.; methodology, B.B.; software, B.B.; validation, B.B.; formal analysis, B.B.; investigation, B.B.; resources, B.B.; data curation, B.B.; writing—original draft preparation, B.B.; writing—review and editing, B.B., G.Z., M.A.M., A.E.O. and M.J.; visualization, B.B., G.Z., M.A.M. and M.J.; supervision, G.Z., M.A.M., M.J. and A.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data used in this study are publicly available from the ICBHI 2017 respiratory sound database.
Acknowledgments
The authors would like to thank the contributors of the ICBHI 2017 respiratory sound database for making the dataset publicly available for research purposes.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Imran, A.; Posokhova, I.; Qureshi, H.N.; Masood, U.; Riaz, M.S.; Ali, K.; John, C.N.; Hussain, M.I.; Nabeel, M. AI4COVID-19: AI Enabled Preliminary Diagnosis for COVID-19 from Cough Samples via an App. Inform. Med. Unlocked 2020, 20, 100378. [Google Scholar] [CrossRef] [PubMed]
- Coppock, H.; Gaskell, A.; Tzirakis, P.; Baird, A.; Jones, L.; Schuller, B. End-to-End Convolutional Neural Network Enables COVID-19 Detection from Breath and Cough Audio: A Pilot Study. BMJ Innov. 2021, 7, 356–362. [Google Scholar] [CrossRef] [PubMed]
- Bouzammour, B.; Zaz, G.; Marktani, M.A.; Chougrad, H.; Touhafi, A.; Jorio, M. Ensemble Deep Learning Models for Respiratory Disease Detection Using Cough Analysis. J. Eng. Res. 2025. [Google Scholar] [CrossRef]
- Zaz, G.; Marktani, M.A.; Elboushaki, A.; Farhane, Y.; Mechaqrane, A.; Jorio, M.; Bekkay, H.; Dosse, S.B.; Mansouri, A.; Ahaitouf, A. Intelligent Multi-Sensor System for Remote Detection of COVID-19. In Smart and Sustainable Technology for Resilient Cities and Communities; Springer Nature: Singapore, 2022; pp. 149–162. [Google Scholar]
- Bouzammour, B.; Zaz, G.; Alami Marktani, M.; Ahaitouf, A.; Jorio, M. Cough Detection for Prevention Against the COVID-19 Pandemic. In International Conference on Electronic Engineering and Renewable Energy Systems; Springer: Singapore, 2022; pp. 421–430. [Google Scholar]
- Aykanat, M.; Kılıç, Ö.; Kurt, B.; Saryal, S. Classification of Lung Sounds Using Convolutional Neural Networks. EURASIP J. Image Video Process. 2017, 2017, 65. [Google Scholar] [CrossRef]
- Bokov, P.; Mahut, B.; Flaud, P.; Delclaux, C. Wheezing Recognition Algorithm Using Recordings of Respiratory Sounds at the Mouth in a Pediatric Population. Comput. Biol. Med. 2016, 70, 40–50. [Google Scholar] [CrossRef]
- Pahar, M.; Theron, G.; Niesler, T. Automatic Tuberculosis Detection in Cough Patterns Using NLP-Style Cough Embeddings. In Proceedings of the 2022 International Conference on Engineering and Emerging Technologies (ICEET), Kuala Lumpur, Malaysia, 27–28 October 2022; pp. 1–6. [Google Scholar]
- McClure, K.; Erdreich, B.; Bates, J.H.T.; McGinnis, R.S.; Masquelin, A.; Wshah, S. Classification and Detection of Breathing Patterns with Wearable Sensors and Deep Learning. Sensors 2020, 20, 6481. [Google Scholar] [CrossRef]
- Kim, B.J.; Kim, B.S.; Mun, J.H.; Lim, C.; Kim, K. An Accurate Deep Learning Model for Wheezing in Children Using Real-World Data. Sci. Rep. 2022, 12, 22465. [Google Scholar] [CrossRef]
- Kim, B.J.; Mun, J.H.; Hwang, D.H.; Suh, D.I.; Lim, C.; Kim, K. An Explainable and Accurate Transformer-Based Deep Learning Model for Wheeze Classification Utilizing Real-World Pediatric Data. Sci. Rep. 2025, 15, 5656. [Google Scholar] [CrossRef]
- Park, J.S.; Kim, K.; Kim, J.H.; Choi, Y.J.; Kim, K.; Suh, D.I. A Machine Learning Approach to the Development and Prospective Evaluation of a Pediatric Lung Sound Classification Model. Sci. Rep. 2023, 13, 1289. [Google Scholar] [CrossRef]
- Mendes, L.; Vogiatzis, I.M.; Perantoni, E.; Kaimakamis, E.; Chouvarda, I.; Maglaveras, N.; Paiva, R.P. Detection of Wheezes Using Their Signature in the Spectrogram Space and Musical Features. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; pp. 5581–5584. [Google Scholar]
- Amose, J.; Manimegalai, P. Classification of Adventitious Lung Sounds: Wheeze, Crackle Using Machine Learning Techniques. Int. J. Intell. Syst. Appl. Eng. 2023, 11, 1143–1152. [Google Scholar]
- Petmezas, G.; Cheimariotis, G.-A.; Stefanopoulos, L.; Rocha, B.; Paiva, R.P.; Katsaggelos, A.K.; Maglaveras, N. Automated Lung Sound Classification Using a Hybrid CNN-LSTM Network and Focal Loss Function. Sensors 2022, 22, 1232. [Google Scholar] [CrossRef]
- Rocha, B.M.; Pessoa, D.; Marques, A.; Carvalho, P.; Paiva, R.P. Automatic Classification of Adventitious Respiratory Sounds: A (Un)Solved Problem? Sensors 2020, 21, 57. [Google Scholar] [CrossRef] [PubMed]
- Polanco-Martagón, S.; Hernández-Mier, Y.; Nuño-Maganda, M.A.; Barrón-Zambrano, J.H.; Magadán-Salazar, A.; Medellín-Vergara, C.A. Comparison of Deep Neural Networks for the Classification of Adventitious Lung Sounds. J. Clin. Med. 2025, 14, 7427. [Google Scholar] [CrossRef]
- Rocha, B.M.; Filos, D.; Mendes, L.; Serbes, G.; Ulukaya, S.; Kahya, Y.P.; Jakovljevic, N.; Turukalo, T.L.; Vogiatzis, I.; Perantoni, E.; et al. An Open Access Database for the Evaluation of Respiratory Sound Classification Algorithms. Physiol. Meas. 2019, 40, 035001. [Google Scholar] [CrossRef] [PubMed]
- Sang, B.; Wen, H.; Junek, G.; Neveu, W.; Di Francesco, L.; Ayazi, F. An Accelerometer-Based Wearable Patch for Robust Respiratory Rate and Wheeze Detection Using Deep Learning. Biosensors 2024, 14, 118. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.; Fazekas, G.; Sandler, M.; Cho, K. A Comparison of Audio Signal Preprocessing Methods for Deep Neural Networks on Music Tagging. In Proceedings of the 2018 26th European Signal Processing Conference (EUSIPCO), Rome, Italy, 3–7 September 2018; pp. 1870–1874. [Google Scholar]
- Abayomi-Alli, O.O.; Damaševičius, R.; Qazi, A.; Adedoyin-Olowe, M.; Misra, S. Data Augmentation and Deep Learning Methods in Sound Classification: A Systematic Review. Electronics 2022, 11, 3795. [Google Scholar] [CrossRef]
- Chu, H.-C.; Zhang, Y.-L.; Chiang, H.-C. A CNN Sound Classification Mechanism Using Data Augmentation. Sensors 2023, 23, 6972. [Google Scholar] [CrossRef]
- Maharana, K.; Mondal, S.; Nemade, B. A Review: Data Pre-Processing and Data Augmentation Techniques. Glob. Transit. Proc. 2022, 3, 91–99. [Google Scholar] [CrossRef]
- Wei, S.; Zou, S.; Liao, F.; Lang, W. A Comparison on Data Augmentation Methods Based on Deep Learning for Audio Classification. J. Phys. Conf. Ser. 2020, 1453, 012085. [Google Scholar] [CrossRef]
- Altinors, A.; Yol, F.; Yaman, O. A Sound Based Method for Fault Detection with Statistical Feature Extraction in UAV Motors. Appl. Acoust. 2021, 183, 108325. [Google Scholar] [CrossRef]
- Gourisaria, M.K.; Agrawal, R.; Sahni, M.; Singh, P.K. Comparative Analysis of Audio Classification with MFCC and STFT Features Using Machine Learning Techniques. Discov. Internet Things 2024, 4, 1. [Google Scholar] [CrossRef]
- Singh, M.K. Multimedia Application for Forensic Automatic Speaker Recognition from Disguised Voices Using MFCC Feature Extraction and Classification Techniques. Multimed. Tools Appl. 2024, 83, 77327–77345. [Google Scholar]
- Abbaskhah, A.; Sedighi, H.; Marvi, H. Infant Cry Classification by MFCC Feature Extraction with MLP and CNN Structures. Biomed. Signal Process. Control 2023, 86, 105261. [Google Scholar] [CrossRef]
- Jain, A.; Samala, P.R.; Mittal, D.; Jyoti, P.; Singh, M. SpliceOut: A Simple and Efficient Audio Augmentation Method. arXiv 2021, arXiv:2110.00046. [Google Scholar] [CrossRef]
- Lee, J.-N.; Lee, J.-Y. An Efficient SMOTE-Based Deep Learning Model for Voice Pathology Detection. Appl. Sci. 2023, 13, 3571. [Google Scholar] [CrossRef]
- Raju, V.V.N.; Saravanakumar, R.; Yusuf, N.; Pradhan, R.; Hamdi, H.; Saravanan, K.A.; Rao, V.S.; Askar, M.A. Enhancing Emotion Prediction Using Deep Learning and Distributed Federated Systems with SMOTE Oversampling Technique. Alex. Eng. J. 2024, 108, 498–508. [Google Scholar] [CrossRef]
- Ali, S.W.; Rashid, M.M.; Yousuf, M.U.; Shams, S.; Asif, M.; Rehan, M.; Ujjan, I.D. Towards the Development of the Clinical Decision Support System for the Identification of Respiration Diseases via Lung Sound Classification Using 1D-CNN. Sensors 2024, 24, 6887. [Google Scholar] [CrossRef]
- Zulfiqar, R.; Majeed, F.; Irfan, R.; Rauf, H.T.; Benkhelifa, E.; Belkacem, A.N. Abnormal Respiratory Sounds Classification Using Deep CNN Through Artificial Noise Addition. Front. Med. 2021, 8, 714811. [Google Scholar] [CrossRef]
- Zhang, P.; Swaminathan, A.; Uddin, A.A. Pulmonary Disease Detection and Classification in Patient Respiratory Audio Files Using Long Short-Term Memory Neural Networks. Front. Med. 2023, 10, 1269784. [Google Scholar] [CrossRef] [PubMed]
- You, M.; Wang, W.; Li, Y.; Liu, J.; Xu, X.; Qiu, Z. Automatic Cough Detection from Realistic Audio Recordings Using C-BiLSTM with Boundary Regression. Biomed. Signal Process. Control 2022, 72, 103304. [Google Scholar] [CrossRef]
- Wall, C.; Zhang, L.; Yu, Y.; Kumar, A.; Gao, R. A Deep Ensemble Neural Network with Attention Mechanisms for Lung Abnormality Classification Using Audio Inputs. Sensors 2022, 22, 5566. [Google Scholar] [CrossRef] [PubMed]
- Khan, R.; Khan, S.U.; Saeed, U.; Koo, I.-S. Auscultation-Based Pulmonary Disease Detection through Parallel Transformation and Deep Learning. Bioengineering 2024, 11, 586. [Google Scholar] [CrossRef] [PubMed]
- Escobar-Pajoy, S.; Ugarte, J.P. Computerized Analysis of Pulmonary Sounds Using Uniform Manifold Projection. Chaos Solitons Fractals 2023, 166, 112930. [Google Scholar] [CrossRef]
- Liao, X.; Wu, Y.; Jiang, N.; Sun, J.; Xu, W.; Gao, S.; Wang, J.; Li, T.; Wang, K.; Li, Q. Automated Detection of Abnormal Respiratory Sound from Electronic Stethoscope and Mobile Phone Using MobileNetV2. Biocybern. Biomed. Eng. 2023, 43, 763–775. [Google Scholar] [CrossRef]
- Zhu, B.; Chen, L.; Li, X.; Zhao, S.; Yu, S.; Sun, Q. ADFF-Net: An Attention-Based Dual-Stream Feature Fusion Network for Respiratory Sound Classification. Technologies 2025, 14, 12. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.